Abstract
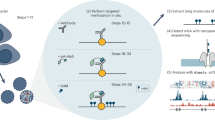
Interaction and co-occurrence of protein and DNA-based epigenetic modifications have become a topic of interest for many fundamental and biomedical questions. We describe within this chapter a protocol that combines two techniques in order to determine the methylation status of the DNA specifically associated with a protein of interest. First, DNA that directly interacts with the selected protein (such as a specific histone modification, a transcription factor, or any other DNA-associated protein) is purified by standard chromatin immunoprecipitation (ChIP). Second, the level of DNA methylation of this immunoprecipitated DNA is measured by bisulfite conversion and Pyrosequencing, a quantitative sequencing-by-synthesis method. This procedure allows determining the methylation status of genomic DNA associated to a specific protein at single nucleotide resolution.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Shanmuganathan R, Basheer NB, Amirthalingam L et al (2013) Conventional and nanotechniques for DNA methylation profiling. J Mol Diagn 15:17–26
Suzuki M, Greally JM (2013) Genome-wide DNA methylation analysis using massively parallel sequencing technologies. Semin Hematol 50:70–77
Mensaert K, Denil S, Trooskens G et al (2014) Next-generation technologies and data analytical approaches for epigenomics. Environ Mol Mutagen 55:155–170
Furey TS (2012) ChIP-seq and beyond: new and improved methodologies to detect and characterize protein-DNA interactions. Nat Rev Genet 13:840–852
Tost J, Dunker J, Gut IG (2003) Analysis and quantification of multiple methylation variable positions in CpG islands by Pyrosequencing. Biotechniques 35:152–156
Tost J, Gut IG (2007) DNA methylation analysis by pyrosequencing. Nat Protoc 2:2265–2275
Dupont JM, Tost J, Jammes H et al (2004) De novo quantitative bisulfite sequencing using the pyrosequencing technology. Anal Biochem 333:119–127
Ahmadian A, Ehn M, Hober S (2006) Pyrosequencing: history, biochemistry and future. Clin Chim Acta 363:83–94
Moison C, Assemat F, Daunay A et al (2014) Synergistic chromatin repression of the tumor suppressor gene RARB in human prostate cancers. Epigenetics 9:477–482
Carey MF, Peterson CL, Smale ST (2009) Chromatin immunoprecipitation (ChIP). Cold Spring Harb Protoc. 2009:pdb prot5279
Kuo MH, Allis CD (1999) In vivo cross-linking and immunoprecipitation for studying dynamic Protein:DNA associations in a chromatin environment. Methods 19:425–433
Orlando V (2000) Mapping chromosomal proteins in vivo by formaldehyde-crosslinked-chromatin immunoprecipitation. Trends Biochem Sci 25:99–104
Johnson KD, Bresnick EH (2002) Dissecting long-range transcriptional mechanisms by chromatin immunoprecipitation. Methods 26:27–36
Ren B, Dynlacht BD (2004) Use of chromatin immunoprecipitation assays in genome-wide location analysis of mammalian transcription factors. Methods Enzymol 376:304–315
Bernstein BE, Humphrey EL, Liu CL et al (2004) The use of chromatin immunoprecipitation assays in genome-wide analyses of histone modifications. Methods Enzymol 376:349–360
Bernstein BE, Mikkelsen TS, Xie X et al (2006) A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125:315–326
Brinkman AB, Pennings SW, Braliou GG et al (2007) DNA methylation immediately adjacent to active histone marking does not silence transcription. Nucleic Acids Res 35:801–811
Statham AL, Robinson MD, Song JZ et al (2012) Bisulfite sequencing of chromatin immunoprecipitated DNA (BisChIP-seq) directly informs methylation status of histone-modified DNA. Genome Res 22:1120–1127
Campan M, Weisenberger DJ, Trinh B et al (2009) MethyLight. Methods Mol Biol 507:325–337
Kramer A, Mailand N, Lukas C et al (2004) Centrosome-associated Chk1 prevents premature activation of cyclin-B-Cdk1 kinase. Nat Cell Biol 6:884–891
Li LC, Dahiya R (2002) MethPrimer: designing primers for methylation PCRs. Bioinformatics 18:1427–1431
Aranyi T, Varadi A, Simon I et al (2006) The BiSearch web server. BMC Bioinformatics 7:431
Moison C, Senamaud-Beaufort C, Fourriere L et al (2013) DNA methylation associated with polycomb repression in retinoic acid receptor beta silencing. FASEB J 27:1468–1478
Kurdistani SK, Grunstein M (2003) In vivo protein-protein and protein-DNA crosslinking for genomewide binding microarray. Methods 31:90–95
Nowak DE, Tian B, Brasier AR (2005) Two-step cross-linking method for identification of NF-kappaB gene network by chromatin immunoprecipitation. Biotechniques 39:715–725
O’Neill LP, Turner BM (2003) Immunoprecipitation of native chromatin: NChIP. Methods 31:76–82
Tost J, El abdalaoui H, Gut IG (2006) Serial pyrosequencing for quantitative DNA methylation analysis. Biotechniques 40:721–726
Acknowledgments
This work was supported by ATIP CNRS and Equipe d’Excellence Région Midi-Pyrenées grant to PBA. CM was recipient of an MNERT fellowship.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Moison, C., Assemat, F., Daunay, A., Arimondo, P.B., Tost, J. (2015). DNA Methylation Analysis of ChIP Products at Single Nucleotide Resolution by Pyrosequencing® . In: Lehmann, U., Tost, J. (eds) Pyrosequencing. Methods in Molecular Biology, vol 1315. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2715-9_22
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2715-9_22
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2714-2
Online ISBN: 978-1-4939-2715-9
eBook Packages: Springer Protocols