Abstract
Cytosine DNA methylation (5-methylcytsone, 5mC) is the major DNA modification found in the genomes of animals and plants. Although the roles of 5mC and its oxidized derivatives in the regulation of gene expression are relatively well attested and extensively explored, a number of recent studies imply that noncytosine DNA modifications may also convey specific biological functions and act as “epigenetic” marks in multicellular organisms. Here we review experimental evidence for the presence of noncytosine epigenetic modifications in metazoans and plants focusing on two “unusual” DNA bases, 5-hydroxymethyluracil (5hmU) and N6-methyladenine (6mA), and suggest potential explanations for inconsistencies in the currently available data on abundance and potential biological roles of these DNA modifications in mammals.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Gommers-Ampt JH, Borst P (1995) Hypermodified bases in DNA. FASEB J 9:1034–1042. https://doi.org/10.1096/fasebj.9.11.7649402
Fu Y, He C (2012) Nucleic acid modifications with epigenetic significance. Curr Opin Chem Biol 16:516–524. https://doi.org/10.1016/j.cbpa.2012.10.002
Kriaucionis S, Heintz N (2009) The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324:929–930. https://doi.org/10.1126/science.1169786
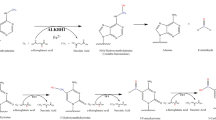
Tahiliani M, Koh K, Shen Y et al (2009) Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324:930–935. https://doi.org/10.1126/science.1170116
Ito S, Shen L, Dai Q et al (2011) Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333:1300–1303. https://doi.org/10.1126/science.1210597
He Y-F, Li B-Z, Li Z et al (2011) Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333:1303–1307. https://doi.org/10.1126/science.1210944
Song J, Pfeifer GP (2016) Are there specific readers of oxidized 5-methylcytosine bases? BioEssays 38:1038–1047. https://doi.org/10.1002/bies.201600126
Klungland A, Robertson AB (2017) Oxidized C5-methyl cytosine bases in DNA: 5-hydroxymethylcytosine; 5-formylcytosine; and 5-carboxycytosine. Free Radic Biol Med 107:62–68. https://doi.org/10.1016/j.freeradbiomed.2016.11.038
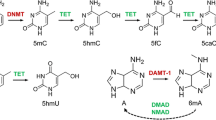
Pfaffeneder T, Spada F, Wagner M et al (2014) Tet oxidizes thymine to 5-hydroxymethyluracil in mouse embryonic stem cell DNA. Nat Chem Biol 10:574–581. https://doi.org/10.1038/nchembio.1532
Mullaart E, Lohman PH, Berends F, Vijg J (1990) DNA damage metabolism and aging. Mutat Res 237:189–210. https://doi.org/10.1016/0921-8734(90)90001-8
Ames BN, Shigenaga MK, Hagen TM (1993) Oxidants, antioxidants, and the degenerative diseases of aging (cancer/mutation/endogenous DNA adducts/oxygen radicals). Proc Natl Acad Sci U S A 90:7915–7922
Cadet J, Wagner JR (2014) Oxidatively generated base damage to cellular DNA by hydroxyl radical and one-electron oxidants: similarities and differences. Arch Biochem Biophys 557:47–54. https://doi.org/10.1016/j.abb.2014.05.001
Olinski R, Starczak M, Gackowski D (2016) Enigmatic 5-hydroxymethyluracil: oxidatively modified base, epigenetic mark or both? Mutat Res Rev Mutat Res 767:59–66. https://doi.org/10.1016/j.mrrev.2016.02.001
Cliffe LJ, Hirsch G, Wang J et al (2012) JBP1 and JBP2 proteins are Fe 2+ /2-oxoglutarate-dependent dioxygenases regulating hydroxylation of thymidine residues in trypanosome DNA. J Biol Chem 287:19886–19895. https://doi.org/10.1074/jbc.M112.341974
