Abstract
Abstract
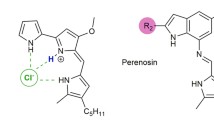
The red-colored prodiginines, exemplified by prodigiosin 1, are secondary metabolites produced by a number of microorganisms, including the bacterium Serratia marcescens. These tripyrrole natural products and their synthetic analogs have received renewed attention over the past deacade, primarily because of their promising immunosuppressive and anticancer activities. One of the hallmarks of prodiginin chemistry is the ability of the monoprotonated ligand to bind anions, including the essential chloride and bicarbonate ions. The resulting lipophilic ion pair is then able to diffuse across the hydrophobic barrier presented by phospholipid bilayers. Thus, prodiginines have been found to be potent transmembrane anion transporters and HCl cotransporters. In this chapter, the author reviews what is known about the solid-state structure of prodiginins and their anion complexes, the solution conformation of prodiginines, and the biochemcal evidence for the ability to bind anions and to transport HCl across cell membranes. Recent progress in making synthetic models of prodiginines and recent results on the ability of prodigiosin to transport HCO 3 − across lipid membranes are discussed.
Graphical Abstract

Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
The prodiginines (prodigiosenes) are heterocyclic natural products with a truly colorful history. Different terrestrial and marine bacteria produce these secondary metabolites. The most well-known microorganism that produces prodiginines is Serratia marcescens (Bacillus prodigiosis), a bacterium that grows well in damp and warm conditions (Fig. 1).
Serratia marcescens owes its name to Bartolomeo Bizio, an Italian microbiologist. Bizio, in an 1823 letter to a Venetian priest, explained the scientific origins of the bloody red color that had appeared on polenta, a corn meal produced annually by the Padua locals [1, 2]. At the time of this occurrence, during the wet and hot year of 1818, the polenta’s strange coloring alarmed the community. As we know now, the polenta’s red color did not arise because of any witchcraft. On the basis of his observations and experiments, Bizio suspected that microorganisms growing on the corn meal had caused the pigmentation. Indeed, it was later shown that the red color was due to the conjugated structure of the prodiginines, exemplified by the parent compound prodigiosin 1 (Fig. 2) (the UV-vis spectrum of prodigiosin 1 was reported in the following paper, although the proposed structure for 1 was incorrect at the time: [3]). There are a number of excellent reviews and book chapters that provide many interesting stories about the rich history and the religious and cultural impact of S. marcescens and their prodiginine metabolites [4–6].
Prodigiosin 1 was first isolated in pure form from S. marcescens in 1929 [7], and its correct structure was established by partial and total synthesis in the early 1960s [8, 9]. In addition to prodigiosin 1, there are many other naturally occurring prodiginines. Some other natural products 2–5 that will be discussed in this chapter are shown in Fig. 3. Early studies on the prodiginine natural products indicated that they were extremely cytotoxic and therefore they were not initially developed as drug candidates. However, over the last 15 years there has been a renewed interest in the chemistry and biochemistry of natural and synthetic prodiginines, in large part because some derivatives have promising anticancer and immunosuppressive activities at concentrations below which they are cytotoxic [10–13].
As depicted in Fig. 2 the protonated prodigiosin 1•H + has a binding pocket for Cl−, using hydrogen bonds and electrostatic interactions to coordinate the anion. As discussed later in this chapter, one hypothesis to explain how prodiginines trigger apoptosis, or programmed cell death, is due to their ability to transport HCl across plasma and intraorganellar membranes and to thus alter intracellular pH [14, 15]. In addition to binding anions, prodiginines also form complexes with transition metal ions [10]. It has been proposed that the redox chemistry of prodiginine–metal complexes may induce apoptosis. Thus, Manderville and colleagues have shown that prodiginines can bind tightly to double-stranded DNA and that prodiginine•Cu2+ complexes can trigger oxidative cleavage of DNA [16, 17]. A description of the metal binding and redox chemistry of the prodiginines is outside the scope of this chapter; the interested reader should refer to Manderville’s 2001 review article on the subject [10]. This chapter’s focus is to review that which is known about the binding and transmembrane transport of anions by prodigiosin 1 and analogs.
One ongoing and important effort is to produce prodiginines that are less toxic than the natural products but that are effective as immunosuppressive and/or anticancer agents. This goal has inspired the chemical syntheses of many new prodiginines [6, 18–22]. A search of the SciFinder database returns structures for hundreds of analogs, many of which are found in an extensive patent literature. Notably, some prodiginines have gone into preclinical and clinical trials (Fig. 4). For example, Aida Pharmaceuticals has prodigiosin 1 in preclinical trials for the treatment of pancreatic cancer [12, 23]. The analog, obatoclax mesylate (GX15-070) 6, [24] developed by Gemin Pharmaceuticals, has undergone Phase I clinical trials for patients suffering from lymphocytic leukemia [25]. The analog PNU-156804 (7), because of its immunosuppressive properties, [26] has been used as an antirejection compound in organ transplants [27].
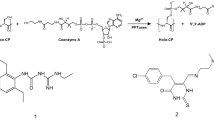
On another front, there has also been a major effort in understanding the biosynthesis of prodiginine natural products. This approach, led by researchers at the University of Cambridge, [28] holds promise for making new analogs by manipulating the biosynthetic machinery. For example, one key intermediate in prodiginine biosynthesis is 4-methoxy-2,2′-bipyrrole-5-carbaldehyde (MBC) 8, which is also often an intermediate in chemical syntheses. As depicted in Fig. 5 condensation of this aldehyde MBC 8 with a variety of pyrroles and amines in the presence of the biosynthetic enzyme PigC can give rise to new prodiginines 9 and a group of derivatives of related natural products known as the tambjamines 10 [29].
The chemistry and biochemistry of the prodiginines, like their history, is quite fascinating. Table 1 lists some of the key review articles that have been written about the prodiginines. The focus of this current chapter is the anion binding and anion transmembrane transport properties of the prodiginines. After a brief review of the structure of the prodiginines, we will focus on the biochemical and chemical data that has been accumulated to show how prodiginines are able to bind anions and to transport anions, especially chloride and bicarbonate, across phospholipid bilayer membranes.
2 Structural Studies of the Prodiginines and Their Anion Complexes
Despite the potent biological activities of many prodiginines, there are surprisingly few structural and computational studies reported in the literature, on either the natural products or their synthetic analogs. Issues concerning tautomeric and rotameric equilibria, solution conformation, and proton affinities of the prodiginines make their structural analysis not at all straightforward. As shown below in Fig. 6, the free base form of prodigiosin 1 may exist in solution as a mixture of tautomers. For example, structures I and II are tautomers that differ in the location of a pyrrole NH proton and the azafulvene’s exocyclic C–C double bond. Indeed, the tautomeric state of the free base has been a point of some contention in the prodiginine literature, with the azafulvene structure being drawn sometimes for the B-ring (tautomer I) and sometimes for the C-ring (tautomer II). As discussed below, X-ray crystal structures of synthetic prodiginines support the contention that isomer I, with the azafulvene located on the B-ring, is the favored tautomer (at least in the solid state). It should be emphasized, however, that protonation of either prodiginine tautomer I or II gives conjugate acids that are simply resonance forms of one another (see structure III).
Structures I and II represent different tautomers for the prodiginine’s free base. Tautomers I and II differ in the location of the pyrrole NH and in the location of the azafulvene’s C–C double bond. Protonation of tautomers I and II gives III, which can be depicted as either one of two major resonance forms
In addition to issues regarding different tautomers and protonation sites, there is even more potential for structural complexity with the prodiginines because of different conformations about the methyne linkage connecting the B- and C-rings. As shown in Fig. 7, both the free base and the protonated form of prodigiosin can adopt either the α-isomer or the β-isomer. The α-isomer 1α is stabilized by a hydrogen bond between the NH proton of the C-ring and the oxygen atom of the B-ring’s methoxy group. For the β-isomer 1β pictured in Fig. 7, which is the conformation that is usually drawn in the literature, the three pyrrole rings are all “cis” to one another, with each nitrogen atom oriented toward the center of a cleft. Transformation of the two isomers 1α and 1β corresponds to a cis–trans isomerization about the double bond connecting the B- and C-rings. While 1α is stabilized by hydrogen bonding between the B- and C-rings, one might well expect that binding an anion into the “all-cis” tripyrrole cleft would shift the equilibrium to favor the β-isomer, especially for the protonated prodiginine cation where each of the three NH protons could hydrogen bond to the anion to give the lipophilic ion pair 1β•H + A --. As described below, both X-ray and NMR data show that anions are indeed bound by the prodiginine β-isomer.
