Abstract
SARS-CoV-2 is a single-stranded RNA virus of ~30 kb genome size which belongs to genus Coronavirus and family Coronaviridae. SARS-CoV-2 has recently emerged and has been declared as a pandemic by the World Health Organization. Genomic characterization of SARS-CoV-2 has shown that it is of zoonotic origin. The structure of SARS-CoV-2 is found to be similar to SARS-CoV with virion size ranging from 70 to 90 nm. Spike, membrane, and envelope surface viral proteins of coronavirus are embedded in host membrane-derived lipid bilayer encapsulating the helical nucleocapsid comprising viral RNA. The genome comprises of 6–11 open reading frames (ORFs) with 5′ and 3′ flanking untranslated regions (UTRs). Sequence variation among SARS-CoV-2 and SARS-CoV revealed no significant difference in ORFs and nsps. The nsps includes two viral cysteine proteases including papain-like protease (nsp3), chymotrypsin-like, 3C-like, or main protease (nsp5), RNA-dependent RNA polymerase (nsp12), helicase (nsp13), and others likely to be involved in the transcription and replication of SARS-CoV-2. The structure of spike glycoprotein structure of SARS-CoV-2 resembles that of the spike protein of SARS-CoV with an root-mean-square deviation (RMSD) of 3.8 Å. Like SARS-CoV, SARS-CoV-2 uses the ACE2 receptor for internalization and TMPRSS2 serine proteases for S protein priming. Histopathological investigation of tissues from SARS-CoV-2 infected patients showed virus-induced cytopathic effect with signs of acute respiratory distress syndrome in lung cells. This chapter discusses about the morphology, genome organization, replication, and pathogenesis of SARS-CoV-2 that may help us understand the disease that may leads to identification of effective antiviral drugs and vaccines.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
3.1 Introduction
Coronaviruses (CoVs) are enveloped single-stranded positive sense RNA viruses that belong to the family Coronaviridae. On the basis of genomic organization and phylogenetic relationship, coronaviruses have been classified into the subfamily Coronavirinae that consists of four genera Alphacoronavirus (αCoV), Betacoronavirus (βCoV), Gammacoronavirus (γCoV), and Deltacoronavirus (δCoV) (Cui et al. 2019). Evolutionary trend analysis of coronaviruses has revealed that αCoV and βCoV originated from bats and rodents, while γCoV and δCoV were found to have originated from avian species (Ge et al. 2017). The ability of CoVs to cross the species barrier has resulted in some of the pathogenic CoVs. HKU1, NL63, OC43, and 229E CoVs are associated with mild symptoms in humans, whereas severe acute respiratory syndrome CoV (SARS-CoV) and Middle East respiratory syndrome CoV (MERS-CoV) are known to cause severe disease (Fehr and Perlman 2015). In 2002–2003, SARS-CoV emerged in China with 8000 clinical cases and 800 deaths. Since 2012, MERS-CoV has caused persistent epidemics in the Arabian Peninsula. Both the viruses have been found to originate from bats and then transmitted into intermediate mammalian host civets in the case of SARS-CoV and camels in the case of MERS-CoV and eventually infected humans (Song et al. 2019).
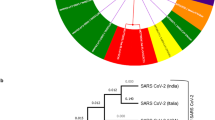
SARS-CoV-2 has been declared as a pandemic, with 1,844,683 confirmed cases and 117,021 deaths globally by 14th April 2020 (World Health Organization 2020). To characterize the novel coronavirus, bronchoalveolar lavage fluid and throat swabs were collected from nine patients who had visited the Wuhan seafood market during the initial outbreak. Special pathogen-free human airway epithelial (HAE) cells were used for virus isolation. The collected samples were inoculated into the HAE cells through the apical surfaces. HAE cells were monitored for cytopathic effects and supernatant was collected to perform RT-PCR assays. Apical samples were collected for next generation sequencing after three passages. The whole-genome sequences of SARS-CoV-2 were generated by a combination of Sanger, Illumina, and Oxford nanopore sequencing (Lu et al. 2020). Phylogenetic analysis has revealed that bats might be at the source of SARS-CoV-2 (Andersen et al. 2020). Additionally, some studies have suggested that the origin of SARS-CoV-2 is associated with pangolins (Li et al. 2020; Shereen et al. 2020). To decipher the mechanism of replication and development of effective preventive and therapeutic strategies, understanding the structure of SARS-CoV-2, genome organization, and replication is crucial. Therefore this chapter focuses on the morphology and structure, genomic organization, and replication cycle of SARS-CoV-2.
