Abstract
Classical studies using several insect species have demonstrated that the principal circadian clock cells that generate circadian oscillations and control behavioral rhythms are located in a specific brain region. The discovery of a clock gene, period (per), in Drosophila melanogaster further facilitated the identification of specific cells by labeling gene expression. Since most of the per-expressing brain neurons display circadian molecular oscillations in the levels of per mRNA and its protein expression, they have conventionally been defined as “circadian clock neurons.” In Drosophila, approximately 150 neurons (out of 200,000 brain neurons) have been identified as clock neurons. However, elucidating the role of clock neurons, even with the Drosophila model, has been a major challenge. In 1995, it was discovered that 16 clock neurons expressed a neuropeptide, pigment-dispersing factor (PDF), the most important neurotransmitter for the insect circadian clock. This was where Drosophila genetics and neuroscience met in chronobiology, leading to a significant development in the functional analysis of clock neurons in Drosophila and the identification of clock neurons in nonmodel insect species. This chapter will summarize the latest findings of the clock neuron network in Drosophila and other insect species.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
The most commonly observed circadian rhythms are shown in behavior, e.g., sleep-wake cycles or nocturnal/diurnal activity. Today, we know that cells containing circadian clocks are widespread throughout the body (Ito and Tomioka 2016; Chap. 6). Needless to say, however, the biggest question in the past was where in the animal’s body the clock was located. During the late 1960s and 1970s, several pioneering studies were conducted on insects to answer this question. The first study was performed by Nishiitsutuji-Uwo and Pittendrigh (1968), who surgically lesioned parts of the brain and measured locomotor activity rhythms in the cockroach Rhyparobia (Leucophaea maderae). The cockroaches in which a brain region called the optic lobe was bilaterally lesioned displayed arrhythmic locomotor activity. Later, the same conclusion was drawn from studies of other insects, such as beetles and crickets (Loher 1972; Fleissner 1982; Tomioka and Chiba 1984). Two significant findings have been made to reinforce the hypothesis that the optic lobe is the locus of the principal circadian pacemaker: (1) Transplantation of the optic lobes into the optic lobeless brain restored the activity rhythm of the donor cockroach in R. maderae (Page 1982). 2) The isolated optic lobes displayed circadian rhythms in neural activity in a self-sustained manner in the cricket Gryllus bimaculatus and R. maderae (Tomioka and Chiba 1986; Colwell and Page 1990; Tomioka and Chiba 1992). Thus, the optic lobes of these insects contain pacemakers that control the activity rhythm.
The optic lobes mainly consist of three neuropils, namely, the lamina, medulla, and lobula, which process visual information from the compound eyes and send the processed information to the midbrain. The anatomical relationship between the light input pathway and the circadian pacemaker is plausible, given the importance of light entrainment. In mammals, light information is conveyed directly from the eye via the retinohypothalamic tract to entrain the mammalian pacemaker located in the suprachiasmatic nucleus (SCN) (Panda et al. 2002; Ruby et al. 2002). This analogy between insects and mammals suggests that the origin of the pacemaker center should be tightly linked to photoreception.
2 Small Ventral Lateral Neurons in Drosophila
Classical lesion experiments have beautifully revealed the brain region important for rhythm generation. However, this method is not suitable for identifying the locus of the pacemaker at the cellular level. The discovery of the period (per) gene in the fruit fly Drosophila melanogaster, a model organism in genetics, overcame this difficulty by using modern cell labeling techniques, which enabled the identification of the cells expressing the per gene in the brain. In situ hybridization and immunohistochemistry, which label mRNA and protein expression, respectively, have revealed that per is expressed in cells distributed in a wide range of brain regions (Liu et al. 1988; Siwicki et al. 1988). Therefore, the difficulty in identifying pacemaker cells persisted.
Since then, developments in fluorescent immunohistochemistry, confocal laser microscopy, and transgenic fly lines, such as the GAL4-UAS system, have contributed significantly to determining the precise anatomical location of per-expressing brain neurons (Kaneko and Hall 2000; Helfrich-Förster 2003). Today, we know that per is expressed in approximately 150 brain neurons classified into nine groups (Fig. 5.1). First, per-expressing neurons are divided mainly into the lateral and dorsal neuron groups. Lateral neurons are located between the optic lobe and midbrain and are further subdivided into small ventral lateral neuron (s-LNv), large ventral lateral neuron (l-LNv), fifth lateral neuron (fifth LN, also known as fifth s-LNv), dorsal lateral neuron (LNd), and lateral posterior neuron (LPN) groups. Dorsal neurons are located in the rim of dorsal brain regions and are further subdivided into anterior dorsal neuron 1 (DN1a), posterior dorsal neuron 1 (DN1p), dorsal neuron 2 (DN2), and dorsal neuron 3 (DN3) groups.
The brain of Drosophila melanogaster and the distribution of clock neurons. The Drosophila central clock consists of approximately 150 neurons in the brain. The clock neurons are divided mainly into nine groups based on their localization and size of cell bodies: small ventral lateral neuron (s-LNv), large ventral lateral neuron (l-LNv), fifth lateral neuron (fifth LN), dorsal lateral neuron (LNd), lateral posterior neuron (LPN), anterior dorsal neuron 1 (DN1a), posterior dorsal neuron 1 (DN1p), dorsal neuron 2 (DN2), and dorsal neuron 3 (DN3)
The disconnected (disco) gene, which encodes a C2H2-type zinc-finger transcription factor, plays a role in nervous system development. disco mutants lack many optic lobe neurons, including per-expressing lateral neurons, and are behaviorally arrhythmic (Dushay et al. 1989; Zerr et al. 1990; Hardin et al. 1992). Since the DN groups are intact in disco mutants, researchers assume that lateral neurons are the central pacemaker neurons that control activity rhythms such as locomotion and eclosion. Within the lateral neuron groups, s-LNv and l-LNv neurons express a neuropeptide, pigment-dispersing factor (PDF) (Helfrich-Förster 1995). Only a few disco mutants retain some of the s-LNv neurons, and they are behaviorally rhythmic (Helfrich-Förster 1998), which further suggests that s-LNv neurons are the essential pacemakers.
The discovery of the Pdf mutant has also significantly advanced the functional analysis of s-LNv neurons. Wild-type Drosophila shows bimodal locomotor activity rhythms with two peaks in the morning and evening in light-dark cycles (LD) and the rhythms free run with a period of approximately 24 h in constant darkness (DD) (Konopka and Benzer 1971; Hamblen-Coyle et al. 1992). Pdf mutants display weak free-running activity rhythms with a period of approximately 22 h in DD for the first few days, and then the rhythms damp out (Renn et al. 1999). These phenotypes are attributed to the loss of PDF in s-LNv neurons, since Pdf knockdown only in s-LNv neurons (but not in l-LNv neurons) reproduces the weak activity rhythm of Pdf mutants (Shafer and Taghert 2009). This s-LNv master pacemaker hypothesis is further supported by the fact that per expression only in s-LNv neurons is sufficient for generating free-running activity rhythms in DD (Grima et al. 2004). Taken together, s-LNv neurons are the most influential clock neurons, and the PDF signaling output from s-LNv neurons conveys important circadian timing information to downstream neurons.
3 Outputs from the Drosophila Clock Neurons
Fourteen neurotransmitters, including PDF, have been identified in Drosophila cerebral clock neurons. Although studies on their functional roles are still in progress, those reported are listed in Table 5.1. Among them, nine neurotransmitters have been reported to be involved in intercellular communication between clock neurons (Fig. 5.2). Here, we summarize what we know about circadian outputs from clock neurons.
Intercellular communication pathways of the Drosophila circadian clock network. The colored clock neurons contain neurotransmitters indicated in each panel and transmit signals to other (or own) clock neurons (arrows). The colors correspond to those used in Fig. 5.1. Since the distributions of the receptors for some ligands are not identified, some correspondences between output and input neurons are uncertain
3.1 Pigment-Dispersing Factor
PDF was found to be the first circadian neurotransmitter (Helfrich-Förster 1995). In 2005, three independent groups identified the PDF receptor (PDFR) gene (Hyun et al. 2005; Mertens et al. 2005; Lear et al. 2005). Interestingly, PDFR is expressed in many clock neurons, including PDF-positive s-LNv neurons, PDF-negative LNd, and other DN groups (Im and Taghert 2010). The function of PDF/PDFR signaling is to synchronize PDF-positive and PDFR-positive clock neurons to adjust the phase of molecular oscillations and Ca2+ rhythms in the clocks (Peng et al. 2003; Lin et al. 2004; Shafer et al. 2008; Yoshii et al. 2009; Liang et al. 2016, 2017; Fig. 5.2a). The mammalian counterpart for PDF/PDFR signaling is vasoactive intestinal polypeptide (VIP)/VIP receptor signaling, which also functions to couple clock neurons in the SCN (Mieda 2020; Ono et al. 2021). Both PDFR and VIP receptors belong to a class II G-protein-coupled receptor family (Mertens et al. 2005), implying an evolutionarily conserved neural mechanism in a wide range of animal species.
PDF is expressed only in s-LNv and l-LNv neurons (Helfrich-Förster 1995). s-LNv and l-LNv neurons are morphologically different, although both groups are located very close to each other in the lateral brain (Helfrich-Förster 1997). s-LNv neurons have smaller cell bodies and send projections into the dorsal brain, where fifth LN, LNd, and DN1 neurons also send their projections, presumably to communicate with each other. l-LNv neurons have larger cell bodies and send the projections in two directions: one goes to the optic lobe with complex arborizations and the other to the contralateral hemisphere. Both s-LNv and l-LNv neurons also send fibers into the accessory medulla (Helfrich-Förster et al. 2007). The role of PDF signaling from s-LNv neurons is to synchronize s-LNv neurons and other PDF receptor-positive clock neurons, such as fifth LN, LNd, and DN1 neurons, which is supposed to be important for the maintenance of free-running rhythms in DD (Peng et al. 2003; Shafer et al. 2008; Yoshii et al. 2009). The projection pattern of l-LNv neurons implies that l-LNv neurons send a signal into the visual processing neurons in the optic lobe and into the contralateral brain for bilateral clock synchronization (Helfrich-Förster et al. 2007). The l-LNv neurons indeed receive a signal from contralateral l-LNv neurons to change membrane potential but via gap junctions, not via PDF (Cao and Nitabach 2008).