Prodiginines, both in their free-base and protonated forms, can exist as so-called α- and β-isomers. The α-isomer is stabilized by an intramolecular hydrogen bond between the B-ring methoxy group and the C-ring NH proton. The monoprotonated β-isomer is stabilized by hydrogen bonds from three NH protons to the bound anion
Rizzo and colleagues used solution NMR spectroscopy to study the rotamer populations of the synthetic prodiginine 7 (PNU-156804), a drug developed in the 1990s by the Pharmacia & Upjohn company [32]. In their study, Rizzo and colleagues used the distinct UV–vis absorption bands for the prodiginine free base (λ max = 460 nm) and for its protonated form (λ max = 525 nm) to conduct a spectrophotometric titration that provided them with a pK a value of 7.20 for the equilibrium mixture of the α- and β-geometric isomers of 7. Significantly, Rizzo and colleagues found that 7 gave two separate peaks in an HPLC trace when the separation was done under acidic conditions. They assigned these separate peaks to two slowly interconverting conformers, namely 7α and 7β. NMR analysis also showed two sets of resonances that varied in their relative ratios depending on the pH or solvent. Using HPLC data, they measured the interconversion rates for the two isomers which enabled them to calculate the pK a values for the two conformers, with pK α = 8.23 ± 0.03 for 7α and pK β = 5.4 ± 0.2 for 7β. This analysis indicated that the α-isomer of the prodiginine free base is much more basic than is the β-isomer, presumably because protonation of the 7α results in the formation of the stabilizing intramolecular hydrogen bond as shown in Fig. 6.
The 1H NMR spectra of 7 in CDCl3 showed a dramatic influence on the nature of the counter-anion, as the 7β•HCl salt gave a single set of resonances that were consistent with the β-isomer. This result is significant because it demonstrated that the Cl− anion could influence ligand conformation in a nonpolar solvent. Thus, in water, the protonated prodiginine 7α•H + adopted the “open” α-isomer, presumably because it was stabilized by the intramolecular hydrogen bond between the B- and C-rings. But in the nonpolar solvent, CDCl3, the Cl− anion can hydrogen bond with the cleft formed by the three pyrrole rings in 7β•H + Cl −. NOE interactions between the three NH-exchangeable protons (NH-1A to NH-1B and NH-1B to NH-1C) and the H-4C to H-1″ NOE confirmed the double-bond geometry for the β-isomer of PNU-156804•HCl in CDCl3. Similar conclusions were obtained in a recent NMR structure determination of the HCl salt of heptylprodigiosin, which also forms the pure β-isomer in CDCl3 [33].
Changing the counter-anion from chloride to the less basic methanesulfonate had a significant influence on the 1H NMR spectra of the ligand. Thus, the PNU-156804•CH3SO3H salt was found to be a mixture of conformers (α/β = 1/2) in CDCl3. This result allowed Rizzo and colleagues to assign the NMR resonances for both isomers and to carry out a detailed conformational analysis. NOESY data indicated that the two geometrical isomers of PNU-156804 7 differed mainly in the torsion angles about the C-2b-C-1″ bonds. For the β-rotamer, this torsion angle was close to 0°, whereas the angle was near 180° for the α-isomer (see Fig. 6). The authors noted that the α-isomer’s geometry coincided nicely with a previous crystal structure of a synthetic prodiginine [34]. Rizzo and colleagues explained that the α-isomer was stabilized by a hydrogen bond between the protonated nitrogen of ring C and the B-ring’s exocyclic oxygen. Importantly, the NMR studies showed NOEs between the CH3 group of the methanesulfonate anion and multiple NH and CH protons on the PNU-156804 core, indicating ion pairing and hydrogen bonding of the CH3SO 3 − anion with the prodiginine receptor, especially for the β-rotamer (see Fig. 8). The authors noted that their observations of distinct α- and β-isomers merited a detailed theoretical study of the prodiginine’s electronic properties. However, a decade later, little computational work on prodiginines has been published.
Recently, Manderville and colleagues published a study describing the conformation, protonation state and metal-binding affinity of synthetic prodiginines 11 and 12 (Fig. 9) analogs that contained isomeric phenols as their A-ring components [35]. These analogs had some unique properties, as compared to natural prodiginines. Thus, 11 and 12 underwent tautomerization in protic solvents to give mixtures of keto tautomers with λ max = 530 nm and enol tautomers with λ max = 460 nm, with the keto tautomers predominating in protic solvents. Conversely, the enol isomers were the exclusive tautomers observed in aprotic solvents. Manderville and colleagues proposed that the special properties for the o-phenol 11 arose because of formation of an intramolecular hydrogen bond between the A-ring’s phenol OH and N1 of the B-ring. They measured a pK a of 6.0 for the protonated analog 11•H+, a value that was 1.4 pK a units below that determined for p-phenol 12. They rationalized that this difference in acidity for analogs 11 and 12 was due to the stabilization provided by the intramolecular hydrogen bond formed by the β-form of 11. They also noted that since the α- and β-forms of natural prodiginines likely have different biological activities, o-phenol 11 may well be a useful model for understanding how prodiginine β-isomers bind to DNA and/or proteins.
3 Further Insight into Prodiginine Conformation from X-Ray and Computational Studies
In 1999, Furstner and colleagues published a study on the total synthesis and structural proof of the macrocyclic nonylprodigiosin 3 (see Fig. 3) [36]. They presented solid-state evidence that a synthetic intermediate on the way to 3 adopted the β-conformation with the B-ring existing as the azafulvene moiety. It should be emphasized that nonylprodigiosin 3 is constrained to adopt a β-conformation because of the carbon chain that links the A- and C-pyrrole rings.
A 2002 study by Manderville and colleagues described a prodiginine 13 that formed a hydrogen-bonded dimer in the solid state (Fig. 10) [37]. Within this dimer, the free base 13 was fixed in the β-conformation with all three pyrrole rings “cis” to one another. Once again, the B-ring of 13 existed as the azafulvene tautomer, which allowed for intermolecular hydrogen bonds between the B-ring nitrogen of one prodiginine 13 and the pyrrole NH atoms on the A- and C-rings of its partner. As described in more detail below, Parr et al. also observed a hydrogen-bonded dimer in the solid-state structure of a similar prodiginine analog [38]. Crystal-packing forces, which facilitate formation of hydrogen-bonded dimers, are the likely reason for the β-isomer being adopted in the solid state. As described below, calculations by Parr and colleagues indicated that the more extended prodiginine α-isomer should be lower in energy than the β-isomer [38].
Prodigiosin analog 13 and depiction of the solid-state structure of its free base form. The X-ray structure shows a noncovalent dimer (13)2 formed by intermolecular hydrogen bonds. Reprinted with permission from [37]
There has been little reported about computational analysis of prodiginines and, to our knowledge, no reports on calculations for prodiginine–anion complexes. The most detailed computational work on prodiginine conformation was reported in 2008 [39]. In that study, Cole and colleagues conducted density functional theory (DFT) calculations on neutral and protonated compounds in order to better understand the mass spectra and fragmentation patterns of protonated prodiginines. These authors noted that prodiginine structures are typically drawn in the β-configuration as depicted in Fig. 6. But, Cole’s DFT calculations showed that the β-isomers are not the lowest-energy structures in the gas phase. Instead, they found that the lowest-energy conformation has the prodiginine A- and C-rings rotated 180° relative to the central B-ring (Fig. 11). They rationalized that this geometry allows the molecule to maintain planarity while placing the lone pairs of electrons on the three nitrogen atoms as far apart as possible, so as to minimize electrostatic repulsion. Ab initio calculations on 2,2′-bipyrrole (a model for the prodiginine A- and B- rings) indicate that the compound’s global energy minimum is a nonplanar conformation that is twisted about 40o out of plane [40]. As reported by Cole, Fig. 11a (I) shows the low-energy geometry of the neutral model compound 14, Fig. 11b (II) shows the chemical structure of this conformation and Fig. 11c (III) shows the compound’s electron density map. Figure 11c (III) indicates that the B-ring’s nitrogen has the highest electron density, suggesting that this nitrogen atom is the preferred site of protonation. These authors also calculated the relative energies of three stable conformers of the [M + H]+ species that resulted from protonation of the neutral 14. As indicated in Fig. 11d, the lowest-energy conformation for 14 contained an intramolecular hydrogen bond between the C-ring’s NH proton and the B-ring’s methoxy group, namely, the α-isomer observed by Rizzo and colleagues in their NMR investigations [32]. In a recent study, DFT calculations by Parr and colleagues also showed that the free base’s lowest-energy conformation had an “all-trans” arrangement for the three pyrrole rings that make up the α-isomer [38]. But, they stressed that this α-isomer conformation was just 4 kJ/mol lower in energy than the β-conformation that had been experimentally observed in their X-ray crystal structure. Parr and colleagues suggested that the ability to form hydrogen bonds, to either a Cl− anion or to another potential electron-rich partner, could readily provide the energy needed to switch the conformation favored by the prodiginine’s tripyrrole unit. Calculations on the related tambjamine natural products (see structure 10 in Fig. 5) also indicate that an intramolecular H-bond is formed between an NH donor and the B-ring’s exocyclic oxygen atom [41]. It would seem that theoretical computations on the structure and dynamics of prodiginines and their complexes with anions and metal cations is one area that is ripe for exploration, especially given these compounds’ growing importance in drug development.
