Abstract
In the recent years of development, deep learning (DL) is very useful in all fields with the growing availability of data. The main goal of DL technology is to make faster, reliable, and good decisions. Because of this ability, DL has found its use in healthcare, particularly with a focus on various types of medical images or images related to patients’ health. These fields have diagnostic processes that depend on gathering and processing large amounts of medical images. This work proposes a deep learning (DL) model based on “Convolutional Neural Network (CNN)” for identifying COVID-19 and Pneumonia using Chest X-Ray images. The result of this processing helps the radiologists to derive insights and make decisions to determine the correct diagnosis of the patient. This model helps in two ways. Firstly, to classify whether a chest X-ray shows any sort of variations with respect to COVID-19 and pneumonia or not. Secondly, to classify with the help of normal chest X-ray images. Experiments were carried out using InceptionV3 (IV3), and VGG16 models on the chest X-ray images. Results revealed that the VGG16 model outperformed IV3 model for COVID-19 and pneumonia identification on multiple performance metrics.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
The “Novel Coronavirus (2019-nCoV)” was originally discovered in Wuhan, China, in December 2019. By the end of March 2020, more than 200 nations worldwide recorded COVID-19 cases that had resulted in more than 7 lakh infections and 30,000+ fatalities when all the afflicted nations were included. Kids, organizations, and the general population are all seriously at risk from this fatal illness. The most prevalent pulmonary lung disease that affects kids most frequently is pneumonia. Pneumonia sickness is frequently contracted by kids under the age of five. Pneumonia puts approximately 7 lakh kids at risk of dying annually. It affects approximately more than 7% of the global total. Even for highly experienced radiologists, it has proven difficult to determine from chest X-rays if a patient has COVID or pneumonia [1].
Therefore, computerized technology is required to forecast these illnesses in individuals. As a result, in this work, it is suggested to use a model that is developed on digitized chest X-ray imagery to effectively diagnose COVID-19 and pneumonia. The decision-making process for radiologists or doctors may be aided by this. A method utilizing various DL models is shown, in which the IV3 and VGG16 models are used to effectively diagnose both disorders. In this supervised learning method, CNN forecasts the outcomes. Appropriate data enhancement strategies are employed to equitably expand the training set. Finally, the model’s efficiency is assessed to determine the extent to which the imagery was properly anticipated. Consequently, the trained model is frequently employed for quick identification of COVID-19 and pneumonia, aiding radiologists in their diagnostic procedure.
1.1 Background
In recent years, deep learning (DL) research has increasingly focused on “computer-aided designs (CAD)”. The current CAD system has already demonstrated its ability to aid the healthcare, particularly regarding the diagnosis of chest radiography, mammography, cancer, etc. While applying the DL approaches to medical scans the critical aspects are essential. Because of this, most earlier methods favored employing custom attributes when creating Software’s that permitted looking at images. However, due to inherent volatility, the hand-crafted components faced restrictions in terms of offering useful functions. In order to derive usable characteristics for image analysis, DL models, particularly CNNs, have shown themselves to be quite valuable [2].
There are transfer learning methods where pre-trained CNN models learn generic information and features on huge scale datasets like ImageNet which are then transferred to the required task. The process of feature extraction requires this type of transfer learning methods. To get a quality feature extraction, availability of pre-trained CNN models like VGG-16 and IV3 play a very important role. In addition to this, the classification used with high-rich extracted features shows improved performance in classifying images [3].
1.2 Research Motivation
The WHO has declared COVID-19 to be a pandemic since it has spread to more than 150 countries around the world. By the completion of the second week in April 2020, there were approximately 19 lakhs confirmed cases of COVID-19, greater than 1 lakh fatalities reported worldwide. Likewise, in the modern era, pneumonia has become the leading cause of death in children all over the world. A WHO survey revealed that each year, pneumonia claims the lives of millions of kids under the age of seven [4]. As a result, it has emerged as one of the major causes of death. We have used a system that is effective and trustworthy for the identification of COVID-19 and pneumonia in our work. The digitized chest X-rays of normal, COVID-19 positive individuals, as well as those with pneumonia were used by the DL model to distinguish between them. The radiologists and other medical professionals can greatly benefit from this model’s decision-making capabilities and ability to improvise computerized diagnoses.