3.2 Morphology of SARS-CoV-2
SARS-CoV-2 isolated from nasopharyngeal and oropharyngeal samples were inoculated on the vero cells. In order to identify SARS-CoV-2, inoculated cells were prefixed using 2% paraformaldehyde and 2.5% glutaraldehyde, and transmission electron microscopy was performed. The structure of SARS-CoV-2 observed by examining infected cells after 3 days post infection. Electron microscopy revealed the coronavirus-specific morphology of SARS-CoV-2 with virus particle sizes ranging from 70 to 90 nm observed under a wide variety of intracellular organelles, most specifically in vesicles (Park et al. 2020). Due to high sequence similarity, the structure of SARS-CoV-2 is speculated to be the same as SARS-CoV (Kumar et al. 2020). The surface viral protein spike, membrane, and envelope of coronavirus are embedded in host membrane-derived lipid bilayer encapsulating the helical nucleocapsid comprising viral RNA (Fig. 3.1) (Finlay et al. 2004). The structure of spike (Yan et al. 2020) and protease of SARS-CoV-2 (Zhang et al. 2020) has been resolved, which provides an opportunity to develop a newer class of drugs for treatment of COVID-19.
Structure of SARS-CoV-2. SARS-CoV-2 has surface viral proteins, namely, spike glycoprotein (S), which mediates interaction with cell surface receptor ACE2. The viral membrane glycoprotein (M) and envelope (E) of SARS-CoV-2 are embedded in host membrane-derived lipid bilayer encapsulating the helical nucleocapsid comprising viral RNA
3.3 Genome Organization of SARS-CoV-2
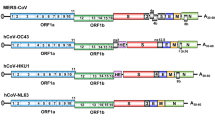
The size of coronavirus genome is in the range of 26 to 32 kb and comprise 6–11 open reading frames (ORFs) encoding 9680 amino acid polyproteins (Guo et al. 2020). The first ORF comprises approximately 67% of the genome that encodes 16 nonstructural proteins (nsps), whereas the remaining ORFs encode for accessory and structural proteins. The genome of SARS-CoV-2 lacks the hemagglutinin-esterase gene. However, it comprises two flanking untranslated regions (UTRs) at 5′ end of 265 and 3′ end of 358 nucleotides. Sequence variation among SARS-CoV-2 and SARS-CoV revealed no significant difference in ORFs and nsps. The nsps includes two viral cysteine proteases including papain-like protease (nsp3), chymotrypsin-like, 3C-like, or main protease (nsp5), RNA-dependent RNA polymerase (nsp12), helicase (nsp13), and others likely to be involved in the transcription and replication of SARS-CoV-2 (Chan et al. 2020). In addition to nsps, four major structural proteins are spike surface glycoprotein (S), membrane, nucleocapsid protein (N), envelope (E) and accessory proteins encoded by ORFs. N-terminal glycosylated ectodomain is present at the N-terminal end of M protein that comprises of three transmembrane domains (TM) and a long C-terminal CT domain (Fig. 3.2).