It has been inferred that l-LNv neurons are less important in free-running conditions. This is because clock protein oscillations are dampened in l-LNv neurons under DD (Yang and Sehgal 2001) and Pdf knockdown in l-LNv neurons does not affect free-running rhythms in DD (Shafer and Taghert 2009). Pdf01 mutants display diminished morning activity and phase-advanced evening activity in LD. The rescue of PDF expression in l-LNv neurons restores the wild-type evening activity (Cusumano et al. 2009; Schlichting et al. 2016; Menegazzi et al. 2017; Schlichting et al. 2019b), suggesting that the role of PDF in l-LNv neurons is to set the phase of evening activity in LD. In contrast, PDF signaling from s-LNv neurons is essential for morning activity (Shafer and Taghert 2009). pdfr mutants also display phase-advanced evening activity (Hyun et al. 2005). The rescue of pdfr expression in DN1p or LNd and fifth LN neurons under the pdfr mutant background is sufficient for the wild-type morning and evening activity (Lear et al. 2009; Zhang et al. 2010a; Schlichting et al. 2016). According to the latest study by Schlichting et al. (2016), l-LNv neurons send PDF signaling to s-LNv neurons (PDF receptor-positive), and then s-LNv neurons pass it on to DN1p, LNd, and fifth LN neurons, which in turn generate wild-type evening activity. Since PDF is the first neuropeptide for the Drosophila circadian clock, it has been extensively studied. The functional analysis of PDF led directly to the analysis of s-LNv and l-LNv neurons, revealing the complexity of the circadian neural network.
3.2 Neuropeptides
To date, 11 neuropeptides expressed in clock neurons have been identified in Drosophila. Johard et al. (2009) found that three neuropeptides, neuropeptide F (NPF), short-neuropeptide F (sNPF), and ion transport peptide (ITP), were expressed in subsets of clock neurons. Drosophila shows sleeplike behavior that is typically characterized as periods of quiescence lasting longer than 5 min (Hendricks et al. 2000; Shaw et al. 2000). NPF-positive clock neurons modulate evening activity and sleep behavior (Hermann et al. 2012; Chung et al. 2017). In addition, NPF signaling from LNd neurons entrains rhythmic gene expression in fat bodies, which are comparable organs to the mammalian liver (Chung et al. 2017).
sNPF and NPF are similar in name, but they are encoded by different genes. Immunohistochemistry using antibodies against the sNPF precursor revealed that it is expressed in s-LNv neurons and two of six LNd neurons (Johard et al. 2009). sNPF signaling from s-LNv neurons negatively affects the Ca2+ level in DN1 neurons, which is correlated with the morning activity peak (Liang et al. 2017; Fig. 5.2b). sNPF also mediates circadian signaling from s-LNv neurons to prothoracicotropic hormone-expressing neurons, which control circadian emergence rhythms (Selcho et al. 2017).
One LNd and fifth LN neurons express the ITP neuropeptide (Johard et al. 2009). The ITP-positive LNd neuron is one of the three NPF-positive LNd neurons. Knockdown of itp expression in clock neurons reduces the evening activity peak in LD and lengthens the free-running period in DD (Hermann-Luibl et al. 2014). Furthermore, simultaneous knockdown of itp and Pdf increases the level of night activity in LD and makes arrhythmic. Taken together, ITP plays a role in the output of clock neurons, especially in relation to PDF signaling.
The neuropeptide diuretic hormone 31 (DH31) is expressed in DN1p and LPN neurons (Kunst et al. 2014; Reinhard et al. 2022). A loss-of-function allele of the DH31 gene increases sleep late at night but shows normal circadian rhythms. However, DH31 does contribute to circadian activity rhythms. The double mutants of Pdf and DH31 are nearly arrhythmic in DD (Goda et al. 2019), suggesting that DH31 plays a role in maintaining rhythms in the absence of PDF. DH31 receptor (DH31R) is expressed in DN1p neurons, and thus DH31 signaling from DN1p neurons may feedback on themselves (Goda et al. 2018; Fig. 5.2c). Flies change their preferred temperature over the course of a day, showing a so-called temperature preference rhythm with a peak in the evening (Kaneko et al. 2012). The DH31-PDFR signaling pathway in DN2 neurons plays a role in the temperature preference rhythm (Goda et al. 2016).
The neuropeptide CCHamide1 (CCHa1) is expressed in DN1a neurons (Fujiwara et al. 2018). The CCHa1 receptor (CCHa1R) is expressed in LNv neurons (Abruzzi et al. 2017; Fujiwara et al. 2018). Since PDFR is expressed in DN1a neurons and s-LNv neurons send projections to the dorsal brain in close proximity to DN1a neurons, DN1a and s-LNv neurons are reciprocally coupled via CCHa1 and PDF signaling (Fig. 5.2a, d). Mutant flies of CCHa1 display diminished morning activity, reduced total activity, and enhanced sleep amount (Fujiwara et al. 2018). The mammalian homolog of CCHa1R is the receptor of gastrin-releasing peptide, which plays a role in the clock neuron network (Mieda 2020; Ono et al. 2021). Thus, similar to PDFR, the receptor (but not the ligand) is well conserved across animal species.
Allatostatin A (AstA) is a neuropeptide expressed in LPN neurons (Ni et al. 2019). Activation of LPN neurons promotes sleep and reduces feeding, and at least the sleep phenotype is partly mediated by AstA signaling (Chen et al. 2016; Ni et al. 2019; Reinhard et al. 2022). Since LPN neurons receive PDF signaling, LPN neurons are downstream of LNv neurons, and AstA signaling from LPN neurons may mediate coupling between LNv clock neurons and sleep-promoting neurons.
Allatostatin C (AstC) neuropeptide expression in clock neurons was first discovered by RNA-sequencing analysis (Abruzzi et al. 2017). Immunohistochemistry using an anti-AstC antibody further revealed that AstC is expressed in four to six DN1p, a subset of LNd, a subset of DN3, and LPN neurons (Díaz et al. 2019; Zhang et al. 2021a; Reinhard et al. 2022; Meiselman et al. 2022). Knockdown of AstC mRNA in clock neurons results in a phase-delayed evening activity, which is mediated by AstC-R2 (one of two AstC receptors) expressed in LNd neurons (Díaz et al. 2019). AstC is also involved in circadian rhythms in the progression of oogenesis in mated females (Allemand 1976). AstC signaling from DN1p neurons outputs to the pars intercerebralis (PI) region, through which circadian oogenesis rhythms are generated (Zhang et al. 2021a, 2021b). Thus, AstC signaling is used in two directions. One is a signal to LNd neurons to communicate between clock neurons (Fig. 5.2e), and the other is an output of temporal information to downstream cells.
Transcriptome analyses in clock neurons reveal that two novel neuropeptides, CNMamide (CNMa) and Trissin, are expressed in DN1p and LNd neurons, respectively (Abruzzi et al. 2017; Ma et al. 2021). DN1p neurons input temperature information and modulate sleep in a temperature-dependent manner (Yadlapalli et al. 2018). CNMa signaling mediates the interaction between DN1p and PI neurons to control temperature-dependent sleep (Jin et al. 2021). In contrast, the function of Trissin has not yet been reported. Transcriptome analysis by Abruzzi et al. (2017) revealed that the receptor of Trissin is expressed in LNd and DN1 neurons, suggesting that Trissin mediates LNd-LNd and LNd-DN1 couplings (Fig. 5.2f).
IPNamide was discovered as the second circadian neuropeptide after PDF (Shafer et al. 2006). IPNamide is expressed in DN1a neurons, but its function has not been reported. IPNamide is encoded by the neuropeptide-like precursor 1 gene (Nplp1), which encodes three other peptides, MTYamide, APK, and VQQ (Baggerman et al. 2002). Thus, it is difficult to analyze the function of IPNamide alone.
3.3 Glutamate, Acetylcholine, and Glycine
Vesicular glutamate transporter (VGlut) is expressed in DN1a, some DN1p, and DN3 neurons, suggesting that these clock neurons use glutamate as a neurotransmitter (Hamasaka et al. 2007). Glutamate signaling is mediated by two main receptors, a glutamate-gated chloride channel, GluCl, and a metabotropic G-protein-coupled receptor, mGluRA. The receptors are expressed in s-LNv, l-LNv, and LNd neurons (Hamasaka et al. 2007; Collins et al. 2012; Guo et al. 2016). Glutamate signaling from DN1p neurons synchronizes DN1p and LNv neurons (Collins et al. 2012; Guo et al. 2016; Fig. 5.2g). Since s-LNv neurons send PDF signaling to DN1p neurons (Shafer et al. 2008), there is reciprocal coupling between s-LNv and DN1p neurons. This coupling is essential for the robustness of molecular oscillations and normal activity rhythms (Hamasaka et al. 2007; Collins et al. 2014). VGlut may also be expressed in LNd and/or fifth LN neurons, as RNA interference for VGlut in LNd and fifth LN neurons influences activity rhythms (Duhart et al. 2020; Fig. 5.2g).
Johard et al. (2009) also reported that acetylcholine (Ach) is used as the circadian neurotransmitter, as choline acetyltransferase (ChAT) is expressed in two of six LNd and fifth LN neurons. ChAT is coexpressed with sNPF in the two LNd neurons, which is different from the three NPF-positive LNd neurons. Thus, six LNd neurons are divided into two sNPF- and ChAT-coexpressing neurons, two NPF-positive neurons, one ITP- and NPF-coexpressing neuron, and one with an unknown neurotransmitter. Cholinergic signaling, which is an excitatory input, from LNd neurons targets s-LNv and l-LNv neurons because its receptor is expressed (McCarthy et al. 2011; Lelito and Shafer 2012; Fig. 5.2h). Knockdowns of ChAT or vesicular acetylcholine transporter (vAchT) in LNd neurons do not change the speed of free-running activity rhythms in DD but reduce the robustness of the rhythms (Duhart et al. 2020).
The other fast neurotransmitter used in the Drosophila clock is glycine (Frenkel et al. 2017). Knockdown of the glycine transporter dGlyT in LNv (s-LNv and l-LNv) neurons results in lengthening of the free-running period in DD. Glycine applications on the cultured brain inhibit the neural activity of DN1p neurons. These results suggest that glycine is used in LNv neurons as a neurotransmitter and that its receptor is expressed in DN1p neurons. Since s-LNv neurons are responsible for the speed of the free-running period, one can assume that glycine mediates the neurotransmission from s-LNv neurons to DN1p neurons (Fig. 5.2i).
3.4 Gap Junctions
Most studies of the neural network of the Drosophila circadian clock are concerned with chemical synapses. Perhaps this is because genetic screening targeting ligands or their receptors is an advantage of Drosophila research. However, insect clock neurons are known to couple at electrical synapses as well (Schneider and Stengl 2006; Li et al. 2018). In the case of Drosophila, the electrical synapse is composed of gap junctions by eight innexin proteins (innexins 1–8). Knockdown of Innexin1 and Innexin2 expression in clock neurons results in longer free-running periods than control strains in DD (Ramakrishnan and Sheeba 2021). Additionally, knockdown of Innexin2 expression leads to a phase shift of PER oscillation and reduces PDF expression in the morning. These results suggest that gap junction-mediated signaling between clock neurons is important for maintaining circadian molecular oscillations.
