Abstract
NK cells represent a very promising source for adoptive cellular approaches for cancer immunotherapy, and extensive research has been conducted, including clinical trials. Gene modification of NK cells can direct their specificity and enhance their function, but the efficiency of gene transfer techniques is very limited. Here we describe two protocols designed to generate mature human NK cells from gene-modified hematopoietic stem cells. These protocols use chimeric antigen receptor as the transgene, but could potentially be modified for the expression any particular transgene in human NK cells.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
Natural Killer (NK) cells are innate immune cells that mediate spontaneous cytotoxicity against tumor and virus-infected cells, and many clinical trials have attempted to harness their properties for cellular therapies [1–6]. Gene transfer technology can enhance the efficacy of such efforts by improving NK cell survival or function, or engineering antigen specificity [7–9].
Chimeric antigen receptors (CAR) are engineered fusion proteins that combine the antigen specificity of antigen-binding moieties of monoclonal antibodies and intracellular activation motifs capable to activate immune cells. Preliminary evidence suggests that NK cells with specificity directed by chimeric antigen receptors may have enhanced cytotoxicity [10–12].
Generation of mature NK cells from hematopoietic stem cells provides the opportunity of generation of younger NK cells and expansion of specific gene-modified clones starting from a smaller number of previously isolated and cryopreserved initial cells, with the added advantage of generation of multiple batches from the same donor [13–16]. In this chapter, we describe a protocol for NK cell differentiation from human hematopoietic stem cells (HSC ) modified to express chimeric antigen receptors using co-culture with a feeder stroma of murine OP9-DL1 cells in presence of human recombinant cytokines [15]. Alternatively, we describe a feeder-free protocol for generation of gene-modified NK cells from human hematopoietic stem cells using insulin-like growth factor 1 (IGF-1) [17]. The transgene used in this protocol is a CD19-specific CAR, but the HSC could potentially receive any type of persistent gene modification for expression in differentiated NK cells.
2 Materials
2.1 Isolation and Cryopreservation of Umbilical Cord Blood HSC
-
1.
Human primary cells from umbilical cord blood collected within 48 h for the CD34 isolation procedure (see Note 1).
-
2.
Ficoll-Paque PLUS (GE Healthcare Life Sciences).
-
3.
CD34 MicroBead Kit UltraPure (Miltenyi Biotec).
-
4.
MidiMACS Starting Kit with LS columns (Miltenyi Biotec).
-
5.
MACS BSA Stock Solution (Miltenyi Biotec).
-
6.
autoMACS Rinsing Solution (Miltenyi Biotec).
-
7.
Dulbecco’s PBS (DPBS).
-
8.
Freezing medium: 10 % DMSO in heat-inactivated fetal bovine serum (Omega).
2.2 Lentiviral Transduction of HSC
-
1.
Cryopreserved primary human CD34-positive hematopoietic stem cells (HSC ) isolated from umbilical cord blood (see Note 1).
-
2.
Transduction medium: Freshly prepared X-Vivo15 medium (Lonza), 50 ng/mL of recombinant human Stem Cell Factor (SCF) (R&D Systems), 50 ng/mL of recombinant human Flt-3 ligand (R&D Systems), and 50 ng/mL of recombinant human thrombopoietin (R&D Systems).
-
3.
Fibronectin fragment CH-296 RetroNectin (Takara Shuzo Co.).
-
4.
48 well non-tissue culture-treated plate, sterile.
-
5.
Third-generation lentiviral vector carrying second-generation chimeric antigen receptor construct (Fig. 1a) (see Note 2).
Fig. 1 Lentiviral vector for transduction of human HSC . (a) Diagram of third-generation lentiviral vector carrying anti-CD19 CAR and EGFP. (b) Co-expression of CAR and EGFP detected by flow cytometry in NK cells differentiated in vitro from human HSC , day 40 of culture. CAR expression was detected by using PE-Texas Red-conjugated polyclonal F(ab′)2 fragment goat antihuman IgG1 Fcχ (Jackson ImmunoResearch Laboratories) [15]
2.3 Differentiation of HSC to NK Cell Lineage
2.3.1 Co-culture with OP9-DL1 Stromal Cells
-
1.
