Abstract
Biomedical Image Processing is becoming an essential field in health care monitoring. Day by day, there is an increase in the number of patients due to pandemic and daily lifestyles. As the disease increases, it becomes burden to doctors to monitoring and diagnosing the diseases. The Biomedical Imaging helps in this aspect. Soft Tissue abnormalities can be easily identified with the help of imaging techniques. Use of contrast materials into the patient’s body traces the difficulties or problems within the tissue. MRI Imaging is one of the techniques that use safe contrast media. But there is a challenging task to differentiate the normal and abnormal parts of the body in the presence of contrast medium. This paper discusses few of the simple and more popular algorithms that can perform the segmentation in contrast and non contrast MRI images. The segmentation performance of each algorithm is compared in terms of performance metrics such as Sensitivity, Specificity, DICE coefficient and Jaccardian Index.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
Introduction
Biomedical Image Processing is the now becoming an emerging field in health monitoring, diagnosis and to treat different kinds of internal organ abnormalities or diseases for scientific study or treatment [1].
Medical Imaging [2] is an in vivo imaging process and it involves visualisation of internal organs within the body of humans or animals using non-invasive imaging modalities such as X-Ray, CT, MRI, PET Scans, etc. with the help of computer aids. Computer Aided Diagnosis (CAD) helps the medical expert to acquire better anatomical structures and to study the Region of Interest (ROI) within the tissue.
CAD tools involve the expert to automate the process [3], to get the results fast and accurately even for large number of cases without fatigue or data overload at minimal cost. They support remote accessing using information technologies through faster communication in case of emergency.
Imaging Modalities and Their Contrast Media
There are many imaging modalities [4] are used in biomedical engineering. They are Computed Tomography (CT), Magnetic Resonance Imaging (MRI), Ultrasound (US) Imaging, Digital Mammography, Positron Emission Tomography (PET) and Thermal Imaging and so on. Each and every modality has its own strengths and weaknesses but they are application specific. For example, X-Ray Imaging is mainly used to visualise hard tissues such as bones within the body whereas CT and MRI imaging modalities are used to observe soft tissues or both. These processes include Ionisation and electromagnetic interferences at microscopic level, respectively.
CT: Computed Tomography (CT) as the name indicates that the computer creates the images sequence by taking the inputs from X-Rays at all possible angles around the object in the form of slices called Tomograms. CTs produce the pictorial information of the object using ionising radiation. It can visualise brightly the high-density matters such as bones, calcium and high densities and darkly the low-density matters such as liquids fats, etc. The Hounsfield units are used to represent the different densities measured. For example, Water typically having 0 HU may be varying between −7 to +7 HU, bones have higher densities greater than 500 HU, soft tissues within the range 10–60 HU and fats with low density between −25 HU to −250 [5].
Modern CT scanners are advantageous that they can scan the abdomen and pelvis very quickly at a slice thickness of less than 1 mm within the span of 6–7 s per view. This is better suitable for the case of children without giving anaesthesia. Detailed information about any tissue is also obtained by other projections such as Sagittal and Coronal. CT scanners can also produce 3D scans [5].
CT scanners produce high-resolution images with minimal noise but are unable to detect inflammation defects.
MRI: MRI [6] is a non-ionising modality in radio imaging and it is extensively used for imaging soft tissues, organs, bone and other internal structures because it offers much greater contrast between the diverse soft tissues of the body. It is mostly used in radiology to diagnose or monitor treatment for abnormal conditions or staging the disease within the chest, abdomen and pelvis. MRIs produce very good contrast images of soft tissues compared to CTs. The human body naturally consists of hydrogen atoms within the body. MRI consists of a powerful rotating magnet, electric field gradients [7] (induction coils) and a digital computer to form images. The protons within the hydrogen atoms of water molecules are having magnetic properties such that all protons are aligned parallel to the main magnetic field. In MRI, a strong magnetic field is applied to line up the nuclear magnetisation of hydrogen atoms of the observing part of the body. An RF signal from the induction coils is applied subsequently in the form of RF pulses to alter the alignment of the protons and reversed to get relaxation energy from the protons. This relaxation energy is detected by MRI scanners to construct the image using K-space [8]. By changing the parameters of this RF pulse sequence, different contrasts may be generated between tissues based on the relaxation properties of the hydrogen atoms there in the tissues.