3.5 Output Modes
Clarifying when and how circadian neurotransmitters are released is still challenging, although some recent advances may have developed experimental methods to approach this long-lasting problem (Leopold et al. 2019; Ding et al. 2019). PDF immunostaining reveals that the PDF level in the terminals of the s-LNv projections cycles with a peak in the morning (Park et al. 2000). Similar observations have been made on ITP, CCHa1, DH31, AstA, and AstC neuropeptides, but the phases of their rhythms are different (Hermann-Luibl et al. 2014; Fujiwara et al. 2018; Díaz et al. 2019; Reinhard et al. 2022). The cycling of the neuropeptide contents may not reflect their synaptic release directly, but it implies that they are released in a circadian manner. A study using a fluorescent sensor for visualizing neuropeptide release has revealed that s-LNv neurons release neuropeptides in the morning with a slight delay from the peak of PDF level at their axonal terminals (Klose et al. 2021). Thus, it is very likely that other circadian neuropeptides are also rhythmically released, by which timing information is transmitted to postsynaptic downstream neurons. However, the importance of rhythmic PDF release to activity rhythms is still controversial (Kula et al. 2006; Prakash et al. 2017).
The cyclic chemical transmissions can be complexly organized by the circadian structure remodeling of clock neurons. Fernández et al. (2008) found that the axonal terminals of s-LNv neurons change morphology, higher complexity during the day and lower complexity during the night, and the daily morphological changes are regulated by the molecular clock. Through rhythmic structure remodeling, s-LNv neurons change synaptic partners throughout the day (Gorostiza et al. 2014). A recent study proposed that s-LNv circadian remodeling is important for integrating light and temperature inputs (Fernandez et al. 2020). Similar morphological changes have been reported in fifth LN, LNd, and DN1a neurons (Duhart et al. 2020; Song et al. 2021). Thus, the circadian remodeling of axonal terminals may be a general property of clock neurons.
4 Functional Differentiation of Individual Clock Neuron Groups in Drosophila
4.1 Morning and Evening Oscillators in the Drosophila Circadian Clock
Drosophila shows two distinct activity peaks in the morning and evening (Helfrich-Förster 2000), and the two activity peaks are controlled by two oscillators with different properties (Yoshii et al. 2012). Grima et al. (2004) and Stoleru et al. (2004) proposed that the two oscillators are separate groups of clock neurons: the morning oscillator (M oscillator) corresponds to s-LNv neurons and the evening oscillator (E oscillator) to LNd neurons. Later, the fifth LN neuron was identified (Rieger et al. 2006) and considered the evening oscillator. The M and E oscillators have different response modes to light. An exposure of dim light, which is equivalent to a light intensity of a quarter moon, in the night phase results in a phase advance of the M peak and a phase delay of the E peak (Bachleitner et al. 2007). This result fits well with the classical two-oscillator model proposed from studies performed in rodents (Pittendrigh and Daan 1976). In this model, the M oscillator accelerates, and the E oscillator decelerates the speed of oscillations upon light exposure. By changing the speed of oscillation depending on light exposure, both oscillators can flexibly adapt to different photoperiods, by which the circadian clock enables the measurement of day length to predict the coming season.
The Drosophila two-oscillator model has certainly inspired the functional analysis of clock neurons. Many studies support this model, but on the other hand, some studies note that it is oversimplified. For example, per0 mutants with a per rescue expression only in a subset of DN1p neurons display relatively normal morning and evening activity (Zhang et al. 2010b). Additionally, flies without s-LNv neurons still display a clear morning activity peak (Sheeba et al. 2010). per0 mutants with a per rescue only in M or E oscillator do not completely restore typical morning and evening activity under various photoperiods (Menegazzi et al. 2020). On the other hand, CRISPR-mediated per or tim gene knockout only in M oscillator causes loss of the morning activity peak (Delventhal et al. 2019). These seemingly contradictory results are because it does not take into account the complex neural network between clock neurons (Jaumouillé et al. 2021). Since LNv and DN1 neurons interact intricately with each other through multiple neurotransmitters, it may be challenging to analyze functions only by manipulating specific clock neuron groups (Yao and Shafer 2014; Yao et al. 2016). Figure 5.3 shows a current M-E two-oscillator model that highlights the importance of the DN1p group for generating both M and E peaks (Chatterjee et al. 2018). In this model, two types of DN1p neurons (M-DN1p and E-DN1p neurons) control the M and E peaks. M-DN1p neurons receive a signal from the M oscillator (s-LNv neurons) and control M peak. E-DN1p neurons and E oscillators (fifth LN and LNd neurons) control the E peak, but they are concurrently influenced by the M oscillator.
Current model for generating morning (M) and evening (E) activity peaks in Drosophila. Drosophila shows two activity peaks in the morning and evening under LD. In principle, s-LNv neurons and fifth LN and LNd neurons are designated as the M oscillator and E oscillator, respectively. DN1p neurons contain the M (M-DN1p) and E oscillators (E-DN1p) and mediate the output from s-LNv neurons
4.2 s-LNv Neurons as the Master Clock?
s-LNv neurons have been considered to be the master pacemaker clock. This hypothesis is based on the following facts: (1) the stability of the free-running rhythm in DD is significantly weakened by the loss of PDF or PDF-positive LNv neurons, and (2) the free-running rhythm is restored by per rescue only in s-LNv neurons (Helfrich-Förster 1998; Renn et al. 1999; Grima et al. 2004; Cusumano et al. 2009). However, several lines of evidence point to different ideas. For example, per0 mutant flies with a per rescue only in the E oscillator (fifth LN and LNd neurons) can display a free-running rhythm under constant dim light conditions (Rieger et al. 2009). Disruption of the molecular clock only in s-LNv neurons is insufficient to render flies arrhythmic in DD (Delventhal et al. 2019; Schlichting et al. 2019a, 2019b; Jaumouillé et al. 2021). Furthermore, silencing of neural activity in M or E oscillators, even with normal molecular oscillations, also renders fly arrhythmic (Bulthuis et al. 2019), which suggests that the disconnection of intercellular communication between clock neurons causes the loss of the free-running rhythm. Thus, s-LNv neurons remain essential for self-sustained DD rhythms, but the importance of other clock neurons has been overlooked.
4.3 DN1p Neurons as Circadian Output Centers
DN1p neurons are composed of heterogeneous neurons. Six of 15 DN1p neurons express CRY and PDFR (Yoshii et al. 2008; Im and Taghert 2010), and they use AstC, DH31, glutamate, and CNMa as neurotransmitters (Kunst et al. 2014; Chatterjee et al. 2018; Ma et al. 2021). CRY-positive DN1p neurons are involved in sexual interactions (Fujii and Amrein 2010; Hanafusa et al. 2013), feeding (Barber et al. 2016), sleep regulation (Guo et al. 2016, 2017; Lamaze et al. 2018), memory extinction (Zhang et al. 2021b), activity rhythms (Nettnin et al. 2021), and reproductive rhythms (Zhang et al. 2021a). These reports suggest that CRY-positive DN1p neurons may be the output center of the circadian clock, which transmits timing information to downstream neurons to generate various behavioral rhythms. This may be why DN1p neurons have many different neurotransmitters.
5 Downstream Neurons of the Drosophila Circadian Clock
PI neurons have been considered the circadian output region for many years (Nishiitsutsuji-Uwo et al. 1967; Cymborowski 1973; Sokolove and Loher 1975; Takekata et al. 2018). In Drosophila, there are three distinct populations of PI neurons that express three different peptides: diuretic hormone 44 (DH44), SIFamide (SIFa), and Drosophila insulin-like peptide (dilp2). DN1p and LNd neurons directly or indirectly contact PI neurons (Cavanaugh et al. 2014; Barber et al. 2016, 2021). DH44-positive and SIFa-positive PI neurons mediate the output pathways to control circadian activity rhythms, whereas SIFa-positive and dilp2-positive PI neurons control feeding rhythms and metabolism (Cavanaugh et al. 2014; Barber et al. 2016, 2021; Dreyer et al. 2019). Some DN1p neurons also contact tubercular-bulbar neurons that, in turn, connect ellipsoid body ring neurons (Guo et al. 2018; Lamaze et al. 2018), which include those that promote sleep and those involved in the output of activity rhythms (Liang et al. 2019).
DN1p neurons are not the only ones coupled to output pathways. As mentioned above, s-LNv neurons communicate with PTTH neurons via sNPF signaling (Selcho et al. 2017). PDF (and sNFP) signaling from s-LNv neurons plays roles in reproductive dormancy mediated by dilp2-positive PI neurons (Nagy et al. 2019) and memory mediated by the mushroom body (Flyer-Adams et al. 2020; Inami et al. 2021). The other output pathway from s-LNv neurons is the neuropeptide leucokinin (LK)-positive neurons (Cavey et al. 2016). Both Lk and Lk receptor mutants reduce the power of activity rhythms in DD. All the output pathways mentioned above have been analyzed morphologically, physiologically, and behaviorally. Recent electron microscopic data further revealed entire postsynaptic neurons of all clock neurons (Scheffer et al. 2020), showing that the output of the circadian clock spreads across a wide range of brain neurons.
6 Clock Neuron Networks in Other Insect Species
In insect species other than Drosophila melanogaster, immunostaining against PDF has provided the most reliable results for identifying putative clock neurons. This is because PDF antibodies are specific to many species due to the high conservation of PDF peptide sequences (Meelkop et al. 2011). Similar to all insect species studied, PDF cells reside in the lateral protocerebrum, and they extend neuronal processes toward the central brain and optic lobes (Helfrich-Förster 2005). In the cockroach R. maderae, ectopic transplantation of the accessory medulla, including PDF neurons, can restore activity rhythms in optic lobeless arrhythmic cockroaches, strongly suggesting the importance of PDF neurons (Reischig and Stengl 2003). Injections of synthetic PDF peptides into the brain phase shift activity rhythms in the cockroach (Petri and Stengl 1997) and cricket (Singaravel et al. 2003). The knockdown of Pdf mRNA expression by RNA interference or the knockout of the Pdf gene by CRISPR/Cas9 results in arrhythmicity or a short free-running period in the German cockroach (Lee et al. 2009), cricket (Hassaneen et al. 2011), and bug (Kotwica-Rolinska et al. 2022). Furthermore, circadian rhythms at the PDF level and the structural changes of the projections from PDF neurons have also been detected (Abdelsalam et al. 2008; Wei and Stengl 2011). Putting all these results together, it is likely that, as in Drosophila, PDF and PDF neurons are essential for circadian activity rhythms in insects. However, things are not so simple.
