Abstract
Candida albicans is a commensal yeast of most healthy individuals, but also one of the most prevalent human fungal pathogens. During adaptation to the mammalian host, C. albicans encounters different niches where it is exposed to several types of stress, including oxidative, nitrosative (e.g., immune system), osmotic (e.g., kidney and oral cavity) stresses and pH variation (e.g., gastrointestinal (GI) tract and vagina). C. albicans has developed the capacity to respond to the environmental changes by modifying its morphology, which comprises the yeast-to-hypha transition, white-opaque switching, and chlamydospore formation. The yeast-to-hypha transition has been very well characterized and was shown to be modulated by several external stimuli that mimic the host environment. For instance, temperature above 37 ℃, serum, alkaline pH, and CO2 concentration are all reported to enhance filamentation. The transition is characterized by the activation of an intricate regulatory network of signaling pathways, involving many transcription factors. The regulatory pathways that control either the stress response or morphogenesis are required for full virulence and promote survival of C. albicans in the host. Many of these transcriptional circuitries have been characterized, highlighting the complexity and the interconnections between the different pathways. Here, we present the major signaling pathways and the main transcription factors involved in the yeast-to-hypha transition. Furthermore, we describe the role of heat shock transcription factors in the morphogenetic transition, providing an edifying example of the complex cross talk between pathways involved in morphogenesis and stress response.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
Fungal pathogens have become responsible for increasing mortality in humans over the past four decades. In developed countries, fungal infections appear as one of the primary causes of hospital-acquired infections, most of them caused by Candida spp.—particularly Candida albicans. Candida infections range from mild superficial infections to life-threatening invasive infections. Despite the use of antifungals and other therapeutic approaches, Candida infections remain associated with a high mortality rate (Kullberg and Arendrup 2016; Pfaller and Diekema 2007).
Numerous studies have shown that morphogenesis is tightly linked to the ability of C. albicans to cause disease. Although C. albicans displays an extended array of morphological transitions (e.g., chlamydospore formation, GUT phenotype, white-opaque switching, gray phenotype), the yeast-to-hypha transition appears to be an important virulence trait (Brown and Gow 1999; Gow et al. 2002; Mukaremera et al. 2017; Nemecek et al. 2006; Peters et al. 2014; Rooney and Klein 2002; Trevijano-Contador et al. 2016). Therefore, particular interest was given to the mechanisms of transition between the unicellular yeast form and either pseudohyphal or hyphal forms, often referred to as filamentous forms (Odds 1987; Sudbery et al. 2004). For instance, the yeast form is required for dissemination into the bloodstream (Bendel et al. 2003; Saville et al. 2003) and adhesion to endothelial cells (Grubb et al. 2009; Yang et al. 2014). In contrast, the filamentous form is needed for tissue invasion and for escaping from immune cells, yielding resistance toward phagocytosis (Erwig and Gow 2016; Fradin et al. 2005; Lorenz et al. 2004; Rooney and Klein 2002).
In this review, we summarize the major signaling pathways that regulate the morphogenetic transition between unicellular yeast form and filamentous forms. We report the differences between C. albicans and the model organism Saccharomyces cerevisiae, highlighting specific features of C. albicans related to morphogenesis and the associated cellular stress response.
2 Stimuli Inducing the Yeast-to-Hypha Transition
The yeast-to-hypha transition is triggered by different environmental cues that reflect the host environment, such as nutrient starvation, N-acetyl glucosamine (GlcNAc), serum, CO2 levels, neutral pH, temperature, and surface contact. In laboratory conditions, several media are used to mimic these environmental cues, such as GlcNAc-containing medium or Spider medium (Liu et al. 1994; Mattia et al. 1982). However, the most widely used inducer of morphogenesis is serum (Ernst 2000; Gow 1997), whose complex composition makes difficult the identification of the factor(s) responsible for filamentation. Early studies have shown that CO2 is an inducer of C. albicans morphogenesis (Persi et al. 1985) and that physiological CO2 levels are essential for C. albicans pathogenicity (Klengel et al. 2005). Filamentation is favored by pH values close to neutral, but is considerably reduced at pH lower than 6, while the yeast form is predominant at pH 4 (Buffo et al. 1984). High temperatures also promote filamentation via the molecular chaperone Hsp90 (O’Meara et al. 2017; Shapiro et al. 2009) and the transcription factor (TF) Sfl2 (Song et al. 2011). Finally, surface growth triggers several behaviors, such as invasion, thigmotropism, and biofilm formation (Kumamoto 2005; Kumamoto 2008; Kumamoto and Vinces 2005).
The yeast-to-hypha transition is also influenced by quorum sensing molecules, such as homoserine lactone (HSL), the 12-carbon backbone molecule dodecanol, farnesol, and tyrosol (Chen et al. 2004; Hogan et al. 2004). These compounds block the hyphal transition by the Ras-cAMP-PKA pathway and impact on cAMP-controlled genes involved in morphogenesis and stress response (Davis-Hanna et al. 2008). In the host, HSL, dodecanol, and farnesol are secreted by bacteria, such as Pseudomonas aeruginosa, and regulate C. albicans morphogenesis through independent mechanisms (Hall et al. 2011). Additionally, farnesol is produced by C. albicans as a by-product of ergosterol biosynthesis (Hornby et al. 2003) and affects many physiological functions, including cell wall maintenance, iron transport and the regulation of genes encoding heat shock proteins (Hsps) (Cao et al. 2005). Recently, it has been reported that farnesol’s effect on hyphal morphogenesis is linked to EED1 gene (see below), since an eed1 null mutant is hypersensitive to this molecule (Polke et al. 2017). Farnesol and HSL inhibit the activity of the adenylate cyclase (AC) Cyr1, and several evidences suggest that farnesol acts downstream of Ras1 (Feng et al. 1999). On the other hand, dodecanol modulates filamentation and cAMP signaling without affecting AC activity, but through a mechanism involving the transcription factor Sfl1 (Hall et al. 2011), while tyrosol is continuously released into the culture medium by C. albicans itself in order to promote germ tube formation (Chen et al. 2004). Finally, several alcohols, such as amyl alcohol, have been shown to inhibit the yeast-to-hypha morphological transition in a dose-dependent manner (Chauhan et al. 2013).
3 Hypha-Associated Genes (HAGs) and Chromatin Remodeling
The genes induced during hyphal formation have been referred to as hypha-specific genes (HSGs), in contrast to genes involved during yeast growth—yeast-specific genes (YSGs). However, it has been shown that HSGs can also be expressed by yeast cells under certain conditions (Fradin et al. 2005) and that hyphal growth does not require the induction of all HSGs (Naseem et al. 2015); thus, they can be referred to as hypha-associated genes (HAGs). During the morphogenetic transition, HAGs are under the control of several TFs, such as Efg1, Tec1, Nrg1, Tup1, Rim101, Bcr1, and Rfg1 (Eckert et al. 2007; Whiteway and Bachewich 2007). During the yeast-to-hypha morphogenetic transition, Efg1 interacts with the histone acetyltransferase (HAT) NuA4 leading to an increase in histone H4 acetylation at the promoter of HAGs (Lu et al. 2008). The Hos2/Set3 histone deacetylase complex (HDAC) acts as a regulator of hyphal development through the cAMP-PKA pathway (Hnisz et al. 2010) and modulates the expression levels of the transcription regulators BRG1, TEC1, NRG1, and EFG1, acting as both repressor and activator (Hnisz et al. 2012). The GATA family TF Brg1 recruits Hda1 to the promoter of HAGs leading to chromatin modifications that prevent the binding of the repressor Nrg1 (Lu et al. 2012; see below).
The Swi/Snf chromatin remodeler complex has been reported to bind to the promoters of HAGs only in hyphae, and this binding is mediated by Efg1 (Lu et al. 2008). Thus, Efg1 recruits NuA4 to the HAGs’ promoters, allowing in turn the recruitment of the Swi/Snf complex in order to activate transcription during the morphogenetic transition (Lu et al. 2008). Moreover, deletion of SWI/SNF or other subunits of the complex, such as Snf6, prevents hyphal growth; a swi1 deletion strain is defective in HAGs expression and is avirulent in a mouse model of systemic infection (Mao et al. 2006; Tebbji et al. 2017).
HAGs form a large group of virulence factors that contribute to pathogenesis. These factors are molecules recognized by the host, such as cell wall proteins (adhesins) and hydrolytic enzymes that contribute to penetration and tissue damage within the host (Hiller et al. 2011). In a reconstituted human epithelium (RHE) model, the expression levels of HAGs increased within 30 min after inoculation, leading to cellular damage (Hiller et al. 2011; Spiering et al. 2010).
In order to penetrate the host tissues, C. albicans produces hydrolytic enzymes, such as phospholipases, lipases, and secreted aspartyl proteinases (SAPs). Genes encoding Sap4, Sap5, and Sap6 were shown to be among the most upregulated HAGs induced by serum and elevated temperature, together with genes encoding a cell wall mannoprotein (HWP1), required for hyphal development and yeast adhesion to epithelium, and the adhesin-like protein 3 (ALS3) (Nantel et al. 2002). The agglutinin-like sequence (ALS) family is composed of glycosylphosphatidylinositol (GPI)-anchored proteins encoded by eight genes (ALS1-7 and ALS9). Among them, ALS3 is the best studied and behaves both as an adhesin and as an invasin. As an adhesin, it mediates the attachment to epithelial and endothelial cells, and to extracellular matrix; as an invasin, it binds to the E-cadherin and N-cadherin host receptors, inducing endocytosis (Phan et al. 2007). Further, C. albicans can use ferritin as a source of iron and Als3 is involved in this process. It has been proposed that C. albicans—in its hyphal form—can bind ferritin by Als3, iron release is then facilitated by acidification of the environment, and uptake is promoted by the reductive pathway (Almeida et al. 2008). Among the HAGs, extent of cell elongation 1 (ECE1) was identified in a screen of yeast and hyphal cells (Birse et al. 1993). Its expression was not detected in yeast, but within 30 min after hyphal induction. However, despite the association between ECE1 expression levels and morphogenesis, ece1 null mutants are not defective for filamentation, showing that ECE1 is not essential for this process (Birse et al. 1993). Recently, it was shown that Ece1 is critical for epithelial damage and C. albicans recognition by the immune system and that ECE1 encodes a cytolytic peptide toxin, called “Candidalysin,” secreted during infection. C. albicans strains lacking this toxin are avirulent in an oropharyngeal candidiasis (OPC) mouse model (Moyes et al. 2016). Hyphal G cyclin 1 (HGC1) is another HAG that encodes a cyclin involved in the cell-cycle regulation that plays a role in polarized growth and represses cell separation of hyphae. Mutants lacking HGC1 are defective for filamentation, but the constitutive expression of HGC1 is not sufficient to activate morphogenesis (Zheng et al. 2004). Furthermore, the TF Ume6 has a role in HGC1 regulation, since the HGC1 expression levels are significantly reduced in ume6 deletion strains, although there is no evidence of a direct regulation mechanism (Fan et al. 2013).
Morphogenesis is important for the ability of C. albicans to cause disease, even if it could be uncoupled to some extent from virulence based on the observation from large-scale studies (Noble et al. 2010; Perez et al. 2013) that some genes that have a role in infectivity do not have an obvious role in morphology and vice versa. Nevertheless, in these large-scale studies, filamentation was tested in only one condition (solid Spider medium), and virulence was only tested in a mouse model of systemic infection (Noble et al. 2010). Therefore, the exact correlation between morphology and virulence remains to be fully appreciated in order to define those factors that contribute independently or simultaneously to these two processes. Yet, the regulation of morphogenesis is critical for C. albicans virulence and a detailed understanding of the molecular pathways that regulate the yeast-to-hypha transition will provide important insights into new therapeutic strategies.
4 Morphogenetic Transition: Major Signaling Pathways
Morphogenesis in C. albicans is promoted mainly by the cAMP-PKA and the mitogen-activated protein kinase (MAPK) pathways described below. However, other signaling pathways such as the target of rapamycin (TOR) and regulation of Ace2 morphogenesis (RAM) pathways are known to regulate morphogenesis in certain environmental conditions, such as matrix embedding and pH responses (Fig. 1).
Major signaling pathways of C. albicans morphogenesis. Several environmental factors are known to activate the yeast-to-hypha transition by affecting different components of the represented pathways. The signaling cascades lead to the activation of transcription factors, such as Efg1, Flo8, Cph1, Czf1, Bcr1, and Ume6 that directly influence the expression of hypha-associated genes. Some of these transcription factors are activated in very specific conditions. For instance, Rim101 cleavage and subsequent activation depend on the environmental pH (blue); Czf1 and Cph1 enhance filamentation in matrix-embedded conditions (purple) and on solid media (dark green), respectively. The cAMP-PKA pathway is represented in orange; the adenylate cyclase Cyr1 is directly activated by serum and CO2. Ras1 activated state also stimulates AC activity, as well as proline and methionine through Gpr1 and Gpa2. Cyr1 associates with Srv2 to increase the intracellular levels of cAMP. cAMP binds to the regulatory subunit of the PKA Bcy1 with the subsequent release of the catalytic subunits Tpk1 and Tpk2, which regulate filamentation in solid and liquid media, respectively. The MAPK pathways are represented in green. The Cek1 pathway leads to the activation of Cph1, the Hog1 pathway whose role in filamentation has not been clarified yet, is involved in the osmotic and oxidative stress response, and the Mkc1 pathway activates several transcription factors, such as Czf1, Bcr1, and Efg1. Activation of the transcription factor Tec1 depends on Cph2 (in yellow) and takes to the SAPs expression, important for virulence. Finally, several stimuli induce the expression of Brg1 (in red), which is recruited at the promoter of HAGs and, together with Hda1, occlude the binding of Nrg1, which represses transcription in a Tup-1-dependent manner. Among the genes subsequently expressed, there is UME6, whose expression depends also on the transcription factor Eed1
The cAMP-PKA pathway. The cyclic adenosine monophosphate (cAMP)-protein kinase A (PKA) pathway is well conserved across eukaryotes and regulates many cellular processes in fungi. PKA is a heterotetramer consisting of two regulatory subunits that bind and inactivate two catalytic subunits. cAMP activates PKA by a conformational change that occurs upon binding. The cAMP intracellular levels are controlled by the AC Cyr1, which synthesizes cAMP from ATP, and by phosphodiesterases, which in contrast hydrolyze cAMP. AC activity is controlled by G protein-coupled receptors. There are two classes of G proteins: the monomeric small GTPases and the heterotrimeric G protein complexes, consisting of three subunits, namely α, β, and γ. Both G protein types are activated upon GTP binding and inactivated when GDP is bound. The exchange between GDP and GTP is catalyzed by GTPase-activating proteins (GAPs) or guanine exchange factors (GEFs).
In S. cerevisiae, the cAMP-PKA pathway is required as the major intermediate by which glucose or other nutrients regulate the cell cycle (Colombo et al. 2017; Fuller and Rhodes 2012). Furthermore, it is involved in pseudohyphal differentiation in response to nitrogen starvation and requires Gpr1, a seven-transmembrane-spanning receptor able to bind glucose (Lorenz et al. 2000), which activates the homolog of the mammalian G-α subunit, Gpa2 (Kubler et al. 1997; Lorenz and Heitman 1997). This signaling culminates in the activation of the TFs Flo8 and Sfl1 (Pan and Heitman 2002). In nitrogen starvation conditions, Gpr1 binds to Gpa2 (Harashima and Heitman 2005; Xue et al. 1998) leading to the activation of cAMP production through the AC Cyr1 (also called Cdc35) upon glucose induction (Colombo et al. 1998; Kraakman et al. 1999; Lorenz et al. 2000). α-Gpa2 does not form a heterotrimeric complex with the known yeast Gβγ subunits Ste4/18, but with Gpb1/2 and Gpg1 proteins (Harashima and Heitman 2005; Kraakman et al. 1999).
