Abstract
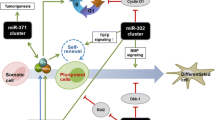
Stem cell differentiation requires a complex coordination of events to transition from a self-renewing to a differentiated cell fate. Stem cells can be pluripotent (capable of giving rise to all embryonic lineages), multipotent (possessing the potential to give rise to multiple lineages) and unipotent (capable of given rise to a single cell lineage). Regardless of their potency all stem cells must silence their self-renewal program during differentiation. The self-renewal program can be defined as the integration of external and internal stimuli that enables a cell to proliferate while maintaining its potency. Two hallmarks of the self-renewal program are a self-reinforcing transcriptional network and a specialized cell-cycle profile. In this chapter we discuss the impact of various microRNAs (miRNAs) to either reinforce or inhibit the self-renewal program of stem cells and how this added regulatory layer provides robustness to cell-fate decisions. We will focus on embryonic stem cells (ESCs) describing miRNA function in self-renewal, differentiation and de-differentiation. We will compare and contrast miRNA functions in ESCs with miRNA function in lineage specific somatic stem cells and in cancer.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Surani MA, Hayashi K, Hajkova P. Genetic and epigenetic regulators of pluripotency. Cell 2007; 128(4):747–762.
Kriegstein A, Alvarez-Buylla A. The glial nature of embryonic and adult neural stem cells. Annu Rev Neurosci 2009; 32(1):149–184.
Mikkola HKA, Orkin SH. The journey of developing hematopoietic stem cells. Development 2006; 133(19):3733–3744.
Arai F, Suda T. Quiescent stem cells in the niche. In: StemBook, ed. The Stem Cell Research Community, StemBook, doi/10.3824/stembook.1.6.1, http://www.stembook.org. 2008.
Jaenisch R, Young R. Stem cells, the molecular circuitry of pluripotency and nuclear reprogramming. Cell 2008; 132(4):567–582.
Cai X, Hagedorn CH, Cullen BR. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 2004; 10(12):1957–1966.
Lee Y, Kim M, Han J et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J 2004; 23(20):4051–4060.
Rodriguez A. Identification of mammalian microRNA host genes and transcription units. Genome Res 2004; 14(10a):1902–1910.
Basyuk E, Suavet F, Doglio A et al. Human let-7 stem-loop precursors harbor features of RNase III cleavage products. Nucleic Acids Res 2003; 31(22):6593–6597.
Gregory RI, Yan K, Amuthan G et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 2004; 432(7014):235–240.
Lee Y, Ahn C, Han J et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003; 425(6956):415–419.
Han J, Lee Y, Yeom K et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev 2004; 18(24):3016–3027.
Denli AM, Tops BBJ, Plasterk RHA et al. Processing of primary microRNAs by the Microprocessor complex. Nature 2004; 432(7014):231–235.
Han J, Lee Y, Yeom K et al. Molecular basis for the recognition of primary microRNAs by the drosha-DGCR8 complex. Cell 2006; 125(5):887–901.
Yi R, Qin Y, Macara IG et al. Exportin-5 mediates the nuclear export of premicroRNAs and short hairpin RNAs. Genes Dev 2003; 17(24):3011–3016.
Bohnsack MT, Czaplinski K, Gorlich D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of premiRNAs. RNA 2004; 10(2):185–191.
Lund E, Güttinger S, Calado A et al. Nuclear export of microRNA precursors. Science 2004; 303(5654):95–98.
Hammond SM. Dicing and slicing: The core machinery of the RNA interference pathway. FEBS Letters 2005; 579(26):5822–5829.
Schwarz DS, Hutvágner G, Du T et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 2003; 115(2):199–208.
Bartel DP. MicroRNAs: Target recognition and regulatory functions. Cell 2009; 136(2):215–233.
Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of posttranscriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 2008; 9(2):102–114.
Wang Y, Baskerville S, Shenoy A et al. Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nat Genet 2008; 40(12):1478–1483.
Bar M, Wyman SK, Fritz BR et al. MicroRNA discovery and profiling in human embryonic stem cells by deep sequencing of small RNA libraries. Stem Cells 2008; 26(10):2496–2505.
Suh M, Lee Y, Kim JY et al. Human embryonic stem cells express a unique set of microRNAs. Dev Biol 2004; 270(2):488–498.
Kanellopoulou C, Muljo SA, Kung AL et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev 2005; 19(4):489–501.
Murchison EP, Partridge JF, Tam OH et al. Characterization of Dicer-deficient murine embryonic stem cells. Proc Natl Acad Sci USA 2005; 102(34):12135–12140.
Wang Y, Medvid R, Melton C et al. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat Genet 2007; 39(3):380–385.
Savatier P, Huang S, Szekely L et al. Contrasting patterns of retinoblastoma protein expression in mouse embryonic stem cells and embryonic fibroblasts. Oncogene 1994; 9(3):809–818.