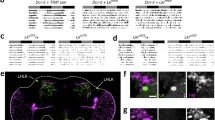
While the identification of PER-expressing cells in the brain has been attempted in several insect species, the locations of the PER-expressing cells are often different from those of Drosophila (Table 5.2; Helfrich-Förster 2005; Beer and Helfrich-Förster 2020). PER is not colocalized in PDF neurons in some insect species (Frisch et al. 1996; Sauman and Reppert 1996; Závodská et al. 2003b; Koide et al. 2021). Furthermore, in the cricket, the surgical lesion of the outer medulla and lamina neuropils results in arrhythmicity even though the accessory medulla, including PDF neurons, is still intact (Okamoto et al. 2001). Therefore, we have to be cautious in simply concluding PDF neurons as clock neurons. It is quite possible that there is a diversity of clock neuron networks, along with the diversity of insect species. Even within the genus Drosophila, there are some variations in PDF/clock protein expression patterns (Hermann et al. 2013; Menegazzi et al. 2017). In contrast, several studies have shown clock neuron networks similar to those of Drosophila in the blowfly Protophormia terraenovae (Shiga and Numata 2009), the honeybee Apis mellifera (Fuchikawa et al. 2017; Beer et al. 2018), the ant Camponotus floridanus (Kay et al. 2018; Fig. 5.4a), and the aphid Acyrthosiphon pisum (Barberà et al. 2017; Colizzi et al. 2021; Fig. 5.4b). In addition, subsets of PER-positive neurons in the blowfly, ant, and honeybee also exhibit PDF, although this is not true in the aphid because it seems that the Pdf gene is lost in the aphid genome (Huybrechts et al. 2010).
The brains of the ant Camponotus floridanus (a) and aphid Acyrthosiphon pisum (b) and their clock neurons. In both insects, clock neurons form clusters similar to Drosophila lateral and dorsal neurons (Fig. 5.1). The ant clock neurons consist of approximately 200 neurons, whereas the aphid clock neurons consist of approximately 40 neurons
7 Bilateral Coupling Between Two Optic Lobe Clocks
Drosophila is not always used as a model of insect chronobiology. The coupling between two clocks residing in the left and right brain is such a subject of study. Even in advanced Drosophila genetics, it is not possible to manipulate one side of the body asymmetrically. The tiny brain of Drosophila also makes it difficult to manipulate the brain surgically. In contrast, robust insect species with larger brains, such as crickets and cockroaches, are good models.
Page and his colleagues demonstrated that excision of one optic lobe (either right or left) in the cockroach R. maderae did not affect the ability to generate free-running activity rhythms, but their periods were longer than those of intact animals (Page et al. 1977; Page 1978). They proposed that two clock components that reside in the left and right optic lobes were mutually coupled and each clock worked to shorten the period of the other clock. The cricket clocks are more intriguing because the coupling between the two optic lobe clocks seems weaker than that of cockroaches. If the optic nerve is unilaterally disconnected from the optic lobe, this optic lobe should be blind and free-run as if it is in DD, while the contralateral optic lobe should be entrained by light cycles unless the two optic lobes exchange the light information. In this situation, crickets display two rhythms simultaneously, a phenomenon called “splitting” (Wiedenmann 1983; Tomioka et al. 1991; Tomioka 1993). The two rhythms do not run completely independently; the free-running period is modulated by the coupling of two optic lobe clocks (Tomioka et al. 1991; Tomioka 1993). Figure 5.5 shows a model of the bilateral optic lobe clocks that interact with each other to exchange zeitgeber and time information. PDF and serotonin are used in this bilateral coupling pathway (Saifullah and Tomioka 2002, 2003). In particular, PDF neurons form commissures projecting in the contralateral optic lobe (Helfrich-Förster 1997; Reischig et al. 2004), which is suitable for the coupling pathway, and its morphology is conserved across many insect species. It should also be mentioned that there are many more interneurons that bridge two sides of the optic lobes and possibly mediate the coupling (Yukizane and Tomioka 1995; Reischig and Stengl 2002).
A model of two clocks located in bilateral optic lobes in the cockroach and cricket. The left and right clocks independently receive light information from the compound eyes on each side. Although the two clocks can separately drive activity rhythms, they mutually interact to exchange time and zeitgeber information, enabling the generation of a coherent activity rhythm
A series of studies on the bilateral coupling of two optic lobe clocks have left the detailed mechanisms unknown (Page et al. 1977; Page 1978; Wiedenmann 1983; Tomioka et al. 1991; Tomioka 1993). However, these studies provide key points into insect circadian networks: (1) coupling may be needed for exchanging zeitgeber and time information, and (2) the strength of coupling may vary among species.
8 Conclusion Remarks
There is still not enough data to summarize the whole picture of insect clock networks. It would be important to try immunostainings with specific antibodies in many more insect species. Even for species that have already been studied previously, it would be significant to perform the latest fluorescent immunostaining with a confocal microscope and newly generated antibodies. In addition, neurotransmitters other than PDF have not yet been focused on nonmodel insects. In Drosophila, the first immunostaining against PER was performed in the late 1980s (Siwicki et al. 1988), but the presently known classification of clock neurons is based on studies conducted approximately in the year 2000. Surprisingly, more detailed and precise classification is still an ongoing subject (Schubert et al. 2018; Reinhard et al. 2022). Thus, the study of clock neuron networks will continue to be an active area of insect chronobiology.
References
Abdelsalam S, Uemura H, Umezaki Y, Saifullah ASM, Shimohigashi M, Tomioka K (2008) Characterization of PDF-immunoreactive neurons in the optic lobe and cerebral lobe of the cricket, Gryllus bimaculatus. J Insect Physiol 54:1205–1212. https://doi.org/10.1016/j.jinsphys.2008.05.001
Abruzzi KC, Zadina A, Luo W, Wiyanto E, Rahman R, Guo F et al (2017) RNA-seq analysis of Drosophila clock and non-clock neurons reveals neuron-specific cycling and novel candidate neuropeptides. PLoS Genet 13:e1006613. https://doi.org/10.1371/journal.pgen.1006613
Allemand R (1976) Influence de modifications des conditions lumineuses sur les rythmes circadiens de vitellogenese et d’ovulation chez Drosophila melanogaster. J Insect Physiol 22:1075–1080. https://doi.org/10.1016/0022-1910(76)90116-5
Bachleitner W, Kempinger L, Wülbeck C, Rieger D, Helfrich-Förster C (2007) Moonlight shifts the endogenous clock of Drosophila melanogaster. Proc Natl Acad Sci U S A 104:3538–3543. https://doi.org/10.1073/pnas.0606870104
Baggerman G, Cerstiaens A, De Loof A, Schoofs L (2002) Peptidomics of the larval Drosophila melanogaster central nervous system. J Biol Chem 277:40368–40374. https://doi.org/10.1074/jbc.M206257200
Barber AF, Erion R, Holmes TC, Sehgal A (2016) Circadian and feeding cues integrate to drive rhythms of physiology in Drosophila insulin-producing cells. Genes Dev 30:2596–2606. https://doi.org/10.1101/gad.288258.116
Barber AF, Fong SY, Kolesnik A, Fetchko M, Sehgal A (2021) Drosophila clock cells use multiple mechanisms to transmit time-of-day signals in the brain. Proc Natl Acad Sci U S A 118:e2019826118. https://doi.org/10.1073/pnas.2019826118
Barberà M, Collantes-Alegre JM, Martínez-Torres D (2017) Characterisation, analysis of expression and localisation of circadian clock genes from the perspective of photoperiodism in the aphid Acyrthosiphon pisum. Insect Biochem Mol Biol 83:54–67. https://doi.org/10.1016/j.ibmb.2017.02.006
Barberà M, Collantes-Alegre JM, Martínez-Torres D (2022) Mapping and quantification of cryptochrome expression in the brain of the pea aphid Acyrthosiphon pisum. Insect Mol Biol 31:159–169. https://doi.org/10.1111/imb.12747
Beer K, Helfrich-Förster C (2020) Model and non-model insects in chronobiology. Front Behav Neurosci 14:601676. https://doi.org/10.3389/fnbeh.2020.601676
Beer K, Kolbe E, Kahana NB, Yayon N, Weiss R, Menegazzi P et al (2018) Pigment-dispersing factor-expressing neurons convey circadian information in the honey bee brain. Open Biol 8:170224. https://doi.org/10.1098/rsob.170224
Bloch G, Solomon SM, Robinson GE, Fahrbach SE (2003) Patterns of PERIOD and pigment-dispersing hormone immunoreactivity in the brain of the European honeybee (Apis mellifera): age- and time-related plasticity. J Comp Neurol 464:269–284. https://doi.org/10.1002/cne.10778
Bulthuis N, Spontak KR, Kleeman B, Cavanaugh DJ (2019) Neuronal activity in non-LNv clock cells is required to produce free-running rest:activity rhythms in Drosophila. J Biol Rhythm 34:249–271. https://doi.org/10.1177/0748730419841468
Cao G, Nitabach MN (2008) Circadian control of membrane excitability in Drosophila melanogaster lateral ventral clock neurons. J Neurosci 28:6493–6501. https://doi.org/10.1523/JNEUROSCI.1503-08.2008
Cavanaugh DJ, Geratowski JD, Wooltorton JRA, Spaethling JM, Hector CE, Zheng X et al (2014) Identification of a circadian output circuit for rest:activity rhythms in Drosophila. Cell 157:689–701. https://doi.org/10.1016/j.cell.2014.02.024
Cavey M, Collins B, Bertet C, Blau J (2016) Circadian rhythms in neuronal activity propagate through output circuits. Nat Neurosci 19:587–595. https://doi.org/10.1038/nn.4263