(a) Optimized geometry of the neutral prodiginine model system 14 at the B3PW91/6-31G* level; (b) chemical structure for 14; (c) electron density map (order of decreasing electron density is purple > blue > green > yellow > red) indicating highest electron density at the nitrogen atom of the central B-ring; (d) the optimized prodiginine conformations and their relative energies calculated by DFT. Reprinted with permission from [39]
4 Solid-State Structural Studies on Prodiginine•HCl and Prodiginine–Metal Ion Complexes
There are only a few X-ray crystal structures available for complexes of protonated prodiginines with anions or for complexes of deprotonated prodiginines with metal cations. In this section, we describe what is known about the solid-state structures of prodiginines and their complexes. While exploring the chemistry of acyclic oligopyrroles Sessler and colleagues synthesized a linear hexapyrrole 15 that is essentially two prodiginine units connected together [42]. Indeed, the structure of the bis-HCl salt of this hexapyrrole 15 can be considered to be a harbinger for understanding how prodiginines bind anions. Thus, the X-ray structure of the bis-HCl salt of 15 showed two chloride anions bound to a planar S-shaped conformation (Fig. 12). Both of the clefts formed by the protonated tripyrrole units had three NH···Cl− hydrogen bonds that fixed the anions in their respective binding sites. This early result, as later described by Sessler and Gale, provided “…direct structural conformation for the mode of binding presumed to be operative in prodigiosin and its analogues” [43].
The X-ray structure of the bis-HCl salt of hexapyrrole 15 shows two chloride anions bound to a planar S-shaped conformation. Reprinted with permission from [42]
Indeed, the Sessler group later obtained solid-state data that confirmed the manner in which prodiginines bind chloride ions [18]. An X-ray crystal structure of the complex 16•HCl showed a chloride anion bound in the prodiginine’s tripyrrole cleft, held there by hydrogen bonds and, presumably, by electrostatic interactions with the protonated ligand (Fig. 13). The prodiginine analog clearly adopts the β-isomer in this structure. The tripyrrole unit in 16•HCl is relatively flat with the C-ring showing a small deviation from planarity. This conformation for 16•HCl is similar to that found in the solid state for the HCl salt of a derivative of nonylprodigiosin, 3•HCl, although as previously mentioned the chain connecting the A- and C-rings in that macrocycle helped to constrain the tripyrromethane’s β-isomer conformation [36].
Left panel: Structure of the prodiginine 16. Right panel: Views of the X-ray structure of the complex formed between the protonated prodiginine 16 and chloride ion; Reprinted with permission from [18]
Parr, Wasserman and colleagues recently reported the X-ray crystal structures of both the free base and the HCl salt of a synthetic prodiginine 17, a compound with a 2,5-dimethyl pyrrole as the C-ring [38]. The X-ray structure of the HCl salt of 17 (Fig. 14) shows a planar tripyrromethane cation with all three N–H hydrogens pointing toward the bound chloride anion. Again, it is these 3 N–H…Cl− hydrogen bonds that stabilize the β-conformation for 17•HCl. This study also revealed the predominant tautomer favored by the free base in the solid state. The X-ray structure of free base 17 showed that the B-ring was not protonated and that this B-ring contained the azafulvene unit. Thus, the C8–C9–C10 methyne linker had a shorter C8–C9 bond (1.350(7) Å), as compared to a C9–C10 bond length of 1.407(7) Å. Crystallized from CH2Cl2, the free base form of 17 forms a noncovalent dimer that is held together by intermolecular H-bonds between NH protons on the A- and C-rings of one molecule of 17 and the B-ring’s nitrogen of the other molecule. This structure is reminiscent of the noncovalent dimer observed in the solid-state structure of analog 13 by Manderville and colleagues [37]. In both of the solid-state dimers formed by 13 and 17, the prodiginine free base adopts the β conformer.
The structure of prodiginine 17 and solid-state structures of the complex 17•HCl and the free base form, which forms a noncovalent dimer (17)2. Reprinted with permission from [38]
5 X-Ray Structures of Prodiginine–Metal Complexes
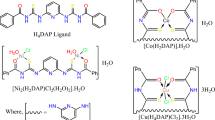
Manderville has provided compelling evidence that prodigiosin (1) can bind to DNA and cleave double-stranded DNA in the presence of transition metals [16, 17]. They have proposed that prodigiosin’s ability to cleave dsDNA may be important for its potent anticancer activity. In a 2003 communication, they reported on the coordination chemistry of prodiginine anions with Zn (II) and Cu (II) [44]. Treatment of prodigiosin 1 with strong base deprotonated the pyrrole NH protons, which then enabled reaction with transition metal salts. Deprotonated prodigiosin 1 bound Zn (II) to generate a 2:1 complex, Zn•(1)2. A solid-state structure of this complex showed it to be a 4-coordinate, neutral species with the B- and C-rings of two prodiginines bound to the metal cation to give a distorted tetrahedral geometry. The 1H NMR spectra of the complex Zn•(1)2 indicated that this structure was maintained in CDCl3 solution. In contrast to the structure obtained with Zn (II), reaction of deprotonated prodigiosin with CuCl2•2H2O gave a 1:1 complex with a distorted square planar geometry, wherein the Cu (II) cation was coordinated to all three pyrrole nitrogen atoms. Remarkably, the C-ring of this prodigiosin•Cu (II) complex was oxidized at its C1 position. Solution NMR experiments indicated that this OH group attached to the prodiginine C-ring was derived from H2O. Manderville and colleagues noted that understanding how prodigiosin was oxidized under these conditions might provide insight into the mechanism of DNA cleavage that was catalyzed by the prodiginine•Cu (II) system (Fig. 15).
Left Panel: Perspective drawing of the solid-state structure of Zn•(1) 2 . The Zn atom is represented by a green sphere. Right Panel: ORTEP plot (50% probability thermal ellipsoids) of the solid-state structure from the reaction of prodigiosin 1 with CuCl2. The observed structure has an –OH group added to the C1 position on prodigiosin’s C-ring. Reprinted with permission from [44]
6 Proton Transport: Early Studies Showing That Prodiginines Alter Intracellular pH
In retrospect, the first indication that prodiginines could move HCl across membranes came during the studies on their influence on the activity of vacuolar H+-ATPase (V-ATPase), a highly conserved enzyme found in eukaryotes. The V-ATPases are responsible for the acidification of certain cell types and intracellular organelles by pumping protons, against their chemical potential, across plasma and intracellular membranes [45]. ATPases catalyze the hydrolysis of adenosine triphosphate (ATP) into adenosine diphosphate (ADP) and phosphate ion. The reaction is highly exergonic and the V-ATPases use the energy from ATP hydrolysis to transport protons across intracellular and plasma membranes. The effect of V-ATPase proton transport activity is the acidification of certain cellular organelles, including lysosomes, Golgi apparatus and other vacuoles. Thus, the V-ATPases are often necessary for proper regulation of intracellular and/or intraorganellar pH and for regulation of numerous cellular functions.
There were two reports in 1995 that prodiginines could destroy the pH gradients that existed across the membranes of certain cellular organelles, and thus serve as de facto inhibitors of V-ATPase [50, 51]. As detailed below, this ability to alter transmembrane pH gradients was later shown to be due to the ability of these natural products to facilitate the cotransport of H+ Cl− (or alternatively to enable exchange of OH− and Cl− anions) ([46, 47]; for a similar study as that described in [47], see also [48, 49]). In the first of their 1995 papers, Kataoka and colleagues showed that prodigiosin 25-C (undecylprodigiosin) 2 was able to neutralize acidic organelles that were located within cytotoxic T cells and, by so doing, negate the cytotoxicity of these T cells [50]. These authors demonstrated that prodigiosin 25-C 2 and the ATPase inhibitor, concanamycin B (CMB), were potent immunosuppressants, both in vitro and in vivo. Thus, they proposed “…because PrG and CMB showed similar immunosuppressive effects in vivo as well as in vitro, it is most likely that the inhibition of intra-organellar acidification is responsible for suppression of CTL in vivo” [50].