Planas-Silva MD, Weinberg RA. The restriction point and control of cell proliferation. Curr Opin Cell Biol 1997; 9(6):768–772.
Giacinti C, Giordano A. RB and cell cycle progression. Oncogene 2006; 25(38):5220–5227.
Savatier P, Lapillonne H, van Grunsven LA et al. Withdrawal of differentiation inhibitory activity/leukemia inhibitory factor up-regulates D-type cyclins and cyclin-dependent kinase inhibitors in mouse embryonic stem cells. Oncogene 1996; 12(2):309–322.
Mittnacht S. Control of pRB phosphorylation. Curr Opin Genet Dev 1998; 8(1):21–27.
Melton C, Judson R, Blelloch R. Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature 2010; Advance Online.
Singh AM, Dalton S. The cell cycle and Myc intersect with mechanisms that regulate pluripotency and reprogramming. Cell Stem Cell 2009; 5(2):141–149.
Cartwright P, McLean C, Sheppard A et al. LIF/STAT3 controls ES cell self-renewal and pluripotency by a Myc-dependent mechanism. Development 2005; 132(5):885–896.
Lin C, Jackson AL, Guo J et al. Myc-regulated microRNAs attenuate embryonic stem cell differentiation. EMBO J 2009; 28(20):3157–3170.
Benetti R, Gonzalo S, Jaco I et al. A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nat Struct Mol Biol 2008; 15(3):268–279.
Sinkkonen L, Hugenschmidt T, Berninger P et al. MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat Struct Mol Biol 2008; 15(3):259–267.
Xu N, Papagiannakopoulos T, Pan G et al. MicroRNA-145 regulates OCT4, SOX2 and KLF4 and represses pluripotency in human embryonic stem cells. Cell 2009; 137(4):647–658.
Tay YM, Tam W, Ang Y et al. MicroRNA-134 modulates the differentiation of mouse embryonic stem cells, where it causes posttranscriptional attenuation of Nanog and LRH1. Stem Cells 2008; 26(1):17–29.
Tay Y, Zhang J, Thomson AM et al. MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 2008; 455(7216):1124–1128.
Johnson CD, Esquela-Kerscher A, Stefani G et al. The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res 2007; 67(16):7713–7722.
Kumar MS, Erkeland SJ, Pester RE et al. Suppression of nonsmall cell lung tumor development by the let-7 microRNA family. Proc Natl Acad Sci USA 2008; 105(10):3903–3908.
Schultz J, Lorenz P, Gross G et al. MicroRNA let-7b targets important cell cycle molecules in malignant melanoma cells and interferes with anchorage-independent growth. Cell Res 2008; 18(5):549–557.
Reinhart BJ, Slack FJ, Basson M et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 2000; 403(6772):901–906.
Pasquinelli AE, Reinhart BJ, Slack F et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 2000; 408(6808):86–89.
Landgraf P, Rusu M, Sheridan R et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007; 129(7):1401–1414.
Griffiths-Jones S. The microRNA Registry. Nucl Acids Res 2004; 32(suppl 1):D109–111.
Griffiths-Jones S, Grocock RJ, van Dongen S et al. miRBase: microRNA sequences, targets and gene nomenclature. Nucl Acids Res 2006; 1;34(Database issue):D140-4
Griffiths-Jones S, Saini HK, van Dongen S et al. miRBase: tools for microRNA genomics. Nucl Acids Res 2008; 36(suppl_1):D154–158.
Hagan JP, Piskounova E, Gregory RI. Lin28 recruits the TUTase Zcchc11 to inhibit let-7 maturation in mouse embryonic stem cells. Nat Struct Mol Biol 2009; 16(10):1021–1025.
Heo I, Joo C, Cho J et al. Lin28 mediates the terminal uridylation of let-7 precursor MicroRNA. Mol Cell 2008; 32(2):276–84.
Heo I, Joo C, Kim Y et al. TUT4 in concert with Lin28 suppresses microRNA biogenesis through premicroRNA uridylation. Cell 2009; 138(4):696–708.
Rybak A, Fuchs H, Smirnova L et al. A feedback loop comprising lin-28 and let-7 controls prelet-7 maturation during neural stem-cell commitment. Nat Cell Biol 2008; 10(8):987–93.
Thomson JM, Newman M, Parker JS et al. Extensive posttranscriptional regulation of microRNAs and its implications for cancer. Genes Dev 2006; 20(16):2202–7.
Viswanathan SR, Daley GQ, Gregory RI. Selective blockade of microRNA processing by Lin28. Science 2008; 320(5872):97–100.
Wu L, Belasco JG. Micro-RNA regulation of the mammalian lin-28 gene during neuronal differentiation of embryonal carcinoma cells. Mol Cell Biol 2005; 25(21):9198–9208.
