Abstract
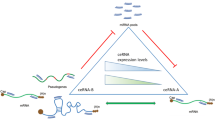
Pools of RNA molecules can act as competing endogenous RNAs (ceRNAs) and indirectly alter their expression levels by competitively binding shared microRNAs. This ceRNA cross talk yields an additional posttranscriptional regulatory layer, which plays key roles in both physiological and pathological processes. MicroRNAs can act as decoys by binding multiple RNAs, as well as RNAs can act as ceRNAs by competing for binding multiple microRNAs, leading to many cross talk interactions that could favor significant large-scale effects in spite of the weakness of single interactions. Identifying and studying these extended ceRNA interaction networks could provide a global view of the fine-tuning gene regulation in a wide range of biological processes and tumor progressions. In this chapter, we review current progress of predicting ceRNA cross talk, by summarizing the most up-to-date databases, which collect computationally predicted and/or experimentally validated miRNA–target and ceRNA–ceRNA interactions, as well as the widespread computational methods for discovering and modeling possible evidences of ceRNA–ceRNA interaction networks. These methods can be grouped in two categories: statistics-based methods exploit multivariate analysis to build ceRNA networks, by considering the miRNA expression levels when evaluating miRNA sponging relationships; mathematical methods build deterministic or stochastic models to analyze and predict the behavior of ceRNA cross talk.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Costa FF (2008) Non-coding RNAs, epigenetics and complexity. Gene 410:9–17. https://doi.org/10.1016/j.gene.2007.12.008
Friedman RC, Farh KK-H, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19:92–105. https://doi.org/10.1101/gr.082701.108
Poliseno L, Salmena L, Zhang J et al (2010) A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465:1033–1038. https://doi.org/10.1038/nature09144
Gu S, Jin L, Zhang F et al (2009) Biological basis for restriction of microRNA targets to the 3′ untranslated region in mammalian mRNAs. Nat Struct Mol Biol 16:144–150. https://doi.org/10.1038/nsmb.1552
Mukherji S, Ebert MS, Zheng GXY et al (2011) MicroRNAs can generate thresholds in target gene expression. Nat Genet 43:854–859. https://doi.org/10.1038/ng.905
Yoon J-H, Abdelmohsen K, Srikantan S et al (2012) LincRNA-p21 suppresses target mRNA translation. Mol Cell 47:648–655. https://doi.org/10.1016/j.molcel.2012.06.027
Wang J, Liu X, Wu H et al (2010) CREB up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res 38:5366–5383
Sumazin P, Yang X, Chiu H-S et al (2011) An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Cell 147:370–381. https://doi.org/10.1016/j.cell.2011.09.041
Tay Y, Kats L, Salmena L et al (2011) Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 147:344–357
Vitiello M, Evangelista M, Zhang Y et al (2020) PTENP1 is a ceRNA for PTEN: it’s CRISPR clear. J Hematol Oncol 13(73). https://doi.org/10.1186/s13045-020-00894-2
Conte F, Fiscon G, Chiara M et al (2017) Role of the long non-coding RNA PVT1 in the dysregulation of the ceRNA-ceRNA network in human breast cancer. PLoS One 12. https://doi.org/10.1371/journal.pone.0171661
Yang L, Peng X, Jin H, Liu J (2019) Long non-coding RNA PVT1 promotes autophagy as ceRNA to target ATG3 by sponging microRNA-365 in hepatocellular carcinoma. Gene 697:94–102. https://doi.org/10.1016/j.gene.2019.02.036
Colombo T, Farina L, Macino G, Paci P (2015) PVT1: a rising star among oncogenic long noncoding RNAs. Biomed Res Int 2015:304208. https://doi.org/10.1155/2015/304208