In their second 1995 publication, Kataoka and colleagues demonstrated that 2 was able to neutralize the proton-translocation activity of V-ATPase, without inhibiting the enzyme’s ATP hydrolysis activity [51]. They found that prodigiosin 25-C 2 inhibited the accumulation of pH-sensitive dyes, such as acridine orange, in the acidic compartments of baby hamster kidney (BHK) cells. In addition, 2 inhibited the proton pump activity of V-ATPase with an IC50 value of ca. 30 nM (see Fig. 16a), but the compound did not alter the ATPase enzyme activity, even at ligand concentrations of up to 1 μM (see Fig. 16b). In marked contrast to the properties displayed by prodigiosin 25-C 2, bafilomycin A, a known inhibitor of V-ATPase, inhibited both the protein’s proton pump activity and its ATP hydrolysis activity. Another group later published similar findings that the immunosuppressant, cycloprodigiosin hydrochloride (cPrG•HCl) 4, inhibited the proton-translocation mediated by V-ATPase without influencing the enzyme’s ATPase activity [52].
Proton pump and ATPase activities in rat liver lysosomes. (a) FITC ∼ dextran containing lysosomes were incubated with prodigiosin 25-C filled circle or bafilomycin AI open circle, and fluorescence quenching was measured after the addition of ATP. (B) V-ATPase activity was measured by the release of inorganic phosphate from ATP. The reaction mixture was incubated with prodigiosin 25-C filled circle or bafilomycin A, open circle. Reprinted with permission from [51]
Significantly, Kataoka and colleagues also used transmission electron microscopy to show that 2 caused a significant swelling of the Golgi apparatus and mitochondria within the BHK cells [51]. These results supported the hypothesis that prodigiosin 25-C 2 raised the pH of acidic compartments through the inhibition of the proton pump activity of V-ATPase, thereby causing the observed morphological changes to the Golgi apparatus and mitochondria. The authors proposed that inhibition of the ATPase’s proton-translocation activity resulted in osmotic imbalances that led to water influx and consequent swelling of the intracellular organelles.
In these 1995 papers, Kataoke and colleagues did not offer any specific molecular mechanism for how prodiginine 2 might disrupt the proton pump activity of V-ATPase, but they later demonstrated how the proton transport properties of prodiginines can have key biological consequences. Thus, Nagai, Kataoka and colleagues showed that both prodigiosin 25-C 2 and metacycloprodigiosin 5 suppressed bone resorption by osteoclasts, cells that are responsible for breaking down bone tissue by digesting minerals and matrix proteins [53]. The osteoclasts require an acidic microenvironment in order to carry out bone digestion and resorption. That acidic pH is regulated by the osteoclast’s V-ATPase, found in the cell’s outer membrane. Nagai and coauthors demonstrated that bone tissue treated with prodiginines had a dramatic decrease in the amount of digestion observed, probably because “…inhibition of the acidification of vacuolar organelles causes the destruction of the acidic microenvironment of osteoclasts and results in significant damage to the osteoclastic function in bone resorption” [53].
7 Prodiginines Alter pH by Facilitating Transmembrane H+ Cl− Symport (or the Functionally Equivalent OH−/Cl− Exchange)
In 1998, two seminal papers in the field were published by Ohkuma, Wasserman and colleagues wherein they demonstrated that prodiginines uncouple the proton-translocation activity of V-ATPase by functioning as transmembrane H+Cl− cotransporters [46, 47]. In these two papers, Ohkuma, Wasserman and colleagues demonstrated that three different members of the prodiginine family, prodigiosin 1, prodigiosin 25-C 2 and metacycloprodigiosin 5 all inhibited the proton-translocation activity of bovine V-ATPase in a process that depended on the presence of chloride anion [46, 47]. They also showed that prodiginines did not cause the formation of a membrane potential across the lysosomes. These two results suggested that the H+ translocation was electroneutral and that chloride was moved across membranes by the prodiginines. On the basis of these observations, they proposed that prodiginines facilitate the cotransport, or symport, of H+Cl− across vesicular membranes. The authors also pointed out that H+Cl− symport was functionally equivalent to OH−/Cl− anion exchange, and that their results could not unequivocally distinguish between the two possible mechanisms. Nonetheless, over the years the prodiginines have been consistently described as transmembrane H+Cl− symporters.
Having demonstrated a Cl− dependence in lysosomal membrane preparations, these authors carried out key experiments that investigated Cl− transport across liposomal membranes, systems devoid of any ion channel proteins that might contribute to anion transport or to proton translocation [46]. First of all, they discovered that the prodiginines could modulate the internal pH of liposomes depending on the Cl− anion gradient that exists across the membrane. Thus, if the extravesicular solution contained excess Cl−, then they found that the addition of the prodiginines would lower the internal pH, presumably by moving H+Cl− into the liposome interior. Conversely, if the intravesicular solution were higher in Cl− concentration, relative to the bulk solution, then the prodiginines would cause an increase in the intra-liposomal pH, again because they cotransported H+Cl− out of the vesicle (or alternatively they promoted transmembrane OH−/Cl− exchange). In another indirect method of demonstrating chloride anion transport, these authors showed that prodiginines were able to induce the Cl−-dependent swelling of liposomes and erythrocytes [47]. In this case, chloride anion permeability was estimated by measuring the swelling of vesicles in buffers containing iso-osmotic ammonium salts. Thus, addition of any of the three prodiginines (namely 2, 3 and 5) caused swelling of phospholipid liposomes, as determined by a reduction in the absorbance of liposomes suspended in 0.2 M NH4Cl, but not in ammonium gluconate [47].
The results described above, although convincing, provided only indirect evidence that prodiginines could transport Cl− anion across lipid membranes. Thus, Ohkuma, Wasserman and colleagues designed an experiment that provided direct proof for transmembrane Cl− transport as mediated by the prodiginines [46]. They demonstrated that prodiginines 2, 3 or 5 each promoted the uptake of radioactive 36Cl from external buffer, with concomitant acidification of the internal compartment, when the initial pH of the extravesicular buffer was more acidic than the liposome’s interior. This was clear evidence that prodiginines could transport H+Cl− when confronted with a transmembrane pH gradient.
Ohkuma, Wasserman and colleagues concluded that the ability of the prodiginines to uncouple the V-ATPase’s proton-translocation activity was due to their ability to facilitate H+ Cl− symport (or OH−/Cl− antiport) across membrane barriers [46–49]. As depicted in Fig. 17, they proposed that prodiginines could form lipophilic ion pairs upon binding H+ Cl− via electrostatic, charge-transfer and hydrogen-bonding interactions. These lipophilic ion pairs would then enable the proton-coupled movement of chloride across a phospholipid membrane. In a recent paper, Ohkuma and colleagues demonstrated that a member of the tambjamine antibiotic family, BE-18591 (18), also uncoupled V-ATPases through its H+Cl−symport activity [54]. As shown in Fig. 17, prodiginines and tambjamines have similar structures. Ohkuma and colleagues found that 4-methoxy-2,2′-bipyrrole-5-carboxyaldehyde 8 (see Fig. 5), the synthetic precursor of BE-18591 (18), did not inhibit the proton pump activity of V-ATPase in membrane vesicles. This finding indicated that at least three nitrogen atoms, with two of the nitrogen atoms in pyrrole rings, are required for H+Cl− symport activity.