OP9 stromal cell line (ATCC) gene-modified to express Delta-like 1 (OP-DL1) (see Note 3).
-
2.
Alpha-20 medium: alpha-Minimum Essential Medium (alpha-MEM) enriched with 20 % of heat-inactivated fetal bovine serum (Omega), 2 mM l-glutamine, 50 U/mL penicillin, and 50 μg/mL streptomycin.
-
3.
NK differentiation medium: alpha-20 medium, enriched with 5 ng/mL of recombinant human SCF (R&D Systems), 5 ng/mL of recombinant human Flt-3 ligand (R&D Systems) , 5 ng/mL of IL-7 (R&D Systems), and 10 ng/mL of IL-15 (R&D Systems).
-
4.
12 well tissue culture-treated plates, sterile.
-
5.
T-75 and T-150 culture flasks.
-
6.
0.05 % trypsin EDTA 1×.
-
7.
Dulbecco’s PBS (DPBS).
-
8.
70 μm cell strainer.
2.3.2 Feeder-Free Differentiation Protocol
-
1.
AIM V CC medium: AIM V Complete medium (Life Technologies), 5 % human AB serum (Corning), and 2 mM GlutaMAX (Life Technologies), freshly supplemented with 30 ng/mL of recombinant human cytokines SCF (R&D Systems), 50 ng/mL of recombinant human Flt-3 ligand (R&D Systems), 50 ng/mL of recombinant human IL-15 (R&D Systems), and 100 ng/mL of recombinant human IGF-1 (R&D Systems).
-
2.
Falcon 48, 12, and 6 well tissue culture-treated microplates, sterile.
3 Methods
3.1 Isolation and Cryopreservation of Umbilical Cord Blood HSC (See Note 1)
Isolation of CD34-positive cells from umbilical cord blood collected within 48 h will increase cell recovery. The cord blood unit can be kept at room temperature until isolation procedure. The isolation procedure should be conducted in a biosafety cabinet to ensure sterility and personal protection.
-
1.
Dilute umbilical cord blood with equal volume of PBS.
-
2.
Pipette 15 mL of Ficoll into a 50 mL conical tube.
-
3.
Layer 35 mL of cord blood onto Ficoll.
-
4.
Centrifuge at 400 × g for 30 min at 20 °C without brake.
-
5.
Prepare isolation buffer by diluting MACS BSA Stock Solution into autoMACS Rinsing Solution according to the manufacturer’s instructions, and keep it ice-cold.
-
6.
Collect the buffy coat of all post-centrifugation Ficoll tubes with a sterile Pasteur pipette or a serological pipette into a new 50 mL conical tube.
-
7.
Dilute such collected buffy coat with equal volume of PBS.
-
8.
Centrifuge at 300 × g for 15 min at 4 °C, with brake on.
-
9.
Remove supernatant.
-
10.
Collect all cell pellets into one 50 mL conical tube and complete to 50 mL with ice-cold isolation buffer.
-
11.
Count cells.
-
12.
Centrifuge at 250 × g for 5 min at 4 °C, with brake on.
-
13.
Remove supernatant and resuspend up to 1 × 108 cells in 300 μL of ice-cold isolation buffer.
-
14.
Follow the manufacturer’s instructions to use the reagents of the Miltenyi Biotec CD34 MicroBead Kit UltraPure and the MidiMACS Kit to perform immunomagnetic positive selection of CD34 HSC using the LS columns.
-
15.
After completion of the isolation protocol, count the CD34-positive HSC and use immediately for transduction, or cryopreserve for posterior use. HSC cells kept in culture with human cytokines will undergo mostly onto myeloid lineage differentiation and lose expression of CD34.
