Abstract
Abiotic stresses, especially drought, high salinity, flooding, and extreme temperatures, have become a big concern due to their high frequency of occurrence and usually beyond human control capacity, as well as their severe impacts on agricultural crop productivities. Under the pressures of climate change and reduction in total cultivated land worldwide for other purposes, sustaining food security to feed an increasing human population while coping with these environmental constraints is a greater challenge than ever. Generating new varieties with better traits based on gene exchange from available sources via conventional breeding methods currently no longer provides an adequate solution in coping with abiotic stresses. Therefore, another research theme attracting the scientists over the past 20 years has been to elucidate molecular mechanisms that the plants employ to defend and adapt to stress conditions. The final aims are to identify and characterize the function of important genes involved in plant responses to stress that can be used for genetic manipulation. Thanks to advances in molecular biotechnology, including gene transfer techniques such as particle bombardment, microinjection, and Agrobacterium-mediated transformation, new varieties with better stress tolerance and yield production could be made by this strategy; thus, in combination with traditional approaches, development of new lines with improved traits has become more practical. According to our current knowledge, transcription factors (TFs) have been recognized to play essential roles in regulating plant responses against adverse abiotic factors. Many TFs belonging to families AP2/EREBP, bZIP, MYB, WRKY, and NAC have been reported to participate in plant responses to various stressors. A number of TFs whose encoding genes are appropriately altered in expression level have shown enhanced tolerance capacity toward drought, salt, and suboptimal temperatures in transgenic model and crop plants. In this chapter, we summarize our current understanding about TF activities in plants under adverse stress conditions and their use in crop improvement.
Xuan Lan Thi Hoang and Nguyen Binh Anh Thu contributed equally to this work.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Abiotic stress is defined as a “non-living environmental factor that can negatively or even harmfully affect the growth and productivity of plant” (Ku et al. 2013). Among the abiotic stressors, drought is the biggest threat to plants in general and to crops in particular. Shortage of water supply leads to reduction of photosynthesis due to lack of intracellular CO2 availability because of stress-induced stomatal closing and reduction of root capacity to absorb nutrients in soil, and thus total energy production of the plant is decreased (Lawlor and Cornic 2002; Lawlor and Tezara 2009; Aroca et al. 2012). Excessive light can exacerbate stress in plants by affecting its photosynthesis. When chloroplasts are exposed to excessive light, photooxidation is induced, which results in the increased production of reactive oxygen species (ROS), such as hydrogen peroxide, superoxide anion, and hydroxyl radicals (Grassi and Magnani 2005; Aguado-Santacruz 2006; Zhou et al. 2007; Miller et al. 2010; Osakabe et al. 2014). These biological molecules can cause negative effects on plant productivity. Moreover, water deficit triggers the stomata closure and a decrease in leaf water potential as well as the downregulation of photosynthesis-related genes and slow diffusion of CO2 (Osakabe et al. 2014). More severe drought exposures can finally lead to plant death.
The invasion of salinity into mainland and higher frequency of flooding as a consequence of ice melt or urbanization process make these the second- and third-ranked concerns to sustainable agriculture. Under high salinity, susceptible plants suffer ion toxicity due to excessive entry of ions, particularly Na+, into the cells, resulting in homeostatic disruption, water loss, and oxidative stress , and thereof inhibition of photosynthesis and enzyme activities, cell division, and plant growth (Munns et al. 2006; Munns and Tester 2008; Parvaiz and Satyawati 2008; Shabala and Munns 2012). It is projected that by 2050, more than half of arable land would encounter salinity problem (Wang et al. 2003; Setia et al. 2011). Opposite to water deficit, waterlogging is another type of water stress whereby it prevents plants, or at least their root part, from accessing oxygen supplied from environment (Mahajan and Tuteja 2005; Calvo-Polanco et al. 2012; Kreuzwieser and Rennenberg 2014). The consequences of this are negative effects on root functions in respiration and taking up nutrients (Mahajan and Tuteja 2005). Meanwhile, exposure to nonoptimal temperature conditions, such as cold (chilling and frost) and heat (high temperature), may cause injuries to cellular structure and functions, and thus significantly affecting plant metabolism, development, and reproduction (Yadav 2010; Theocharis et al. 2012; Hasanuzzaman et al. 2013). For example, cellular ice formation, dehydration, and fluidity movement reduction in the phospholipid membrane are among early effects caused by low temperatures (Mahajan and Tuteja 2005; Wang et al. 2006).
It has been known that chilling, freezing, heat, drought, and salinity all lead to hyperosmotic condition as a secondary stress in plants. Depletion in cellular water is the common consequence caused by each of these stress factors. Hyperosmosis is explained as a condition in which there is an increase in concentration of extracellular solutes due to water deficit, and thus triggering the movement of water out of the cells into the environment (Aguado-Santacruz 2006). As a result, cellular turgor cannot be maintained and intracellular concentration of solutes is increased (Aguado-Santacruz 2006; Solanke and Sharma 2008; Chaves et al. 2009). It is important to mention that although some of the molecular responses to these types of stressors overlap as they produce similar effects in the water status of cells, stress-specific responses are also present depending on which particular stress the plant is facing with (Aguado-Santacruz 2006).
In addition to the well-mentioned abiotic stressors, such as drought , salinity, heat, cold, and flooding, there are other abiotic stressors appearing with less frequency, including UV radiation, heavy metal toxicity, and inorganic nutrient deficiency (Clemens 2006; Hossain et al. 2012; Hideg et al. 2013; Liang et al. 2013). Nowadays, abiotic stresses are not purely considered as natural causes since their occurrence and severity are under the influence of human activities such as not only inappropriate practices in agriculture like intensive land use without break and excessive organic fertilization but also in other areas, such as deforestation and industrial gas release into the atmosphere. As a consequence of these human errors, climate change has been a hot topic for discussion, and of course this exacerbates unpredictability of abiotic stresses, making them more serious threats to the food security of humankind.
To reduce agricultural loss by the impact of abiotic stresses, scientists have been trying to widen our current understanding of the mechanisms of plant tolerance to various abiotic stressors by using different methodologies. Although how expression of genes alters under stress conditions can be easily determined, working out their roles in the stress responses and identifying their relationship with other members in the whole picture of stress acclimation are not easy tasks (Chaves et al. 2009). That may be the reason why the number of successes in the development of crop plants coping with the major stressors, including water scarcity, salinity, and temperature stresses, is still limited. To accelerate this process, a combination of modern techniques in molecular biology, advances in phenotyping, quantitative trait locus (QTL) mapping, and conventional breeding strategies has been used. The purposes of this chapter are to summarize our current knowledge about stress-adaptive mechanisms in plants, mainly focusing on major abiotic stresses (drought, salinity, and extreme temperatures) and transcription factors (TFs)—important members in the stress-adaptive pathways—and then review the progress in applications of TFs in improving abiotic stress tolerance in crop plants.
2 Mechanisms in Plant Responses to Abiotic Stresses
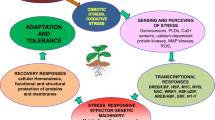
It is predicted that abiotic stresses will continue to be major constraints affecting global crop yields (Sharma and Lavanya 2002; Osakabe et al. 2013b), and thus, through evolution plants have developed various mechanisms to cope with them. These mechanisms have been classified into three groups: (1) stress escape by adjusting plant development and reproduction in accordance with the environmental conditions; (2) stress avoidance by developing morphologically and physiologically advantageous traits; and (3) stress tolerance by regulating morphological, physiological, biochemical, and molecular activities to minimize the stress impact (Turner et al. 2001; Mittler 2002; Chaves et al. 2003). The last defending strategy attracts the largest research attention and hence will be discussed in detail later in this section.
2.1 Physiological and Biochemical Responses
Adaptation to abiotic stress is undoubtedly one of the most complex biological processes in plants since it involves multigenic traits. Reponses that are usually seen at the physiological level are closure of stomata, shedding of leaves, decrease in light absorbance and photosynthetic activities, adjustment in relative water content, increase in root growth, and reduction in shoot growth (Manavalan et al. 2009; Shanker and Venkateswarlu 2011; Thu et al. 2014; Ha et al. 2013). For example, under high CO2 and abiotic stress conditions, excess C may favor root growth over that of photosynthetic organs (Hachiya et al. 2014). Shoot-growth restriction could be considered as an advantage to help plants preserve limited carbohydrate resource for maintaining the most important metabolism activities to ensure their survival over the stress period and their capacity to recover quickly after the demission of the stress (Aguado-Santacruz 2006). In contrast, continuation of root growth is favorably promoted to search for water from deeper soil layers (Alsina et al. 2011; Aroca et al. 2012). Examination of various root characteristics, such as dry weight, total length, and number of lateral roots, enables us to quickly assess the tolerant capacity of a particular plant variety under stress conditions (Liu et al. 2005; Gowda et al. 2011; Kumar et al. 2012). Other physiological symptoms can be observed under abiotic stress, including decrease in leaf expansion, wilting, discoloration, chlorosis, surface lesion and/or sterility at reproduction stage, senescence, stunted shoots, and lower biomass production (Wen et al. 2002; Yadav 2010; Jaarsma et al. 2013). Hence, many common parameters, such as survival rate, productivity, and relative growth rate, are used for the evaluation of plant tolerance to various stresses (Ashraf and Harris 2004; Parvaiz and Satyawati 2008).
The most important, commonly seen response of plants to major abiotic stresses at biochemical level is perhaps the osmotic adjustment. A wide variety of solutes are involved in this regulation, including amino acids and amino acid-associated compounds (proline, aspartate, glutamate, and glycine betaine etc.), sugars (sucrose, fructose, fructans, and trehalose etc.), and polyols (mannitol, and sorbitol etc.) (Chaves et al. 2003; Koyro et al. 2012). These metabolites are also called osmo-protective substances that contribute to maintaining the cell turgor and prevention of water loss. Additionally, increase of other products, such as antioxidant substances (e.g., ascorbate, glutathione, carotenoids, anthocyanins, and α-tocopherol), detoxifying enzymes (e.g., catalase, superoxide dismutase, peroxidase, ascorbate peroxidase, glutathione peroxidase, and glutathione reductase), heat shock proteins (HSPs), and late embryogenesis abundant (LEA) proteins, is also observed under stresses (Goyal et al. 2005; Chakrabortee et al. 2010; Aguado-Santacruz 2006). Moreover, under stress conditions, such as drought and salinity, transportation activities of anions and cations, including Cl−, Na+, and K+, as well as the participation of water transport systems play important roles to maintain an ion balance between tissues and cells in stress adaptation (Osakabe et al. 2013a; Osakabe et al. 2014).
2.2 Signal Perception, Transduction, and Regulating Pathways Involved in Plant Responses to Abiotic Stresses
Regulation of molecular activities such as gene expression is considered the most critical factor determining the response degree of a plant as a whole toward a specific stress condition. In this process, being able to sense changes in the surrounding environment is the first important step for plants to respond, survive, and adapt to living conditions. Disruption of normal intracellular homeostasis in plants caused by stress is regarded as early alerting signals that include alterations of cellular size, solute concentration, and cellular turgidity and/or increase in ROS levels (Loutfy et al. 2012; Suzuki et al. 2012). A number of stress detectors have been identified in plants and most of them are located on the cell membrane (Mahajan and Tuteja 2005). Under cold, drought, or salinity stress conditions, Ca2+ ions have been shown to be shuttled into the cell cytoplasm, indicating participation of Ca2+ permeable channels in the stress-responsive networks as stress signal receptors (Boudsocq and Sheen 2013). Calcium ions have been indicated to interact with kinase enzymes, including Ca-dependent protein kinases (CDPKs), calmodulin-binding protein kinases, and Ca2+/calmodulin-dependent protein kinases that are known to participate in signaling pathways via phosphorylation activities. Taking a strategy used by plants to cope with salt stress as an example, the salt overly sensitive (SOS)-responsive pathway, which aids the reestablishment of intracellular ionic balance, includes a sensor Ca-binding protein SOS3 that is able to mediate the activity of kinase SOS2 (Boudsocq and Laurière 2005; Thapa et al. 2011). These activated kinases will in turn directly regulate activities of TFs in the downstream transduction pathway (Chinnusamy et al. 2004). It was reported that an increase in ROS concentration during stress also led to the enhancement of activity of these Ca channels (Mori and Schroeder 2004).
