Abstract
Primary Sjögren’s syndrome (SjS) is a chronic and systemic autoimmune epithelitis with predominant female incidence, which is characterized by exocrine gland dysfunction. Incompletely understood, the etiology of SjS is multi-factorial and evidence is growing to consider that epigenetic factors are playing a crucial role in its development. Independent from DNA sequence mutations, epigenetics is described as inheritable and reversible processes that modify gene expression. Epigenetic modifications reported in minor salivary gland and lymphocytes from SjS patients are related to (i) an abnormal DNA methylation process inducing in turn defective control of normally repressed genes involving such matters as autoantigens, retrotransposons, and the X chromosome in women; (ii) altered nucleosome positioning associated with autoantibody production; and (iii) altered control of microRNA. Results from epigenome-wide association studies have further revealed the importance of the interferon pathway in disease progression, the calcium signaling pathway for controlling fluid secretions, and a cell-specific cross talk with risk factors associated with SjS. Importantly, epigenetic modifications are reversible thus opening opportunities for therapeutic procedures in this currently incurable disease.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Primary Sjögren’s syndrome (SjS) is an autoimmune systemic disease characterized by lymphocytic infiltration of the exocrine glands with tropism for the salivary, lacrimal, upper respiratory, and vaginal glands of women (Brito-Zeron et al. 2016a). The clinical picture of SjS is based on evidence of xerostomia and xerophthalmia, which define a syndrome of dryness, and association with intense fatigue and widespread pain that leads to a profound alteration in the patients’ quality of life (Ramos-Casals et al. 2012). In one-third of patients, there are systemic manifestations, i.e., extra-glandular, which can affect the kidneys, liver, lungs, and thyroid.
The severity of the disease is generally associated with visceral abnormalities and the development of B lymphoma in 5% of patients (Brito-Zeron et al. 2016b; Nocturne and Mariette 2018). The prevalence of SjS ranges from 0.01 to 0.72% (Kabasakal et al. 2006; Maldini et al. 2014). In its primary form, SjS primarily affects women (9:1), with an average age of onset of about 50 years (Ramos-Casals et al. 2015). The secondary forms (50%), generally of lesser intensity, coexist with systemic lupus erythematosus (SLE), rheumatoid arthritis (RA), thyroiditis, and even primary biliary cirrhosis.
In addition to the fact that SjS is characterized by an autoimmune epithelitis, its physiopathology raises numerous questions (Brito-Zeron et al. 2016a; Sandhya et al. 2017). Based on data acquired from the analysis of the exocrine glands, in particular minor salivary glands (MSG), several steps were highlighted. First of all, activation of the epithelium, which leads to lymphocyte infiltration, consisting mainly of T cells and more particularly CD4+ T cells (Christodoulou et al. 2010; Verstappen et al. 2018). Then, and concomitantly with epithelial activation and disease progression, new cell populations appear in MSG such as follicular T cells, TH17 cells, dendritic (interferon producing) cells, and B lymphocytes which gradually become predominant and organize themselves into ectopic germinal centers. B lymphocyte hyperactivation is accompanied by local production of autoantibodies (auto-Ab) and, in particular, of sicca syndrome type A (SSA/Ro) or type B (SSB/La) auto-Ab.
At a peripheral level, the detection of anti-SSA/SSB auto-Ab is frequently associated with the detection of rheumatoid factor, hypergammaglobulinemia, and hypocomplementemia that reflects an active immunological profile (Capaldo et al. 2016). This hyperactivation is accompanied by abnormalities of peripheral B lymphocyte subpopulations, reflecting the attraction of memory B cells into tissues (Alonso et al. 2010; Cornec et al. 2014; Simonin et al. 2016; Renaudineau 2017). In the MSG, there is also an increase in the size of the glands and significant ultra-sonographic changes (Le Goff et al. 2017).
The etiology and pathogenesis of SjS are multi-factorial and consist of an aggregate of genetic predispositions with environmental factors (Renaudineau and Ballestar 2016). Recent data further support the important contribution of epigenetic mechanisms in SjS no longer solely to manage epithelial system development, but also as key regulators for controlling cell cycle, cell differentiation, immune system recruitment, and activation and, last but not least, to control the response to external factors such as various treatment modalities. From the cellular point of view, the definition of epigenetics has advanced and now includes adjustments in gene expression that do not entail modifications in the DNA sequence and are inheritable and reversible. This chapter summarizes implications of epigenetic modifications in SjS and their consequences.
2 Environmental, Genetic, and Epigenetic Factors
2.1 Environmental Factors
Due to the frequent presence of severe fatigue and arthralgia at diagnosis, a viral cause is suspected, notably Herpes family members such as Epstein–Barr virus (EBV) and human herpes virus 6 (HHV6) (Lucchesi et al. 2014). Several indirect arguments favor this direction, such as the evidence of a higher prevalence of EBV- and HHV6-specific IgG in this disease (Kivity et al. 2014). A cross-reactivity is described for these two viruses with the SSB/La protein due to molecular mimicry (Haaheim et al. 1996; Hajjar et al. 1995), and a correlation between anti-SSB/La auto-Ab and IgG levels against the early antigens of EBV was recently reported (Agmon-Levin et al. 2017). In this study, a correlation was also reported between anti-Ro/SSA auto-Ab and IgG/M directed against EBV, cytomegalovirus (CMV), and toxoplasma.
However, no direct evidence has been reported on the direct role of these viruses which are widely distributed in the general population, and several studies did not find this association. As a result, other viruses have been implicated, again without direct evidence, including HTLV1 (human T cell lymphotropic virus 1), hepatitis B and C viruses, retrovirus, and coxsackievirus (Hajjar et al. 1995; Wattiaux et al. 1995; Brauner et al. 2017; Nakamura et al. 2018). Another explanation is related to the capacity of SjS patients to have a higher immune response to infectious agents as observed with the H1N1 influenza vaccine (Brauner et al. 2017). An alteration of the microbiome is also reported in SjS with dysbiosis reported in the intestinal tract, and in ocular and oral flora that are suspected of influencing the immune system (De Luca and Shoenfeld 2018; Tsigalou et al. 2018).
Other factors associated with SjS include tobacco, vitamin D deficiency, ultraviolet radiation, and chemical agents whose exposure appears to be correlated with the appearance of SjS (Busche et al. 2015; Garcia-Carrasco et al. 2017). The demographic components of SjS reinforce the involvement of environmental factors with a higher reported prevalence of SjS in Nordic countries (Shapira et al. 2010) Another argument in support of this hypothesis is based on the study of homozygous twins for whom the concordance rate with SjS, which defines the genetic share of a disease, ranges between 15 and 25% (Brooks et al. 2010).
2.2 Genetic Factors
With the development of GWAS (genome-wide association study) projects, more than 40 risk factors have been associated with SjS but they have only a slight effect on the risk of developing SjS (Lessard et al. 2012; Konsta et al. 2014). First, there are risk factors involved in antigen presentation with certain regions of the HLA class I and II system or upstream with factors involved in the regulation and expression of HLA molecules. The presence of anti-SSA/SSB auto-Ab and labial salivary gland focus score is associated with HLA-DQA1 and HLA-DQB1 in Europeans, an association not confirmed in Asians, which contrasts with HLA-DPB1 that was observed in both ethnic groups (Taylor et al. 2017).
For the other risk factors associated with SjS, there are risk factors involved in innate and acquired immunity such as genes associated with interferon (IFN) signatures (irf5 [interferon regulatory factor 5], il12a [interleukin 12A], ncr3 [natural cytotoxicity triggering receptor 3], and stat4 [signal transducer and activator of transcription 4]), genes associated with T and B cell functions (tnfaip3 [TNF-alpha induced protein 3], tnip1 [TNFAIP3-interacting protein 1], cxcr5 [C-X-C chemokine receptor type 5], blk [B lymphocyte kinase], baff [B cell activating factor], ebf1 [early B cell factor 1], gtf2i [general transcription factor IIi], tnsff4 [Tumor Necrosis Factor Superfamily Member 4], lta [Lymphotoxin-α], and ccl11 [C-C motif chemokine 11]), and genes with other functions (htt [solute carrier family 6 member 4]) (Lessard et al. 2013; Burbelo et al. 2014; Konsta et al. 2015; Teos and Alevizos 2017). The mode of action of these SjS associated risk factors is complex because few of them have been attributed to the normal functions of the proteins. However, recent studies revealed that they are not randomly located but present in cell-specific and epigenetic regulatory zones to control transcription (Konsta et al. 2015). Consequently, a better understanding of the cross talk between risk factors and epigenetic factors to control cell-specific gene expression can lead to a better understanding of the pathophysiology of the disease.
