Abstract
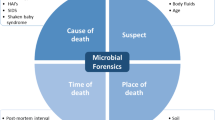
An incredible life is present beyond the scope of the naked eye. The one present underneath the lens of a microscope is the life of “microorganisms.” Microorganisms are present everywhere in the environment. Human body serves as a host to a wide variety of microbes including bacteria and viruses. Though most of the microorganisms are harmless to humans, they can cause diseases in humans as well as in animals and plants. Microorganisms play a vital role in food and dairy industries; production of enzymes, amino acids, vitamins, antibiotics, etc.; genetic engineering; biotechnology; and so on. With advancement in science and technology, the antisocial elements are manufacturing bioweapons by using the microorganisms and the toxins produced by them. Microbial forensics refers to deployment of scientific principles to analyze microbial evidence. Technologies like massive parallel sequencing (MPS) and next-generation sequencing (NSG) help scientists to understand the role of microorganisms in the origin of biocrimes, cause and time of death, sexual assault, homicide, agricultural contamination, and medical malpractice. The microbial forensic investigation results in linking the causative agent with a specific group or an individual by following special procedures.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Forensic microbiology
- Microbial biodiversity
- Biocrimes
- Bioterrorism
- Postmortem interval
- Sexual assault
- Forensic investigation
1 Introduction
The term forensic science broadly refers to the science pertaining to the administration of law and justice. Forensic science basically incorporates and utilizes the techniques and principles of basic sciences for the purpose of administration of law. Forensic science works in conjunction with the experts from many subjects like physics, chemistry, biology, zoology, botany, microbiology, etc. Thus, it is an interdisciplinary or a multidisciplinary field, where, depending upon the need of the case, scientists of different expertise assist the court of law. Almost every forensic science laboratory in the world has well-established disciplines like forensic ballistics, physics, chemistry, toxicology, biology, entomology, etc., but microbial forensics is comparatively new and a young field for the forensic laboratories and scientists as well.
Microorganisms are ubiquitous organisms and are attentive observers to our world. Microorganisms are present everywhere, including the air we breathe and the food we eat. Human population is host to a wide variety of microbes, including bacteria and viruses that are present on our skin, in our alimentary and respiratory tract (mouth and nose), conjunctiva, and so on. Specifically, the human skin is a habitat of countless numbers of eukaryotic microbes including fungi and protists and represents a diverse and dynamic ecosystem which has evolved to colonize almost every available habitat on the human body as well as within the integumentary system (Roth and James 1988; Grice et al. 2009). The presence of a large number of microorganisms on human skin enables the analysis of forensically interesting body sites (Costello et al. 2009; Sender et al. 2016).
According to reports, microbes are present in the ratio of 4:3 to the cells of human body and weigh about 0.2 kg of a 70-kg human body (Rana and Manhas 2018). The number of microbial fauna varies from <10 to >107 microorganisms per square cm (Bojar and Holland 2002). In addition to the microbial fauna found in the environment, forensic scientists have been exploring the microbes present in and on the human body. Apart from being present in high numbers, the microbial communities associated with humans are highly diverse at various body sites (Costello et al. 2009). This is because of the different thriving conditions present on the human body. For instance, the skin is dry, moist, or sebaceous at different sites, thereby causing different kinds of microbes to thrive on these different microenvironments (Grice et al. 2009). Hence, the interaction of microbes with humans is inevitable.
Though most of the microorganisms are harmless to us, however, their activities can produce diseases in humans, as well as in animals and plants. They need a host mechanism to survive and proliferate. Diseases are caused due to this adaptive strategy of microbes for replication and survival within the host (Cummings and Relman 2002). However, diseases may also be induced by a forced interaction between microbes and a host or by altering or manipulating the genome of the microbe. This interaction may be induced as a result of the acts of bioterrorism (Cummings and Relman 2002) which involves malafide human intention.
In the forensic scenario as well, microorganisms play a vital role and have a variety of forensic applications, the foremost being the use of microbes as biological weapons in biocrimes or bioterrorism. However, due to advancements in recent technology, specifically the massive parallel sequencing (MPS) and the next-generation sequencing (NSG), the scope of microbial forensics has been expanded, and microbial analysis can be used to understand the role of microorganisms in the process of decomposition, thereby aiding in estimation of postmortem interval. Additionally, microbes are present in the soil as forensic biomarkers; helps in personal identification from body fluids like saliva; assist in understanding the epidemiology of a disease, thereby aiding in linking a victim and a suspect through microbial diseases (Oliveira and Amorim 2018).
2 Microbes as Biological Agents in Biocrimes
Biocrime or bioterrorism refers to the threat raised due to use of microorganisms and their toxins and related products to commit acts of crime in order to terrorize people. Such acts of crime can be harmful to humans as well as animals and plants and can lead to the outbreak and spread of various microbial infections and diseases which may ultimately lead to an epidemic.
The Centers for Disease Control and Prevention (CDC) defines biocrimes or bioterrorism as the crimes that involve the deliberate usage of living organisms or the toxins produced by them to harm animals, plants, humans, or environment so as to disrupt the social stability and cause panic in the society (Jansen et al. 2014). The National Center for Biotechnology Information (NCBI) defines a biocrime as “an assault crime, except, instead of a gun or knife, the weapon used is a pathogen or a toxin” (Schutzer et al. 2005). Although commonly termed “bioterrorism,” the purposes of such attacks may not necessarily be intended and confined to terrorize the established government structures but can also be influenced by religious, political, or ecological beliefs (Carus 2001).
The traditional terrorists use violence as a means to an end, while bioterrorists view biological agents as specialized tools to cause mass destruction by spreading diseases.
On the basis of risk capability of the microbes, the public health agency like CDC of the United States (2002) classified bioweapons into three categories: categories A, B, and C. Category A includes the agents responsible for high mortality, easy dissemination, and a high potential to cause major health concerns, while categories B and C are moderate and easy to disperse, respectively. Category A includes organisms like Francisella tularensis, Bacillus anthracis, Yersinia pestis, and hemorrhagic Ebola virus and botulinum neurotoxins, while category B includes Brucella species, abrin, ricin, Clostridium perfringens E. toxin, and Burkholderia mallei. Hantavirus and Nipah viruses are examples of microbes in category C (Smart 1997; Grundmann 2014; Jansen et al. 2014; Kaur et al. 2014; Janik et al. 2019).
The concept of biocrime is not new. It can be traced well throughout history but was not scientifically advanced (Bhatia et al. 2016). Biocrimes can be witnessed from the time of Bible. The Bible documents that God demanded the freedom of Israelite slaves by the Pharaoh. However, when the Pharaoh did not agree, God sent ten deadly plagues upon Egypt. One of them was Shkhim (the sixth plague) which is a skin disease causing boils on the whole body. It is believed that anthrax was a cause of these boils, and it spreads when Moses sprinkles soot in the air (Mickley 2010).
An incident considered to be the most plausible was the use of bioweapon in 1346 by the Mongols. The Tartar army besieged the city of Caffa in the fourteenth century. The soldiers of the Tartar army started to die due to plague, and the survivors threw the infected bodies over the wall into the city in order to spread the disease among the residents, which caused an epidemic resulting in the residents leaving the place and the troops taking over the city of Caffa (Carus 2017).