Ulbert S, Eide L, Seeberg E, Borst P (2004) Base J, found in nuclear DNA of Trypanosoma brucei, is not a target for DNA glycosylases. DNA Repair (Amst) 3:145–154. https://doi.org/10.1016/j.dnarep.2003.10.009
van Leeuwen F, Kieft R, Cross M, Borst P (1998) Biosynthesis and function of the modified DNA base β-d-glucosyl-hydroxymethyluracil in Trypanosoma brucei. Mol Cell Biol 18:5643–5651. https://doi.org/10.1128/mcb.18.10.5643
Borst P, Sabatini R (2008) Base J: discovery, biosynthesis, and possible functions. Annu Rev Microbiol 62:235–251. https://doi.org/10.1146/annurev.micro.62.081307.162750
Van Luenen HGAM, Farris C, Jan S et al (2012) Glucosylated hydroxymethyluracil, DNA base J, prevents transcriptional readthrough in Leishmania. Cell 150:909–921. https://doi.org/10.1016/j.cell.2012.07.030
Kawasaki F, Beraldi D, Hardisty RE et al (2017) Genome-wide mapping of 5-hydroxymethyluracil in the eukaryote parasite Leishmania. Genome Biol 18:1–8. https://doi.org/10.1186/s13059-017-1150-1
Guo JU, Su Y, Zhong C et al (2011) Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell 145:423–434. https://doi.org/10.1016/j.cell.2011.03.022
Cortellino S, Xu J, Sannai M et al (2011) Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination-base excision repair. Cell 146:67–79
Hollstein MC, Brooks P, Linn S, Ames BN (1984) Hydroxymethyluracil DNA glycosylase in mammalian cells. Proc Natl Acad Sci U S A 81:4003–4007. https://doi.org/10.1073/pnas.81.13.4003
Boorstein RJ, Levy DD, Teebor GW (1987) 5-Hydroxymethyluracil-DNA glycosylase activity may be a differentiated mammalian function. Mutat Res 183:257–263. https://doi.org/10.1016/0167-8817(87)90008-3
Rusmintratip V, Sowers LC (2000) An unexpectedly high excision capacity for mispaired 5-hydroxymethyluracil in human cell extracts. Proc Natl Acad Sci U S A 97:14183–14187. https://doi.org/10.1073/pnas.97.26.14183
Kemmerich K, Dingler FA, Rada C, Neuberger MS (2012) Germline ablation of SMUG1 DNA glycosylase causes loss of 5-hydroxymethyluracil-and UNG-backup uracil-excision activities and increases cancer predisposition of Ung−/−Msh2−/− mice. Nucleic Acids Res 40:6016–6025. https://doi.org/10.1093/nar/gks259
Doseth B, Visnes T, Wallenius A et al (2011) Uracil-DNA glycosylase in base excision repair and adaptive immunity: species differences between man and mouse. J Biol Chem 286:16669–16680. https://doi.org/10.1074/jbc.M111.230052
Alsøe L, Sarno A, Carracedo S et al (2017) Uracil accumulation and mutagenesis dominated by cytosine deamination in CpG Dinucleotides in mice lacking UNG and SMUG1. Sci Rep 7:7199. https://doi.org/10.1038/s41598-017-07314-5
Nabel CS, Manning SA, Kohli RM (2012) The curious chemical biology of cytosine: deamination, methylation, and oxidation as modulators of genomic potential. ACS Chem Biol 7:20–30. https://doi.org/10.1021/cb2002895
Franchini DM, Petersen-Mahrt SK (2014) AID and APOBEC deaminases: balancing DNA damage in epigenetics and immunity. Epigenomics 6:427–443. https://doi.org/10.2217/epi.14.35
Grin I, Ishchenko AA (2016) An interplay of the base excision repair and mismatch repair pathways in active DNA demethylation. Nucleic Acids Res 44:3713–3727. https://doi.org/10.1093/nar/gkw059
