Abstract
For over two decades, genetic studies have been used to assist in the conservation and management of both Greater Sage-grouse (Centrocercus urophasianus) and Gunnison Sage-grouse (C. minimus), addressing a wide variety of topics including taxonomy, parentage, population connectivity, and demography. The field of conservation genetics has been transformed by dramatic improvements in sequencing technology, facilitating genomic studies in many wildlife species. The quality and amount of data generated by genomic methods vastly exceed that of traditional genetic studies, allowing for increased precision in estimating genetic parameters of interest. Perhaps more importantly, genomic methods can provide insight into non-neutral evolution such as adaptive divergence. Here we recount the shift from genetic to genomic methods using two wildlife species of substantial conservation interest, focusing on the improved capabilities and advantages of genomic methods. For instance, reassessment of divergence in sage-grouse using genomic methods confirmed strong differentiation between the two species and revealed that a small population in the state of Washington was more genetically distinct than previously recognized. Further, new genomic resources and approaches have been used to identify a family of genes linked to local dietary adaptation suggesting that sage-grouse may possess digestive and metabolic adaptations that mitigate the effects of consuming plant secondary metabolites like those found in sagebrush. Genetic variation among populations in these gene regions is thought to be involved with local dietary adaptations, and therefore maintaining the tie between sage-grouse and the chemistry of local sagebrush may be an important management consideration. We posit that the integration of newly developed genomic resources combined with the vast wealth of ecological and behavioral data for sage-grouse has the potential to shed light on mechanistic relationships that ultimately are vital to the conservation and management of these species.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Adaptive genetic variation
- Centrocercus
- Conservation genetics
- Landscape genetics
- Whole-genome sequencing
1 Introduction
1.1 Background
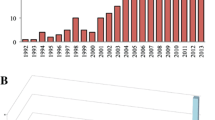
Molecular genetic methods (i.e., those methods involving a small number of anonymous and presumed neutral markers) have been used to address a wide variety of conservation and management issues for both Greater Sage-grouse (Centrocercus urophasianus) and Gunnison Sage-grouse (C. minimus). Both species have been well-studied from a genetic perspective with over 25 peer-reviewed publications in the past 20 years. The earliest research began in the mid-1990s examining taxonomy and distinct populations (Kahn et al. 1999; Oyler-McCance et al. 1999; Young et al. 2000; Benedict et al. 2003). Since those early papers, the range in topics tackled by genetic studies has been broad, examining questions ranging from lek formation and mating system (Gibson et al. 2005; Semple et al. 2001; Bird et al. 2012) to detecting gene flow and identifying landscape features impacting population connectivity (Bush et al. 2011; Oyler-McCance et al. 2005a, b; Cross et al. 2016; Row et al. 2015). Dramatic improvements in DNA sequencing technology (Mardis 2008; Shendure and Ji 2008; Metzker 2010) have facilitated the ability to collect genomic data for virtually any organism and the ability to parse anonymous versus putative adaptive genetic variation (Allendorf et al. 2010; Luikart et al. 2019; Hohenlohe et al. 2019). Such genomic approaches have recently been applied to sage-grouse, broadening our understanding about evolutionary history, current genomic structure, and potential adaptation – all of which are important for successful management and conservation. The aim of this chapter is to discuss the 20-year progression of molecular studies on Greater and Gunnison Sage-grouse, highlighting the expanded capabilities and advantages of genomic approaches and considering future research directions.
1.2 Conservation Status of Sage-Grouse
The distributions of both Greater and Gunnison Sage-grouse have contracted considerably across North America since the time of European settlement (Fig. 1). As of 2004, Greater Sage-grouse were thought to occupy roughly 56% of their historical distribution, while Gunnison Sage-grouse occupy only 10% (Schroeder et al. 2004). The causes of range contraction vary in different parts of the ranges yet likely involve habitat fragmentation, degradation, and loss associated with agriculture, resource extraction, livestock grazing, fencing, powerlines, invasive plants, and changes in the fire cycle (Connelly and Braun 1997; Braun 1998; Oyler-McCance et al. 2001; Knick et al. 2003; Connelly et al. 2004; Green et al. 2017; Monroe et al. 2017). Despite significant range contraction, the Greater Sage-grouse persists across much of the remaining western North American landscape that is dominated by sagebrush (Artemisia spp.). Some populations (e.g., in the states of Washington and Utah and the Jackson Hole population in the state of Wyoming) are small and isolated, while others persist in relatively continuous habitat (Fig. 1). Conservation and management efforts often cross state and federal boundaries and, by necessity, focus on large-scale processes. Both species have been petitioned to be listed under the US Endangered Species Act, with Gunnison Sage-grouse currently listed as threatened under US law (USFWS 2014) and Greater Sage-grouse listed as endangered under the Canadian Federal Species at Risk Act (Environment Canada 2014) for the northernmost populations in Alberta and Saskatchewan. The threat of potential listing has facilitated a plethora of studies on habitat requirements, population trends, impacts of management actions, and causes of decline, particularly for Greater Sage-grouse.
Current and presettlement distribution of sage-grouse (modified from Schroeder et al. 2004). The boundary for the Bi-State population of Greater Sage-grouse as well as the Washington populations is delineated by dotted lines, whereas the boundary for the Gunnison Sage-grouse distribution is delineated by a solid line. The numbers represent sampling locations for the whole-genome resequencing study (1, Alberta; 2, Jackson Hole; 3, Bi-State; 4, Washington; 5, Piceance Basin; 6, Gunnison Sage-grouse) of Oh et al. (2019)
1.3 Sagebrush Specialist
Sage-grouse are considered sagebrush obligate species (Beever and Aldridge 2011), depending on sagebrush throughout their entire life cycle (Patterson 1952). They require sagebrush for cover and nesting, and while they forage on sagebrush throughout the year, they rely on it exclusively for food in the winter months (Patterson 1952; Dalke et al. 1963; Wallestad et al. 1975; Braun et al. 1976; Connelly et al. 2000; Young et al. 2000). There are six main species and subspecies of sagebrush that are important for sage-grouse (described in Connelly et al. 2000), and their occurrence varies widely across the landscape due to differences in climate, soil type, topography, and disturbance (West 1983; Miller et al. 2011). Sagebrush leaves contain high levels of plant secondary metabolites (PSMs) such as monoterpenes (Kelsey et al. 1982) that act as a defense against herbivory by inhibiting digestive enzymes in herbivores (Kohl et al. 2015). Different varieties of sagebrush have distinct combinations and concentrations of toxins that vary across the landscape (Frye et al. 2013). Sage-grouse are dietary specialists and consume sagebrush leaves selectively, targeting leaves with higher nutrient content and lower concentration of PSMs (Remington and Braun 1985; Welch et al. 1988; Frye et al. 2013) and have coevolved mechanisms to deal with the inhibition of digestive enzymes associated with PSMs (Kohl et al. 2015).
1.4 Mating System
Both species of sage-grouse have a polygynous mating system that has been the focus of numerous studies over many decades (Wiley 1974; Wittenberger 1978; Gibson and Bradbury 1986; Bergerud 1988; Gibson et al. 1991; Young et al. 2000). In the spring, males congregate on leks, where they engage in an elaborate strutting display to attract females. Males establish territories on leks and defend them throughout the breeding season (Gibson and Bradbury 1986). Behavioral observations suggest that females arrive at leks later in the breeding season and typically mate with one of the dominant males on the lek (Wiley 1974; Gibson and Bradbury 1986; Gibson et al. 1991). Thus, reproductive success is highly variable among males, with a small proportion of males monopolizing all matings, which has important implications for management and conservation. Such highly skewed mating success among males implies strong sexual selection which can lead to rapid changes in morphology and behavior that can facilitate divergence and speciation (Ellsworth et al. 1994; Uy and Borgia 2000; Panhuis et al. 2001; Spaulding 2007; Oyler-McCance et al. 2010), processes that favor the formation of evolutionarily significant units that are important to identify and protect. Further, this mating skew decreases the overall genetic diversity and effective population size of sage-grouse (Stiver et al. 2008), which may be important for surviving and adapting to future stressors such as novel diseases or environmental change. Importantly, lekking behavior and the leks themselves provide a predictable time and location for biologists to count, monitor, and sample sage-grouse for research.