2.3 Epigenetic Factors
Epigenetics plays an essential role in acting at key stages of differentiation and activation of the immune system. But there are actually several epigenetic mechanisms involved, such as post-translational histone modifications (to alter DNA compaction or decondensation), non-coding RNAs (which can modulate gene expression via sense/anti-sense interactions with other transcripts), and DNA methylation (modulates actions of transcription factors and gene repressors) (Fig. 11.1). This latter mechanism occurs by the transfer of a methyl group on the 5th carbon of the CpG dinucleotide cytosine of DNA (5-mCyt) under the action of DNA methyltransferases (DNMTs) with S-adenosylmethionine (SAM) being the key source of the methyl groups (Renaudineau and Youinou 2011; Brooks and Renaudineau 2015). Important modifications of these processes, specific for a given cell type, have been observed in pathology and in particular in cancers and autoimmune diseases (Brooks et al. 2010; Bagacean et al. 2017a, b). The first argument implicating epigenetic mechanisms in SjS derives from the observation that oral administration of DNA demethylating drugs such as hydralazine or isoniazid promotes sicca development in mice treated over several weeks together with immunological elements of an SLE-like disease (Bordron et al. 2018). From these initial experiments, it was also reported that the animal strain, age, and sex were important for the pathological process, and that this effect disappeared when the demethylating drug was withdrawn.
Next, both the Richardson and Zouali teams demonstrated that forcing DNA hypomethylation in both CD4+ T cells and CD19+ B cells promotes autoreactivity (Quddus et al. 1993; Mazari et al. 2007). Indeed, when CD4+ T cells or CD19+ B cells pretreated with DNA methyltransferase inhibitors are passively transferred into mice, the engrafted mice produce auto-Ab including anti-dsDNA Ab. One further step was to characterize in SLE patients the defective pathways that lead to decreased DNMT1 expression in both CD4+ T cells and CD19+ B cells, which includes the MAPk/Erk pathways (Gorelik et al. 2015). From the epigenetic point of view, the most important difference between SLE and SjS is related to the fact that epithelial cells, and to a lesser extent lymphocytes, have a defective DNA methylation process (Dantec et al. 2015).
3 Epigenetic Defects
3.1 DNA Methylation
3.1.1 Global Methylation Analysis
-
1.
Demethylation of DNA in SjS
Overall, DNA methylation acts on gene transcription either directly by preventing transcription factors from binding or indirectly by recruiting enzymes responsible for chromatin compaction such as histone deacetylases (HDAC) and histone methyltransferases (HMT). In mammals, chromatin is normally methylated and compacted and it is the regulatory and transcriptionally active zones that are demethylated. The DNA demethylation process is either passive during cell division or active and is in this case initiated by TET (ten-eleven translocation) oxidation enzymes (Bagacean et al. 2018). TETs oxidize 5-mCyt into 5-hydroxymethylcytosine (5-hmCyt) and subsequently, in a less efficient manner, into 5-formylcytosine (5-fCyt) and 5-carboxylcytosine (5-CaCyt) with the use of α-ketoglutarate (α-KG), molecular oxygen and iron as cofactors. The overall rate of 5mCyt found in mammals varies according to cell type from 70 to 80%, with low levels in immune system cells.
Analysis of histological sections from MSG and epithelial cells cultured and isolated from patients’ MSGs, as well as analysis of peripheral B and T lymphocyte populations (Table 11.1) and analysis of the overall DNA methylation state made it possible to highlight in SjS: (i) a significant reduction in overall 5-mCyt status in the MSG and which predominates in epithelial cells; (ii) that this defect was preserved in culture and that it was associated with suppression of expression of the methylation enzyme DNMT1 as reported in long term purified salivary gland epithelial cells (SGEC); and (iii) conversely, no difference in overall DNA methylation could be observed in B and T cells of patients’ and controls’ peripheral blood, pointing out that epigenetic changes preferentially affect the epithelial cells.
MSG 5-mCyt reduction in SjS and DNMT1/3b reductions in SjS patients with lymphoma were recently confirmed (Lagos et al. 2018; Mavragani et al. 2018), and Lagos et al. have further underlined an active DNA demethylation process based on the observation that 5hmCyt and TET2 levels are increased in epithelial cells from MSG sections.
-
2.
Involvement of lymphocytes and pro-inflammatory cytokines in the demethylation of DNA from epithelial cells of minor salivary glands
The presence of a significant lymphocyte infiltration appears to be associated with 5-mCyt reduction in MSG since the methylation level is inversely proportional to the Tarpley and focus scores which reflects the inflammatory state of MSG (Konsta et al. 2016b; Lagos et al. 2018). An inverse association with anti-SSB/La auto-Ab positivity is also reported in these two studies. On one hand, the role of B cells is suspected to interfere with the Erk/DNMT1 pathway leading to 5-mCyt reduction in the HSG (human salivary gland) cell line when this cell line is co-cultivated with B cells (Thabet et al. 2013). This process could be reversed as observed in MSG from patients that have recovered a normal DNA methylation level 4 months after receiving a treatment with rituximab, an anti-B lymphocyte immunotherapy (Thabet et al. 2013). On the other hand, HSG stimulation with the pro-inflammatory cytokines interferon (IFN)-γ and the tumor necrosis factor (TNF)-α promote TET2 overexpression that in turn increases 5-hmCyt and decreases 5-mCyt (Lagos et al. 2018).
3.1.2 Complete Methylome Analysis
-
1.
Methylation analysis of CpG units of peripheral blood cell DNA
Several teams have used the HM450K technology from Illumina to study DNA methylation across the genome in peripheral blood mononuclear cells (PBMCs) which include a mixture of T, B cells, and monocytes, or better by using purified T cells (total and naive) and B cells of patients with SjS (Altorok et al. 2014; Imgenberg-Kreuz et al. 2016; Miceli-Richard et al. 2016). Apart from the different methods of analysis, we can highlight several points. First, hypermethylated regions of DNA are found in HLA type I and II antigen presentation genes but also on certain genes regulated by interferon type I (IFI44L, IFITMI). Second, specific signaling pathways are found in T cells (solute-carrying proteins and the transcription factor RUN2X) and in B cells (B cell receptors, developmental genes). Third, the effect predominates in patients with anti-SSA/SSB auto-Ab and in B cells because the methylation changes are 50 times greater than in T cells (Miceli-Richard et al. 2016).
-
2.
Methylation analysis of CpG units of minor salivary gland DNA
Two teams analyzed DNA methylation using HM450K chips in MSG of patients with SjS (Cole et al. 2016; Imgenberg-Kreuz et al. 2016). However, the limitation of this approach, as reported for PBMC, is related to the heterogeneity of the samples since the glands studied include both acinar and tubular epithelial cells which are found in patients and controls, but also immune cells present in patients. Despite this limitation, the importance of the IFN pathway is reported, again with, in particular, the detection of a gene inducible by the IFN such as oas2, which is demethylated. The other differentially methylated regions concern psmd8 (26S proteasome non-ATPase regulatory subunit 8), tap1 (antigen peptide transporter 1), and microRNAs as well as a wide variety of genes involved in cell activation, antigen presentation and autoantigen production (Cole et al. 2016; Renaudineau and Ballestar 2016).
-
3.
Methylation analysis of CpG units of epithelial cells in minor salivary glands
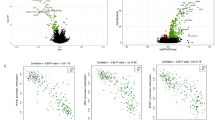
As previously indicated, the study of CpG patterns on MSG was carried out on a cellular mixture, and therefore does not fully reflect the impact of DNA modifications on MSG epithelial cells. This limitation was removed by using SGEC from MSG cultured for 3–4 weeks to obtain a pure cell population to use as the biological source (Charras et al. 2017). Significant differences were observed when comparing SjS patients with a control population using HM450K chips. These differences concern a large number of genes regulated by IFN (61% of genes). In addition, the calcium pathway (involved in salivary flow control) was demethylated while the Wnt pathway (involved in epithelial cell survival and differentiation) was methylated in this study (Fig. 11.2). From such analysis, the phosphatidyl inositol (PI3)-kinase pathway was also associated with hydroxychloroquine intake.