Another example of bioweapon which could be quoted here is of “ricin.” Ricin is a plant toxin that is extracted from castor plant Ricinus communis. The CDC has classified it as a category B pathogen that is easy to disseminate and has low mortality. Many cases have been reported in the past where conviction has been made for the possession or misuse of ricin. In a well-known case, a Bulgarian exile, Georgi Markov who was a novelist and a play writer, had published anticommunist views. On September 7, 1978, while standing at the bus stop, he felt a stab in his right thigh. It was found that a man had injected a ricin pallet in his thigh using an umbrella-like weapon (Crompton and Gall 1980). Subsequently, he suffered from gastroenteritis and fever and reportedly died within 3 days of the occurrence of the event. Autopsy report concluded that the death was caused by ricin (Duncan and Smith 2012). The bioterrorist attacks have been a matter of concern in the present century since 2001. In 2001, following the attack on World Trade Center, an unknown person sent bacterium B. anthracis spores through letters to news and congressional offices in the United States. This agent is the causative agent of the disease anthrax. In this attack, 5 people died and 17 others were infected due to inhalation of anthrax spores. The Federal Bureau of Investigation (FBI) investigated the case, and Bruce Ivins was found to be the perpetrator of this attack. But, unfortunately, he committed suicide before he could face the trial in the court. This attack actually threw spotlight on the field of microbial forensics.
The bioweapons or microorganisms used for the commission of biocrimes may be natural, wild-type strains or genetically engineered microbes (Arora et al. 2002). The use of microbes as agents of bioterrorism is becoming a choice due to the ease of production and high toxicity (Rossodivita et al. 2019). Moreover, these are cost-effective weapons, need no expertise, and are less sophisticated but are more lethal and powerful (Budowle 2003; Moghaddasi et al. 2018). Further, most of the pathogens are endemically found and can be disseminated with ease. Furthermore, most of the organisms are quite stable and are transmittable (e.g., smallpox, anthrax, HIV, etc.), thereby causing potential threat with just a few initial outbreaks. Microbial forensics seeks to answer the basic questions including identification of the threat agent and linking it to a probable source, if possible. Due to the potential risks involved, the forensic scientists need to develop methods for identification and attribution of the microbes used in biocrimes. However, this becomes challenging as there are millions of species of microbes with a unique microbiome of their own. This makes it even more complicated to identify and individualize the causative microbe involved in any acts of biocrimes (Taylor et al. 2001).
The results of biowarfare often lead to the death of the host organism or to the toxicity of water or soil (Green et al. 2019; Janik et al. 2019). Apart from humans, the next potential threat is borne by agriculture and food, as both are exceptionally tempting targets of attack due to economic, social, as well as political impact they have (Rogers et al. 1999; Wilson et al. 2001; Budowle et al. 2005). The agents causing foodborne diseases can also be used as agents of bioterrorism, as no sophisticated equipment is required to spread the microbes involved; rather, the perpetrator generally needs a readily available, infested agent.
Microbial forensics, as the name suggests, refers to the deploying of scientific principles and techniques to analyze microbial evidence. Budowle (2003) defines microbial forensics as “a scientific discipline dedicated to analyze evidence from an act of bioterrorism, biocrime or inadvertent microorganism or toxin release for attribution purposes.” Microbial evidence is the evidence produced from any act of bioterrorism, biocrime, or an inadvertent release of toxin or microorganism (Budowle et al. 2003).
The misuse of biological agents poses severe threat to the society as well as the general health and well-being of the individuals. As said by Murch (2003), “Science and technology are used to serve as independent ‘witnesses’ in criminal or civil matters, intelligence, and policy.” Microbial forensics, hence, helps in the attribution of an offense, based upon the analysis of biological and microbial evidence. The disease-causing microbes and pathogens and the toxins released by them are identified and distinguished by the law enforcement agencies in collaboration with scientists from various fields like microbiology, genetics, public health, and so on.
3 Role of Microorganisms in the Decomposition Process
Decomposition is a natural process and refers to the breakdown of dead organic matter to smaller particles, thereby resulting in release of carbon dioxide, water, minerals, and energy into the atmosphere. It is a complex process in which both biotic and abiotic factors come into play and is aided by two different groups of organisms, namely, the scavengers and the microorganisms, which are considered as true decomposers (Saraswat et al. 2008).
The biotic factors include but are definitely not confined to the stature, size, and weight of the cadaver and presence of intrinsic and extrinsic bacteria, other microbes, insects, and scavengers. The abiotic factors include climatic conditions and humidity (Evans 1963; Vass 2001; Carter et al. 2008; Janaway et al. 2009; Pechal et al. 2013).
When an animal dies, a series of events take place, beginning from the shutdown of its immune system leading to a change in the internal temperature of the body and subsequently the initiation of the growth of microorganisms (Weiss et al. 2016). A dead body, at this stage, can in fact be compared to an ecosystem hosting a variety of bacteria, insects, and fungi (Hyde et al. 2013).
The increase in the internal temperature causes the skin of the cadaver to rupture, thereby allowing the interchange of air, microbes, and body fluids. A carcass releases large amounts of nitrogen into the environment, mostly in the form of ammonia, along with other elements like carbon and phosphorus. Decomposition is influenced by various taphonomic processes, which are in turn influenced by the geographical area where the cadaver has been discovered (Goff 1993).
Initially, the bacteria begin with the digestion of the intestine, thereby causing decomposition of the body from inside to the outside (Janaway et al. 2009). Autolysis of the cells of the cadaver also begins, and the internal bacteria undergo anaerobic respiration to release putrefying gases like hydrogen sulfide, methane, cadaverine, and putrescine (Vass 2001). However, if the cadaver is frozen immediately following the death, then the decomposition occurs from outside to inside, after the body is brought to a temperature above 0 °C. Due to the accumulation of gases in the internal organs, the body bloats. Putrefaction begins during the bloat stage, and the cadaver looks inflated like a balloon. The cadaver becomes a site of attraction as well as infestation for the adult blowflies (Calliphoridae).
Next to bloating is the decay stage, which is marked by the rupture of the outer layer of the skin resultantly leading to deflation of the body as the gases present in the abdomen escape and the decay initiates. The strong odor associated with decomposition is evident, and the cadaver at this stage becomes the breeding ground for large masses of Diptera larvae. Most of the Calliphoridae present generally complete their development and leave the decomposing remains. By the end of the decay stage, only the skin and the cartilages are visible (Catts and Haskell 1990; Hewadikaram and Goff 1991), and this marks the beginning of the post-decay stage, where Diptera are no longer the principal organisms in the cadaver. In dry and arid xerophytic and mesophytic conditions, Diptera generally gets replaced by the adult Coleoptera, who feeds on the cadaver, thereby removing the remaining flesh and cartilage so as to expose the clean bones. The post-decay stage is followed by the skeletal stage, wherein a number of soil-dwelling microorganisms can be seen for an initial period.
In order to determine the postmortem interval (PMI), the stages of decomposition form an important parameter. Several studies have reported the stages of decomposition as a function of temperature to find out the time intervals for decomposition. Megyesi et al. (2005) devised a scoring system to evaluate decomposition as a function of time by using bloating and purging of the head and trunk to mark the end of early decomposition. However, it has been reported that bloat and purge are quite proficient to evaluate all the stages of decomposition (Hyde et al. 2013).