Genomic organization of SARS-CoV-2. The size of coronavirus genome ranges from 26 to 32 kb and comprises 6–11 open reading frames (ORFs) encoding 9680 amino acid polyprotein. The first ORF comprises of approximately 67% of the genome that encodes 16 nonstructural proteins (nsps), whereas the remaining ORFs encode for accessory and structural proteins. The nsps includes two viral cysteine proteases, including papain-like protease (nsp3), chymotrypsin-like, 3C-like, or main protease (nsp5), RNA-dependent RNA polymerase (nsp12), helicase (nsp13), and others likely to be involved in the transcription and replication of SARS-CoV-2. In addition to nsps, the genome encodes for four major structural proteins including spike surface glycoprotein (S), membrane, nucleocapsid protein (N), envelope (E) and accessory proteins like ORFs
The M and E proteins are required for virus morphogenesis, assembly, and budding, whereas S glycoprotein is a fusion viral protein comprising two subunits S1 and S2. The S1 subunit, which shares 70% sequence identity with bat SARS-like CoVs and human SARS-CoV, comprises signal peptide, N-terminal domain (NTD), and receptor-binding domain (RBD) (Walls et al. 2020). Most of the differences were found in the external subdomain that is primarily responsible for interaction of spike with the ACE2 receptor. The ectodomain of spike protein (1–1208 amino acid residues) was cloned, expressed and crystallize to solve the spike glycoprotein structure of SARS-CoV-2. The structure of spike glycoprotein structure of SARS-CoV-2 resembles the spike protein of SARS-CoV with an RMSD of 3.8 Å. The study also reveals that the receptor-binding region (RBD) exhibited the highest structural divergence (Wrapp et al. 2020). The S2 subunit that shares 99% sequence identity with bat SARS-like CoVs and human SARS-CoV comprises two heptad repeat regions known as HR-N and HR-C, which form the coiled-coil structures surrounded by the protein ectodomain. The S protein has been found to exhibit a furin cleavage site (PRRARS’V) at the interface between S1 and S2 subunits that is processed during the biogenesis (Coutard et al. 2020).
3.4 Entry and Replication of SARS-CoV-2 in Host Cells
Entry of coronaviruses into host target cells depends on the binding of spike glycoprotein to the cellular receptor and priming of S protein by host cell proteases. Like SARS-CoV, SARS-CoV-2 uses the ACE2 receptor for internalization and TMPRSS2 serine proteases for S protein priming (Hoffmann et al. 2020). Similar to SARS-CoV, the extrapulmonary spread of SARS-CoV-2 may be seen due to the widespread tissue expression of the ACE2 receptor. In addition, studies revealed that the spike protein of SARS-CoV-2 exhibits 10–20 times higher affinity as compared to that of SARS-CoV (Wrapp et al. 2020). Binding of spike protein to the ACE2 receptor results in conformational changes in spike protein that leads to the fusion of viral envelop protein with host cell membrane following entry via endosomal pathway (Coutard et al. 2020; Matsuyama and Taguchi 2009). This event is followed by the release of viral RNA into the host cytoplasm that undergoes translation and generates replicase polyproteins pp1a and pp1b that further cleaved by virus encoded proteinases into small proteins. The replication of coronavirus involves ribosomal frame shifting during the translation process and generates both genomic and multiple copies of subgenomic RNA species by discontinuous transcription that encodes for relevant viral proteins. Assembly of virion takes place via interaction of viral RNA and protein at endoplasmic reticulum (ER) and Golgi complex. These virions are subsequently released out of the cells via vesicles (Fig. 3.3) (Hoffmann et al. 2020).
Entry and replication of SARS-CoV-2 in host cells. Entry of SARS-CoV-2 into host target cells depends on the binding of spike glycoprotein to the cellular receptor ACE2 for internalization. Internalization results in uncoating of viral RNA into cytoplasm that undergoes translation and generates replicase polyproteins pp1a and pp1b, which is further cleaved by virus-encoded proteinases into small proteins. The replication of SARS-CoV-2 involves ribosomal frame shifting during the translation process and generates both genomic and multiple copies of subgenomic RNA species by discontinuous transcription required for relevant viral proteins. Assembly of virion takes place via interaction of viral RNA and protein at endoplasmic reticulum (ER) and Golgi complex. These virions are subsequently released out of the cells via vesicles via exocytosis
3.5 Pathogenesis of SARS-CoV-2
The pathological findings of SARS-CoV-2 infected patients highly resemble that of SARS-CoV and MERS-CoV infected patients. Flow cytometric analysis of peripheral blood samples showed significant reduction of CD4 and CD8 T cell counts, and their status was found to be hyperactivated as higher proportion of dual positive (HLA-DR and CD38) was seen. Rapid progression of pneumonia was seen in chest X-ray images with some differences between the right and left lung. Histopathological investigation of lung, liver, and heart tissue was performed. Lung biopsy showed cellular fibromyxoid exudates with bilateral diffuse alveolar damage. The right lung showed prominent desquamation of pneumocytes and formation of hyaline membrane, indicating signs of acute respiratory distress syndrome (ARDS), whereas the left lung showed pulmonary edema with formation of hyaline membrane (Xu et al. 2020). In addition, both lungs were found to exhibit interstitial mononuclear patchy inflammatory infiltrates dominated specifically by lymphocytes (Tian et al. 2020). The intra-alveolar spaces were characterized by multinucleated syncytial cells with atypical enlarged pneumocytes showing virus-induced cytopathic effect. Liver biopsy of patients infected with SARS-CoV-2 showed moderate microvesicular steatosis and mild portal and lobular activity, suggesting that injury might have been caused by the virus or drug induced. A few interstitial mononuclear inflammatory infiltrates were observed in the heart tissue. These pathological changes may provide new insights into the pathogenesis of pneumonia induced by SARS-CoV-2 that may help clinicians to effectively deal with COVID-19 patients.