Furthermore, the G protein Ras2 can activate cAMP generation by Cyr1 (Minato et al. 1994). In fungi, Ras signaling has been associated to cell shape (Cullen and Sprague 2012), stress response (de la Torre-Ruiz et al. 2010), mating (Alspaugh et al. 2000), and nutrient sensing (Broach 2012). In S. cerevisiae, the role of Ras1 and Ras2 is not totally clear: current model suggests that low-level sugar phosphorylation activates the Ras-mediated localization of Cyr1 to the plasma membrane, where Gpr1-Gpa2 can trigger the signaling (Colombo et al. 2004; Thevelein and de Winde 1999). In contrast, Pde1 and Pde2, respectively low- and high-affinity cAMP phosphodiesterases, negatively regulate cAMP levels (Hu et al. 2010; Ma et al. 1999). PKA comprises a single regulatory subunit, Bcy1 (or Sra1) (Toda et al. 1987), and three catalytic subunits Tpk1, Tpk2, and Tpk3 (Toda et al. 1987). The Tpk2 subunit activates pseudohyphal differentiation, whereas Tpk1 and Tpk3 play a negative role, inhibiting filamentous growth (Pan and Heitman 1999; Robertson and Fink 1998). Tpk2 mediates the binding of the TF Flo8 and inhibits the binding of the negative regulator Sfl1 to the promoter of the FLO11 gene, which encodes a GPI-anchored cell surface adhesin (Kobayashi et al. 1996; Rupp et al. 1999), through Flo8 and Sfl1 phosphorylation (Pan and Heitman 2002).
In C. albicans, the cAMP-PKA pathway mediates different cellular processes, such as cell-cycle regulation and stress response (Cao et al. 2017; Giacometti et al. 2009), but it is one of the major pathways that positively regulate filamentation, involving the TFs Efg1 and Flo8 (Bockmuhl and Ernst 2001; Cao et al. 2006) (Fig. 1). Indeed, reduced activity of PKA results in defects in filamentation (Bockmuhl et al. 2001; Sonneborn et al. 2000), and a flo8 deletion mutant is blocked in filamentation and in the expression of HAGs, leading to avirulence in a mouse model of systemic infection (Cao et al. 2006). Many factors have been described that activate filamentation through this pathway. For instance, amino acids such as proline or methionine stimulate hyphal growth via Gpr1 and Gpa2 (Maidan et al. 2005; Miwa et al. 2004). CO2 also activates AC by directly stimulating Cyr1 catalytic domain (Klengel et al. 2005; Parrino et al. 2017). Furthermore, serum strongly induces filamentation through the muramyl peptide, which binds to Cyr1 and stimulates AC activity (Xu et al. 2008). The cAMP levels are negatively regulated by the phosphodiesterases Pde1 and Pde2, as reported in S. cerevisiae (Hoyer et al. 1994; Jung and Stateva 2003). Ras1 is activated by the GAP Csc25 (also known as Cdc25). The Ras1 active state stimulates the AC activity of Cyr1, which interacts with the AC-associated protein Srv2 (also known as cyclase-associated protein Cap1) in order to increase the intracellular cAMP levels (Fig. 1). Cyr1 physically interacts with Ras1 through the Ras association (RA) domain in the presence of serum (Fang and Wang 2006). The increased cAMP binds to Bcy1 leading to the release of the two PKA catalytic subunits encoded by TPK1 and TPK2 (there is no Tpk3 in C. albicans). Either Tpk1 or Tpk2 can activate TFs such as Efg1, by phosphorylation (Bockmuhl and Ernst 2001) (Fig. 1). Both subunits have been linked to different cues: Tpk1 is reported to be critical for hyphal development on solid-inducing medium, while Tpk2 has a role in filamentation only in liquid medium (Bockmuhl et al. 2001). Although the mechanisms that activate Ras1-mediated hyphal growth are not fully understood, it is known that incubation at 37 ℃ induces filamentous growth through the molecular chaperone Hsp90, which has an important role in the regulation of Ras1 and AC activity in response to temperature. However, Hsp90 works via effectors downstream of PKA and distinct from Efg1 (Shapiro et al. 2009).
The Ras-cAMP-PKA pathway has been involved in virulence (Bahn et al. 2003; Hogan and Sundstrom 2009; Leberer et al. 2001). For instance, Tpk2 is required for germination and engulfment in epithelial cells and for virulence in a mouse model of OPC (Sanchez AA 2005). This pathway regulates Efg1, which is considered as the major regulator of the morphogenetic transition in C. albicans (Braun and Johnson 2000). EFG1 encodes a basic helix-loop-helix (bHLH) TF that belongs to the APSES family of fungus-specific regulators (Doedt et al. 2004). The role of Efg1 in filamentation is complex: Loss of EFG1 results in a defect in filamentation under hypha-inducing conditions (Lo et al. 1997); however, under microaerophilic or embedded conditions, hyphal development is normal in an efg1 deletion strain (Giusani et al. 2002). Therefore, under hypoxic conditions, Efg1 functions as a repressor of hyphal growth on agar at temperatures below 35 ℃ (Setiadi et al. 2006), while under normoxia, Efg1 is a strong inducer of hyphal growth (Lo et al. 1997; Stoldt et al. 1997). The hypoxic repressor function is mediated by modifications at the Efg1 N-terminus (Desai et al. 2015). Genome-wide chromatin immunoprecipitation (ChIP) analysis showed that the promoter regions bound by Efg1 in yeast-form cells are different from those bound in hyphal cells (Lassak et al. 2011). These evidences suggest a double role for Efg1, either as a repressor or as an activator of filamentation, depending on the environmental cues. Indeed, while in a model of systemic bloodstream infection (increased oxygen levels) the efg1 mutant was much less virulent than the WT (Bendel et al. 2003; Lo et al. 1997), in hypoxic niches, as in the mouse gut, the efg1 mutant was hyperproliferative (Pande et al. 2013; White et al. 2007). Additionally, it was reported that the changes in EFG1 expression, depending on histone modifying enzymes, are important to allow C. albicans to better adapt in the host, establishing optimal survival strategies (Tyc et al. 2016). A transcriptomic analysis showed an overlap between the targets of Flo8 and Efg1, and a physical interaction between these two proteins was demonstrated (Cao et al. 2006).
The MAPK pathways. The mitogen-activated protein kinase (MAPK) signal transduction pathway comprises a conserved module of three kinases: the MAP kinase kinase kinase (MAPKKK), the MAP kinase kinase (MAPKK), and the MAP kinase (MAPK). The signaling depends on three phosphotransfer steps, where the MAPKKK becomes phosphorylated and phosphorylates the MAPKK, which in turn phosphorylates the MAPK.
In S. cerevisiae, the MAPK cascade is used to mediate various cellular functions, such as osmolarity adaptation (Hohmann 2009), mating (Schrick et al. 1997), cell wall integrity (Levin 2005), and filamentation. Indeed, the genome of S. cerevisiae encodes multiple MAPKs. In order to activate morphogenesis, the G protein Ras2 stimulates the activity of the G protein Cdc42, upon nitrogen starvation signals. Cdc42 binds to GTP, becomes active, and interacts with the p21-activated kinase (PAK) Ste20, which needs to be dissociated from its negative regulator Hsl17 (Fujita et al. 1999). The Cdc42-Ste20 dimer activates the MAPK cascade: Ste11 (MAPKKK), Ste7 (MAPKK), and Kss1 (MAPK) (Cook et al. 1996; Madhani et al. 1997). Once activated, Kss1 removes repressor activity of Dig1/Dig2 dimer (also known as Rst1/Rst2) on the TF Ste12 through Dig1/Dig2 phosphorylation (Bardwell et al. 1998a, b; Cook et al. 1996). Ste12 cooperates with the TF Tec1 to bind the enhancers called filamentation and invasion response elements (FREs) and to start the transcriptional program required for filamentous growth by the activation of genes, such as FLO11 (Chou et al. 2006; Lo and Dranginis 1998; Madhani and Fink 1997). In C. albicans, three different MAPK pathways have been described: the Hog1-MAPK, the Mkc1-MAPK, and Cek1/Cek2 (see below). All the MAPK signaling pathways serve as a pattern of cascades essential for C. albicans morphogenesis, but they are also required under different conditions and they are activated by different stimuli. Hence, in contrast to S. cerevisiae the MAPK pathways of C. albicans show a broader function, and all of them play a role in morphogenesis, suggesting the importance of phenotypic plasticity for C. albicans in order to adapt and survive in different host niches.
The Hog1-MAPK pathway. Activation of the Hog1-MAPK pathway culminates in the phosphorylation, activation, and nuclear translocation of Hog1. In S. cerevisiae, this pathway is involved in osmolarity adaptation. Two separate branches have been described, where independent transmembrane osmosensors converge at the MAPKK Pbs2, which activates the Hog1 MAPK (Hohmann 2009). The Sln1 branch, the so-called two component system, is controlled by the plasma membrane-localized sensor Sln1, a histidine kinase active under low osmolarity conditions. Sln1 inactivates Ssk1 via the phosphorelay protein Ypd1. When osmolarity increases, Sln1 is inactivated, the phosphorylation steps are interrupted, and the unphosphorylated form of Ssk1 accumulates. Ssk1 binds to the regulatory domain of Ssk2 and Ssk22 MAPKKKs, triggering their autophosphorylation and activation (Posas and Saito 1998). Then, Ssk2 and Ssk22 phosphorylate and activate Pbs2 MAPKK, which in turn phosphorylate the MAPK Hog1 (de Nadal et al. 2002; Hohmann 2002; Maeda et al. 1995; Posas et al. 1996). The Sho1 branch is controlled by two mucin-like transmembrane sensors, namely Msb2 and Hkr1 (de Nadal et al. 2007; Tatebayashi et al. 2007). These sensors might monitor the movements between the cell wall and the plasma membrane, since mucins connect the cell interior with the cell wall. Sho1 is a membrane-localized scaffold protein, recruiting components to the cell surface where remodeling is necessary (Reiser et al. 2000). Sho1 is activated under high osmolarity conditions, and the signaling to Pbs2 requires the Ste11 MAPKKK, which interacts with Ste50, Ste20, and the small GTPase Cdc42 (de Nadal et al. 2002; Posas et al. 2000).
S. cerevisiae harbors only one transmembrane histidine kinase (Sln1), but three are found in C. albicans, namely Sln1, Nik1, and Chk1 (Calera et al. 1999; Nagahashi et al. 1998) (Fig. 1). Deletion of CaSLN1 only slightly affects the osmotic stress tolerance (Nagahashi et al. 1998). The second putative kinase, Nik1, is the homolog of Neurospora crassa Nik1 and is required for osmotolerance and filamentation (Alex et al. 1996; Alex et al. 1998; Srikantha et al. 1998). Finally, CaChk1 modulates the expression of cell surface components (Kruppa et al. 2004), showing a role in virulence and morphogenesis (Calera et al. 1999; Calera and Calderone 1999).
Homologs of S. cerevisiae components of the osmostress pathway (PBS2, SSK1 and SHO1) have also been identified in C. albicans, but they exhibit different functions as compared to their counterparts in S. cerevisiae. Deletion of PBS2 prevents the relay of both osmotic and oxidative stress signals to Hog1 (Arana et al. 2005). In contrast, deletion of SSK1 prevents activation of Hog1 in response to oxidative stress. This contrasts with the observations in S. cerevisiae where inactivation of both the Sho1- and the Sln1 osmosensing branches prevents relay of osmotic stress signals to Hog1 (Maeda et al. 1995). Furthermore, sho1 mutants are sensitive to oxidative stress, but Sho1 has a minor role in the transmission of the phosphorylation signal to Hog1 in response to oxidative stress, which probably occurs through the Sln1-Ssk1 branch (Roman et al. 2005). However, which histidine kinase regulates Ssk1 in response to oxidative stress is unclear, since sln1 mutant are still able to activate the pathway in response to H2O2 (Li et al. 2004; Roman et al. 2005). Furthermore, a sho1 null mutant is sensitive to cell wall interfering compounds, showing modifications in the cell wall architecture (Roman et al. 2005). Therefore, in C. albicans the Hog1 pathway is involved not only in both osmotic (Arana et al. 2005; San Jose et al. 1996) and oxidative (Alonso-Monge et al. 2003; Correia et al. 2017; Du et al. 2005) stresses, but also in cell wall biosynthesis (Alonso-Monge et al. 1999, 2001; Monge et al. 2006; O’Rourke and Herskowitz 1998). Additionally, it plays a role in morphogenesis: hog1 and pbs2 mutants show enhanced filamentation in non-inducing conditions (Alonso-Monge et al. 1999; Arana et al. 2005). However, a direct role of Hog1 in the regulation of morphogenetic gene targets has not been established yet.
The Mkc1-MAPK pathway. In S. cerevisiae, protein kinase C (Pkc1) is involved in response to a large number of stresses, including osmotic (Garcia-Rodriguez et al. 2005) and nutritional (Vilella et al. 2005) stresses. Pkc1 activation leads to a MAPK module consisting of the bypass of C protein kinase (Bck1) MAPKKK that activates the Mkk1 MAPKK, which in turn phosphorylates the Mpk1 MAPK. In C. albicans, pkc1 mutants are sensitive to osmotic stress, but do not display any filamentation defect (Paravicini et al. 1996), whereas Pkc1 activates the Mkc1 pathway that in contrast has a role in morphogenesis (Navarro-Garcia et al. 1995). The MAPK Mkc1, the homolog of ScMpk1, plays a role in cellular integrity and cell wall formation (Navarro-Garcia et al. 1995) and is essential for invasive growth under embedded conditions and for biofilm formation (Kumamoto 2005). This suggests that Mkc1 signaling is relevant for processes dependent on physical interaction with external surfaces (Eisman et al. 2006). Further, mkc1 mutants are sensitive to high temperatures, lytic enzyme preparations, and cell wall antifungals, indicating an additional role of Mkc1 in the construction of the cell wall (Navarro-Garcia et al. 1995; Navarro-Garcia et al. 1998). Mkc1 is also phosphorylated upon oxidative stress (as described for Hog1), osmotic stress, antifungal treatment, in the presence of calcium and at low temperatures. The targeted TFs or those indirectly induced by Mkc1 remain to be identified, but possible candidates are Efg1, Czf1, and Bcr1 (Biswas et al. 2007) (Fig. 1). Bcr1 is involved in C. albicans hyphal development and virulence; its expression depends on the hyphal regulator Tec1 (Nobile and Mitchell 2005), a member of the TEA/ATTS family of TFs. Tec1 regulates virulence because it is required for the expression of the SAPs and for filamentation in order to escape macrophages (Schweizer et al. 2000). The SAP gene family encodes at least nine members (Sap1 to Sap9), considered putative virulence factors. Sap activity has been demonstrated to be important for attachment and penetration in the host during infection (Borg and Ruchel 1988; Ollert et al. 1993). The Sap4, Sap5, and Sap6 proteins are activated during hyphal formation (Hube et al. 1994), while the sap4 sap5 sap6 triple mutant is avirulent in vivo (Sanglard et al. 1997). TEC1 expression has been shown to be regulated by Efg1, providing an edifying example of how C. albicans can converge different signals to regulate a common set of differentially expressed genes (Lane et al. 2001) (Fig. 1).
In C. albicans, TEC1 transcription might depend on a TF named Cph2 (Lane et al. 2001). Cph2 belongs to the myc subfamily of bHLH proteins and regulates hyphal growth independently of either the MAPK or the cAMP-PKA pathways. A cph2 deletion strain shows medium-specific impairment in hyphal growth (Lane et al. 2001). Further, ectopic expression of TEC1 suppressed the filamentation defects in cph2 null mutant, suggesting that Cph2 function in hyphal growth is mediated, in part, through Tec1. Furthermore, Cph2 may also play a direct role in the transcriptional activation of some HAGs (Lane et al. 2001).