Though bloat has been profusely used as a parameter to estimate time since death, limited literature is accounted for the action of internal microorganisms (Melvin et al. 1984; Hopkins et al. 2000; Howard et al. 2010). There are, however, many studies to show the role of microorganisms in decomposition in general (Melvin et al. 1984; Kakizaki et al. 2008; Meyers and Foran 2008; Stokes et al. 2009; Howard et al. 2010; Lenz and Foran 2010; Dickson et al. 2011; Butzbach et al. 2013; Schoenen and Schoenen 2013).
The geographical conditions like climate and topography also affect the process of decomposition. In dry and arid regions, with strong air current, the cadaver tends to mummify (Wescott 2018), while in warm and humid regions, the decomposition is quick (Powers et al. 2009; Forbes et al. 2014). Hence, the plentitude and variety and dissemination of the microorganisms in the cadaver are highly significant parameters in the determination of the postmortem interval.
4 Soil Microbes as Forensic Biomarkers
The study of soil is known as “pedology.” Soil is a complex substance which contains minerals as well as decaying organic matter. This organic matter supports microbial life, the presence of which causes significant changes in the soil profile. Soil analysis is potent for linking a person, animal, or plant to a specific place. Off lately, most of the forensic studies on soil were based upon the chemical as well as the mineral profile of the soil (Tibbett and Carter 2009; Ruffell 2010). This required large amounts of soil sample and lacked a good mapped soil database (Zala 2007). However, due to extremely diverse microbial communities present in the soil, the focus is now shifting to the microbes for identification purposes.
The soil microbes thrive well in a specific pH, the alteration to which can affect the microbial density in a particular soil type. Generally, with an increase in the pH, the microbial biomass decreases (Santiago-Rodriguez TM 2016). Changes in pH may further lead to alterations in the cellular processes like DNA replication and transcription, thereby resulting in specific changes which may allow the specific population of microbes to survive in varied pHs (Eilers et al. 2012).
Biological factors including anthropogenic activities like agriculture and discharge of feces and sewage also influence the soil microbial diversity (Brookes and Mcgrath 1984; Elliott 1986; Stockdale and Brookes 2006). There are other external sources of alteration such as decomposition of a cadaver which can alter the soil composition. When soil comes in contact with a decomposing cadaver, the microbes in the immediately surrounding areas are likely to get highly affected. As a result of decomposition, significant quantity of decomposing organic matter from the cadaver may leak into the soil (Vass et al. 2004), thereby causing changes in the soil profile. These changes may be quantified and may assist in the crime scene reconstruction by relating the cadaver to the site of its presence by way of the analysis of soil microbes.
Most of the studies relating to soil microbiology have been undertaken using 16S ribosomal RNA gene sequencing using terminal restriction fragment length polymorphism analysis (T-RFLP) (Quaak and Kuiper 2011) or amplicon length heterogeneity-polymerase chain reaction (ALH-PCR) (Moreno et al. 2006). The rRNA gene is only about 1.5 kilobases long, due to which it is easy to sequence and is cost-effective (Gunn 2009). The study of soil microbes is advantageous as a very small sample size is required owing to the plentitude of soil microbes. Further, the techniques involved are comparatively cheap and automated (Hill et al. 2007; Sensabaugh 2009).
5 Personal Identification from Microbes Present in Saliva
Saliva is a fluid secreted by the salivary glands found in the mouth of humans and other animals. In addition to the salivary fluid, it contains exfoliated epithelial cells of the mouth as well as the microorganisms. Saliva is of utmost importance to forensic scientists as it is easy to sample and the technique used for collection is less invasive than sampling blood or urine.
Saliva may often be encountered in sexual offenses along with bite marks or lip prints and can aid in personal identification of the culprit. The methods of identification mainly depend upon analysis of human DNA. Nevertheless, the human DNA may be degraded or masked and may be present in less quantity. The bacterial DNA, therefore, is of immense utility as it is comparatively better resistant to degradation than human DNA (Leake et al. 2016). The differentiation between identical twins may also be possible by using bacterial DNA (Stahringer et al. 2012). Almost 99% of the environmental bacteria cannot be cultured in the lab (Handelsman 2004). However, with the advent of modern technological aids like the next-generation sequencing (NGS), the bacteria can be employed for a variety of forensic purposes. Studies have reported that it is possible to recover live bacteria from bite sites if they are undisturbed for 24 h (Borgula et al. 2003). It has further been reported that microbes present on the skin can be potent to link an individual to the objects touched by them (Fierer et al. 2010).
A lot of studies have shown the composition of the bacterial communities present in the saliva (Costello et al. 2009; Lazarevic et al. 2009; Zaura et al. 2009; Lazarevic et al. 2010; Caporaso et al. 2011), but the extent up to which it can differentiate between two individuals needs to be explored. A lot of research is required to establish the reliability of bacterial microbiome in personal identification.
6 Role of Microbes in Reconstruction of Crime Scene
Crime scene reconstruction is the process that helps the investigators to interpret the evidences found at the crime scene. Law enforcement agencies play a crucial role in assessing and investigating the evidences, wherein microbial forensics comes as an aid to analyze and characterize the microbial evidence for crime scene reconstruction.
6.1 Role in Epidemiology
The World Health Organization (WHO) defines epidemiology as the study of the distribution and determinants of health-related states or events (including disease) and the application of this study to control the diseases and other health problems. Epidemiology studies the occurrence of disease as well as factors responsible for the occurrence of disease in populations. Biocrimes also come under the purview of the general principles of epidemiologic investigation involving a biological agent (Flowers et al. 2002). In case of an outbreak of a disease, certain important points have to be taken into consideration, namely, determination of the occurrence of an outbreak, identification of the population at risk, determination of the method of dispersal and dissemination of disease, and last but not the least, identification and characterization of the causal agent (Morse and Budowle 2006). In order to link the donor and recipient of an infectious disease, the associated genetic marker, exhibiting the following important criteria, has to be assessed:
-
(a)
Sufficient variation must be exhibited by the genetic markers so as to exclude the unlinked individuals.
-
(b)
Such variation must be easily identifiable.
-
(c)
The genetic variation should neither arise at a very slow or a very fast pace so as to compromise one’s ability to link the infected people together (Gunn 2009).
The use of epidemiologic methods for investigating the suspicious health problems/diseases or evidences relating to intentional acts or criminal behavior is called forensic epidemiology. In fact, forensic science and epidemiology must be treated as an integrated discipline, as many of the methods and procedures followed in epidemiology are similar to the ones used in microbial forensics (Goodman et al. 2003). The investigation under microbial forensics differs from epidemiological investigations on the point that the former are scrutinized and evaluated by the scientists as well as the law enforcement agencies while the latter is evaluated by scientists alone.
6.2 Cause and Manner of Death
When a person dies, one of the main objectives is to determine the cause and manner of death. Microbes can be of immense utility in some cases to determine whether an individual died due to a natural cause, such as disease, or a biological attack (Gunn and Sarah 2012). If the cause of death remains unknown till the time the individual was alive, then postmortem analysis along with serological and/or genomic analysis may help to deduce the underlying cause of death. If the disease-causing agent is identified prior to death, the analysis confirming the agent to be the cause of death may still be desired post death. Knowledge about the cause of death is helpful in treating other patients having similar ailment; and it may also help in preventing the dissemination of disease by either warning the public or quarantining a suspicious contaminated agent/source.