Proposed mechanism for how prodigiosin 1 and the related tambjamine 18 alter the intravesicular pH within a liposome. The lipophilic ion pair 1•H + Cl − formed by prodigiosin 1 and HCl can be transported across the membrane (a symport process shown in red). An alternative to explain the change in intravesicular pH change would be if the monoprotonated form of prodigiosin 1 were able to catalyze exchange of the Cl− and OH− anions (antiport process shown in black)
8 HCl Transport Mediated by Prodiginines Has Biological Consequences
Plant vacuoles contain two distinct proteins that function as H+ pumps: H+-ATPase and an H+-translocating pyrophosphatase (H+-PPase) [55]. Pyrophosphatase is an enzyme that converts one molecule of pyrophosphate into two phosphate ions in an exergonic reaction (ΔG = −34 KJ). This reaction, when coupled to the translocation of protons across the membrane, enables an acidic pH to be maintained within the plant vacuole. Shimmen et al. showed that nanomolar concentrations of cycloprodigiosin hydrochloride (cPrG•HCl) 4 blocked acidification of vacuolar vesicles by H+-PPase [56]. At the time of this study, there were no known inhibitors of this H+-PPase pump. Significantly, this inhibition of vacuolar acidification by cPrG•HCl 4 was anion-specific, as it occurred in buffers containing chloride anion but not in media containing the more hydrophilic sulfate anion (SO 4 2− ). Shimmen’s paper provided one of the first reports about the attenuated activity of a prodiginine as a function of the anion (Cl− vs. SO 4 2− ).
These authors prepared isolated vesicles from vacuolar membranes that contained the H+-PPase proton pump enzyme, but that had other plasma and ER membrane components removed. They used cPrG•HCl 4 purified from Pseudoalteromonas denitrificans to show that this compound markedly suppressed the H+ transport activity of the H+-PPase without influencing the enzyme’s phosphatase activity. This ligand-mediated inhibition of acidification occurred only in buffers containing chloride anion, but not in the sulfate buffers lacking chloride. In solutions containing 50 mM KCl, the prodigiosin cPrG•HCl 4 (10 nM) inhibited the effects of the proton transport activity of H+-PPase. In a separate experiment, these authors also showed that cPrG•HCl 4 rapidly destroyed any pH gradient that had been previously generated across the vacuolar membrane by the action of H+-PPase. In marked contrast, cPrG•HCl 4 showed no effect on the intravacuolar pH in an assay medium that lacked chloride anion, but instead contained 25 mM K2SO4. These observations supported the earlier proposal made by Sato et al. that prodiginines function as H+/Cl− symporters [46, 47]. Presumably, sulfate anion was too hydrophilic for cPrG 4 to be able to bind and move across the vacuolar membrane. These authors noted that the concentration of Cl− is high enough in plant cells that cPrG•HCl 4 should be able to destroy any pH gradients that are built up across the vacuolar membrane within plant cells. Thus, they suggested that the H+uncoupling activity of cPrG•HCl 4 might ultimately be useful for studying the physiology of vacuoles in living plants. Indeed, one year later these same authors showed that cPrG•HCl 4 could promote acidification of vacuoles in living plant cells in the presence of Cl− anion [57].
9 Prodiginines, H+Cl− Cotransport and Apoptosis
Cell shrinkage, membrane blebbing, condensation of nuclear chromatin and fragmentation of DNA characterize apoptosis, or programmed cell death. Various studies indicate that intracellular acidification can induce apoptosis in cancer cells [58]. The intracellular pH (pHi) of cancer cells is often more alkaline than that of normal cells, and maintenance of this alkaline pHi is required for growth, transformation, and metabolism of these cancer cells. Therefore, a potential antitumor therapy would be to trigger apoptosis of cancer cells by perturbing the mechanisms that control the pHi of cancer cells. As described above, prodiginines have recently been shown to deacidify lysosomes and, thus, negate the proton-translocation activities of V-ATPase without any measurable inhibition of ATP hydrolysis [50, 51]. This uncoupling of V-ATPase by prodiginines was attributed to the compounds’ ability to promote H+/Cl− symport (or the functionally equivalent OH−/Cl− antiport) across vesicular membranes that experienced a transmembrane pH gradient.
In this study, Yamamoto and colleagues examined the impact of cycloprodigiosin hydrochloride (cPrG•HCl) 4 on cancer cell lines both in vitro and in vivo [14]. They found that cPrG•HCl 4 inhibited the growth of six different liver cancer cell lines in a dose- and time-dependent fashion. In sharp contrast, treatment of normal rat hepatocytes with cPrG•HCl 4 had no effect on cell growth. In one case, in vitro treatment of Huh-7 liver cancer cells with cPrG•HCl 4 caused apoptosis. This group also showed that cPrG•HCl 4 raised the pH of acidic organelles within these Huh-7 cancer cells and also lowered the intracellular pHi (below pHi = 6.8). Figure 18 shows Huh-7 cells loaded with the pH-sensitive dye, acridine orange, before and after treatment of these cells with cPrG•HCl 4. In the upper frame, the orange fluorescence is due to acridine orange located within the acidic lysosomes. As seen in the lower frame, after treatment with cPrG•HCl 4 for 1 h the characteristic orange fluorescence was not observed, indicating deacidification of the lysosomes. These authors also used flow cytometry to determine that the intracellular pH of the Huh-7 cells decreased from pHi = 7.3 to pHi = 6.8 after treatment with cPrG•HCl 4 for 24 h. For the in vivo experiments, nude mice containing xenografted Huh-7 cells received 2 weeks of treatment with cPrG•HCl 4. After 8 days of treatment with compound 4, they observed that tumor growth was significantly inhibited. Importantly, a histological examination showed that apoptosis had occurred in the tumor cells treated with cPrG•HCl 4. Although these authors cautioned that the molecular mechanisms underlying the effects of cPrG•HCl 4 on tumor cells needed further attention, they suggested that prodiginine analogs had much potential to be effective anticancer drugsFootnote 1 [59].
Acridine orange staining of (a) untreated Huh-7 liver cancer cells and (b) Huh-7 cells treated with 276.5 nmol/L of cPrG•HCl 4 for 1 h. The red color in (a) is indicative of the acid form of the pH-sensitive acridine dye. Treatment of the cells with cPrG•HCl 4 results in neutralization of the acidic compartments within these cells. Reprinted with permission from [14]
In a later paper, Nakashima and colleagues showed that a bacterial prodiginine, one that they dubbed PG-L-1, was cytotoxic to human myeloid leukemia (U937) cells [15]. This compound caused the typical increase in acidity in intracellular compartments, presumably because of H+ Cl− symport. The authors proposed that this acidification of the cytoplasm triggered apoptosis. As shown in Fig. 19, incubation with the prodigiosin PG-L-1 caused significant morphological changes in the U937 cells. Fluorescent staining of these treated U937 cells showed that PG-L-1 caused diagnostic changes in the nuclear material that was typical of apoptosis.
Effect of prodiginine PG-L-1 on cellular and nuclear morphology in U937 cells. Cells (6 × 105 cells/dish) were treated with 0 mg/mL (A, E), 0.1 mg/mL (B, F), 1.5 mg/mL (C, G), and 50 mg/mL (D, H) of PG-L-1 and then the cells were observed under a phase contrast microscope (A, B, C, D), or a fluorescence microscope (E, F, G, H) after being stained with Hoechst 33258 (40 mM) for 10 min. The bar indicates 10 mm. Reprinted with permission from [15]
For more details about the ability of prodiginines to trigger cellular apoptosis and on the use of prodiginines as anticancer therapeutics, the interested reader should consult recent reviews and primary articles [10–15, 59–61].
10 Synthetic Prodiginine Analogs Shown to Bind Cl− in Solution, Transport Cl− Across Lipid Membranes and Possess Anticancer Activity
An important study regarding the potential anticancer properties of synthetic prodiginines was reported by Sessler, Magda and colleagues in 2005 [18]. In that paper, they reported that the transmembrane rate of Cl− transport correlated with the compound’s in vitro anticancer activity. As mentioned earlier, X-ray crystal structures showed that some of these synthetic analogs bound Cl− in the solid state (see compound 16 in Fig. 13). Sessler, Magda and colleagues also used isothermal titration calorimetry (ITC) to demonstrate that the protonated forms of the prodiginines had a strong affinity for binding chloride in CH3CN solution. While the free base forms of the tripyrroles bound Cl− only weakly, even below the limits of detection by ITC, the protonated forms of the prodiginine analogs showed substantial affinities for Cl−. Thus, the apparent binding constants were in the range of K a = 5.9 × 105M−1 to 1.0 × 105 M−1for the formation of Cl− adducts by these protonated tripyrroles.
These authors also measured the prodiginines’ efficiency for transmembrane transport of chloride ions across synthetic phospholipid membranes, using an ion-selective electrode to monitor Cl− efflux from the liposomes. They discovered that the synthetic analog 13 (see Fig. 10) with the structure closest to that of the natural product 1 showed the fastest rate of Cl− efflux from liposomes. The authors concluded, from a series of transport experiments using different transmembrane pH gradients, that these synthetic compounds operated via the H+/Cl− cotransport mechanism.