1.5 Why Sage-Grouse Are Good Candidates for Genetic and Genomic Research
Unlike many species of conservation concern, sage-grouse have been closely monitored for decades as they are game birds that have been sought by hunters and are therefore actively managed by governmental wildlife agencies. Research and monitoring efforts by these agencies have produced a wealth of data on population trends and habitat needs (see volume edited by Knick and Connely 2011), and concern over listing has prompted further research to better understand threats to sage-grouse and their habitat. This resulted in an abundance of information regarding sage-grouse populations, habitats, and threats, providing an extensive baseline into which molecular data can be integrated. The collection of samples for genetic analyses has been relatively straightforward as wings from hunter-killed grouse are collected each fall by most state agencies to determine demographic information, and DNA can be extracted from the muscle tissue of those wings (Oyler-McCance et al. 1999, 2005a; Benedict et al. 2003). In addition, blood collected from birds in radio telemetry studies has served as a good source of DNA (Oyler-McCance et al. 2005b, 2014; Bush et al. 2011). More recently, DNA extracted from feathers and fecal pellets collected noninvasively on leks during the breeding season has successfully been used in genetic analyses (Bush et al. 2005, 2010; Baumgardt et al. 2013; Row et al. 2015; Cross et al. 2016; Shyvers et al. 2019). While many genomic methods (e.g., whole-genome or reduced representation (re)sequencing) require relatively large quantities of high molecular weight DNA from tissue or blood (see Oyler-McCance et al. 2016 for a discussion of DNA quantity and quality in genomic applications), a few (e.g., targeted sequence capture; for review, see Jones and Good (2016)) have successfully generated genome-wide SNP markers using low-quality DNA samples such as from preserved museum specimens (Bi et al. 2013), suggesting that feather or fecal-derived DNA may be suitable for some genomic applications. Finally, sage-grouse are closely related to two agriculturally important galliform species (domestic turkey [Meleagris gallopavo; Dalloul et al. 2010] and chicken [Gallus gallus; International Chicken Genome Sequencing Consortium 2004]), thereby providing extensive genomic resources, including functional and structural genomic datasets and experimental validation that can serve as invaluable resources for assigning putative gene functions to sage-grouse orthologs.
2 How Traditional Genetics and the Shift to Genomics Help Conserve Sage-Grouse
2.1 Delineating Units for Conservation and Management
Historically, sage-grouse were considered to be one species. Research in the 1990s revealed dramatic morphological (Hupp and Braun 1991) and behavioral (Young et al. 1994) differences between sage-grouse in southwestern Colorado and southeastern Utah compared to the rest of the range, raising the possibility that this group of sage-grouse may be a new species. Genetic analyses using mitochondrial DNA and microsatellite loci were then employed to explore genetic differentiation between sage-grouse in northern and southern Colorado (Kahn et al. 1999; Oyler-McCance et al. 1999), comparing across the purported species boundary located within Colorado. These studies revealed a lack of gene flow between the two morphologically and behaviorally distinct groups of sage-grouse in Colorado, consistent with the idea that sage-grouse in southwestern Colorado were a distinct species. This new species was subsequently named, described (Young et al. 2000), and recognized by the American Ornithologists’ Union (2000). Further, these studies revealed that the newly described Gunnison Sage-grouse had much less genetic diversity than was found within Greater Sage-grouse in northern Colorado (Kahn et al. 1999; Oyler-McCance et al. 1999).
The recognition of the Gunnison Sage-grouse as a separate species led to the renaming of all other sage-grouse as Greater Sage-grouse and a correction to its range distribution (Fig. 1). Within the revised large range of the Greater Sage-grouse, the species had historically been divided into two taxonomic groups; an eastern (C. u. urophasianus) and a western (C. u. phaios) subspecies (Aldrich 1946). This delineation was based on plumage and coloration differences in 11 individual Greater Sage-grouse collected from Washington, Oregon, and California (Aldrich 1946). The western subspecies presumably occurred in southern British Columbia (Aldridge and Brigham 2003), central Washington, east-central Oregon, and northeastern California (Aldrich 1946), although Aldrich and Duvall (1955) considered the birds in California to be intermediate. Populations in other areas of the range were considered to be the eastern subspecies. The validity of the subspecies distinction was later questioned (Johnsgard 1983). Using genetic techniques similar to those used to evaluate the validity of the Gunnison Sage-grouse, Benedict et al. (2003) and Oyler-McCance et al. (2005a) examined the subspecific boundary and found no genetic evidence to support the original subspecies distinction (Fig. 1). Instead, they found several populations that were notable for other reasons. Along the border between Nevada and California, the “Bi-State” population (alternately referred to as “Lyon/Mono”) was found to be unusual, with mitochondrial DNA sequences largely unique compared to the rest of the range (Benedict et al. 2003). Further examination of the Bi-State population revealed that, unlike Gunnison Sage-grouse, they are neither morphologically nor behaviorally distinct from other Greater Sage-grouse (Taylor and Young 2006; Schroeder 2008). Interestingly, Benedict et al. (2003) also found that the two populations in Washington contained the lowest level of haplotype diversity observed (with one of the few haplotypes a common, widespread haplotype), perhaps resulting from a recent genetic bottleneck given that these populations now occupy just 8–10% of their original range and have shown significant declines in population size (Schroeder et al. 2000).
Managing populations with unique genetic diversity could be extremely important if those populations are to be conserved. Benedict et al. (2003) suggested that the unique allelic composition of the Bi-State population might be of particular importance for conservation. Since the likelihood that distinctiveness of anonymous genetic markers extends to genes under adaptive selection, they suggested this population should be managed independently, avoiding translocation of other Greater Sage-grouse into this area. They also surmised that the probable loss of genetic variation in Washington should be addressed, recommending that translocation of birds from neighboring populations may be justified to ensure continued persistence of the populations in Washington (Benedict et al. 2003). A subsequent study spanning the species’ entire range using both mitochondrial DNA and nuclear microsatellites came to similar conclusions (Oyler-McCance et al. 2005a).
Compared to other Greater Sage-grouse populations, the Bi-State population has a somewhat similar amount of genetic divergence (based on anonymous neutral markers) as the Gunnison Sage-grouse, yet it lacks the morphological and behavioral differences present between the two species (Taylor and Young 2006; Schroeder 2008; Oyler-McCance et al. 2014). This has led to lingering confusion over the taxonomic status of the Bi-State population. A shift to genomic markers has helped to resolve this taxonomic uncertainty. Using a reduced representation approach (RAD-Seq), Oyler-McCance et al. (2015a) identified over 11,000 single-nucleotide polymorphisms (SNPs) among three groups: Gunnison Sage-grouse, Bi-State, and the southern portion of the range of Greater Sage-grouse. Contrary to previous findings with microsatellites and mitochondrial DNA, they found much higher differentiation between Gunnison and Greater Sage-grouse than within Greater Sage-grouse (e.g., Bi-State population versus populations in the remainder of the species’ range). When each SNP site was mapped onto the chicken genome, the most highly divergent SNPs between Greater and Gunnison Sage-grouse were located on the Z chromosome (sex-determining macrochromosome in birds), and for both species, genetic diversity on the Z chromosome was reduced compared to autosomes (Oyler-McCance et al. 2015b). Greater divergence on the Z chromosome could be the result of selection (including sexual selection) or genetic drift associated with a genetic bottleneck related to the speciation event. These recent findings highlight the added value of genomic approaches, which help to better characterize patterns of genetic variation in sage-grouse and add insights into the mechanisms underlying speciation in these birds.