Implication of DNA methylation in the pathogenesis of Sjogren’s syndrome (Charras et al. 2017)
3.2 Histones
Histones are small globular proteins (11–15 kD) with flexible N-terminal tails that project from the nucleosome core. Histone N-terminal tails play an important role in controlling gene transcription and expression and this is regulated by post-translational modifications at lysine, arginine, and serine residues. Modifications at these residues can be acetylation, methylation, phosphorylation, ubiquitination/sumoylation, ADP ribosylation, deimination/citrullination, protein conjugation, or β-N-acetylglucosamination.
Using a chromatin immunoprecipitation (ChIP) approach a decrease in H4 acetylation was observed at the AQP5 (aquaporin 5) promoter and there was overexpression from the gene in human salivary gland acinar cells after treatment with TNF-α (Yamamura et al. 2012). Such an observation is in line with the report of Imgenberg-Kreuz et al. in which they investigated the distribution of hypomethylated, hypermethylated, and differentially methylated cytosines (DMC) in T and B cells. In both cell types, hypomethylated DMCs are located within areas of H3K4me1 and H3J27Ac, while it is in association with H3K26 that is reported for hypermethylated DMCs (Imgenberg-Kreuz et al. 2016). Interestingly, the main alteration in the salivary proteome of SjS patients is related to the abnormal presence of histones (Hall et al. 2017), probably through an accelerating apoptotic process, explaining why anti-histone auto-Ab are reported in SjS (Hu et al. 2011).
In addition, treating the SjS non-obese diabetic (NOD) mouse model with resveratrol, which enhances NAD(+)-dependent histone deacetylase activity through sirtuin 1, improves saliva secretion and expression of the anti-inflammatory cytokine IL-10 in salivary glands without affecting inflammatory cell infiltration (Inoue et al. 2016). SAHA (suberoylanilide hydroxamic acid, a specific histone deacetylase inhibitor) reduces inflammation in dry eye disease and this may have applications in SjS (Ratay et al. 2018). Such discrepancies may be explained in part by the fact that the two drugs are not acting on the same cell subset.
3.3 miRNA
Small non-coding and single-strand RNA of 19–22 nucleotides in length, microRNAs (miRNAs) adjust gene expression at the post-transcriptional level and are crucial in a wide array of physiological and pathological processes. In the nucleus, fundamental miRNA transcripts are generated through RNA polymerase II, and cleaved by an RNAse III enzyme, referred to as Drosha. After cleavage, miRNAs are transported to the cytoplasm via exportin 5 for processing using Dicer to generate mature miRNA duplexes. Duplexes are then separated into single strands at the core of the multiprotein RNA-induced silencing complex (RISC) which includes argonaute proteins. Most miRNAs bind to the 3′ untranslated vicinity (UTR) of the targeted mRNAs which leads to mRNA translation or repression or degradation (Zare-Shahabadi et al. 2013). Auto-Ab to the miRNA-binding protein argonaute 2 (Su antigen) enriched in the cytoplasmic GW/P bodies are described in 10–20% of patients with SjS and neurological diseases (Bhanji et al. 2007; Satoh et al. 2013).
miRNA expression profile analysis reveals distinct profiles when comparing MSG from control subjects and SjS patients. Alevizios et al. have also reported that miRNA upregulation is more important in the group with decreased salivary functions (Alevizos et al. 2011). In addition, let7b, miR16, miR181a, miR223, and miR483-5p levels are positively correlated with Ro52/TRIM21-mRNA in MSG, while in SGEC miR181a and miR200b-3p are negatively correlated with Ro52/TRIM21 and Ro60/TROVE2 mRNAs, respectively, whereas let7b, miR200b-5p, and miR223 are associated with La/SSB-mRNA (Gourzi et al. 2015).
With regards to PBMCs instead of PBMC’s subsets, a limited number of miRNAs, including miR-146a and miR-155, are reported when comparing SjS patients with sicca-complaining controls (Pauley et al. 2011; Peng et al. 2014; Shi et al. 2014; Gourzi et al. 2015). In contrast, when considering purified T and B cells from SjS patients, there are major differences corresponding to 21 miRNAs in T cells (9 upregulated and 12 downregulated) and 24 miRNAs in B cells (11 upregulated and 12 downregulated) (Wang-Renault et al. 2018). In this study, regulation through DNA methylation at promoters was excluded and differential expression patterns were observed according to the anti-SSA auto-Ab status. The most interesting pathways associated with differentially expressed miRNAs are related to the PI3K pathway (T and B cells), the transforming growth factor (TGF)-β pathway (T cells), and the Wnt pathway (B cells).
In monocytes from SjS patients, miRNAs are upregulated and they preferentially target the TGF-β signaling pathway and, to a lesser extent, the Janus kinase/signal transducer and activator of transcription (JAK-STAT) signaling cascades (Williams et al. 2016). The EBV-specific xeno-miRNA, EBV-miR-BART13-3p, can be transferred from SjS B cells, through exosomes, to SGEC (Gallo et al. 2016). This xeno-miRNA controls calcium signaling and salivary flux by targeting the stromal interacting molecule 1 (STIM1) (Mukherjee et al. 2017).
4 Epigenetic Reprogramming and Consequences
4.1 Methylation Modifications
4.1.1 Retrotransposons
More than 50% of our genome is composed of retrotransposons that correspond to DNA sequences that have the ability of multiplying to spread in the genome. Retrotransposons can be divided into different families, including Alu (10%), LINE (17%), and endogenous human retroviruses (HERV, 8%). In order to protect themselves from the action of these elements, they are regulated by DNA methylation.
Alu transcripts are found to be increased in SjS, and interferon type I allows Alu production as well as that of other pro-inflammatory cytokines (Mavragani and Crow 2010; Hung et al. 2015). Alu elements are also capable of binding to the SSA/Ro60 auto-Ag that can lead to the formation of an immune complex when associated with anti-Ro60 auto-Ab that is widely found in SjS and SLE. As a consequence this provides an explanation for the SSA/Ro60-associated RNA complex to initiate Toll-like receptors (TLR)-7/8 dependent pro-inflammatory cytokine release (Reed and Gordon 2016). Similar observations were made with LINE-1 which is upregulated in MSG from SjS patients, in relation to a defective DNA methylation process as recently reported by Mavragani et al. (2018). As a consequence, inappropriate LINE-1 expression in SjS is believed to contribute to the pathophysiology of SjS through the activation of the TLR-7/8-IFN-type I pathway as observed in plasmacytoid dendritic cells transfected with LINE-1 sequences (Balada et al. 2010; Mavragani et al. 2016).
In MSG (Table 11.2), HERV have been found to be overexpressed, such as HERV-K113, HERV-5, and the HERV-E family including clone 4-1 (Moyes et al. 2005; Le Dantec et al. 2012). For the most part, HERV genes have evolved to become non-functional due to deletions, presence of frame shifts or stop codons. However, a few copies have retained their functionality to generate viral proteins and to promote the expression of fusion transcripts with neighboring genes (Renaudineau et al. 2005; Le Dantec et al. 2015). For this reason, HERV elements are repressed by DNA methylation as demonstrated with HRES-1 (human T cell leukemia related endogenous retrovirus), inserted at the long arm of chromosome 1 at position 1q42 (Fali et al. 2014). When this control is lost, the HRES-1 Gag p38 auto-Ag is expressed and induces the production of anti-Gag p38 autoantibodies (Banki et al. 1992). Auto-Ab against HRES-1 Gag p38 is detected in 10% of patients with SjS versus 1.5% in healthy controls. Another HERV-E element overexpressed in SjS is HERV-E clone 4-1. HERV 4-1 p30 gag auto-Ab has been detected in 35% of SjS sufferers and is absent in healthy donors (Hishikawa et al. 1997).
4.1.2 Autoantigens
Demethylation of the SSB/La promoter is observed in patients with SjS, which generates overexpression of the transcript and protein (Konsta et al. 2016b). This effect is even more true in those SjS patients that express anti-SSB/La auto-Ab and can be reproduced by treating the HSG cell line with 5-azacytidine (5-Aza). Similar observations have been made with ICA1, another autoantigen (Charras et al. 2017).
4.1.3 Other Genes
Treatment with 5-Aza increases the expression of cytokeratin 19 in the HSG line (Konsta et al. 2016a) and also the aquaporin 5 gene (aqp5) with an increase in salivary flow in a human ductal salivary gland line (Motegi et al. 2005). Analysis by bisulfite sequencing of the aqp5 promoter shows hypermethylation of the CpG islets at the Sp1 transcription factor binding sites, sites that can be demethylated with 5-Aza, allowing the Sp1 transcription factor to bind to DNA and initiate transcription of this gene.