The Cek1-MAPK pathway. The Candida ERK-like kinase (Cek1) MAPK pathway was first discovered in C. albicans because of its ability to confer a cell-cycle-specific arrest when overexpressed in S. cerevisiae (Whiteway et al. 1992). S. cerevisiae homolog of Cek1, Fus3, is part of a pathway participating in mating (Elion 2000), invasive growth (Palecek et al. 2002), and vegetative growth (Cullen et al. 2000; Lee and Elion 1999). In C. albicans, the Cek1-MAPK pathway ends up with the activation of the TF Cph1, a Ste12 homolog (Liu et al. 1994) (Fig. 1). Interestingly, Cph1 is required for filamentation on solid media, but not in liquid (Leberer et al. 1996; Liu et al. 1994) and is dispensable for virulence in a murine model of systemic candidiasis (Liu et al. 1994). The efg1 cph1 double mutant fails to form filaments in hypha-inducing conditions and is avirulent in a systemic mouse model of candidiasis (Lo et al. 1997). However, under certain conditions, such as matrix-embedded, the efg1 cph1 double mutant is still able to filament, suggesting the presence of other pathways for hyphal development that involve other TFs (e.g., Czf1). Several studies have highlighted the involvement of the Cek1-MAPK pathway in morphogenesis and filamentation. For instance, deletion of CEK1 results in filamentation defects in media where nitrogen source is limiting (Csank et al. 1998). As mentioned before, nitrogen starvation is one of the signals that trigger the morphogenetic transition (Csank et al. 1998; Tripathi et al. 2002). C. albicans has two genes encoding ammonium permeases (MEP1 and MEP2) that, in addition to their role as transporters, enable growth when limiting concentrations of ammonium are the only available nitrogen source. Mep2 has also a central role in the induction of filamentation on solid surface under limiting nitrogen conditions (Biswas and Morschhauser 2005). It has been proposed that Mep2 triggers the Cph1/Cek1-mediated pathway in a Ras1-dependent manner (Biswas and Morschhauser 2005), and mutations of components of this pathway lead to reduced virulence (Marcil et al. 2002). Finally, Cek1-MAPK is also activated by growth signals (Roman et al. 2005), by quorum sensing (Sato et al. 2004), and is involved in cell wall biogenesis (Roman et al. 2005) and in mating (Magee et al. 2002).
Negative regulators of morphogenesis and chromatin remodeling. The hyphal induction program is also under negative regulation by the TFs Nrg1, Rfg1 (Kadosh and Johnson 2005), and Sfl1 (see below). Another negative regulator is Tup1, but this protein does not directly bind DNA. In S. cerevisiae, Tup1 regulates genes involved in glucose homeostasis, DNA damage, and oxygen stress response (DeRisi et al. 1997), forming a transcriptional corepressor complex with Ssn6 (Smith and Johnson 2000). In C. albicans, the tup1 mutant constitutively grows in a filamentous form, in the absence of hypha-inducing conditions (Braun and Johnson 1997). However, Tup1 is not only a repressor of filamentation, since the tup1 deletion strain has misshapen cell wall and is not able to grow at 42 ℃ (Braun and Johnson 2000).
SSN6 encodes a global transcriptional repressor, whose overexpression increases filamentation and decreases virulence (Hwang et al. 2003). As stated above, the Ssn6–Tup1 corepressor complex does not interact directly with DNA, but is targeted to specific promoters through interaction with sequence-specific DNA-binding proteins (i.e., TFs) (Keleher et al. 1992; Smith and Johnson 2000). An independent role of Ssn6 has been reported, namely through its interaction with the HDAC Rpd31. The ssn6 rpd31 double mutant is able to form elongated cells, but is unable to extend filaments, suggesting that Ssn6 has a dual role in filamentation depending on the interaction with Rpd31 (Lee et al. 2015).
The Nrg1 TF contains a highly conserved zinc finger domain. In C. albicans, Nrg1 represses transcription in a Tup1-dependent manner (Braun and Johnson 2000; Murad et al. 2001). Ectopic overexpression of NRG1 blocks hyphal growth in all conditions (Sanchez AA 2005) and attenuates virulence in a systemic infection mouse model (Saville et al. 2006). Another TF interacting with the Tup1-Ssn6 complex is repressor of filamentous growth 1 (Rfg1), homolog of ScRox1 (Khalaf and Zitomer 2001). However, RFG1 overexpression does not inhibit hyphal development in vitro or in vivo and drives pseudohyphal formation under yeast-growth conditions (Cleary et al. 2010).
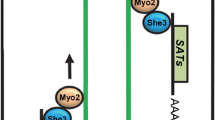
Molecular mechanisms involved in the initiation and extension of hyphal development. The two phases of hyphal development, namely initiation and maintenance, are sustained in C. albicans by two parallel pathways active either in air or in hypoxia plus CO2 (Fig. 2) (Lu et al. 2013). Serum, GlcNAc, or nutrient limitation induce the expression of the activator Brg1, which is recruited to the promoter of HAGs through changes in chromatin that occlude the binding of the repressor Nrg1 (Fig. 2). Both hyphal elongation pathways require the initiation step with a transient downregulation of Nrg1 via the cAMP-dependent PKA pathway; at the same time, Brg1 and the hypha-specific TF Ume6 are upregulated (Fig. 2). Recently, Hir1, a subunit of the HIR histone chaperone complex, has been shown to be involved in hyphal initiation, providing evidence of the tight control of gene expression during yeast-to-hypha transition (Jenull et al. 2017). Afterward, a window of opportunity permits to choose the maintenance phase: either involving Brg1 and Hda1 (air), or initiating a positive feedback loop by stabilizing Ume6 (hypoxia plus high CO2 levels) (Fig. 2) (Lu et al. 2013). In the first case, maintenance needs the HDAC Hda1 recruitment to promoters, allowing a chromatin remodeling that blocks Nrg1 access at the promoters of HAGs (Lu et al. 2011, 2014). Brg1 recruits Hda1 to promoters of HAGs under reduced Tor1 signaling for hyphal maintenance (Lu et al. 2012). In the feed-forward regulation of hyphal development, Brg1 regulates the expression of Ume6 (Lu et al. 2012). Hyphal extension and maintenance depend on the TF Ume6 and the Eed1/Def1 protein.
Hyphal initiation and maintenance. Both hyphal elongation pathways require the initiation step characterized by a transient downregulation of Nrg1 via the cAMP-dependent PKA pathway, and upregulation of Brg1 and Ume6. Serum, GlcNAc, or nutrient limitation induce the expression of Brg1. Hir1, a subunit of the HIR histone chaperone complex, has been shown to be involved in hyphal initiation, but its exact function remains to be elucidated. The maintenance phase consists of two parallel pathways active either in normoxia or hypoxia + CO2. In normoxia, Brg1 recruits the HDAC Hda1 to the promoters of HAGs, allowing a chromatin remodeling that prevents Nrg1 binding to these promoters. Brg1 regulates the expression of Ume6, which is unstable under normoxia and is degraded (dashed lines). Under hypoxia and in the presence of 5% CO2, the oxygen sensor Ofd1 and an uncharacterized CO2-sensing pathway stabilize Ume6 (thick full line), leading to hyphal development through a positive feedback loop where Ume6 binds to its own promoter, regulates HAGs expression, and inhibits NRG1 expression. Furthermore, Ume6 is regulated by Eed1
In C. albicans, UME6 encodes a filament-specific transcription regulator sufficient to direct hyphal growth in the absence of hypha-inducing conditions (Zeidler et al. 2009). This differs greatly from S. cerevisiae Ume6, whose function is to regulate arginine catabolism (Messenguy et al. 2000), DNA repair (Sweet et al. 1997), and meiosis (Rubin-Bejerano et al. 1996). C. albicans ume6 deletion mutants are defective in hyphal growth and are attenuated in virulence (Banerjee et al. 2008), while constitutive overexpression promotes hyperfilamentation in the absence of filament-inducing conditions, tissue invasion, and virulence (Banerjee et al. 2008; Carlisle et al. 2009; Zeidler et al. 2009). Ectopic overexpression of UME6 is able to rescue the hyphal growth defects observed in brg1 and hda1 mutants (Lu et al. 2012).
As described before, hyphal maintenance can occur through two different pathways (air or hypoxia plus CO2). In response to hypoxia and 5% CO2, the oxygen sensor Ofd1 and an uncharacterized CO2-sensing pathway stabilize Ume6, leading to hyphal development through a positive feedback loop where Ume6 binds to its own promoter (Fig. 2). Furthermore, Ume6 is degraded under normoxia, while very stable in hypoxia or hypercapnia: it was reported that deleting Ofd1 resulted in stabilizing Ume6 in 5% CO2, but not in air (Lu et al. 2013). Additionally, UME6 has one of the longest 5’ untranslated regions (UTRs) identified in fungi, which is predicted to form a complex and stable secondary structure. The 5’ UTR region functions to inhibit Ume6 protein expression under several filamentous-inducing conditions. Overall, the translational efficiency mechanism has evolved to fine-tune morphogenesis, and in this case, the 5’ UTR region of UME6 is found to regulate the protein translational efficiency (Childers et al. 2014). Furthermore, Ume6 is regulated posttranslationally by the cell-cycle kinase Cdc28/Cdk1, which reduces Ume6 activity, and the cyclin Cln3, an essential promoter of yeast proliferation, suppresses hyphal induction through Ume6 (Mendelsohn et al. 2017).
In addition to the TFs already described, UME6 expression depends on epithelial escape and dissemination (EED1, also known as DEF1) (Martin et al. 2011) (Fig. 1). EED1 was identified in a genome-wide analysis in a RHE model. It was shown that an eed1 mutant strain was still able to invade superficial oral epithelial cells, but in these cells the mutant grew as yeasts, without dissemination in tissues (Zakikhany et al. 2007). EED1 is unique to C. albicans, and no homolog has been detected in the genomes of the species that belong to the CUG clade, including species closely related to C. albicans (Butler et al. 2009), even if orthologs of its flanking genes exist in other fungi (Martin et al. 2011). Furthermore, it seems that EED1 does not have any DNA-binding domain; however, it is one of the key outputs of the signal transduction pathways that lead to activation of HAGs (Sudbery 2011). The reason why EED1 has evolved specifically in C. albicans could be linked to its importance in the biology of filamentation and possibly to pathogenesis. In fact, Eed1 is important for hyphal development on solid media and for the interaction with host cells, but ectopic overexpression of UME6 restored hyphal defects of eed1 loss of function (Martin et al. 2011). The suggested model proposes that Eed1 expression is regulated by Efg1 (Martin et al. 2011) and by Nrg1 (Doedt et al. 2004; Martin et al. 2011), and this is the primary element of a cycle controlling hyphal extension. The second step involves UME6 activation by Eed1, and this may be required to keep NRG1 expression at low levels to allow hyphal maintenance (Martin et al. 2011). The mechanism by which Eed1 regulates Ume6 remains unclear, and it might require additional players.
5 Morphogenetic Transition: Other Signaling Pathways
The matrix-embedded response. The ability to respond to the matrix-embedding environment is significant for C. albicans growing within host tissues. As already mentioned, the TF Czf1 is involved in the regulation of hyphal growth under these conditions (Brown et al. 1999), but the master regulator Efg1 and the TF Cph1 are also associated with the response to the matrix (Fig. 1). Ectopic expression of CZF1 in embedded cells promotes hyphal formation. However, czf1 mutants exhibit a defect in filamentation only in some embedding conditions (low temperature, in the absence of strong inducers of hyphal growth). Mutants lacking both EFG1 and CPH1, although defective in germ tube formation under many laboratory conditions (Lo et al. 1997), were still able to filament in the tongues of immunosuppressed gnotobiotic piglets and in embedded conditions (Riggle et al. 1999). In contrast, strains lacking both CPH1 and CZF1 displayed the most defective phenotype in a matrix-embedding environment, suggesting that these genes play an important role in hyphal growth under these conditions (Brown et al. 1999). Efg1 is a positive regulator of CZF1 expression, as it binds to its promoter, and deletion of EFG1 abolishes CZF1 expression. On the other hand, CZF1 expression is negatively autoregulated, through direct binding to own promoter (Vinces et al. 2006). This suggests that a different genetic program is involved in filamentous growth in embedded conditions (Vinces et al. 2006).
Another signaling pathway regulating filamentation in embedded conditions involves the G protein Rac1 (Hope et al. 2008) (Fig. 1). Rac1 is required for filamentous growth in an agar matrix (Bassilana and Arkowitz 2006) and is activated upon interaction with the GEF Dck1 (Hope et al. 2008). Rac1 and Dck1 function together with Lmo1, a protein similar to the human engulfment and cell motility (ELMO1) protein, upstream the Cek1 and Mkc1 MAP kinase pathways for invasive filamentous growth and cell wall integrity (Hope et al. 2010) (Fig. 1).
The pH-dependent response. C. albicans is highly tolerant to a wide range of pH conditions encountered in several niches within the human body. As already mentioned, pH fluctuations drive the morphological transition; moreover, C. albicans actively neutralizes the environment from either acidic or alkaline pHs. Under acidic conditions, the fungus is able to raise the pH from 4 to >7 in less than 12 h, resulting also in yeast-to-hypha transition (Vylkova et al. 2011).
In the pH-dependent filamentation, the Rim101 TF (also called Prr2 for pH response regulator) has a central role, by inducing the expression of alkaline-specific genes and repressing the expression of acid-specific genes (Fonzi 1999; Ramon et al. 1999). The extracellular pH is sensed by the transmembrane proteins Rim21 and Dfg16 (Barwell et al. 2005; Penalva et al. 2002) (Fig. 1). The protein Rim8 (or Prr1) has a crucial role, since its expression is activated in low pH conditions and mutants lacking RIM8 are defective in pH-dependent regulation of gene expression (Porta et al. 1999). Rim20 and Rim13 are also part of the pathway: They form the endosomal sorting complexes required for transport (ESCRT), required for endocytic trafficking, and together with Vps32 (also called Snf7) activate the transcription factor Rim101 by proteolytic cleavage of its C-terminal domain (Cornet et al. 2005; Kullas et al. 2004) (Fig. 1). Under alkaline conditions, Rim101 is active in its shorter form, while in acidic conditions, it is processed in a longer form whose function is not known (Li et al. 2004). Once activated, Rim101 binds to the promoter regions of different genes, activating PRA1 and PHR1 and repressing PHR2 (Ramon et al. 1999; Ramon and Fonzi 2003; Baek et al. 2006). PHR1 and PHR2 are required for normal morphology at pH above or below 5.5, respectively. These genes encode GPI-anchored glycosidases that localize at the plasma membrane and are involved in cross-linking cell wall glucans (Fonzi 1999; Muhlschlegel and Fonzi 1997). PRA1 encodes a cell wall protein whose deletion is associated to morphological defects in alkaline-induced filamentation (Sentandreu et al. 1998) and host zinc acquisition (Citiulo et al. 2012). The pH response represents another example of the dynamic interaction and adaptation of C. albicans to its host environment.
The Target of Rapamycin (TOR) pathway. Rapamycin, produced by the bacterium Streptomyces hygroscopicus, was discovered in a screen for antimicrobial activity against C. albicans and later found to have a potent immunosuppressive activity (Baker et al. 1978; Vezina et al. 1975). Rapamycin binds to its intracellular receptor FKBP (FK506-binding protein), and this complex inhibits the activity of a phosphatidylinositol 3-kinase-related kinase, the target of rapamycin (TOR), which is important for cell growth (Schmelzle and Hall 2000). There are two highly homologous TOR proteins in S. cerevisiae, Tor1 and Tor2 (Kunz et al. 1993). In addition to rapamycin-dependent function, Tor2 also regulates cytoskeletal dynamics during polarized cell growth (Helliwell et al. 1998). Furthermore, two different TOR complexes have been found in S. cerevisiae (Loewith et al. 2002; Reinke et al. 2004; Wullschleger et al. 2005): target of rapamycin complex 1 (TORC1), which contains either Tor1 or Tor2 and mediates the TOR-shared, rapamycin-sensitive pathway, and target of rapamycin complex 2 (TORC2), which contains Tor2 and mediates a TOR2-unique, rapamycin-insensitive, pathway. Within the TORC1 complex, Tor1 is conserved from yeast to humans. In C. albicans, Tor1 participates in a signaling pathway involved in cellular growth in response to nutrient availability (Rohde et al. 2008; Rohde and Cardenas 2004) and it mediates repression in cell–cell adhesion (Bastidas et al. 2009). In fact, Tor1 controls the expression of adhesin genes by interacting with the transcriptional activators Bcr1 and Efg1. In addition, it promotes the expression of the transcriptional repressors Nrg1 and Tup1 (Fig. 3) (Bastidas et al. 2009). Tor1 signaling during hyphal initiation activates the expression of the transcription factor Brg1, which recruits the HDAC Hda1 to the promoters of hypha-specific genes (Fig. 3) (Su et al. 2013). One of the TORC1 substrates is the kinase Sch9, which in C. albicans is involved in cellular growth, stress response, and filamentation under certain conditions. In particular, CaSch9 regulates filamentation differentially in liquid and solid media (Liu et al. 2010). This observation appears consistent with the fact that CaTor1 is required for filamentation on agar surface, but not in liquid medium (Bastidas et al. 2009). Further, Sch9 controls hypha formation in response to oxygen and CO2 levels: sch9 mutants showed a striking hyperfilamentation under hypoxia, but only in the presence of elevated CO2 levels (>1%) and at a high temperature (>37 ℃). Notably, to generate this hyperfilamentous phenotype, the transcription factors Czf1 and Ace2, as well as both protein kinase A isoforms, were required (Stichternoth et al. 2011).