Another type of membrane protein sensors involved in the perception of drought , high salinity, and low temperature, which deserves to be mentioned, is histidine kinases (HKs) that are members of two-component system (TCS). In Arabidopsis thaliana, a number of HKs (AHKs) have been identified (Schaller et al. 2008). HKs functioning as osmosensors, such as Sln1 and Sho1, are also present in yeast in which both of them are membrane spanning but have no similarity in structure. In Arabidopsis, a homolog of Sln1, AtHK1/AHK1, has been identified which is able to suppress the salt-sensitive phenotype of the yeast double-mutant sln1∆ sho1∆ that lacked both osmosensors (Urao et al. 1999). The interaction between AHK1 and the downstream Arabidopsis histidine phosphotransfers (AHPs) in the multistep phosphorelay has been identified (Urao et al. 2000; Kumar et al. 2013). In addition to these above-described sensors, G protein-associated receptors may function in the perception of a secondary signal derived from these stresses (Perfus-Barbeoch et al. 2004; Misra et al. 2007). They might also serve as a kind of membrane-bound receptors for abscisic acid (ABA; Pandey et al. 2009).
It is important to emphasize that a specific abiotic stress can generate more than one type of signals that can be detected by one or more sensors (Gong et al. 2013). For example, drought causes both osmotic and oxidative stresses to plants. Oxidative stress and appearance of photorespiration as secondary stress effects from drought can lead to increased production of ROS, possibly deriving from organelles such as chloroplasts, mitochondria, glyoxysomes, and peroxisomes, and from activities of β-oxidation of lipids and several different oxidases (Apel and Hirt 2004; Miller et al. 2010; Voss et al. 2013). ROS inhibits the function of many enzymes, increases protein and DNA susceptibility to degradation due to structural modifications, reduces permeability of plasma membrane, and stimulates the activity of Ca-dependent proteases and nucleases (Sharma et al. 2012).
The participators working as second messengers can be divided into three groups, including: (1) diacylglycerol, inositol triphosphate, such as IP3, and phosphate idylinositols (group of hydrophobic molecules) which are located on the membrane and are able to pass the signal to membrane-associated effector proteins; (2) cAMP, cGMP, sugars, such as sucrose, glucose, and fructose, and Ca2+ (group of hydrophilic molecules) which are presented within the cytosol; and (3) nitric oxide and carbon monoxide (group of gas molecules) which can diffuse through cytosol and cellular membranes (Bhargava and Sawant 2013; Aguado-Santacruz 2006; Chaves et al. 2009). Additionally, ROS can also be assigned as another member of second messenger since its damage effects to cellular components during stress switch on specific genes involved in the activation of mitogen-activated protein kinase (MAPKs) as well as the production of antioxidants , HSPs, and ROS-scavenging enzymes.
Several second messengers, such as IP3, can regulate intracellular Ca2+ levels, while the Ca2+ ion itself is also regarded as a second messenger. Ca2+ has been known to be involved in the closure of stomatal aperture (Zhang et al. 2011a), and often initiates a protein phosphorylation cascade that regulates expression of downstream regulatory and functional genes, contributing to stress tolerance (Gong et al. 2013). It is noticed that in order to successfully transfer the signal message through the whole signal transduction pathway, extra assistants that are in charge of modification, delivery, or assembly of signaling components are required as well. Examples of these molecules are protein modifiers involved in protein lipidation, methylation, glycosylation, and ubiquitination; scaffolds; and adaptors (Xiong et al. 2002).
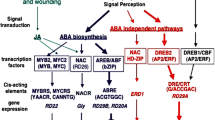
A great variety of other elements are also recruited in the networks, leading to acclimative reactions to stress, such as plant growth regulators, including phytohormones, polyamines, and Ca, and various proteins, including CDPKs, MAPKs (including MAPKKKs, MAP kinase kinase kinases; MAPKKs, MAP kinase kinases), and TFs (Ohnishi et al. 2008; Huang et al. 2012). Particularly, involvement of ABA and its induced biosynthesis in response to various abiotic stresses, such as salinity, drought, and cold, have been well reported in literature (Fujita et al. 2011; Danquah et al. 2014). Many stress-responsive functional genes, such as RD22 (responsive to dehydration 22), RD29A, COR15 (cold responsive 15), COR47, and P5CS (-Δ-1-pyrroline-5-carboxylate synthase), and regulatory genes, including dehydration-responsive element-binding protein (DREB), ABA-responsive element (AREB), myeloblastosis (MYB), and NAC (NAM, no apical meristem; ATAF, Arabidopsis transcription activation factor; and CUC, cup-shaped cotyledon), whose expression is regulated by ABA, have been identified to be involved in regulating plant responses to stresses (Xiong et al. 2001; Dalal et al. 2009; Fujita et al. 2011; Nakashima et al. 2009; Tran et al. 2010).
Protein phosphorylation is an important and effective mechanism that plants use in the signal transduction process to trigger stress responses as quickly as possible. In plants, in addition to the TCSs, activities of the two classes of stress-activated protein kinases, MAPKs and CDPKs, are also performed via phosphorylation at specific amino acid residues present in the structure (Schaller et al. 2008; Huang et al. 2012). In the working module of MAPK members, an MAPKKK is activated by phosphorylation which in turn phosphorylates the activity of MAPKK (Bhargava and Sawant 2013). At the end of the phosphorylation cascade, activation of a cytoplasmic MAPK often results in its translocation into the nucleus to regulate gene expression via controlling TFs also by phosphorylation (Danquah et al. 2014). Alternatively, these terminal MAPKs remain in the cytoplasma where they phosphorylate enzymes or cytoskeleton components (Danquah et al. 2014).
In further progression of the signaling process, TFs should be highlighted as the key participators. They work as final transducers in the transduction module and directly mediate gene expression since they can bind to regulatory regions of specific promoters (e.g., cis-acting elements). There is a fact that expression of a gene can be regulated by several mechanisms of which some are still unknown (Weake and Workman 2010). Nonetheless, the regulation of gene expression via the interaction of TFs and promoters are well documented (Yamaguchi-Shinozaki and Shinozaki 2006). Detailed information about TFs will be discussed in the next section of this chapter.
3 Plant TFs and their Roles in Abiotic Stress Responses
In plant genomes, approximately 7 % of the coding sequences are assigned to TFs (Udvardi et al. 2007), which are divided into families based on their distinct signatures in structure. Major families are well known to be involved in various abiotic stresses either in ABA-dependent (MYB/MYC, bZIP) (Zhang et al. 2009a; Fujita et al. 2011), ABA-independent pathway (WRKY) (Chen et al. 2012; Rushton et al. 2008; Umezawa et al. 2006), or in both pathways (NAC, AP2/EREBP) (Souer et al. 1996; Olsen et al. 2005; Nakashima et al. 2012; Puranik et al. 2012). With the aim to generate new varieties, which are able to cope with abiotic stress(es) more efficiently, genetic engineering has been considered as a powerful approach in addition to conventional breeding methods. Thanks to recent advances in technologies, including real-time quantitative polymerase chain reaction (RT-qPCR), omic technologies, cloning, transformation, gene overexpressing and silencing, and other biochemical, molecular, and physiological methods employed in stress-response analyses at physiological and molecular levels, essential candidate genes contributing to plant adaptation to stress have been identified. These genes include those encoding members functioning in various stages of stress signal transduction cascade. Among these, TFs have drawn particular attention due to their important function in stress regulation and their potential in genetic engineering. Therefore, numerous attempts have been made all around the world, mainly by overexpressing the TF-encoding genes, and several promising results have been reported. Table 14.1 summarizes a number of studies using TFs to create transgenic model and crop plants with improved abiotic stress tolerance within the past 5 years.
3.1 The AP2/EREBP Family
In plants, AP2/EREBP (APETALA2/ethylene-responsive element-binding protein) has been known as a large family of TF genes which contains the highly conserved AP2/ethylene-responsive element-binding factor (ERF) DNA-binding domain (Riechmann and Meyerowitz 1998), and has been identified in various species, such as Arabidopsis, tobacco (Nicotiana tabacum), and tomato (Solanum lycopersicum) (Fischer and Dröge-Laser 2004; Oñate-Sánchez and Singh 2002; Tournier et al. 2003). Based on the number of AP2/ERF domains and the gene structure, the AP2/EREBP gene family can be divided into four subfamilies AP2, RAV (related to ABI3/VP1), DREB, and ERF (Sakuma et al. 2002; Sharoni et al. 2011). Among these, the ERF and DREB TF subfamilies were discovered in various plant species, including rice (Oryza sativa) (Quan et al. 2010), Arabidopsis (Sakuma et al. 2006), and tobacco (Agarwal et al. 2010), and their functions in the plant responses to biotic and abiotic stresses have been extensively studied (Agarwal et al. 2006; Agarwal et al. 2010).
The DREB-type TFs contain a conserved DNA-binding domain of 58–60 amino acids, and thus are able to bind to the PyCCGACAT cis-elements named as DRE/C-repeat (CRT) motifs located in the promoter regions of target genes to activate or suppress their transcription for achieving stress adaptation (Park et al. 2001; Ito et al. 2006; Jaglo et al. 2001; Kasuga et al. 1999; Liu et al. 1998; Agarwal et al. 2006; Sakuma et al. 2002). According to Sakuma et al. (2002), DREB subfamily has been further classified into six subgroups termed A-1 to A-6, of which A-1 and A-2 are the two largest groups. DREB1/C-repeat-binding factor (CBF; A-1) subgroup of Arabidopsis with three major regulator factors, DREB1A/CBF3, DREB1B/CBF1, and DREB1C/CBF2, has shown its importance in plant responses to cold stress. Overexpression of RD29A:DREB1 or 35S:CBF genes showed significantly improved tolerance to freezing, drought, and high salinity in Arabidopsis (Jaglo-Ottosen et al. 1998; Liu et al. 1998; Mizoi et al. 2012). Furthermore, the DREB1s can be used to produce transgenic crops with higher tolerance to drought, high salt, high temperature, and cold stress. Bouaziz et al. (2013) reported that transgenic potato plants overexpressing 35S:StDREB1, which had been isolated from potato (S. tuberosum) and classified in the A-4 group of DREB subfamily based on multiple sequence alignments and phylogenetic characterization, had enhanced salt and drought tolerance. Moreover, StDREB1 was significantly induced by NaCl, drought, low temperature, and ABA conditions in potato. In Populus trichocarpa, the development of Populus varieties with a greater tolerance to many adverse environments has been aided by understanding the characterization of the DREB subfamily (Chen et al. 2013b). Based on the expression analysis of 15 selected PtrDREB genes under abiotic stress conditions in Populus, Chen et al. (2013b) claimed that there were 14 genes with induced expression under different abiotic stresses. GmDREB2 isolated from soybean is a novel DREB gene of A-5 subgroup which was found to have induced expression by drought, high salt, and low temperature stresses (Chen et al. 2007). Overexpression of GmDREB2 by constitutive 35S promoter or stress-inducible RD29A promoter resulted in upregulation of downstream genes in transgenic Arabidopsis (Chen et al. 2007). As a result, the transgenic plants showed enhanced tolerance to drought and high salinity without any growth retardation even with constitutive overexpression (Chen et al. 2007), as was observed with constitutive overexpression of OsDREB1A (Dubouzet et al. 2003). LeDREB2 was discovered from tomato genome and classified into an A-2 group member of the DREB family. The expression of this gene was induced by high salinity, drought, and cold (Islam and Wang 2012). In another independent research on expression analysis in Pisum sativum, PsDREB2A was reported to be induced in pea roots and leaves under water deficit (Jovanovic et al. 2013). Mallikarjuna et al. (2011) developed transgenic rice lines overexpressing OsDREB2A under control of stress-inducible RD29A promoter, which resulted in the enhanced growth performance and significant tolerance to osmotic, salt, and dehydration stresses during simulated stress conditions as compared with the wild type. Later on, Zhang et al. (2013) used this gene to enhance salt tolerance of soybeans. The authors reported that salt tolerance was enhanced in the 35S:OsDREB2A transgenic soybean plants due to accumulation of osmolytes, such as soluble sugars and free proline, as well as induced expression of several stress-responsive TFs and key genes (Zhang et al. 2013). LcDREB3a from the drought-tolerant forage grass Leymus chinensis was shown to function in the improvement of drought and salt tolerance in Arabidopsis overexpressing 35S:LcDREB3a without causing growth retardation by inducing expression of stress tolerance genes when compared to control (Peng et al. 2011). Besides, transgenic expression of another LcDREB member (LcDREB2) in combination with its downstream gene (S-adenosyl-methionine decarboxylase, LcSAMDC2,-encoding gene) obtained from L. chinensis under the control of the 35S promoter could improve the salt tolerance in Arabidopsis (Peng et al. 2013). In Malus domestica, RT-qPCR analysis showed significant upregulation of some putative MdDREB genes under various abiotic stress treatments, which proved their vital roles during stress adaptation (Zhao et al. 2012).