The in vitro anticancer activity of these synthetic analogs was determined from a cell proliferation assay using both A549 human lung cancer and PC3 human prostate cell lines. All of the tested compounds, including 13 (Fig. 10) and 16 (Fig. 13),exhibited significant cytotoxic activity, with 100% of the cancer cells killed using concentrations of 40 mM of the analogs. In examining the combined data, the authors noted that the in vitro anticancer activity correlated best with the Cl− transport rates, rather than with the association constant (K a) for Cl− binding. These results led Sessler, Magda and colleagues to conclude that “… the strong correlation between transport rates and anticancer activity in vitro, in conjunction with evidence for anion binding in the solid state and solution phase, lead us to suggest that the H+/Cl− ion symport mechanism proposed by Ohkuma, Wasserman, and co-workers is chemically reasonable” [18].
11 Prodiginines Can Also Facilitate Anion Exchange (Antiport) Across Phospholipid Membranes
On the basis of the pioneering studies of Ohkuma, Wassermann and colleagues, prodigiosin 1 has typically been described in the literature as an H+/Cl− cotransporter. However, Ohkuma et al. indicated that the observed pH changes in vesicles caused by the addition of 1 could also be due to a Cl−/OH− anion exchange mechanism [46, 47]. A 2005 study by Seganish and Davis showed that prodigiosin 1 could indeed move Cl− anions across phospholipid membranes via an anion exchange, or antiport, mechanism [62]. Thus, these authors showed that prodigiosin 1 was able to exchange Cl− anion for NO 3 − anions during transmembrane transport, without any observed change in the internal pH of the liposomes, a result that was entirely consistent with prodigiosin 1 functioning as an antiporter.
In their study, Seganish and Davis began by showing that prodigiosin 1 operates as an anion carrier, rather than as an ion channel. To distinguish a carrier mechanism from channel formation, they measured Cl− influx into liposomes made from dipalmitoyl-phosphatidylcholine (DPPC) at temperatures that included those that were above and below DPPC’s gel-to-liquid crystalline transition temperature (41°C). These DPPC liposomes contained a fluoresent dye, lucigenin, that is selective for interaction with Cl− anion [63]. Previous studies have shown that an ion carrier’s efficiency is limited by diffusion through the membrane and that the transport rate is greatly diminished in the “frozen” gel state [64]. Ion channel activity, on the other hand, is not as sensitive to the lipid phase and the observed transport rate for channels is not as temperature-dependent as for mobile carriers. Indeed, the transmembrane transport of Cl− by prodigiosin 1 across the DPPC liposomes was quite sensitive to the lipid phase. Thus, the influx of Cl− was rapid at 43°C, a temperature above DPPC’s phase transition, wheras little Cl− influx was observed at 37°C, a temperature at which DPPC exists in its gel state (Fig. 20). The authors indicated that these transport results were consistent with prodigiosin 1 functioning as a chloride anion carrier.
Chloride influx into DPPC liposomes at 43°C and 37°C. The data at each temperature is the average of three runs using 0.004 mol% of prodigiosin 1. The trace shown in gray represents a DMSO blank at 37°C. Reprinted with permission from [62]
Seganish and Davis also discovered that the ability of prodigiosin 1 to transport Cl− into phospholipid vesicles depended greatly on the nature of the anion that was initially inside the liposome. In experiments with vesicles that contained either Na2SO4 or NaNO3, they measured both the internal pH of the liposomes (using the pH-sensitive dye pyranine) [65] and Cl− influx (using the lucigenin dye). When the liposomes were filled with the hydrophilic sulfate anion (ΔG hyd = −1,080 kJ/mol), the authors found that the addition of prodigiosin 1 resulted in immediate acidification inside the liposomes but Cl− influx occurred to only 20% of the value that would be expected for complete equilibration. In contrast, when similar experiments were conducted with liposomes that were loaded with the less hydrophilic nitrate anion (ΔG hyd = −300 kJ/mol) the authors observed rapid and complete exchange of Cl− and NO 3 −, but with no concomitant change in intravesicular pH (Fig. 21). Seganish and Davis concluded that prodigiosin 1 was able to facilitate Cl−/NO 3 − anion exchange (antiport) in the experiments with nitrate-loaded liposomes. In contrast, prodigiosin 1 was able to promote some initial H+/Cl− symport (or OH−/Cl− exchange) when the hydrophilic sulfate anion was present in the liposomes. Thus, this paper was important because it showed, for the first time, that prodigiosin 1 could facilitate transmembrane anion exchange. Moreover, the study also demonstrated that prodigiosin’s mechanism for Cl− transport might well change from Cl−/A− antiport to H+/Cl− symport, depending on the environmental conditions.
Chloride gradient assays on 100 nm EYPC liposomes in 10 mM sodium phosphate (pH 6.4) containing either 75 mM Na2SO4 (a and b) or 100 mM NaNO3 (c and d). The chloride gradient was initiated by adding NaCl to give an external concentration of 25 mM. Chloride concentration inside the vesicles (panels a and c) was calculated from the fluorescence of entrapped lucigenin dye. The pH (panels b and d) is calculated from the fluorescence ratio of HPTS dye emitted at 510 nm when excited at 403 and 460 nm in a dual wavelength assay. The trace shown in gray represents DMSO blanks. Reprinted with permission from [62]
Thompson, Davis and colleagues later used this Cl−/NO 3 − transmembrane exchange assay, with the lucigenin dye, to investigate the ability of a series of synthetic prodiginines to transport chloride ions across liposomal membranes made from egg yolk l-phosphatidylcholine (EYPC) [66]. They found that most of these synthetic prodiginines retained significant chloride ion transport ability at low ligand: lipid concentrations (0.1 mol%). For example, the β-substituted prodiginines 19 and 20, shown in Fig. 22, while less active than prodigiosin 1, still retained a significant ability to move Cl− anion across EYPC lipid membranes. The authors noted that the modest decreases in Cl− transport rates exhibited by these synthetic prodiginines may be due to either a reduced ability to partition into the lipid membrane, a reduced rate of diffusion across the bilayer or because of a change in binding affinity for Cl− anion. Nonetheless, the most important finding of these transport experiments was that different side chains could be attached to the prodiginine’s tripyrrole unit core without losing the ability to transport chloride ions across phospholipid membranes.
12 H+ Cl− Transport by Synthetic Receptors Designed to Mimic Prodiginine Function
Recently, the groups of Gale and Smith reported intriguing studies on neutral ligands designed to mimic prodigiosin’s ability to transport HCl across bilayer membranes [67, 68]. Thus, compound 21 in Fig. 23 contains two hydrogen bond donors, a pyrrole NH and an amide NH, for interaction with anions. Moreover, compound 21 has a basic imidazole side chain that can be protonated and then enhance hydrogen bond interactions with a bound Cl− anion. In this way, receptor 21 was designed so that it might cotransport H+ Cl− across a bilayer that is experiencing a pH gradient. The working hypothesis was that compound 21 should be able to move HCl from an acidic solution into a more basic solution where ligand deprotonation and Cl− decomplexation would then be favored. An X-ray crystal structure of 21•HCl showed a [2 + 2] dimer, with each chloride involved in three hydrogen bonds. Two of the NH…Cl hydrogen bonds involved the pyrrole NH and amide NH of one molecule and the third hydrogen bond was with the imidazolium unit of another molecule of 21. The authors noted that HCl bound within such a [2 + 2] dimer would be protected by the structure’s lipophilic exterior. If the solid-state structure were relevant to function, then such an organization would provide a nice way to sequester HCl for transport across the hydrophobic membrane.
A synthetic prodiginine mimic 21 and the crystal structure of its HCl complex showing formation of a “2 + 2” hydrogen-bonded dimer. Reprinted with permission from [67]
Using a Cl− electrode, the authors found that 21 transported Cl− across vesicles made from 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) and cholesterol. The authors showed that the rates of Cl− transport were pH-dependent. The greatest rate of Cl− efflux from the NaCl-filled liposomes occurred when the intravesicular solution was acidic (pH 4.0) and the extravesicular buffer was near neutral (pH 6.7). In biomimetic fashion, the synthetic compound 21 could also deacidify liposomes, a property that is a hallmark of the prodiginines. Thus, addition of compound 21 to liposomes loaded with the acid-sensitive dye, Oregon Green 514, destroyed the initial pH gradient across the membrane. Gale and Smith proposed that the free base of 21 that partitioned into the membrane could diffuse across the bilayer where it bound HCl at the interface of the lipid membrane and the intravesicular solution. The resulting lipophilic ion pair could then move back across the bilayer to deposit HCl into the more basic external solution.