TOR pathway during morphogenesis. Rapamycin binds to its intracellular receptor FKBP forming a complex that inhibits the activity of Tor1. Tor1 regulates adhesins genes expression by interacting with Bcr1 and Efg1 and promotes the expression of Nrg1 and Tup1. During hyphal initiation, Tor1 signaling activates the expression of Brg1, which recruits the HDAC Hda1 to the promoters of HAGs
The Regulation of Ace2 and Morphogenesis (RAM) pathway. The RAM signaling pathway is involved in the regulation of polarized morphogenesis, cellular separation, and bud development in S. cerevisiae (Nelson et al. 2003). This pathway converges in the activation of Ace2, a TF whose activity is regulated by a set of six proteins that include the Cbk1 kinase, its activator Mob2, Tao3 (Pag1), Hym1, Sog2, and the Ste20-like protein kinase Kic1 (Fig. 4) (Bidlingmaier et al. 2001; Colman-Lerner et al. 2001; Du and Novick 2002; Nelson et al. 2003; Weiss et al. 2002). During cellular division, the Cbk1–Mob2 complex blocks Ace2 in the nucleus of the daughter cell where Ace2 accumulates and drives the expression of genes that are not expressed in the mother cell (Colman-Lerner et al. 2001; Mazanka et al. 2008; Weiss et al. 2002). In C. albicans, the RAM pathway is also involved in morphogenesis and cell polarity (McNemar and Fonzi 2002; Song et al. 2008). It was shown that mutations of the RAM genes lead to hypersensitivity to cell wall or membrane stresses, defects in cell separation, and loss of cell polarity. In particular, the Cbk1–Mob2 interaction is a key event for hyphal growth (Song et al. 2008). In addition, Ace2 localization depends on the protein phosphatase Cdc14 in both yeasts (Fig. 4) (Clemente-Blanco et al. 2006; Weiss et al. 2002). Furthermore, deletion of CaACE2 caused an increased invasion of solid agar medium and led to abnormal pseudohyphal growth, even in the absence of external inducers. The mutant cells were also impaired for adherence to plastic surface, biofilm formation, and virulence (Kelly et al. 2004). Finally, Ace2 is found in two isoforms: a full-length Ace2L, involved in the septin ring formation during hyphal development that avoids cell separation, and a shorter form Ace2S, that regulates the expression of cell separation genes (Calderon-Norena et al. 2015).
Activation of RAM pathway during morphogenesis. During morphogenesis, the TF Ace2 is regulated by the kinase Cbk1 associated to its activator Mob2. This complex, whose formation is a key event for hyphal growth, is regulated by Hym1, Sog2 and the Ste20-like protein kinase Kic1 complex, together with Tao3. In addition, Ace2 localization depends on the protein phosphatase Cdc14
6 Role of the Heat Shock Factors in C. albicans Morphogenetic Transition
In response to heat stress, cells activate an ancient signaling pathway leading to the transient expression of heat shock or heat stress proteins (Hsps). Hsps display very complex protection mechanisms, but the most conserved Hsps are molecular chaperones that prevent protein misfolding and the formation of non-specific protein aggregates, and assist proteins in the acquisition of their native structures (Richter et al. 2010).
Specific heat shock TFs are required to upregulate HSPs. Heat shock TFs share a common structure characterized by an N-terminal DNA-binding domain (DBD) followed by a cluster of hydrophobic amino acids organized in heptad repeats that form leucine zippers, and C-terminal heptad repeats (Su et al. 2013). In eukaryotes, HSP expression depends on the heat shock TF Hsf1, conserved from yeasts to humans (Sorger and Pelham 1987; Wiederrecht et al. 1987). In response to temperature upshift, Hsf1 binds to a short highly conserved DNA sequence, namely the heat shock element (HSE), consisting of short tandem repeats of the sequence AGAAN (N is any nucleotide) found in the promoters of many heat shock genes (Fernandes et al. 1995).
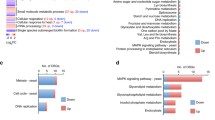
Heat shock transcription factors in eukaryotes contain a helix-turn-helix DBD. In S. cerevisiae, the DBD is important not only to bind DNA, but also for regulation of Hsf1 function. In C. albicans, in addition to Hsf1, there are three transcription factors containing an HSF-type DBD, namely Sfl1, Sfl2, and Skn7 (Fig. 5). These transcription factors play a role in morphogenesis, as reflected in the overrepresentation of genes involved in the regulation of filamentous growth among their common targets (Fig. 5b).
a HSF-type regulators transcriptional network. Sfl1 and Sfl2 regulate the expression of yeast-associated genes, and genes encoding repressors and activators of hyphal growth (blue boxes indicate common targets), by direct binding or together with Efg1 and Ndt80 (gray, dashed lines indicate hypothetical interaction). Sfl1 (dark green) induces the expression of yeast-associated genes (RME1, RHD1, YWP1) and of genes encoding repressors of hyphal growth (SSN6, NRG1) and represses genes encoding activators of hyphal growth (BRG1, UME6, TEC1). Sfl2 (red) is activated by temperature increase and negatively regulates yeast-associated genes, that are either common targets of Sfl1 (RME1, RHD1, YWP1) or unique targets of Sfl2 (PIR1, RDH3). Conversely, Sfl2 negatively regulates the expression of genes encoding repressor of hyphal growth, also regulated by Sfl1 (SSN6, NRG1, RFG1), and positively the expression of activators of hyphal growth also regulated by Sfl1 (UME6, TEC1), as well as hypha-associated genes (HGC1, HWP1, HYR1, ECE1, SAP4, FAV2, IHD1, RBT4, ALS3). Sfl1 and Sfl2 negatively regulate each other. Skn7 (dark blue) is activated during morphogenesis and oxidative stress; Skn7 induces the expression of hypha-associated genes, also targets of Sfl2 (IHD1, RBT4), negatively regulates yeast-associated genes also regulated by Sfl2 (RME1, RHD1, YWP1), and regulates the expression of several oxidative stress response genes (TSA1, TSA1B, GPX2, SSU81, TRR1). Hsf1 (dark gray) is activated upon temperature increase as Sfl2 and shares common targets with Sfl1 (SSN6), Sfl2 (ALS3), and Skn7 (TSA1, TSA1B, GPX2). b Transcriptional circuitry of HSF-type Regulators Hsf1, Sfl2, Sfl1 and Skn7. Hsf1 (gray sphere), Sfl2 (red sphere), Sfl1 (orange sphere), and Skn7 (blue sphere) share common direct transcriptional targets (open spheres, highlighted with green shading for Sfl1 and Sfl2 common targets and yellow shading for targets involving at least 3 HSF-type regulators). Targets that are specific to each regulator are highlighted with red (Sfl2), blue (Skn7), and gray (Hsf1) shadings. The associated functional categories (GO terms) that are significantly enriched are shown at the bottom right part of the figure. GO term enrichment analyses were conducted using the GO Term Finder algorithm (http://www.candidagenome.org/cgi-bin/GO/goTermFinder) at the Candida Genome Database. Binding data were taken from references (Leach et al. 2016) (Hsf1) (Znaidi et al. 2013) (Sfl2, Sfl1) and (Basso et al. 2017) (Skn7). Transcriptional network was constructed using Cytoscape version 3.4.0
Hsf1. In C. albicans, Hsf1 is required for virulence and HSE-containing genes are activated upon systemic infection (Nicholls et al. 2011), demonstrating that the connection between heat shock response and pathogenicity is crucial. In response to an acute temperature upshift, C. albicans induces classical heat shock genes and some other stress-regulated genes (Fig. 5a and b) (Enjalbert et al. 2003). In C. albicans as in other yeasts, Hsf1 is also essential for viability (Nicholls et al. 2009), and the Hsf1-HSE regulon is critical for the modulation of genes involved in protein folding under basal conditions, including the chaperone-encoding HSP70, HSP90, and HSP104 genes, as well as in response to heat stress (Leach et al. 2012; Nicholls et al. 2009).
Hsf1 physically interacts with the heat shock proteins Hsp70 and Hsp90 (Leach et al. 2012). Hsp70 and Hsp90 contribute to the Hsf1 autoregulatory circuit: in the absence of stress, Hsp90 represses Hsf1, but under thermal stress, the repression is released and Hsf1 can become active, leading to an increase in Hsp90 levels (Jarosz et al. 2010; Leach et al. 2012).
Heat shock regulation also influences C. albicans morphogenesis, since a mild heat shock is normally used to stimulate the yeast-to-hypha transition in experimental conditions (Fig. 1) (Odds 1988). The levels of the chaperone Hsp90 are increased during the yeast-to-hypha transition (Swoboda et al. 1995), and Hsp90 was demonstrated to regulate morphogenesis, since compromising its function induces the yeast-to-hypha transition via the Ras1-PKA pathway and attenuates virulence in systemic infection mouse model (Shapiro et al. 2009). Recently, it has been shown that hsf1 mutants display defective filamentation on various solid hyphal inducing media (Nair et al. 2017) and that tuning the expression level of HSF1 has a significant impact on morphogenesis (Veri et al. 2018). Indeed, overexpression of HSF1 was sufficient to drive expression of the morphogenetic regulators UME6 and BRG1, while loss of either UME6 or BRG1 was sufficient to block filamentation in response to HSF1 overexpression (Song and Carlson 1998).
Sfl1 and Sfl2. Suppressor gene for flocculation (SFL1) was first identified in S. cerevisiae where it encodes a repressor for invasion and pseudohyphal formation (Robertson and Fink 1998), and it binds specifically to GAA triplet motif of the HSE elements (Zhu et al. 2009). Sfl1 behaves as a repressor of flocculation-related genes, such as FLO11, STA1, and SUC2 (Kim et al. 2004; Robertson and Fink 1998; Song and Carlson 1998). However, this transcription factor can also act as a positive regulator of stress-responsive genes (Conlan and Tzamarias 2001; Galeote et al. 2007). Inhibition of transcription by Sfl1 is likely due to its interaction with the Ssn6–Tup1 corepressor complex. Besides, repression of gene expression by Sfl1 and Ssn6–Tup1 might involve a negative regulation of RNA polymerase II, since some components of the RNA polymerase II mediator complex, such as Sin4 and Srb10, are needed for full repression by Sfl1 (Conlan and Tzamarias 2001; Song and Carlson 1998). Tpk2 inhibits Sfl1 binding to DNA (Conlan and Tzamarias 2001; Pan and Heitman 2002) and, in parallel, activates Flo8 in the cAMP-signaling pathway (Conlan and Tzamarias 2001; Fujita et al. 1989). In C. albicans, two structural homologs of ScSfl1 have been identified, namely Sfl1 and Sfl2, and their role in morphogenesis and virulence were proven (Bauer and Wendland 2007; Song et al. 2011; Spiering et al. 2010). In fact, deletion of CaSFL1 leads to flocculation and promotes hyphal development, through HAGs and cell adhesion genes induction, while its overexpression blocks filamentation (Bauer and Wendland 2007; Li et al. 2007). Therefore, Sfl1 and Flo8 antagonistically regulate hyphal development in a mechanism similar to the one described in S. cerevisiae, where they can bind the same regions depending on phosphorylation by Tpk2 (Li et al. 2007). However, in the absence of Flo8, Sfl1 acts as a repressor at high temperature, and as an activator at low temperature, proving its dual function: in embedded agar at 24 ℃, overexpression of SFL1 enhances filamentation, whereas deletion of SFL1 blocks filamentous growth in a flo8 deletion mutant (Li et al. 2007). Interestingly, both deletion and overexpression of Sfl1 attenuate C. albicans virulence during systemic infection (Li et al. 2007).
Sfl2 is a positive regulator of filamentous growth in C. albicans, displaying high amino-acid sequence homology with Sfl1 (Song et al. 2011). Both TFs are probably the products of a gene duplication event from a shared unique ancestor and have diverged to exert antagonistic functions in filamentous growth. Interestingly, exchanging the HSF-DNA-binding domains of Sfl1 and Sfl2 almost completely reversed the repressor/activator functions of the hybrids generated from the domain swapping (Song et al. 2011), suggesting that their antagonistic functions are conferred by their ability to bind DNA. A comprehensive model for Sfl1 and Sfl2 transcriptional network was recently provided (Fig. 5a and b) (Znaidi et al. 2013). According to this model, Sfl1 and Sfl2 positively and negatively regulate a common set of targets that can be divided into three categories: repressors of hyphal growth (SSN6, NRG1, RFG1), activators of hyphal development (UME6, BRG1, TEC1), and yeast-specific genes (YSGs) (RME1, RHD1, YWP1). In particular, while the two transcription factors can act both as repressor and activator for the first two gene categories described above, only Sfl1 upregulates and Sfl2 downregulates the yeast-specific gene category (Fig. 5a). Furthermore, Sfl1 and Sfl2 directly repress each other’s expression. Lastly, Sfl1 and Sfl2 behave as «switch on/off» regulators, with Sfl1 turning off the expression of hyphal positive regulators expression and turning on the YSGs and the hyphal repressors, whereas Sfl2 turns off the YSGs and negative regulators of hyphal development and turns on activators of hyphal growth, as well as a set of HAGs, not regulated by Sfl1 (Fig. 5a). Moreover, motif discovery analyses suggest an interaction between Sfl1 and Sfl2 with the transcriptional regulators Ndt80 and Efg1, proving the complexity of the circuitry that regulates morphogenesis (Znaidi et al. 2013). Finally, evidences of the reminiscent heat shock response of these two transcription factors are reported. For instance, Sfl1 and Sfl2 can bind to the promoter of HSP104 and HSP70 (Znaidi et al. 2013), and one potential Sfl1 binding motif is similar to that for ScHsf1 (MacIsaac et al. 2006; Morozov and Siggia 2007). Taken together, these data suggest the specific needs of C. albicans to survive and adapt in warm-blooded animals, converting temperature-sensing inputs into morphogenesis output by the HSF-type transcription factors.
Skn7. The third C. albicans transcription factor that shares the HSF-DBD homology with Hsf1, Sfl1, and Sfl2 is named Skn7. Skn7 is highly conserved among fungi, displaying a uniform architecture consisting in an N-terminal HSF-DBD and a C-terminal receiver domain (Fassler and West 2011). Within the HSF domain, two residues known to be involved in contacting DNA and critical in combination for the protein function are conserved, namely phenylalanine 76 and leucine 83 (Basso et al. 2017; Fassler and West 2011). However, Skn7 presents a very interesting peculiarity when compared with the other HSF-type transcription factors: the presence of a response-regulator domain. This domain confers an additional regulation for transcriptional activity via phosphorylation of conserved residues within the protein and distinguishes Skn7 from the other HSF-type DBD proteins (Hsf1, Sfl1, and Sfl2). In C. albicans, the mechanisms that regulate Skn7 are not known. Nevertheless, it has been reported that Skn7 is involved in the oxidative stress response (Basso et al. 2017; Singh et al. 2004) and morphogenesis (Basso et al. 2017; Chauvel et al. 2012; Singh et al. 2004) (Fig. 5a and b), but how it regulates either the adaptation to oxidative stress or the morphogenetic transition has not been defined so far. C. albicans adaptation to oxidative stress depends on the TF Cap1 (Wang et al. 2006; Zhang et al. 2000), and Skn7 does not share common targets with Cap1 (Basso et al. 2017; Znaidi et al. 2009), suggesting that the role of these TFs in the oxidative stress response is different. However, Skn7 seems to prevent the accumulation of reactive oxygen species (ROS) occurring during filamentation on solid medium and, interestingly, the morphogenesis regulation appears uncoupled from the protection against intracellular ROS (Basso et al. 2017). Furthermore, Skn7 positively regulates the expression of other TFs, such as Brg1, Cph1, Czf1, Eed1, Sfl2, Tec1, and Ume6, and is required for Cph1, Tec1, and Ume6 function (Basso et al. 2017), highlighting its deep interconnection with other regulators of morphogenesis.