In another subfamily of AP2/EREBP TFs, ERFs have been indicative of their participation in plant responses to biotic and abiotic stresses by recognizing the cis-acting element AGCCGCC, known as the GCC box (Hao et al. 1998; Ohme-Takagi and Shinshi 1995; Fujimoto et al. 2000). Based on the phylogenetic analyses of 125 AP2/ERF members in Arabidopsis, the ERF subfamily could be divided into six subgroups, from ERF-B1 to ERF-B6 (Sakuma et al. 2002). In wheat (Triticum aestivum), 47 ERF-encoding genes have already been identified (Zhuang et al. 2011). Among these, constitutively overexpressing TaPIE1 controlled by maize ubiquitin promoter in wheat exhibited significantly enhanced resistance to both pathogen (triggered by Rhizoctonia cerealis) and freezing stress, whereas constitutive knockdown wheat plants by a recombinant construct between barley stripe mosaic virus (BSMV) and TaPIE1 were more susceptible to both stresses relative to control plants (Zhu et al. 2014). Functional analysis of TSRF1, a member of tomato ERF TFs, demonstrated that overexpression of 35S:TSRF1 improved the osmotic and drought tolerance of rice seedlings without growth retardation as indicated by physiological analyses of root and leaf growth, leaf water loss, and survival rate under stress conditions compared to control (Quan et al. 2010). In another study in rice, Joo et al. (2013) reported that ERF genes, including OsERF4a and OsERF10a, had an important contribution in conferring drought stress tolerance. Both constitutive and ABA-inducible expression of the ERF-associated amphiphilic repression (EAR) domain-containing protein-encoding OsERF4a showed increased drought tolerance as a consequence of suppression of a putative repressor Silent information regulator 2 (Sir2) involved in response to drought. By using a yeast-one hybrid system, OsAP23 belonging to the B3 group of the ERF subfamily was isolated from rice (Zhuang et al. 2013). When exposed to high salt concentrations, several stress-responsive genes were induced significantly in the wild-type lines compared to Arabidopsis overexpressing 35S:OsAP23, suggesting a negative regulatory role of OsAP23 in salt stress response (Zhuang et al. 2013). Besides, characterization of an ERF gene from soybean, GmERF3, showed its inducible expression in soybean by high salinity, drought, ABA, salicylic acid (SA), jasmonic acid (JA), ethylene (ET), and soybean mosaic virus (SMV), whereas GmERF3 mRNA was not significantly accumulated under cold stress treatment (Zhang et al. 2009b). Transgenic tobacco plants with overexpressed 35S:GmERF3 displayed not only enhanced resistance against infection by Ralstonia solanacearum, Alternaria alternata, and tobacco mosaic virus (TMV) but also higher tolerance to high salinity and dehydration. Functional analyses of a tomato ERF (JERF3-jasmonate and ethylene responsive factor 3) in transgenic tobacco overexpressing 35S:JERF3 indicated that expression of this gene could enhance the tolerance to salt, drought, and freezing (Wang et al. 2004; Wu et al. 2008). Also in further report of this group, transgenic rice plants overexpressing 35S:JERF3 exhibited better tolerance to drought and osmotic stress in comparison with non-transgenic rice seedlings (Zhang et al. 2010).
3.2 The bZIP Family
The bZIP (basic leucine zipper) family is another large group of TFs in plants. At present, most reports on stress responses have shown that bZIP TFs regulate stress response in ABA-dependent manner through interaction with specific ABA-responsive cis-acting elements (ABRE) to promote transcription of downstream target genes (Kobayashi et al. 2005; Kim 2006; Zou et al. 2008; Shinozaki and Yamaguchi-Shinozaki 2007; Uno et al. 2000). In Arabidopsis, AtbZIP1-knockout mutants showed a decrease in salt and osmotic stress tolerance, suggesting its positive regulation of plant response to these stresses (Sun et al. 2012). Transgenic Arabidopsis overexpressing the maize ABP9 gene under the control of 35S promoter, which encodes a bZIP TF, enhanced tolerance to multiple stresses (Zhang et al. 2011b). Another bZIP from maize was identified as 35S:ZmbZIP72 whose overexpression improved drought and partial salt tolerance of transgenic Arabidopsis plants (Ying et al. 2012). Meanwhile, expression analysis of LrbZIP in tips of lotus roots (Nelumbo nucifera) showed strong upregulation by low temperature, salt, and ABA treatments (Cheng et al. 2013a). Transgenic tobacco transformed with 35S:LrbZIP exhibited higher salt tolerance comparing to the control under salt stress (Cheng et al. 2013a). Lu et al. (2009) identified OsbZIP72 as a positive regulator whose constitutive overexpression increased hypersensitivity to ABA and transcript level of ABA-responsive genes to improve drought tolerance in transgenic rice. Several soybean bZIPs, including GmbZIP44, GmbZIP62, and GmbZIP78, displayed notable roles in stress acclimation . These TFs functioned as negative regulators of ABA signaling and plant responses to salt and freezing tolerance (Liao et al. 2008b). Gao et al. (2011) also indicated the positive role of GmbZIP1 in the enhancement of multiple abiotic stress tolerance, including drought, salinity, and cold stresses in transgenic Arabidopsis by stimulating the expression of ABA- or stress-related genes. Overexpression of this gene by ubiquitin promoter also resulted in enhanced drought, salt, and freezing tolerance in transgenic wheat (Gao et al. 2011). The authors indicated that Arabidopsis and tobacco overexpressing GmbZIP1 by stress-inducible RD29A or 35S promoter also showed increased tolerance under similar stresses. ScAREB1, SpAREB1, and SlAREB1 belonging to the bZIP family of S. chilense, S. peruvianum, and S. lycopersicum, respectively, were upregulated by salt stress (Yáñez et al. 2009). Moreover, expression of SlAREB1 was induced by other stresses, such as drought and cold, and ABA in S. lycopersicum, and its encoded TF upregulated stress-responsive genes in 35S:SlAREB1-overexpressing transgenic tobacco and tomato plants (Yáñez et al. 2009).
3.3 The MYB Family
The MYB TFs have also been known to form one of the largest TF families, which interact with one or more of the two stress-inducible cis-elements known as MYB-binding sites (MBS) that contain consensus sequences CNGTT(A/G) (MBSI) or C(G/T)T(A/T)GTT(A/G) (MBSII) to activate their downstream genes (Abe et al. 1997; Pabo and Sauer 1992; Riechmann and Ratcliffe 2000). The MYB TFs possess a MYB domain containing 1–4 imperfect tandem repeats (MYB repeat) located near the N-terminus and thus showing their distinctive characteristics (Ambawat et al. 2013). Based on the number of adjacent repeat(s) in the MYB domain, the MYB family is divided into different types, including 4R-MYB (four repeats), 3R-MYB (R1R2R3-MYB) (three consecutive repeats), R2R3-MYB (two repeats), and the MYB-related type with just a single repeat (Rosinski and Atchley 1998; Jin and Martin 1999; Dubos et al. 2010). Members belonging to this family have been found in different plant species, such as 204 members in Arabidopsis, 218 members in rice, 279 members in grapevine, and 197 members in Populus (Wilkins et al. 2009; Velasco et al. 2007; Chen et al. 2006). The roles of MYB proteins have been indicated in many physiological and biochemical processes which include regulation of primary and secondary metabolism, control of cell development and cell cycle, hormone synthesis, and signal transduction. The MYBs are also involved in plant responses to various biotic and abiotic stresses (Dubos et al. 2010; Feller et al. 2011; Stracke et al. 2001). Mochida et al. (2009) also found approximately 160 GmMYBs in soybean, of which 43 out of 48 analyzed genes showed expression changes at least by one of the following treatments: ABA, high salinity, drought, and cold.
A number of studies have indicated the potential of MYB TFs in genetic engineering for improved stress tolerance. When the rice OsMYB3R-2 and the wheat TaPIMP1 genes were overexpressed in Arabidopsis and tobacco using 35S promoter, respectively, the transgenic lines displayed increased drought tolerance (Liu et al. 2011b; Dai et al. 2007). Besides, salt and freezing tolerance was elevated significantly in Arabidopsis overexpressing either 35S:GmMYB76 or 35S:GmMYB177 (Liao et al. 2008a). In addition, while GmMYB177 was upregulated by both drought and NaCl treatments, GmMYB76 was induced by NaCl treatment only. Jaradat et al. (2013) characterized the function of AtMYBR1 by using mybr1-mutant and 35S:AtMYBR1-overexpressing Arabidopsis lines. Its negative regulatory functions in drought response and senescence, as well as in the downregulation of many ABA-responsive genes involved in abiotic stresses, were revealed in their study. Moreover, expression of Arabidopsis AtMYB44 gene has been shown to improve salt and drought stress tolerance in soybean and Arabidopsis by preventing excessive ROS accumulation (Persak and Pitzschke 2014; Seo et al. 2012). Expression analysis of the OsMyb4 in rice suggested that this gene might be involved in rice response to dehydration and cold stress (Baldoni et al. 2013). Moreover, the authors found induction of OsMyb4-like genes in wheat and Arabidopsis under similar stress treatments. Overexpression of 35S:OsMyb4 in apple could improve adaptive responses to drought and cold stresses (Pasquali et al. 2008).
3.4 The WRKY Family
The WRKY TFs were first reported in the study of Ishiguro and Nakamura (1994) and the name WRKY (pronounced “worky”) was coined with the identification of WRKY1, WRKY2, and WRKY3 from parsley (Petroselinum crispum) (Rushton et al. 1996). The WRKY family is among the largest families of TFs in higher plants (Rushton et al. 2010). The WRKY domain is about 60 residues in length, and based on the number of WRKY domains, WRKY TFs were divided into three groups, group I (two domains), group II (one domain), and group III (one domain with structure of zinc fingers C2HC) (Eulgem et al. 2000). Till date, the functions of WRKY TFs have been intensively studied in not only biotic stress responses but also abiotic stress responses, as well as in seed germination, flower development, and senescence (Tripathi et al. 2014; Rushton et al. 2012; Thao and Tran 2012).