In light of these studies, there is ongoing interest in better understanding the genetic distinctiveness of sage-grouse populations, particularly those with small populations that exist on the margins of the species range, often constrained to relatively isolated patches of suitable sagebrush habitat that may limit gene flow from neighboring populations. From a conservation perspective, an important question is whether the patterns of genetic differentiation observed with anonymous markers in such populations have any functional genetic significance that might suggest local adaptation. While homology-based approaches provide a convenient option, species-specific genomic resources are necessary to fully characterize genetic variation observed. Recent efforts to enhance sage-grouse genomic studies have been bolstered by the de novo assembly of a high-coverage (ca. 170×) reference genome for Gunnison Sage-grouse (Oh et al. 2019). Comparative genomic analysis shows that 98% of scaffolds mapped with high confidence (e-value <1e-50) to the chicken genome, with all chicken chromosomes covered by three or more scaffolds. Preliminary genome annotation was also performed, utilizing both ab initio gene prediction and homology-based methods, resulting in a draft annotation containing 18,565 protein-coding genes. Taken together, this reference genome represents arguably the most comprehensive set of genomic resources available for a non-domesticated galliform species to date and was used to facilitate a whole-genome resequencing study (Oh et al. 2019) aimed at investigating anonymous and adaptive differentiation for several small, isolated, and potentially unique Greater Sage-grouse populations (Fig. 1): (1) at the northern extent of the contemporary species range in southeastern Alberta (Bush et al. 2011); (2) an isolated population near Jackson Hole, Wyoming (Schulwitz et al. 2014); (3) the Bi-State population (Benedict et al. 2003; Oyler-McCance et al. 2005a); and (4) a population in southern Washington (Benedict et al. 2003; Oyler-McCance et al. 2005a). Fifteen individual genomes were resequenced from each population, in addition to 15 samples from Greater Sage-grouse from the Piceance Basin in northwestern Colorado, which were expected to be more genetically representative of the largest populations occurring across relatively contiguous habitat of the Wyoming Basin (Oyler-McCance et al. 2005a). Fifteen Gunnison Sage-grouse samples were also included to evaluate previous interspecific genetic comparisons at a finer resolution. Analyses of approximately 1.5 million SNPs in the resulting dataset suggested distinct clustering by population (Fig. 2), with a largely hierarchical population structure, consistent with a pattern of postglacial recolonization from multiple refugia (Taberlet et al. 1998; Oyler-McCance et al. 2005a; Meirmans 2012). Evaluation of divergence at the whole-genome level (Oh et al. 2019) suggested greatest levels of differentiation at the interspecific level (mean pairwise FST at autosomal SNPs for C. urophasianus × C. minimus populations = 0.460), largely corroborating previous results. Interestingly, comparisons among Greater Sage-grouse populations indicated relatively high levels of divergence in pairwise contrasts involving Washington (mean pairwise FST at autosomal SNPs = 0.231) compared to the mean values among Greater Sage-grouse populations in the northeastern core of the species range (Alberta, Jackson Hole, and Piceance Basin: mean pairwise FST at autosomal SNPs = 0.103). The Bi-State population also showed comparatively elevated levels of genome-wide differentiation (mean pairwise FST at autosomal SNPs = 0.137). Importantly, because tests of population structure over relatively large geographic ranges can be biased by limited dispersal (i.e., isolation-by-distance), a partial Mantel test confirmed the evidence of genetic clustering, while controlling for interpopulation distance (Meirmans 2012). While evidence from previous genetic analyses of Washington sage-grouse has been consistent with a history of isolation and dramatic reductions in population size (Oyler-McCance et al. 2005a), these results at the whole-genome level provide new quantitative evidence for greater genetic distinctiveness of Washington birds than previously appreciated, which likely has implications for management priorities (Oh et al. 2019).
Principal component analysis of complete dataset representing five populations of Greater Sage-grouse (AL, Alberta, Canada; PI, Piceance Basin, Colorado; JH, Jackson Hole, Wyoming; BI, Bi-State population spanning the border between California and Nevada; WA, Washington) and the Gunnison Sage-grouse (GU in southwestern Colorado), based on 1,500,781 nuclear SNPs. Axes represent first (PC1) and second (PC2) principal components, with percentage of total genetic variance explained by each component shown in parentheses (Oh et al. 2019, reprinted with permission from Oxford University Press, Genome Biology and Evolution)
Whole-genome sequences were also used to investigate historical demographic trends in both sage-grouse species. Utilizing the Gunnison Sage-grouse genome along with a reference genome for Greater Sage-grouse (sequenced to a moderate depth of ~27× and then aligned to the C. minimus reference), Oh et al. (2019) used the pairwise sequentially Markovian coalescent model (Li and Durbin 2011) to infer changes in effective population size (Ne) over the past ~4 million years. Both species showed evidence of declines in Ne (Fig. 3), part of which coincides with the last glacial period in North America (c. 110,000–11,700 BP). However, while the Greater Sage-grouse genome revealed some evidence of population size stabilization (consistent with postglacial range expansion), the inferred Ne for Gunnison Sage-grouse exhibited consistent decline, suggesting that the ancestral population to this species may have been demographically isolated from other sage-grouse populations and undergone a more severe bottleneck, perhaps contributing to initial population divergence and the speciation process (Oh et al. 2019).
Inferred changes in ancestral effective population size for Greater Sage-grouse (blue) and Gunnison Sage-grouse (red) from Oh et al. (2019) (Reprinted with permission from Oxford University Press, Genome Biology and Evolution). Thick lines are median estimate from PSMC analysis of autosomes from a single individual of each species; lighter lines are from 100 bootstrap replicates. Values were scaled by generation time (g) (Stiver et al. 2008) and lineage-specific estimated mutation rate (μ) (Nam et al. 2010). The median divergence time for the two species is estimated to be roughly 1.24 million years ago (range 0.58–1.64 million years ago, Kumar et al. 2017; Jetz et al. 2012)
2.2 Population Connectivity and the Factors Influencing Gene Flow
Dispersal of individuals has important effects on population dynamics and persistence, as well as patterns of diversity and population structure (e.g., Garant et al. 2005; Row et al. 2010, 2016; Fedy et al. 2017). Thus, documenting effective dispersal (i.e., dispersal that results in gene flow) across landscapes can inform management actions on how to improve or maintain population connectivity. Genetic studies have examined gene flow at both large (Oyler-McCance et al. 2005a, b; Cross et al. 2018; Row et al. 2018) and small scales (Bush et al. 2011; Oyler-McCance et al. 2014; Schulwitz et al. 2014; Davis et al. 2015; Cross et al. 2016; Row et al. 2016). A common theme that emerges from these studies is that sage-grouse follow an isolation-by-distance pattern where neighboring populations tend to be more closely related than those that are separated by larger geographic distances (i.e., gene flow occurs more readily among neighboring populations). Further, populations in discrete patches of habitat isolated from other populations in more continuous sagebrush (e.g., satellite populations of Gunnison Sage-grouse, Washington, Jackson Hole, Bi-State populations within Greater Sage-grouse) are less connected than populations in more contiguous habitat (Oyler-McCance et al. 2005a, b; Schulwitz et al. 2014).
While documenting levels of gene flow among populations is an important first step, understanding how different landscapes actually influence gene flow provides a logical progression and can be critical for management and prioritization of areas for protection. The field of landscape genetics combines landscape modelling and genetic data to better comprehend how landscape features influence gene flow across a given region (Manel et al. 2003; Storfer et al. 2007; Forester et al. 2018). Several studies have used microsatellite data to examine such relationships in both Greater and Gunnison Sage-grouse at vastly different extents (Shirk et al. 2015; Row et al. 2015, 2018; Zimmerman 2019). Landscape features that impact gene flow in Greater Sage-grouse are scale-dependent and vary across the range (Row et al. 2018). Similarly, the habitat composition also had a scale-dependent facilitation of gene flow for Gunnison Sage-grouse, with the presence of sagebrush habitat facilitating gene flow among populations and high-quality nesting habitat and a tall shrub component facilitating gene flow among leks within the largest population (Zimmerman 2019). In general, sage-grouse gene flow tends to be greatest in areas of high-quality breeding habitat, yet conifers, rugged terrain, and agriculture impeded gene flow in many areas (Shirk et al. 2015; Row et al. 2015, 2018; Zimmerman 2019). Thresholds can be identified for the amount of breeding habitat or other important variables (positive or negative) that might influence gene flow (see Row et al. 2018), providing guidance on how to best manage landscapes to promote connectivity and gene flow. Genomic methods have the potential to add precision to landscape genetic studies due to the large number of markers. For instance, Jahner et al. (2016) analyzed variation at 27,866 SNPs in 140 male Greater Sage-grouse in a small region in central Nevada and found that geographic distance and suitable habitat best predicted genetic differentiation. Landscape genetic studies have produced maps that depict the strength and redundancy of connectivity that can help inform conservation actions that maintain and restore functional connectivity for sage-grouse. The added precision from genomic studies could further refine such efforts (Forester et al. 2018). Moreover, genomic methods could greatly expand the types of landscape genetic research questions being asked for sage-grouse by including adaptive loci. For instance, ties between adaptive genetic loci and environmental gradients could be examined (Waits and Storfer 2016) and used to predict potential responses to changing habitats under differing climate change scenarios.