The major goal of this research was to develop a COVID-19 and pneumonia disease detection algorithm employing real-time data input DL. The DL approaches such as IV3 and VGG-16 models were used in this work. The work also delivered a web application for uploading chest X-ray images and predicting whether an individual is normal or infected with COVID/pneumonia.
1.3 Research Objectives
The major goal was to use DL models like IV3 and VGG16 to predict COVID-19 and pneumonia illness. The following actions have been taken in order to achieve this goal.
-
Pre-processing and information characterization: In this step, chest X-ray imagery from individuals was gathered from [5]. It included pre-processing the data by preparing the images for data augmentation and balancing. These distinct X-ray images were separated into train, test, and validation datasets during data pre-processing. They were enhanced and subject to normalization.
-
Model building: In this stage, images were classified into three classes—normal, COVID-19, and pneumonia—using the IV3 and VGG-16 models. To retrieve different information from the images, CNN employed a variety of adaptive algorithms. Most of the chest X-ray images are in gray scale. The pre-trained IV3 and VGG-16 models were applied for this purpose.
-
Model evaluation and web application deployment: The trained IV3 and VGG-16 models were evaluated using accuracy and loss on the test images of chest X-rays. The python framework “Django” was used to publish the model as a web application.
1.4 Main Contributions
-
Developed a deep learning model for detecting pneumonia and COVID-19 on chest X-ray images obtained from the repository [5].
-
Comparison of deep learning models with respect to accuracy and loss on test dataset.
-
Multiclass classification of images into normal, COVID affected, and pneumonia affected is performed.
-
Developed a web application. The result of this processing helps the radiologists to derive insights and make decisions to determine the correct diagnosis of the patient.
-
This model helps in two ways. Firstly, to classify whether a chest X-ray shows any sort of variations with respect to COVID-19 and pneumonia or not. Secondly, to classify with the help of normal chest X-ray images.
2 Related Work
All are aware that RTPCR procedures are employed to identify COVID-19, however carrying out RTPCR to identify COVID is unsafe since swabs testing that enter the throat through the nostril produce choking that releases viruses into the air and endangers the lives of medical personnel. Consequently, studies suggest that CT scans are significantly safer than this kind of swab examination. In addition, it is advised that a person who tests positive for COVID undergo a CT scan testing after the RTPCR testing. To the present day, hiring a radiologist to write radiological assessments for all the important discoveries or characteristics in chest X-rays is the finest method institutions use to detect COVID-19 and pneumonia. The latest AI systems using DL models are capable of reporting key COVID and pneumonia findings which are used by chest experts [6].
As a DL approach utilized in a range of image evaluation and interpretation, CNNs are well suited for the interpretation of chest X-ray images. Utilizing the images of the chest X-rays, we employed the CNN models IV3 and VGG-16 models to determine whether a person has COVID-19 or pneumonia. Several researchers have investigated the problem of accurately classifying images using DL techniques. Numerous research publications have contributed to the model’s enhancement to address the issue of the identification of COVID-19 and pneumonia. This section discusses some of the relevant work carried out in the domain of COVID-19 and pneumonia detection using machine learning (ML) / deep learning (DL) techniques.
The research [7] developed an ML model with more than 80% accuracy that can aid clinicians in diagnosing COVID-19. A preexisting ML model was trained on the data to make predictions. Without changing or making the FN rate in the dataset lower, they employed machine learning techniques to get an AUC of 0.84. The authors did not consider chest X-ray images during their experimentation. Only textual data with limited size was considered. Pneumonia detection was also not performed in their work.