Overexpression of the heat- and drought-inducible rice OsWRKY11 gene mediated using the heat-inducible HSP101 promoter showed significant heat and drought tolerances in transgenic rice plants (Wu et al. 2009). Mochida et al. (2009) identified more than 210 putative WRKY TF-encoding genes in soybean. Under various abiotic stresses, 24 of 64 examined GmWRKY genes were found to be induced by drought (Zhou et al. 2008). Zhou et al. (2008) reported that 35S:GmWRKY21-overexpressing Arabidopsis plants exhibited improved tolerance to cold stress in comparison with wild type. The same authors also demonstrated that transgenic Arabidopsis plants overexpressing 35S:GmWRKY54 were more tolerant to drought and salt stress, whereas those overexpressing 35S:GmWRKY13 were more sensitive to salt stress (Zhou et al. 2008), suggesting these two WRKY TFs have opposite functions in plant responses to salt stress. Overexpression of the rice 35S:OsWRKY45 enhanced salt and drought tolerance of Arabidopsis transgenic plants in addition to increased disease resistance (Qiu and Yu 2009). The grapevine VvWRKY11 TF played a role in osmotic stress tolerance as improved tolerance of 35S:VvWRKY11-overexpressing transgenic Arabidopsis seedlings to mannitol-induced osmotic stress was observed in comparison with wild-type plants (Liu et al. 2011a). Recently, overexpression of the 35S:AtWRKY30 in Arabidopsis showed enhanced abiotic stress tolerance during early growth stages due to the binding of the TF to W-boxes in the promoter region of many stress/development-related genes, leading to the activation of their expression (Scarpeci et al. 2013). Results of RT-qPCR analyses showed that ZmWRKY33 of maize belonging to the group I subfamily was induced by high salt, dehydration, cold, and ABA treatments. Overexpression of this gene under control of 35S promoter in Arabidopsis activated stress-responsive genes, such as RD29A, under both normal growth and stress conditions, thereby improving tolerance of transgenic plants to salt stress (Li et al. 2013). Babitha et al. (2013) overexpressed 35S:AtWRKY28 in Arabidopsis and observed enhanced tolerance of transgenic plants to high NaCl, high mannitol, and oxidative stress. Additionally, higher root growth was observed in transgenic lines under mannitol-induced stress conditions. These transgenic plants showed their capacity in growth recovery to normal level after an 8-day drought exposure period followed by 6 days of rewatering.
3.5 The NAC Family
The plant-specific NAC TF family was initially described in Petunia and Arabidopsis more than 15 years ago (Aida et al. 1997; Souer et al. 1996). In plants, the NAC TFs have been reported to regulate diverse biological processes, such as flowering (Yoo et al. 2007), regulation of secondary cell wall synthesis and cell division (Zhong et al. 2007), embryo development (Duval et al. 2002), auxin signaling and lateral root formation (Xie et al. 2000), senescence (Kjaersgaard et al. 2011), as well as biotic and abiotic stress responses (Olsen et al. 2005; Puranik et al. 2012). The typical features of an NAC TF include an N-terminal conserved DNA-binding domain involving nucleus-oriented localization and a variable domain located at the C-terminal end which is essential for transcriptional activation (Fang et al. 2008; Riechmann and Ratcliffe 2000; Hao et al. 2011). Alignments of Arabidopsis and rice NAC domains suggested eight NAC subfamilies (from NAC-a to NAC-h), mainly distinguished by their unique structures in motif at the C-terminal of NAC domain (Fujita et al. 2004; Fang et al. 2008; Shen et al. 2009). Till date, NAC TFs have been systematically identified in many plant species thanks to the availability of their sequenced genomes. For instance, at least 117, 151, 163, 152, and 200 NAC genes have been identified in Arabidopsis, rice (Nuruzzaman et al. 2010), poplar (Hu et al. 2010), tobacco (Rushton et al. 2008), and soybean (Mochida et al. 2009), respectively. In drought signaling, NAC TFs were reported to function in both ABA-dependent and ABA-independent pathways (Shinozaki and Yamaguchi-Shinozaki 2007). The role of NACs in relation to drought response was initially proposed in a study of overexpression of either ANAC019, ANAC055, or ANAC072 in Arabidopsis which led to considerable increase in drought tolerance of transgenic plants (Tran et al. 2004). Thereafter, stress-related NAC genes have been detected in other plant species, such as rice (OsNAC5, OsNAC6, SNAC1, and ONAC45) (Puranik et al. 2012; Nakashima et al. 2007; Hu et al. 2006; Zheng et al. 2009; Song et al. 2011; Takasaki et al. 2010), wheat (TaNAC4, TaNAC69, and TaNAC2a) (Xue et al. 2011; Tang et al. 2012; Xia et al. 2010), oilseed rape (Brassica napus) (BnNAC2 and BnNAC5) (Zhong et al. 2012), and peanut (Arachis hypogaea) (AhNAC3) (Liu et al. 2013), which showed strong potential for genetic engineering of improved biotic and/or abiotic stress-tolerant crops.
Transgenic Arabidopsis displayed enhanced tolerance to drought stress without growth retardation when overexpressing the rice OsNAC52 using 35S promoter (Gao et al. 2010). In another independent study of rice, ONAC063 expression was highly induced in roots by high salinity as well as by high osmotic pressure and ROS levels (Yokotani et al. 2009). 35S:ONAC063-overexpressing transgenic Arabidopsis also displayed enhanced tolerance to high salinity and osmotic pressure (Yokotani et al. 2009). In a study of Lu et al. (2012), a maize NAC gene, ZmSNAC1, was cloned and functionally characterized. Low temperature, high salinity, drought stress, and ABA treatment strongly induced the expression of this gene. Overexpression of 35S:ZmSNAC1 in Arabidopsis resulted in hypersensitivity of transgenic plants to ABA and osmotic stress at the germination stage, but enhanced dehydration tolerance at the seedling stage as compared with non-transgenic control (Lu et al. 2012). Recently, a novel TF named JcNAC1 from the new model woody plant, Jatropha curcas, was reported to have function in responses to abiotic stresses and pathogen infection as overexpression of this gene under control of 35S promoter not only changed the expression of stress-related genes but also increased tolerance of transgenic J. curcas to drought (Qin et al. 2014). Another novel NAC TF, EcNAC from finger millet (Eleusine coracana), was overexpressed under control of either 35S promoter or synthetic 4xABRE stress-inducible promoter in tobacco, and both transgenic lines led to enhanced tolerance to various abiotic stresses, including stresses induced by polyethylene glycol (PEG) and mannitol, as well as high salinity (Ramegowda et al. 2012). In alfalfa (Medicago sativa), a NAC TF involved in response to abiotic stress was identified, and the expression of this gene was shown to be induced by drought, high salinity, and ABA (Wang 2013). Results revealed that transgenic Arabidopsis overexpressing this TF using 35S promoter had better drought tolerance than the wild type (Wang 2013). Baloglu et al. (2012) performed expression analysis of TaNAC69-1 and TtNAMB-2 in durum wheat (T. turgidum) under different abiotic stress conditions. Specifically, TaNAC69-1 was upregulated after 3 h of salt treatment, and the highest level of expression was observed at 24 and 48 h of post-treatment with heat and salinity, respectively. On the other hand, TtNAMB-2 was significantly induced by salt and low temperature stresses. In soybean, the first GmNAC genes identified were GmNAC1–6 in a study conducted by Meng et al. (2007). Subsequently, expression of these genes in response to various stress and hormone treatments, including ABA, JA, high salinity, and PEG-induced osmotic stress, was analyzed in detail by another group (Pinheiro et al. 2009). Later on, Tran et al. (2009) initiated a study of the GmNAC family at a wider scale, covering the expression analysis of 31 GmNAC genes at seedling stage and under different abiotic stress conditions, including dehydration, salinity, cold, and ABA treatment, as well as examination of their transcriptional activity. According to the results, nine genes were shown to be upregulated by at least one of the tested treatments. Except GmNAC028, all remaining genes (GmNAC002, 003, 004, 010, 012, 013, 015, and 020) also had transcriptional activation activity as shown by a yeast one-hybrid assay. More recently, Tran’s laboratory studied 152 GmNAC genes, which could be detected in soybean genome with full-length cDNA, and proposed a comprehensive nomenclature for the GmNAC members (Le et al. 2011). Furthermore, the authors reported that 31 genes displayed significantly altered expression upon dehydration, with 25 up- and 6 downregulated genes. Additionally, the same research group demonstrated the complexity in the dynamics of drought-responsive expression of the GmNAC genes as they indicated that expression of several GmNAC genes were tissue- and/or development stage-dependent (Le et al. 2012). More recently, Thao et al. (2013) found differential expression of a subset of drought-responsive GmNACs in soybean cultivars differing in drought tolerance, and identified positive correlation between GmNAC expression levels in these cultivars and their drought-tolerant degree. On the basis of their results, the authors also suggested a number of promising candidate GmNAC genes with potential application in genetic engineering of improved drought-tolerant soybean varieties (Thao et al. 2013).
4 Conclusions and Future Perspectives
Given the fact that using conventional breeding methods to create better stress-tolerant cultivars is not really effective since it is applied for varieties with close relationship only, creating transgenic plants by genetic technologies is a much more powerful approach. Thanks to advanced development in molecular cloning and plant transformation methods, barriers in gene transfer across species can be easily overcome. Therefore, the main challenge for scientists is to gain more and more in-depth understanding of mechanisms that plants employ to respond to stresses, especially to conditions similarly to the field environment. By doing this, important genes involved in plant adaptation to environmental stresses can be identified and used for crop improvement . Till date, many genes have been assigned as crucial contributors and uncountable attempts have been made to evaluate their roles in transgenic plants regarding the tolerance capacity toward abiotic stresses. The results of applications of TF-encoding genes so far have been quite promising. Accordingly, TFs bear a high potential for crop improvement using genetic engineering, and thus their characterization should deserve even more attention from the research community in the coming years.