Genetic data have frequently been used to estimate diversity within and differentiation between populations. Although one key feature of genomic data is being able to evaluate functional genetic regions, using thousands of anonymous loci can increase the precision of population parameter estimates (Allendorf et al. 2010). For instance, Gunnison Sage-grouse samples have been used to compare population parameter estimates from two datasets, one composed of 22 microsatellite loci from 254 individuals across populations and another composed of 14,072 SNP loci from 60 individuals (a subset of the 254) across populations (Zimmerman et al. 2019b). Both datasets generally showed the same pattern of differentiation, diversity, and clustering, although the SNP data had some increased precision of estimates and identification of distinct groups, as expected (Fig. 4). However, this increased precision was not always realized with differentiation metrics (see FST; Fig. 4). As other studies have demonstrated (Willing et al. 2012; Defaveri et al. 2013), precision in bootstrapped confidence intervals for pairwise differentiation is impacted by how many SNPs are used in combination with the number of individuals sampled for each population (more of both results in greater precision).
Increased precision in Gunnison Sage-grouse population genetic metrics for 14,072 putatively neutral SNP loci from 60 individuals versus 22 microsatellite loci from 254 individuals for multiple measures (Figure created from data presented in Zimmerman et al. 2019b). For (a) differentiation metrics (FST (Weir and Cockerham 1984); DJost (Jost 2008); GST (Hedrick 2005); calculated in diveRsity R package (Keenan et al. 2013) with 1,000 bootstraps) and (b) diversity metrics (AR = allelic richness, HE = expected heterozygosity, FIS = inbreeding coefficient; calculated in diveRsity R package), increased precision is illustrated as the difference in 95% confidence interval width for estimates calculated from microsatellites and SNPs (>0 indicates SNPs have a smaller width). Populations in pairwise comparisons for differentiation metrics (a) are abbreviated along the x-axis: CM Cimarron, CR Crawford, DC Dove Creek, GB Gunnison Basin, PM Piñon Mesa, SM San Miguel; CM.CR FST between Cimarron and Crawford. Clustering approaches (hclust function in R, the complete method and with dissimilarity matrix (Nei and Kumar 2000) and 1,000 bootstraps) show a clear increase in precision of identifiable groups when using SNPs (d) as opposed to microsatellites (c)
2.3 Managing Genetic Diversity
Small and isolated populations often face a risk of severe inbreeding and the resulting expression of deleterious recessive alleles. Additionally, for species with ranges that span diverse habitats, natural selection may promote highly localized adaptations that could warrant consideration when setting conservation priorities or contemplating certain management practices such as translocation or captive breeding. Thus, an important challenge for conservation genetics is to balance the maintenance of genetic diversity with the retention of potentially locally adapted genetic variants. The transition to genomic studies in sage-grouse conservation has provided important advances toward this goal.
Previous genetic analyses revealed low genetic diversity in Gunnison Sage-grouse satellite populations in comparison to the larger Gunnison Basin population (Oyler-McCance et al. 2005b). One of the management actions taken to mitigate both population size and genetic diversity was to translocate individuals from Gunnison Basin to satellite populations (Fig. 5). Recently, genetic data from 22 microsatellite loci were used to estimate change in diversity, differentiation, and population admixture among samples collected before, and 9 years after translocation efforts began (Zimmerman et al. 2019a). Satellite populations that received translocated birds all had increased genetic diversity, decreased genetic differentiation from the larger Gunnison Basin population, and showed signals of population admixture within individuals, indicating reproduction between Gunnison Basin transplants and resident satellite population birds. Though this work was completed using microsatellite loci, large numbers of anonymous loci from genomic techniques would likely identify finer signatures of change as a result of translocation. For one of the datasets, Zimmerman et al. (2019a) used a large number of noninvasively collected genetic samples, which were low in quality and unsuitable for many genomic techniques. Once anonymous loci are identified from high-quality samples, additional samples collected noninvasively could be used to continue tracking change as a result of the management action.
Range of Gunnison Sage-grouse in southwestern Colorado and southeastern Utah (modified from Zimmerman et al. 2019a). The largest (core) population is the Gunnison Basin. All other populations are considered to be satellite populations. The blue arrows represent translocation of Gunnison Sage-grouse from the Gunnison Basin to satellite populations, and the numbers represent the number of birds that were translocated between 2005 and spring of 2014 (USFWS 2014). Although birds were translocated into the Cimarron population, none of those birds survived. Analysis of pre- and post-translocation genetic data generally revealed increased genetic variation in the satellite populations and a decrease in differentiation between satellites and the Gunnison Basin population (Zimmerman et al. 2019a)
Despite relatively close geographic proximity, Gunnison Sage-grouse satellite populations have relatively high levels of genetic differentiation (Oyler-McCance et al. 2005b) with conspicuous differences in habitat quantity, quality, and ecological composition (GSRCC 2005) leading to the possibility of local adaptation across populations. Understanding the underlying genetic basis of such adaptations could be important for implementing conservation measures (Savolainen et al. 2013) such as translocations. The small effective population size of satellite populations could present a risk of translocations overwhelming any locally adapted alleles with variation from the larger Gunnison Basin. Zimmerman et al. (2019b) used 15,033 SNP genotypes in genomic outlier analyses, genotype-environment associations, and gene ontology (GO) enrichment analyses to examine patterns of putatively adaptive genetic differentiation in six of the Gunnison Sage-grouse populations. A total of 411 loci linked to 289 putative genes associated with biological functions that were overrepresented in the assemblage of outliers were identified. Of particular interest was the identification of candidate SNPs linked to four genes which are members of the cytochrome P450 gene family (CYP4V2, CYP2R1, CYP2C23B, CYP4B1) which could indicate adaptive divergence for genes involved in sagebrush PSM metabolism and candidate loci that were linked to genes potentially involved in antiviral response (DEAD box helicase gene family and SETX) (Zimmerman et al. 2019b). Additionally, seven of the candidate SNPs corresponded to predicted non-synonymous amino acid substitutions in putative genes; this included two putative genes associated with antiviral response (DDX60 and SETX), as well as one (CYB5R4) that was previously associated with heat stress response (Zimmerman et al. 2019b; Zimmerman 2019).
Genomic methods have also been applied to investigate potentially adaptive genetic variation in small and isolated Greater Sage-grouse populations (Oh et al. 2019). Utilizing the whole-genome resequencing datasets (see above), a population genomics study was carried out to identify SNPs that bear the signature of selection (Oh et al. 2019). Briefly, the analysis utilized a Bayesian method that first estimates overall genetic covariance among populations and then identifies outlier loci that deviate from the expectations of this background population structure (Gautier 2015). This analysis identified 8,630 outlier SNPs that exhibited extreme differentiation among populations (i.e., exceeded 1% probability threshold). Of these, 147 SNPs occurred within exons of predicted protein-coding genes, with 50 identified as causing non-synonymous changes. Another 2,099 SNPs occurred within 5 kb up- or downstream regions of genes, thus suggesting potential regulatory effects on nearby genes. Gene ontology analysis of predicted genes associated with outlier SNPs revealed participation in diverse organismal functions, including spermatogenesis (HOOK1, MYCBP-associated protein) and immune function (CFI, GAB3), suggesting a potential role of sexual and pathogen-mediated natural selection, respectively, in shaping patterns of protein variation. In a parallel approach, the same study tested for evidence of positive natural selection on cytochrome P450 genes, along with a panel of candidate genes that are likely related to metabolism of PSMs, identified from pharmacological literature. Multiple genomic regions containing outlier SNPs that were associated with candidate genes related to the metabolism of xenobiotic compounds were identified, suggesting that interpopulation variation could underlie consequential local dietary adaptations (Oh et al. 2019). These potential links between sage-grouse and the chemistry of the local sagebrush plants within which they reside are highly relevant to consider for conservation and management strategies. For instance, sagebrush restoration efforts could consider using local sagebrush material to avoid mismatches in PSMs with the local sage-grouse population. Additionally, it may be prudent for translocation efforts to carefully consider the sagebrush communities associated with source and recipient sage-grouse populations.