Xue W, Chen J, Liu X et al (2018) PVT1 regulates the malignant behaviors of human glioma cells by targeting miR-190a-5p and miR-488-3p. Biochim Biophys Acta (BBA) Mol Basis Dis 1864:1783–1794. https://doi.org/10.1016/j.bbadis.2018.02.022
The oncogenic role of circPVT1 in head and neck squamous cell carcinoma is mediated through the mutant p53/YAP/TEAD transcription-competent complex. Genome Biol. https://genomebiology.biomedcentral.com/articles/10.1186/s13059-017-1368-y. Accessed 24 Feb 2020
Paci P, Colombo T, Farina L (2014) Computational analysis identifies a sponge interaction network between long non-coding RNAs and messenger RNAs in human breast cancer. BMC Syst Biol 8:83. https://doi.org/10.1186/1752-0509-8-83
Tay FC, Lim JK, Zhu H et al (2015) Using artificial microRNA sponges to achieve microRNA loss-of-function in cancer cells. Adv Drug Deliv Rev 81:117–127. https://doi.org/10.1016/j.addr.2014.05.010
Thomson DW, Dinger ME (2016) Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet 17:272–283
Agarwal V, Bell GW, Nam J-W, Bartel DP (2015) Predicting effective microRNA target sites in mammalian mRNAs. elife 4:e05005
Wong N, Wang X (2015) miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Res 43:D146–D152. https://doi.org/10.1093/nar/gku1104
Liu W, Wang X (2019) Prediction of functional microRNA targets by integrative modeling of microRNA binding and target expression data. Genome Biol 20(18). https://doi.org/10.1186/s13059-019-1629-z
Karagkouni D, Paraskevopoulou MD, Chatzopoulos S et al (2018) DIANA-TarBase v8: a decade-long collection of experimentally supported miRNA–gene interactions. Nucleic Acids Res 46:D239–D245. https://doi.org/10.1093/nar/gkx1141
Huang H-Y, Lin Y-C-D, Li J et al (2020) miRTarBase 2020: updates to the experimentally validated microRNA–target interaction database. Nucleic Acids Res 48:D148–D154. https://doi.org/10.1093/nar/gkz896
Paraskevopoulou MD, Vlachos IS, Karagkouni D et al (2016) DIANA-LncBase v2: indexing microRNA targets on non-coding transcripts. Nucleic Acids Res 44:D231–D238. https://doi.org/10.1093/nar/gkv1270
Sticht C, Torre CDL, Parveen A, Gretz N (2018) miRWalk: an online resource for prediction of microRNA binding sites. PLoS One 13:e0206239. https://doi.org/10.1371/journal.pone.0206239
MIENTURNET: an interactive web tool for microRNA-target enrichment and network-based analysis. BMC Bioinformatics 20(1):545
Sarver AL, Subramanian S (2012) Competing endogenous RNA database. Bioinformation 8:731–733
Das S, Ghosal S, Sen R, Chakrabarti J (2014) ln Ce DB: database of human long noncoding RNA acting as competing endogenous RNA. PLoS One 9:e98965
Furió-Tarí P, Tarazona S, Gabaldón T et al (2016) spongeScan: a web for detecting microRNA binding elements in lncRNA sequences. Nucleic Acids Res 44:W176–W180. https://doi.org/10.1093/nar/gkw443
Jeggari A, Marks DS, Larsson E (2012) miRcode: a map of putative microRNA target sites in the long non-coding transcriptome. Bioinformatics 28:2062–2063. https://doi.org/10.1093/bioinformatics/bts344
Wang P, Zhi H, Zhang Y et al (2015) MiRSponge: a manually curated database for experimentally supported miRNA sponges and ceRNAs database 2015. https://doi.org/10.1093/database/bav098
Wang P, Li X, Gao Y et al (2019) LncACTdb 2.0: an updated database of experimentally supported ceRNA interactions curated from low- and high-throughput experiments. Nucleic Acids Res 47:D121–D127. https://doi.org/10.1093/nar/gky1144
Yang J-H, Li J-H, Shao P et al (2011) starBase: a database for exploring microRNA-mRNA interaction maps from Argonaute CLIP-Seq and Degradome-Seq data. Nucleic Acids Res 39:D202–D209. https://doi.org/10.1093/nar/gkq1056
Li J-H, Liu S, Zhou H et al (2013) starBase v2. 0: decoding miRNA-ceRNA, miRNA-ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 2013:gkt1248