13 Prodigiosin 1 Facilitates Transmembrane Transport of Bicarbonate Anion
Recently, Davis, Gale, Quesada and colleagues showed that prodigiosin 1 can facilitate the transmembrane exchange of Cl− and HCO 3 − anions [69]. These authors suggested that prodigiosin’s specific pattern of hydrogen-bond donors and acceptors might well allow it to function as a complementary receptor for binding and membrane transport of bicarbonate (Fig. 24). Indeed, it was reported over 50 years ago that prodigiosin 1 reacts with carbonic acid to give a protonated adduct, 1•HCO 3 − [70]. Davis and colleagues showed, using NMR spectroscopy, that prodigiosin 1 can bind HCO 3 − anion in the nonpolar solvent CD2Cl2. Significantly, the 1H NMR signals for 1 that were most influenced upon addition of tetraethyl-ammonium bicarbonate were the H2 proton on the A-ring and the methyl group on the C-ring. The authors noted that these would be the protons closest to the anion-binding cleft formed by the “all-cis” β-isomer of prodigiosin 1 (Fig. 1). Although the authors did not report a stability constant for the 1•HCO 3 − complex, their 1H NMR data demonstrated that the free base form of prodigiosin 1 could bind HCO 3 − in solution.
In addition to showing evidence for binding, Davis and colleagues also demonstrated that prodigiosin 1 was a potent transmembrane transporter of HCO 3 − using independent assays. Earlier studies by Seganish and Davis had shown that prodigiosin 1 could facilitate the anion exchange of nitrate and bicarbonate in phospholipid membranes [62]. Although nitrate and bicarbonate anions have similar sizes and shapes, bicarbonate is more hydrophilic than nitrate and thus it is more challenging to transport bicarbonate across a lipid bilayer [71]. In the first set of experiments, phospholipid vesicles filled with NaCl solution were suspended in a sulfate-containing buffer and prodigiosin 1 was added to the solution (see Fig. 25). The efflux of Cl− anion from the liposomes was monitored using a Cl−-selective electrode and under these conditions there was no significant movement of Cl− from within the liposomes. After 2 min, a solution of sodium bicarbonate was added to the extravesicular solution and it was immediately evident that this addition turned on chloride efflux, suggesting that prodigiosin 1 could facilitate Cl−/HCO 3 − antiport exchange across these liposomal membranes.
The experiments depicted in Fig. 25 provided strong, yet indirect, evidence that prodigiosin 1 could transport bicarbonate across lipid membranes, since it was the Cl− and not the HCO 3 − anion that was being monitored. The authors next used 13C NMR spectroscopy to verify the prodigiosin-mediated transmembrane transport of HCO 3 −. In these NMR assays, the paramagnetic Mn2+ ion was used to bleach any 13C NMR signal that originated from extravesicular H13CO 3 − anion, whereas the 13C NMR signals for intravesicular H13CO 3 − would be unaffected by the membrane-impermeable Mn2+. Thus, these NMR experiments allowed for the direct discrimination of extravesicular and intravesicular H13CO 3 − and enabled the transmembrane movement of this anion to be firmly established (Fig. 26).
Results of chloride transport (measured using a chloride-selective electrode) that commenced when a bicarbonate pulse was added to the external solution. Below t = 120 s (i), chloride efflux is promoted by the addition of prodigiosin 1 (diamond) (0.04% molar carrier to lipid) to unilamellar POPC vesicles loaded with 451 mM NaCl and 20 mM phosphate buffer, pH 7.2, dispersed in 150 mM Na2SO4 and 20 mM phosphate buffer, pH 7.2. At t = 120 s, a solution of NaHCO3 was added to give a 40 mM external concentration. At t = 420 s, the vesicles were lysed with detergent and the final reading at t = 480 s was considered to equal 100% chloride efflux. Figure modified from [69]
Figure 26 shows data that illustrates prodigiosin-mediated HCO 3 − /Cl− exchange. EYPC liposomes (5 mM) filled with 450 mM NaCl were suspended in a sulfate solution and 50 mM NaH13CO3 was added to this solution. A sharp 13C NMR signal for the extravesicular H13CO 3 − was observed at δ 161 ppm. Addition of Mn2+ to this solution caused complete broadening of the 13C NMR signal for extravesicular bicarbonate. After the addition of prodigiosin 1, a sharp 13C NMR signal for H13CO 3 − reemerged due to ligand-mediated transport of bicarbonate into the liposome.
13C-NMR experiments demonstrate that prodigiosin 1 is able to facilitate chloride/bicarbonate antiport. (a) Titration sequence for monitoring the transmembrane transport of bicarbonate ions in H13CO 3 − loaded EYPC liposomes by prodigiosin 1 and DMSO. (b–d) 13C-NMR spectra (i) before and (ii) after the addition of the 50 mM NaCl pulse to EYPC vesicles loaded with 100 mM NaH13CO3, 20 mM HEPES buffer, pH 7.5, and dispersed in 75 mM Na2SO4, 20 mM HEPES buffer, pH 7.5; (iii) after the addition of prodigiosin 1 or DMSO (1, 0.1 mol%; DMSO, 870 mol%); (iv) after the addition of 0.5 mM Mn2+ (1:100 Mn2+/Cl− ratio). Figure modified from [69]
In this paper, the authors demonstrated that “small” molecules such as the natural product prodigiosin 1 could facilitate Cl−/HCO 3 − anion exchange, a process that is typically mediated by membrane proteins. The authors noted that this demonstration of Cl−/HCO 3 − antiport “…may well present an alternative mechanism by which prodigiosin 1 can influence biological systems” [69]. Furthermore, they proposed that synthetic Cl−/HCO 3 − antiporters based on prodigiosin’s tripyrrole framework might prove to be useful tools for biomembrane research.
14 Conclusions and Outlook
It is now clear from both the biochemical literature and from a variety of solid-state and solution studies that the protonated prodiginines are able to bind to anions. The resulting lipophilic ion pairs are also able to diffuse across the hydrophobic barrier that makes up phospholipid bilayer. In this way, the prodiginines are able to function as transmembrane anion transporters and, depending on the conditions, they can also function as de facto H+Cl− transporters. In particular, the ability to coordinate and transport both Cl− and HCO 3 − anions makes the prodiginines potentially valuable for mediating important biological processes that depend on the proper balance of these two essential anions. The author believes that the combination of organic synthesis and the use of biosynthetic machinery will, in the future, continue to produce new prodiginine analogs that are less cytotoxic but better able to be used as anticancer agents and other pharmaceuticals. The coming decade will undoubtedly reveal more about the mechanism of action of these pyrrole-based natural products and also reveal more about how their anion binding and anion transport abilities is translated into a biological function.
Notes
- 1.
The same group later reported similar studies as described in [14].