References
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought-and abscisic acid-regulated gene expression. Plant Cell 9(10):1859–1868
Abogadallah GM, Nada RM, Malinowski R, Quick P (2011) Overexpression of HARDY, an AP2/ERF gene from Arabidopsis, improves drought and salt tolerance by reducing transpiration and sodium uptake in transgenic Trifolium alexandrinum L. Planta 233(6):1265–1276
Agarwal PK, Agarwal P, Reddy M, Sopory SK (2006) Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep 25(12):1263–1274
Agarwal P, Agarwal PK, Joshi AJ, Sopory SK, Reddy MK (2010) Overexpression of PgDREB2A transcription factor enhances abiotic stress tolerance and activates downstream stress-responsive genes. Mol Biol Rep 37(2):1125–1135
Aguado-Santacruz GA (2006) Genetic manipulation of plants for increased drought tolerance. Adv Agr Food Biotechnol 2006:71–98
Aida M, Ishida T, Fukaki H, Fujisawa H, Tasaka M (1997) Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9(6):841–857
Alsina MM, Smart DR, Bauerle T, De Herralde F, Biel C, Stockert C, Negron C, Save R (2011) Seasonal changes of whole root system conductance by a drought-tolerant grape root system. J Exp Bot 62(1):99–109
Ambawat S, Sharma P, Yadav NR, Yadav RC (2013) MYB transcription factor genes as regulators for plant responses: an overview. Physiol Mol Biol Plants 19(3):307–321
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Aroca R, Porcel R, Ruiz-Lozano JM (2012) Regulation of root water uptake under abiotic stress conditions. J Exp Bot 63(1):43–57
Ashraf M, Harris P (2004) Potential biochemical indicators of salinity tolerance in plants. Plant Sci 166(1):3–16
Babitha K, Ramu S, Pruthvi V, Mahesh P, Nataraja KN, Udayakumar M (2013) Co-expression of AtbHLH17 and AtWRKY28 confers resistance to abiotic stress in Arabidopsis. Transgenic Res 22(2):327–341
Baldoni E, Genga A, Medici A, Coraggio I, Locatelli F (2013) The OsMyb4 gene family: stress response and transcriptional auto-regulation mechanisms. Biol Plant 57(4):691–700
Baloglu MC, Oz MT, Oktem HA, Yucel M (2012) Expression analysis of TaNAC69-1 and TtNAMB-2, wheat NAC family transcription factor genes under abiotic stress conditions in durum wheat (Triticum turgidum). Plant Mol Biol Rep 30(5):1246–1252
Bhargava S, Sawant K (2013) Drought stress adaptation: metabolic adjustment and regulation of gene expression. Plant Breed 132(1):21–32
Bouaziz D, Pirrello J, Charfeddine M, Hammami A, Jbir R, Dhieb A, Bouzayen M, Gargouri-Bouzid R (2013) Overexpression of StDREB1 transcription factor increases tolerance to salt in transgenic potato plants. Mol Biotechnol 54(3):803–817
Boudsocq M, Laurière C (2005) Osmotic signaling in plants. Multiple pathways mediated by emerging kinase families. Plant Physiol 138(3):1185–1194
Boudsocq M, Sheen J (2013) CDPKs in immune and stress signaling. Trends Plant Sci 18(1):30–40
Calvo-Polanco M, Señorans J, Zwiazek JJ (2012) Role of adventitious roots in water relations of tamarack (Larix laricina) seedlings exposed to flooding. BMC Plant Biol 12(1):99–108
Chakrabortee S, Meersman F, Schierle GSK, Bertoncini CW, McGee B, Kaminski CF, Tunnacliffe A (2010) Catalytic and chaperone-like functions in an intrinsically disordered protein associated with desiccation tolerance. Proc Natl Acad Sci U S A 107(37):16084–16089
Chaves MM, Maroco JP, Pereira JS (2003) Understanding plant responses to drought-from genes to the whole plant. Funct Plant Biol 30(3):239–264
Chaves M, Flexas J, Pinheiro C (2009) Photosynthesis under drought and salt stress: regulation mechanisms from whole plant to cell. Ann Bot 103(4):551–560
Chen Y, Yang X, He K, Liu M, Li J, Gao Z, Lin Z, Zhang Y, Wang X, Qiu X (2006) The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60(1):107–124
Chen M, Wang Q-Y, Cheng X-G, Xu Z-S, Li L-C, Ye X-G, Xia L-Q, Ma Y-Z (2007) GmDREB2, a soybean DRE-binding transcription factor, conferred drought and high-salt tolerance in transgenic plants. Biochem Biophys Res Commun 353(2):299–305
Chen L, Song Y, Li S, Zhang L, Zou C, Yu D (2012) The role of WRKY transcription factors in plant abiotic stresses. Biochim Biophys Acta 1819(2):120–128
Chen Y, Chen Z, Kang J, Kang D, Gu H, Qin G (2013a) AtMYB14 regulates cold tolerance in Arabidopsis. Plant Mol Biol Rep 31(1):87–97
Chen Y, Yang J, Wang Z, Zhang H, Mao X, Li C (2013b) Gene structures, classification, and expression models of the DREB transcription factor subfamily in Populus trichocarpa. Scientific World Journal 2013(2013):1–12
Chen X, Wang Y, Lv B, Li J, Luo L, Lu S, Zhang X, Ma H, Ming F (2014) The NAC family transcription factor OsNAP confers abiotic stress response through the ABA pathway. Plant Cell Physiol. 55(3):604-619
Cheng L, Li S, Hussain J, Xu X, Yin J, Zhang Y, Chen X, Li L (2013a) Isolation and functional characterization of a salt responsive transcriptional factor, LrbZIP from lotus root (Nelumbo nucifera Gaertn). Mol Biol Rep 40(6):4033–4045
Cheng L, Li X, Huang X, Ma T, Liang Y, Ma X, Peng X, Jia J, Chen S, Chen Y (2013b) Overexpression of sheepgrass R1-MYB transcription factor LcMYB1 confers salt tolerance in transgenic Arabidopsis. Plant Physiol Biochem 70:252–260
Chinnusamy V, Schumaker K, Zhu J-K (2004) Molecular genetic perspectives on cross-talk and specificity in abiotic stress signalling in plants. J Exp Bot 55(395):225–236
Clemens S (2006) Toxic metal accumulation, responses to exposure and mechanisms of tolerance in plants. Biochimie 88(11):1707–1719
Dai X, Xu Y, Ma Q, Xu W, Wang T, Xue Y, Chong K (2007) Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol 143(4):1739–1751
Dalal M, Tayal D, Chinnusamy V, Bansal KC (2009) Abiotic stress and ABA-inducible Group 4 LEA from Brassica napus plays a key role in salt and drought tolerance. J Biotechnol 139(2):137–145
Danquah A, de Zelicourt A, Colcombet J, Hirt H (2014) The role of ABA and MAPK signaling pathways in plant abiotic stress responses. Biotechnol Adv 32(1):40–52
Ding Z, Li S, An X, Liu X, Qin H, Wang D (2009) Transgenic expression of MYB15 confers enhanced sensitivity to abscisic acid and improved drought tolerance in Arabidopsis thaliana. J Genet Genomics 36(1):17–29
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15(10):573–581
Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi‐Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33(4):751–763
Duval M, Hsieh T-F, Kim SY, Thomas TL (2002) Molecular characterization of AtNAM: a member of the Arabidopsis NAC domain superfamily. Plant Mol Biol 50(2):237–248
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5(5):199–206
Fang Y, You J, Xie K, Xie W, Xiong L (2008) Systematic sequence analysis and identification of tissue-specific or stress-responsive genes of NAC transcription factor family in rice. Mol Genet Genomics 280(6):547–563
Feller A, Machemer K, Braun EL, Grotewold E (2011) Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J 66(1):94–116
Fischer U, Dröge-Laser W (2004) Overexpression of NtERF5, a new member of the tobacco ethylene response transcription factor family enhances resistance to tobacco mosaic virus. Mol Plant Microbe Interact 17(10):1162–1171
Fujimoto SY, Ohta M, Usui A, Shinshi H, Ohme-Takagi M (2000) Arabidopsis ethylene-responsive element binding factors act as transcriptional activators or repressors of GCC box-mediated gene expression. Plant Cell 12(3):393–404
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran L-SP, Yamaguchi-Shinozaki K, Shinozaki K (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39(6):863–876
Fujita Y, Fujita M, Shinozaki K, Yamaguchi-Shinozaki K (2011) ABA-mediated transcriptional regulation in response to osmotic stress in plants. J Plant Res 124(4):509–525
Gao F, Xiong A, Peng R, Jin X, Xu J, Zhu B, Chen J, Yao Q (2010) OsNAC52, a rice NAC transcription factor, potentially responds to ABA and confers drought tolerance in transgenic plants. Plant Cell Tiss Org 100(3):255–262
Gao S-Q, Chen M, Xu Z-S, Zhao C-P, Li L, Xu H-j, Tang Y-m, Zhao X, Ma Y-Z (2011) The soybean GmbZIP1 transcription factor enhances multiple abiotic stress tolerances in transgenic plants. Plant Mol Biol 75(6):537–553
Gong Y, Rao L, Yu D (2013) Abiotic stress in plants. In: Stoytcheva M, Zlatev R (eds) Agricultural chemistry. InTech, Rijeka. doi:10.5772/55163
Gowda VR, Henry A, Yamauchi A, Shashidhar H, Serraj R (2011) Root biology and genetic improvement for drought avoidance in rice. Field Crop Res 122(1):1–13
Goyal K, Walton L, Tunnacliffe A (2005) LEA proteins prevent protein aggregation due to water stress. Biochem J 388:151–157
Grassi G, Magnani F (2005) Stomatal, mesophyll conductance and biochemical limitations to photosynthesis as affected by drought and leaf ontogeny in ash and oak trees. Plant Cell Environ 28(7):834–849
Ha CV, Le DT, Nishiyama R, Watanabe Y, Tran UT, Dong NV, Tran L-SP (2013) Characterization of the newly developed soybean cultivar DT2008 in relation to the model variety W82 reveals a new genetic resource for comparative and functional genomics for improved drought tolerance. Biomed Res Int 2013:1–8
Hachiya T, Sugiura D, Kojima M, Sato S, Yanagisawa S, Sakakibara H, Terashima I, Noguchi K (2014) High CO2 triggers preferential root growth of Arabidopsis thaliana via two distinct systems at low pH and low N stresses. Plant Cell Physiol 55(2):269–280
Hao D, Ohme-Takagi M, Sarai A (1998) Unique mode of GCC box recognition by the DNA-binding domain of ethylene-responsive element-binding factor (ERF domain) in plant. J Biol Chem 273(41):26857–26861
Hao YJ, Wei W, Song QX, Chen HW, Zhang YQ, Wang F, Zou HF, Lei G, Tian AG, Zhang WK (2011) Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants. Plant J 68(2):302–313
Hasanuzzaman M, Nahar K, Alam MM, Roychowdhury R, Fujita M (2013) Physiological, biochemical, and molecular mechanisms of heat stress tolerance in plants. Int J Mol Sci 14(5):9643–9684
Hideg É, Jansen MA, Strid Å (2013) UV-B exposure, ROS, and stress: inseparable companions or loosely linked associates? Trends Plant Sci 18(2):107–115
Hossain MA, Piyatida P, da Silva JAT, Fujita M (2012) Molecular mechanism of heavy metal toxicity and tolerance in plants: central role of glutathione in detoxification of reactive oxygen species and methylglyoxal and in heavy metal chelation. J Bot 2012(2012):1-37
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci U S A 103(35):12987–12992
Hu R, Qi G, Kong Y, Kong D, Gao Q, Zhou G (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 10(1):145
Huang X-S, Liu J-H, Chen X-J (2010) Overexpression of PtrABF gene, a bZIP transcription factor isolated from Poncirus trifoliata, enhances dehydration and drought tolerance in tobacco via scavenging ROS and modulating expression of stress-responsive genes. BMC Plant Biol 10(1):230