Chatterjee A, Lamaze A, De J, Mena W, Chélot E, Martin B et al (2018) Reconfiguration of a multi-oscillator network by light in the Drosophila circadian clock. Curr Biol 28:2007–2017.e4. https://doi.org/10.1016/j.cub.2018.04.064
Chen J, Reiher W, Hermann-Luibl C, Sellami A, Cognigni P, Kondo S et al (2016) Allatostatin A signalling in Drosophila regulates feeding and sleep and is modulated by PDF. PLoS Genet 12:e1006346. https://doi.org/10.1371/journal.pgen.1006346
Chung BY, Ro J, Hutter SA, Miller KM, Guduguntla LS, Kondo S et al (2017) Drosophila neuropeptide F signaling independently regulates feeding and sleep-wake behavior. Cell Rep 19:2441–2450. https://doi.org/10.1016/j.celrep.2017.05.085
Colizzi FS, Beer K, Cuti P, Deppisch P, Martínez Torres D, Yoshii T et al (2021) Antibodies against the clock proteins Period and Cryptochrome reveal the neuronal organization of the circadian clock in the pea aphid. Front Physiol 12:705048. https://doi.org/10.3389/fphys.2021.705048
Collins B, Kane EA, Reeves DC, Akabas MH, Blau J (2012) Balance of activity between LN(v)s and glutamatergic dorsal clock neurons promotes robust circadian rhythms in Drosophila. Neuron 74:706–718. https://doi.org/10.1016/j.neuron.2012.02.034
Collins B, Kaplan HS, Cavey M, Lelito KR, Bahle AH, Zhu Z et al (2014) Differentially timed extracellular signals synchronize pacemaker neuron clocks. PLoS Biol 12:e1001959. https://doi.org/10.1371/journal.pbio.1001959
Colwell CS, Page TL (1990) A circadian rhythm in neural activity can be recorded from the central nervous system of the cockroach. J Comp Physiol A 166:643–649. https://doi.org/10.1007/BF00240014
Cusumano P, Klarsfeld A, Chelot E, Picot M, Richier B, Rouyer F (2009) PDF-modulated visual inputs and cryptochrome define diurnal behavior in Drosophila. Nat Neurosci 12:1431–1437. https://doi.org/10.1038/nn.2429
Cymborowski B (1973) Control of the circadian rhythm of locomotor activity in the house cricket. J Insect Physiol 19:1423–1440. https://doi.org/10.1016/0022-1910(73)90173-X
Delventhal R, O’Connor RM, Pantalia MM, Ulgherait M, Kim HX, Basturk MK et al (2019) Dissection of central clock function in Drosophila through cell-specific CRISPR-mediated clock gene disruption. elife 8:e48308. https://doi.org/10.7554/eLife.48308
Díaz MM, Schlichting M, Abruzzi KC, Long X, Rosbash M (2019) Allatostatin-C/AstC-R2 is a novel pathway to modulate the circadian activity pattern in Drosophila. Curr Biol 29:13–22.e3. https://doi.org/10.1016/j.cub.2018.11.005
Ding K, Han Y, Seid TW, Buser C, Karigo T, Zhang S et al (2019) Imaging neuropeptide release at synapses with a genetically engineered reporter. elife 8:e46421. https://doi.org/10.7554/eLife.46421
Dreyer AP, Martin MM, Fulgham CV, Jabr DA, Bai L, Beshel J et al (2019) A circadian output center controlling feeding:fasting rhythms in Drosophila. PLoS Genet 15:e1008478. https://doi.org/10.1371/journal.pgen.1008478
Duhart JM, Herrero A, de la Cruz G, Ispizua JI, Pírez N, Ceriani MF (2020) Circadian structural plasticity drives remodeling of E cell output. Curr Biol 30:5040–5048.e5. https://doi.org/10.1016/j.cub.2020.09.057
Dushay MS, Rosbash M, Hall JC (1989) The disconnected visual system mutations in Drosophila melanogaster drastically disrupt circadian rhythms. J Biol Rhythm 4:1–27. https://doi.org/10.1177/074873048900400101
Fernández MP, Berni J, Ceriani MF (2008) Circadian remodeling of neuronal circuits involved in rhythmic behavior. PLoS Biol 6:e69. https://doi.org/10.1371/journal.pbio.0060069
Fernandez MP, Pettibone HL, Bogart JT, Roell CJ, Davey CE, Pranevicius A et al (2020) Sites of circadian clock Neuron plasticity mediate sensory integration and entrainment. Curr Biol 30:2225–2237.e5. https://doi.org/10.1016/j.cub.2020.04.025
Fleissner G (1982) Isolation of an insect circadian clock. J Comp Physiol 149:311–316. https://doi.org/10.1007/BF00619146
Flyer-Adams JG, Rivera-Rodriguez EJ, Yu J, Mardovin JD, Reed ML, Griffith LC (2020) Regulation of olfactory associative memory by the circadian clock output signal Pigment-Dispersing Factor (PDF). J Neurosci 40:9066–9077. https://doi.org/10.1523/JNEUROSCI.0782-20.2020
Frenkel L, Muraro NI, Beltrán González AN, Marcora MS, Bernabó G, Hermann-Luibl C et al (2017) Organization of circadian behavior relies on glycinergic transmission. Cell Rep 19:72–85. https://doi.org/10.1016/j.celrep.2017.03.034
Frisch B, Fleissner GG, Brandes C, Hall JC (1996) Staining in the brain of Pachymorpha sexguttata mediated by an antibody against a Drosophila clock-gene product: labeling of cells with possible importance for the beetle’s circadian rhythms. Cell Tissue Res 286:411–429. https://doi.org/10.1007/s004410050711
Fuchikawa T, Beer K, Linke-Winnebeck C, Ben-David R, Kotowoy A, Tsang VWK et al (2017) Neuronal circadian clock protein oscillations are similar in behaviourally rhythmic forager honeybees and in arrhythmic nurses. Open Biol 7:170047. https://doi.org/10.1098/rsob.170047
Fujii S, Amrein H (2010) Ventral lateral and DN1 clock neurons mediate distinct properties of male sex drive rhythm in Drosophila. Proc Natl Acad Sci U S A 107:10590–10595. https://doi.org/10.1073/pnas.0912457107
Fujiwara Y, Hermann-Luibl C, Katsura M, Sekiguchi M, Ida T, Helfrich-Förster C et al (2018) The CCHamide1 neuropeptide expressed in the anterior dorsal neuron 1 conveys a circadian signal to the ventral lateral neurons in Drosophila melanogaster. Front Physiol 9:1276. https://doi.org/10.3389/fphys.2018.01276
Goda T, Doi M, Umezaki Y, Murai I, Shimatani H, Chu ML et al (2018) Calcitonin receptors are ancient modulators for rhythms of preferential temperature in insects and body temperature in mammals. Genes Dev 32:140–155. https://doi.org/10.1101/gad.307884.117
Goda T, Tang X, Umezaki Y, Chu ML, Kunst M, Nitabach MN et al (2016) Drosophila DH31 neuropeptide and PDF receptor regulate night-onset temperature preference. J Neurosci 36:11739–11754. https://doi.org/10.1523/JNEUROSCI.0964-16.2016
Goda T, Umezaki Y, Alwattari F, Seo HW, Hamada FN (2019) Neuropeptides PDF and DH31 hierarchically regulate free-running rhythmicity in Drosophila circadian locomotor activity. Sci Rep 9:838. https://doi.org/10.1038/s41598-018-37107-3
Gorostiza EA, Depetris-Chauvin A, Frenkel L, Pírez N, Ceriani MF (2014) Circadian pacemaker neurons change synaptic contacts across the day. Curr Biol 24:2161–2167. https://doi.org/10.1016/j.cub.2014.07.063
Grima B, Chelot E, Xia R, Rouyer F (2004) Morning and evening peaks of activity rely on different clock neurons of the Drosophila brain. Nature 431:869–873. https://doi.org/10.1038/nature02935
Guo F, Chen X, Rosbash M (2017) Temporal calcium profiling of specific circadian neurons in freely moving flies. Proc Natl Acad Sci U S A 114:E8780–E8787. https://doi.org/10.1073/pnas.1706608114
Guo F, Holla M, Díaz MM, Rosbash M (2018) A circadian output circuit controls sleep-wake arousal in Drosophila. Neuron 100:624–635.e4. https://doi.org/10.1016/j.neuron.2018.09.002
Guo F, Yu J, Jung HJ, Abruzzi KC, Luo W, Griffith LC et al (2016) Circadian neuron feedback controls the Drosophila sleep-activity profile. Nature 536:292–297. https://doi.org/10.1038/nature19097
Hamasaka Y, Rieger D, Parmentier ML, Grau Y, Helfrich-Förster C, Nässel DR (2007) Glutamate and its metabotropic receptor in Drosophila clock neuron circuits. J Comp Neurol 505:32–45. https://doi.org/10.1002/cne.21471
Hamblen-Coyle MJ, Wheeler DA, Rutila JE, Rosbash M, Hall JC (1992) Behavior of period-altered circadian rhythm mutants of Drosophila in light: dark cycles (Diptera: Drosophilidae). J Insect Behav 5:417–446. https://doi.org/10.1007/BF01058189
Hanafusa S, Kawaguchi T, Umezaki Y, Tomioka K, Yoshii T (2013) Sexual interactions influence the molecular oscillations in DN1 pacemaker neurons in Drosophila melanogaster. PLoS One 8:e84495. https://doi.org/10.1371/journal.pone.0084495
Hardin PE, Hall JC, Rosbash M (1992) Behavioral and molecular analyses suggest that circadian output is disrupted by disconnected mutants in D. melanogaster. EMBO J 11:1–6. https://doi.org/10.1002/j.1460-2075.1992.tb05020.x
Hassaneen E, El-Din Sallam A, Abo-Ghalia A, Moriyama Y, Karpova SG, Abdelsalam S et al (2011) Pigment-dispersing factor affects nocturnal activity rhythms, photic entrainment, and the free-running period of the circadian clock in the cricket Gryllus bimaculatus. J Biol Rhythm 26:3–13. https://doi.org/10.1177/0748730410388746
Helfrich-Förster C (1995) The period clock gene is expressed in central nervous system neurons which also produce a neuropeptide that reveals the projections of circadian pacemaker cells within the brain of Drosophila melanogaster. Proc Natl Acad Sci U S A 92:612–616. https://doi.org/10.1073/pnas.92.2.612
Helfrich-Förster C (1997) Development of pigment-dispersing hormone-immunoreactive neurons in the nervous system of Drosophila melanogaster. J Comp Neurol 380:335–354. https://doi.org/10.1002/(sici)1096-9861(19970414)380:3<335::aid-cne4>3.0.co;2-3
Helfrich-Förster C (1998) Robust circadian rhythmicity of Drosophila melanogaster requires the presence of lateral neurons: a brain-behavioral study of disconnected mutants. J Comp Physiol A 182:435–453. https://doi.org/10.1007/s003590050192
Helfrich-Förster C (2000) Differential control of morning and evening components in the activity rhythm of Drosophila melanogaster—sex-specific differences suggest a different quality of activity. J Biol Rhythm 15:135–154. https://doi.org/10.1177/074873040001500208
Helfrich-Förster C (2003) The neuroarchitecture of the circadian clock in the brain of Drosophila melanogaster. Microsc Res Tech 62:94–102. https://doi.org/10.1002/jemt.10357
Helfrich-Förster C (2005) Organization of endogenous clocks in insects. Biochem Soc Trans 33:957–961. https://doi.org/10.1042/BST20050957