The “Corona Hack-Chest X-Ray” Dataset, which is a publicly accessible dataset, is the first one that the authors [8] investigated. The second set of data comes from a facility in British Columbia, Canada, called “Vancouver General Hospital”. To reach accuracy of about 88%, they utilized pre-trained ANNs such as DarkNet19, Alex Net, and ResNet50. Techniques that are quick, precise, and powerful are required to diagnose COVID-19 testing. Swab tests result in expensive, time-consuming, and wasteful conventional lab procedures that are now performed. However, the authors did not focus on distinguishing between pneumonia and COVID affected patients. COVID-19 and pneumonia were detected with more than 95% accuracy in our work using DL methods and on chest X-ray images. Regardless of the widespread accessibility of X-rays, numerous limited hospitals in remote regions lack the radiological expertise needed to review and understand these images.
In [9], chest X-ray scans were chosen depending on their age and then classified. By entering the chest X-ray into the system, the model also made a prediction regarding the person’s age. The researchers separated the pediatric data from the adult population scans into other classes, such as pneumonia vs. normal vs. COVID-19. This aids in the DL-based models’ biased development. They had encouraging outcomes. In this investigation, chest X-ray imageries from distinct classes representing wildly dissimilar or various age groups—namely, pediatric population chest X-ray images—have been used. For the creation of effective different classifiers, DL or intelligence training requires appropriate varieties of medical scans, such as chest X-ray scans from identical age categories. Timely identification of COVID-19 and other serious viral disorders can be facilitated by DL-based categorization, which is particularly beneficial for radiologists.
The pre-trained XceptionNet on the ImageNet database was employed in the research [10]. They integrated two collections that were used to categorize individuals into normal and ill groups. By employing huge filtering sizes, the scientists were able to achieve the same outcome as conventional convolution with less operations. They succeeded in achieving 90% accuracy, 80% recall, and an F1-score higher than 85%. Nevertheless, the networks produce base Maps around the major bronchial region because of inflammation, which is not always a COVID-19 exclusive symptom. Chest X-rays from the two distinct datasets have a range of sizes and scaling the images can result in deformation.
EfficientNet was employed by the researchers [11] as the DL approach for COVID-19 identification. They showed that Xception, AmoebaNet—C, and ResNet were inferior to EfficientNet. From EfficientNet-B0 to EfficientNet-B7, they illustrated how EfficientNet evolved. The effectiveness of the algorithm enhanced as the parameter improved. The calculation, nevertheless, proved costly. Due to the idea of compounded modeling scalability, the EfficientNet could perform more effectively.
In [12], the authors compared Inception V3, Xception, and ResNeXt models and examined their accuracy for COVID-19 detection using chest X-ray images. They observed that the Xception model performed with superior accuracy. Their work mainly concentrates on probable techniques of identifying COVID-19 infected people and doesn’t promise significant clinical validity.
In [13], a deep learning model, the COVID-19 detection neural network (COVNet), was built to retrieve imagery data from cubic lung CT scans for the diagnosis of COVID-19. Diagnostic tests of community-acquired pneumonia (CAP) as well as other quasi anomalies were added to assess the model’s robustness. Diagnostics performance was evaluated utilizing the area under the receiver operating characteristic curve, sensitivity, and specificity.
The research [14] intends to create and assess an Ai system for the identification of COVID-19 on chest CT using data from a diversified multinational, inter-organizational dataset. The authors demonstrated that the robust models could attain up to 90% accuracy in independent assessment groups, preserving good specificity in non-COVID-19 associated pneumonias, and displaying acceptable generalization to unknown clinical populaces.
After carefully reading and analyzing each of these earlier studies, several of their findings have certain limits. Diagnostic DL frameworks are utilized in certain research articles for determining if a person possesses COVID-19/pneumonia/healthy. However, most of the works have performed binary classification in which they have identified whether a person has COVID-19 or not/pneumonia or not. Most of the efforts have not distinguished between COVID-19 and pneumonia using chest X-ray images. Comparison of the DL models is also not performed in many works. In addition to being extremely useful in clinical settings, automatic illness identification from chest X-rays at the level of professional radiologists will also be essential in providing healthcare to people with insufficient access to experts. To address this, we devised a solution that involves comparing two algorithms and predicting using the best model.