The role of microbes to determine the cause of death can be understood by quoting the example of SIDS (sudden infant death syndrome), which is one of the leading causes of post-neonatal infant mortality in the world. It has been suggested that microorganisms play a crucial role in many of the cases of SIDS (Gleeson and Cripps 2004). Studies have reported that viral infections caused by human herpesvirus-6, Epstein–Barr virus, and cytomegalovirus were present in infants that died from SIDS (Alvarez-Lafuente et al. 2008). SIDS has also been reported to be associated with bacteria like Staphylococcus aureus, Escherichia coli, Streptococcus spp., and Haemophilus influenzae (Blackwell 2004). The cause of death was inferred by relating the presence of specific microbes in the deceased individuals.
Attribution of microbial infection as a cause of death must be done with utmost care because numerous bacteria are otherwise present in the human body in a sterile state. However, these sterile bacteria may start taking advantage of the lack of a healthy immune system in case of death of the host (Janaway et al. 2009).
6.3 Drowning Cases
In cases of drowning, microbial communities present in the water may provide useful evidence about the cause of death as well as the place of drowning. The diatoms are one of the most common microorganisms which are indicative of drowning as the cause of death (Timperman 1972). Diatoms are unicellular, microscopic phytoplanktons containing silica cell walls. The silica cell wall is resistant to degradation from acid, enzymes, and temperature, due to which they are present in the drowned bodies (Lin et al. 2014; Rana and Manhas 2018). They have been found in lungs, blood, bone marrow, and other internal organs (Ludes et al. 1999). Diatoms are exclusively present in cases where the person drowned in natural water body like sea, lake, or river. The water bodies containing treated water like a swimming pool generally lack the diatoms due to the process of water treatment (Lin et al. 2014).
6.4 Sexual Assault Cases
Cases involving sexual assault are another category where microbial forensics helps link the victim to the suspect. In cases of sexual assault, a victim can generally furnish a statement about the occurrence of the crime. However, in certain instances, the victim cannot speak for himself, as in cases of child sexual abuse. In such cases, microbial forensic analysis is of utmost significance as it may help link the victim to the culprit by detecting the sexually transmitted pathogens. This may be done by using various methods like nucleic acid amplification tests (NAATs) (Jaureguy et al. 2016) and cultures to analyze the body fluids collected by way of swabs. The bed sheets, blankets, clothing, hair, as well as fibers may also be additionally analyzed. A number of pathogens are reported to cause sexually transmitted diseases (STDs), including Neisseria gonorrhoeae, Chlamydia trachomatis, and Trichomonas vaginalis, to name a few. STDs can be transmitted during sexual assault and can be of forensic significance in cases involving transmission of an infectious disease as a weapon of biocrime.
6.5 Human Immunodeficiency Virus (HIV)
HIV is another example of a natural disease-causing pathogen that has been reported as a means of commission of a biocrime (Ou et al. 1988; Ou et al. 1992; Robbins et al. 2003). Though viruses can infect all life forms, they cannot replicate without the host mechanism. A basic virus is comprised of a nucleic acid genome which is present inside a protein coat. Viruses may ether be composed of DNA or RNA. HIV is an RNA virus, which evolves expeditiously (Carrillo and Rock 2005) and may exhibit high rates of mutation (Drake et al. 1998; Preston et al. 1988; Sala and Wain-Hobson 2000; Svarovskaia et al. 2004). Therefore, there is a high probability that two HIV samples having common origin differ at a number of nucleotides within the genome. Hence, it is impossible to obtain an exact match between the virus molecular profile from an alleged donor and recipient.
Investigations in HIV cases involve both gathering the factual information referred to as epidemiological investigation as well as the genetic and/or serologic data. Further, phylogenetic analysis comes as an aid in supporting or contradicting the relationships of samples that have an alleged common origin in comparison to those which do not.
7 Collection of Microbial Evidence
Evidence collection is of paramount importance for any forensic investigation, including microbial forensic investigation. Degradation or contamination of evidence may prove detrimental for the investigation and in turn for the whole process of attribution (Budowle et al. 2003). Investigations pertaining to microbial forensics are potentially similar to other forensic investigations, wherein similar steps of crime scene investigation, evidence collection and preservation, analysis of evidence, and maintaining a chain of custody are to be followed. Microbiological evidence may include viable samples of the microbes, their toxins, nucleic acids, specimens from victims, laboratory equipment, dissemination devices and their contents, environmental samples, contaminated clothing, or trace evidence specific to the process that produced the particular biological agent. Therefore, the method of collection should be sensitive and reliable.
The biocrimes are generally reported by the first responders including health-care providers, police, paramedics, firefighters, or even the general public. The first responders are followed by the crime scene investigators. For most of the cases involving microorganisms, a plan describing the detailed strategy must be devised prior to the initial sampling. This is so because the microbes involved in that very particular case may be hazardous and fatal for the crime scene investigators. The investigators must wear protective equipment so as to protect them from the hazards related to the microbial evidences (CDC 2002).
The samples to be collected must include every material found on the scene which must be well labeled indicating the time and site of collection. The name of the person who has collected the sample should also be mentioned.
Evidence collection may be done following three general approaches, the foremost being collection of the whole item and transporting it to a safer place for further sampling and analysis. In case the whole item is too large to be collected and transported, a part of the item can be sampled and preserved by using techniques like vacuuming or filtration. For trace materials, swabbing or wiping the contaminated surfaces with suitable sample collection devices is suggested (Smith 2011). The sample collection may be done with the help of either dry or wet swabs, wipes, vacuums, filters, aspirating – needles, and so on (Buttner et al. 2001; Buttner et al. 2005). However, the sample collection methods as well as the devices have certain limitations, the primary being the lack of proper validation of the methods in use. Further, it cannot be said with certainty as to which method would be the most effective method for investigation of that particular evidence.
The general methodology for processing the microbial samples is outlined as follows:
-
Sample collection
-
Sample transportation
-
Sample extraction
-
Sample analysis (Junkins et al. 2017)
It is obvious that the procedure for each stage will depend upon the organisms to be collected. Extraction techniques will further determine what analyses can be done on each sample. In some cases, the quantity of the sample available may be limited. Moreover, the data on the techniques which could be efficient may or may not be available to the investigator. Further, owing to the diversity of microbes, there is a great deal of uncertainty associated with the analysis. A technique might work well for one organism but at the same time may be deleterious for another (Schutzer et al. 2005). Another critical point to take into account is that the procedures used for sample collection must be developed so as to preserve the traditional forensic evidences like hair, documents, fingerprints, DNA, etc., wherever possible. Minimal disturbance of evidence is appreciated in a forensic setup; hence, the methods of collection must preferably be noninvasive like sterile swabbing methods. Generally, microbes are present in larger numbers in the gastrointestinal (GI) tract and other body openings like buccal cavity and the anal orifice; therefore, collection from these regions is comparatively easier. It has been reported that in general, 0.25–0.50 g of microbial GI material per sample is sufficient for DNA isolation. It has further been suggested to collect replicate samples, if possible (Pechal et al. 2017).
There are many other points of significance which must be followed while on the field, the prime being that the investigator must be aware of as well as careful of the other animals present on the field that may either feed upon the cadaver or may even be harmful to the investigator. One must take precautions on the field so as to protect the cadaver as well as oneself from the necrophagous organisms present on the field. This may generally be achieved by using protective cage made of sturdy material covered by a layer of strong mesh so as to protect the cadaver from scavengers. The large mesh may be further lined by a fine net if required, to prevent the small insects from entering.