3 Future Directions in Sage-Grouse Genomics
3.1 Identifying and Conserving Adaptive Genetic Variation
The genetic and genomic research described above highlights the many ways that molecular data have contributed to the management of sage-grouse. The new genomic resources available for both species of sage-grouse greatly expand the types of questions that can now be answered, with a particular focus on understanding and conserving adaptation. Given the recently discovered relationship between specific sagebrush varieties and the sage-grouse that coevolved with them, genomic methods could further explore this relationship. A comprehensive analysis of diet, for example, can be obtained using metabarcoding approaches (Jarman et al. 2004; Deagle et al. 2009; Pompanon et al. 2012) and is particularly compelling as it can be completed noninvasively through analysis of fecal pellets. Moreover, the role of sage-grouse gut microbiome in metabolizing PSMs in sagebrush may be important (Kohl et al. 2015) and could be further investigated using genomic techniques. Adaptive genetic variation can also be identified by testing for associations between genomic variation and environmental variables important for sage-grouse. As the range of Greater Sage-grouse remains large, encompassing a wide variety of habitat and environmental conditions, this type of analysis could be particularly useful. Finally, genomic approaches can provide insights into the susceptibility of sage-grouse to disease. Rudimentary exploration of genetic diversity at immune genes has shown that both species of sage-grouse have lower levels of diversity compared to other prairie grouse and that Gunnison Sage-grouse is particularly low (Minias et al. 2016, 2018). Newly developed genomic resources for both species (Oh et al. 2019) should facilitate the expansion of this line of research.
3.2 Exploring the Impacts of Low Genetic Diversity
Both species of sage-grouse have experienced significant population declines that can result in loss of genetic diversity, which may decrease evolutionary adaptive potential and increase the likelihood of inbreeding depression (Allendorf et al. 2010; Steiner et al. 2013). Issues with low genetic diversity and inbreeding depression have been documented in a close relative, the greater prairie-chicken (Tympanuchus cupido; Westemeier et al. 1998), and was suspected in at least one population of Gunnison Sage-grouse (Stiver et al. 2008). Although the range and overall number of Greater Sage-grouse (>100,000 individuals) are relatively large, some populations (e.g., two populations in Washington and one in Strawberry Valley, Utah) show low levels of genetic diversity (Oyler-McCance et al. 2005a; Oh et al. 2019). Gunnison Sage-grouse, on the other hand, have a small and fragmented range, small number of individuals (<5,000), and much lower genetic diversity (Oyler-McCance et al. 2015a; Oh et al. 2019), which may make impacts of low genetic diversity more problematic. Genomic methods can elucidate the underlying genetic basis of inbreeding depression and provide a mechanistic link between phenotypes and the molecular processes behind them (Steiner et al. 2013). Identifying genes that contribute to inbreeding depression can be achieved through genome-wide association studies comparing genomic variation of individuals with different fitness levels. This approach is currently being investigated in captive Attwater’s prairie-chicken (T. cupido attwateri, J Johnson, pers. comm) and could potentially be useful in sage-grouse. Captive breeding of both species of sage-grouse has been attempted (Pyrah 1964; Johnson and Boyce 1991; Thompson et al. 2015; Apa and Wiechman 2015) and potentially could be used as a tool to augment wild sage-grouse populations as is taking place in Canada (D McKinnon, pers. comm), increasing both the size and genetic diversity of populations. Genomic analyses can provide more precise estimates of relatedness and inbreeding coefficients both in captive and wild populations that may assist in release, translocation, and genetic rescue efforts (Kardos et al. 2015).
3.3 Document and Better Understand Physiological Response to Stress
To date, an understanding of how sage-grouse respond to stress has relied on measuring corticosteroid metabolites in fecal samples (Jankowski et al. 2009, 2014; Blickley et al. 2012). Genomic methods have the potential to expand such investigations through gene expression experiments involving transcriptome sequencing. Such investigations could examine the physiological response of sage-grouse to biotic and abiotic stressors that occur both naturally (e.g., seasonal changes in temperature) and due to anthropogenic origin (e.g., noise) (Kleist et al. 2018). By providing measures of relative changes in gene expression in response to exposure to stressors, these analyses can yield insight not only into the molecular basis of these responses but may also serve as biological indicators for monitoring ecosystem health (Isaksson 2015). The main limitation for gene expression studies is that they require systems that are amenable to experimental manipulation, something that has proven to be difficult for sage-grouse. Improvements in our ability to maintain sage-grouse in captivity from captive breeding programs may provide an avenue to move forward with such experimental studies, such as testing adaptability to different food resources and thus variation in sagebrush PSMs.
3.4 Incorporate Genomic Data into Comprehensive Monitoring Programs
Baseline microsatellite data across the range of both species have been collected (Zimmerman et al. 2019a; Cross et al. 2018; Row et al. 2018), providing current information on connectivity among populations and levels of anonymous genetic diversity within them. While these microsatellite datasets provide useful information, reliance on these markers for future monitoring may not be ideal for several reasons. First, microsatellite allelic variation is based on fragment size, typically estimated from electrophoretic methods, which often vary among different techniques and conditions, thereby requiring constant standardization across instruments and laboratories, and may sometimes be influenced by subjectivity in scoring. Second, while microsatellite markers may be useful for documenting connectivity and levels of anonymous genetic diversity, they are typically not useful alone for identifying and monitoring adaptive genetic variation that may be important for conservation efforts. Finally, genomic genotyping methods are likely now less expensive and more repeatable than traditional microsatellite genotyping. Thus, a standardized set of genome-wide SNPs representing both anonymous and adaptive processes could be developed from existing genomic resources and archived samples and used as a baseline for future monitoring programs.
In the past, monitoring of sage-grouse populations relied on yearly lek counts in the field and analyses of trends that were tracked through time by individual states. Recent more comprehensive monitoring efforts now coordinate monitoring at different hierarchical scales (e.g., lek, lek cluster, region, or management zone) across the range of Greater Sage-grouse (Coates et al. 2017; Edmunds et al. 2017). These efforts identify when a lek or lek cluster is declining, identifying when trends deviate from a broader-scale pattern(s), and ultimately will link causal mechanisms to those declines which will identify potential management actions. Genomic monitoring could be incorporated into such programs as feathers could be collected periodically from a subset of leks and analyzed to watch for changes in connectivity, isolation of populations, or loss of anonymous or adaptive genetic diversity, evaluating potential links to population trends.
4 Conclusion
Information from genetic studies has informed conservation and management of sage-grouse for nearly two decades, addressing a broad variety of questions from taxonomy and gene flow to investigations of mating systems and unique identification of individuals for demographic analyses. Genomic methods, however, can build significantly on these foundations, greatly expanding the types of questions that can now be addressed. Novel genomic techniques coupled with the recently developed genomic resources for sage-grouse facilitate more precise estimates of parameters of interest (e.g., gene flow, inbreeding coefficients) and provide a more comprehensive understanding of the genetic basis of adaptation in sage-grouse. The integration of these new genomic resources with existing ecological and behavioral data for sage-grouse promises to shed light on mechanistic relationships that ultimately are vital for the conservation and management of these species.
References
Aldrich JW. New subspecies of birds from western North America. Proc Biol Soc Wash. 1946;59:129–36.
Aldrich JW, Duvall AJ. Distribution of American gallinaceous game birds. USFWS Circular 34, Washington, DC; 1955.
Aldridge CL, Brigham RM. Distribution, status and abundance of greater sage-grouse, Centrocercus urophasianus, in Canada. Can Field Nat. 2003;117:25–34.
Allendorf FW, Hohenlohe PA, Luikart G. Genomics and the future of conservation genetics. Nat Rev Genet. 2010;11:697–709.
American Ornithologist’s Union. Forty-second supplement to the American Ornithologists Union check-list for North American birds. Auk. 2000;117(3):847–58.
Apa AD, Wiechman LA. Captive-rearing of Gunnison sage-grouse from egg collection to adulthood to foster proactive conservation and recovery of a conservation-reliant species. Zoo Biol. 2015;34:438–52.
Baumgardt JA, Goldberg CS, Reese KP, Connelly JW, Musil DD, Garton EO, Waits LP. A method for estimating sex ratio for sage-grouse using noninvasive genetic samples. Mol Ecol Resour. 2013;13:393–402.
Beever EA, Aldridge CL. Influences of free-roaming equids on sagebrush ecosystems, with a focus on greater sage-grouse. In: Knick ST, Connelly JW, editors. Greater sage-grouse: ecology and conservation of a landscape species and its habitats, Studies in avian biology, vol. 38. Berkley: University of California Press; 2011. p. 273–90.
Benedict NG, Oyler-McCance SJ, Taylor SE, Braun CE, Quinn TW. Evaluation of the eastern (Centrocercus urophasianus urphasianus) and western (Centrocercus urophasianus phaios) subspecies of Sage-grouse using mitochondrial control-region sequence data. Conserv Genet. 2003;4:201–310.