Depending on relaxation times and dynamic contrast variations [9], the intensities within the image are correlating with the tissue characteristics. It can produce morphological and functional information about the tissue without any ionising radiation.
PET: Positron Emission Tomography (PET) [10] is a functional imaging technique used to obtain the information about the changes in metabolic process, oxygen through the blood flow and tissues or organs’ working conditions. It is the combination of nuclear medicine and biochemical interactions with the body of observation. PET scans are mostly used for monitoring the functioning of brain disorders, heart problems and cancer cells within the body. PETs employ radioactive tracers that were swallowed as gas or injected into the body parts being examined. These tracers chemically react differently with the different parts of the body. These differential changes will be collected by the PET scanners as bright spots if they were infected. PET scanners can able to measure the oxygen [11] and sugar levels of the body and early stages of defects which are not possible by other scanners at cellular level. In PETs, gamma cameras are used to produce and to collect gamma rays as similar to X-rays. High chemical reaction of tracers will be shown as hot spots with greater intensity and less chemical reactions as cold spots with light intensities. Cancer cells [12] have high metabolic rate viewed as hot spots than non-cancer cells. Healthy tissues have more tracer absorption than the diseased one. Thus the level of heart problems was detected based the degree of interaction and colour differentiation. PET scanners are also used for detecting problems through the interaction of tracer with the glucose such as Alzheimer’s disease [13], depression, epilepsy, head and Parkinson’s disease in central nervous system. PETs also used with CT/MRI to get more information about disease.
PET scans are less accurate in case of small-sized tumours and at high blood sugar levels.
Microscopy [14] and PA imaging techniques [15] with contrasts are useful to detect defects from the defected organs due to deeper penetration.
According to our findings [16], ultrasonography examination could be used as a preliminary assessment for stratifying patients based on their risk of ADPKD progression. Because of the low accuracy and reproducibility of ultrasound, this estimate should only be used for patients with kidney volumes that are close to normal. A first magnetic resonance imaging (MRI) scan is recommended for dimensional increases in kidney size.
Contrast Materials: To improve medical imaging, various types of contrast media have been utilised. These contrast materials can be injected into veins or arteries, spinal discs or fluid spaces and other bodily cavities. Contrast materials come in a variety of forms:
-
1.
X-ray and computed tomography (CT) imaging exams use iodine-based and barium-sulfate chemicals. The most often used contrast substance is barium-sulfate, which is administered orally. It is also administered rectally and comes in a variety of forms. Iodine-based and barium-sulfate contrast materials block or limit the ability of X-rays to pass through a specific area of the body. As a result, blood vessels, organs and other bodily tissue containing iodine-based or barium compounds on X-Ray or CT pictures change appearance.
-
2.
Gadolinium is a key component of the most used magnetic resonance (MR) contrast material. When this chemical is present in the body, it changes the magnetic properties of adjacent water molecules, improving the quality of magnetic resonance imaging (MRI) images. Thus, internal soft organs, such as the heart, lungs, liver, adrenal glands, kidneys, pancreas, gall bladder, spleen, uterus and bladder are monitored.
-
3.
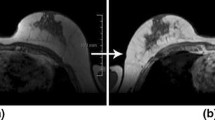
In imaging studies, saline (salt water) and gas (such as air) are also employed as contrast materials. Ultrasound imaging exams, particularly those of the heart, have used microbubbles and microspheres. Blood perfusion in organs is assessed using contrast-enhanced ultrasonography with microbubbles [17], thrombosis, such as in myocardial infarction, cardiac anomalies, liver and kidney tumours, inflammatory activity in inflammatory bowel illness and reaction to chemotherapy treatment (Fig. 9.1).