In the study of microbes for estimating PMI, insects that feed on the decomposing body as well as those on and under the soil surrounding the cadaver may be of interest (Pechal et al. 2014; Weiss et al. 2016). While collecting soil microbes associated with the decomposing cadaver, the placement of the cadaver on the soil is significant. It is so because the soil present under and around the cadaver is a mixture of microbes originally present in the soil and those which have leached from the decomposing cadaver (Carter et al. 2007). This in turn influences the diversity and density of soil microbes; hence, it is appropriate to sample soil from different depths and document properly the layer of soil which has been sampled (Ranjard et al. 2001; Kakirde et al. 2010). Further, the quantity of the soil sample must be adjusted so as to house the desired amount of microbiome. Soil samples collected within 5 cm from the surface generally requires 5–10 g of soil, while the soil present on the subsurface may require a larger volume so as to be sufficiently rich in biomass for DNA/RNA extraction. Separate sterile devices like tubes or pipes must be used for collecting different samples (Kakirde et al. 2010).
It is advisable to process the samples as soon as possible (within 2 h) so as to prevent any material changes like oxygenation in them. However, in practical field scenario, this may not be accomplished with ease (Rochelle et al. 1994). Therefore, it is advisable to protect the samples from extreme changes in temperature, pH, and ionic strength of a solution, if applicable (Wilfinger et al. 1997). Use of airtight containers must be done cautiously depending upon the type of microbes to be stored. For instance, storing the aerobic samples in airtight containers could affect the microbial composition as it will restrict the exchange of air and gasses. Conversely, storing the anaerobic communities in excess of air might contaminate the sample (Rochelle et al. 1994). When immediate transfer and processing of samples are not possible, it is advisable to have immediate, sample storage facility on the field to facilitate short-term sample storage. This may include innovations like ice packs, coolers, etc. Storage at −20 °C is recommended. Though freezing is used as an aid to preserve the sample in a majority of studies, however, some studies suggest that freezing changes the structure of microbial community (Kakirde et al. 2010; Carroll et al. 2012). Once frozen, the samples must not be thawed unless and until extraction has to be done. Repeated cycles of freezing and thawing must be avoided so as to avoid fragmentation, which may in turn affect characterization of the microbiome (Männistö et al. 2009; Cardona et al. 2012). It is needless to say that a checklist as well as a proper chain of custody must be maintained prior and subsequent to sample collection.
It is needless to say that the samples collected must be transported to the lab as soon as possible on ice or in liquid nitrogen and must be stored at −20 or − 80 °C till the time they are processed.
8 Analysis of Microbial Evidence
Classical technique for the identification of microbes involves the biochemical testing of the microorganism in pure culture in different growth media (Maccallum and Hastings 1899). After a microorganism is isolated and obtained in pure culture, it can be analyzed phenotypically, genotypically, or both. Phenotype refers to the set of observable characteristics like the color, colony morphology, growth pattern on agar and in broth, susceptibility to antibiotics, etc., of an individual which results from the interaction of its genotype with the environment. Genotype refers to the specific genetic combination which leads to the phenotype (Slonczewski and Foster 2013). Some microbes can even be identified by the biochemical processes performed by them, e.g., the ability to create the enzyme “catalase,” which helps in the breaking down of hydrogen peroxide to oxygen and water (Slonczewski and Foster 2013).
However, though the classical methods are a valuable source for the identification of microorganisms, there are certain limitations of this method. It has been reported that every microorganism cannot be cultured in the lab. In fact, only 1–2% of the microorganisms are reported to be culturable (Amann et al. 1995; Wilson et al. 1997). Further, most of the organisms are fastidious and have specific environmental and nutritional requirements. Furthermore, the organism to be cultivated must be live, and the methods of commercial identification that are not optimized can therefore lead to misidentification. In view of the above limitations, it is very difficult, if not impossible, to obtain a forensically significant culture.
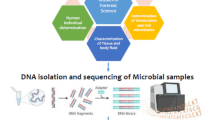
8.1 Extraction of DNA for Analysis
The method used for extraction of DNA tends to have a significant impact on the interpretation of results of microbial structure and function (Henderson et al. 2013; Wagner Mackenzie et al. 2015). The method of DNA extraction must be such so as to enable the efficient recovery of both the prokaryotic and the eukaryotic DNA. The microbial DNA can be extracted from the environment either directly, whereby the entire sample is lysed during cell lysis, or indirectly, where the microbial cells first get separated and subsequently undergo lysis during the cell lysis (Ogram et al. 1987; Parachin et al. 2010; Delmont et al. 2011). As both the methods mentioned here yield similar results (Delmont et al. 2011), the choice of method would depend upon the experimental requirement and setup. For example, if the aim of the experiment is to construct large-insert clone libraries, the indirect method would be a better choice as the direct method would lead to the shearing of the genomic DNA. On the other hand, if the aim involves deploying the DNA for 16 s rDNA or shotgun sequencing where small sample sizes are sufficient, the direct method would be better as it is less strenuous and quicker and yields high DNA content. However, the direct method has a disadvantage that it cannot remove PCR inhibitors from some environmental samples efficiently. Due to the development of commercial DNA purification kits, which can remove common PCR inhibitors, the direct method for DNA extraction has become quite popular in recent years.
The quantity as well as the quality of the extracted DNA may be evaluated using a number of different techniques. The purity of DNA is assessed by using UV spectroscopy; the presence of inhibitors is evaluated using a PCR inhibition assay, while the intactness is evaluated using gel electrophoresis. The quantification of DNA is mostly done by using quantitative polymerase chain reaction (qPCR) (Budowle et al. 2005).
Many real-time PCR assays are highly specific as well as sensitive and thereby shorten the time required for analysis when compared to the conventional PCR protocols, cultivation, as well as biochemical methods of identification. The next-generation sequencing which targets the nucleic acids is recently being deployed as a popular method to study genetic material from mixed microbial populations for analysis of microbial communities. The ability of the technique to identify the species up to the strain level from both live and dead microorganisms which are nonculturable has been very advantageous for the field of microbiology (Woese et al. 2000). Common sequencing techniques include 16S or 18S ribosomal RNA gene sequencing; next-generation sequencing (NGS), namely, pyrosequencing; and pulsed field gel electrophoresis (Gu et al. 2015). Techniques involving non-DNA-based analysis such as studying microbial chemistry and matrix using mass spectrometry can also be utilized (Seng et al. 2013).
9 Conclusion
Microorganisms are ubiquitous diverse organisms, harbored by the humans majorly on their skin as well as on the gastrointestinal and respiratory tract. The technological advances in microbial profiling are very significant for forensic purposes as they help in crime scene reconstruction by linking people, animals, plants, and objects to one another and/or to the scene of crime. The analysis and evaluation of the microbial profiles could be of immense use in estimating the postmortem interval (PMI). The soil profile of a crime scene may help to trace the location of the cadaver by studying the microbes present in the soil. Microorganisms have the potential to solve the cases related to origin of biocrimes, sexual assaults, homicides, agricultural contaminations, and medical malpractices. Microbial forensics quantifies molecular variations to find out the origin and the route of transmission of a particular microbial strain.