Other gene promoters have been analyzed, in particular the ire1α (inositol-requiring enzyme 1α), the xbp-1 (X box-binding protein 1 (XBP-1), and the dst (dystonin) genes (Gonzalez et al. 2011; Sepulveda et al. 2018). Decreased mRNA levels for IRE1α and XBP-1 can be explained in part by an IFN-dependent hypermethylation pathway controlling their promoters. When demethylated, the dst promoter leads to the low expression of an epithelial alternative splicing variant, an auto-Ag, the bp320 pemphigoid bullous antigen 1. In CD4+ T cells of patients with SjS, demethylation at CD70 (TNFSF7) promoter contributes to CD70 overexpression (Yin et al. 2010), while hypermethylation in the FOXP3 promoter leads to its repression (Yu et al. 2013).
4.1.4 X Chromosome in SjS
In women, in order to balance X-linked gene dosage, one of the two X chromosomes is inactivated in each somatic cell. This epigenetic control is suspected to be defective in SjS (Brooks and Renaudineau 2015). Several arguments support this assertion such as the predominant female sex bias observed in SjS and reports showing that trisomy X (47, XXX), a superb female phenotype (mosaic of XXXXX/XXXX/XXX/XX/XO), or Klinefelter’s syndrome (47, XXY) increase the chance of developing SjS (Harris et al. 2016; Liu et al. 2016; Sharma et al. 2017). Observing that the X-linked CD40 ligand (CD40L, Xq26.3) was overexpressed in CD4+ T cells from SjS females, Belkhir et al. failed to link CD40L expression levels with the DNA methylation status of its regulatory areas, in contrast to what is observed in SLE (Lu et al. 2007; Belkhir et al. 2014).
X chromosome inactivation (XCI) occurs early in female mammalian development to transcriptionally silence all but one of the X chromosomes in each cell through increased DNA methylation, thereby achieving dosage equivalency with the one X chromosome in males (XY). As a consequence, differences in X-linked gene expression between SjS and controls may underlie an abnormal control of genes following X chromosome inactivation (XCI) in SjS women with normal 46, XX genotype, as proposed first by Brooks et al. in response to disruption by a nearby nucleolus during stress (Brooks 2010, 2017) and validated in silico by Mougeot et al. (2018).
The X inactivation specific transcript (XIST), a lncRNA, along with LINE-1 genes in the X chromosome are involved in establishing XCI. However, XIST and LINE-1 sequences, two demethylation sensors, are overexpressed in SjS as well as the polycomb repressive complexes (PRC)2 genes EED and EZH1 that can be recruited by XIST to silence target genes. Altogether, these suggest an active but probably ineffective XCI process and difficulties in maintaining XCI during SjS, a defect that can be explained in part by the X chromosome nucleolus nexus hypothesis (Brooks 2017).
4.2 Histone Modifications in SjS
Imgenberg-Kreuz et al. demonstrated that several differentially methylated CpG (DMC) sites were hypomethylated and enriched in histone enhancers (H3K4me1 and H3K27ac) allowing access to the chromatin. On the other hand, hypermethylated DMCs in patients were underrepresented in enhancer regions (H3K4me3) but instead were enriched in the histone marker (H3K36) for actively transcribed genes (Imgenberg-Kreuz et al. 2016).
4.3 miRNA in SjS
4.3.1 miRNA in Minor Salivary Glands from SS Patients
miRNAs have been investigated in MSG revealing that miRNA expression is differentially expressed between SjS patients and controls, and that miRNA is involved in the control of salivation through neurologic pathways. Downregulation of the miRNA let-7b in MSG from SjS patients is also suspected of contributing to the lack of transcriptional control of the auto-Ag SSA/Ro and SSB/La.
4.3.2 miRNA in PBMC and Exosomes from SjS Patients
The two main miRNAs associated with SjS, miR146a, and miR155, are upregulated in response to the adaptive immune response when testing PBMC from SjS patients and from SS-prone mouse models (Pauley et al. 2011). Authors have shown, in SjS patients, that miR146a expands prior to disease onset in PBMCs, and at a more advanced stage in MSG from SjS patients. In addition, detected in PBMC and MSG, miRNAs are also present in exosomes which are microvesicles secreted by a large variety of cells including lymphocytes.
Mir146a is important for control of the phagocytic process and to repress inflammatory cytokine production in human monocytic THP1 cells. Mir146a is activated through the transcription factor NF-kappa B that controls the TLR/INF pathway through TNF-associated component 6 (TRAF6), IL-1 receptor-associated kinase (IRAK1), STAT1, and IRF5. Additionally, Zilahi et al. have measured the expression of miR146a and miR146b, and their target genes IRAK1, IRAK4, TRAF6 in PBMCs of patients with SjS and from controls (Zilahi et al. 2012). By quantitative RT-PCR they identified miR146a/b and the gene of TRAF6, as being overexpressed in SjS patients, whereas the expression of IRAK1 was significantly decreased. They proposed that the TRAF6 gene contributes to the increased activation of the NF-κB pathway by the involvement of PKCξ present typically in the disease, and that possibly the TRAF6 gene is a new biomarker of SjS.
Experiments for miR155 have shown an influence on the response of toll-like receptors (TLRs) and interleukin-1 receptors (TIRs) that can have an additional effect on the immune response. FoxP3 transcription factor, which is detected in a subset of T cells infiltrating SjS MSG, has been shown to result in miR155 expression (Wang-Renault et al. 2018).
The expression of B cell activating factor (BAFF) is increased in B cells from SjS patients and such expression is inversely correlated with miR-30b expression The utilization of an antagomir (miRNA inhibitor) for miR-30 increases BAFF expression after transfection as observed in the THP-1 cell line (Wang-Renault et al. 2018).
5 Genetic Risk Factors Associated with SjS and Related to Epigenetic Factors
5.1 Methylation Modification
Recent data demonstrate highly significant correlations between DNA methylation modifications and the most important risk factors associated with SjS (Table 11.3). This was described for the HLA region and IRF5-TNPO3 locus with methylation quantitative trait loci (metQTL) by Imgenberg-Kreuz et al. using PBMC (Imgenberg-Kreuz et al. 2016, 2018). The same observation between DNA methylation and genetic risk loci was reported by Miceli-Richard et al. using peripheral B cells from SjS patients for HLA-DRA, HLA-DQB1, HLA-DQA1, HLA-DPB1, IRF5, CXCR5, BLK, PRDM1, ITSN2, GTF2I, and COL11A2 (Miceli-Richard et al. 2016). LTA and GSTM1 were reported by Altorok et al. in CD4+ T cells, and CXCR5, BLK, LTA, and BAFF in MSGs (Altorok et al. 2014).
In SGEC, we have reported that seven SjS genetic risk factors presented at least one differentially methylated site: CXCR5, GTF2I, ICA1, NRLP3, SLC25A10, TNF, and MBL2 (Charras et al. 2017). However, it should be kept in mind that the demonstration of correlations may be difficult to establish as reported by Gestermann et al. who have tried to establish a link between the CpG polymorphism in the promoter of irf5 and DNA methylation by comparing CD4+ T cells, B lymphocytes, and monocytes from 19 SjS patients and 24 healthy controls (Gestermann et al. 2012).
5.2 Histone Modifications
In general, single nucleotide polymorphisms (SNP) related to SjS risk factors are enriched at 29.2% in promoters (RNA polymerase 2A site), at 56.9% in enhancers (H3K27Ac, H3K36me3, and H3K27me3), and at 6.9% in insulators (CTCF binding) (Konsta et al. 2015). In addition, we have also reported a cell-specific effect between SjS risk factors and the histone markers in monocyte enhancers (H3K36me3) and in B cell promoters (H3K4me2, H3K4me3, and H3K9Ac) and enhancer (H3K36me3) cells.
5.3 miRNA Modification
Targets of miRNA may include SjS risk factors such as BAFF, which is overexpressed in B cells and whose 3′ UTR is targeted by hsa-miR-30b-5p (Wang-Renault et al. 2018). Another example is TRIM21 as reported in Table 11.2 (Gourzi et al. 2015; Yang et al. 2016).
6 Conclusions
Epigenetic research conducted on SjS in the last decade has contributed to better understanding of this complex disease. More breakthroughs are expected in the near future. Future research may be focused on selecting pure cell subsets for analysis, understanding the mechanisms that control epigenetic defects, and coupling epigenetic analysis with other OMIC approaches (RNA-Seq, GWAS, proteomic). Finally, epigenetic research provides us the opportunity to develop new drugs in order to prevent/cure not only SjS but also lymphoma associated with SjS.