3.2 Lentiviral Transduction of HSC
3.2.1 High-Titer Lentivirus for Modification of HSC
Our protocol for packaging and titer analysis of high-titer lentiviral vectors for gene modification of primary human cells is very extensive and has been published elsewhere [15, 18]. In brief, such vectors should have titer of at least 5 × 107–1 × 108 TU/mL to ensure very efficient transduction, what requires post-packaging procedures to obtain 100–1000-fold concentration. The vector used in this protocol is a third-generation replication-incompetent HIV-based lentivirus carrying a CD19-specific CAR and enhanced green fluorescent protein (EGFP) (Fig. 1) (see Note 2).
3.2.2 Lentiviral Transduction of HSC
Before an NK differentiation protocol is initiated, the investigators should evaluate in preliminary experiments the optimal transduction conditions using the available preparations of lentiviral vectors in human HSC [18] (see Note 4). Gene-modified HSC should be placed in NK differentiation culture conditions immediately after lentiviral transduction, and full expression of integrated vector copies will take place around 10 days after transduction. We routinely use overnight cytokine pre-stimulation of HSC in plates coated with fibronectin fragment CH-296 RetroNectin before treatment with the lentiviral vector [15, 18].
-
1.
RetroNectin coating of transduction plate: fill the wells of a non-tissue culture-treated 48 well plate with 500 μL/well of RetroNectin 20 μg/mL in PBS and incubate for 2 h in room temperature; after aspirating the RetroNectin solution, fill the wells with 1 mL/well of blocking solution with 2 % fetal bovine serum (FBS) in PBS and incubate for 30 min in room temperature; remove the blocking solution and wash twice with 1 mL/well of wash buffer with 0.025 M HEPES in PBS (see Note 5).
-
2.
Resuspend human CD34-positive cells at 1 × 106 cells/mL in transduction medium.
-
3.
Plate 400 μL/well in the RetroNectin coated 48 well transduction plate (from step 1).
-
4.
Incubate the plate for 14 h in 5 % CO2 at 37 °C for pre-stimulation; the cells will attach to the bottom coated with RetroNectin.
-
5.
After pre-stimulation, add the necessary volume of lentiviral vector to ensure transduction efficiency at a viral vector copy number of 1–3 copies/cell [15, 18] as determined in preliminary experiments using available vector preparations (see Note 4).
-
6.
Incubate transduction plate with added lentiviral vectors for 24 h in 5 % CO2 at 37 °C before transferring HSC to differentiation cultures.
3.3 Culture and Passaging of OP9-DL1 Stromal Cells
It is fundamental to keep healthy, low-passage, stromal cells for co-culture with the primary human cells (see Notes 3 and 6).
-
1.
Start OP9-DL1 stromal cell culture with 2 × 106–5 × 106 cells in a T-75 flask using alpha-20 culture medium without any cytokines (see Note 7).
-
2.
To passage OP9-DL1, first carefully aspirate and remove the culture medium; rinse with 20 mL of PBS at room temperature, and completely remove PBS.
-
3.
Add 12 mL of trypsin for a T-75 flask (or 30 mL for a T-150 flask) and incubate at 37 °C for 5–15 min until cell layer is dispersed.
-
4.
Add at least the same volume of alpha-20 medium to resuspend cells in order to inactivate the trypsin.
-
5.
Count the cells and add 2 × 106–5 × 106 cells to a T-75 flask, or 5 × 106–10 × 106 cells to a T-150 flask, for continued culture (see Note 8).
-
6.
For the co-culture with human cells, plate 2 × 105–5 × 105 per well in a tissue culture-treated 12 well plate, in alpha-20 medium (see Note 9).
3.4 Differentiation of HSC to NK Cell Lineage
3.4.1 Differentiation by Co-culture with OP9-DL1 Stromal Cells
All co-cultures should be performed using freshly prepared NK differentiation medium from the first day.
-
1.
Vigorously pipette the HSC in the transduction plate after adding 0.5–1 mL of NK differentiation medium per well, in order to detach all cells adherent to the bottom.
-
2.
Count cells and adjust concentration to 5 × 105 cells/mL in NK differentiation medium.
-
3.
Carefully remove the alpha-20 medium from the confluent OP-DL1 stroma cultured in the 12 well plates (see Note 9).
-
4.