In order to be effectively applicable in the court of law, the techniques involved in the analysis of microbes must be standardized and validated. There is an utmost need of national and international collaborations to create a database on bioterrorism for the development of bioterrorism information system that can help to prevent the threat of bioattack. To combat the menace of biocrimes rapidly and effectively, there is a need to allocate funds for the establishment of hi-tech microbial forensic laboratories for research and development. Therefore, though forensic microbiology has vast applications and utility in criminal investigations and administration of law, however, a great deal of research in the subject is required so that its potential can be fully explored.
References
Álvarez-Lafuente R, Aguilera B, Suárez-Mier MP et al (2008) Detection of human herpesvirus-6, Epstein-Barr virus and cytomegalovirus in formalin-fixed tissues from sudden infant death: a study with quantitative real-time PCR. Forensic Sci Int 178(2–3):106–111
Amann R, Ludwig W, Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59(1):143–169
Arora DR, Gautam V, Arora B (2002) Biological warfare: bioterrorism. Indian J Med Microbiol 20(1):6–11
Bhatia M, Mishra B, Thakur A et al (2016) Concept of forensic microbiology and its applications. SMU Med J 3(1):275–294
Blackwell C (2004) Infection, inflammation and SIDS. FEMS Pathogens Dis 42(1):1–2
Bojar RA, Holland KT (2002) The human cutaneous microflora and factors controlling colonisation. World J Microbiol Biotechnol 18(9):889–903
Borgula LM, Robinson FG, Rahimi M, Chew KEK, Birchmeier KR, Owens SG, Kieser JA, Tompkins GR (2003) Recovery of oral bacteria from experimental bite marks. J Forensic Odontostomato 21:23–30
Brookes PC, McGrath SP (1984) Effect of metal toxicity on the size of the soil microbial biomass. J Soil Sci 35(2):341–346
Budowle B (2003) Defining a new forensic discipline: microbial forensics. Profiles in DNA 6(1):7–10
Budowle B, Schutzer SE, Einseln A et al (2003) Building microbial forensics as a response to bioterrorism. Science 301(5641):1852–1853
Budowle B, Johnson MD, Fraser CM et al (2005) Genetic analysis and attribution of microbial forensics evidence. Crit Rev Microbiol 31(4):233–254
Buttner MP, Cruz-Perez P, Stetzenbach LD (2001) Enhanced detection of surface-associated bacteria in indoor environments by quantitative PCR. Appl Environ Microbiol 67(6):2564–2570
Buttner MP, Cruz P, Stetzenbach LD et al (2005) Evaluation of the biological sampling kit (BiSKit) for large-area surface sampling. Appl Environ Microbiol 70(12):7040–7045
Butzbach DM, Stockham PC, Kobus HJ et al (2013) Bacterial degradation of risperidone and paliperidone in decomposing blood. J Forensic Sci 58(1):90–100
Caporaso JG, Lauber CL, Costello EK et al (2011) Moving pictures of the human microbiome. Genome Biol 12(5):R50
Cardona OD, van Aalst MK, Birkmann J, Fordham M, McGregor G, Mechler R (2012) Determinants of risk: exposure and vulnerability. Cambridge University Press, Cambridge
Carrillo C, Rock DL (2005) Molecular epidemiology and forensics of RNA viruses. In: Breeze R, Budowle B, Schutzer S (eds) Microbial forensics. Academic Press, Amsterdam
Carter DO, Yellowlees D, Tibbett M (2007) Cadaver decomposition in terrestrial ecosystems. Naturwissenschaften 94(1):12–24
Carter DO, Yellowlees D, Tibbett M (2008) Temperature affects microbial decomposition of cadavers (Rattus rattus) in contrasting soils. Appl Soil Ecol 40(1):129–137
Carroll IM, Ringel-Kulka T, Siddle JP et al (2012) Characterization of the fecal microbiota using high-throughput sequencing reveals a stable microbial community during storage. PLoS One 7(10):e46953
Carus WS (2001) Bioterrorism and biocrimes: the illicit use of biological agents since 1900. National Defense University, Washington, DC
Carus WS (2017) A short history of biological warfare: from from pre-history to the 21st century. National Defense University Press, Washington, DC
Catts EP, Haskell NH (1990) Entomology and death: a procedural guide. Joyce’s Print Shop. Inc., Clemson, p 182
Centers for Disease Control and Prevention (CDC) (2002) Occupational health guidelines for remediation workers at Bacillus anthracis-contaminated sites–United States, 2001–2002. MMWR. Morbidity and mortality weekly report. 51(35):786
Costello EK, Lauber CL, Hamady M et al (2009) Bacterial community variation in human body habitats across space and time. Science 326(5960):1694–1697
Crompton R, Gall D (1980) Georgi Markov—death in a pellet. Med Leg J 48(2):51–62
Cummings CA, Relman DA (2002). Microbial Forensics-“Cross-Examining Pathogens”. Science’s Compass 296:1976–77
Delmont TO, Robe P, Clark I, Simonet P, Vogel TM (2011) Metagenomic comparison of direct and indirect soil DNA extraction approaches. J Microbiol Methods 86(3):397–400
Dickson GC, Poulter RT, Maas EW et al (2011) Marine bacterial succession as a potential indicator of postmortem submersion interval. Forensic Sci Int 209(1–3):1–10
Drake JW, Charlesworth B, Charlesworth D (1998) Rates of spontaneous mutation. Genetics 148(4):1667–1686
Duncan RVI, Smith LA (2012) Of beans and beads: ricin and abrin in bioterrorism and biocrime. J Bioterr Biodef S2:002. https://doi.org/10.4172/2157-2526.S2-002
Eilers KG, Debenport S, Anderson S et al (2012) Digging deeper to find unique microbial communities: the strong effect of depth on the structure of bacterial and archaeal communities in soil. Soil Biol Biochem 50:58–65
Elliott ET (1986) Aggregate structure and carbon, nitrogen, and phosphorus in native and cultivated soils 1. Soil Sci Soc Am J 50(3):627–633
Evans WE (1963) Adipocere formation in a relatively dry environment. Med Sci Law 3:145–153
Fierer N, Lauber CL, Zhou N et al (2010) Forensic identification using skin bacterial communities. Proc Natl Acad Sci U S A 107(14):6477–6481
Flowers LK, Mothershead JL, Blackwell TH (2002) Bioterrorism preparedness. II: the community and emergency medical services systems. Emerg Med Clin North Am 20(2):457–476
Forbes SL, Perrault KA, Stefanuto PH et al (2014) Comparison of the decomposition VOC profile during winter and summer in a moist, mid-latitude (Cfb) climate. PLoS One 9(11):e113681
Gleeson M, Cripps A (2004) Development of mucosal immunity in the first year of life and relationship to sudden infant death syndrome. FEMS Immunol Med Microbiol 42(1):21–33
Goff ML (1993) Estimation of postmortem interval using arthropod development and successional patterns. Forensic Sci Rev 5:81–94
Goodman RA, Munson JW, Dammers K et al (2003) Forensic epidemiology: law at the intersection of public health and criminal investigations. J Law Med Ethics 31(4):684–700
Green MS, LeDuc J, Cohen D et al (2019) Confronting the threat of bioterrorism: realities, challenges, and defensive strategies. Lancet Infect Dis 19(1):e2–e13