Bergerud AT. Mating systems in grouse. In: Bergerud AT, Gratson MW, editors. Adaptive strategies and population ecology of northern grouse. Minneapolis: University of Minnesota Press; 1988. p. 439–72.
Bi K, Linderoth T, Vanderpool D, Good JM, Nielsen R, Moritz C. Unlocking the vault: next-generation museum population genomics. Mol Ecol. 2013;22:6018–32.
Bird KL, Aldridge CL, Carpenter JE, Paszkowski CA, Boyce MS, Coltman DW. The secret sex lives of sage-grouse: multiple paternity and intraspecific nest parasitism revealed through genetic analysis. Behav Ecol. 2012;24:29–38.
Blickley JL, Word KR, Krakauer AH, Phillips JL, Sells SN, Taff CC, Wingfield JC, Patricelli GL. Experimental chronic noise is related to elevated fecal corticosteroid metabolites in lekking male greater sage-grouse (Centrocercus urophasianus). PLOS One. 2012;7(11):e50462. https://doi.org/10.1371/journal.pone.0050462.
Braun CE. Sage grouse declines in western North America: what are the problems? Proc West Assoc of State Fish Wildl Agencies. 1998;78:139–56.
Braun CE, Baker MF, Eng RL, Gashwiler JS, Schroeder MH. Conservation committee report on effects of alteration of sagebrush communities on the associated avifauna. Wilson Bull. 1976;88:165–71.
Bush KL, Vinsky MD, Aldridge CL, Paszkowski CA. A comparison of sample types varying in invasiveness for use in DNA sex determination in an endangered population of greater sage-grouse (Centrocercus urophasianus). Conserv Genet. 2005;6:867–70.
Bush KL, Aldridge CL, Carpenter JE, Paszkowski CA, Boyce MS, Coltman DW. Birds of a feather do not always lek together: genetic diversity and kinship structure of greater sage-grouse (Centrocercus urophasianus) in Alberta. Auk. 2010;127:343–53.
Bush KL, Dyte CK, Moynahan BJ, Aldridge CL, Sauls HS, Battazzo AM, Walker BL, Doherty KE, Tack J, Carlson J, Eslinger D, Nicholson J, Boyce MS, Naugle DE, Paszkowski CA, Coltman DW. Population structure and genetic diversity of greater sage-grouse (Centrocercus urophasianus) in fragmented landscapes at the northern edge of their range. Conserv Genet. 2011;12:527–42.
Coates PS, Prochazka BG, Ricca MA, Wann GT, Aldridge CL, Hanser SE, Doherty KE, O’Donnell MS, Edmunds DE, Espinosa SP. Hierarchical population monitoring of greater sage-grouse (Centrocercus urophasianus) in Nevada and California – identifying populations for management at the appropriate spatial scale. U.S. Geological Survey Open-File Report 2017-1089.2017.
Connelly JW, Braun CE. Long-term changes in sage grouse Centrocercus urophasianus populations in western North America. Wildl Biol. 1997;3:229–34.
Connelly JW, Schroeder MA, Sands AR, Braun CE. Guidelines to manage sage grouse populations and their habitats. Wildl Soc Bull. 2000;28:967–85.
Connelly JW, Knick ST, Schroeder MA, Stiver SJ. Conservation assessment of greater sage-grouse and sagebrush habitats. Western Association of Fish and Wildlife Agencies: Cheyenne; 2004.
Cross TB, Naugle DE, Carlson JC, Schwartz MK. Hierarchical population structure in greater sage-grouse provides insight into management boundary delineation. Conserv Genet. 2016;17:1417–33.
Cross TB, Naugle DE, Oyler-McCance SJ, Row JR, Fedy BC, Schwartz MK. The genetic network of greater sage-grouse: range-wide identification of keystone hubs of connectivity. Ecol Evol. 2018;8:5394–412.
Dalke PD, Pyrah DB, Stanton DC, Crawford JE, Schlatterer EF. Ecology, productivity, and management of sage grouse in Idaho. J Wildl Manag. 1963;27:811–41.
Dalloul RA, Long JA, Zimin AV, Aslam L, Beal K, et al. Multi-platform next-generation sequencing of the domestic Turkey (Meleagris gallopavo): genome assembly and analysis. PLoS Biol. 2010;8:e1000475.
Davis DM, Reese KP, Gardner SC, Bird KL. Genetic structure of greater sage-grouse (Centrocercus urophasianus) in a declining, peripheral population. Condor Ornithol Appl. 2015;117:530–44.
Deagle BE, Kirkwood R, Jarman SN. Analysis of Australian fur seal diet by pyrosequencing prey DNA in faeces. Mol Ecol. 2009;18:2022–38.
Defaveri J, Viitaniemi H, Leder E, Merilä J. Characterizing genic and nongenic molecular markers: comparison of microsatellites and SNPs. Mol Ecol Resour. 2013;13:377–92.
Edmunds DR, Aldridge CL, O’Donnell MS, Monroe AP. Greater sage-grouse population trends across Wyoming. J Wildl Manag. 2017;82:397–412.
Ellsworth DL, Honeycutt RL, Silvy NJ, Rittenhouse KD, Smith MH. Mitochondrial-DNA and nuclear-gene differentiation in North American prairie grouse (genus Tympanuchus). Auk. 1994;111:661–71.
Environment Canada. Amended recovery strategy for the greater sage-grouse (Centrocercus urophasianus uophasianus) in Canada, Species at risk act recovery strategy series. Ottawa: Environment Canada; 2014.
Fedy BC, Row JR, Oyler-McCance SJ. Integration of genetic and demographic data to assess population risk in a continuously distributed species. Conserv Genet. 2017;18:89–104.
Forester BR, Landguth EL, Hand BK, Balkenhol N. Landscape genomics for wildlife research. In: Hohenlohe PA, Rajora OP, editors. Population genomics: wildlife. Cham: Springer International Publishing AG; 2018. https://doi.org/10.1007/13836_2018_56.
Frye GG, Connelly JW, Musil DD, Forbey JS. Phytochemistry predicts habitat selection by an avian herbivore at multiple spatial scales. Ecology. 2013;94:308–14.
Garant D, Kruuk LEB, Wilkin TA, McCleery RH, Sheldon BC. Evolution driven by differential dispersal within a wild bird population. Nature. 2005;433:60–5.
Gautier M. Genome-wide scan for adaptive divergence and association with population-specific covariates. Genetics. 2015;201:1555–79.
Gibson RM, Bradbury JW. Male and female mating strategies on sage grouse leks. In: Rubenstein DI, Wrangham RW, editors. Ecological aspects of social evolution. Princeton: Princeton University Press; 1986. p. 379–98.
Gibson RM, Bradbury JW, Vehrencamp SL. Mate choice in lekking sage grouse revisited: the roles of vocal display, female site fidelity and copying. Behav Ecol. 1991;2:165–80.
Gibson RM, Pires D, Delaney KS, Wayne RK. Microsatellite DNA analysis shows that greater sage-grouse leks are not kin groups. Mol Ecol. 2005;14:4453–9.
Green AW, Aldridge CL, O’Donnell MS. Investigating impacts of oil and gas development on greater sage-grouse. J Wildl Manag. 2017;81:46–57.
Gunnison Sage-grouse Rangewide Steering Committee (GSRSC). Gunnison sage-grouse rangewide conservation plan. Denver: Colorado Division of Wildlife; 2005.
Hedrick PW. A standardized genetic differentiation measure. Evolution. 2005;59:1633–8.
Hohenlohe PA, Hand BK, Andrews KR, Luikart G. Population genomics provides key insights in ecology and evolution. In: Rajora OP, editor. Population genomics: concepts, approaches and applications. Cham: Springer Nature Switzerland AG; 2019. p. 483–510.
Hupp JW, Braun CE. Geographic variation among sage grouse in Colorado. Wilson Bull. 1991;103:255–61.
International Chicken Genome Sequencing Consortium. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature. 2004;432:695–716.
Isaksson C. Urbanization, oxidative stress and inflammation: a question of evolving, acclimatizing or coping with urban environmental stress. Funct Ecol. 2015;29:913–23.
Jahner JP, Gibson D, Weitzman CL, Bloomberg EJ, Sedinger JS, Parchman TL. Fine scale genetic structure among greater sage-grouse leks in Central Nevada. BMC Evol Biol. 2016;16:127.