Seed the transduced HSC cells suspended in the NK differentiation medium by slow and gentle pipetting against the wall, at 3 × 105–5 × 105 cells per well.
-
5.
Gently add freshly prepared NK differentiation medium by slow pipetting against the wall for a total volume of 2 mL per well.
-
6.
Split HSC every 3–4 days; vigorously pipette using P-1000 to resuspend cells and detach stroma; remove stromal cells using a 70 μm cell strainer unto a 50 mL conical tube, then repeat steps 2–5 in this section (see Note 10).
-
7.
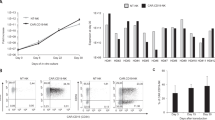
Maintain co-culture of transduced HSC with stromal cells for 35–40 days to obtain functional NK cells (see Notes 11 and 12) (Fig. 2a). CD56+ NK cells can be sorted via fluorescence activated cell sorting (FACS) or immunomagnetic bead selection for continued culture or functional assays such as cytotoxicity assessment.
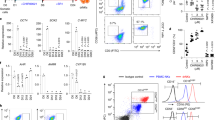
Fig. 2 Expression of NK-specific surface markers along progression of in vitro differentiation culture from gene-modified human HSC . (a) Co-culture with OP9-DL1 stromal cells. (b) Feeder-free culture. Representative plots were taken at days 14, 29, and 40 of both NK differentiation culture protocols. Gated human cells positive for EGFP were evaluated for the CD56, CD16, CD94, CD57, CD158, and CD335 surface markers
3.4.2 Feeder-Free Differentiation Culture (Alternate Protocol)
All media changes and splits should be performed using AIM V CC from the first day (see Note 13).
-
1.
Vigorously pipette the HSC in the transduction plate after adding 0.5–1 mL of AIM V CC per well, in order to detach all cells adherent to the bottom.
-
2.
Transfer cells to an appropriately sized conical tube. Centrifuge at 300 × g for 10 min. Remove supernatant and resuspend cell pellet by flicking the tube (see Note 14). Add 1 mL (or appropriate volume) of AIM V CC.
-
3.
Count cells using a viability dye (e.g., trypan blue) and adjust viable cell concentration to 0.5 × 106 cells/mL in AIM V CC.
-
4.
Plate cells into appropriately sized wells of tissue culture-treated microplates. For example, plate as many as 0.5 × cells in 1 mL of AIM V CC into one well of a 24 well plate, or up to 3 × 106 cells in 6 mL of AIM V CC into the well of a 6 well plate.
-
5.
Cells are incubated at 37 °C in a humidified atmosphere with 5 % CO2.
-
6.
Differentiating cells will need to be split up to twice a week. Pipet to detach gently adherent cells and transfer half the media and cells to a neighboring well that has not been used or to a new equally sized plate. Replenish the media to the original volume in both wells with fresh AIM V CC (see Note 15).
-
7.
In order to remove cellular debris, every 10–12 days the cells should be washed and completely replenished with fresh media and cytokines. Pipet to detach the slightly adherent cells, ignoring the strongly adherent cell populations. Transfer the non-adherent cells to conical tubes and centrifuge at 300 × g for 10 min. Remove supernatant and resuspend cell pellet in an appropriate volume of AIM V CC for counting cells. Count cells using a viability dye and adjust viable cell concentration to 1 × 106 cells/mL in AIM V CC. Plate cells into appropriately sized wells of Falcon tissue culture-treated microplates. For example, plate as many as 5 × 106 cells in 5 mL AIM V CC into one well of a 6 well plate (see Note 15).
-
8.
CD56+ NK cells will begin to appear around Day 14, peak around Day 28, and begin to decline thereafter (Fig. 2b).
-
9.
Between days 14 and 28, CD56+ NK cells can be sorted via fluorescence activated cell sorting (FACS) or immunomagnetic bead selection for continued culture or functional assays such as cytotoxicity assessment. Sorted NK cells can be cultured at 1 × 106 cells/mL, but will no longer appreciably expand, in AIM V medium (Life Technologies) supplemented with 5 % human AB serum (Corning) and 2 mM GlutaMAX (Life Technologies) and 20 ng/mL of recombinant human IL-15 (R&D Systems). In our hands, sorted NK cells in this media remain cytotoxic for approximately 2 weeks longer.