References
Bizio B (1823) Biblioteca Italiana o sia Giornale di Letteratura, Scienze e Arti Tomo 30:275–295
Merlino CP (1924) J Bacteriol 9:527–543
Hubbard R, Rimington C (1950) Biochem J 46:200–225
Gaughran ER (1969) Trans NY Acad Sci 31:3–24
Bennett JW (2000) Adv Appl Microbiol 47:1–32
Fürstner A (2003) Angew Chem Int Ed 42:3582–3603
Wrede IF, Hettche O (1929) Ber Deut Chem Ges 62:2678–2687
Wasserman HH, McKeon JE, Smith L, Forgione P (1960) J Am Chem Soc 82:506–507
Rapoport H, Holden KG (1962) J Am Chem Soc 84:634–642
Manderville RA (2001) Curr Med Chem Anticancer Agents 1:195–218
Perez-Tomas R, Montaner B, Llagostera E, Soto-Cerrato V (2003) Biochem Pharm 66:1447–1452
Williamson NR, Fineran PC, Gristwood T, Chawrai SR, Leeper FJ, Salmond GPC (2007) Future Microbiol 2:605–618
Pandey R, Chander R, Sainis KB (2009) Curr Pharm Design 15:732–741
Yamamoto C, Takemoto H, Kuno K, Yamamoto D, Tsubaru A, Kamata K, Hirata H, Yamamoto A, Kano H, Seki T, Inoue K (1999) Hepatology 30:894–902
Nakashima T, Tamura T, Kurachi M, Yamaguchi K, Oda T (2005) Biol Pharm Bull 28:2289–2295
Melvin MS, Ferguson DC, Lindquist N, Manderville RA (1999) J Org Chem 64:6861–6869
Melvin MS, Tomlinson JT, Saluta GR, Kucera GL, Lindquist N, Manderville RA (2000) J Am Chem Soc 122:6333–6334
Sessler JL, Eller LR, Cho WS, Nicolaou S, Aguilar A, Lee JT, Lynch VM, Magda DJ (2005) Angew Chem Int Ed 44:5989–5992
Baldino CM, Parr J, Wilson CJ, Ng SC, Yohannes D, Wasserman HH (2006) Bioorg Med Chem Lett 16:701–704
Fürstner A, Radkowski K, Peters H, Seidel G, Wirtz C, Mynott R, Lehmann CW (2007) Chem Eur J 13:1929–1945
Diaz RIS, Bennett SM, Thompson A (2009) Chem Med Chem 4:742–745
Clift MD, Thomson RJ (2009) J Am Chem Soc 131:14579–14583
For more information on Aida Pharmaceuticals and their development of prodigiosin 1 as an anti-cancer agent, see the website: http://www.crunchbase.com/company/aida-pharmaceuticals
Dairi K, Yao Y, Faley M, Tripathy S, Rioux E, Billot X, Rabouin D, Gonzalez G, Lavallee JF, Attardo G (2007) Org Process Res Dev 11:1051–1054
O’Brien SM, Claxton DF, Crump M, Faderl S, Kipps T, Keating MJ, Viallet J, Cheson BD (2009) Blood 113:299–305
D’Alessio R, Bargiotti A, Carlini O, Colotta F, Ferrari M, Gnocchi P, Isetta A, Mongelli N, Motta P, Rossi A, Rossi M, Tibolla M, Vanotti E (2000) J Med Chem 43:2557–2565
Stepkowski SM, Erwin-Cohen RA, Behbod F, Wang ME, Qu X, Tejpal N, Nagy ZS, Kahan BD, Kirken RA (2002) Blood 99:680–689
Williamson NR, Fineran PC, Leeper FJ, Salmond GPC (2006) Nat Rev Microbiol 4:887–899
Chawrai SR, Williamson NR, Salmond GPC, Leeper FJ (2008) Chem Commun:1862–1864
Gerber NN (1975) Prodigiosin-like pigments. CRC Crit Rev Microbiol 3:469–485
Kim D, Kim J, Yim JH, Kwon SK, Lee CH, Lee HK (2008) J Microbiol Biotech 18:1621–1629
Rizzo V, Morelli A, Pinciroli V, Sciangula D, D’Alessio R (1999) J Pharm Sci 88:73–78
Sertan-de Guzman AA, Predicala RZ, Bernardo EB, Neilan BA, Elardo SP, Mangalindan GC, Tasdemir D, Ireland CM, Barraquio WL, Concepcion GP (2007) FEMS Microbiol Lett 277:188–196
Blake AJ, Hunter GA, McNab H (1990) Chem Commun:734–736
La JQH, Michaelides AA, Manderville RA (2007) J Phys Chem B 111:11803–11811
Fürstner A, Grabowski J, Lehmann CW (1999) J Org Chem 64:8275–8280
Melvin MS, Tomlinson JT, Park G, Day CS, Saluta GR, Kucera GL, Manderville RA (2002) Chem Res Toxicol 15:734–741
Jenkins S, Incarvito CD, Parr J, Wasserman HH (2009) CrystEngComm 11:242–245
Chen K, Rannulu NS, Cai Y, Lane P, Liebl AL, Rees BB, Corre C, Challis GL, Cole RB (2008) J Am Soc Mass Spectrom 19:1856–1866
Duarte HA, Duani H, De Almeida WB (2003) Chem Phys Lett 369:114–124
Skawinski WJ, Venanzi TJ, Venanzi CA (2004) A molecular orbital study of tambjamine E and analogues. J Phys Chem A108:4542–4550
Sessler JL, Weghorn SJ, Lynch V, Fransson K (1994) J Chem Soc Chem Commun:1289–1290
Sessler JL, Camiolo S, Gale PA (2003) Coord Chem Rev 240:17–55
Park G, Tomlinson JT, Melvin MS, Wright MW, Day CS, Manderville RA (2003) Org Lett 5:113–116
Nelson N, Perzov N, Cohen A, Hagai K, Padler V, Nelson H (2000) J Exp Biol 203:89–95
Sato T, Konno H, Tanaka Y, Kataoka T, Nagai K, Wasserman HH, Ohkuma S (1998) J Biol Chem 273:21455–21462
Ohkuma S, Sato T, Okamoto M, Matsuya H, Arai K, Katakoa T, Nagai K, Wasserman HH (1998) Biochem J 334:731–741
Konno H, Matsuya H, Okamoto M, Sato T, Tanaka Y, Yokoyama K, Katakoa T, Nagai K, Wasserman HH, Ohkuma S (1998) J Biochem 124:547–556
Matsuya H, Okamoto M, Ochi T, Nishikawa A, Shimizu S, Katakoa T, Nagai K, Wasserman HH, Ohkuma S (2000) Biochem Pharm 60:1855–1863
Lee MH, Kataoka T, Magae J, Nagai K (1995) Biosci Biotechnol Biochem 59:1417–1421
Kataoka T, Muroi M, Ohkuma S, Waritani T, Magae J, Takatsuki A, Kondo S, Yamasaki M, Nagai K (1995) FEBS Lett 359:53–59
Kawauchi K, Shibutani K, Yagisawa H, Nakatsuji S, Anzai H, Yokoyama Y, Ikegami Y, Moriyama Y, Hirata H (1997) Biochem Biophys Res Commun 237:543–547
Woo JT, Ohba Y, Tagami K, Sumitani K, Kataoka T, Nagai K (1997) Biosci Biotech Biochem 61:400–402
Tanigaki K, Sato T,Tanaka Y, Ochi T,Nishikawa A, Nagai K, Kawashima H, Ohkuma S(2002) FEBS Lett.524:37-42
Rea PA, Poole RJ (1993) Annu Rev Plant Physiol Plant Mol 44:157–180
Maeshima M, Nakayasu T, Kawauchi K, Hirata H, Shimmen T (1999) Plant Cell Physiol 40:439–442
Nakayasu T, Kawauchi K, Hirata H, Shimmen T (2000) Plant Cell Physiol 41:857–863
Lagadic-Gossmann D, Huc L, Lecureur V (2004) Cell Death Differ 11:953–961
Yamamoto C, Takemoto H, Kuno K, Yamamoto D, Nakai K, Baden T, Kamata K, Hirata H, Watanabe T, Inoue K (2001) Oncology 8:821–824
Francisco R, Pérez-Tomás R, Gimènez-Bonafé P, Soto-Cerrato V, Giménez-Xavier P, Ambrosio S (2007) Eur J Pharmcol 572:111–119
Kawauchi K, Tobiume K, Iwashita K, Inagaki H, Morikawa T, Shibukawa Y, Moriyama Y, Hirata H, Kamata H (2008) Biosci Biotechnol Biochem 72:1564–1570
Seganish JL, Davis JT (2005) Chem Commun:5781–5783
McNally BA, Koulov AV, Smith BD, Joos JB, Davis AP (2005) Chem Commun:1087–1089
Krasne S, Eisenman G, Szabo G (1971) Science 174:412–414
Kano K, Fendler JH (1978) Biochim Biophys Acta 509:289–299
Sáez Díaz RI, Regourd J, Santacroce PV, Davis JT, Jakeman DL, Thompson A (2007) Chem Commun:2701–2703
Gale PA, Light ME, McNally B, Navakhun K, Sliwinski KE, Smith BD (2005) Chem Commun:3773–3775
Gale PA (2005) Chem Commun:3761–3772
Davis JT, Gale PA, Okunola OA, Prados P, Iglesias-Sánchez JC, Torroba T, Quesada R (2009) Nat Chem 1:138–144
Stefayne D (1960) J Org Chem 25:1261–1262
Davis AP, Sheppard DN, Smith BD (2007) Chem Soc Rev 36:348–357
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2010 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Davis, J.T. (2010). Anion Binding and Transport by Prodigiosin and Its Analogs. In: Gale, P., Dehaen, W. (eds) Anion Recognition in Supramolecular Chemistry. Topics in Heterocyclic Chemistry, vol 24. Springer, Berlin, Heidelberg. https://doi.org/10.1007/7081_2010_29
Download citation
DOI: https://doi.org/10.1007/7081_2010_29
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-15443-0
Online ISBN: 978-3-642-15444-7
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)