Huang G-T, Ma S-L, Bai L-P, Zhang L, Ma H, Jia P, Liu J, Zhong M, Guo Z-F (2012) Signal transduction during cold, salt, and drought stresses in plants. Mol Biol Rep 39(2):969–987
Ishiguro S, Nakamura K (1994) Characterization of a cDNA encoding a novel DNA-binding protein, SPF1, that recognizes SP8 sequences in the 5′ upstream regions of genes coding for sporamin and β-amylase from sweet potato. Mol Gen Genet 244(6):563–571
Islam M, Wang M (2012) Expression patterns of an abiotic stress-inducible dehydration responsive element binding protein-2 (DREB2) gene in tomato. Bangladesh J Plant Breed Genet 25(1):01–09
Ito Y, Katsura K, Maruyama K, Taji T, Kobayashi M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006) Functional analysis of rice DREB1/CBF-type transcription factors involved in cold-responsive gene expression in transgenic rice. Plant Cell Physiol 47(1):141–153
Jaarsma R, de Vries RS, de Boer AH (2013) Effect of salt stress on growth, Na + accumulation and proline metabolism in potato (Solanum tuberosum) cultivars. PloS ONE 8(3):e60183
Jaglo-Ottosen KR, Gilmour SJ, Zarka DG, Schabenberger O, Thomashow MF (1998) Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280(5360):104–106
Jaglo KR, Kleff S, Amundsen KL, Zhang X, Haake V, Zhang JZ, Deits T, Thomashow MF (2001) Components of the Arabidopsis C-repeat/dehydration-responsive element binding factor cold-response pathway are conserved inbrassica napus and other plant species. Plant Physiol 127(3):910–917
Jaradat MR, Feurtado JA, Huang D, Lu Y, Cutler AJ (2013) Multiple roles of the transcription factor AtMYBR1/AtMYB44 in ABA signaling, stress responses, and leaf senescence. BMC Plant Biol 13(1):192
Jin H, Martin C (1999) Multifunctionality and diversity within the plant MYB-gene family. Plant Mol Biol 41(5):577–585
Joo J, Choi HJ, Lee YH, Kim Y-K, Song SI (2013) A transcriptional repressor of the ERF family confers drought tolerance to rice and regulates genes preferentially located on chromosome 11. Planta 238(1):155–170
Jovanovic Z, Stanisavljevic N, Mikic A, Radovic S, Maksimovic V (2013) The expression of drought responsive element binding protein (DREB2 A) related gene from pea (Pisum sativum L.) as affected by water stress. Aust J Crop Sci 7(10):1590–1596
Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1999) Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17(3):287–291
Kim SY (2006) The role of ABF family bZIP class transcription factors in stress response. Physiol Plant 126(4):519–527
Kjaersgaard T, Jensen MK, Christiansen MW, Gregersen P, Kragelund BB, Skriver K (2011) Senescence-associated barley NAC (NAM, ATAF1, 2, CUC) transcription factor interacts with radical-induced cell death 1 through a disordered regulatory domain. J Biol Chem 286(41):35418–35429
Kobayashi Y, Murata M, Minami H, Yamamoto S, Kagaya Y, Hobo T, Yamamoto A, Hattori T (2005) Abscisic acid-activated SNRK2 protein kinases function in the gene-regulation pathway of ABA signal transduction by phosphorylating ABA response element-binding factors. Plant J 44(6):939–949
Koyro H-W, Ahmad P, Geissler N (2012) Abiotic stress responses in plants: an overview. In: Ahmad P, Prasad MNV (eds) Environmental adaptations and stress tolerance of plants in the era of climate change. Springer, New York, pp 1–28
Kreuzwieser J, Rennenberg H (2014) Molecular and physiological responses of trees to waterlogging stress. Plant Cell Environ. doi:10.1111/pce.12310
Ku Y-S, Au-Yeung W-K, Yung Y-L, Li M-W, Wen C-Q, Liu X, Lam H-M (2013) Drought stress and tolerance in soybean. In: Board JE (ed) A comprehensive survey of internaitonal soybean research—genetics, physiology, agronomy and nitrogen relationships. InTech, New York, pp 209–237
Kumar N, Nandwal AS, Waldia RS, Singh S, Devi S, Sharma KD, Kumar A (2012) Drought tolerance in chickpea as evaluated by root characteristics, plant water status, membrane integrity and chlorophyll fluorescence techniques. Exp Agric 48(03):378–387
Kumar MN, Jane W-N, Verslues PE (2013) Role of the putative osmosensor arabidopsis histidine kinase1 in dehydration avoidance and low-water-potential response. Plant Physiol 161(2):942–953
Lawlor D, Cornic G (2002) Photosynthetic carbon assimilation and associated metabolism in relation to water deficits in higher plants. Plant Cell Environ 25(2):275–294
Lawlor DW, Tezara W (2009) Causes of decreased photosynthetic rate and metabolic capacity in water-deficient leaf cells: a critical evaluation of mechanisms and integration of processes. Ann Bot 103(4):561–579
Le DT, Nishiyama R, Watanabe Y, Mochida K, Yamaguchi-Shinozaki K, Shinozaki K, Tran LSP (2011) Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress. DNA Res 18(4):263–276
Le DT, Nishiyama R, Watanabe Y, Tanaka M, Seki M, Yamaguchi-Shinozaki K, Shinozaki K, Tran L-SP (2012) Differential gene expression in soybean leaf tissues at late developmental stages under drought stress revealed by genome-wide transcriptome analysis. PloS ONE 7(11):e49522
Li H, Gao Y, Xu H, Dai Y, Deng D, Chen J (2013) ZmWRKY33, a WRKY maize transcription factor conferring enhanced salt stress tolerances in Arabidopsis. Plant Growth Regul 70(3):207–216
Liang C, Tian J, Liao H (2013) Proteomics dissection of plant responses to mineral nutrient deficiency. Proteomics 13(3-4):624–636
Liao Y, Zou H-F, Wang H-W, Zhang W-K, Ma B, Zhang J-S, Chen S-Y (2008a) Soybean GmMYB76, GmMYB92, and GmMYB177 genes confer stress tolerance in transgenic Arabidopsis plants. Cell Res 18(10):1047–1060
Liao Y, Zou H-F, Wei W, Hao Y-J, Tian A-G, Huang J, Liu Y-F, Zhang J-S, Chen S-Y (2008b) Soybean GmbZIP44, GmbZIP62 and GmbZIP78 genes function as negative regulator of ABA signaling and confer salt and freezing tolerance in transgenic Arabidopsis. Planta 228(2):225–240
Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought-and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 10(8):1391–1406
Liu F, Liu Q, Liang X, Huang H, Zhang S (2005) Morphological, anatomical, and physiological assessment of ramie [Boehmeria nivea (L.) Gaud.] tolerance to soil drought. Genet Resour Crop Evol 52(5):497–506
Liu H, Yang W, Liu D, Han Y, Zhang A, Li S (2011a) Ectopic expression of a grapevine transcription factor VvWRKY11 contributes to osmotic stress tolerance in Arabidopsis. Mol Biol Rep 38(1):417–427
Liu H, Zhou X, Dong N, Liu X, Zhang H, Zhang Z (2011b) Expression of a wheat MYB gene in transgenic tobacco enhances resistance to Ralstonia solanacearum, and to drought and salt stresses. Funct Integr Genomics 11(3):431–443
Liu X, Liu S, Wu J, Zhang B, Li X, Yan Y, Li L (2013) Overexpression of Arachis hypogaea NAC3 in tobacco enhances dehydration and drought tolerance by increasing superoxide scavenging. Plant Physiol Biochem 70:354–359
Liu C, Mao B, Ou S, Wang W, Liu L, Wu Y, Chu C, Wang X (2014a) OsbZIP71, a bZIP transcription factor, confers salinity and drought tolerance in rice. Plant Mol Biol 84(1-2):19–36
Liu G, Li X, Jin S, Liu X, Zhu L, Nie Y, Zhang X (2014b) Overexpression of rice NAC gene SNAC1 improves drought and salt tolerance by enhancing root development and reducing transpiration rate in transgenic cotton. PloS ONE 9(1):e86895
Loutfy N, El-Tayeb MA, Hassanen AM, Moustafa MF, Sakuma Y, Inouhe M (2012) Changes in the water status and osmotic solute contents in response to drought and salicylic acid treatments in four different cultivars of wheat (Triticum aestivum). J Plant Res 125(1):173–184
Lu G, Gao C, Zheng X, Han B (2009) Identification of OsbZIP72 as a positive regulator of ABA response and drought tolerance in rice. Planta 229(3):605–615
Lu M, Ying S, Zhang D-F, Shi Y-S, Song Y-C, Wang T-Y, Li Y (2012) A maize stress-responsive NAC transcription factor, ZmSNAC1, confers enhanced tolerance to dehydration in transgenic Arabidopsis. Plant Cell Rep 31(9):1701–1711
Mahajan S, Tuteja N (2005) Cold, salinity and drought stresses: an overview. Arch Biochem Biophys 444(2):139–158
Mallikarjuna G, Mallikarjuna K, Reddy M, Kaul T (2011) Expression of OsDREB2A transcription factor confers enhanced dehydration and salt stress tolerance in rice (Oryza sativa L.). Biotechnol Lett 33(8):1689–1697
Manavalan LP, Guttikonda SK, Tran LSP, Nguyen HT (2009) Physiological and molecular approaches to improve drought resistance in soybean. Plant Cell Physiol 50(7):1260–1276
Mao X, Chen S, Li A, Zhai C, Jing R (2014) Novel NAC transcription factor TaNAC67 confers enhanced multi abiotic stress tolerances in Arabidopsis. PloS ONE 9(1):e84359
Meng Q, Zhang C, Gai J, Yu D (2007) Molecular cloning, sequence characterization and tissue-specific expression of six NAC-like genes in soybean (Glycine max (L.) Merr.). J Plant Physiol 164(8):1002–1012
Miller G, Suzuki N, Ciftci-yilmaz S, Mittler R (2010) Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ 33(4):453–467
Misra S, Wu Y, Venkataraman G, Sopory SK, Tuteja N (2007) Heterotrimeric G-protein complex and G-protein-coupled receptor from a legume (Pisum sativum): role in salinity and heat stress and cross-talk with phospholipase C. Plant J 51(4):656–669
Mittler R (2002) Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci 7(9):405–410
Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) AP2/ERF family transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819(2):86–96
Mochida K, Yoshida T, Sakurai T, Yamaguchi-Shinozaki K, Shinozaki K, Tran LSP (2009) In silico analysis of transcription factor repertoire and prediction of stress responsive transcription factors in soybean. DNA Res 16(6):353–369
Mori IC, Schroeder JI (2004) Reactive oxygen species activation of plant Ca2 + channels. A signaling mechanism in polar growth, hormone transduction, stress signaling, and hypothetically mechanotransduction. Plant Physiol 135(2):702–708
Munns R, James RA, Läuchli A (2006) Approaches to increasing the salt tolerance of wheat and other cereals. J Exp Bot 57(5):1025–1043
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Nakashima K, Tran LSP, Van Nguyen D, Fujita M, Maruyama K, Todaka D, Ito Y, Hayashi N, Shinozaki K, Yamaguchi‐Shinozaki K (2007) Functional analysis of a NAC‐type transcription factor OsNAC6 involved in abiotic and biotic stress‐responsive gene expression in rice. Plant J 51(4):617–630
Nakashima K, Ito Y, Yamaguchi-Shinozaki K (2009) Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol 149(1):88–95
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819(2):97–103
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465(1):30–44
Ohme-Takagi M, Shinshi H (1995) Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 7(2):173–182
Ohnishi T, Nakazono M, Tsutsumi N (2008) Abiotic stress. In: Hirano H-Y, Sano Y, Hirai A, Sasaki T (eds) Rice biology in the genomics era. Springer, Berlin, pp 337–355