Helfrich-Förster C, Shafer OT, Wülbeck C, Grieshaber E, Rieger D, Taghert P (2007) Development and morphology of the clock-gene-expressing lateral neurons of Drosophila melanogaster. J Comp Neurol 500:47–70. https://doi.org/10.1002/cne.21146
Hendricks JC, Finn SM, Panckeri KA, Chavkin J, Williams JA, Sehgal A et al (2000) Rest in Drosophila is a sleep-like state. Neuron 25:129–138. https://doi.org/10.1016/s0896-6273(00)80877-6
Hermann C, Saccon R, Senthilan PR, Domnik L, Dircksen H, Yoshii T et al (2013) The circadian clock network in the brain of different Drosophila species. J Comp Neurol 521:367–388. https://doi.org/10.1002/cne.23178
Hermann C, Yoshii T, Dusik V, Helfrich-Förster C (2012) Neuropeptide F immunoreactive clock neurons modify evening locomotor activity and free-running period in Drosophila melanogaster. J Comp Neurol 520:970–987. https://doi.org/10.1002/cne.22742
Hermann-Luibl C, Yoshii T, Senthilan PR, Dircksen H, Helfrich-Förster C (2014) The ion transport peptide is a new functional clock neuropeptide in the fruit fly Drosophila melanogaster. J Neurosci 34:9522–9536. https://doi.org/10.1523/JNEUROSCI.0111-14.2014
Huybrechts J, Bonhomme J, Minoli S, Prunier-Leterme N, Dombrovsky A, Abdel-Latief M et al (2010) Neuropeptide and neurohormone precursors in the pea aphid, Acyrthosiphon pisum. Insect Mol Biol 19:87–95. https://doi.org/10.1111/j.1365-2583.2009.00951.x
Hyun S, Lee Y, Hong ST, Bang S, Paik D, Kang J et al (2005) Drosophila GPCR Han is a receptor for the circadian clock neuropeptide PDF. Neuron 48:267–278. https://doi.org/10.1016/j.neuron.2005.08.025
Im SH, Taghert PH (2010) PDF receptor expression reveals direct interactions between circadian oscillators in Drosophila. J Comp Neurol 518:1925–1945. https://doi.org/10.1002/cne.22311
Inami S, Sato T, Kurata Y, Suzuki Y, Kitamoto T, Sakai T (2021) Consolidation and maintenance of long-term memory involve dual functions of the developmental regulator Apterous in clock neurons and mushroom bodies in the Drosophila brain. PLoS Biol 19:e3001459. https://doi.org/10.1371/journal.pbio.3001459
Ito C, Tomioka K (2016) Heterogeneity of the peripheral circadian systems in Drosophila melanogaster: a review. Front Physiol 7:8. https://doi.org/10.3389/fphy.2016.00008
Jaumouillé E, Koch R, Nagoshi E (2021) Uncovering the roles of clocks and neural transmission in the resilience of Drosophila circadian network. Front Physiol 12:663339. https://doi.org/10.3389/fphys.2021.663339
Jin X, Tian Y, Zhang ZC, Gu P, Liu C, Han J (2021) A subset of DN1p neurons integrates thermosensory inputs to promote wakefulness via CNMa signaling. Curr Biol 31:2075–2087.e6. https://doi.org/10.1016/j.cub.2021.02.048
Johard HA, Yoishii T, Dircksen H, Cusumano P, Rouyer F, Helfrich-Förster C et al (2009) Peptidergic clock neurons in Drosophila: ion transport peptide and short neuropeptide F in subsets of dorsal and ventral lateral neurons. J Comp Neurol 516:59–73. https://doi.org/10.1002/cne.22099
Kaneko H, Head LM, Ling J, Tang X, Liu Y, Hardin PE et al (2012) Circadian rhythm of temperature preference and its neural control in Drosophila. Curr Biol 22:1851–1857. https://doi.org/10.1016/j.cub.2012.08.006
Kaneko M, Hall JC (2000) Neuroanatomy of cells expressing clock genes in Drosophila: transgenic manipulation of the period and timeless genes to mark the perikarya of circadian pacemaker neurons and their projections. J Comp Neurol 422:66–94. https://doi.org/10.1002/(sici)1096-9861(20000619)422:1<66::aid-cne5>3.0.co;2-2
Kay J, Menegazzi P, Mildner S, Roces F, Helfrich-Förster C (2018) The circadian clock of the ant Camponotus floridanus is localized in dorsal and lateral neurons of the brain. J Biol Rhythm 33:255–271. https://doi.org/10.1177/0748730418764738
Klose MK, Bruchez MP, Deitcher DL, Levitan ES (2021) Temporally and spatially partitioned neuropeptide release from individual clock neurons. Proc Natl Acad Sci U S A 118:e2101818118. https://doi.org/10.1073/pnas.2101818118
Kobelková A, Závodská R, Sauman I, Bazalová O, Dolezel D (2015) Expression of clock genes period and timeless in the central nervous system of the Mediterranean flour moth, Ephestia kuehniella. J Biol Rhythm 30:104–116. https://doi.org/10.1177/0748730414568430
Koide R, Xi J, Hamanaka Y, Shiga S (2021) Mapping PERIOD-immunoreactive cells with neurons relevant to photoperiodic response in the bean bug Riptortus pedestris. Cell Tissue Res 385:571–583. https://doi.org/10.1007/s00441-021-03451-6
Konopka RJ, Benzer S (1971) Clock mutants of Drosophila melanogaster. Proc Natl Acad Sci U S A 68:2112–2116. https://doi.org/10.1073/pnas.68.9.2112
Kotwica-Rolinska J, Damulewicz M, Chodakova L, Kristofova L, Dolezel D (2022) Pigment dispersing factor is a circadian clock output and regulates photoperiodic response in the linden bug, Pyrrhocoris apterus. Front Physiol 13:884909. https://doi.org/10.3389/fphys.2022.884909
Kula E, Levitan ES, Pyza E, Rosbash M (2006) PDF cycling in the dorsal protocerebrum of the Drosophila brain is not necessary for circadian clock function. J Biol Rhythm 21:104–117. https://doi.org/10.1177/0748730405285715
Kunst M, Hughes ME, Raccuglia D, Felix M, Li M, Barnett G et al (2014) Calcitonin gene-related peptide neurons mediate sleep-specific circadian output in Drosophila. Curr Biol 24:2652–2664. https://doi.org/10.1016/j.cub.2014.09.077
Kutaragi Y, Tokuoka A, Tomiyama Y, Nose M, Watanabe T, Bando T et al (2018) A novel photic entrainment mechanism for the circadian clock in an insect: involvement of c-fos and cryptochromes. Zool Lett 4:26. https://doi.org/10.1186/s40851-018-0109-8
Lamaze A, Krätschmer P, Chen KF, Lowe S, Jepson JEC (2018) A wake-promoting circadian output circuit in Drosophila. Curr Biol 28:3098–3105.e3. https://doi.org/10.1016/j.cub.2018.07.024
Lear BC, Merrill CE, Lin JM, Schroeder A, Zhang L, Allada R (2005) A G protein-coupled receptor, groom-of-PDF, is required for PDF neuron action in circadian behavior. Neuron 48:221–227. https://doi.org/10.1016/j.neuron.2005.09.008
Lear BC, Zhang L, Allada R (2009) The neuropeptide PDF acts directly on evening pacemaker neurons to regulate multiple features of circadian behavior. PLoS Biol 7:e1000154. https://doi.org/10.1371/journal.pbio.1000154
Lee CM, Su MT, Lee HJ (2009) Pigment dispersing factor: an output regulator of the circadian clock in the German cockroach. J Biol Rhythm 24:35–43. https://doi.org/10.1177/0748730408327909
Lelito KR, Shafer OT (2012) Reciprocal cholinergic and GABAergic modulation of the small ventrolateral pacemaker neurons of Drosophila’s circadian clock neuron network. J Neurophysiol 107:2096–2108. https://doi.org/10.1152/jn.00931.2011
Leopold AV, Shcherbakova DM, Verkhusha VV (2019) Fluorescent biosensors for neurotransmission and neuromodulation: engineering and applications. Front Cell Neurosci 13:474. https://doi.org/10.3389/fncel.2019.00474
Li MT, Cao LH, Xiao N, Tang M, Deng B, Yang T et al (2018) Hub-organized parallel circuits of central circadian pacemaker neurons for visual photoentrainment in Drosophila. Nat Commun 9:4247. https://doi.org/10.1038/s41467-018-06506-5
Liang X, Ho MCW, Zhang Y, Li Y, Wu MN, Holy TE et al (2019) Morning and evening circadian pacemakers independently drive premotor centers via a specific dopamine relay. Neuron 102:843–857.e4. https://doi.org/10.1016/j.neuron.2019.03.028
Liang X, Holy TE, Taghert PH (2016) Synchronous Drosophila circadian pacemakers display nonsynchronous Ca2+ rhythms in vivo. Science 351:976–981. https://doi.org/10.1126/science.aad3997
Liang X, Holy TE, Taghert PH (2017) A series of suppressive signals within the Drosophila circadian neural circuit generates sequential daily outputs. Neuron 94:1173–1189.e4. https://doi.org/10.1016/j.neuron.2017.05.007
Lin Y, Stormo GD, Taghert PH (2004) The neuropeptide pigment-dispersing factor coordinates pacemaker interactions in the Drosophila circadian system. J Neurosci 24:7951–7957. https://doi.org/10.1523/JNEUROSCI.2370-04.2004
Liu X, Lorenz L, Yu QN, Hall JC, Rosbash M (1988) Spatial and temporal expression of the period gene in Drosophila melanogaster. Genes Dev 2:228–238. https://doi.org/10.1101/gad.2.2.228
Loher W (1972) Circadian control of stridulation in the cricket Teleogryllus commodus Walker. J Comp Physiol 79:173–190. https://doi.org/10.1007/BF00697770
Lupien M, Marshall S, Leser W, Pollack GS, Honegger HW (2003) Antibodies against the PER protein of Drosophila label neurons in the optic lobe, central brain, and thoracic ganglia of the crickets Teleogryllus commodus and Teleogryllus oceanicus. Cell Tissue Res 312:377–391. https://doi.org/10.1007/s00441-003-0720-6
Ma D, Przybylski D, Abruzzi KC, Schlichting M, Li Q, Long X et al (2021) A transcriptomic taxonomy of Drosophila circadian neurons around the clock. elife 10:e63056. https://doi.org/10.7554/eLife.63056
McCarthy EV, Wu Y, Decarvalho T, Brandt C, Cao G, Nitabach MN (2011) Synchronized bilateral synaptic inputs to Drosophila melanogaster neuropeptidergic rest/arousal neurons. J Neurosci 31:8181–8193. https://doi.org/10.1523/JNEUROSCI.2017-10.2011
Meelkop E, Temmerman L, Schoofs L, Janssen T (2011) Signalling through pigment dispersing hormone-like peptides in invertebrates. Prog Neurobiol 93:125–147. https://doi.org/10.1016/j.pneurobio.2010.10.004
Meiselman MR, Alpert MH, Cui X, Shea J, Gregg I, Gallio M et al (2022) Recovery from cold-induced reproductive dormancy is regulated by temperature-dependent AstC signaling. Curr Biol 32:1362–1375. https://doi.org/10.1016/j.cub.2022.01.061
Menegazzi P, Beer K, Grebler V, Schlichting M, Schubert FK, Helfrich-Förster C (2020) A functional clock within the main morning and evening neurons of D. melanogaster is not sufficient for wild-type locomotor activity under changing day length. Front Physiol 11:229. https://doi.org/10.3389/fphys.2020.00229