References
Agmon-Levin N, Dagan A, Peri Y, Anaya JM, Selmi C, Tincani A, Bizzaro N, Stojanovich L, Damoiseaux J, Cohen Tervaert JW, Mosca M, Cervera R, Shoenfeld Y (2017) The interaction between anti-Ro/SSA and anti-La/SSB autoantibodies and anti-infectious antibodies in a wide spectrum of auto-immune diseases: another angle of the autoimmune mosaic. Clin Exp Rheumatol 35(6):929–935
Alevizos I, Alexander S, Turner RJ, Illei GG (2011) MicroRNA expression profiles as biomarkers of minor salivary gland inflammation and dysfunction in Sjogren’s syndrome. Arthritis Rheum 63(2):535–544
Alonso R, Buors C, Le Dantec C, Hillion S, Pers JO, Saraux A, Montero E, Marianowski R, Loisel S, Devauchelle V, Youinou P, Renaudineau Y (2010) Aberrant expression of CD6 on B-cell subsets from patients with Sjogren’s syndrome. J Autoimmun 35(4):336–341
Altorok N, Coit P, Hughes T, Koelsch KA, Stone DU, Rasmussen A, Radfar L, Scofield RH, Sivils KL, Farris AD, Sawalha AH (2014) Genome-wide DNA methylation patterns in naive CD4+ T cells from patients with primary Sjogren’s syndrome. Arthritis Rheumatol 66(3):731–739
Bagacean C, Le Dantec C, Berthou C, Tempescul A, Saad H, Bordron A, Zdrenghea M, Cristea V, Douet-Guilbert N, Renaudineau Y (2017a) Combining cytogenetic and epigenetic approaches in chronic lymphocytic leukemia improves prognosis prediction for patients with isolated 13q deletion. Clin Epigenetics 9:122
Bagacean C, Tempescul A, Le Dantec C, Bordron A, Mohr A, Saad H, Olivier V, Zdrenghea M, Cristea V, Cartron PF, Douet-Guilbert N, Berthou C, Renaudineau Y (2017b) Alterations in DNA methylation/demethylation intermediates predict clinical outcome in chronic lymphocytic leukemia. Oncotarget 8(39):65699–65716
Bagacean C, Zdrenghea M, Dantec CL, Tempescul A, Berthou C, Renaudineau Y (2018) DNA demethylation marks in chronic lymphocytic leukemia: it is time to let the cat out of the bag. Future Sci OA 4(2):FSO265
Balada E, Vilardell-Tarres M, Ordi-Ros J (2010) Implication of human endogenous retroviruses in the development of autoimmune diseases. Int Rev Immunol 29(4):351–370
Banki K, Maceda J, Hurley E, Ablonczy E, Mattson DH, Szegedy L, Hung C, Perl A (1992) Human T-cell lymphotropic virus (HTLV)-related endogenous sequence, HRES-1, encodes a 28-kDa protein: a possible autoantigen for HTLV-I gag-reactive autoantibodies. Proc Natl Acad Sci USA 89(5):1939–1943
Belkhir R, Gestermann N, Koutero M, Seror R, Tost J, Mariette X, Miceli-Richard C (2014) Upregulation of membrane-bound CD40L on CD4+ T cells in women with primary Sjogren’s syndrome. Scand J Immunol 79(1):37–42
Bhanji RA, Eystathioy T, Chan EK, Bloch DB, Fritzler MJ (2007) Clinical and serological features of patients with autoantibodies to GW/P bodies. Clin Immunol 125(3):247–256
Bordron A, Charras A, Le Dantec C, Renaudineau Y (2018) Influence of epigenetic in Sjogren’s syndrome. Rev Med Interne 39(5):346–351
Brauner S, Folkersen L, Kvarnstrom M, Meisgen S, Petersen S, Franzen-Malmros M, Mofors J, Brokstad KA, Klareskog L, Jonsson R, Westerberg LS, Trollmo C, Malmstrom V, Ambrosi A, Kuchroo VK, Nordmark G, Wahren-Herlenius M (2017) H1N1 vaccination in Sjogren’s syndrome triggers polyclonal B cell activation and promotes autoantibody production. Ann Rheum Dis 76(10):1755–1763
Brito-Zeron P, Baldini C, Bootsma H, Bowman SJ, Jonsson R, Mariette X, Sivils K, Theander E, Tzioufas A, Ramos-Casals M (2016a) Sjogren syndrome. Nat Rev Dis Primers 2:16047
Brito-Zeron P, Kostov B, Solans R, Fraile C, Suarez-Cuervo C, Casanovas A, Rascon FJ, Qanneta R, Perez-Alvarez R, Ripoll M, Akasbi M, Pinilla B, Bosch JA, Nava-Mateos J, Diaz-Lopez B, Morera-Morales ML, Gheitasi H, Retamozo S, Ramos-Casals M, SS Study Group, Autoimmune Diseases Study Group (GEAS), Spanish Society of Internal Medicine (2016b) Systemic activity and mortality in primary Sjogren syndrome: predicting survival using the EULAR-SS disease activity index (ESSDAI) in 1045 patients. Ann Rheum Dis 75(2):348–355
Brooks WH (2010) X chromosome inactivation and autoimmunity. Clin Rev Allergy Immunol 39(1):20–29
Brooks WH (2017) Viral impact in autoimmune diseases: expanding the “X Chromosome-Nucleolus Nexus” hypothesis. Front Immunol 8:1657
Brooks WH, Renaudineau Y (2015) Epigenetics and autoimmune diseases: the X chromosome-nucleolus nexus. Front Genet 6:22
Brooks WH, Le Dantec C, Pers JO, Youinou P, Renaudineau Y (2010) Epigenetics and autoimmunity. J Autoimmun 34(3):J207–J219
Burbelo PD, Ambatipudi K, Alevizos I (2014) Genome-wide association studies in Sjogren’s syndrome: what do the genes tell us about disease pathogenesis?” Autoimmun Rev 13(7):756–761
Busche S, Shao X, Caron M, Kwan T, Allum F, Cheung WA, Ge B, Westfall S, Simon MM, Multiple Tissue Human Expression Resource, Barrett A, Bell JT, McCarthy MI, Deloukas P, Blanchette M, Bourque G, Spector TD, Lathrop M, Pastinen T, Grundberg E (2015) Population whole-genome bisulfite sequencing across two tissues highlights the environment as the principal source of human methylome variation. Genome Biol 16(1):290
Capaldo C, Carvajal Alegria G, Cornec D, Jousse-Joulin S, Devauchelle-Pensec V, Renaudineau Y (2016) The active immunological profile in patients with primary Sjogren’s syndrome is restricted to typically encountered autoantibodies. Clin Exp Rheumatol 34(4):722
Charras A, Konsta OD, Le Dantec C, Bagacean C, Kapsogeorgou EK, Tzioufas AG, Pers JO, Bordron A, Renaudineau Y (2017) Cell-specific epigenome-wide DNA methylation profile in long-term cultured minor salivary gland epithelial cells from patients with Sjogren’s syndrome. Ann Rheum Dis 76(3):625–628
Christodoulou MI, Kapsogeorgou EK, Moutsopoulos HM (2010) Characteristics of the minor salivary gland infiltrates in Sjogren’s syndrome. J Autoimmun 34(4):400–407
Cole MB, Quach H, Quach D, Baker A, Taylor KE, Barcellos LF, Criswell LA (2016) Epigenetic signatures of salivary gland inflammation in Sjogren’s syndrome. Arthritis Rheumatol 68(12):2936–2944
Cornec D, Saraux A, Cochener B, Pers JO, Jousse-Joulin S, Renaudineau Y, Marhadour T, Devauchelle-Pensec V (2014) Level of agreement between 2002 American-European Consensus Group and 2012 American College of Rheumatology classification criteria for Sjogren’s syndrome and reasons for discrepancies. Arthritis Res Ther 16(2):R74
Dantec L, Charras A, Brooks W, Renaudineau Y (2015) Similarities and differences of epigenetic mechanisms in lupus and Sjögren’s syndrome. Lupus Open Access 1:e101
De Luca F, Shoenfeld Y (2018) The microbiome in autoimmune diseases. Clin Exp Immunol 195(1):74–85