7 Conclusion
C. albicans exhibits remarkable morphological features of adaption to host niches that differ in terms of temperature, pH, oxygen/CO2 levels, and nutrient availability. Such environmental fluctuations are a source of stress, requiring the activation of regulatory pathways and transcriptional circuits that impact on C. albicans morphogenesis and promote survival within body organs. Some of the transcriptional circuitries governing C. albicans morphogenesis have been characterized in detail. They already highlight the complexity that lies behind modulation of their activity. As stated above, turning on and off gene expression by TFs in response to a variety of stimuli is tightly orchestrated by a series of cascades and highly interconnected signal transduction pathways, including the cAMP/PKA, the Hog1/Mkc1/Cek1-MAPK pathways or those responding to matrix embedding, pH variation, and heat shock. Through deciphering the components of the cascades activated upstream of TFs, one could provide important clues on how signals that trigger filamentous growth and stress response are sensed and transduced. On the other hand, through studying TF function and modeling the transcriptional circuitries that operate during morphogenesis, one could identify important effectors and determinants of morphological switching that act downstream of TFs, as illustrated in Fig. 5b. The knowledge gathered from deciphering the regulatory circuitries involved in morphogenesis and/or stress response in C. albicans could open up new avenues for identifying potential molecular targets for future antifungal drug development.
References
Alex LA, Borkovich KA, Simon MI (1996) Hyphal development in Neurospora crassa: involvement of a two-component histidine kinase. Proc Natl Acad Sci U S A 93(8):3416–3421
Alex LA, Korch C, Selitrennikoff CP, Simon MI (1998) COS1, a two-component histidine kinase that is involved in hyphal development in the opportunistic pathogen Candida albicans. Proc Natl Acad Sci U S A 95(12):7069–7073
Almeida RS, Brunke S, Albrecht A, Thewes S, Laue M, Edwards JE et al (2008) The hyphal-associated adhesin and invasin Als3 of Candida albicans mediates iron acquisition from host ferritin. PLoS Pathog 4(11):e1000217
Alonso-Monge R, Navarro-Garcia F, Molero G, Diez-Orejas R, Gustin M, Pla J et al (1999) Role of the mitogen-activated protein kinase Hog1p in morphogenesis and virulence of Candida albicans. J Bacteriol 181(10):3058–3068
Alonso-Monge R, Real E, Wojda I, Bebelman JP, Mager WH, Siderius M (2001) Hyperosmotic stress response and regulation of cell wall integrity in Saccharomyces cerevisiae share common functional aspects. Mol Microbiol 41(3):717–730
Alonso-Monge R, Navarro-Garcia F, Roman E, Negredo AI, Eisman B, Nombela C et al (2003) The Hog1 mitogen-activated protein kinase is essential in the oxidative stress response and chlamydospore formation in Candida albicans. Eukaryot Cell 2(2):351–361
Alspaugh JA, Cavallo LM, Perfect JR, Heitman J (2000) RAS1 regulates filamentation, mating and growth at high temperature of Cryptococcus neoformans. Mol Microbiol 36(2):352–365
Arana DM, Nombela C, Alonso-Monge R, Pla J (2005) The Pbs2 MAP kinase kinase is essential for the oxidative-stress response in the fungal pathogen Candida albicans. Microbiology 151(Pt 4):1033–1049
Baek YU, Martin SJ, Davis DA (2006) Evidence for novel pH-dependent regulation of Candida albicans Rim101, a direct transcriptional repressor of the cell wall beta-glycosidase Phr2. Eukaryot Cell 5(9):1550–1559
Bahn YS, Staab J, Sundstrom P (2003) Increased high-affinity phosphodiesterase PDE2 gene expression in germ tubes counteracts CAP1-dependent synthesis of cyclic AMP, limits hypha production and promotes virulence of Candida albicans. Mol Microbiol 50(2):391–409
Baker H, Sidorowicz A, Sehgal SN, Vezina C (1978) Rapamycin (AY-22,989), a new antifungal antibiotic. III. In vitro and in vivo evaluation. J Antibiot (Tokyo) 31(6):539–45
Banerjee M, Thompson DS, Lazzell A, Carlisle PL, Pierce C, Monteagudo C et al (2008) UME6, a novel filament-specific regulator of Candida albicans hyphal extension and virulence. Mol Biol Cell 19(4):1354–1365
Bardwell L, Cook JG, Voora D, Baggott DM, Martinez AR, Thorner J (1998a) Repression of yeast Ste12 transcription factor by direct binding of unphosphorylated Kss1 MAPK and its regulation by the Ste7 MEK. Genes Dev 12(18):2887–2898
Bardwell L, Cook JG, Zhu-Shimoni JX, Voora D, Thorner J (1998b) Differential regulation of transcription: repression by unactivated mitogen-activated protein kinase Kss1 requires the Dig1 and Dig2 proteins. Proc Natl Acad Sci U S A 95(26):15400–15405
Barwell KJ, Boysen JH, Xu W, Mitchell AP (2005) Relationship of DFG16 to the Rim101p pH response pathway in Saccharomyces cerevisiae and Candida albicans. Eukaryot Cell 4(5):890–899
Bassilana M, Arkowitz RA (2006) Rac1 and Cdc42 have different roles in Candida albicans development. Eukaryot Cell 5(2):321–329
Basso V, Znaidi S, Lagage V, Cabral V, Schoenherr F, LeibundGut-Landmann S et al (2017) The two-component response regulator Skn7 belongs to a network of transcription factors regulating morphogenesis in Candida albicans and independently limits morphogenesis-induced ROS accumulation. Mol Microbiol 106(1):157–182
Bastidas RJ, Heitman J, Cardenas ME (2009) The protein kinase Tor1 regulates adhesin gene expression in Candida albicans. PLoS Pathog 5(2):e1000294
Bauer J, Wendland J (2007) Candida albicans Sfl1 suppresses flocculation and filamentation. Eukaryot Cell 6(10):1736–1744
Bendel CM, Hess DJ, Garni RM, Henry-Stanley M, Wells CL (2003) Comparative virulence of Candida albicans yeast and filamentous forms in orally and intravenously inoculated mice. Crit Care Med 31(2):501–507
Bidlingmaier S, Weiss EL, Seidel C, Drubin DG, Snyder M (2001) The Cbk1p pathway is important for polarized cell growth and cell separation in Saccharomyces cerevisiae. Mol Cell Biol 21(7):2449–2462
Birse CE, Irwin MY, Fonzi WA, Sypherd PS (1993) Cloning and characterization of ECE1, a gene expressed in association with cell elongation of the dimorphic pathogen Candida albicans. Infect Immun 61(9):3648–3655
Biswas K, Morschhauser J (2005) The Mep2p ammonium permease controls nitrogen starvation-induced filamentous growth in Candida albicans. Mol Microbiol 56(3):649–669
Biswas S, Van Dijck P, Datta A (2007) Environmental sensing and signal transduction pathways regulating morphopathogenic determinants of Candida albicans. Microbiol Mol Biol Rev 71(2):348–376
Bockmuhl DP, Ernst JF (2001) A potential phosphorylation site for an A-type kinase in the Efg1 regulator protein contributes to hyphal morphogenesis of Candida albicans. Genetics 157(4):1523–1530
Bockmuhl DP, Krishnamurthy S, Gerads M, Sonneborn A, Ernst JF (2001) Distinct and redundant roles of the two protein kinase A isoforms Tpk1p and Tpk2p in morphogenesis and growth of Candida albicans. Mol Microbiol 42(5):1243–1257
Borg M, Ruchel R (1988) Expression of extracellular acid proteinase by proteolytic Candida spp. during experimental infection of oral mucosa. Infect Immun 56(3):626–31
Braun BR, Johnson AD (1997) Control of filament formation in Candida albicans by the transcriptional repressor TUP1. Science 277(5322):105–109
Braun BR, Johnson AD (2000) TUP1, CPH1 and EFG1 make independent contributions to filamentation in Candida albicans. Genetics 155(1):57–67
Broach JR (2012) Nutritional control of growth and development in yeast. Genetics 192(1):73–105
Brown AJ, Gow NA (1999) Regulatory networks controlling Candida albicans morphogenesis. Trends Microbiol 7(8):333–338
Brown DH Jr, Giusani AD, Chen X, Kumamoto CA (1999) Filamentous growth of Candida albicans in response to physical environmental cues and its regulation by the unique CZF1 gene. Mol Microbiol 34(4):651–662
Buffo J, Herman MA, Soll DR (1984) A characterization of pH-regulated dimorphism in Candida albicans. Mycopathologia 85(1–2):21–30
Butler G, Rasmussen MD, Lin MF, Santos MA, Sakthikumar S, Munro CA et al (2009) Evolution of pathogenicity and sexual reproduction in eight Candida genomes. Nature 459(7247):657–662
Calderon-Norena DM, Gonzalez-Novo A, Orellana-Munoz S, Gutierrez-Escribano P, Arnaiz-Pita Y, Duenas-Santero E et al (2015) A single nucleotide polymorphism uncovers a novel function for the transcription factor Ace2 during Candida albicans hyphal development. PLoS Genet 11(4):e1005152
Calera JA, Calderone R (1999) Flocculation of hyphae is associated with a deletion in the putative CaHK1 two-component histidine kinase gene from Candida albicans. Microbiology 145(Pt 6):1431–1442
Calera JA, Zhao XJ, De Bernardis F, Sheridan M, Calderone R (1999) Avirulence of Candida albicans CaHK1 mutants in a murine model of hematogenously disseminated candidiasis. Infect Immun 67(8):4280–4284
Cao YY, Cao YB, Xu Z, Ying K, Li Y, Xie Y et al (2005) cDNA microarray analysis of differential gene expression in Candida albicans biofilm exposed to farnesol. Antimicrob Agents Chemother 49(2):584–589
Cao F, Lane S, Raniga PP, Lu Y, Zhou Z, Ramon K et al (2006) The Flo8 transcription factor is essential for hyphal development and virulence in Candida albicans. Mol Biol Cell 17(1):295–307
Cao C, Wu M, Bing J, Tao L, Ding X, Liu X et al (2017) Global regulatory roles of the cAMP/PKA pathway revealed by phenotypic, transcriptomic and phosphoproteomic analyses in a null mutant of the PKA catalytic subunit in Candida albicans. Mol Microbiol 105(1):46–64
Carlisle PL, Banerjee M, Lazzell A, Monteagudo C, Lopez-Ribot JL, Kadosh D (2009) Expression levels of a filament-specific transcriptional regulator are sufficient to determine Candida albicans morphology and virulence. Proc Natl Acad Sci U S A 106(2):599–604
Chauhan NM, Shinde RB, Karuppayil SM (2013) Effect of alcohols on filamentation, growth, viability and biofilm development in Candida albicans. Braz J Microbiol 44(4):1315–1320
Chauvel M, Nesseir A, Cabral V, Znaidi S, Goyard S, Bachellier-Bassi S et al (2012) A versatile overexpression strategy in the pathogenic yeast Candida albicans: identification of regulators of morphogenesis and fitness. PLoS ONE 7(9):e45912
Chen H, Fujita M, Feng Q, Clardy J, Fink GR (2004) Tyrosol is a quorum-sensing molecule in Candida albicans. Proc Natl Acad Sci U S A 101(14):5048–5052
Childers DS, Mundodi V, Banerjee M, Kadosh D (2014) A 5’ UTR-mediated translational efficiency mechanism inhibits the Candida albicans morphological transition. Mol Microbiol 92(3):570–585
Chou S, Lane S, Liu H (2006) Regulation of mating and filamentation genes by two distinct Ste12 complexes in Saccharomyces cerevisiae. Mol Cell Biol 26(13):4794–4805
Citiulo F, Jacobsen ID, Miramon P, Schild L, Brunke S, Zipfel P et al (2012) Candida albicans scavenges host zinc via Pra1 during endothelial invasion. PLoS Pathog 8(6):e1002777
Cleary IA, Mulabagal P, Reinhard SM, Yadev NP, Murdoch C, Thornhill MH et al (2010) Pseudohyphal regulation by the transcription factor Rfg1p in Candida albicans. Eukaryot Cell 9(9):1363–1373
Clemente-Blanco A, Gonzalez-Novo A, Machin F, Caballero-Lima D, Aragon L, Sanchez M et al (2006) The Cdc14p phosphatase affects late cell-cycle events and morphogenesis in Candida albicans. J Cell Sci 119(Pt 6):1130–1143
Colman-Lerner A, Chin TE, Brent R (2001) Yeast Cbk1 and Mob2 activate daughter-specific genetic programs to induce asymmetric cell fates. Cell 107(6):739–750
Colombo S, Ma P, Cauwenberg L, Winderickx J, Crauwels M, Teunissen A et al (1998) Involvement of distinct G-proteins, Gpa2 and Ras, in glucose- and intracellular acidification-induced cAMP signalling in the yeast Saccharomyces cerevisiae. EMBO J 17(12):3326–3341
Colombo S, Ronchetti D, Thevelein JM, Winderickx J, Martegani E (2004) Activation state of the Ras2 protein and glucose-induced signaling in Saccharomyces cerevisiae. J Biol Chem 279(45):46715–46722
Colombo S, Broggi S, Collini M, D’Alfonso L, Chirico G, Martegani E (2017) Detection of cAMP and of PKA activity in Saccharomyces cerevisiae single cells using Fluorescence Resonance Energy Transfer (FRET) probes. Biochem Biophys Res Commun 487(3):594–599
Conlan RS, Tzamarias D (2001) Sfl1 functions via the co-repressor Ssn6-Tup1 and the cAMP-dependent protein kinase Tpk2. J Mol Biol 309(5):1007–1015
Cook JG, Bardwell L, Kron SJ, Thorner J (1996) Two novel targets of the MAP kinase Kss1 are negative regulators of invasive growth in the yeast Saccharomyces cerevisiae. Genes Dev 10(22):2831–2848
Cornet M, Bidard F, Schwarz P, Da Costa G, Blanchin-Roland S, Dromer F et al (2005) Deletions of endocytic components VPS28 and VPS32 affect growth at alkaline pH and virulence through both RIM101-dependent and RIM101-independent pathways in Candida albicans. Infect Immun 73(12):7977–7987
Correia I, Alonso-Monge R, Pla J (2017) Corrigendum: the Hog1 MAP kinase promotes the recovery from cell cycle arrest induced by hydrogen peroxide in Candida albicans. Front Microbiol 8:555
Csank C, Schroppel K, Leberer E, Harcus D, Mohamed O, Meloche S et al (1998) Roles of the Candida albicans mitogen-activated protein kinase homolog, Cek1p, in hyphal development and systemic candidiasis. Infect Immun 66(6):2713–2721
Cullen PJ, Sprague GF Jr (2012) The regulation of filamentous growth in yeast. Genetics 190(1):23–49
Cullen PJ, Schultz J, Horecka J, Stevenson BJ, Jigami Y, Sprague GF Jr (2000) Defects in protein glycosylation cause SHO1-dependent activation of a STE12 signaling pathway in yeast. Genetics 155(3):1005–1018
Davis-Hanna A, Piispanen AE, Stateva LI, Hogan DA (2008) Farnesol and dodecanol effects on the Candida albicans Ras1-cAMP signalling pathway and the regulation of morphogenesis. Mol Microbiol 67(1):47–62
de la Torre-Ruiz MA, Mozo-Villarias A, Pujol N, Petkova MI (2010) How budding yeast sense and transduce the oxidative stress signal and the impact in cell growth and morphogenesis. Curr Protein Pept Sci 11(8):669–679