Menegazzi P, Dalla Benetta E, Beauchamp M, Schlichting M, Steffan-Dewenter I, Helfrich-Förster C (2017) Adaptation of circadian neuronal network to photoperiod in high-latitude European Drosophilids. Curr Biol 27:833–839. https://doi.org/10.1016/j.cub.2017.01.036
Mertens I, Vandingenen A, Johnson EC, Shafer OT, Li W, Trigg JS et al (2005) PDF receptor signaling in Drosophila contributes to both circadian and geotactic behaviors. Neuron 48:213–219. https://doi.org/10.1016/j.neuron.2005.09.009
Mieda M (2020) The central circadian clock of the suprachiasmatic nucleus as an ensemble of multiple oscillatory neurons. Neurosci Res 156:24–31. https://doi.org/10.1016/j.neures.2019.08.003
Mohamed AA, Wang Q, Bembenek J, Ichihara N, Hiragaki S, Suzuki T et al (2014) N-acetyltransferase (nat) is a critical conjunct of photoperiodism between the circadian system and endocrine axis in Antheraea pernyi. PLoS One 9:e92680. https://doi.org/10.1371/journal.pone.0092680
Nagy D, Cusumano P, Andreatta G, Anduaga AM, Hermann-Luibl C, Reinhard N et al (2019) Peptidergic signaling from clock neurons regulates reproductive dormancy in Drosophila melanogaster. PLoS Genet 15:e1008158. https://doi.org/10.1371/journal.pgen.1008158
Nettnin EA, Sallese TR, Nasseri A, Saurabh S, Cavanaugh DJ (2021) Dorsal clock neurons in Drosophila sculpt locomotor outputs but are dispensable for circadian activity rhythms. iScience 24:103001. https://doi.org/10.1016/j.isci.2021.103001
Ni JD, Gurav AS, Liu W, Ogunmowo TH, Hackbart H, Elsheikh A et al (2019) Differential regulation of the Drosophila sleep homeostat by circadian and arousal inputs. elife 8:e40487. https://doi.org/10.7554/eLife.40487
Nishiitsutsuji-Uwo J, Petropulos SF, Pittendrigh CS (1967) Central nervous system control of circadian rhythmicity in the cockroach. I. Role of the pars intercerebralis. Biol Bull 133:679–696. https://doi.org/10.2307/1539928
Nishiitsutsuji-Uwo J, Pittendrigh C (1968) Central nervous system control of circadian rhythmicity in the cockroach. III. The optic lobes, locus of the driving oscillation? Z Vergl Physiol 58:14–46. https://doi.org/10.1007/BF00302434
Okamoto A, Mori H, Tomioka K (2001) The role of the optic lobe in circadian locomotor rhythm generation in the cricket, Gryllus bimaculatus, with special reference to PDH-immunoreactive neurons. J Insect Physiol 47:889–895. https://doi.org/10.1016/s0022-1910(01)00061-0
Ono D, Honma KI, Honma S (2021) Roles of neuropeptides, VIP and AVP, in the mammalian central circadian clock. Front Neurosci 15:650154. https://doi.org/10.3389/fnins.2021.650154
Page TL (1978) Interactions between bilaterally paired components of the cockroach circadian system. J Comp Physiol 124:225–236. https://doi.org/10.1007/BF00657054
Page TL (1982) Transplantation of the cockroach circadian pacemaker. Science 216:73–75. https://doi.org/10.1126/science.216.4541.73
Page TL, Caldarola PC, Pittendrigh CS (1977) Mutual entrainment of bilaterally distributed circadian pacemaker. Proc Natl Acad Sci U S A 74:1277–1281. https://doi.org/10.1073/pnas.74.3.1277
Panda S, Sato TK, Castrucci AM, Rollag MD, DeGrip WJ, Hogenesch JB et al (2002) Melanopsin (Opn4) requirement for normal light-induced circadian phase shifting. Science 298:2213–2216. https://doi.org/10.1126/science.1076848
Park JH, Helfrich-Förster C, Lee G, Liu L, Rosbash M, Hall JC (2000) Differential regulation of circadian pacemaker output by separate clock genes in Drosophila. Proc Natl Acad Sci U S A 97:3608–3613. https://doi.org/10.1073/pnas.070036197
Peng Y, Stoleru D, Levine JD, Hall JC, Rosbash M (2003) Drosophila free-running rhythms require intercellular communication. PLoS Biol 1:32–40. https://doi.org/10.1371/journal.pbio.0000013
Petri B, Stengl M (1997) Pigment-dispersing hormone shifts the phase of the circadian pacemaker of the cockroach Leucophaea maderae. J Neurosci 17:4087–4093. https://doi.org/10.1523/JNEUROSCI.17-11-04087.1997
Pittendrigh CS, Daan S (1976) A functional analysis of circadian pacemakers in nocturnal rodents. V. Pacemaker structure: a clock for all seasons. J Comp Physiol A 106:333–355. https://doi.org/10.1007/BF01417860
Prakash P, Nambiar A, Sheeba V (2017) Oscillating PDF in termini of circadian pacemaker neurons and synchronous molecular clocks in downstream neurons are not sufficient for sustenance of activity rhythms in constant darkness. PLoS One 12:e0175073. https://doi.org/10.1371/journal.pone.0175073
Ramakrishnan A, Sheeba V (2021) Gap junction protein Innexin2 modulates the period of free-running rhythms in Drosophila melanogaster. iScience 24:103011. https://doi.org/10.1016/j.isci.2021.103011
Reinhard N, Bertolini E, Saito A, Sekiguchi M, Yoshii T, Rieger D et al (2022) The lateral posterior clock neurons of Drosophila melanogaster express three neuropeptides and have multiple connections within the circadian clock network and beyond. J Comp Neurol 530:1507–1529. https://doi.org/10.1002/cne.25294
Reischig T, Petri B, Stengl M (2004) Pigment-dispersing hormone (PDH)-immunoreactive neurons form a direct coupling pathway between the bilaterally symmetric circadian pacemakers of the cockroach Leucophaea maderae. Cell Tissue Res 318:553–564. https://doi.org/10.1007/s00441-004-0927-1
Reischig T, Stengl M (2002) Optic lobe commissures in a three-dimensional brain model of the cockroach Leucophaea maderae: a search for the circadian coupling pathways. J Comp Neurol 443:388–400. https://doi.org/10.1002/cne.10133
Reischig T, Stengl M (2003) Ectopic transplantation of the accessory medulla restores circadian locomotor rhythms in arrhythmic cockroaches (Leucophaea maderae). J Exp Biol 206:1877–1886. https://doi.org/10.1242/jeb.00373
Renn SC, Park JH, Rosbash M, Hall JC, Taghert PH (1999) A pdf neuropeptide gene mutation and ablation of PDF neurons each cause severe abnormalities of behavioral circadian rhythms in Drosophila. Cell 99:791–802. https://doi.org/10.1016/s0092-8674(00)81676-1
Rieger D, Shafer OT, Tomioka K, Helfrich-Förster C (2006) Functional analysis of circadian pacemaker neurons in Drosophila melanogaster. J Neurosci 26:2531–2543. https://doi.org/10.1523/JNEUROSCI.1234-05.2006
Rieger D, Wülbeck C, Rouyer F, Helfrich-Förster C (2009) Period gene expression in four neurons is sufficient for rhythmic activity of Drosophila melanogaster under dim light conditions. J Biol Rhythm 24:271–282. https://doi.org/10.1177/0748730409338508
Ruby NF, Brennan TJ, Xie X, Cao V, Franken P, Heller HC et al (2002) Role of melanopsin in circadian responses to light. Science 298:2211–2213. https://doi.org/10.1126/science.1076701
Saifullah ASM, Tomioka K (2002) Serotonin sets the day state in the neurons that control coupling between the optic lobe circadian pacemakers in the cricket Gryllus bimaculatus. J Exp Biol 205:1305–1314. https://doi.org/10.1242/jeb.205.9.1305
Saifullah ASM, Tomioka K (2003) Pigment-dispersing factor sets the night state of the medulla bilateral neurons in the optic lobe of the cricket, Gryllus bimaculatus. J Insect Physiol 49:231–239. https://doi.org/10.1016/s0022-1910(02)00270-6
Sauman I, Briscoe AD, Zhu H, Shi D, Froy O, Stalleicken J et al (2005) Connecting the navigational clock to sun compass input in monarch butterfly brain. Neuron 46:457–467. https://doi.org/10.1016/j.neuron.2005.03.014
Sauman I, Reppert SM (1996) Circadian clock neurons in the silkmoth Antheraea pernyi: novel mechanisms of Period protein regulation. Neuron 17:889–900. https://doi.org/10.1016/s0896-6273(00)80220-2
Scheffer LK, Xu CS, Januszewski M, Lu Z, Takemura SY, Hayworth KJ et al (2020) A connectome and analysis of the adult Drosophila central brain. elife 9:e57443. https://doi.org/10.7554/eLife.57443
Schlichting M, Díaz MM, Xin J, Rosbash M (2019a) Neuron-specific knockouts indicate the importance of network communication to Drosophila rhythmicity. elife 8:e48301. https://doi.org/10.7554/eLife.48301
Schlichting M, Menegazzi P, Lelito KR, Yao Z, Buhl E, Dalla Benetta E et al (2016) A neural network underlying circadian entrainment and photoperiodic adjustment of sleep and activity in Drosophila. J Neurosci 36:9084–9096. https://doi.org/10.1523/JNEUROSCI.0992-16.2016
Schlichting M, Weidner P, Diaz M, Menegazzi P, Dalla Benetta E, Helfrich-Förster C et al (2019b) Light-mediated circuit switching in the Drosophila neuronal clock network. Curr Biol 29:3266–3276.e3. https://doi.org/10.1016/j.cub.2019.08.033
Schneider NL, Stengl M (2006) Gap junctions between accessory medulla neurons appear to synchronize circadian clock cells of the cockroach Leucophaea maderae. J Neurophysiol 95:1996–2002. https://doi.org/10.1152/jn.00835.2005
Schubert FK, Hagedorn N, Yoshii T, Helfrich-Förster C, Rieger D (2018) Neuroanatomical details of the lateral neurons of Drosophila melanogaster support their functional role in the circadian system. J Comp Neurol 526:1209–1231. https://doi.org/10.1002/cne.24406
Sehadová H, Markova EP, Sehnal F, Takeda M (2004) Distribution of circadian clock-related proteins in the cephalic nervous system of the silkworm, Bombyx mori. J Biol Rhythm 19:466–482. https://doi.org/10.1177/0748730404269153
Selcho M, Millán C, Palacios-Muñoz A, Ruf F, Ubillo L, Chen J et al (2017) Central and peripheral clocks are coupled by a neuropeptide pathway in Drosophila. Nat Commun 8:15563. https://doi.org/10.1038/ncomms15563
Shafer OT, Helfrich-Förster C, Renn SC, Taghert PH (2006) Reevaluation of Drosophila melanogaster’s neuronal circadian pacemakers reveals new neuronal classes. J Comp Neurol 498:180–193. https://doi.org/10.1002/cne.21021