Fali T, Le Dantec C, Thabet Y, Jousse S, Hanrotel C, Youinou P, Brooks WH, Perl A, Renaudineau Y (2014) DNA methylation modulates HRES1/p28 expression in B cells from patients with lupus. Autoimmunity 47(4):265–271
Gallo A, Jang SI, Ong HL, Perez P, Tandon M, Ambudkar I, Illei G, Alevizos I (2016) Targeting the Ca(2+) sensor STIM1 by exosomal transfer of Ebv-miR-BART13-3p is associated with Sjogren’s syndrome. EBioMedicine 10:216–226
Garcia-Carrasco M, Jimenez-Herrera EA, Galvez-Romero JL, de Lara LV, Mendoza-Pinto C, Etchegaray-Morales I, Munguia-Realpozo P, Ruiz-Arguelles A, Jose R, Vera-Recabarren M, Cervera R (2017) Vitamin D and Sjogren syndrome. Autoimmun Rev 16(6):587–593
Gestermann N, Koutero M, Belkhir R, Tost J, Mariette X, Miceli-Richard C (2012) Methylation profile of the promoter region of IRF5 in primary Sjogren’s syndrome. Eur Cytokine Netw 23(4):166–172
Gonzalez S, Aguilera S, Alliende C, Urzua U, Quest AF, Herrera L, Molina C, Hermoso M, Ewert P, Brito M, Romo R, Leyton C, Perez P, Gonzalez MJ (2011) Alterations in type I hemidesmosome components suggestive of epigenetic control in the salivary glands of patients with Sjogren’s syndrome. Arthritis Rheum 63(4):1106–1115
Gorelik G, Sawalha AH, Patel D, Johnson K, Richardson B (2015) T cell PKCdelta kinase inactivation induces lupus-like autoimmunity in mice. Clin Immunol 158(2):193–203
Gourzi VC, Kapsogeorgou EK, Kyriakidis NC, Tzioufas AG (2015) Study of microRNAs (miRNAs) that are predicted to target the autoantigens Ro/SSA and La/SSB in primary Sjogren’s syndrome. Clin Exp Immunol 182(1):14–22
Haaheim LR, Halse AK, Kvakestad R, Stern B, Normann O, Jonsson R (1996) Serum antibodies from patients with primary Sjogren’s syndrome and systemic lupus erythematosus recognize multiple epitopes on the La(SS-B) autoantigen resembling viral protein sequences. Scand J Immunol 43(1):115–121
Hajjar C, Sainte-Foie S, Savin J, Lacave J, Berlet F, Teron-Aboud B, Batelier L, Guillemin B (1995) HTLV1 infection and Sicca syndrome. J Fr Ophtalmol 18(10):597–602
Hall SC, Hassis ME, Williams KE, Albertolle ME, Prakobphol A, Dykstra AB, Laurance M, Ona K, Niles RK, Prasad N, Gormley M, Shiboski C, Criswell LA, Witkowska HE, Fisher SJ (2017) Alterations in the salivary proteome and N-glycome of Sjogren’s syndrome patients. J Proteome Res 16(4):1693–1705
Harris VM, Sharma R, Cavett J, Kurien BT, Liu K, Koelsch KA, Rasmussen A, Radfar L, Lewis D, Stone DU, Kaufman CE, Li S, Segal B, Wallace DJ, Weisman MH, Venuturupalli S, Kelly JA, Alarcon-Riquelme ME, Pons-Estel B, Jonsson R, Lu X, Gottenberg JE, Anaya JM, Cunninghame-Graham DS, Huang AJW, Brennan MT, Hughes P, Alevizos I, Miceli-Richard C, Keystone EC, Bykerk VP, Hirschfield G, Xie G, Ng WF, Nordmark G, Bucher SM, Eriksson P, Omdal R, Rhodus NL, Rischmueller M, Rohrer M, Wahren-Herlenius M, Witte T, Mariette X, Lessard CJ, Harley JB, Sivils KL, Scofield RH (2016) Klinefelter’s syndrome (47, XXY) is in excess among men with Sjogren’s syndrome. Clin Immunol 168:25–29
Hishikawa T, Ogasawara H, Kaneko H, Shirasawa T, Matsuura Y, Sekigawa I, Takasaki Y, Hashimoto H, Hirose S, Handa S, Nagasawa R, Maruyama N (1997) Detection of antibodies to a recombinant gag protein derived from human endogenous retrovirus clone 4-1 in autoimmune diseases. Viral Immunol 10(3):137–147
Housekeeping Genes. http://www.cgen.com/supp_info/Housekeeping_genes.html
Hu S, Vissink A, Arellano M, Roozendaal C, Zhou H, Kallenberg CG, Wong DT (2011) Identification of autoantibody biomarkers for primary Sjogren’s syndrome using protein microarrays. Proteomics 11(8):1499–1507
Hung T, Pratt GA, Sundararaman B, Townsend MJ, Chaivorapol C, Bhangale T, Graham RR, Ortmann W, Criswell LA, Yeo GW, Behrens TW (2015) The Ro60 autoantigen binds endogenous retroelements and regulates inflammatory gene expression. Science 350(6259):455–459
Imgenberg-Kreuz J, Sandling JK, Almlof JC, Nordlund J, Signer L, Norheim KB, Omdal R, Ronnblom L, Eloranta ML, Syvanen AC, Nordmark G (2016) Genome-wide DNA methylation analysis in multiple tissues in primary Sjogren’s syndrome reveals regulatory effects at interferon-induced genes. Ann Rheum Dis 75(11):2029–2036
Imgenberg-Kreuz J, Sandling JK, Nordmark G (2018) Epigenetic alterations in primary Sjogren’s syndrome—an overview. Clin Immunol 196:12–20
Inoue H, Kishimoto A, Ushikoshi-Nakayama R, Hasaka A, Takahashi A, Ryo K, Muramatsu T, Ide F, Mishima K, Saito I (2016) Resveratrol improves salivary dysfunction in a non-obese diabetic (NOD) mouse model of Sjogren’s syndrome. J Clin Biochem Nutr 59(2):107–112
Kabasakal Y, Kitapcioglu G, Turk T, Oder G, Durusoy R, Mete N, Egrilmez S, Akalin T (2006) The prevalence of Sjogren’s syndrome in adult women. Scand J Rheumatol 35(5):379–383
Kivity S, Arango MT, Ehrenfeld M, Tehori O, Shoenfeld Y, Anaya JM, Agmon-Levin N (2014) Infection and autoimmunity in Sjogren’s syndrome: a clinical study and comprehensive review. J Autoimmun 51:17–22
Konsta OD, Thabet Y, Le Dantec C, Brooks WH, Tzioufas AG, Pers JO, Renaudineau Y (2014) The contribution of epigenetics in Sjogren’s syndrome. Front Genet 5:71
Konsta OD, Le Dantec C, Charras A, Brooks WH, Arleevskaya MI, Bordron A, Renaudineau Y (2015) An in silico approach reveals associations between genetic and epigenetic factors within regulatory elements in B cells from primary Sjogren’s syndrome patients. Front Immunol 6:437
Konsta OD, Charras A, Le Dantec C, Kapsogeorgeou E, Bordron A, Brooks WH, Tzioufas AG, Pers JO, Renaudineau Y (2016a) Epigenetic modifications in salivary glands from patients with Sjogren’s syndrome affect cytokeratin 19 expression. Bull Group Int Rech Sci Stomatol Odontol 53(1):e01
Konsta OD, Le Dantec C, Charras A, Cornec D, Kapsogeorgou EK, Tzioufas AG, Pers JO, Renaudineau Y (2016b) Defective DNA methylation in salivary gland epithelial acini from patients with Sjogren’s syndrome is associated with SSB gene expression, anti-SSB/LA detection, and lymphocyte infiltration. J Autoimmun 68:30–38
Lagos C, Carvajal P, Castro I, Jara D, Gonzalez S, Aguilera S, Barrera MJ, Quest AFG, Bahamondes V, Molina C, Urzua U, Hermoso MA, Leyton C, Gonzalez MJ (2018) Association of high 5-hydroxymethylcytosine levels with ten eleven translocation 2 overexpression and inflammation in Sjogren’s syndrome patients. Clin Immunol 196:85–96
Le Dantec C, Vallet S, Brooks WH, Renaudineau Y (2015) Human endogenous retrovirus group E and its involvement in diseases. Viruses 7(3):1238–1257
Le Dantec C, Varin MM, Brooks WH, Pers JO, Youinou P, Renaudineau Y (2012) Epigenetics and Sjogren’s syndrome. Curr Pharm Biotechnol 13(10):2046–2053