3 Dataset
As stated in Table 1, we used the database from [5] for this research that contains scans of normal, COVID, and pneumonia persons. The total size of the dataset was 6432 images.
As shown in Table 1, the total number of images in the training data was 5144 and the test data had 1288 images. Figure 1 shows the distribution of records with respect to each class in the dataset. In both training and test data, the number of pneumonia images were more.
IV3 and VGG-16 models were trained using the training dataset and then evaluated on test dataset. Multiclass classification into normal, COVID, and pneumonia classes was performed. Figure 2 shows the sample of normal, pneumonia, and COVID-19 X-ray images present in the dataset.
4 Framework and System Design
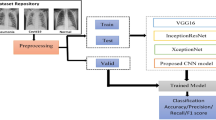
This section discusses the proposed framework for COVID-19 and pneumonia detection. Figure 3 shows the CNN architecture for the detection of pneumonia and COVID-19.
Here we are passing images from 3 classes that is COVID, pneumonia and normal chest X-ray and feed it to our pre-trained CNN model. In the CNN model pooling takes place which converts the images to 2D array. This is then passed to the dropout layer which helps reduce overfitting. After this it is fed to the fully connected layer and SoftMax layer for final prediction. Figure 4 depicts the class hierarchy of the CNN model used in this work.
The 3 categories of chest X-ray images are divided into 3 classes, i.e., 0 for normal, 1 for COVID, and 2 for pneumonia. These three classes are classified and predicted with the help of CNN algorithms (IV3 and VGG-16). The performance of the DL models was evaluated using performance metrics by comparing the predicted and actual classes.
Figure 5 shows the sequence model for COVID, and pneumonia detection used in this work.
The chest X-ray images are cropped first to the required size and it is preprocessed to the required form. In the pre-processing stage, “Contrast limited adaptive histogram equalization (CLAHE)” was applied to enhance the visibility intensity of the chest X-ray images. The resulting image was the Feature Set-1. The raw images were sharpened, and canny image was formed. This resulted in Feature Set-2. Both Feature Set-1 and Feature Set-2 were combined to form the updated dataset. Once the images were preprocessed, the classification phase was performed. In this phase, the updated dataset was split into training and test sets. The hyper-tuning of the IV3 and the VGG-16 models was performed on the training images to build the trained IV3/VGG-16 model. The trained IV3/VGG-16 model was evaluated on the test set. The results demonstrated that the VGG-16 model performed better than the IV3 model in classifying the chest X-ray images into normal, COVID-19, and pneumonia classes.
4.1 Inception V3 Model
Figure 6 depicts the Inception V3 model used in this work.
For image categorization, the Inception V3 model, a DL prototype based on CNN, is employed. It is an improved version of the Inception V1 that Google unveiled in 2014. These simulations, known as GoogleNet, were created by Google. Convolutional and pooling layers make up its construction. Using a kernel over every pixel while altering its nearby neighbors throughout the image sequence is the technique of convolution. The characteristic map’s size is reduced by the usage of pooling. The maximum and average pooling strategies are the most popular.
In Fig. 6, the dataset consisting of normal, COVID-19, and pneumonia images are loaded into the IV3 model. Inception V3 is a CNN model comprising of 48 layers for processing information. An Inception V3 network’s architecture is developed gradually and methodically, as follows:
-
With factorized convolutions, the number of networking variables can be decreased, which improves computing performance. It monitors the effectiveness of the system as well.
-
Training times can be reduced by switching to shorter convolutions instead of larger ones. A 5 × 5 filter, for example, contains 25 variables, while the same combination performed using twin 3 × 3 filters only have 18 (3 * 3 + 3 * 3).
