Abstract
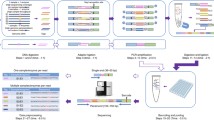
Molecular inversion probes (MIPs) in combination with massively parallel DNA sequencing represent a versatile, yet economical tool for targeted sequencing of genomic DNA. Several thousand genomic targets can be selectively captured using long oligonucleotides containing unique targeting arms and universal linkers. The ability to append sequencing adaptors and sample-specific barcodes allows large-scale pooling and subsequent high-throughput sequencing at relatively low cost per sample. Here, we describe a “wet bench” protocol detailing the capture and subsequent sequencing of >2000 genomic targets from 192 samples, representative of a single lane on the Illumina HiSeq 2000 platform.
These authors contributed equally to this work.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Mamanova L, Coffey AJ, Scott CE et al (2010) Target-enrichment strategies for next-generation sequencing. Nat Methods 7:111–118
Hardenbol P, Baner J, Jain M et al (2003) Multiplexed genotyping with sequence-tagged molecular inversion probes. Nat Biotechnol 21:673–678
Porreca GJ, Zhang K, Li JB et al (2007) Multiplex amplification of large sets of human exons. Nat Methods 4:931–936
Turner EH, Lee C, Ng SB et al (2009) Massively parallel exon capture and library-free resequencing across 16 genomes. Nat Methods 6:315–316
O’Roak BJ, Vives L, Fu W et al (2012) Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338:1619–1622
Boyle EA, O’Roak BJ, Martin BK et al (2014) MIPgen: optimized modeling and design of molecular inversion probes for targeted resequencing. Bioinformatics 30:2670–2672
Hiatt JB, Pritchard CC, Salipante SJ et al (2013) Single molecule molecular inversion probes for targeted, high-accuracy detection of low-frequency variation. Genome Res 23:843–854
O'Roak BJ, Stessman HA, Boyle EA et al (2014) Recurrent de novo mutations implicate novel genes underlying simplex autism risk. Nat Commun 5(5595):1–6
Qubit® Assays: Quick Reference Guide: Pub.no: MAN0010876
Qubit® 2.0 Fluorometer: MAN0003231
MiSeq: Denature and Dilute Libraries Guide: 15039740v1
MiSeq: Reagent Kit v3- Preparation Guide: Part#15044983
MiSeq System User Guide: part # 15027617
Acknowledgments
We thank Bradley P. Coe for his critical review of the manuscript and Tonia Brown for assistance with the manuscript preparation. We thank Brian J. O’Roak, Beth Martin, Evan A. Boyle, and Joseph B. Hiatt for their overall contributions to developing the MIP protocol. S.C. is supported by a National Health and Medical Research Council (NHMRC) CJ Martin Biomedical Fellowship (#1073726). H.A.S. is supported, in part, by the NHGRI Interdisciplinary Training in Genome Science Grant (T32HG00035). E.E.E. is an investigator of the Howard Hughes Medical Institute. J.S. is an investigator of the Howard Hughes Medical Institute.
Competing financial Interests
E.E.E. is on the scientific advisory board (SAB) of DNAnexus, Inc., and is a consultant for the Kunming University of Science and Technology (KUST) as part of the 1000 China Talent Program.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Cantsilieris, S., Stessman, H.A., Shendure, J., Eichler, E.E. (2017). Targeted Capture and High-Throughput Sequencing Using Molecular Inversion Probes (MIPs). In: White, S., Cantsilieris, S. (eds) Genotyping. Methods in Molecular Biology, vol 1492. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6442-0_6
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6442-0_6
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6440-6
Online ISBN: 978-1-4939-6442-0
eBook Packages: Springer Protocols