3.6 Conclusions
Phylogenetic analysis revealed that SARS-CoV-2 might have originated from bats or pangolins. Structural investigations of virus-infected cells reveal the coronavirus-specific morphology of SARS-CoV-2 and the size of the virus (70–90 nm). The size of SARS-CoV-2 genome ranges from 26 to 32 kb and comprises 6–11 ORFs which lacks hemagglutinin-esterase gene. However, it comprises of 5′ and 3′ flanking untranslated regions (UTRs). The spike glycoprotein structure of SARS-CoV-2 resembles the spike protein of SARS-CoV with an RMSD of 3.8 Å. Like SARS-CoV, SARS-CoV-2 uses the ACE2 receptor for internalization and TMPRSS2 serine proteases for S protein priming. Histopathological investigation of tissues from SARS-CoV-2 infected patients showed virus-induced cytopathic effect with signs of acute respiratory distress syndrome in lung cells.
3.7 Future Perspectives
SARS-CoV-2 has recently emerged and has been declared as a pandemic by the World Health Organization. Based on the genomic sequences submitted to NCBI database, the scientific community has analyzed the samples and suggested preventive and therapeutic strategies. Therefore, investigation of genomic diversity in the collected specimens from around the globe needs to be conducted in order to design common, effective therapies and vaccines. In addition, genomic characterization helps us accurately identify the origin and evolution of the virus. Deciphering the mechanism of SARS-CoV-2 replication in various cell-based models may help us understand the pathogenesis and identify specific targets to develop effective antiviral drugs.
References
Andersen KG, Rambaut A, Lipkin WI et al (2020) The proximal origin of SARS-CoV-2. Nat Med 26(4):450–452. https://doi.org/10.1038/s41591-020-0820-9
Chan JF, Kok KH, Zhu Z, Chu H, To KK, Yuan S, Yuen KY (2020) Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microbes Infect 9(1):221–236. https://doi.org/10.1080/22221751.2020.1719902. eCollection 2020
Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E (2020) The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir Res 176:104742. https://doi.org/10.1016/j.antiviral.2020.104742
Cui J, Li F, Shi ZL (2019) Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol 17(3):181–192. https://doi.org/10.1038/s41579-018-0118-9
Fehr AR, Perlman S (2015) Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol 1282:1–23. https://doi.org/10.1007/978-1-4939-2438-7_1
Finlay BB, See RH, Brunham RC (2004) Rapid response research to emerging infectious diseases: lessons from SARS. Nat Rev Microbiol 2(7):602–607
Ge XY, Yang WH, Zhou JH, Li B, Zhang W, Shi ZL, Zhang YZ (2017) Detection of alpha- and betacoronaviruses in rodents from Yunnan, China. Virol J 14(1):98. https://doi.org/10.1186/s12985-017-0766-9
Guo YR, Cao QD, Hong ZS, Tan YY, Chen SD, Jin HJ, Tan KS, Wang DY, Yan Y (2020) The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak—an update on the status. Mil Med Res 7(1):11. https://doi.org/10.1186/s40779-020-00240-0
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S (2020) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181:1–10. https://doi.org/10.1016/j.cell.2020.02.052. [Epub ahead of print]
Kumar S, Maurya VK, Prasad AK et al (2020) Structural, glycosylation and antigenic variation between 2019 novel coronavirus (2019-nCoV) and SARS coronavirus (SARS-CoV). VirusDis 31(1):13–21. https://doi.org/10.1007/s13337-020-00571-5