-
A 3 × 3 convolution can be changed to a 1 × 3 convolution succeeded by a 3 × 1 convolution thanks to asymmetrical obfuscations. For example, the suggested asymmetrical combination uses less variables than might be needed if a 3 × 3 convolution were substituted by a 2 × 2 convolution.
-
An auxiliary encoder, which is a lightweight convolutional neural network (CNN) placed across levels in training and whose loss is contributed to the primary network’s loss. Auxiliary filters were employed to create a larger structure in GoogleNet, while in Inception V3 they serve as a regularization term.
-
Pooling activities are typically used to shrink the size of a grid.
The IV3 components in Fig. 6 summarize all the previously mentioned ideas. The completely linked layer receives the result of the IV3 component. Feed-forward neural nets sum up a fully connected layer. The final few segments of a Net are tightly bound. The result of the last pooling or convolutional layer is smoothed and supplied into the fully linked layer as its input. The SoftMax layer receives the results from the fully linked level. The IV3 model suggests a multinomial probabilistic model using the SoftMax activation function as the activation function in the output nodes. In this work, SoftMax was used as the last layer since multiclass classification (3 classes: normal, pneumonia, and COVID-19) had to be performed. In other words, when determining class membership on more than 2-class labels, SoftMax is employed as the activation function. The benefits of Inception V3 are that it is quite effective; it has greater depth of network than Inception V1 and V2 and 42 layers make up the structure.
4.2 VGG-16 Model
Figure 7 shows the VGG-16 model used for COVID and pneumonia detection.
A CNN model with 16 layers is called VGG16. It can classify images because it is a pre-trained algorithm. The input supplied to the model has predefined RGB visual size of 224 × 224. Firstly, the image is sent through the stacking of fully connected layers in which it records the lowest idea of the chosen fields. Following this, 5 maximum pooling layers complete the pooling process. Following this, 3 fully associated levels are utilized, each of which has 1000 channels and the final two 4096 channels each. As the last layer, SoftMax provides probability for every class label. The RELU is present in every hidden unit, assisting in preventing the exponential increase needed to run the NN. To determine whether algorithm is more effective in classifying the COVID-19 and pneumonia diseases, we compared Inception V3 and VGG16 using accuracy and validation loss.
5 Results
In this section we demonstrate the working of the system. It includes a comprehensible summary of the results of all critical tests that were carried out. The Inception V3 and VGG which are DL models based on CNN, for image classification and can be used for prediction purposes as well. We have compared the performance of both the algorithms. The editor used to code the model is Jupyter Notebook in the beginning and shifted to Google Collab. Django is an API that is used to connect the front-end UI with the back-end CNN model. Django is the open-source Python web platform that is used for developing our user interface, i.e., the front-end. We have used Django as it is more user-friendly, efficient and makes the website look much better.
Figure 8 gives the comparison of the DL models with respect to accuracy and loss for COVID and pneumonia detection.
As can be observed, the VGG-16 model performed with an accuracy improvement of 5.88%, sensitivity improvement of 3.88%, specificity improvement of 4.92%, precision improvement of 9%, and recall improvement of 7.41% over the Inception V3 model. Similarly, the VGG-16 model achieved lesser loss of 0.7046 over the Inception V3 model on the test dataset.
Figure 9 shows the accuracy and loss plots of the VGG-16 model on training and validation datasets at different epochs.
It can be observed that the highest validation accuracy and lowest model validation loss was obtained at epochs 16 and 25. Figure 10 shows the confusion matrix of the best performing VGG-16 model.
In Fig. 10, 0 stands for normal, 1 for COVID, and 2 for pneumonia classes. The VGG16 model was able to classify 17 records, 15 records, and 15 records accurately as belonging to normal, COVID, and pneumonia classes. However, it misclassified 2 images belonging to COVID class as pneumonia and 2 images belonging to pneumonia as COVID class.