Le TD, Zhang J, Liu L, Li J (2016) Computational methods for identifying miRNA sponge interactions. Brief Bioinform 2016:bbw042
Li Y, Jin X, Wang Z et al (2019) Systematic review of computational methods for identifying miRNA-mediated RNA-RNA crosstalk. Brief Bioinform 20:1193–1204. https://doi.org/10.1093/bib/bbx137
Kell DB, Oliver SG (2004) Here is the evidence, now what is the hypothesis? The complementary roles of inductive and hypothesis-driven science in the post-genomic era. BioEssays 26:99–105. https://doi.org/10.1002/bies.10385
Tomczak K, Czerwinska P, Wiznerowicz M, others (2015) The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol Pozn 19:A68–A77
Zhang Y, Xu Y, Feng L et al (2016) Comprehensive characterization of lncRNA-mRNA related ceRNA network across 12 major cancers. Oncotarget 7:64148–64167. https://doi.org/10.18632/oncotarget.11637
Do D, Bozdag S (2018) Cancerin: a computational pipeline to infer cancer-associated ceRNA interaction networks. PLoS Comput Biol 14:e1006318. https://doi.org/10.1371/journal.pcbi.1006318
List M, Dehghani Amirabad A, Kostka D, Schulz MH (2019) Large-scale inference of competing endogenous RNA networks with sparse partial correlation. Bioinforma Oxf Engl 35:i596–i604. https://doi.org/10.1093/bioinformatics/btz314
Wang J-B, Liu F-H, Chen J-H et al (2017) Identifying survival-associated modules from the dysregulated triplet network in glioblastoma multiforme. J Cancer Res Clin Oncol 143:661–671. https://doi.org/10.1007/s00432-016-2332-z
Sardina DS, Alaimo S, Ferro A et al (2017) A novel computational method for inferring competing endogenous interactions. Brief Bioinform 18:1071–1081. https://doi.org/10.1093/bib/bbw084
Zhang J, Le TD, Liu L, Li J (2017) Identifying miRNA sponge modules using biclustering and regulatory scores. BMC Bioinformatics 18(44). https://doi.org/10.1186/s12859-017-1467-5
Tong Y, Ru B, Zhang J (2018) miRNACancerMAP: an integrative web server inferring miRNA regulation network for cancer. Bioinformatics 34:3211–3213. https://doi.org/10.1093/bioinformatics/bty320
Zhang J, Liu L, Xu T et al (2019) miRspongeR: an R/bioconductor package for the identification and analysis of miRNA sponge interaction networks and modules. BMC Bioinformatics 20:235. https://doi.org/10.1186/s12859-019-2861-y
Figliuzzi M, Marinari E, De Martino A (2013) MicroRNAs as a selective channel of communication between competing RNAs: a steady-state theory. Biophys J 104:1203–1213
Ala U, Karreth FA, Bosia C et al (2013) Integrated transcriptional and competitive endogenous RNA networks are cross-regulated in permissive molecular environments. Proc Natl Acad Sci 110:7154–7159
Bosia C, Pagnani A, Zecchina R (2013) Modelling competing endogenous RNA networks. PLoS One 8:e66609
Chiu H-S, Martínez MR, Komissarova EV et al (2018) The number of titrated microRNA species dictates ceRNA regulation. Nucleic Acids Res 46:4354–4369. https://doi.org/10.1093/nar/gky286
Miotto M, Marinari E, De Martino A (2019) Competing endogenous RNA crosstalk at system level. PLoS Comput Biol 15:e1007474. https://doi.org/10.1371/journal.pcbi.1007474
Tibshirani: the lasso problem and uniqueness
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Conte, F., Fiscon, G., Sibilio, P., Licursi, V., Paci, P. (2021). An Overview of the Computational Models Dealing with the Regulatory ceRNA Mechanism and ceRNA Deregulation in Cancer. In: Poliseno, L. (eds) Pseudogenes. Methods in Molecular Biology, vol 2324. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1503-4_10
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1503-4_10
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1502-7
Online ISBN: 978-1-0716-1503-4
eBook Packages: Springer Protocols