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10(2):79–87
Oñate-Sánchez L, Singh KB (2002) Identification of Arabidopsis ethylene-responsive element binding factors with distinct induction kinetics after pathogen infection. Plant Physiol 128(4):1313–1322
Osakabe Y, Yamaguchi-Shinozaki K, Shinozaki K, Tran L-SP (2013a) ABA control of plant macroelement membrane transport systems in response to water deficit and high salinity. New Phytol 202(1):35–49
Osakabe Y, Yamaguchi-Shinozaki K, Shinozaki K, Tran L-SP (2013b) Sensing the environment: key roles of membrane-localized kinases in plant perception and response to abiotic stress. J Exp Bot 64(2):445–458
Osakabe Y, Osakabe K, Shinozaki K, Tran L-SP (2014) Response of plants to water stress. Front Plant Sci 5. doi:10.3389/fpls.2014.00086
Pabo CO, Sauer RT (1992) Transcription factors: structural families and principles of DNA recognition. Annu Rev Biochem 61(1):1053–1095
Pandey S, Nelson DC, Assmann SM (2009) Two novel GPCR-type G proteins are abscisic acid receptors in Arabidopsis. Cell 136(1):136–148
Park JM, Park C-J, Lee S-B, Ham B-K, Shin R, Paek K-H (2001) Overexpression of the tobacco Tsi1 gene encoding an EREBP/AP2-type transcription factor enhances resistance against pathogen attack and osmotic stress in tobacco. Plant Cell 13(5):1035–1046
Parvaiz A, Satyawati S (2008) Salt stress and phyto-biochemical responses of plants-a review. Plant Soil Environ 54(3):89–99
Pasquali G, Biricolti S, Locatelli F, Baldoni E, Mattana M (2008) Osmyb4 expression improves adaptive responses to drought and cold stress in transgenic apples. Plant Cell Rep 27(10):1677–1686
Peng X, Ma X, Fan W, Su M, Cheng L, Iftekhar A, Lee B-H, Qi D, Shen S, Liu G (2011) Improved drought and salt tolerance of Arabidopsis thaliana by transgenic expression of a novel DREB gene from Leymus chinensis. Plant Cell Rep 30(8):1493–1502
Peng X, Zhang L, Zhang L, Liu Z, Cheng L, Yang Y, Shen S, Chen S, Liu G (2013) The transcriptional factor LcDREB2 cooperates with LcSAMDC2 to contribute to salt tolerance in Leymus chinensis. Plant Cell Tissue Org 113(2):245–256
Perfus-Barbeoch L, Jones AM, Assmann SM (2004) Plant heterotrimeric G protein function: insights from Arabidopsis and rice mutants. Curr Opin Plant Biol 7(6):719–731
Persak H, Pitzschke A (2014) Dominant repression by Arabidopsis transcription factor MYB44 causes oxidative damage and hypersensitivity to abiotic stress. Int J Mol Sci 15(2):2517–2537
Pinheiro GL, Marques CS, Costa MDBL, Reis PAB, Alves MS, Carvalho CM, Fietto LG, Fontes EPB (2009) Complete inventory of soybean NAC transcription factors: sequence conservation and expression analysis uncover their distinct roles in stress response. Gene 444(1):10–23
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17(6):369–381
Qin Y, Tian Y, Han L, Yang X (2013) Constitutive expression of a salinity-induced wheat WRKY transcription factor enhances salinity and ionic stress tolerance in transgenic Arabidopsis thaliana. Biochem Biophys Res Commun 441(2):476–481
Qin X, Zheng X, Huang X, Lii Y, Shao C, Xu Y, Chen F (2014) A novel transcription factor JcNAC1 response to stress in new model woody plant Jatropha curcas. Planta 239(2):511–520
Qiu Y, Yu D (2009) Over-expression of the stress-induced OsWRKY45 enhances disease resistance and drought tolerance in Arabidopsis. Environ Exp Bot 65(1):35–47
Quan R, Hu S, Zhang Z, Zhang H, Zhang Z, Huang R (2010) Overexpression of an ERF transcription factor TSRF1 improves rice drought tolerance. Plant Biotechnol J 8(4):476–488
Ramegowda V, Senthil-Kumar M, Nataraja KN, Reddy MK, Mysore KS, Udayakumar M (2012) Expression of a finger millet transcription factor, EcNAC1, in tobacco confers abiotic stress-tolerance. PloS ONE 7(7):e40397
Ravikumar G, Manimaran P, Voleti S, Subrahmanyam D, Sundaram R, Bansal K, Viraktamath B, Balachandran S (2014) Stress-inducible expression of AtDREB1A transcription factor greatly improves drought stress tolerance in transgenic indica rice. Transgenic Res:1–19
Riechmann JL, Meyerowitz EM (1998) The AP2/EREBP family of plant transcription factors. Biol Chem 379:633–646
Riechmann JL, Ratcliffe OJ (2000) A genomic perspective on plant transcription factors. Curr Opin Plant Biol 3(5):423–434
Rong W, Qi L, Wang A, Ye X, Du L, Liang H, Xin Z, Zhang Z (2014) The ERF transcription factor TaERF3 promotes tolerance to salt and drought stresses in wheat. Plant Biotechnol J. 12(4): 468–479
Rosinski JA, Atchley WR (1998) Molecular evolution of the Myb family of transcription factors: evidence for polyphyletic origin. J Mol Evol 46(1):74–83
Rushton PJ, Torres JT, Parniske M, Wernert P, Hahlbrock K, Somssich I (1996) Interaction of elicitor-induced DNA-binding proteins with elicitor response elements in the promoters of parsley PR1 genes. EMBO J 15(20):5690
Rushton PJ, Bokowiec MT, Han S, Zhang H, Brannock JF, Chen X, Laudeman TW, Timko MP (2008) Tobacco transcription factors: novel insights into transcriptional regulation in the Solanaceae. Plant Physiol 147(1):280–295
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends Plant Sci 15(5):247–258
Rushton DL, Tripathi P, Rabara RC, Lin J, Ringler P, Boken AK, Langum TJ, Smidt L, Boomsma DD, Emme NJ (2012) WRKY transcription factors: key components in abscisic acid signalling. Plant Biotechnol J 10(1):2–11
Saad ASI, Li X, Li H-P, Huang T, Gao C-S, Guo M-W, Cheng W, Zhao G-Y, Liao Y-C (2013) A rice stress-responsive NAC gene enhances tolerance of transgenic wheat to drought and salt stresses. Plant Sci 203:33–40
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration-and cold-inducible gene expression. Biochem Biophys Res Commun 290(3):998–1009
Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006) Functional analysis of an Arabidopsis transcription factor, DREB2A involved in drought-responsive gene expression. Plant Cell 18(5):1292–1309
Scarpeci TE, Zanor MI, Mueller-Roeber B, Valle EM (2013) Overexpression of AtWRKY30 enhances abiotic stress tolerance during early growth stages in Arabidopsis thaliana. Plant Mol Biol 83(3):265–277
Schaller GE, Kieber JJ, Shiu S-H (2008) Two-component signaling elements and histidyl-aspartyl phosphorelays. In: The Arabidopsis book, vol 6. p e0112
Seo JS, Sohn HB, Noh K, Jung C, An JH, Donovan CM, Somers DA, Kim DI, Jeong S-C, Kim C-G (2012) Expression of the Arabidopsis AtMYB44 gene confers drought/salt-stress tolerance in transgenic soybean. Mol Breed 29(3):601–608
Setia R, Lewis M, Marschner P, Raja Segaran R, Summers D, Chittleborough D (2011) Severity of salinity accurately detected and classified on a paddock scale with high resolution multispectral satellite imagery. Land Degrad Dev 24(4):375–384
Shabala S, Munns R (2012) Salinity stress: physiological constraints and adaptive mechanisms. In: Shabala S (ed) Plant stress physiology. CAB International, Oxford, pp 59–93
Shanker A, Venkateswarlu B (2011) Abiotic stress response in plants-physiological, biochemical and genetic perspectives. InTech, Rijeka. doi:10.5772/1762
Sharma KK, Lavanya M (2002) Recent developments in transgenics for abiotic stress in legumes of the semi-arid tropics. JIRCAS Working Report 23:61–73
Sharma P, Jha AB, Dubey RS, Pessarakli M (2012) Reactive oxygen species, oxidative damage, and antioxidative defense mechanism in plants under stressful conditions. J Bot 2012 (2012):217037
Sharoni AM, Nuruzzaman M, Satoh K, Shimizu T, Kondoh H, Sasaya T, Choi I-R, Omura T, Kikuchi S (2011) Gene structures, classification and expression models of the AP2/EREBP transcription factor family in rice. Plant Cell Physiol 52(2):344–360
Shekhawat UKS, Ganapathi TR, Srinivas L (2011) Cloning and characterization of a novel stress-responsive WRKY transcription factor gene (MusaWRKY71) from Musa spp. cv. Karibale Monthan (ABB group) using transformed banana cells. Mol Biol Rep 38(6):4023–4035
Shen H, Yin Y, Chen F, Xu Y, Dixon RA (2009) A bioinformatic analysis of NAC genes for plant cell wall development in relation to lignocellulosic bioenergy production. Bioenerg Res 2(4):217–232
Shi W, Liu D, Hao L, Wu C-a, Guo X, Li H (2014) GhWRKY39, a member of the WRKY transcription factor family in cotton, has a positive role in disease resistance and salt stress tolerance. Plant Cell Tiss Org 1–16
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58(2):221–227
Solanke AU, Sharma AK (2008) Signal transduction during cold stress in plants. Physiol Mol Biol Plants 14(1-2):69–79
Song S-Y, Chen Y, Chen J, Dai X-Y, Zhang W-H (2011) Physiological mechanisms underlying OsNAC5-dependent tolerance of rice plants to abiotic stress. Planta 234(2):331–345
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85(2):159–170
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol 4(5):447–456
Su L, Li J, Liu D, Zhai Y, Zhang H, Li X, Zhang Q, Wang Y, Wang Q (2014) A novel MYB transcription factor, GmMYBJ1, from soybean confers drought and cold tolerance in Arabidopsis thaliana. Gene. doi:10.1016/j.gene.2014.01.024
Sun X, Li Y, Cai H, Bai X, Ji W, Ding X, Zhu Y (2012) The Arabidopsis AtbZIP1 transcription factor is a positive regulator of plant tolerance to salt, osmotic and drought stresses. J Plant Res 125(3):429–438
Suzuki N, Koussevitzky S, Mittler R, Miller G (2012) ROS and redox signalling in the response of plants to abiotic stress. Plant Cell Environ 35(2):259–270
Takasaki H, Maruyama K, Kidokoro S, Ito Y, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K, Nakashima K (2010) The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol Genet Genomics 284(3):173–183
Tang Y, Liu M, Gao S, Zhang Z, Zhao X, Zhao C, Zhang F, Chen X (2012) Molecular characterization of novel TaNAC genes in wheat and overexpression of TaNAC2a confers drought tolerance in tobacco. Physiol Plant 144(3):210–224
Thao NP, Tran L-SP (2012) Potentials toward genetic engineering of drought-tolerant soybean. Crit Rev Biotechnol 32(4):349–362
Thao NP, Thu NBA, Hoang XLT, Ha VC, Tran LSP (2013) Differential expression analysis of a subset of drought-responsive GmNAC genes in two soybean cultivars differing in drought tolerance. Int J Mol Sci 14(12):23828–23841
Thapa G, Dey M, Sahoo L, Panda S (2011) An insight into the drought stress induced alterations in plants. Biol Plant 55(4):603–613
Theocharis A, Clément C, Barka EA (2012) Physiological and molecular changes in plants grown at low temperatures. Planta 235(6):1091–1105
Thu NBA, Nguyen QT, Hoang XLT, Thao NP, Tran LSP (2014) Evaluation of drought tolerance of the Vietnamese soybean cultivars provides potential resources for soybean production and genetic engineering. Biomed Res Int 2014(2014):809736.