Li X, Zai J, Zhao Q, Nie Q, Li Y, Foley BT, Chaillon A (2020) Evolutionary history, potential intermediate animal host, and cross-species analyses of SARS-CoV-2. J Med Virol 2020:1–10. https://doi.org/10.1002/jmv.25731
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395(10224):565–574. https://doi.org/10.1016/S0140-6736(20)30251-8
Matsuyama S, Taguchi F (2009) Two-step conformational changes in a coronavirus envelope glycoprotein mediated by receptor binding and proteolysis. J Virol 83(21):11133–11141. https://doi.org/10.1128/JVI.00959-09
Park WB, Kwon NJ, Choi SJ, Kang CK, Choe PG, Kim JY, Yun J, Lee GW, Seong MW, Kim NJ, Seo JS, Oh MD (2020) Virus isolation from the first patient with SARS-CoV-2 in Korea. J Korean Med Sci 35(7):e84. https://doi.org/10.3346/jkms.2020.35.e84
Shereen MA, Khan S, Kazmi A, Bashir N, Siddique R (2020) COVID-19 infection: origin, transmission, and characteristics of human coronaviruses. J Adv Res 24:91–98. https://doi.org/10.1016/j.jare.2020.03.005
Song Z, Xu Y, Bao L, Zhang L, Yu P, Qu Y, Zhu H, Zhao W, Han Y, Qin C (2019) From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses. 11(1). pii: E59. https://doi.org/10.3390/v11010059
Tian S, Hu W, Niu L, Liu H, Xu H, Xiao SY (2020) Pulmonary pathology of early-phase 2019 novel coronavirus (COVID-19) pneumonia in two patients with lung cancer. J Thorac Oncol. pii: S1556-0864(20)30132-5. https://doi.org/10.1016/j.jtho.2020.02.010. [Epub ahead of print]
Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D (2020) Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 180:1–12. https://doi.org/10.1016/j.cell.2020.02.058
World Health Organization (2020) Coronavirus disease 2019 (COVID-19) situation report—85. World Health Organization. https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200414-sitrep-85-covid-19.pdf?sfvrsn=7b8629bb_4. Accessed 14 Apr 2020
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS (2020) Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367(6483):1260–1263. https://doi.org/10.1126/science.abb2507
Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C, Liu S, Zhao P, Liu H, Zhu L, Tai Y, Bai C, Gao T, Song J, Xia P, Dong J, Zhao J, Wang FS (2020) Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med 8(4):420–422. https://doi.org/10.1016/S2213-2600(20)30076-X. [Epub ahead of print]
Yan R, Zhang Y, Li Y, Xia L, Guo Y, Zhou Q (2020) Structural basis for the recognition of the SARS-CoV-2 by full-length human ACE2. Science 367(6485):1444–1448. https://doi.org/10.1126/science.abb2762
Zhang L, Lin D, Sun X, Curth U, Drosten C, Sauerhering L, Becker S, Rox K, Hilgenfeld R (2020) Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science. pii: eabb3405. https://doi.org/10.1126/science.abb3405
Acknowledgments
The authors are grateful to the Vice Chancellor, King George’s Medical University (KGMU), Lucknow, India, for the encouragement of this work. The authors have no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 The Editor(s) (if applicable) and The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Kumar, S., Nyodu, R., Maurya, V.K., Saxena, S.K. (2020). Morphology, Genome Organization, Replication, and Pathogenesis of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). In: Saxena, S. (eds) Coronavirus Disease 2019 (COVID-19). Medical Virology: From Pathogenesis to Disease Control. Springer, Singapore. https://doi.org/10.1007/978-981-15-4814-7_3
Download citation
DOI: https://doi.org/10.1007/978-981-15-4814-7_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-4813-0
Online ISBN: 978-981-15-4814-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)