Le Goff M, Cornec D, Jousse-Joulin S, Guellec D, Costa S, Marhadour T, Le Berre R, Genestet S, Cochener B, Boisrame-Gastrin S, Renaudineau Y, Pers JO, Saraux A, Devauchelle-Pensec V (2017) Comparison of 2002 AECG and 2016 ACR/EULAR classification criteria and added value of salivary gland ultrasonography in a patient cohort with suspected primary Sjogren’s syndrome. Arthritis Res Ther 19(1):269
Lessard CJ, Ice JA, Adrianto I, Wiley GB, Kelly JA, Gaffney PM, Montgomery CG, Moser KL (2012) The genomics of autoimmune disease in the era of genome-wide association studies and beyond. Autoimmun Rev 11(4):267–275
Lessard CJ, Li H, Adrianto I, Ice JA, Rasmussen A, Grundahl KM, Kelly JA, Dozmorov MG, Miceli-Richard C, Bowman S, Lester S, Eriksson P, Eloranta ML, Brun JG, Goransson LG, Harboe E, Guthridge JM, Kaufman KM, Kvarnstrom M, Jazebi H, Cunninghame Graham DS, Grandits ME, Nazmul-Hossain AN, Patel K, Adler AJ, Maier-Moore JS, Farris AD, Brennan MT, Lessard JA, Chodosh J, Gopalakrishnan R, Hefner KS, Houston GD, Huang AJ, Hughes PJ, Lewis DM, Radfar L, Rohrer MD, Stone DU, Wren JD, Vyse TJ, Gaffney PM, James JA, Omdal R, Wahren-Herlenius M, Illei GG, Witte T, Jonsson R, Rischmueller M, Ronnblom L, Nordmark G, Ng WF, UK Primary Sjögren’s Syndrome Registry, Mariette X, Anaya JM, Rhodus NL, Segal BM, Scofield RH, Montgomery CG, Harley JB, Sivils KL (2013) Variants at multiple loci implicated in both innate and adaptive immune responses are associated with Sjogren’s syndrome. Nat Genet 45(11):1284–1292
Liu K, Kurien BT, Zimmerman SL, Kaufman KM, Taft DH, Kottyan LC, Lazaro S, Weaver CA, Ice JA, Adler AJ, Chodosh J, Radfar L, Rasmussen A, Stone DU, Lewis DM, Li S, Koelsch KA, Igoe A, Talsania M, Kumar J, Maier-Moore JS, Harris VM, Gopalakrishnan R, Jonsson R, Lessard JA, Lu X, Gottenberg JE, Anaya JM, Cunninghame-Graham DS, Huang AJW, Brennan MT, Hughes P, Illei GG, Miceli-Richard C, Keystone EC, Bykerk VP, Hirschfield G, Xie G, Ng WF, Nordmark G, Eriksson P, Omdal R, Rhodus NL, Rischmueller M, Rohrer M, Segal BM, Vyse TJ, Wahren-Herlenius M, Witte T, Pons-Estel B, Alarcon-Riquelme ME, Guthridge JM, James JA, Lessard CJ, Kelly JA, Thompson SD, Gaffney PM, Montgomery CG, Edberg JC, Kimberly RP, Alarcon GS, Langefeld CL, Gilkeson GS, Kamen DL, Tsao BP, McCune WJ, Salmon JE, Merrill JT, Weisman MH, Wallace DJ, Utset TO, Bottinger EP, Amos CI, Siminovitch KA, Mariette X, Sivils KL, Harley JB, Scofield RH (2016) X chromosome dose and sex bias in autoimmune diseases: increased prevalence of 47, XXX in systemic lupus erythematosus and Sjogren’s syndrome. Arthritis Rheumatol 68(5):1290–1300
Lu Q, Wu A, Tesmer L, Ray D, Yousif N, Richardson B (2007) Demethylation of CD40LG on the inactive X in T cells from women with lupus. J Immunol 179(9):6352–6358
Lucchesi D, Pitzalis C, Bombardieri M (2014) EBV and other viruses as triggers of tertiary lymphoid structures in primary Sjogren’s syndrome. Expert Rev Clin Immunol 10(4):445–455
Maldini C, Seror R, Fain O, Dhote R, Amoura Z, De Bandt M, Delassus JL, Falgarone G, Guillevin L, Le Guern V, Lhote F, Meyer O, Ramanoelina J, Sacre K, Uzunhan Y, Leroux JL, Mariette X, Mahr A (2014) Epidemiology of primary Sjogren’s syndrome in a French multiracial/multiethnic area. Arthritis Care Res (Hoboken) 66(3):454–463
Martini D, Gallo A, Vella S, Sernissi F, Cecchettini A, Luciano N, Polizzi E, Conaldi PG, Mosca M, Baldini C (2017) Cystatin S-a candidate biomarker for severity of submandibular gland involvement in Sjogren’s syndrome. Rheumatology (Oxford) 56(6):1031–1038
Mavragani CP, Crow MK (2010) Activation of the type I interferon pathway in primary Sjogren’s syndrome. J Autoimmun 35(3):225–231
Mavragani CP, Nezos A, Sagalovskiy I, Seshan S, Kirou KA, Crow MK (2018) Defective regulation of L1 endogenous retroelements in primary Sjogren’s syndrome and systemic lupus erythematosus: role of methylating enzymes. J Autoimmun 88:75–82
Mavragani CP, Sagalovskiy I, Guo Q, Nezos A, Kapsogeorgou EK, Lu P, Liang Zhou J, Kirou KA, Seshan SV, Moutsopoulos HM, Crow MK (2016) Expression of long interspersed nuclear element 1 retroelements and induction of type i interferon in patients with systemic autoimmune disease. Arthritis Rheumatol 68(11):2686–2696
Mazari L, Ouarzane M, Zouali M (2007) Subversion of B lymphocyte tolerance by hydralazine, a potential mechanism for drug-induced lupus. Proc Natl Acad Sci USA 104(15):6317–6322
Miceli-Richard C, Wang-Renault SF, Boudaoud S, Busato F, Lallemand C, Bethune K, Belkhir R, Nocturne G, Mariette X, Tost J (2016) Overlap between differentially methylated DNA regions in blood B lymphocytes and genetic at-risk loci in primary Sjogren’s syndrome. Ann Rheum Dis 75(5):933–940
Motegi K, Azuma M, Tamatani T, Ashida Y, Sato M (2005) Expression of aquaporin-5 in and fluid secretion from immortalized human salivary gland ductal cells by treatment with 5-aza-2′-deoxycytidine: a possibility for improvement of xerostomia in patients with Sjogren’s syndrome. Lab Invest 85(3):342–353
Mougeot JL, Noll BD, Bahrani Mougeot FK (2018) Sjogren’s syndrome X-chromosome dose effect: an epigenetic perspective. Oral Dis 25(2):372–384
Moyes DL, Martin A, Sawcer S, Temperton N, Worthington J, Griffiths DJ, Venables PJ (2005) The distribution of the endogenous retroviruses HERV-K113 and HERV-K115 in health and disease. Genomics 86(3):337–341
Mukherjee S, Karolak A, Debant M, Buscaglia P, Renaudineau Y, Mignen O, Guida WC, Brooks WH (2017) Molecular dynamics simulations of membrane-bound STIM1 to investigate conformational changes during STIM1 activation upon calcium release. J Chem Inf Model 57(2):335–344
Nakamura H, Hasegawa H, Sasaki D, Takatani A, Shimizu T, Kurushima S, Horai Y, Nakashima Y, Nakamura T, Fukuoka J, Kawakami A (2018) Detection of human T lymphotropic virus type-I bZIP factor and tax in the salivary glands of Sjogren’s syndrome patients. Clin Exp Rheumatol 112(3):51–60
Nocturne G, Mariette X (2018) B cells in the pathogenesis of primary Sjogren syndrome. Nat Rev Rheumatol 14(3):133–145
Pauley KM, Stewart CM, Gauna AE, Dupre LC, Kuklani R, Chan AL, Pauley BA, Reeves WH, Chan EK, Cha S (2011) Altered miR-146a expression in Sjogren’s syndrome and its functional role in innate immunity. Eur J Immunol 41(7):2029–2039
Peng L, Ma W, Yi F, Yang YJ, Lin W, Chen H, Zhang X, Zhang LH, Zhang F, Du Q (2014) MicroRNA profiling in Chinese patients with primary Sjogren syndrome reveals elevated miRNA-181a in peripheral blood mononuclear cells. J Rheumatol 41(11):2208–2213
