Abstract
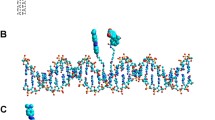
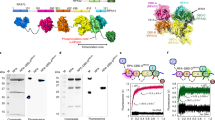
Replication protein A (RPA) is an essential single-stranded DNA (ssDNA)-binding protein that sequesters ssDNA and protects it from nucleolytic degradation. The RPA-ssDNA nucleoprotein acts as a hub to recruit over two dozen DNA metabolic enzymes onto ssDNA to coordinate DNA replication, repair, and recombination. RPA functions as a heterotrimer composed of RPA70, RPA32, and RPA14 subunits and has multiple DNA-binding and protein-interaction domains. Several of these domains are connected by disordered linkers allowing RPA to adopt a wide variety of conformations on ssDNA. Here we describe a fluorescence-based tool to monitor the dynamics of select DNA-binding domains of RPA. Noncanonical amino acids are utilized to site-specifically engineer fluorescent probes in Saccharomyces cerevisiae RPA heterologously expressed in BL21 (DE3) and its derivatives. A procedure to synthesize 4-azido-l-phenylalanine (4AZP), a noncanonical amino acid, is also described. Sites for fluorophore positioning that produce a measurable change in fluorescence upon binding to ssDNA are detailed. This fluorescence enhancement through noncanonical amino acid (FEncAA) approach can also be applied to other DNA-binding proteins to investigate the dynamics of protein-nucleic acid interactions.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wold MS (1997) Replication protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu Rev Biochem 66:61–92. https://doi.org/10.1146/annurev.biochem.66.1.61
Chen R, Wold MS (2014) Replication protein A: single-stranded DNA's first responder: dynamic DNA-interactions allow replication protein A to direct single-strand DNA intermediates into different pathways for synthesis or repair. BioEssays 36(12):1156–1161. https://doi.org/10.1002/bies.201400107
Fanning E, Klimovich V, Nager AR (2006) A dynamic model for replication protein A (RPA) function in DNA processing pathways. Nucleic Acids Res 34(15):4126–4137. https://doi.org/10.1093/nar/gkl550
Liu S, Opiyo SO, Manthey K, Glanzer JG, Ashley AK, Amerin C, Troksa K, Shrivastav M, Nickoloff JA, Oakley GG (2012) Distinct roles for DNA-PK, ATM and ATR in RPA phosphorylation and checkpoint activation in response to replication stress. Nucleic Acids Res 40(21):10780–10794. https://doi.org/10.1093/nar/gks849
Marechal A, Zou L (2015) RPA-coated single-stranded DNA as a platform for post-translational modifications in the DNA damage response. Cell Res 25(1):9–23. https://doi.org/10.1038/cr.2014.147
Zou Y, Liu Y, Wu X, Shell SM (2006) Functions of human replication protein A (RPA): from DNA replication to DNA damage and stress responses. J Cell Physiol 208(2):267–273. https://doi.org/10.1002/jcp.20622
Pokhrel N, Origanti S, Davenport EP, Gandhi D, Kaniecki K, Mehl RA, Greene EC, Dockendorff C, Antony E (2017) Monitoring replication protein A (RPA) dynamics in homologous recombination through site-specific incorporation of non-canonical amino acids. Nucleic Acids Res 45(16):9413–9426. https://doi.org/10.1093/nar/gkx598
Yates LA, Aramayo RJ, Pokhrel N, Caldwell CC, Kaplan JA, Perera RL, Spies M, Antony E, Zhang X (2018) A structural and dynamic model for the assembly of replication protein A on single-stranded DNA. Nat Commun 9(1):5447. https://doi.org/10.1038/s41467-018-07883-7
Pokhrel N, Caldwell CC, Corless EI, Tillison EA, Tibbs J, Jocic N, Tabei SMA, Wold MS, Spies M, Antony E (2019) Dynamics and selective remodeling of the DNA-binding domains of RPA. Nat Struct Mol Biol 26(2):129–136. https://doi.org/10.1038/s41594-018-0181-y
Brosey CA, Soss SE, Brooks S, Yan C, Ivanov I, Dorai K, Chazin WJ (2015) Functional dynamics in replication protein A DNA binding and protein recruitment domains. Structure 23(6):1028–1038. https://doi.org/10.1016/j.str.2015.04.008
Brosey CA, Yan C, Tsutakawa SE, Heller WT, Rambo RP, Tainer JA, Ivanov I, Chazin WJ (2013) A new structural framework for integrating replication protein A into DNA processing machinery. Nucleic Acids Res 41(4):2313–2327. https://doi.org/10.1093/nar/gks1332
Mer G, Bochkarev A, Gupta R, Bochkareva E, Frappier L, Ingles CJ, Edwards AM, Chazin WJ (2000) Structural basis for the recognition of DNA repair proteins UNG2, XPA, and RAD52 by replication factor RPA. Cell 103(3):449–456. https://doi.org/10.1016/s0092-8674(00)00136-7
Sibenaller ZA, Sorensen BR, Wold MS (1998) The 32- and 14-kilodalton subunits of replication protein A are responsible for species-specific interactions with single-stranded DNA. Biochemistry 37(36):12496–12506. https://doi.org/10.1021/bi981110+
Miyake-Stoner SJ, Miller AM, Hammill JT, Peeler JC, Hess KR, Mehl RA, Brewer SH (2009) Probing protein folding using site-specifically encoded unnatural amino acids as FRET donors with tryptophan. Biochemistry 48(25):5953–5962. https://doi.org/10.1021/bi900426d
Hammill JT, Miyake-Stoner S, Hazen JL, Jackson JC, Mehl RA (2007) Preparation of site-specifically labeled fluorinated proteins for 19F-NMR structural characterization. Nat Protoc 2(10):2601–2607. https://doi.org/10.1038/nprot.2007.379
Acknowledgments
This work was supported in part by grants from the NIH to E.A. (R01GM130746 and R01GM133967) and S.O. (R15GM126477).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Kuppa, S., Pokhrel, N., Corless, E., Origanti, S., Antony, E. (2021). Generation of Fluorescent Versions of Saccharomyces cerevisiae RPA to Study the Conformational Dynamics of Its ssDNA-Binding Domains. In: Oliveira, M.T. (eds) Single Stranded DNA Binding Proteins. Methods in Molecular Biology, vol 2281. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1290-3_9
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1290-3_9
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1289-7
Online ISBN: 978-1-0716-1290-3
eBook Packages: Springer Protocols