Fig. 9.1 Illustration example for non-contrast and Gd Contrast MRI images.
Proposed Methodology
In this paper, the proposed block diagram as shown in Fig. 9.2 is a new combination of denoising algorithms used to improve the image quality and to enhance the image contrast before segmenting the MRI Image. The image quality is improved by reducing the noise present within the image using Adaptive NLM Algorithm and contrast is enhanced using CLAHE algorithm. The Region of Interest (ROI) to be segmented in contrast-enhanced image is compared with the same in contrast image obtained from the subject injecting the contrast agent. The segmentation performance is compared in terms of its characteristics such as Sensitivity or Recall, Specificity, DICE and Jaccardian Coefficients.
Preprocessing of MRI Images
There are several image denoising techniques available, then Non-local filters are widely used in the area of MR imaging [18, 19].
Advantages of NLM Algorithms
-
The NLM noise reduction algorithm, in particular, was devised to minimise the loss of underlying image information while selectively removing only the noise.
-
After setting areas with the same sized mask positioned around the region of interest, this technique assesses the similarity of the intensity and edge information in a picture (ROI).
-
Furthermore, the higher the allocated weight employed during picture processing, the higher the degree of similarity.
In terms of PSNR, MSE and SSIM, experimental results shown in [20] on standard images indicate that the adaptive NLM algorithm outperforms the traditional NLM algorithm.
Contrast Limited Adaptive Histogram Equalisation (CLAHE)
A CLAHE algorithm [21] limit the amplification by clipping the histogram. It has two key parameters: Block Size (BS) and Clip Limit (CL). These two parameters improve the brightness and contrast of the image, respectively.
The CLAHE method to enhance the original image consists of the following steps:
Step 1: initially the input Image is dividing into non-overlapping contextual regions equal to M × N.
Step 2: Histogram is calculated for each contextual region.
Step 3: Calculate the Clip Limit (CL) for each region as given below
where \(N_{{{\text{avg}}}}\) is the average number of pixels, \(N_{{{\text{gray}}}}\) is the number of gray levels in the contextual region, \(N_{r} \left( x \right)\) and \(N_{r} \left( y \right)\) are the numbers of pixels in the X dimension and Y dimension of the contextual region. Then the actual CL is
where \(N_{{{\text{clip}}}}\) is the normalised CL in the range of [0, 1]. If the number of pixels is greater than \(N_{{{\text{CL}}}}\) , the pixels will be clipped. The total number of clipped pixels is defined as \(N_{{\sum {\text{clip}}}}\) then the average of the remain pixels to distribute to each gray level is
The histogram clipping rule is given by the following statements
where \(H_{{{\text{region}}}} \left( i \right)\) and \(H_{{{\text{region\_clip}}}} \left( i \right)\) are original histogram and clipped histogram of each region at i-th gray level.
Step 4: Redistribute the remaining pixels until the remaining pixels have been all distributed. The step of redistribution of pixels is given by
where \(N_{{{\text{remaining}}}}\) is the remaining number of clipped pixels. Step is positive integer at least 1.
Step 5: Then intensity values are distributed according to Rayleigh distribution to enhance the contrast and re-scaled using linear contrast stretching.
Step 6: To eliminate artefacts in the image, new gray level is calculated and assigned to the pixels in contextual region using bilinear interpolation.
Selection and Segmentation of ROI
The Region of Interest to be segmented within the MRI Image is selected manually and Chan-Vese Active Contours [22] was applied to get the object of the image. Manually the Mask will be drawn surrounding the ROI of the object for both CLAHE-enhanced and Contrast-enhanced images. Then efficiency of algorithm is estimated in terms of image characteristics.