The better performance of the VGG-16 model can be attributed to the fact that the VGG-16 model has more generalization power and less overfitting in comparison to the Inception V3 model. Furthermore, VGG-16 model has multiple stacked smaller size kernel which is better than the one with a larger size kernel because multiple non-linear layers increase the depth of the network enabling it to learn more complex features, and that too at a lower cost. 3 × 3 kernels help in retaining finer level properties of the image.
Figures 11, 12, 13, and 14 depict the snapshots of the web application developed in this work for COVID-19 and pneumonia detection.
Figure 11 shows the user interface describing the details of the COVID-19 and pneumonia illnesses.
Figure 12 depicts the user interface showing the chest X-rays of a normal person along with individuals affected with COVID-19 and pneumonia diseases.
Figure 13 shows the user interface describing the tips to be followed for preventing COVID-19 and pneumonia.
In Fig. 14, the end user such as a radiologist will upload the chest X-ray image. Our deep learning algorithm trained on chest X-rays will treat this uploaded chest X-ray image as the test image. It will predict whether the person is normal or is suffering from COVID-19/pneumonia. The best performing VGG-16 model was used as the back-end DL model for prediction.
6 Conclusion and Future Scope
The study uses CNN architectures and DL techniques to use computer vision to identify COVID and pneumonia illness. The architecture for fine-tuning and extraction of features is often used as each discovered convolutional neural network model is evaluated experimentally. From the repository, we have obtained a dataset containing imagery of persons with COVID and pneumonia as well as images of regular chest X-rays. Our analysis enabled us to identify the most potent algorithm for both COVID and pneumonia detection among the two. We found that the VGG16 model outperformed the IV3 models that corroborate the authors’ findings, with an accuracy improvement of 5%. As we can see, most algorithms did a good job of identifying normal, COVID, and pneumonia chest X-rays. A novel method of identifying and diagnosing COVID and pneumonia that could be useful in giving hospital services is suggested as a result of this work. The adjustment of the hyper-parameters should be one of the factors considered to increase the accuracy of the model in later studies where the adaption of alternative CNN architectures, such as shuffling Net and MobileNet designs, for the detection might be adopted. This research may also assist healthcare practitioners, such as doctors and other healthcare professionals, in making decisions about the use of more sophisticated prediction in genuine pneumonia detection as well as the possibility for detecting COVID and pneumonia using deep learning methods.
References
Hu B, Guo H, Zhou P et al (2021) Characteristics of SARS-CoV-2 and COVID-19. Nat Rev Microbiol 19:141–154. https://doi.org/10.1038/s41579-020-00459-7
Manoj Kumar MV, Atalla S, Almuraqab N, Moonesar IA (2022) Detection of COVID-19 using deep learning techniques and cost effectiveness evaluation: a survey. Front Artif Intell 5:912022. https://doi.org/10.3389/frai.2022.912022
Nguyen D, Kay F, Tan J, Yan Y, Ng YS, Iyengar P, Peshock R, Jiang S (2021) Deep learning-based COVID-19 pneumonia classification using chest CT images: model generalizability. Front Artif Intell 29(4):694875. https://doi.org/10.3389/frai.2021.694875
Sindhu VS, Dixit R, Sreedevi I (2022) Pneumonia detection using improved convolutional neural network. In: 2022 8th International conference on advanced computing and communication systems (ICACCS), pp 472–475. https://doi.org/10.1109/ICACCS54159.2022.9785127