Shafer OT, Kim DJ, Dunbar-Yaffe R, Nikolaev VO, Lohse MJ, Taghert PH (2008) Widespread receptivity to neuropeptide PDF throughout the neuronal circadian clock network of Drosophila revealed by real-time cyclic AMP imaging. Neuron 58:223–237. https://doi.org/10.1016/j.neuron.2008.02.018
Shafer OT, Taghert PH (2009) RNA-interference knockdown of Drosophila pigment dispersing factor in neuronal subsets: the anatomical basis of a neuropeptide’s circadian functions. PLoS One 4:e8298. https://doi.org/10.1371/journal.pone.0008298
Shao QM, Bembenek J, Trang le TD, Hiragaki S, Takeda M (2008a) Molecular structure, expression patterns, and localization of the circadian transcription modulator CYCLE in the cricket, Dianemobius nigrofasciatus. J Insect Physiol 54:403–413. https://doi.org/10.1016/j.jinsphys.2007.10.013
Shao QM, Hiragaki S, Takeda M (2008b) Co-localization and unique distributions of two clock proteins CYCLE and CLOCK in the cephalic ganglia of the ground cricket, Allonemobius allardi. Cell Tissue Res 331:435–446. https://doi.org/10.1007/s00441-007-0534-Z
Shao QM, Sehadová H, Ichihara N, Sehnal F, Takeda M (2006) Immunoreactivities to three circadian clock proteins in two ground crickets suggest interspecific diversity of the circadian clock structure. J Biol Rhythm 21:118–131. https://doi.org/10.1177/0748730405283660
Shaw PJ, Cirelli C, Greenspan RJ, Tononi G (2000) Correlates of sleep and waking in Drosophila melanogaster. Science 287:1834–1837. https://doi.org/10.1126/science.287.5459.1834
Sheeba V, Fogle KJ, Holmes TC (2010) Persistence of morning anticipation behavior and high amplitude morning startle response following functional loss of small ventral lateral neurons in Drosophila. PLoS One 5:e11628. https://doi.org/10.1371/journal.pone.0011628
Shiga S, Numata H (2009) Roles of PER immunoreactive neurons in circadian rhythms and photoperiodism in the blow fly, Protophormia terraenovae. J Exp Biol 212:867–877. https://doi.org/10.1242/jeb.027003
Singaravel M, Fujisawa Y, Hisada M, Saifullah AS, Tomioka K (2003) Phase shifts of the circadian locomotor rhythm induced by pigment-dispersing factor in the cricket Gryllus bimaculatus. Zool Sci 20:1347–1354. https://doi.org/10.2108/zsj.20.1347
Siwicki KK, Eastman C, Petersen G, Rosbash M, Hall JC (1988) Antibodies to the period gene product of Drosophila reveal diverse tissue distribution and rhythmic changes in the visual system. Neuron 1:141–150. https://doi.org/10.1016/0896-6273(88)90198-5
Sokolove PG, Loher W (1975) Rôle of eyes, optic lobes, and pars intercerebralis in locomotory and stridulatory circadian rhythms of Teleogryllus commodus. J Insect Physiol 21:785–799. https://doi.org/10.1016/0022-1910(75)90009-8
Song BJ, Sharp SJ, Rogulja D (2021) Daily rewiring of a neural circuit generates a predictive model of environmental light. Sci Adv 7:eabe4284. https://doi.org/10.1126/sciadv.abe4284
Stoleru D, Peng Y, Agosto J, Rosbash M (2004) Coupled oscillators control morning and evening locomotor behaviour of Drosophila. Nature 431:862–868. https://doi.org/10.1038/nature02926
Takekata H, Numata H, Shiga S (2018) Effects of pars intercerebralis removal on circatidal rhythm in the mangrove cricket, Apteronemobius asahinai. J Comp Physiol A 204:801–810. https://doi.org/10.1007/s00359-018-1281-1
Tomioka K (1993) Analysis of coupling between optic lobe circadian pacemakers in the cricket Gryllus bimaculatus. J Comp Physiol A 172:401–408. https://doi.org/10.1007/BF00213522
Tomioka K, Chiba Y (1984) Effects of nymphal stage optic nerve severance or optic lobe removal on the circadian locomotor rhythm of the cricket, Gryllus bimaculatus. Zool Sci 1:385–394. https://doi.org/10.34425/zs000039
Tomioka K, Chiba Y (1986) Circadian rhythm in the neurally isolated lamina-medulla-complex of the cricket, Gryllus bimaculatus. J Insect Physiol 32:747–755. https://doi.org/10.1016/0022-1910(86)90077-6
Tomioka K, Chiba Y (1992) Characterization of optic lobe circadian pacemaker by in situ and in vitro recording of neuronal activity in the cricket Gryllus bimaculatus. J Comp Physiol A 171:1–7. https://doi.org/10.1007/BF00195955
Tomioka K, Yamada K, Yokoyama S, Chiba Y (1991) Mutual interactions between optic lobe circadian pacemakers in the cricket Gryllus bimaculatus. J Comp Physiol A 169:291–298. https://doi.org/10.1007/BF00206993
Vafopoulou X, Terry KL, Steel CG (2010) The circadian timing system in the brain of the fifth larval instar of Rhodnius prolixus (hemiptera). J Comp Neurol 518:1264–1282. https://doi.org/10.1002/cne.22274
Wei H, Stengl M (2011) Light affects the branching pattern of peptidergic circadian pacemaker neurons in the brain of the cockroach Leucophaea maderae. J Biol Rhythm 26:507–517. https://doi.org/10.1177/0748730411419968
Wen CJ, Lee HJ (2008) Mapping the cellular network of the circadian clock in two cockroach species. Arch Insect Biochem Physiol 68:215–231. https://doi.org/10.1002/arch.20236
Wiedenmann G (1983) Splitting in a circadian activity rhythm: the expression of bilaterally paired oscillators. J Comp Physiol 150:51–60. https://doi.org/10.1007/BF00605287
Wise S, Davis NT, Tyndale E, Noveral J, Folwell MG, Bedian V et al (2002) Neuroanatomical studies of period gene expression in the hawkmoth, Manduca sexta. J Comp Neurol 447:366–380. https://doi.org/10.1002/cne.10242
Yadlapalli S, Jiang C, Bahle A, Reddy P, Meyhofer E, Shafer OT (2018) Circadian clock neurons constantly monitor environmental temperature to set sleep timing. Nature 555:98–102. https://doi.org/10.1038/nature25740
Yang Z, Sehgal A (2001) Role of molecular oscillations in generating behavioral rhythms in Drosophila. Neuron 29:453–467. https://doi.org/10.1016/s0896-6273(01)00218-5
Yao Z, Bennett AJ, Clem JL, Shafer OT (2016) The Drosophila clock neuron network features diverse coupling modes and requires network-wide coherence for robust circadian rhythms. Cell Rep 17:2873–2881. https://doi.org/10.1016/j.celrep.2016.11.053
Yao Z, Shafer OT (2014) The Drosophila circadian clock is a variably coupled network of multiple peptidergic units. Science 343:1516–1520. https://doi.org/10.1126/science.1251285
Yoshii T, Rieger D, Helfrich-Förster C (2012) Two clocks in the brain: an update of the morning and evening oscillator model in Drosophila. Prog Brain Res 199:59–82. https://doi.org/10.1016/B978-0-444-59427-3.00027-7
Yoshii T, Todo T, Wülbeck C, Stanewsky R, Helfrich-Förster C (2008) Cryptochrome is present in the compound eyes and a subset of Drosophila’s clock neurons. J Comp Neurol 508:952–966. https://doi.org/10.1002/cne.21702
Yoshii T, Wülbeck C, Sehadova H, Veleri S, Bichler D, Stanewsky R et al (2009) The neuropeptide pigment-dispersing factor adjusts period and phase of Drosophila’s clock. J Neurosci 29:2597–2610. https://doi.org/10.1523/JNEUROSCI.5439-08.2009
Yukizane M, Tomioka K (1995) Neural pathways involved in mutual interactions between optic lobe circadian pacemakers in the cricket Gryllus bimaculatus. J Comp Physiol A 176:601–610. https://doi.org/10.1007/BF01021580
Závodská R, Sauman I, Sehnal F (2003a) The cycling and distribution of PER-like antigen in relation to neurons recognized by the antisera to PTTH and EH in Thermobia domestica. Insect Biochem Mol Biol 33:1227–1238. https://doi.org/10.1016/j.ibmb.2003.06.009
Závodská R, Šauman I, Sehnal F (2003b) Distribution of PER protein, pigment-dispersing hormone, prothoracicotropic hormone, and eclosion hormone in the cephalic nervous system of insects. J Biol Rhythm 18:106–122. https://doi.org/10.1177/0748730403251711
Závodská R, Sehadová H, Sauman I, Sehnal F (2005) Light-dependent PER-like proteins in the cephalic ganglia of an apterygote and a pterygote insect species. Histochem Cell Biol 123:407–418. https://doi.org/10.1007/s00418-004-0728-3
Zerr DM, Hall JC, Rosbash M, Siwicki KK (1990) Circadian fluctuations of period protein immunoreactivity in the CNS and the visual system of Drosophila. J Neurosci 10:2749–2762. https://doi.org/10.1523/JNEUROSCI.10-08-02749.1990
Zhang C, Daubnerova I, Jang YH, Kondo S, Žitňan D, Kim YJ (2021a) The neuropeptide allatostatin C from clock-associated DN1p neurons generates the circadian rhythm for oogenesis. Proc Natl Acad Sci U S A 118:e2016878118. https://doi.org/10.1073/pnas.2016878118
Zhang L, Chung BY, Lear BC, Kilman VL, Liu Y, Mahesh G et al (2010a) DN1(p) circadian neurons coordinate acute light and PDF inputs to produce robust daily behavior in Drosophila. Curr Biol 20:591–599. https://doi.org/10.1016/j.cub.2010.02.056
Zhang Y, Liu Y, Bilodeau-Wentworth D, Hardin PE, Emery P (2010b) Light and temperature control the contribution of specific DN1 neurons to Drosophila circadian behavior. Curr Biol 20:600–605. https://doi.org/10.1016/j.cub.2010.02.044
Zhang Y, Zhou Y, Zhang X, Wang L, Zhong Y (2021b) Clock neurons gate memory extinction in Drosophila. Curr Biol 31:1337–1343.e4. https://doi.org/10.1016/j.cub.2021.01.008
Zhu H, Sauman I, Yuan Q, Casselman A, Emery-Le M, Emery P et al (2008) Cryptochromes define a novel circadian clock mechanism in monarch butterflies that may underlie sun compass navigation. PLoS Biol 6:e4. https://doi.org/10.1371/journal.pbio.0060004
Acknowledgments
We thank Misako Yoshii for figure drawing and Charlotte Helfrich-Förster for comments on figures. This work was supported by JSPS KAKENHI (19H03265).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Yoshii, T., Fukuda, A. (2023). Neurocircuitry of Circadian Clocks. In: Numata, H., Tomioka, K. (eds) Insect Chronobiology. Entomology Monographs. Springer, Singapore. https://doi.org/10.1007/978-981-99-0726-7_5
Download citation
DOI: https://doi.org/10.1007/978-981-99-0726-7_5
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-0725-0
Online ISBN: 978-981-99-0726-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)