Tournier B, Sanchez-Ballesta MT, Jones B, Pesquet E, Regad F, Latché A, Pech J-C, Bouzayen M (2003) New members of the tomato ERF family show specific expression pattern and diverse DNA-binding capacity to the GCC box element. FEBS Lett 550(1):149–154
Tran L-SP, Nakashima K, Sakuma Y, Simpson SD, Fujita Y, Maruyama K, Fujita M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2004) Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 16(9):2481–2498
Tran L-SP, Quach TN, Guttikonda SK, Aldrich DL, Kumar R, Neelakandan A, Valliyodan B, Nguyen HT (2009) Molecular characterization of stress-inducible GmNAC genes in soybean. Mol Genet Genomics 281(6):647–664
Tran L-SP, Nishiyama R, Yamaguchi-Shinozaki K, Shinozaki K (2010) Potential utilization of NAC transcription factors to enhance abiotic stress tolerance in plants by biotechnological approach. GM Crops 1(1):32–39
Tripathi P, Rabara RC, Rushton PJ (2014) A systems biology perspective on the role of WRKY transcription factors in drought responses in plants. Planta 239(2):255–266
Turner NC, Wright GC, Siddique K (2001) Adaptation of grain legumes (pulses) to water-limited environments. Adv Agron 71:193-231
Udvardi MK, Kakar K, Wandrey M, Montanari O, Murray J, Andriankaja A, Zhang J-Y, Benedito V, Hofer JM, Chueng F (2007) Legume transcription factors: global regulators of plant development and response to the environment. Plant Physiol 144(2):538–549
Umezawa T, Fujita M, Fujita Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Engineering drought tolerance in plants: discovering and tailoring genes to unlock the future. Curr Opin Biotechnol 17(2):113–122
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci U S A 97(21):11632–11637
Urao T, Yakubov B, Satoh R, Yamaguchi-Shinozaki K, Seki M, Hirayama T, Shinozaki K (1999) A transmembrane hybrid-type histidine kinase in Arabidopsis functions as an osmosensor. Plant Cell 11(9):1743–1754
Urao T, Miyata S, Yamaguchi-Shinozaki K, Shinozaki K (2000) Possible His to Asp phosphorelay signaling in an Arabidopsis two-component system. FEBS Lett 478(3):227–232
Velasco R, Zharkikh A, Troggio M, Cartwright DA, Cestaro A, Pruss D, Pindo M, FitzGerald LM, Vezzulli S, Reid J (2007) A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PloS ONE 2(12):e1326
Voss I, Sunil B, Scheibe R, Raghavendra A (2013) Emerging concept for the role of photorespiration as an important part of abiotic stress response. Plant Biol 15(4):713–722
Wang W, Vinocur B, Altman A (2003) Plant responses to drought, salinity and extreme temperatures: towards genetic engineering for stress tolerance. Planta 218(1):1–14
Wang H, Huang Z, Chen Q, Zhang Z, Zhang H, Wu Y, Huang D, Huang R (2004) Ectopic overexpression of tomato JERF3 in tobacco activates downstream gene expression and enhances salt tolerance. Plant Mol Biol 55(2):183–192
Wang X, Li W, Li M, Welti R (2006) Profiling lipid changes in plant response to low temperatures. Physiol Plant 126(1):90–96
Wang YX (2013) Characterization of a novel Medicago sativa NAC transcription factor gene involved in response to drought stress. Mol Biol Rep 40(11):6451–6458
Wang C, Deng P, Chen L, Wang X, Ma H, Hu W, Yao N, Feng Y, Chai R, Yang G (2013) A wheat WRKY transcription factor TaWRKY10 confers tolerance to multiple abiotic stresses in transgenic tobacco. PloS ONE 8(6):e65120
Weake VM, Workman JL (2010) Inducible gene expression: diverse regulatory mechanisms. Nat Rev Genet 11(6):426–437
Wen J-Q, Oono K, Imai R (2002) Two novel mitogen-activated protein signaling components, OsMEK1 and OsMAP1, are involved in a moderate low-temperature signaling pathway in rice. Plant Physiol 129(4):1880–1891
Wilkins O, Nahal H, Foong J, Provart NJ, Campbell MM (2009) Expansion and diversification of the Populus R2R3-MYB family of transcription factors. Plant Physiol 149(2):981–993
Wu L, Zhang Z, Zhang H, Wang X-C, Huang R (2008) Transcriptional modulation of ethylene response factor protein JERF3 in the oxidative stress response enhances tolerance of tobacco seedlings to salt, drought, and freezing. Plant Physiol 148(4):1953–1963
Wu X, Shiroto Y, Kishitani S, Ito Y, Toriyama K (2009) Enhanced heat and drought tolerance in transgenic rice seedlings overexpressing OsWRKY11 under the control of HSP101 promoter. Plant Cell Rep 28(1):21–30
Xia N, Zhang G, Liu X-Y, Deng L, Cai G-L, Zhang Y, Wang X-J, Zhao J, Huang L-L, Kang Z-S (2010) Characterization of a novel wheat NAC transcription factor gene involved in defense response against stripe rust pathogen infection and abiotic stresses. Mol Biol Rep 37(8):3703–3712
Xie Q, Frugis G, Colgan D, Chua N-H (2000) Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development. Genes Dev 14(23):3024–3036
Xiong L, Ishitani M, Lee H, Zhu J-K (2001) The Arabidopsis LOS5/ABA3 locus encodes a molybdenum cofactor sulfurase and modulates cold stress-and osmotic stress-responsive gene expression. Plant Cell 13(9):2063–2083
Xiong L, Lee H, Ishitani M, Zhu J-K (2002) Regulation of osmotic stress-responsive gene expression by the LOS6/ABA1 locus in Arabidopsis. J Biol Chem 277(10):8588–8596
Xue G-P, Way HM, Richardson T, Drenth J, Joyce PA, McIntyre CL (2011) Overexpression of TaNAC69 leads to enhanced transcript levels of stress up-regulated genes and dehydration tolerance in bread wheat. Mol Plant 4(4):697–712
Yadav SK (2010) Cold stress tolerance mechanisms in plants. A review. Agron Sustain Dev 30(3):515–527
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yáñez M, Cáceres S, Orellana S, Bastías A, Verdugo I, Ruiz-Lara S, Casaretto JA (2009) An abiotic stress-responsive bZIP transcription factor from wild and cultivated tomatoes regulates stress-related genes. Plant Cell Rep 28(10):1497–1507
Ying S, Zhang D-F, Fu J, Shi Y-S, Song Y-C, Wang T-Y, Li Y (2012) Cloning and characterization of a maize bZIP transcription factor, ZmbZIP72, confers drought and salt tolerance in transgenic Arabidopsis. Planta 235(2):253–266
Yokotani N, Ichikawa T, Kondou Y, Matsui M, Hirochika H, Iwabuchi M, Oda K (2009) Tolerance to various environmental stresses conferred by the salt-responsive rice gene ONAC063 in transgenic Arabidopsis. Planta 229(5):1065–1075
Yoo SY, Kim Y, Kim SY, Lee JS, Ahn JH (2007) Control of flowering time and cold response by a NAC-domain protein in Arabidopsis. PloS ONE 2(7):1–10
Zhai Y, Wang Y, Li Y, Lei T, Yan F, Su L, Li X, Zhao Y, Sun X, Li J (2013) Isolation and molecular characterization of GmERF7, a soybean ethylene-response factor that increases salt stress tolerance in tobacco. Gene 513(1):174–183
Zhang C-L, He X-Y, He Z-H, Wang L-H, Xia X-C (2009a) Cloning of TaCYP707A1 gene that encodes ABA 8′-hydroxylase in common wheat (Triticum aestivum L.). Agr Sci China 8(8):902–909
Zhang G, Chen M, Li L, Xu Z, Chen X, Guo J, Ma Y (2009b) Overexpression of the soybean GmERF3 gene, an AP2/ERF type transcription factor for increased tolerances to salt, drought, and diseases in transgenic tobacco. J Exp Bot 60(13):3781–3796
Zhang H, Liu W, Wan L, Li F, Dai L, Li D, Zhang Z, Huang R (2010) Functional analyses of ethylene response factor JERF3 with the aim of improving tolerance to drought and osmotic stress in transgenic rice. Transgenic Res 19(5):809–818
Zhang W, Jeon BW, Assmann SM (2011a) Heterotrimeric G-protein regulation of ROS signalling and calcium currents in Arabidopsis guard cells. J Exp Bot 62(7):2371–2379
Zhang X, Wang L, Meng H, Wen H, Fan Y, Zhao J (2011b) Maize ABP9 enhances tolerance to multiple stresses in transgenic Arabidopsis by modulating ABA signaling and cellular levels of reactive oxygen species. Plant Mol Biol 75(4-5):365–378
Zhang XX, Tang YJ, Ma QB, Yang CY, Mu YH, Suo HC, Luo LH, Nian H (2013) OsDREB2A, a rice transcription factor, significantly affects salt tolerance in transgenic soybean. PloS ONE 8(12):e83011
Zhao T, Liang D, Wang P, Liu J, Ma F (2012) Genome-wide analysis and expression profiling of the DREB transcription factor gene family in Malus under abiotic stress. Mol Genet Genomics 287(5):423–436
Zheng X, Chen B, Lu G, Han B (2009) Overexpression of a NAC transcription factor enhances rice drought and salt tolerance. Biochem Biophys Res Commun 379(4):985–989
Zhong R, Richardson EA, Ye Z-H (2007) Two NAC domain transcription factors, SND1 and NST1, function redundantly in regulation of secondary wall synthesis in fibers of Arabidopsis. Planta 225(6):1603–1611
Zhong H, Guo Q-Q, Chen L, Ren F, Wang Q-Q, Zheng Y, Li X-B (2012) Two Brassica napus genes encoding NAC transcription factors are involved in response to high-salinity stress. Plant Cell Rep 31(11):1991–2003
Zhou Y, Lam HM, Zhang J (2007) Inhibition of photosynthesis and energy dissipation induced by water and high light stresses in rice. J Exp Bot 58(5):1207–1217
Zhou QY, Tian AG, Zou HF, Xie ZM, Lei G, Huang J, Wang CM, Wang HW, Zhang JS, Chen SY (2008) Soybean WRKY-type transcription factor genes, GmWRKY13, GmWRKY21, and GmWRKY54, confer differential tolerance to abiotic stresses in transgenic Arabidopsis plants. Plant Biotechnol J 6(5):486–503
Zhu XL, Qi L, Liu X, Cai SB, Xu HJ, Huang RF, Li JR, Wei XN, Zhang ZY (2014) The wheat ERF transcription factor TaPIE1 mediates host responses to both the necrotrophic pathogen Rhizoctonia cerealis and freezing stresses. Plant Physiol. 164(3):1499-514
Zhuang J, Chen JM, Yao QH, Xiong F, Sun CC, Zhou XR, Zhang J, Xiong AS (2011) Discovery and expression profile analysis of AP2/ERF family genes from Triticum aestivum. Mol Biol Rep 38(2):745–753
Zhuang J, Jiang HH, Wang F, Peng RH, Yao QH, Xiong AS (2013) A rice OsAP23, functioning as an AP2/ERF transcription factor, reduces salt tolerance in transgenic Arabidopsis. Plant Mol Biol Rep 31(6):1336–1345
Zou M, Guan Y, Ren H, Zhang F, Chen F (2008) A bZIP transcription factor, OsABI5, is involved in rice fertility and stress tolerance. Plant Mol Biol 66(6):675–683
Acknowledgments
This work is funded by Vietnam National Foundation for Science and Technology Development (NAFOSTED) under grant number 106.16-2011.37 to Nguyen Phuong Thao.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Hoang, X., Thu, N., Thao, N., Tran, LS. (2014). Transcription Factors in Abiotic Stress Responses: Their Potentials in Crop Improvement. In: Ahmad, P., Wani, M., Azooz, M., Phan Tran, LS. (eds) Improvement of Crops in the Era of Climatic Changes. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-8824-8_14
Download citation
DOI: https://doi.org/10.1007/978-1-4614-8824-8_14
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-8823-1
Online ISBN: 978-1-4614-8824-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)