Boorstein RJ, Chiu LN, Teebor GW (1989) Phylogenetic evidence of a role for 5-hydroxymethyluracil-DNA glycosylase in the maintenance of 5-methylcytosine in DNA. Nucleic Acids Res 17:7653–7661
Birnbaum GI, Deslauriers R, Lin TS et al (1980) A novel intramolecular hydrogen bond in the crystal structure of 5-hydroxymethyl-2′-deoxyuridine, an antiviral and antineoplastic nucleoside. Conformational analysis of the deoxyribose ring. J Am Chem Soc 102:4236–4241. https://doi.org/10.1021/ja00532a041
Klose RJ, Sarraf SA, Schmiedeberg L et al (2005) DNA binding selectivity of MeCP2 due to a requirement for A/T sequences adjacent to methyl-CpG. Mol Cell 19:667–678
Greer EL, Blanco MA, Gu L et al (2015) DNA methylation on N6-adenine in C. elegans. Cell 161:868–878. https://doi.org/10.1016/j.cell.2015.04.005
Zhang G, Huang H, Liu D et al (2015) N6-methyladenine DNA modification in Drosophila. Cell 161:893–906. https://doi.org/10.1016/j.cell.2015.04.018
Yao B, Li Y, Wang Z et al (2018) Active N 6 -methyladenine demethylation by DMAD regulates gene expression by coordinating with Polycomb protein in neurons. Mol Cell 71:848–857.e6. https://doi.org/10.1016/j.molcel.2018.07.005
Shah K, Cao W, Ellison CE (2019) Adenine methylation in drosophila is associated with the tissue-specific expression of developmental and regulatory genes. G3 (Bethesda) 9:1893–1900. https://doi.org/10.1534/g3.119.400023
Liu J, Zhu Y, Luo GZ et al (2016) Abundant DNA 6mA methylation during early embryogenesis of zebrafish and pig. Nat Commun 7:1–7. https://doi.org/10.1038/ncomms13052
Wu TP, Wang T, Seetin MG et al (2016) DNA methylation on N6-adenine in mammalian embryonic stem cells. Nature 532:329–333. https://doi.org/10.1038/nature17640
Xiao C-L, Zhu S, He M et al (2018) N(6)-methyladenine DNA modification in the human genome. Mol Cell 71:306–318. https://doi.org/10.1016/j.molcel.2018.06.015
Xie Q, Wu TP, Gimple RC et al (2018) N6-methyladenine DNA modification in glioblastoma. Cell 175:1228–1243.e20. https://doi.org/10.1016/j.cell.2018.10.006
Koziol MJ, Bradshaw CR, Allen GE et al (2016) Identification of methylated deoxyadenosines in vertebrates reveals diversity in DNA modifications. Nat Struct Mol Biol 23:24–30. https://doi.org/10.1038/nsmb.3145
O’Brown ZK, Greer EL (2016) N6-Methyladenine: a conserved and dynamic DNA mark. Adv Exp Med Biol 945:213–246. https://doi.org/10.1007/978-3-319-43624-1_10
Vanyushin BF, Belozersky AN, Kokurina NA, Kadirova DX (1968) 5-Methylcytosine and 6-methylaminopurine in bacterial DNA. Nature 218:1066–1067. https://doi.org/10.1038/2181066a0
Dunn DB, Smith JD (1958) The occurrence of 6-methylaminopurine in deoxyribonucleic acids. Biochem J 68:627–636. https://doi.org/10.1042/bj0680627
Marinus MG, Morris NR (1973) Isolation of deoxyribonucleic acid methylase mutants of Escherichia coli K-12. J Bacteriol 114:1143–1150
Russell DW, Hirata RK (1989) The detection of extremely rare DNA modifications. Methylation in dam- and hsd- Escherichia coli strains. J Biol Chem 264:10787–10794
Campbell JL, Kleckner N (1990) E. coli oriC and the dnaA gene promoter are sequestered from dam methyltransferase following the passage of the chromosomal replication fork. Cell 62:967–979. https://doi.org/10.1016/0092-8674(90)90271-F