Popular Segmentation Algorithms
Here we have taken the threshold-based and clustering algorithms which are useful for Medical Image segmentation applications. They are simple and popular algorithms, such as Otsu Algorithm [23], Active Contours without edges, Level sets [24], K-Means Segmentation [25] and Fuzzy C-Means Segmentation Algorithm [26]. They all are familiar and very simple to understand. Each algorithm has its own advantages and they are application specific.
Performance Metrics
This paper taken spatial overlap-based metrics [27] into account to evaluate the segmentation performance of each algorithm. These are derived from the cardinalities of confusion matrix such as namely the true positives (TP), the false positives (FP), the true negatives (TN) and the false negatives (FN). Then the derives metrics from these cardinalities are mostly used for segmentation evaluation are given by
This is also called True Positive Rate (TPR). Similarly, the True Negative Rate (TNR) is given by
Then
This is called Overlap Index mostly used for validating Medical volume Segmentations. For the generalised segmentation with multiple labels, Jaccardian Index is considered.
In this analysis, we have considered the above-mentioned four parameters to concise evaluation of segmentation.
Results and Discussions
The datasets were collected from publicly available TCIA database [28] browsing on the preferences as Imaging Modality: MR, Anatomical site: Kidney and Brain and Species: Homo sapiens excluding Phantoms. Totally there are 1043 subjects that were browsed. There are 82 categories of collections of kidney volumes in different projections and 961 categories of collections of Brain structures were obtained from different MRI machines such as GE MEDICAL SYSTEMS, SIEMENS, TOSHIBA_MEC and Philips Medical Systems with different subject IDs. We are mainly concentrate on Axial and coronal projections and selected 32 volumes with sizes 512 × 512 and 256 × 256, thicknesses of 4 mm, 5 mm,6 mm,7 mm, 8 mm and corresponding the spacing of 1.5 mm,2 mm,2.5 mm,3 mm,6 mm and 8 mm, respectively with and without Contrast media. From image volume sequence the slice with maximum tissue area was obtained from different series and then converted those into jpeg format. Each image used in this paper is mentioned with its ID, sequence number and slice number. The sequence/series number differentiates the projection series.
The kidney image shown in Fig. 9.3a is extracted for male subjects at an age of 41 years using GE MEDICAL SYSTEMS, COR KIDNEY1 PRE sequence with slice number 11 out of 20. It was obtained at a magnetic field strength of 1.5 T and Coronal 90 degrees flip angle projection with repetition time 160 s and echo time 1.344 s. The slice is with 256 × 256 size and the thickness of each slice is 5 mm with spacing 6 mm between them. This image is a non-contrasted image shown in Fig. 9.3i, it is preprocessed through Adaptive denoising algorithm to remove unwanted noise shown in Fig. 9.3ii, then contrast is enhanced and gray levels are distributed uniformly using CLAHE algorithm shown in Fig. 9.3iii. A Chan-Vese Active contour Algorithm is used to select the ROI manually and the proposed algorithms are employed for segmentation for both contrast and non contrast images.
Similarly, the kidney image shown in Fig. 9.4a is extracted for male subjects at an age of 45 years using SEIMENS MEDICAL SYSTEMS, COR KIDNEY1 PRE sequence with slice number 30 out of 34.
It was obtained at a magnetic field strength of 1.5 T and Axial 90° flip angle projection with repetition time 120 s and echo time 1.156 s. The slice is with 256 × 256 sizes and the thickness of each slice is 5 mm with spacing 6 mm between them.
Conclusions
The proposed methodology is a new combination of Adaptive NLM Algorithm to reduce the noise in MRI images and CLAHE Algorithm is used to enhance the image contrast before segmenting the MRI Image. The Region of Interest (ROI) to be segmented in contrast-enhanced image is compared with the same in contrast image obtained from the subject injecting the contrast agent. The segmentation performance is compared in terms of its characteristics such as Sensitivity or Recall, Specificity, DICE and Jaccardian Coefficients as shown in Table 9.1 and their graphical evaluation is shown in Figs. 9.5 and 9.6.