Jankowski MD, Wittwer DJ, Heisey DM, Franson JC, Hofmeister EK. The adrenocortical response of greater sage-grouse (Centrocercus urophasianus) to capture ACTH injection, and confinement, as measured in fecal samples. Physiol Biochem Zool. 2009;82:190–201.
Jankowski MD, Russell RE, Franson JC, Dusek RJ, Hines MK, Gree M, Hofmeister EK. Corticosterone metaboloite concentrations in greater sage-grouse are positively associated with the presence of cattle grazing. Rangeland Ecol Manag. 2014;67:237–46.
Jarman SN, Deagle BE, Gales NJ. Group-specific polymerase chain reaction for DNA-based analysis of species diversity and identity in dietary samples. Mol Ecol. 2004;13:1313–22.
Jetz W, Thomas GH, Joy JB, Hartmann K, Mooers AO. The global diversity of birds in space and time. Nature. 2012;491:444–8.
Johnsgard PA. The grouse of the world. Lincoln: University of Nebraska Press; 1983.
Johnson GD, Boyce MS. Survival, growth, and reproduction of captive reared sage grouse. Wildl Soc Bull. 1991;18:88–93.
Jones MR, Good JM. Targeted capture in evolutionary and ecological genomics. Mol Ecol. 2016;25:185–202.
Jost L. GST and its relatives do not measure differentiation. Mol Ecol. 2008;17:4015–26.
Kahn NW, Braun CE, Young JR, Wood S, Mata DR, Quinn TW. Molecular analysis of genetic variation among large- and small-bodied Sage Grouse using mitochondrial control-region sequences. Auk. 1999;116:819–24.
Kardos M, Luikart G, Allendorf FW. Measuring individual inbreeding in the age of genomics: marker-based measures are better than pedigrees. Heredity. 2015;115:63–72.
Keenan K, McGinnity P, Cross TF, Crozier WW, Prodohl PA. diveRsity: an R package for the estimation of population genetics parameters and their associated errors. Methods Ecol Evol. 2013;4:782–8.
Kelsey RG, Stephens JR, Shafizadeh F. The chemical constituents of sagebrush foliage and their isolation. J Range Manag. 1982;35:617–22.
Kleist NJ, Guralnick RP, Cruz A, Lowry CA, Francis CD. Chronic anthropogenic noise disrupts glucocorticoid signaling and has multiple effects on fitness in an avian community. Proc Natl Acad Sci. 2018;115:E648–57.
Knick ST, Connely JW. Greater sage-grouse: ecology and conservation of a landscape species and its habitats, Studies in avian biology, vol. 38. Berkley: University of California Press; 2011.
Knick ST, Dobkin DS, Rotenberry JT, Schroeder MA, Vander Haegen WM, van Ripper C III. Teetering on the edge or too late? Conservation and research issues for avifauna of sagebrush habitats. Condor. 2003;105:611–34.
Kohl KD, Pitman E, Robb BC, Connelly JW, Dearing MD, Forbey JS. Monoterpenes as inhibitors of digestive enzymes and counter-adaptations in a specialist avian herbivore. J Comp Physiol B. 2015;185:425–34.
Kumar S, Stecher G, Suleski M, Hedges SB. Timetree: a resource for timelines, timetrees, and divergence times. Mol Biol Evol. 2017;4:1812–9.
Li H, Durbin R. Inference of human population history from individual whole-genome sequences. Nature. 2011;475:493–6.
Luikart G, Kardos M, Hand BK, Rajora OP, Aitken SN, Hohenlohe PA. Population genomics: advancing understanding of nature. In: Rajora OP, editor. Population genomics: concepts, approaches and applications. Cham: Springer Nature Switzerland AG; 2019. p. 3–79.
Manel S, Schwartz MK, Luikart G, Taberlet P. Landscape genetics: combining landscape ecology and population genetics. Trends Ecol Evol. 2003;18:189–97.
Mardis ER. Next-generation DNA sequencing methods. Annu Rev Genomics Hum Genet. 2008;9:387–402.
Meirmans PG. The trouble with isolation by distance. Mol Ecol. 2012;21:2839–46.
Metzker ML. Sequencing technologies – the next generation. Nat Rev Genet. 2010;11:31–46.
Miller RF, Knick ST, Pyke DA, Meike CW, Hanser SE, Wisdom MJ, Hild AL. Characteristics of sagebrush habitats and limitations to long-term conservation. In: Knick SJ, Connelly JW, editors. Greater sage-grouse: ecology and conservation of a landscape species and its habitats, Studies in avian biology, vol. 38. Berkley: University of California Press; 2011. p. 145–84.
Minias P, Bateson ZW, Whittingham LA, Johnson JA, Oyler-McCance SJ, Dunn PO. Contrasting evolutionary histories of MHC class I and class II loci in grouse – effects of selection and gene conversion. Heredity. 2016;116:466–76.
Minias P, Bateson ZW, Whittingham LA, Johnson JA, Oyler-McCance SJ, Dunn PO. Extensive shared polymorphism at non-MHC immune genes in recently diverged North American prairie-grouse. Immunogenetics. 2018;70:195–204.
Monroe AP, Aldridge CL, Assal TJ, Veblen KE, Pyke DA, Casazza ML. Patterns in greater sage-grouse population dynamics correspond with public grazing records at broad scales. Ecol Appl. 2017;27:1096–107.
Nam K, Mugal C, Nabholz B, Schielzeth H, Wolf JBW, Backström N, Künstner A, Balakrishnan CN, Heger A, Ponting CP, Clayton DF, Ellegren H. Molecular evolution of genes in avian genomes. Genome Biol. 2010;11:R68.
Nei M, Kumar S. Molecular evolution and phylogenetics. New York: Oxford University Press; 2000.
Oh KP, Aldridge CL, Forbey JS, Dadabay CY, Oyler-McCance SJ. Genomic insights into neutral and adaptive variation in sage-grouse: implications for ecology and conservation. Genome Biol Evol. 2019;11:2023–2034.
Oyler-McCance SJ, Kahn NW, Burnham KP, Braun CE, Quinn TW. A population genetic comparison of large- and small-bodied sage grouse in Colorado using microsatellite and mitochondrial DNA markers. Mol Ecol. 1999;8:1457–65.
Oyler-McCance SJ, Burnham KP, Braun CE. Influence of changes in sagebrush on Gunnison sage grouse in southwestern Colorado. Southwest Nat. 2001;46:323–31.
Oyler-McCance SJ, Taylor SE, Quinn TW. A multilocus population genetic survey of the greater sage-grouse across their range. Mol Ecol. 2005a;14:1293–310.
Oyler-McCance SJ, St. John J, Taylor SE, Apa AD, Quinn TQ. Population genetics of Gunnison sage-grouse: implications for management. J Wildl Manag. 2005b;69:630–7.
Oyler-McCance SJ, St. John J, Quinn TW. Rapid evolution in lekking grouse. Ornithol Monogr. 2010;67:114–22.
Oyler-McCance SJ, Casazza ML, Fike JA, Coates PS. Hierarchical spatial genetic structure in a distinct population segment of greater sage-grouse. Conserv Genet. 2014;15:1299–311.
Oyler-McCance SJ, Cornman RS, Jones KL, Fike JA. Genomic single-nucleotide polymorphisms confirm that Gunnison and greater sage-grouse are genetically well differentiated and that the bi-state population is distinct. Condor. 2015a;117:217–27.
Oyler-McCance SJ, Cornman RS, Jones KL, Fike JA. Z chromosome divergence, polymorphism and relative effective population size in a genus of lekking birds. Heredity. 2015b;115:452–9.
Oyler-McCance SJ, Oh KP, Langin KM, Aldridge CL. A field ornithologist’s guide to genomics: practical considerations for ecology and conservation. Auk Ornithol Adv. 2016;4:626–48.
Panhuis TM, Butlin R, Zuk M, Tregenza T. Sexual selection and speciation. Trends Ecol Evol. 2001;16:364–71.
Patterson RL. The sage grouse in Wyoming. Denver: Sage Books; 1952.
Pompanon F, Deagle BE, Symondson WOC, Brown DS, Jarman SN, Taberlet P. Who is eating what: diet assessment using next generation sequencing. Mol Ecol. 2012;21:1931–50.