Yamaki H, Ohtsubo E, Nagai K, Maeda Y (1988) The oriC unwinding by dam methylation in Escherichia coli. Nucleic Acids Res 16:5067–5073. https://doi.org/10.1093/nar/16.11.5067
Wallecha A, Munster V, Correnti J et al (2002) Dam- and OxyR-dependent phase variation of agn43: essential elements and evidence for a new role of DNA methylation. J Bacteriol 184:3338–3347. https://doi.org/10.1128/JB.184.12.3338-3347.2002
Robbins-Manke JL, Zdraveski ZZ, Marinus M, Essigmann JM (2005) Analysis of global gene expression and double-strand-break formation in DNA adenine methyltransferase- and mismatch repair-deficient Escherichia coli. J Bacteriol 187:7027–7037. https://doi.org/10.1128/JB.187.20.7027-7037.2005
Roberts D (1985) IS10 transposition IS regulated by DNA adenine methylation. Cell 43:117–130. https://doi.org/10.1016/0092-8674(85)90017-0
Pukkila PJ, Peterson J, Herman G et al (1983) Effects of high levels of DNA adenine methylation on methyl-directed mismatch repair in Escherichia coli. Genetics 104:571–582
Messer W, Noyer-Weidner M (1988) Timing and targeting: the biological functions of Dam methylation in E. coli. Cell 54:735–737. https://doi.org/10.1016/S0092-8674(88)90911-7
Sarnacki SH, Castañeda M del RA, Llana MN et al (2013) Dam methylation participates in the regulation of PmrA/PmrB and RcsC/RcsD/RcsB two component regulatory systems in Salmonella enterica Serovar Enteritidis. PLoS One 8:e56474. https://doi.org/10.1371/journal.pone.0056474
Luria SE, Human ML (1952) A nonhereditary, host-induced variation of bacterial viruses. J Bacteriol 64:557–569
Meselson M, Yuan R (1968) DNA restriction enzyme from E. coli. Nature 217:1110–1114. https://doi.org/10.1038/2171110a0
Linn S, Arber W (1968) Host specificity of DNA produced by Escherichia coli, X. In vitro restriction of phage fd replicative form. Proc Natl Acad Sci 59:1300–1306. https://doi.org/10.1073/pnas.59.4.1300
Smith JD, Arber W, Kühnlein U (1972) Host specificity of DNA produced by Escherichia coli. J Mol Biol 63:1–8. https://doi.org/10.1016/0022-2836(72)90517-7
Zaleski P, Wojciechowski M, Piekarowicz A (2005) The role of Dam methylation in phase variation of Haemophilus influenzae genes involved in defence against phage infection. Microbiology 151:3361–3369. https://doi.org/10.1099/mic.0.28184-0
Rae PMM, Spear BB (1978) Macronuclear DNA of the hypotrichous ciliate Oxytricha fallax. Proc Natl Acad Sci 75:4992–4996. https://doi.org/10.1073/pnas.75.10.4992
Gorovsky MA, Hattman S, Pleger GL (1973) (6 N)methyl adenine in the nuclear DNA of a eucaryote, Tetrahymena pyriformis. J Cell Biol 56:697–701. https://doi.org/10.1083/jcb.56.3.697
Ammermann D, Steinbrück G, Baur R, Wohlert H (1981) Methylated bases in the DNA of the ciliate Stylonychia mytilus. Eur J Cell Biol 24:154–156
Fu Y, Luo G-Z, Chen K et al (2015) N6-methyldeoxyadenosine marks active transcription start sites in Chlamydomonas. Cell 161:879–892. https://doi.org/10.1016/j.cell.2015.04.010
Zhou C, Wang C, Liu H et al (2018) Identification and analysis of adenine N 6-methylation sites in the rice genome. Nat Plants 4:554–563. https://doi.org/10.1038/s41477-018-0214-x
Seidl MF (2017) Adenine N6-methylation in diverse fungi. Nat Genet 49:823–824. https://doi.org/10.1038/ng.3873