de Nadal E, Alepuz PM, Posas F (2002) Dealing with osmostress through MAP kinase activation. EMBO Rep 3(8):735–740
de Nadal E, Real FX, Posas F (2007) Mucins, osmosensors in eukaryotic cells? Trends Cell Biol 17(12):571–574
DeRisi JL, Iyer VR, Brown PO (1997) Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278(5338):680–686
Desai PR, van Wijlick L, Kurtz D, Juchimiuk M, Ernst JF (2015) Hypoxia and Temperature Regulated Morphogenesis in Candida albicans. PLoS Genet 11(8):e1005447
Doedt T, Krishnamurthy S, Bockmuhl DP, Tebarth B, Stempel C, Russell CL et al (2004) APSES proteins regulate morphogenesis and metabolism in Candida albicans. Mol Biol Cell 15(7):3167–3180
Du LL, Novick P (2002) Pag1p, a novel protein associated with protein kinase Cbk1p, is required for cell morphogenesis and proliferation in Saccharomyces cerevisiae. Mol Biol Cell 13(2):503–514
Du C, Calderone R, Richert J, Li D (2005) Deletion of the SSK1 response regulator gene in Candida albicans contributes to enhanced killing by human polymorphonuclear neutrophils. Infect Immun 73(2):865–871
Eckert SE, Heinz WJ, Zakikhany K, Thewes S, Haynes K, Hube B et al (2007) PGA4, a GAS homologue from Candida albicans, is up-regulated early in infection processes. Fungal Genet Biol 44(5):368–377
Eisman B, Alonso-Monge R, Roman E, Arana D, Nombela C, Pla J (2006) The Cek1 and Hog1 mitogen-activated protein kinases play complementary roles in cell wall biogenesis and chlamydospore formation in the fungal pathogen Candida albicans. Eukaryot Cell 5(2):347–358
Elion EA (2000) Pheromone response, mating and cell biology. Curr Opin Microbiol 3(6):573–581
Enjalbert B, Nantel A, Whiteway M (2003) Stress-induced gene expression in Candida albicans: absence of a general stress response. Mol Biol Cell 14(4):1460–1467
Ernst JF (2000) Regulation of dimorphism in Candida albicans. Contrib Microbiol 5:98–111
Erwig LP, Gow NA (2016) Interactions of fungal pathogens with phagocytes. Nat Rev Microbiol 14(3):163–176
Fan Y, He H, Dong Y, Pan H (2013) Hyphae-specific genes HGC1, ALS3, HWP1, and ECE1 and relevant signaling pathways in Candida albicans. Mycopathologia 176(5–6):329–335
Fang HM, Wang Y (2006) RA domain-mediated interaction of Cdc35 with Ras1 is essential for increasing cellular cAMP level for Candida albicans hyphal development. Mol Microbiol 61(2):484–496
Fassler JS, West AH (2011) Fungal Skn7 stress responses and their relationship to virulence. Eukaryot Cell 10(2):156–167
Feng Q, Summers E, Guo B, Fink G (1999) Ras signaling is required for serum-induced hyphal differentiation in Candida albicans. J Bacteriol 181(20):6339–6346
Fernandes M, Xiao H, Lis JT (1995) Binding of heat shock factor to and transcriptional activation of heat shock genes in Drosophila. Nucleic Acids Res 23(23):4799–4804
Fonzi WA (1999) PHR1 and PHR2 of Candida albicans encode putative glycosidases required for proper cross-linking of beta-1,3- and beta-1,6-glucans. J Bacteriol 181(22):7070–7079
Fradin C, De Groot P, MacCallum D, Schaller M, Klis F, Odds FC et al (2005) Granulocytes govern the transcriptional response, morphology and proliferation of Candida albicans in human blood. Mol Microbiol 56(2):397–415
Fujita A, Kikuchi Y, Kuhara S, Misumi Y, Matsumoto S, Kobayashi H (1989) Domains of the SFL1 protein of yeasts are homologous to Myc oncoproteins or yeast heat-shock transcription factor. Gene 85(2):321–328
Fujita A, Tonouchi A, Hiroko T, Inose F, Nagashima T, Satoh R et al (1999) Hsl7p, a negative regulator of Ste20p protein kinase in the Saccharomyces cerevisiae filamentous growth-signaling pathway. Proc Natl Acad Sci U S A 96(15):8522–8527
Fuller KK, Rhodes JC (2012) Protein kinase A and fungal virulence: a sinister side to a conserved nutrient sensing pathway. Virulence 3(2):109–121
Galeote VA, Alexandre H, Bach B, Delobel P, Dequin S, Blondin B (2007) Sfl1p acts as an activator of the HSP30 gene in Saccharomyces cerevisiae. Curr Genet 52(2):55–63
Garcia-Rodriguez LJ, Valle R, Duran A, Roncero C (2005) Cell integrity signaling activation in response to hyperosmotic shock in yeast. FEBS Lett 579(27):6186–6190
Giacometti R, Kronberg F, Biondi RM, Passeron S (2009) Catalytic isoforms Tpk1 and Tpk2 of Candida albicans PKA have non-redundant roles in stress response and glycogen storage. Yeast 26(5):273–285
Giusani AD, Vinces M, Kumamoto CA (2002) Invasive filamentous growth of Candida albicans is promoted by Czf1p-dependent relief of Efg1p-mediated repression. Genetics 160(4):1749–1753
Gow NA (1997) Germ tube growth of Candida albicans. Curr Top Med Mycol 8(1–2):43–55
Gow NA, Brown AJ, Odds FC (2002) Fungal morphogenesis and host invasion. Curr Opin Microbiol 5(4):366–371
Grubb SE, Murdoch C, Sudbery PE, Saville SP, Lopez-Ribot JL, Thornhill MH (2009) Adhesion of Candida albicans to endothelial cells under physiological conditions of flow. Infect Immun 77(9):3872–3878
Hall RA, Turner KJ, Chaloupka J, Cottier F, De Sordi L, Sanglard D et al (2011) The quorum-sensing molecules farnesol/homoserine lactone and dodecanol operate via distinct modes of action in Candida albicans. Eukaryot Cell 10(8):1034–1042
Harashima T, Heitman J (2005) G subunit Gpa2 recruits kelch repeat subunits that inhibit receptor-G protein coupling during cAMP-induced dimorphic transitions in Saccharomyces cerevisiae. Mol Biol Cell 16(10):4557–4571
Helliwell SB, Howald I, Barbet N, Hall MN (1998) TOR2 is part of two related signaling pathways coordinating cell growth in Saccharomyces cerevisiae. Genetics 148(1):99–112
Hiller E, Zavrel M, Hauser N, Sohn K, Burger-Kentischer A, Lemuth K et al (2011) Adaptation, adhesion and invasion during interaction of Candida albicans with the host–focus on the function of cell wall proteins. Int J Med Microbiol 301(5):384–389
Hnisz D, Majer O, Frohner IE, Komnenovic V, Kuchler K (2010) The Set3/Hos2 histone deacetylase complex attenuates cAMP/PKA signaling to regulate morphogenesis and virulence of Candida albicans. PLoS Pathog 6(5):e1000889
Hnisz D, Bardet AF, Nobile CJ, Petryshyn A, Glaser W, Schock U et al (2012) A histone deacetylase adjusts transcription kinetics at coding sequences during Candida albicans morphogenesis. PLoS Genet 8(12):e1003118
Hogan DA, Sundstrom P (2009) The Ras/cAMP/PKA signaling pathway and virulence in Candida albicans. Future Microbiol. 4(10):1263–1270
Hogan DA, Vik A, Kolter R (2004) A Pseudomonas aeruginosa quorum-sensing molecule influences Candida albicans morphology. Mol Microbiol 54(5):1212–1223
Hohmann S (2002) Osmotic stress signaling and osmoadaptation in yeasts. Microbiol Mol Biol Rev 66(2):300–372
Hohmann S (2009) Control of high osmolarity signalling in the yeast Saccharomyces cerevisiae. FEBS Lett 583(24):4025–4029
Hope H, Bogliolo S, Arkowitz RA, Bassilana M (2008) Activation of Rac1 by the guanine nucleotide exchange factor Dck1 is required for invasive filamentous growth in the pathogen Candida albicans. Mol Biol Cell 19(9):3638–3651
Hope H, Schmauch C, Arkowitz RA, Bassilana M (2010) The Candida albicans ELMO homologue functions together with Rac1 and Dck1, upstream of the MAP Kinase Cek1, in invasive filamentous growth. Mol Microbiol 76(6):1572–1590
Hornby JM, Kebaara BW, Nickerson KW (2003) Farnesol biosynthesis in Candida albicans: cellular response to sterol inhibition by zaragozic acid B. Antimicrob Agents Chemother 47(7):2366–2369
Hoyer LL, Cieslinski LB, McLaughlin MM, Torphy TJ, Shatzman AR, Livi GP (1994) A Candida albicans cyclic nucleotide phosphodiesterase: cloning and expression in Saccharomyces cerevisiae and biochemical characterization of the recombinant enzyme. Microbiology 140(Pt 7):1533–1542
Hu Y, Liu E, Bai X, Zhang A (2010) The localization and concentration of the PDE2-encoded high-affinity cAMP phosphodiesterase is regulated by cAMP-dependent protein kinase A in the yeast Saccharomyces cerevisiae. FEMS Yeast Res 10(2):177–187
Hube B, Monod M, Schofield DA, Brown AJ, Gow NA (1994) Expression of seven members of the gene family encoding secretory aspartyl proteinases in Candida albicans. Mol Microbiol 14(1):87–99
Hwang CS, Oh JH, Huh WK, Yim HS, Kang SO (2003) Ssn6, an important factor of morphological conversion and virulence in Candida albicans. Mol Microbiol 47(4):1029–1043
Jarosz DF, Taipale M, Lindquist S (2010) Protein homeostasis and the phenotypic manifestation of genetic diversity: principles and mechanisms. Annu Rev Genet 44:189–216
Jenull S, Tscherner M, Gulati M, Nobile CJ, Chauhan N, Kuchler K (2017) The Candida albicans HIR histone chaperone regulates the yeast-to-hyphae transition by controlling the sensitivity to morphogenesis signals. Sci Rep 7(1):8308
Jung WH, Stateva LI (2003) The cAMP phosphodiesterase encoded by CaPDE2 is required for hyphal development in Candida albicans. Microbiology 149(Pt 10):2961–2976
Kadosh D, Johnson AD (2005) Induction of the Candida albicans filamentous growth program by relief of transcriptional repression: a genome-wide analysis. Mol Biol Cell 16(6):2903–2912
Keleher CA, Redd MJ, Schultz J, Carlson M, Johnson AD (1992) Ssn6-Tup1 is a general repressor of transcription in yeast. Cell 68(4):709–719
Kelly MT, MacCallum DM, Clancy SD, Odds FC, Brown AJ, Butler G (2004) The Candida albicans CaACE2 gene affects morphogenesis, adherence and virulence. Mol Microbiol 53(3):969–983
Khalaf RA, Zitomer RS (2001) The DNA binding protein Rfg1 is a repressor of filamentation in Candida albicans. Genetics 157(4):1503–1512
Kim TS, Lee SB, Kang HS (2004) Glucose repression of STA1 expression is mediated by the Nrg1 and Sfl1 repressors and the Srb8-11 complex. Mol Cell Biol 24(17):7695–7706
Klengel T, Liang WJ, Chaloupka J, Ruoff C, Schroppel K, Naglik JR et al (2005) Fungal adenylyl cyclase integrates CO2 sensing with cAMP signaling and virulence. Curr Biol 15(22):2021–2026
Kobayashi O, Suda H, Ohtani T, Sone H (1996) Molecular cloning and analysis of the dominant flocculation gene FLO8 from Saccharomyces cerevisiae. Mol Gen Genet 251(6):707–715
Kraakman L, Lemaire K, Ma P, Teunissen AW, Donaton MC, Van Dijck P et al (1999) A Saccharomyces cerevisiae G-protein coupled receptor, Gpr1, is specifically required for glucose activation of the cAMP pathway during the transition to growth on glucose. Mol Microbiol 32(5):1002–1012
Kruppa M, Jabra-Rizk MA, Meiller TF, Calderone R (2004) The histidine kinases of Candida albicans: regulation of cell wall mannan biosynthesis. FEMS Yeast Res 4(4–5):409–416
Kubler E, Mosch HU, Rupp S, Lisanti MP (1997) Gpa2p, a G-protein alpha-subunit, regulates growth and pseudohyphal development in Saccharomyces cerevisiae via a cAMP-dependent mechanism. J Biol Chem 272(33):20321–20323
Kullas AL, Li M, Davis DA (2004) Snf7p, a component of the ESCRT-III protein complex, is an upstream member of the RIM101 pathway in Candida albicans. Eukaryot Cell 3(6):1609–1618
Kullberg BJ, Arendrup MC (2016) Invasive candidiasis. N Engl J Med 374(8):794–795
Kumamoto CA (2005) A contact-activated kinase signals Candida albicans invasive growth and biofilm development. Proc Natl Acad Sci USA 102(15):5576–5581
Kumamoto CA (2008) Molecular mechanisms of mechanosensing and their roles in fungal contact sensing. Nat Rev Microbiol 6(9):667–673
Kumamoto CA, Vinces MD (2005) Alternative Candida albicans lifestyles: growth on surfaces. Annu Rev Microbiol 59:113–133
Kunz J, Henriquez R, Schneider U, Deuter-Reinhard M, Movva NR, Hall MN (1993) Target of rapamycin in yeast, TOR2, is an essential phosphatidylinositol kinase homolog required for G1 progression. Cell 73(3):585–596
Lane S, Birse C, Zhou S, Matson R, Liu H (2001a) DNA array studies demonstrate convergent regulation of virulence factors by Cph1, Cph2, and Efg1 in Candida albicans. J Biol Chem 276(52):48988–48996
Lane S, Zhou S, Pan T, Dai Q, Liu H (2001b) The basic helix-loop-helix transcription factor Cph2 regulates hyphal development in Candida albicans partly via TEC1. Mol Cell Biol 21(19):6418–6428
Lassak T, Schneider E, Bussmann M, Kurtz D, Manak JR, Srikantha T et al (2011) Target specificity of the Candida albicans Efg1 regulator. Mol Microbiol 82(3):602–618
Leach MD, Tyc KM, Brown AJ, Klipp E (2012) Modelling the regulation of thermal adaptation in Candida albicans, a major fungal pathogen of humans. PLoS ONE 7(3):e32467
Leach MD, Farrer RA, Tank K, Miao Z, Walker LA, Cuomo CA et al (2016) Hsf1 and Hsp90 orchestrate temperature-dependent global transcriptional remodelling and chromatin architecture in Candida albicans. Nat Commun. 26(7):11704
Leberer E, Harcus D, Broadbent ID, Clark KL, Dignard D, Ziegelbauer K et al (1996) Signal transduction through homologs of the Ste20p and Ste7p protein kinases can trigger hyphal formation in the pathogenic fungus Candida albicans. Proc Natl Acad Sci U S A 93(23):13217–13222
Leberer E, Harcus D, Dignard D, Johnson L, Ushinsky S, Thomas DY et al (2001) Ras links cellular morphogenesis to virulence by regulation of the MAP kinase and cAMP signalling pathways in the pathogenic fungus Candida albicans. Mol Microbiol 42(3):673–687
Lee BN, Elion EA (1999) The MAPKKK Ste11 regulates vegetative growth through a kinase cascade of shared signaling components. Proc Natl Acad Sci U S A 96(22):12679–12684
Lee JE, Oh JH, Ku M, Kim J, Lee JS, Kang SO (2015) Ssn6 has dual roles in Candida albicans filament development through the interaction with Rpd31. FEBS Lett 589(4):513–520
Levin DE (2005) Cell wall integrity signaling in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 69(2):262–291
Li D, Gurkovska V, Sheridan M, Calderone R, Chauhan N (2004a) Studies on the regulation of the two-component histidine kinase gene CHK1 in Candida albicans using the heterologous lacZ reporter gene. Microbiology 150(Pt 10):3305–3313
Li M, Martin SJ, Bruno VM, Mitchell AP, Davis DA (2004b) Candida albicans Rim13p, a protease required for Rim101p processing at acidic and alkaline pHs. Eukaryot Cell 3(3):741–751