Grice EA, Kong HH, Conlan S et al (2009) Topographical and temporal diversity of the human skin microbiome. Science 324(5931):1190–1192
Grundmann O (2014) The current state of bioterrorist attack surveillance and preparedness in the US. Risk Manage Healthc Policy 7:177–187
Gu Y, Mao X, Zha L, Hou Y, Yun L (2015) Development of a candidate method for forensic microbial genotyping using multiplex pyrosequencing combined with a universal biotinylated primer. Forensic Sci Int 246:e1–e6
Gunn A (2009) Essential forensic biology, 2nd edn. John Wiley & Sons
Gunn A, Sarah JP (2012) Microbes as forensic indicators. Trop Biomed 29(3):311–330
Handelsman J (2004) Metagenomics: application of genomics to uncultured microorganisms. Microbiol Mol Biol Rev 68(4):669–685
Henderson G, Cox F, Kittelmann S et al (2013) Effect of DNA extraction methods and sampling techniques on the apparent structure of cow and sheep rumen microbial communities. PLoS One 8(9):e74787
Hewadikaram KA, Goff ML (1991) Effect of carcass size on rate of decomposition and arthropod succession patterns. Am J Forensic Med Pathol 12(3):235–240
Hill PW, Marshall C, Williams CG, Blum H, Harmens H, Jones DL, Farrar JF (2007) The fate of photosynthetically-fixed carbon in grassland as modified by elevated CO and sward management. New Phytol 173(4):766–777
Hopkins DW, Wiltshire PE, Turner BD (2000) Microbial characteristics of soils from graves: an investigation at the interface of soil microbiology and forensic science. Appl Soil Ecol 14(3):283–288
Howard GT, Duos B, Watson-Horzelski EJ (2010) Characterization of the soil microbial community associated with the decomposition of a swine carcass. Int Biodeterior Biodegradation 64:300–304
Hyde ER, Haarmann DP, Lynne AM et al (2013) The living dead: bacterial community structure of a cadaver at the onset and end of the bloat stage of decomposition. PloS one 8(10):e77733
Janaway RC, Percival SL, Wilson AS (2009) Decomposition of human remains. In: Microbiology and aging. Humana Press, Totowa, pp 313–334
Janik E, Ceremuga M, Saluk-Bijak J et al (2019) Biological toxins as the potential tools for bioterrorism. Int J Mol Sci 20(5):1181
Jansen HJ, Breeveld FJ, Stijnis C (2014) Biological warfare, bioterrorism, and biocrime. Clin Microbiol Infect 20(6):488–496
Jaureguy F, Chariot P, Vessieres A, Picard B (2016) Prevalence of chlamydia trachomatis and Neisseria gonorrhoeae infections detected by real-time PCR among individuals reporting sexual assaults in the Paris, France area. Forensic Sci Int 266:130–133
Junkins EN, Embriette RH, David OC (2017) Culture and long-term storage of microorganisms for forensic science. In: Carter DO, Tomberlin JK, Benbow ME, Metcalf JL (eds). John Wiley & Sons, New York
Kakirde K, Parsley L, Liles M (2010) Size does matter: application-driven approaches for soil metagenomics. Soil Biol Biochem 42(11):1911–1923
Kakizaki E, Takahama K, Seo Y et al (2008) Marine bacteria comprise a possible indicator of drowning in seawater. Forensic Sci Int 176(2–3):236–247
Kaur M, Gupte S, Aggarwal P et al (2014) Methods in microbial forensics. J Punjab Acad Forensic Med Toxicol 14(1):51–54
Lazarevic V, Whiteson K, Huse S et al (2009) Metagenomic study of the oral microbiota by Illumina high-throughput sequencing. J Microbiol Methods 79(3):266–271
Lazarevic V, Whiteson K, Hernandez D et al (2010) Study of inter-and intra-individual variations in the salivary microbiota. BMC Genomics 11(1):523
Leake SL, Pagni M, Falquet L (2016) The salivary microbiome for differentiating individuals: proof of principle. Microbes Infect 18(6):399–405
Lenz EJ, Foran DR (2010) Bacterial profiling of soil using genus-specific markers and multidimensional scaling. J Forensic Sci 55(6):1437–1442
Lin CY, Yen WC, Hsieh HM et al (2014) Diatomological investigation in sphenoid sinus fluid and lung tissue from cases of suspected drowning. Forensic Sci Int 244:111–1115
Ludes B, Coste M, North N, Doray S, Tracqui A, Kintz P (1999) Diatom analysis in victim’s tissues as an indicator of the site of drowning. Int J Legal Med 112(3):163–166
Maccallum WG, Hastings TW (1899) A case of acute endocarditis caused by micrococcus zymogenes (Nov. Spec.), with a description of the microorganism. J Exp Med 4(5–6):521–534
Männistö MK, Tiirola M, Häggblom MM (2009) Effect of freeze-thaw cycles on bacterial communities of Arctic tundra soil. Microb Ecol 58(3):621–631
Megyesi MS, Nawrocki SP, Haskell NH (2005) Using accumulated degree-days to estimate the postmortem interval from decomposed human remains. J Forensic Sci 50(3):1–9
Melvin JR, Cronholm LS, Simson LR et al (1984) Bacterial transmigration as an indicator of time of death. J Forensic Sci 29(2):412–417
Meyers MS, Foran DR (2008) Spatial and temporal influences on bacterial profiling of forensic soil samples. J Forensic Sci 53(3):652–660
Mickley A (2010) Preventing the sixth plague: microbial forensics in the war against terrorism. Honors College Theses 1:91
Moghaddasi H, Arani LS, Zarghi A (2018) Features of bioterrorism information system. J Bioterror Biodefense 9:1–6
Moreno LI, Mills DK, Entry J, Sautter RT, Mathee K (2006) Microbial metagenome profiling using amplicon length heterogeneity-polymerase chain reaction proves more effective than elemental analysis in discriminating soil specimens. J Forensic Sci 51(6):1315–1322
Morse SA, Budowle B (2006) Microbial forensics: application to bioterrorism preparedness and response. Infect Dis Clin 20(2):455–473
Murch RS (2003) Microbial forensics: building a national capacity to investigate bioterrorism. Biosecur Bioterr Biodef Strategy Pract Sci 1(2):117–122
Ogram A, Sayler GS, Barkay T (1987) The extraction and purification of microbial DNA from sediments. J Microbiol Methods 7(2–3):57–66
Oliveira M, Amorim A (2018) Microbial forensics: new breakthroughs and future prospects. Appl Microbiol Biotechnol 102(24):10377–10391
Ou CY, Kwok S, Mitchell SW et al (1988) DNA amplification for direct detection of HIV-1 in DNA of peripheral blood mononuclear cells. Science 239(4837):295–297
Ou CY, Ciesielski CA, Myers G et al (1992) Molecular epidemiology of HIV transmission in a dental practice. Science 256(5060):1165–1171
Parachin NS, Schelin J, Norling B, Rådström P, Gorwa-Grauslund MF (2010) Flotation as a tool for indirect DNA extraction from soil. Appl Microbiol Biotechnol 87(5):1927–1933
Pechal JL, Crippen TL, Tarone AM et al (2013) Microbial community functional change during vertebrate carrion decomposition. PLoS One 8(11):e79035
Pechal JL, Crippen TL, Benbow ME et al (2014) The potential use of bacterial community succession in forensics as described by high throughput metagenomic sequencing. Int J Legal Med 128(1):193–205
Pechal JL, Schmidt CJ, Jordan HR, Benbow ME (2017) Frozen: thawing and its effect on the postmortem microbiome in two pediatric cases. J Forensic Sci 62(5):1399–1405