The execution time in Fig. 9.6a shows that Otsu, K-Means and Fuzzy C-Means Algorithms are faster for both Non Contrast and Contrast MRI Images compared to the remaining algorithms. The sensitivity is less for Fuzzy C-Means due to soft computing and it is high for Otsu algorithms as shown in Fig. 9.6b. The specificity is more for contrast images in case of Fuzzy C-Means and it is less for Non Contrast image in case of Level set Segmentation. Finally, the DICE Coefficient and Jaccardian Indexes are similar for both Non Contrast and Contrast MRI Images and they are high for segmentation using Otsu Algorithm.
References
Abdallah, Y.: Improvement of sonographic appearance using HATTOP methods. Int. J. Sci. Res. (IJSR) 4(2), 2425–2430 (2015). https://doi.org/10.14738/jbemi.55.5283
Abdallah, O.M.Y., Alqahtani, T.: Research in Medical Imaging Using Image Processing Techniques, Medical Imaging—Principles and Applications, Yongxia Zhou, Intech Open (2019). https://doi.org/10.5772/intechopen.84360
Sharma, N., Aggarwal, L.M.: Automated medical image segmentation techniques. J. Med. Phys. 35(1), 3–14 (2010). https://doi.org/10.4103/0971-6203.58777
Morris, P.: Biomedical Imaging, Woodhead Publishing, pp. xix–xxi (2014). ISBN 9780857091277
Hiorns, M.P.: Imaging of the urinary tract: the role of CT and MRI. Pediatr. Nephrol. 26(1), 59–68 (2011). https://doi.org/10.1007/s00467-010-1645-4
Grattan-Smith, J.D., Jones, R.A.: MR urography in children. Pediatr. Radiol. 36, 1229–1232 (2006). https://doi.org/10.1007/s00247-006-0222-2
Weishaupt, D., Koechli, V.D., Marincek, B.: How Does MRI Work?: An Introduction to the Physics and Function of Magnetic Resonance Imaging. Springer Science & Business Media (2013). ISBN 978-3-662-07805-1
Ljunggren, S.: A simple graphical representation of fourier-based imaging methods. J. Magn. Reson. 54(2), 338–343 (1983)
Yankeelov, T.E., Gore, J.C.: Dynamic contrast enhanced magnetic resonance imaging in oncology: theory, data acquisition, analysis, and examples. Curr. Med. Imaging Rev. 3(2), 91–107 (2009). https://doi.org/10.2174/157340507780619179
Bailey, D.L., Townsend, D.W., Valk, P.E., Maisy, M.N.: Positron Emission Tomography: Basic Sciences. Springer, Secaucus, NJ. 978-1-85233-798-8 (2005)
Azad, G.K., Siddique, M., Taylor, B., Green, A., O’Doherty, J., Gariani, J., et al.: 18F-Fluoride PET/CT SUV? J. Nucl. Med. 60(3), 322–327 (2019)
Galway, K., Black, A., Cantwell, M., Cardwell, C.R., Mills, M., Donnelly, M.: Psychosocial interventions to improve quality of life and emotional wellbeing for recently diagnosed cancer patients. Cochrane Database Syst. Rev. 11, CD007064 (2012). https://doi.org/10.1002/14651858.cd007064
Dementia: Quick Reference Guide (PDF). (UK) National Institute for Health and Clinical Excellence, London. November 2006. ISBN 978-1-84629-312-2
Thomas, G., et al.: Measuring the mechanical properties of living cells using atomic force microscopy. J. Visualized Experiments: JoVE 76, 50497 (2013). https://doi.org/10.3791/50497
Jimmy, L.S., Karpiouk, A.B.,, Wang, B., Emelianov, S.Y.: Photoacoustic imaging of clinical metal needles in tissue. J. Biomed. Opt. 15(2), 021309 (2010). https://doi.org/10.1117/1.3368686
