Abstract
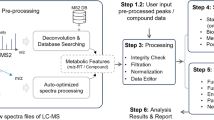
The informatics pipeline for making sense of untargeted LC–MS or GC–MS data starts with preprocessing the raw data. Results from data preprocessing undergo statistical analysis and subsequently mapped to metabolic pathways for placing untargeted metabolomics data in the biological context. ADAP is a suite of computational algorithms that has been developed specifically for preprocessing LC–MS and GC–MS data. It consists of two separate computational workflows that extract compound-relevant information from raw LC–MS and GC–MS data, respectively. Computational steps include construction of extracted ion chromatograms, detection of chromatographic peaks, spectral deconvolution, and alignment. The two workflows have been incorporated into the cross-platform and graphical MZmine 2 framework and ADAP-specific graphical user interfaces have been developed for using ADAP with ease. This chapter summarizes the algorithmic principles underlying key steps in the two workflows and illustrates how to apply ADAP to preprocess LC–MS and GC–MS data.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Pluskal T, Castillo S, Villar-Briones A, Oresic M (2010) MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinf 11:395
Jiang W, Qiu Y, Ni Y, Su M, Jia W, Du X (2010) An automated data analysis pipeline for GC-TOF-MS metabonomics studies. J Proteome Res 9(11):5974–5981
Ni Y, Qiu Y, Jiang W, Suttlemyre K, Su M, Zhang W, Jia W, Du X (2012) ADAP-GC 2.0: deconvolution of coeluting metabolites from GC/TOF-MS data for metabolomics studies. Anal Chem 84(15):6619–6629
Ni Y, Su M, Qiu Y, Jia W, Du X (2016) ADAP-GC 3.0: improved peak detection and deconvolution of co-eluting metabolites from GC/TOF-MS data for metabolomics studies. Anal Chem 88(17):8802–8811
Smirnov A, Jia W, Walker DI, Jones DP, Du X (2018) ADAP-GC 3.2: graphical software tool for efficient spectral deconvolution of gas chromatography-high-resolution mass spectrometry metabolomics data. J Proteome Res 17(1):470–478
Coble JB, Fraga CG (2014) Comparative evaluation of preprocessing freeware on chromatography/mass spectrometry data for signature discovery. J Chromatogr A 1358:155–164
Rafiei A, Sleno L (2015) Comparison of peak-picking workflows for untargeted liquid chromatography/high-resolution mass spectrometry metabolomics data analysis. Rapid Commun Mass Spectrom 29(1):119–127
Chambers MC, Maclean B, Burke R, Amodei D, Ruderman DL, Neumann S, Gatto L, Fischer B, Pratt B, Egertson J, Hoff K, Kessner D, Tasman N, Shulman N, Frewen B, Baker TA, Brusniak MY, Paulse C, Creasy D, Flashner L, Kani K, Moulding C, Seymour SL, Nuwaysir LM, Lefebvre B, Kuhlmann F, Roark J, Rainer P, Detlev S, Hemenway T, Huhmer A, Langridge J, Connolly B, Chadick T, Holly K, Eckels J, Deutsch EW, Moritz RL, Katz JE, Agus DB, MacCoss M, Tabb DL, Mallick P (2012) A cross-platform toolkit for mass spectrometry and proteomics. Nat Biotechnol 30(10):918–920
Myers OD, Sumner SJ, Li S, Barnes S, Du X (2017) One step forward for reducing false positive and false negative compound identifications from mass spectrometry metabolomics data: new algorithms for constructing extracted ion chromatograms and detecting chromatographic peaks. Anal Chem 89(17):8696–8703
Acknowledgements
We thank the USA National Science Foundation award 1262416 and National Institutes of Health/National Cancer Institute grant U01CA235507 for funding the research and development of ADAP.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Du, X., Smirnov, A., Pluskal, T., Jia, W., Sumner, S. (2020). Metabolomics Data Preprocessing Using ADAP and MZmine 2. In: Li, S. (eds) Computational Methods and Data Analysis for Metabolomics. Methods in Molecular Biology, vol 2104. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0239-3_3
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0239-3_3
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0238-6
Online ISBN: 978-1-0716-0239-3
eBook Packages: Springer Protocols