Kermany DS, Goldbaum M, Cai W, Valentim CCS, Liang H, Baxter SL, McKeown A, Yang G, Wu X, Yan F, Dong J, Prasadha MK, Pei J, Ting MYL, Zhu J, Li C, Hewett S, Dong J, Ziyar I, Shi A, Zhang R, Zheng L, Hou R, Shi W, Fu X, Duan Y, Huu VAN, Wen C, Zhang ED, Zhang CL, Li O, Wang X, Singer MA, Sun X, Xu J, Tafreshi A, Anthony Lewis M, Xia H, Zhang K (2018) Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell 172(5):1122–1131. https://doi.org/10.1016/j.cell.2018.02.010
Nishio M, Kobayashi D, Nishioka E et al (2022) Deep learning model for the automatic classification of COVID-19 pneumonia, non-COVID-19 pneumonia, and the healthy: a multi-centre retrospective study. Sci Rep 12:8214. https://doi.org/10.1038/s41598-022-11990-3
Ferrari D, Milic J, Tonelli R, Ghinelli F, Meschiari M, Volpi S, Faltoni M, Franceschi G, Iadisernia V, Yaacoub D, Ciusa G, Bacca E, Rogati C, Tutone M, Burastero G, Raimondi A, Menozzi M, Franceschini E, Cuomo G, Corradi L, Orlando G, Santoro A, Digaetano M, Puzzolante C, Carli F, Borghi V, Bedini A, Fantini R, Tabbì L, Castaniere I, Busani S, Clini E, Girardis M, Sarti M, Cossarizza A, Mussini C, Mandreoli F, Missier P, Guaraldi G (2020) Machine learning in predicting respiratory failure in patients with COVID-19 pneumonia—challenges, strengths, and opportunities in a global health emergency. PLOS ONE 15:e0239172
Elgendi M, Nasir MU, Tang Q, Fletcher RR, Howard N, Menon C, Ward R, Parker W, Nicolaou S (2020) The performance of deep neural networks in differentiating chest X-rays of COVID-19 patients from other bacterial and viral pneumonias. Front Med 7
Sharma A, Rani S, Gupta D (2020) Artificial intelligence-based classification of chest X-ray images into COVID-19 and other infectious diseases. Int J Biomed Imaging 2020:1–10
Luján-García JE, Moreno-Ibarra MA, Villuendas-Rey Y, Yáñez-Márquez C (2020) Fast COVID-19 and pneumonia classification using chest X-ray images. Mathematics 8:1423
Liu C, Wang X, Liu C, Sun Q, Peng W (2020) Differentiating novel coronavirus pneumonia from general pneumonia based on machine learning. BioMedical Eng OnLine, 19
Jain R, Gupta M, Taneja S et al (2021) Deep learning-based detection and analysis of COVID-19 on chest X-ray images. Appl Intell 51:1690–1700. https://doi.org/10.1007/s10489-020-01902-1
Li L, Qin L, Xu Z, Yin Y, Wang X, Kong B, Bai J, Lu Y, Fang Z, Song Q, Cao K, Liu D, Wang G, Xu Q, Fang X, Zhang S, Xia J, Xia J (2020) Using artificial intelligence to detect COVID19 and community-acquired pneumonia based on pulmonary CT: evaluation of the diagnostic accuracy. Radiology 296:E65–E71
Harmon SA, Sanford TH, Xu S, Turkbey EB, Roth H, Xu Z, Yang D, Myronenko A, Anderson V, Amalou A, Blain M, Kassin M, Long D, Varble N, Walker SM, Bagci U, Ierardi AM, Stellato E, Plensich GG, Franceschelli G, Girlando C, Irmici G, Labella D, Hammoud D, Malayeri A, Jones E, Summers RM, Choyke PL, Xu D, Flores M, Tamura K, Obinata H, Mori H, Patella F, Cariati M, Carrafiello G, An P, Wood BJ, Turkbey B (2020) Artificial intelligence for the detection of COVID-19 pneumonia on chest CT using multinational datasets. Nat Commun 11
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Aditya Shastry, K., Manjunatha, B.A., Mohan, M., Kiran, N. (2024). Deep Learning Models for COVID-19 and Pneumonia Detection. In: Shetty, N.R., Prasad, N.H., Nalini, N. (eds) Advances in Computing and Information. ERCICA 2023. Lecture Notes in Electrical Engineering, vol 1104. Springer, Singapore. https://doi.org/10.1007/978-981-99-7622-5_7
Download citation
DOI: https://doi.org/10.1007/978-981-99-7622-5_7
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-7621-8
Online ISBN: 978-981-99-7622-5
eBook Packages: EngineeringEngineering (R0)