Li Y, Su C, Mao X, Cao F, Chen J (2007) Roles of Candida albicans Sfl1 in hyphal development. Eukaryot Cell 6(11):2112–2121
Liu H, Kohler J, Fink GR (1994) Suppression of hyphal formation in Candida albicans by mutation of a STE12 homolog. Science 266(5191):1723–1726
Liu W, Zhao J, Li X, Li Y, Jiang L (2010) The protein kinase CaSch9p is required for the cell growth, filamentation and virulence in the human fungal pathogen Candida albicans. FEMS Yeast Res 10(4):462–470
Lo WS, Dranginis AM (1998) The cell surface flocculin Flo11 is required for pseudohyphae formation and invasion by Saccharomyces cerevisiae. Mol Biol Cell 9(1):161–171
Lo HJ, Kohler JR, DiDomenico B, Loebenberg D, Cacciapuoti A, Fink GR (1997) Nonfilamentous C. albicans mutants are avirulent. Cell 90(5):939–49
Loewith R, Jacinto E, Wullschleger S, Lorberg A, Crespo JL, Bonenfant D et al (2002) Two TOR complexes, only one of which is rapamycin sensitive, have distinct roles in cell growth control. Mol Cell 10(3):457–468
Lorenz MC, Heitman J (1997) Yeast pseudohyphal growth is regulated by GPA2, a G protein alpha homolog. EMBO J 16(23):7008–7018
Lorenz MC, Pan X, Harashima T, Cardenas ME, Xue Y, Hirsch JP et al (2000) The G protein-coupled receptor gpr1 is a nutrient sensor that regulates pseudohyphal differentiation in Saccharomyces cerevisiae. Genetics 154(2):609–622
Lorenz MC, Bender JA, Fink GR (2004) Transcriptional response of Candida albicans upon internalization by macrophages. Eukaryot Cell 3(5):1076–1087
Lu Y, Su C, Mao X, Raniga PP, Liu H, Chen J (2008) Efg1-mediated recruitment of NuA4 to promoters is required for hypha-specific Swi/Snf binding and activation in Candida albicans. Mol Biol Cell 19(10):4260–4272
Lu Y, Su C, Wang A, Liu H (2011) Hyphal development in Candida albicans requires two temporally linked changes in promoter chromatin for initiation and maintenance. PLoS Biol 9(7):e1001105
Lu Y, Su C, Liu H (2012) A GATA transcription factor recruits Hda1 in response to reduced Tor1 signaling to establish a hyphal chromatin state in Candida albicans. PLoS Pathog 8(4):e1002663
Lu Y, Su C, Solis NV, Filler SG, Liu H (2013) Synergistic regulation of hyphal elongation by hypoxia, CO2, and nutrient conditions controls the virulence of Candida albicans. Cell Host Microbe 14(5):499–509
Lu Y, Su C, Liu H (2014) Candida albicans hyphal initiation and elongation. Trends Microbiol 22(12):707–714
Ma P, Wera S, Van Dijck P, Thevelein JM (1999) The PDE1-encoded low-affinity phosphodiesterase in the yeast Saccharomyces cerevisiae has a specific function in controlling agonist-induced cAMP signaling. Mol Biol Cell 10(1):91–104
MacIsaac KD, Wang T, Gordon DB, Gifford DK, Stormo GD, Fraenkel E (2006) An improved map of conserved regulatory sites for Saccharomyces cerevisiae. BMC Bioinformatics 7:113
Madhani HD, Fink GR (1997) Combinatorial control required for the specificity of yeast MAPK signaling. Science 275(5304):1314–1317
Madhani HD, Styles CA, Fink GR (1997) MAP kinases with distinct inhibitory functions impart signaling specificity during yeast differentiation. Cell 91(5):673–684
Maeda T, Takekawa M, Saito H (1995) Activation of yeast PBS2 MAPKK by MAPKKKs or by binding of an SH3-containing osmosensor. Science 269(5223):554–558
Magee BB, Legrand M, Alarco AM, Raymond M, Magee PT (2002) Many of the genes required for mating in Saccharomyces cerevisiae are also required for mating in Candida albicans. Mol Microbiol 46(5):1345–1351
Maidan MM, Thevelein JM, Van Dijck P (2005) Carbon source induced yeast-to-hypha transition in Candida albicans is dependent on the presence of amino acids and on the G-protein-coupled receptor Gpr1. Biochem Soc Trans 33(Pt 1):291–293
Mao X, Cao F, Nie X, Liu H, Chen J (2006) The Swi/Snf chromatin remodeling complex is essential for hyphal development in Candida albicans. FEBS Lett 580(11):2615–2622
Marcil A, Harcus D, Thomas DY, Whiteway M. Candida albicans killing by RAW 264.7 mouse macrophage cells: effects of Candida genotype, infection ratios, and gamma interferon treatment. Infect Immun. 2002;70(11):6319–29
Martin R, Moran GP, Jacobsen ID, Heyken A, Domey J, Sullivan DJ et al (2011) The Candida albicans-specific gene EED1 encodes a key regulator of hyphal extension. PLoS ONE 6(4):e18394
Mattia E, Carruba G, Angiolella L, Cassone A (1982) Induction of germ tube formation by N-acetyl-D-glucosamine in Candida albicans: uptake of inducer and germinative response. J Bacteriol 152(2):555–562
Mazanka E, Alexander J, Yeh BJ, Charoenpong P, Lowery DM, Yaffe M et al (2008) The NDR/LATS family kinase Cbk1 directly controls transcriptional asymmetry. PLoS Biol 6(8):e203
McNemar MD, Fonzi WA (2002) Conserved serine/threonine kinase encoded by CBK1 regulates expression of several hypha-associated transcripts and genes encoding cell wall proteins in Candida albicans. J Bacteriol 184(7):2058–2061
Mendelsohn S, Pinsky M, Weissman Z, Kornitzer D (2017) Regulation of the Candida albicans hypha-inducing transcription factor Ume6 by the CDK1 Cyclins Cln3 and Hgc1. mSpehe 2(2):e00248-16
Messenguy F, Vierendeels F, Scherens B, Dubois E (2000) In Saccharomyces cerevisiae, expression of arginine catabolic genes CAR1 and CAR2 in response to exogenous nitrogen availability is mediated by the Ume6 (CargRI)-Sin3 (CargRII)-Rpd3 (CargRIII) complex. J Bacteriol 182(11):3158–3164
Minato T, Wang J, Akasaka K, Okada T, Suzuki N, Kataoka T (1994) Quantitative analysis of mutually competitive binding of human Raf-1 and yeast adenylyl cyclase to Ras proteins. J Biol Chem 269(33):20845–20851
Miwa T, Takagi Y, Shinozaki M, Yun CW, Schell WA, Perfect JR et al (2004) Gpr1, a putative G-protein-coupled receptor, regulates morphogenesis and hypha formation in the pathogenic fungus Candida albicans. Eukaryot Cell 3(4):919–931
Monge RA, Roman E, Nombela C, Pla J (2006) The MAP kinase signal transduction network in Candida albicans. Microbiology 152(Pt 4):905–912
Morozov AV, Siggia ED (2007) Connecting protein structure with predictions of regulatory sites. Proc Natl Acad Sci U S A 104(17):7068–7073
Moyes DL, Wilson D, Richardson JP, Mogavero S, Tang SX, Wernecke J et al (2016) Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 532(7597):64–68
Muhlschlegel FA, Fonzi WA (1997) PHR2 of Candida albicans encodes a functional homolog of the pH-regulated gene PHR1 with an inverted pattern of pH-dependent expression. Mol Cell Biol 17(10):5960–5967
Mukaremera L, Lee KK, Mora-Montes HM, Gow NAR (2017) Candida albicans yeast, pseudohyphal, and hyphal morphogenesis differentially affects immune recognition. Front Immunol 8:629
Murad AM, Leng P, Straffon M, Wishart J, Macaskill S, MacCallum D et al (2001) NRG1 represses yeast-hypha morphogenesis and hypha-specific gene expression in Candida albicans. EMBO J 20(17):4742–4752
Nagahashi S, Mio T, Ono N, Yamada-Okabe T, Arisawa M, Bussey H et al (1998) Isolation of CaSLN1 and CaNIK1, the genes for osmosensing histidine kinase homologues, from the pathogenic fungus Candida albicans. Microbiology 144(Pt 2):425–432
Nair R, Shariq M, Dhamgaye S, Mukhopadhyay CK, Shaikh S, Prasad R (2017) Non-heat shock responsive roles of HSF1 in Candida albicans are essential under iron deprivation and drug defense. Biochim Biophys Acta 1864(2):345–354
Nantel A, Dignard D, Bachewich C, Harcus D, Marcil A, Bouin AP et al (2002) Transcription profiling of Candida albicans cells undergoing the yeast-to-hyphal transition. Mol Biol Cell 13(10):3452–3465
Naseem S, Araya E, Konopka JB (2015) Hyphal growth in Candida albicans does not require induction of hyphal-specific gene expression. Mol Biol Cell 26(6):1174–1187
Navarro-Garcia F, Sanchez M, Pla J, Nombela C (1995) Functional characterization of the MKC1 gene of Candida albicans, which encodes a mitogen-activated protein kinase homolog related to cell integrity. Mol Cell Biol 15(4):2197–2206
Navarro-Garcia F, Alonso-Monge R, Rico H, Pla J, Sentandreu R, Nombela C (1998) A role for the MAP kinase gene MKC1 in cell wall construction and morphological transitions in Candida albicans. Microbiology 144(Pt 2):411–424
Nelson B, Kurischko C, Horecka J, Mody M, Nair P, Pratt L et al (2003) RAM: a conserved signaling network that regulates Ace2p transcriptional activity and polarized morphogenesis. Mol Biol Cell 14(9):3782–3803
Nemecek JC, Wuthrich M, Klein BS (2006) Global control of dimorphism and virulence in fungi. Science 312(5773):583–588
Nicholls S, Leach MD, Priest CL, Brown AJ (2009) Role of the heat shock transcription factor, Hsf1, in a major fungal pathogen that is obligately associated with warm-blooded animals. Mol Microbiol 74(4):844–861
Nicholls S, MacCallum DM, Kaffarnik FA, Selway L, Peck SC, Brown AJ (2011) Activation of the heat shock transcription factor Hsf1 is essential for the full virulence of the fungal pathogen Candida albicans. Fungal Genet Biol 48(3):297–305
Nobile CJ, Mitchell AP (2005) Regulation of cell-surface genes and biofilm formation by the C. albicans transcription factor Bcr1p. Curr Biol 15(12):1150–5
Noble SM, French S, Kohn LA, Chen V, Johnson AD (2010) Systematic screens of a Candida albicans homozygous deletion library decouple morphogenetic switching and pathogenicity. Nat Genet 42(7):590–598
Odds FC (1987) Candida infections: an overview. Crit Rev Microbiol 15(1):1–5
Odds FC (1988) Candida and Candidosis: a review and bibliography, 2nd edn
Ollert MW, Sohnchen R, Korting HC, Ollert U, Brautigam S, Brautigam W (1993) Mechanisms of adherence of Candida albicans to cultured human epidermal keratinocytes. Infect Immun 61(11):4560–4568
O’Meara TR, Robbins N, Cowen LE (2017) The Hsp90 chaperone network modulates Candida virulence traits. Trends Microbiol 25(10):809–819
O’Rourke SM, Herskowitz I (1998) The Hog1 MAPK prevents cross talk between the HOG and pheromone response MAPK pathways in Saccharomyces cerevisiae. Genes Dev 12(18):2874–2886
Palecek SP, Parikh AS, Kron SJ (2002) Sensing, signalling and integrating physical processes during Saccharomyces cerevisiae invasive and filamentous growth. Microbiology 148(Pt 4):893–907
Pan X, Heitman J (1999) Cyclic AMP-dependent protein kinase regulates pseudohyphal differentiation in Saccharomyces cerevisiae. Mol Cell Biol 19(7):4874–4887
Pan X, Heitman J (2002) Protein kinase A operates a molecular switch that governs yeast pseudohyphal differentiation. Mol Cell Biol 22(12):3981–3993
Pande K, Chen C, Noble SM (2013) Passage through the mammalian gut triggers a phenotypic switch that promotes Candida albicans commensalism. Nat Genet 45(9):1088–1091
Paravicini G, Mendoza A, Antonsson B, Cooper M, Losberger C, Payton MA (1996) The Candida albicans PKC1 gene encodes a protein kinase C homolog necessary for cellular integrity but not dimorphism. Yeast 12(8):741–756
Parrino SM, Si H, Naseem S, Groudan K, Gardin J, Konopka JB (2017) cAMP-independent signal pathways stimulate hyphal morphogenesis in Candida albicans. Mol Microbiol 103(5):764–779
Penalva MA, Arst HN, Jr (2002) Regulation of gene expression by ambient pH in filamentous fungi and yeasts. Microbiol Mol Biol Rev 66(3):426–46 (table of contents)
Perez JC, Kumamoto CA, Johnson AD (2013) Candida albicans commensalism and pathogenicity are intertwined traits directed by a tightly knit transcriptional regulatory circuit. PLoS Biol 11(3):e1001510
Persi MA, Burnham JC, Duhring JL (1985) Effects of carbon dioxide and pH on adhesion of Candida albicans to vaginal epithelial cells. Infect Immun 50(1):82–90
Peters BM, Palmer GE, Nash AK, Lilly EA, Fidel PL Jr, Noverr MC (2014) Fungal morphogenetic pathways are required for the hallmark inflammatory response during Candida albicans vaginitis. Infect Immun 82(2):532–543
Pfaller MA, Diekema DJ (2007) Epidemiology of invasive candidiasis: a persistent public health problem. Clin Microbiol Rev 20(1):133–163
Phan QT, Myers CL, Fu Y, Sheppard DC, Yeaman MR, Welch WH et al (2007) Als3 is a Candida albicans invasin that binds to cadherins and induces endocytosis by host cells. PLoS Biol 5(3):e64
Polke M, Sprenger M, Scherlach K, Alban-Proano MC, Martin R, Hertweck C et al (2017) A functional link between hyphal maintenance and quorum sensing in Candida albicans. Mol Microbiol 103(4):595–617
Porta A, Ramon AM, Fonzi WA (1999) PRR1, a homolog of Aspergillus nidulans palF, controls pH-dependent gene expression and filamentation in Candida albicans. J Bacteriol 181(24):7516–7523
Posas F, Saito H (1998) Activation of the yeast SSK2 MAP kinase kinase kinase by the SSK1 two-component response regulator. EMBO J 17(5):1385–1394
Posas F, Wurgler-Murphy SM, Maeda T, Witten EA, Thai TC, Saito H (1996) Yeast HOG1 MAP kinase cascade is regulated by a multistep phosphorelay mechanism in the SLN1-YPD1-SSK1 “two-component” osmosensor. Cell 86(6):865–875
Posas F, Chambers JR, Heyman JA, Hoeffler JP, de Nadal E, Arino J (2000) The transcriptional response of yeast to saline stress. J Biol Chem 275(23):17249–17255
Ramon AM, Fonzi WA (2003) Diverged binding specificity of Rim101p, the Candida albicans ortholog of PacC. Eukaryot Cell 2(4):718–728
Ramon AM, Porta A, Fonzi WA (1999) Effect of environmental pH on morphological development of Candida albicans is mediated via the PacC-related transcription factor encoded by PRR2. J Bacteriol 181(24):7524–7530
Reinke A, Anderson S, McCaffery JM, Yates J 3rd, Aronova S, Chu S et al (2004) TOR complex 1 includes a novel component, Tco89p (YPL180w), and cooperates with Ssd1p to maintain cellular integrity in Saccharomyces cerevisiae. J Biol Chem 279(15):14752–14762
Reiser V, Salah SM, Ammerer G (2000) Polarized localization of yeast Pbs2 depends on osmostress, the membrane protein Sho1 and Cdc42. Nat Cell Biol 2(9):620–627
Richter K, Haslbeck M, Buchner J (2010) The heat shock response: life on the verge of death. Mol Cell 40(2):253–266