Pyrah DB. Sage chickens in captivity. Game Bird Breed Pheas Fanciers Avicultur Gaz. 1964;13:10–1.
Remington TE, Braun CE. Sage grouse food selection in winter, North Park, Colorado. J Wildl Manag. 1985;49:1055–61.
Row JR, Blouin-Demers G, Lougheed SC. Habitat distribution influences dispersal and fine-scale genetic population structure of eastern foxsnakes (Mintonius gloydi) across a fragmented landscape. Mol Ecol. 2010;19:5157–71.
Row JR, Oyler-McCance SJ, Fike JA, O’Donnell MS, Doherty KE, Aldridge CL, Bowen ZH, Fedy BC. Landscape characteristics influencing the genetic structure of greater sage-grouse within the stronghold of their range: a holistic modeling approach. Ecol Evol. 2015;5:1955–69.
Row JR, Oyler-McCance SJ, Fedy BC. Differential influences of local subpopulations on regional diversity and differentiation for greater sage-grouse (Centrocercus urophasianus). Mol Ecol. 2016;25:4424–37.
Row JR, Doherty K, Cross T, Schwartz M, Oyler-McCance SJ, Naugle D, Knick ST, Fedy B. Quantifying functional connectivity: the role of breeding habitat, abundance, and landscape features on range-wide gene flow in sage-grouse. Evol Appl. 2018;11:1305–21.
Savolainen O, Lascoux M, Merila J. Ecological genomics of local adaptation. Nat Rev Genet. 2013;14:807–20.
Schroeder MA. Variation in greater sage-grouse morphology by region and population. Spokane: US Fish and Wildlife Service Report; 2008.
Schroeder MA, Hays DW, Livingston MF, Stream LE, Jacobson JE, Pierce DJ. Changes in the distribution and abundance of sage grouse in Washington. Northwest Nat. 2000;81:104–12.
Schroeder MA, Aldridge CL, Apa AD, Bohne JR, Braun CE, Bunnel SD, Connelly JW, Deibert PA, Gardner SC, Hilliard MA, Kobringer GD, McAdam SM, McCarthy CW, McCarthy JJ, Mitchell DL, Rickerson EV, Stiver SJ. Distribution of sage-grouse in North America. Condor. 2004;106:363–76.
Schulwitz S, Bedrosian B, Johnson JA. Low anonymous genetic diversity in isolated greater sage-grouse (Centrocercus urophasianus) populations in northwest Wyoming. Condor Ornith Appl. 2014;116:560–73.
Semple K, Wayne RK, Gibson RM. Microsatellite analysis of female mating behavior in lek-breeding sage grouse. Mol Ecol. 2001;10:2043–8.
Shendure J, Ji H. Next-generation DNA sequencing. Nat Biotech. 2008;26:1135–45.
Shirk AJ, Schroeder MA, Robb LA, Cushman SA. Empirical validation of landscape resistance models: insights from the greater sage-grouse (Centrocercus urophasianus). Landsc Ecol. 2015;30(10):1837–50. https://doi.org/10.1007/s10980-015-0214-4.
Shyvers JL, Walker BL, Oyler-McCance SJ, Fike JA, Noon BR. Genetic mark-recapture analysis of winter faecal pellets allows estimation of population size in Sage Grouse Centrocercus urophasianus. Ibis. 2019 (in press).
Spaulding A. Rapid courtship evolution in grouse (Tetraonidae): contrasting patterns of acceleration between the Eurasian and North American polygynous clades. Proc R Soc London B. 2007;274:1079–86.
Steiner CC, Putnam AS, Hoeck PEA, Ryder OA. Conservation genomics of threatened animal species. Annu Rev Anim Biosci. 2013;1:261–81.
Stiver JR, Apa AD, Remington TE, Gibson RM. Polygyny and female breeding failure reduce effective population size in the lekking Gunnison sage-grouse. Biol Conserv. 2008;141:472–81.
Storfer A, Murphy MA, Evans JS, Goldberg CS, Robinson S, Spear SF, Dezzani R, Delmelle E, Vierling L, Waits LP. Putting the “landscape” in landscape genetics. Heredity. 2007;98:128–42.
Taberlet P, Fumagalli L, Wust-Saucy AG, Cosson JF. Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol. 1998;7:453–64.
Taylor SE, Young JR. A comparative behavioral study of three greater sage-grouse populations. Wilson J Ornithol. 2006;118:36–41.
Thompson TR, Apa AD, Reese KP, Tadvick KM. Captive rearing sage-grouse for augmentation of surrogate wild broods: evidence for success. J Wildl Manag. 2015;79:998–1013.
USFWS. Endangered and threatened wildlife and plants; determination for the Gunnison Sage-grouse as a threatened or endangered species. Fed Regist. 2014;79:69192–310.
Uy JAC, Borgia G. Sexual selection drives rapid divergence in bowerbird display traits. Evolution. 2000;63:153–64.
Waits LP, Storfer A. Basics of population genetics: quantifying neutral and adaptive genetic variation for landscape genetic studies. In: Balkenhol N, Cushman SA, editors. Landscape genetics: concepts, methods, applications. Hoboken: Wiley; 2016. p. 35–57.
Wallestad R, Peterson JG, Eng RL. Foods of adult sage grouse in central Montana. J Wildl Manag. 1975;39:628–30.
Weir BS, Cockerham CC. Estimating F-statistics for the analysis of population structure. Evolution. 1984;38:1358–70.
Welch BL, Pederson JC, Rodriquez RL. Selection of big sagebrush by sage grouse. Great Basin Nat. 1988;48:274–9.
West NE. Western intermountain sagebrush steppe. In: West NE, editor. Ecosystems of the world, Temperate deserts and semi-deserts, vol. 5. New York: Elsevier; 1983. p. 331–49.
Westemeier RL, Brawn JD, Simpson SA, Esker TL, Jansen RW, Walk JW, Kershner EL, Bouzat JL, Paige KN. Tracking the long-term decline and recovery of an isolated population. Science. 1998;282:1695–8.
Wiley RH. Evolution of social organization and life-history patterns among grouse. Q Rev Biol. 1974;49:201–27.
Willing E-M, Dreyer C, van Oosterhout C. Estimates of genetic differentiation measured by FST do not necessarily require large sample sizes when using many SNP markers. PLOS One. 2012;7:1–7.
Wittenberger JF. The evolution of mating systems in grouse. Condor. 1978;80:126–37.
Young JR, Hupp JW, Bradbury JW, Braun CE. Phenotypic divergence of secondary sexual traits among sage grouse populations. Anim Behav. 1994;47:1353–62.
Young JR, Braun CE, Oyler-McCance SJ, Hupp JW, Quinn TW. A new species of sage-grouse (Pahsianidae: Centrocercus) from southwestern Colorado. Wilson Bull. 2000;112:445–53.
Zimmerman, SJ. Genetics, genomics, grouse, and conservation: use of genetic and genomic data to evaluate conservation actions and characterize popualtions of Gunnison Sage-grouse. PhD dissertation, Colorado State University, Fort Collins; 2019.
Zimmerman SJ, Aldridge CL, Apa AD, Oyler-McCance SJ. Evaluation of the genetic change from translocation among Gunnison Sage-grouse (Centrocercus minimus) populations. Condor. 2019a;121:1–14.
Zimmerman SJ, Aldridge CL, Oyler-McCance SJ. Signatures of adaptive divergence among populations of an avian species of conservation concern. Evol Appl. 2019b; https://doi.org/10.1111/eva.12825.
Acknowledgments
We thank M. Schroeder, J. Fike, and the editors of this volume for their helpful comments on this chapter and J. Saher for her help with figures. Any use of trade, firm, or product names is for descriptive purposes only and does not imply endorsement by the US Government.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 This is a U.S. government work and not under copyright protection in the U.S.; foreign copyright protection may apply
About this chapter
Cite this chapter
Oyler-McCance, S.J., Oh, K.P., Zimmerman, S.J., Aldridge, C.L. (2020). The Transformative Impact of Genomics on Sage-Grouse Conservation and Management. In: Hohenlohe, P.A., Rajora, O.P. (eds) Population Genomics: Wildlife. Population Genomics. Springer, Cham. https://doi.org/10.1007/13836_2019_65
Download citation
DOI: https://doi.org/10.1007/13836_2019_65
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-63488-9
Online ISBN: 978-3-030-63489-6
eBook Packages: Computer ScienceComputer Science (R0)