4 Notes
-
1.
The presented protocols can be applied to any source of human CD34-positive cells [14]. However, the final yield of differentiated NK cells is higher using fetal liver or umbilical cord blood, as opposed to bone marrow or mobilized peripheral stem cells [19].
-
2.
The presence of a co-delivered marker, such as enhanced green fluorescent protein (EGFP) or a selection component, will allow enrichment for gene-modified NK cell precursors at any step of the differentiation protocol.
-
3.
It is not clear if DL1 expression is necessary for successful generation of mature and functional NK cells. Preliminary evidence suggests that OP9-DL1 co-culture promotes faster differentiation and higher cell proliferation [13, 15, 20].
-
4.
In clinically relevant experiments, the viral vector copy number per cell in HSC should be kept within 1–3 copies/cell, in order to avoid genotoxicity.
-
5.
A plate coated with RetroNectin can be preserved at 4 °C for up to 7 days well filled with 1 mL of wash buffer.
-
6.
In the co-culture protocol, we have used non-irradiated OP9-DL1 stromal cells.
-
7.
Start OP9-DL1 stromal cell culture 1–2 weeks before HSC transduction, in order to have at least a full T-75 flask with confluent stroma before starting the protocol.
-
8.
Passage OP9-DL1 cells cultured in flasks every 3–5 days and when no more confluent than 90 % split cells at 1:5–1:10, i.e., using one-fifth to one-tenth of the volume of cell suspension from one flask to reseed a new flask. Fully confluent stroma decays in about 10–14 days. Timing of the passage of OP9-DL1 cells with the seeding of plates for the co-culture with HSC will optimize the protocol and avoid waste.
-
9.
OP9-DL1 cells should be at 90 % confluence before seeding the human cells, what usually takes 2–4 days after seeding on a tissue culture-treated 12 well plate.
-
10.
Seed the HSC over fresh confluent OP9-DL1 stromal cells each time HSC are passaged. During the first 2 weeks, the human cells will proliferate faster and may require passaging more often.
-
11.
In the OP9-DL1 co-culture protocol, differentiated NK cells will develop cytotoxicity against the stromal cells around day 35 of culture, as a surrogate of successful differentiation.
-
12.
The use of a NK cell expansion protocol will increase the final yield of the cell generation when used in sequence to the protocols described in this chapter. Cells should be transferred to the expansion protocol at day 30 or later, when cells have higher expression of CD16 and other mature NK cell-specific markers.
-
13.
In preparation of AIM V CC, human AB serum must be heat inactivated at 56 °C for 30 min and sterile filtered to remove excess lipids. Alternatively, the AIM V Complete medium can be sterile filtered prior to the addition of cytokines.
-
14.
In order to resuspend the cell pellet, it is important to flick the tube to disperse the cells rather than to directly pipet the pellet, which can cause shearing and decrease viability of cells at this point.
-
15.
As HSC differentiate, adherent cells such as endothelial cells and macrophages will become observable. NK and NK progenitors, however, are not strongly adherent; therefore, during splits, pipet to detach cells that are easily detachable and do not attempt to remove strongly adherent cells. If the well becomes overgrown with adherent cells, do not continue to use it for NK differentiation and expansion .