Riggle PJ, Andrutis KA, Chen X, Tzipori SR, Kumamoto CA (1999) Invasive lesions containing filamentous forms produced by a Candida albicans mutant that is defective in filamentous growth in culture. Infect Immun 67(7):3649–3652
Robertson LS, Fink GR (1998) The three yeast A kinases have specific signaling functions in pseudohyphal growth. Proc Natl Acad Sci U S A 95(23):13783–13787
Rohde JR, Cardenas ME (2004) Nutrient signaling through TOR kinases controls gene expression and cellular differentiation in fungi. Curr Top Microbiol Immunol 279:53–72
Rohde JR, Bastidas R, Puria R, Cardenas ME (2008) Nutritional control via Tor signaling in Saccharomyces cerevisiae. Curr Opin Microbiol 11(2):153–160
Roman E, Nombela C, Pla J (2005) The Sho1 adaptor protein links oxidative stress to morphogenesis and cell wall biosynthesis in the fungal pathogen Candida albicans. Mol Cell Biol 25(23):10611–10627
Rooney PJ, Klein BS (2002) Linking fungal morphogenesis with virulence. Cell Microbiol 4(3):127–137
Rubin-Bejerano I, Mandel S, Robzyk K, Kassir Y (1996) Induction of meiosis in Saccharomyces cerevisiae depends on conversion of the transcriptional represssor Ume6 to a positive regulator by its regulated association with the transcriptional activator Ime1. Mol Cell Biol 16(5):2518–2526
Rupp S, Summers E, Lo HJ, Madhani H, Fink G (1999) MAP kinase and cAMP filamentation signaling pathways converge on the unusually large promoter of the yeast FLO11 gene. EMBO J 18(5):1257–1269
San Jose C, Monge RA, Perez-Diaz R, Pla J, Nombela C (1996) The mitogen-activated protein kinase homolog HOG1 gene controls glycerol accumulation in the pathogenic fungus Candida albicans. J Bacteriol 178(19):5850–5852
Park H, Myers CL, Sheppard DC, Phan QT, Sanchez AA, J EE et al (2005) Role of the fungal Ras-protein kinase A pathway in governing epithelial cell interactions during oropharyngeal candidiasis. Cell Microbiol 7(4):499–510
Sanglard D, Hube B, Monod M, Odds FC, Gow NA (1997) A triple deletion of the secreted aspartyl proteinase genes SAP4, SAP5, and SAP6 of Candida albicans causes attenuated virulence. Infect Immun 65(9):3539–3546
Sato T, Watanabe T, Mikami T, Matsumoto T (2004) Farnesol, a morphogenetic autoregulatory substance in the dimorphic fungus Candida albicans, inhibits hyphae growth through suppression of a mitogen-activated protein kinase cascade. Biol Pharm Bull 27(5):751–752
Saville SP, Lazzell AL, Monteagudo C, Lopez-Ribot JL (2003) Engineered control of cell morphology in vivo reveals distinct roles for yeast and filamentous forms of Candida albicans during infection. Eukaryot Cell 2(5):1053–1060
Saville SP, Lazzell AL, Bryant AP, Fretzen A, Monreal A, Solberg EO et al (2006) Inhibition of filamentation can be used to treat disseminated candidiasis. Antimicrob Agents Chemother 50(10):3312–3316
Schmelzle T, Hall MN (2000) TOR, a central controller of cell growth. Cell 103(2):253–262
Schrick K, Garvik B, Hartwell LH (1997) Mating in Saccharomyces cerevisiae: the role of the pheromone signal transduction pathway in the chemotropic response to pheromone. Genetics 147(1):19–32
Schweizer A, Rupp S, Taylor BN, Rollinghoff M, Schroppel K (2000) The TEA/ATTS transcription factor CaTec1p regulates hyphal development and virulence in Candida albicans. Mol Microbiol 38(3):435–445
Sentandreu M, Elorza MV, Sentandreu R, Fonzi WA (1998) Cloning and characterization of PRA1, a gene encoding a novel pH-regulated antigen of Candida albicans. J Bacteriol 180(2):282–289
Setiadi ER, Doedt T, Cottier F, Noffz C, Ernst JF (2006) Transcriptional response of Candida albicans to hypoxia: linkage of oxygen sensing and Efg1p-regulatory networks. J Mol Biol 361(3):399–411
Shapiro RS, Uppuluri P, Zaas AK, Collins C, Senn H, Perfect JR et al (2009) Hsp90 orchestrates temperature-dependent Candida albicans morphogenesis via Ras1-PKA signaling. Curr Biol 19(8):621–629
Singh P, Chauhan N, Ghosh A, Dixon F, Calderone R (2004) SKN7 of Candida albicans: mutant construction and phenotype analysis. Infect Immun 72(4):2390–2394
Smith RL, Johnson AD (2000) Turning genes off by Ssn6-Tup1: a conserved system of transcriptional repression in eukaryotes. Trends Biochem Sci 25(7):325–330
Song W, Carlson M (1998) Srb/mediator proteins interact functionally and physically with transcriptional repressor Sfl1. EMBO J 17(19):5757–5765
Song Y, Cheon SA, Lee KE, Lee SY, Lee BK, Oh DB et al (2008) Role of the RAM network in cell polarity and hyphal morphogenesis in Candida albicans. Mol Biol Cell 19(12):5456–5477
Song W, Wang H, Chen J (2011) Candida albicans Sfl2, a temperature-induced transcriptional regulator, is required for virulence in a murine gastrointestinal infection model. FEMS Yeast Res 11(2):209–222
Sonneborn A, Bockmuhl DP, Gerads M, Kurpanek K, Sanglard D, Ernst JF (2000) Protein kinase A encoded by TPK2 regulates dimorphism of Candida albicans. Mol Microbiol 35(2):386–396
Sorger PK, Pelham HR (1987) Purification and characterization of a heat-shock element binding protein from yeast. EMBO J 6(10):3035–3041
Spiering MJ, Moran GP, Chauvel M, Maccallum DM, Higgins J, Hokamp K et al (2010) Comparative transcript profiling of Candida albicans and Candida dubliniensis identifies SFL2, a C. albicans gene required for virulence in a reconstituted epithelial infection model. Eukaryot Cell. 9(2):251–65
Srikantha T, Tsai L, Daniels K, Enger L, Highley K, Soll DR (1998) The two-component hybrid kinase regulator CaNIK1 of Candida albicans. Microbiology 144(Pt 10):2715–2729
Stichternoth C, Fraund A, Setiadi E, Giasson L, Vecchiarelli A, Ernst JF (2011) Sch9 kinase integrates hypoxia and CO2 sensing to suppress hyphal morphogenesis in Candida albicans. Eukaryot Cell 10(4):502–511
Stoldt VR, Sonneborn A, Leuker CE, Ernst JF (1997) Efg1p, an essential regulator of morphogenesis of the human pathogen Candida albicans, is a member of a conserved class of bHLH proteins regulating morphogenetic processes in fungi. EMBO J 16(8):1982–91
Su C, Lu Y, Liu H (2013) Reduced TOR signaling sustains hyphal development in Candida albicans by lowering Hog1 basal activity. Mol Biol Cell 24(3):385–397
Sudbery PE (2011) Growth of Candida albicans hyphae. Nat Rev Microbiol 9(10):737–748
Sudbery P, Gow N, Berman J (2004) The distinct morphogenic states of Candida albicans. Trends Microbiol 12(7):317–324
Sweet DH, Jang YK, Sancar GB (1997) Role of UME6 in transcriptional regulation of a DNA repair gene in Saccharomyces cerevisiae. Mol Cell Biol 17(11):6223–6235
Swoboda RK, Bertram G, Budge S, Gooday GW, Gow NA, Brown AJ (1995) Structure and regulation of the HSP90 gene from the pathogenic fungus Candida albicans. Infect Immun 63(11):4506–4514
Tatebayashi K, Tanaka K, Yang HY, Yamamoto K, Matsushita Y, Tomida T et al (2007) Transmembrane mucins Hkr1 and Msb2 are putative osmosensors in the SHO1 branch of yeast HOG pathway. EMBO J 26(15):3521–3533
Tebbji F, Chen Y, Sellam A, Whiteway M (2017) The genomic landscape of the fungus-specific SWI/SNF complex subunit, Snf6, in Candida albicans. mSphere 2(6)
Thevelein JM, de Winde JH (1999) Novel sensing mechanisms and targets for the cAMP-protein kinase A pathway in the yeast Saccharomyces cerevisiae. Mol Microbiol 33(5):904–918
Toda T, Cameron S, Sass P, Zoller M, Scott JD, McMullen B et al (1987a) Cloning and characterization of BCY1, a locus encoding a regulatory subunit of the cyclic AMP-dependent protein kinase in Saccharomyces cerevisiae. Mol Cell Biol 7(4):1371–1377
Toda T, Cameron S, Sass P, Zoller M, Wigler M (1987) Three different genes in S. cerevisiae encode the catalytic subunits of the cAMP-dependent protein kinase. Cell. 50(2):277–87
Trevijano-Contador N, Rueda C, Zaragoza O (2016) Fungal morphogenetic changes inside the mammalian host. Semin Cell Dev Biol 57:100–109
Tripathi G, Wiltshire C, Macaskill S, Tournu H, Budge S, Brown AJ (2002) Gcn4 co-ordinates morphogenetic and metabolic responses to amino acid starvation in Candida albicans. EMBO J 21(20):5448–5456
Tyc KM, Herwald SE, Hogan JA, Pierce JV, Klipp E, Kumamoto CA (2016) The game theory of Candida albicans colonization dynamics reveals host status-responsive gene expression. BMC Syst Biol 10:20
Veri AO, Miao Z, Shapiro RS, Tebbji F, O’Meara TR, Kim SH et al (2018) Tuning Hsf1 levels drives distinct fungal morphogenetic programs with depletion impairing Hsp90 function and overexpression expanding the target space. PLoS Genet 14(3):e1007270
Vezina C, Kudelski A, Sehgal SN (1975) Rapamycin (AY-22,989), a new antifungal antibiotic. I. Taxonomy of the producing streptomycete and isolation of the active principle. J Antibiot (Tokyo) 28(10):721–6
Vilella F, Herrero E, Torres J, de la Torre-Ruiz MA (2005) Pkc1 and the upstream elements of the cell integrity pathway in Saccharomyces cerevisiae, Rom2 and Mtl1, are required for cellular responses to oxidative stress. J Biol Chem 280(10):9149–9159
Vinces MD, Haas C, Kumamoto CA (2006) Expression of the Candida albicans morphogenesis regulator gene CZF1 and its regulation by Efg1p and Czf1p. Eukaryot Cell 5(5):825–835
Vylkova S, Carman AJ, Danhof HA, Collette JR, Zhou H, Lorenz MC (2011) The fungal pathogen Candida albicans autoinduces hyphal morphogenesis by raising extracellular pH. MBio 2(3):e00055–11
Wang Y, Cao YY, Jia XM, Cao YB, Gao PH, Fu XP et al (2006) Cap1p is involved in multiple pathways of oxidative stress response in Candida albicans. Free Radic Biol Med 40(7):1201–1209
Weiss EL, Kurischko C, Zhang C, Shokat K, Drubin DG, Luca FC (2002) The Saccharomyces cerevisiae Mob2p-Cbk1p kinase complex promotes polarized growth and acts with the mitotic exit network to facilitate daughter cell-specific localization of Ace2p transcription factor. J Cell Biol 158(5):885–900
White SJ, Rosenbach A, Lephart P, Nguyen D, Benjamin A, Tzipori S et al (2007) Self-regulation of Candida albicans population size during GI colonization. PLoS Pathog 3(12):e184
Whiteway M, Bachewich C (2007) Morphogenesis in Candida albicans. Annu Rev Microbiol 61:529–553
Whiteway M, Dignard D, Thomas DY (1992) Dominant negative selection of heterologous genes: isolation of Candida albicans genes that interfere with Saccharomyces cerevisiae mating factor-induced cell cycle arrest. Proc Natl Acad Sci U S A 89(20):9410–9414
Wiederrecht G, Shuey DJ, Kibbe WA, Parker CS (1987) The Saccharomyces and Drosophila heat shock transcription factors are identical in size and DNA binding properties. Cell 48(3):507–515
Wullschleger S, Loewith R, Oppliger W, Hall MN (2005) Molecular organization of target of rapamycin complex 2. J Biol Chem 280(35):30697–30704
Xu XL, Lee RT, Fang HM, Wang YM, Li R, Zou H et al (2008) Bacterial peptidoglycan triggers Candida albicans hyphal growth by directly activating the adenylyl cyclase Cyr1p. Cell Host Microbe 4(1):28–39
Xue Y, Batlle M, Hirsch JP (1998) GPR1 encodes a putative G protein-coupled receptor that associates with the Gpa2p G subunit and functions in a Ras-independent pathway. EMBO J 17(7):1996–2007
Yang W, Yan L, Wu C, Zhao X, Tang J (2014) Fungal invasion of epithelial cells. Microbiol Res 169(11):803–810
Zakikhany K, Naglik JR, Schmidt-Westhausen A, Holland G, Schaller M, Hube B (2007) In vivo transcript profiling of Candida albicans identifies a gene essential for interepithelial dissemination. Cell Microbiol 9(12):2938–2954
Zeidler U, Lettner T, Lassnig C, Muller M, Lajko R, Hintner H et al (2009) UME6 is a crucial downstream target of other transcriptional regulators of true hyphal development in Candida albicans. FEMS Yeast Res 9(1):126–142
Zhang X, De Micheli M, Coleman ST, Sanglard D, Moye-Rowley WS (2000) Analysis of the oxidative stress regulation of the Candida albicans transcription factor, Cap1p. Mol Microbiol 36(3):618–629
Zheng X, Wang Y, Wang Y (2004) Hgc1, a novel hypha-specific G1 cyclin-related protein regulates Candida albicans hyphal morphogenesis. EMBO J 23(8):1845–1856
Zhu C, Byers KJ, McCord RP, Shi Z, Berger MF, Newburger DE et al (2009) High-resolution DNA-binding specificity analysis of yeast transcription factors. Genome Res 19(4):556–566
Znaidi S, Barker KS, Weber S, Alarco AM, Liu TT, Boucher G et al (2009) Identification of the Candida albicans Cap1p regulon. Eukaryot Cell 8(6):806–820
Znaidi S, Nesseir A, Chauvel M, Rossignol T, d’Enfert C (2013) A comprehensive functional portrait of two heat shock factor-type transcriptional regulators involved in Candida albicans morphogenesis and virulence. PLoS Pathog 9(8):e1003519
Acknowledgements
This work has been supported by grants from the Agence Nationale de la Recherche (CANDIHUB, ANR-14-CE-0018), the French Government’s Investissement d’Avenir program (Laboratoire d’Excellence Integrative Biology of Emerging Infectious Diseases, ANR-10-LABX-62-IBEID), the European Commission (FinSysB PITN-GA-2008-214004), and the Wellcome Trust (The Candida albicans ORFeome project, WT088858MA). S.Z. is an Institut Pasteur International Network Affiliate Program Fellow. S.Z. was the recipient of postdoctoral fellowships from the European Commission (FINSysB, PITN-GA-2008-214004), the Agence Nationale de la Recherche (KANJI, ANR-08-MIE-033-01), and the French Government’s Investissement d’Avenir program (Institut de Recherche Technologique BIOASTER, ANR-10-AIRT-03). V.B. was supported by a grant from the Pasteur-Paris University (PPU) International PhD program and the “Fondation Daniel et Nina Carasso.”
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Basso, V., d’Enfert, C., Znaidi, S., Bachellier-Bassi, S. (2018). From Genes to Networks: The Regulatory Circuitry Controlling Candida albicans Morphogenesis. In: Rodrigues, M. (eds) Fungal Physiology and Immunopathogenesis . Current Topics in Microbiology and Immunology, vol 422. Springer, Cham. https://doi.org/10.1007/82_2018_144
Download citation
DOI: https://doi.org/10.1007/82_2018_144
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-30236-8
Online ISBN: 978-3-030-30237-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)