Abstract
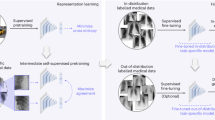
Deep Learning provides exciting solutions to problems in medical image analysis and is regarded as a key method for future applications. However, only a few annotated medical image datasets exist compared to numerous natural images. A solution to this problem is transfer learning using ImageNet. However, because the domain of ImageNet is different from that of medical images, the results of transfer learning are not always good. Therefore, we propose a model to investigate transfer learning by self-supervised learning using medical images. It is widely known that the results of Computerized Tomography (CT) scan are 3D volume images. There are lots of slices in CT or Magnetic Resonance Imaging scan images. So why not make these slices to a class? It is imperative to formulate this intuition as a self-supervised feature learning at the case-level. The results of our experiment demonstrated that, under self-supervised feature learning settings, our method surpasses the transfer learning method that uses ImageNet for classification. By experimenting with unannotated datasets, our method is remarkable for consistently improving test performance with a few annotated data. By fine-tuning the learned features, we obtained competitive results for self-supervised learning and classification tasks.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Henley, S.J., et al.: Annual report to the nation on the status of cancer, part i: national cancer statistics. Cancer 126(10), 2225–2249 (2020)
Roy, S., et al.: Three-dimensional spatiotemporal features for fast content-based retrieval of focal liver lesions. IEEE Transactions on Biomedical Engineering 61(11), 2768–2778 (2014)
Diamant, I., et al.: Improved patch-based automated liver lesion classification by separate analysis of the interior and boundary regions. IEEE J. Biomed. Health Inform. 20(6), 1585–1594 (2015)
Xu, Y., et al.: Texture-specific bag of visual words model and spatial cone matching-based method for the retrieval of focal liver lesions using multiphase contrast-enhanced CT images. Int. J. Comput. Assist. Radiol. Surg. 13(1) (2018)
Wang, J., et al.: Tensor-based sparse representations of multi-phase medical images for classification of focal liver lesions. Pattern Recognit. Lett. 130, 207–215 (2020)
Maayan, F.-A., et al.: Modeling the intra-class variability for liver lesion detection using a multi-class patch-based CNN. In: Patch-Based Techniques in Medical Imaging, pp. 129–137. Springer International Publishing, Cham (2017)
Yasaka, K., et al.: Deep learning with convolutional neural network for differentiation of liver masses at dynamic contrast-enhanced CT: a preliminary study. Radiology 286(3), 887–896 (2018)
Liang, D., et al.: Combining convolutional and recurrent neural networks for classification of focal liver lesions in multi-phase CT images. In: International Conference on Medical Image Computing and Computer-Assisted Intervention, pp. 666–675. Springer (2018)
Wang, W., et al.: Classification of focal liver lesions using deep learning with fine-tuning. In: Proceedings of the 2018 International Conference on Digital Medicine and Image Processing, pp. 56–60 (2018)
Wang, W., et al.: Deep fusion models of multi-phase CT and selected clinical data for preoperative prediction of early recurrence in hepatocellular carcinoma. IEEE Access 8, 139212–139220 (2020)
Russakovsky, O., et al.: ImageNet large scale visual recognition challenge. Int. J. Comput. Vis. (IJCV) 115(3), 211–252 (2015)
Wu, Z., et al.: Unsupervised feature learning via non-parametric instance discrimination. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition(CVPR), pp. 3733–3742 (2018)
Kaiming, H., et al.: Momentum contrast for unsupervised visual representation learning. In: Conference on Computer Vision and Pattern Recognition (CVPR) (2020)
Doersch, C., et al.: Unsupervised visual representation learning by context prediction. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 1422–1430 (2015)
Gidaris, S., et al.: Unsupervised representation learning by predicting image rotations (2018). arXiv:1803.07728
Noroozi, M., et al.: Unsupervised learning of visual representations by solving jigsaw puzzles. In: European Conference on Computer Vision, pp. 69–84. Springer (2016)
Ting, C., et al.: A simple framework for contrastive learning of visual representations (2020). arXiv:2002.05709
Kaiming, H., et al.: Deep residual learning for image recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2016)
Acknowledgements
We would like to thank Sir Run Run Shaw Hospital for providing medical data and helpful advice on this research. This work is supported in part by the Grantin Aid for Scientific Research from the Japanese Ministry for Education, Science, Culture and Sports (MEXT) under the Grant No. 20KK0234, No. 20K21821, No.18H03267, in part by Zhejiang Lab Program under the Grant No. 2020ND8AD01.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Dong, H. et al. (2021). Case Discrimination: Self-supervised Feature Learning for the Classification of Focal Liver Lesions. In: Chen, YW., Tanaka, S., Howlett, R.J., Jain, L.C. (eds) Innovation in Medicine and Healthcare. Smart Innovation, Systems and Technologies, vol 242. Springer, Singapore. https://doi.org/10.1007/978-981-16-3013-2_20
Download citation
DOI: https://doi.org/10.1007/978-981-16-3013-2_20
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-3012-5
Online ISBN: 978-981-16-3013-2
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)