References
Campbell KS, Hasegawa J (2013) Natural killer cell biology: an update and future directions. J Allergy Clin Immunol 132:536–544
Childs RW, Berg M (2013) Bringing natural killer cells to the clinic: ex vivo manipulation. Hematology Am Soc Hematol Educ Program 2013:234–246
Davies JOJ, Stringaris K, Barrett JA et al (2014) Opportunities and limitations of natural killer cells as adoptive therapy for malignant disease. Cytotherapy 16:1453–1466
Leung W (2014) Infusions of allogeneic natural killer cells as cancer therapy. Clin Cancer Res 20:3390–3400
Moretta L, Montaldo E, Vacca P et al (2014) Human natural killer cells: origin, receptors, function, and clinical applications. Int Arch Allergy Immunol 164:253–264
Knorr DA, Bachanova V, Verneris MR et al (2014) Clinical utility of natural killer cells in cancer therapy and transplantation. Semin Immunol 26:161–172
Micucci F, Zingoni A, Piccoli M et al (2006) High-efficient lentiviral vector-mediated gene transfer into primary human NK cells. Exp Hematol 34:1344–1352
Shook DR, Campana D (2011) Natural killer cell engineering for cellular therapy of cancer. Tissue Antigens 78:409–415
Chu J, Deng Y, Benson DM et al (2014) CS1-specific chimeric antigen receptor (CAR)-engineered natural killer cells enhance in vitro and in vivo antitumor activity against human multiple myeloma. Leukemia 28:917–927
Boissel L, Betancur M, Wels WS et al (2009) Transfection with mRNA for CD19 specific chimeric antigen receptor restores NK cell mediated killing of CLL cells. Leuk Res 33:1255–1259
Boissel L, Betancur-Boissel M, Lu W et al (2013) Retargeting NK-92 cells by means of CD19- and CD20-specific chimeric antigen receptors compares favorably with antibody-dependent cellular cytotoxicity. Oncoimmunology 2, e26527
Glienke W, Esser R, Priesner C et al (2015) Advantages and applications of CAR-expressing natural killer cells. Front Pharmacol 6:1–7
De Smedt M, Taghon T, Van de Walle I et al (2007) Notch signaling induces cytoplasmic CD3 epsilon expression in human differentiating NK cells. Blood 110:2696–2703
Luevano M, Madrigal A, Saudemont A (2012) Generation of natural killer cells from hematopoietic stem cells in vitro for immunotherapy. Cellular Mol Immunol 9:310–320
De Oliveira SN, Ryan C, Giannoni F et al (2013) Modification of hematopoietic stem/progenitor cells with CD19-specific chimeric antigen receptors as a novel approach for cancer immunotherapy. Hum Gene Ther 24:824–839
Cany J, van der Waart AB, Tordoir M et al (2013) Natural killer cells generated from cord blood hematopoietic progenitor cells efficiently target bone marrow-residing human leukemia cells in NOD/SCID/IL2Rgnull mice. PLoS One 8:1–11
Ni F, Sun R, Fu B et al (2013) IGF-1 promotes the development and cytotoxic activity of human NK cells. Nat Commun 4:1479
Cooper AR, Patel S, Senadheera S et al (2011) Highly efficient large-scale lentiviral vector concentration by tandem tangential flow filtration. J Virol Methods 177:1–9
Luevano M, Domogala A, Blundell M et al (2014) Frozen cord blood hematopoietic stem cells differentiate into higher numbers of functional natural killer cells in vitro than mobilized hematopoietic stem cells or freshly isolated cord blood hematopoietic stem cells. PLoS One 9:e87086
La Motte-mohs RN, Herer E, Zu JC (2005) Induction of T-cell development from human cord blood hematopoietic stem cells by Delta-like 1 in vitro. In Vitro 105:1431–1439
Acknowledgment
Support was provided by the UCLA Department of Pediatrics (K-12 UCLA Child Health Research Center Development Award (CHRCDA)), Gwynne Hazen Cherry Memorial Fund, Pediatric Cancer Research Foundation, UCLA Children’s Discovery and Innovation Institute, UCLA Jonsson Comprehensive Cancer Center, UCLA/CFAR Virology Core Lab, UCLA Clinical and Translational Science Institute Grant UL1TR000124, Lights Camera Cure and St. Baldrick’s Foundation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Lowe, E., Truscott, L.C., De Oliveira, S.N. (2016). In Vitro Generation of Human NK Cells Expressing Chimeric Antigen Receptor Through Differentiation of Gene-Modified Hematopoietic Stem Cells. In: Somanchi, S. (eds) Natural Killer Cells. Methods in Molecular Biology, vol 1441. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3684-7_20
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3684-7_20
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3682-3
Online ISBN: 978-1-4939-3684-7
eBook Packages: Springer Protocols