Mondo SJ, Dannebaum RO, Kuo RC et al (2017) Widespread adenine N6-methylation of active genes in fungi. Nat Genet 49:964–968. https://doi.org/10.1038/ng.3859
Cummings DJ, Tait A, Goddard JM (1974) Methylated bases in DNA from Paramecium aurelia. Biochim Biophys Acta 374:1–11. https://doi.org/10.1016/0005-2787(74)90194-4
Van Etten JL, Schuster AM, Girton L et al (1985) DNA methylation of viruses infecting a eukaryotic chlorella -like green alga. Nucleic Acids Res 13:3471–3478. https://doi.org/10.1093/nar/13.10.3471
Liang Z, Shen L, Cui X et al (2018) DNA N6-adenine methylation in Arabidopsis thaliana. Dev Cell 45:406–416.e3. https://doi.org/10.1016/j.devcel.2018.03.012
Zilberman D, Gehring M, Tran RK et al (2007) Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nat Genet 39:61–69. https://doi.org/10.1038/ng1929
Yao B, Cheng Y, Wang Z et al (2017) DNA N6-methyladenine is dynamically regulated in the mouse brain following environmental stress. Nat Commun 8:1–10. https://doi.org/10.1038/s41467-017-01195-y
Kweon SM, Chen Y, Moon E et al (2019) An adversarial DNA N6-Methyladenine-sensor network preserves polycomb silencing. Mol Cell 74:1138–1147.e6. https://doi.org/10.1016/j.molcel.2019.03.018
Ratel D, Ravanat J-L, Berger F, Wion D (2006) N6-methyladenine: the other methylated base of DNA. BioEssays 28:309–315. https://doi.org/10.1002/bies.20342
Schiffers S, Ebert C, Rahimoff R et al (2017) Quantitative LC-MS provides no evidence for m 6 dA or m 4 dC in the genome of mouse embryonic stem cells and tissues. Angew Chemie Int Ed Engl 56:11268–11271. https://doi.org/10.1002/anie.201700424
Lentini A, Lagerwall C, Vikingsson S et al (2018) A reassessment of DNA-immunoprecipitation-based genomic profiling. Nat Methods 15:499–504. https://doi.org/10.1038/s41592-018-0038-7
O’Brown ZK, Boulias K, Wang J et al (2019) Sources of artifact in measurements of 6mA and 4mC abundance in eukaryotic genomic DNA. BMC Genomics 20:445. https://doi.org/10.1186/s12864-019-5754-6
Charles MP, Ravanat JL, Adamski D et al (2004) N(6)-Methyldeoxyadenosine, a nucleoside commonly found in prokaryotes, induces C2C12 myogenic differentiation. Biochem Biophys Res Commun 314:476–482. https://doi.org/10.1016/j.bbrc.2003.12.132
Abakir A, Giles TC, Cristini A et al (2020) N6-methyladenosine regulates the stability of RNA:DNA hybrids in human cells. Nat Genet 52:48–55. https://doi.org/10.1038/s41588-019-0549-x
Liu J, Dou X, Chen C et al (2020) N (6)-methyladenosine of chromosome-associated regulatory RNA regulates chromatin state and transcription. Science 367(6477):580–586. https://doi.org/10.1126/science.aay6018
Acknowledgments
A.R.’s lab is supported by Biotechnology and Biological Sciences Research Council [grant number BB/N005759/1] to A.R.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Editor(s) (if applicable) and The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Lowe, P., Olinski, R., Ruzov, A. (2021). Evidence for Noncytosine Epigenetic DNA Modifications in Multicellular Eukaryotes: An Overview. In: Ruzov, A., Gering, M. (eds) DNA Modifications. Methods in Molecular Biology, vol 2198. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0876-0_2
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0876-0_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0875-3
Online ISBN: 978-1-0716-0876-0
eBook Packages: Springer Protocols