Yang D, Moss EG. Temporally regulated expression of Lin-28 in diverse tissues of the developing mouse. Gene Expression Patterns 2003; 3(6):719–726.
Marson A, Levine SS, Cole MF et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 2008; 134(3):521–533.
Burdon T, Smith A, Savatier P. Signalling, cell cycle and pluripotency in embryonic stem cells. Trends Cell Biol 2002; 12(9):432–438.
Judson RL, Babiarz JE, Venere M et al. Embryonic stem cell-specific microRNAs promote induced pluripotency. Nat Biotechnol 2009; 27(5):459–461.
Schwamborn JC, Berezikov E, Knoblich JA. The TRIM-NHL Protein TRIM32 activates microRNAs and prevents self-renewal in mouse neural progenitors. Cell 2009; 136(5):913–925.
Maller Schulman BR, Liang X, Stahlhut C et al. The let-7 microRNA target gene, Mlin41/Trim71 is required for mouse embryonic survival and neural tube closure. Cell Cycle 2008; 7(24):3935–3942.
Rybak A, Fuchs H, Hadian K et al. The let-7 target gene mouse lin-41 is a stem cell specific E3 ubiquitin ligase for the miRNA pathway protein Ago2. Nat Cell Biol 2009; 11(12):1411–1420.
Neumuller RA, Betschinger J, Fischer A et al. Mei-P26 regulates microRNAs and cell growth in the Drosophila ovarian stem cell lineage. Nature 2008; 454(7201):241–245.
Hochedlinger K, Plath K. Epigenetic reprogramming and induced pluripotency. Development 2009; 136(4):509–523.
Kumar MS, Lu J, Mercer KL et al. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet 2007; 39(5):673–677.
Ivanovska I, Ball AS, Diaz RL et al. MicroRNAs in the miR-106b Family Regulate p21/CDKN1A and Promote Cell Cycle Progression. Mol Cell Biol 2008; 28(7):2167–2174.
Petrocca F, Visone R, Onelli MR et al. E2F1-Regulated microRNAs impair TGF[beta]-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 2008; 13(3):272–286.
He L, Thomson JM, Hemann MT et al. A microRNA polycistron as a potential human oncogene. Nature 2005; 435(7043):828–833.
Mendell JT. miRiad roles for the miR-17-92 cluster in development and disease. Cell 2008; 133(2):217–222.
Dews M, Homayouni A, Yu D et al. Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nat Genet 2006; 38(9):1060–1065.
Voorhoeve PM, le Sage C, Schrier M et al. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell 2006; 124(6):1169–1181.
Yu F, Yao H, Zhu P et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 2007; 131:1109–1123.
Trang P, Medina PP, Wiggins JF et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene [Internet] 2009 [cited 2009 Dec 16]; Available from: http://www.ncbi.nlm.nih.gov/pubmed/19966857.
Kumar MS, Erkeland SJ, Pester RE et al. Suppression of nonsmall cell lung tumor development by the let-7 microRNA family. Proc Natl Acad Sci USA 2008; 105(10):3903–3908.
Esquela-Kerscher A, Trang P, Wiggins JF et al. The let-7 microRNA reduces tumor growth in mouse models of lung cancer. Cell Cycle 2008; 7(6):759–764.
Chang T, Zeitels LR, Hwang H et al. Lin-28B transactivation is necessary for Myc-mediated let-7 repression and proliferation. Proc Natl Acad Sci USA 2009; 106(9):3384–3389.
Iliopoulos D, Hirsch HA, Struhl K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA and IL6 links inflammation to cell transformation. Cell 2009; 139(4):693–706.
Viswanathan SR, Powers JT, Einhorn W et al. Lin28 promotes transformation and is associated with advanced human malignancies. Nat Genet 2009; 41(7):843–848.
Dangi-Garimella S, Yun J, Eves EM et al. Raf kinase inhibitory protein suppresses a metastasis signalling cascade involving LIN28 and let-7. EMBO J 2009; 28(4):347–358.
Büssing I, Slack FJ, Großhans H. let-7 microRNAs in development, stem cells and cancer. Trends Mol Med 2008; 14(9):400–409.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2010 Landes Bioscience and Springer Science+Business Media
About this chapter
Cite this chapter
Melton, C., Blelloch, R. (2010). MicroRNA Regulation of Embryonic Stem Cell Self-Renewal and Differentiation. In: Meshorer, E., Plath, K. (eds) The Cell Biology of Stem Cells. Advances in Experimental Medicine and Biology, vol 695. Springer, Boston, MA. https://doi.org/10.1007/978-1-4419-7037-4_8
Download citation
DOI: https://doi.org/10.1007/978-1-4419-7037-4_8
Publisher Name: Springer, Boston, MA
Print ISBN: 978-1-4419-7036-7
Online ISBN: 978-1-4419-7037-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)