Powers JS, Montgomery RA, Adair EC et al (2009) Decomposition in tropical forests: a pan-tropical study of the effects of litter type, litter placement and mesofaunal exclusion across a precipitation gradient. J Ecol 97(4):801–811
Preston BD, Poiesz BJ, Loeb LA (1988) Fidelity of HIV-1 reverse transcriptase. Science 242(4882):1168–1171
Quaak FCA, Kuiper I (2011). Statistical data analysis of bacterial t-RFLP profiles in forensic soil comparisons. Forensic Sci Int 210:96–101
Rana AK, Manhas S (2018) The future of forensic biology. J Biomed 3:13–18
Ranjard L, Poly F, Lata JC et al (2001) Characterization of bacterial and fungal soil communities by automated ribosomal intergenic spacer analysis fingerprints: biological and methodological variability. Appl Environ Microbiol 67(10):4479–4487
Robbins KE, Lemey P, Pybus OG et al (2003) US human immunodeficiency virus type 1 epidemic: date of origin, population history, and characterization of early strains. J Virol 77(11):6359–6366
Rochelle P, Cragg B, Fry J et al (1994) Effect of sample handling on estimation of bacterial diversity in marine sediments by 16S rRNA gene sequence analysis. FEMS Microbiol Ecol 15(1–2):215–225
Rogers P, Whitby S, Dando M (1999) Biological warfare against crops. Sci Am 280(6):70–75
Rossodivita A, Visconti A, Saporito T et al (2019) Bioterrorism: toxins as potential biological weapons-an emerging global health threat. Int J Infect Dis 79:55
Roth RR, James WD (1988) Microbial ecology of the skin. Annu Rev Microbiol 42(1):441–464
Ruffell A (2010) Forensic pedology, forensic geology, forensic geoscience, geoforensics and soil forensics. Forensic Sci Int 202(1–3):9–12
Sala M, Wain-Hobson S (2000) Are RNA viruses adapting or merely changing? J Mol Evol 51(1):12–20
Santiago-Rodriguez TM, Cano RJ (2016) Soil microbial forensics. Microbiol Spectrum. 4(4). https://doi.org/10.1128/microbiolspec.EMF-0007-2015
Saraswat PK, Nirwan PS, Saraswat S et al (2008) Biodegradation of dead bodies including human cadavers and their safe disposal with reference to mortuary practice. J Indian Acad Forensic Med 30:273–280
Schoenen D, Schoenen H (2013) Adipocere formation—the result of insufficient microbial degradation. Forensic Sci Int 226(1–3):301e1–301e6
Schutzer SE, Budowle B, Atlas RM (2005) Biocrimes, microbial forensics, and the physician. PLoS Med 2(12):e337
Sender R, Fuchs S, Milo R (2016) Revised estimates for the number of human and bacteria cells in the body. PLoS Biol 14(8):e1002533
Seng P, Abat C, Rolain JM et al (2013) Identification of rare pathogenic bacteria in a clinical microbiology laboratory: impact of matrix-assisted laser desorption ionization–time of flight mass spectrometry. J Clin Microbiol 51(7):2182–2194
Sensabaugh G (2009) Microbial community profiling for the characterisation of soil evidence: forensic considerations. In: Ritz K, Dawson LA, Miller D (eds) Criminal and environmental soil forensics. Springer, London, p 49e60
Slonczewski JL, Foster JW (2013) Microbiology: an evolving science. W.W. Norton and Company, New York
Smart JK (1997) History of chemical and biological warfare: an American perspective. In: Medical aspects of chemical and biological warfare. Office of the Surgeon General, Washington, DC, pp 9–86
Smith JA (2011) Collection and preservation of microbial forensic samples. In: Microbial Forensics. Academic, London, pp 379–392
Stahringer SS, Clemente JC, Corley RP (2012) Nurture trumps nature in a longitudinal survey of salivary bacterial communities in twins from early adolescence to early adulthood. Genome Res 22(11):2146–2152
Stockdale EA, Brookes PC (2006) Detection and quantification of the soil microbial biomass–impacts on the management of agricultural soils. J Agric Sci 144(4):285–302
Stokes KL, Forbes SL, Tibbett M (2009) Freezing skeletal muscle tissue does not affect its decomposition in soil: evidence from temporal changes in tissue mass, microbial activity and soil chemistry based on excised samples. Forensic Sci Int 183(1–3):6–13
Svarovskaia ES, Xu H, Mbisa JL et al (2004) Human apolipoprotein B mRNA-editing enzyme-catalytic polypeptide-like 3G (APOBEC3G) is incorporated into HIV-1 virions through interactions with viral and nonviral RNAs. J Biol Chem 279(34):35822–35828
Taylor LH, Latham SM, Woolhouse ME (2001) Risk factors for human disease emergence. Philos Trans R Soc Lond Ser B Biol Sci 356(1411):983–989
Tibbett M, Carter DO (2009) Research in forensic taphonomy: a soil-based perspective. In: Criminal and environmental soil forensics. Springer, Dordrecht, pp 317–331
Timperman J (1972) The diagnosis of drowning. A review. Forensic Sci 1(4):397–409
Vass AA (2001) Beyond the grave-understanding human decomposition. Microbiol Today 28:190–193
Vass AA, Smith RR, Thompson CV (2004) Decompositional odor analysis database. J Forensic Sci 49(4):1–10
Wagner Mackenzie B, Waite DW, Taylor MW (2015) Evaluating variation in human gut microbiota profiles due to DNA extraction method and inter-subject differences. Front Microbiol 6:130
Weiss S, Carter DO, Metcalf JL et al (2016) Carcass mass has little influence on the structure of gravesoil microbial communities. Int J Legal Med 130(1):253–263
Wescott DJ (2018) Recent advances in forensic anthropology: decomposition research. Forensic Sci Res 3(4):278–293
Wilfinger WW, Mackey K, Chomczynski P (1997) Effect of pH and ionic strength on the spectrophotometric assessment of nucleic acid purity. BioTechniques 22(3):474–481
Wilson MJ, Weightman AJ, Wade WG (1997) Applications of molecular ecology in the characterization of uncultured microorganisms associated with human disease. Rev Med Microbiol 8(2):91–103
Wilson TM, Gregg DA, King DJ et al (2001) Agroterrorism, biological crimes, and biowarfare targeting animal agriculture: the clinical, pathologic, diagnostic, and epidemiologic features of some important animal diseases. Clin Lab Med 21(3):549–592
Woese CR, Olsen GJ, Ibba M, Söll D (2000) Aminoacyl-tRNA synthetases, the genetic code, and the evolutionary process. Microbiol Mol Biol Rev 64(1):202–236
Zala K (2007) Dirty science: soil forensics digs into new techniques. Science 318(5849):386–387
Zaura E, Keijser BJ, Huse SM et al (2009) Defining the healthy “core microbiome” of oral microbial communities. BMC Microbiol 9(1):259
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Aeri, R., Arora, J., Kapoor, P., Sharma, A.D. (2020). An Introduction to Microbial Forensics. In: Sharma, S., Sharma, N., Sharma, M. (eds) Microbial Diversity, Interventions and Scope. Springer, Singapore. https://doi.org/10.1007/978-981-15-4099-8_21
Download citation
DOI: https://doi.org/10.1007/978-981-15-4099-8_21
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-4098-1
Online ISBN: 978-981-15-4099-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)