Raghavendra, P., Pullaiah, T.: Advances in Cell and Molecular Diagnostics. Academic Press, pp. 85–111 (2018). ISBN 9780128136799. https://doi.org/10.1016/B978-0-12-813679-9.00004-X
Meloni, M.F., Smolock, A., Cantisani, V., et al.: Contrast enhanced ultrasound in the evaluation and percutaneous treatment of hepatic and renal tumors. Eur. J. Radiol. 84(9), 1666–1674 (2015)
Bhujle, H.V., Vadavadagi, B.H.: NLM based magnetic resonance image denoising—a review. Biomed. Sign. Process. Control 47, 252–261 (2019)
Buades, A., Coll, B., Morel, J.-M.: A non-local algorithm for image denoising. IEEE Comp. Soc. Conf. Comp. Vis. Pattrn. Recog. 2, 60–65 (2005)
Verma, R., Pandey, R.: Non local means algorithm with adaptive isotropic search window size for image denoising. Ann. IEEE India Conf. (INDICON) 2015, 1–5 (2015). https://doi.org/10.1109/INDICON.2015.7443193
Ma, J., Fan, X., Yang, S.X., Zhang, X., Zhu, X.: Contrast limited adaptive histogram equalization-based fusion in YIQ and HSI color spaces for underwater image enhancement. Int. J. Pattern Recognit. Artif. Intell. 32, 1–26 (2018)
Chan, T.F., Vese, L.A.: Active contours without edges. IEEE Trans. Image Process. 10(1), 266–277 (2001). https://doi.org/10.1109/83.902291
Wang, H., Dong, Y.: An improved image segmentation algorithm based on Otsu method. In: International Symposium on Photoelectronic Detection and Imaging 2007: Related Technologies and Applications, vol. 6625 (2008)
Li, C., Huang, R., Ding, Z., Gatenby, J.C., Metaxas, D.N., Gore, J.C.: A level set method for image segmentation in the presence of intensity inhomogeneities with application to MRI. IEEE Trans. Image Process. 20(7), 2007–2016 (2011). https://doi.org/10.1109/TIP.2011.2146190
Venkatachalam, K., Reddy, V.P., Amudhan, M., Raguraman, A., Mohan, E.: An implementation of K-means clustering for efficient image segmentation. In: 2021 10th IEEE international conference on Communication Systems and Network Technologies (CSNT), pp. 224–229 (2021). https://doi.org/10.1109/CSNT51715.2021.9509680
Rahman, T., Islam, M.S.: Image segmentation based on fuzzy C means clustering algorithm and morphological reconstruction. In: 2021 International Conference on Information and Communication Technology for Sustainable Development (ICICT4SD), pp. 259–263 (2021). https://doi.org/10.1109/ICICT4SD50815.2021.9396873
Taha, A.A., Hanbury, A.: Metrics for evaluating 3D medical image segmentation: analysis, selection, and tool. BMC Med. Imaging 15, 29 (2015). https://doi.org/10.1186/s12880-015-0068-x
TCIA Database. https://nbia.cancerimagingarchive.net/nbia-search/
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Prabhu Das, S., Jagadesh, B.N., Prabhakara Rao, B. (2022). Performance Evaluation of Segmentation Algorithms in Non Contrast and Contrast MRI Images for Region of Interest. In: Satyanarayana, C., Gao, XZ., Ting, CY., Muppalaneni, N.B. (eds) Proceedings of the International Conference on Computer Vision, High Performance Computing, Smart Devices and Networks. Advanced Technologies and Societal Change. Springer, Singapore. https://doi.org/10.1007/978-981-19-4044-6_10
Download citation
DOI: https://doi.org/10.1007/978-981-19-4044-6_10
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-4043-9
Online ISBN: 978-981-19-4044-6
eBook Packages: Computer ScienceComputer Science (R0)