Quddus J, Johnson KJ, Gavalchin J, Amento EP, Chrisp CE, Yung RL, Richardson BC (1993) Treating activated CD4+ T cells with either of two distinct DNA methyltransferase inhibitors, 5-azacytidine or procainamide, is sufficient to cause a lupus-like disease in syngeneic mice. J Clin Invest 92(1):38–53
Ramos-Casals M, Brito-Zeron P, Siso-Almirall A, Bosch X (2012) Primary Sjogren syndrome. BMJ 344:e3821
Ramos-Casals M, Brito-Zeron P, Kostov B, Siso-Almirall A, Bosch X, Buss D, Trilla A, Stone JH, Khamashta MA, Shoenfeld Y (2015) Google-driven search for big data in autoimmune geoepidemiology: analysis of 394,827 patients with systemic autoimmune diseases. Autoimmun Rev 14(8):670–679
Ratay ML, Balmert SC, Bassin EJ, Little SR (2018) Controlled release of an HDAC inhibitor for reduction of inflammation in dry eye disease. Acta Biomater 71:261–270
Reed JH, Gordon TP (2016) Autoimmunity: Ro60-associated RNA takes its toll on disease pathogenesis. Nat Rev Rheumatol 12(3):136–138
Renaudineau Y (2017) Immunophenotyping as a new tool for classification and monitoring of systemic autoimmune diseases. Clin Rev Allergy Immunol 53(2):177–180
Renaudineau Y, Ballestar E (2016) Epigenetics: DNA methylation signatures in Sjogren syndrome. Nat Rev Rheumatol 12(10):565–566
Renaudineau Y, Youinou P (2011) Epigenetics and autoimmunity, with special emphasis on methylation. Keio J Med 60(1):10–16
Renaudineau Y, Vallet S, Le Dantec C, Hillion S, Saraux A, Youinou P (2005) Characterization of the human CD5 endogenous retrovirus-E in B lymphocytes. Genes Immun 6(8):663–671
Sandhya P, Kurien BT, Danda D, Scofield RH (2017) Update on pathogenesis of Sjogren’s syndrome. Curr Rheumatol Rev 13(1):5–22
Satoh M, Chan JY, Ceribelli A, Vazquez del-Mercado M, Chan EK (2013) Autoantibodies to Argonaute 2 (Su antigen). Adv Exp Med Biol 768:45–59
Sepulveda D, Barrera MJ, Castro I, Aguilera S, Carvajal P, Lagos C, Gonzalez S, Albornoz N, Bahamondes V, Quest AFG, Urzua U, Molina C, Leyton C, Hermoso MA, Gonzalez MJ (2018) Impaired IRE1alpha/XBP-1 pathway associated to DNA methylation might contribute to salivary gland dysfunction in Sjogren’s syndrome patients. Rheumatology (Oxford) 57(6):1021–1032
Shapira Y, Agmon-Levin N, Shoenfeld Y (2010) Geoepidemiology of autoimmune rheumatic diseases. Nat Rev Rheumatol 6(8):468–476
Sharma R, Harris VM, Cavett J, Kurien BT, Liu K, Koelsch KA, Fayaaz A, Chaudhari KS, Radfar L, Lewis D, Stone DU, Kaufman CE, Li S, Segal B, Wallace DJ, Weisman MH, Venuturupalli S, Kelly JA, Pons-Estel B, Jonsson R, Lu X, Gottenberg JE, Anaya JM, Cunninghame-Graham DS, Huang AJW, Brennan MT, Hughes P, Alevizos I, Miceli-Richard C, Keystone EC, Bykerk VP, Hirschfield G, Nordmark G, Bucher SM, Eriksson P, Omdal R, Rhodus NL, Rischmueller M, Rohrer M, Wahren-Herlenius M, Witte T, Alarcon-Riquelme M, Mariette X, Lessard CJ, Harley JB, Ng WF, Rasmussen A, Sivils KL, Scofield RH (2017) Rare X chromosome abnormalities in systemic lupus erythematosus and Sjogren’s syndrome. Arthritis Rheumatol 69(11):2187–2192
Shi H, Zheng LY, Zhang P, Yu CQ (2014) miR-146a and miR-155 expression in PBMCs from patients with Sjogren’s syndrome. J Oral Pathol Med 43(10):792–797
Simonin L, Pasquier E, Leroyer C, Cornec D, Lemerle J, Bendaoud B, Hillion S, Pers JO, Couturaud F, Renaudineau Y (2016) Lymphocyte disturbances in primary antiphospholipid syndrome and application to venous thromboembolism follow-up. Clin Rev Allergy Immunol 53(1):14–27
Taylor KE, Wong Q, Levine DM, McHugh C, Laurie C, Doheny K, Lam MY, Baer AN, Challacombe S, Lanfranchi H, Schiodt M, Srinivasan M, Umehara H, Vivino FB, Zhao Y, Shiboski SC, Daniels TE, Greenspan JS, Shiboski CH, Criswell LA (2017) Genome-wide association analysis reveals genetic heterogeneity of Sjogren’s syndrome according to ancestry. Arthritis Rheumatol 69(6):1294–1305
Teos LY, Alevizos I (2017) Genetics of Sjogren’s syndrome. Clin Immunol 182:41–47
Thabet Y, Le Dantec C, Ghedira I, Devauchelle V, Cornec D, Pers JO, Renaudineau Y (2013) Epigenetic dysregulation in salivary glands from patients with primary Sjogren’s syndrome may be ascribed to infiltrating B cells. J Autoimmun 41:175–181
Tsigalou C, Stavropoulou E, Bezirtzoglou E (2018) Current insights in microbiome shifts in Sjogren’s syndrome and possible therapeutic interventions. Front Immunol 9:1106
Verstappen GM, Corneth OBJ, Bootsma H, Kroese FGM (2018) Th17 cells in primary Sjogren’s syndrome: pathogenicity and plasticity. J Autoimmun 87:16–25
Wang-Renault SF, Boudaoud S, Nocturne G, Roche E, Sigrist N, Daviaud C, Bugge Tinggaard A, Renault V, Deleuze JF, Mariette X, Tost J (2018) Deregulation of microRNA expression in purified T and B lymphocytes from patients with primary Sjogren’s syndrome. Ann Rheum Dis 77(1):133–140
Wattiaux MJ, Jouan-Flahault C, Youinou P, Cabane J, Andreani T, Serfaty L, Imbert JC (1995) Association of Gougerot-Sjogren syndrome and viral hepatitis C. Apropos of 6 cases. Ann Med Interne (Paris) 146(4):247–250
Williams AE, Choi K, Chan AL, Lee YJ, Reeves WH, Bubb MR, Stewart CM, Cha S (2016) Sjogren’s syndrome-associated microRNAs in CD14(+) monocytes unveils targeted TGFbeta signaling. Arthritis Res Ther 18(1):95
Yamamura Y, Motegi K, Kani K, Takano H, Momota Y, Aota K, Yamanoi T, Azuma M (2012) TNF-alpha inhibits aquaporin 5 expression in human salivary gland acinar cells via suppression of histone H4 acetylation. J Cell Mol Med 16(8):1766–1775
Yang Y, Peng L, Ma W, Yi F, Zhang Z, Chen H, Guo Y, Wang L, Zhao LD, Zheng W, Li J, Zhang F, Du Q (2016) Autoantigen-targeting microRNAs in Sjogren’s syndrome. Clin Rheumatol 35(4):911–917
Yin H, Zhao M, Wu X, Gao F, Luo Y, Ma L, Liu S, Zhang G, Chen J, Li F, Zuo X, Lu Q (2010) Hypomethylation and overexpression of CD70 (TNFSF7) in CD4+ T cells of patients with primary Sjogren’s syndrome. J Dermatol Sci 59(3):198–203
Yu X, Liang G, Yin H, Ngalamika O, Li F, Zhao M, Lu Q (2013) DNA hypermethylation leads to lower FOXP3 expression in CD4+ T cells of patients with primary Sjogren’s syndrome. Clin Immunol 148(2):254–257
Zare-Shahabadi A, Renaudineau Y, Rezaei N (2013) MicroRNAs and multiple sclerosis: from physiopathology toward therapy. Expert Opin Ther Targets 17(12):1497–1507
Zilahi E, Tarr T, Papp G, Griger Z, Sipka S, Zeher M (2012) Increased microRNA-146a/b, TRAF6 gene and decreased IRAK1 gene expressions in the peripheral mononuclear cells of patients with Sjogren’s syndrome. Immunol Lett 141(2):165–168
Acknowledgements
We are grateful to the “Association Française du Gougerot-Sjögren et des Syndromes Secs” for their support, and to Genevieve Michel and Simone Forest for their help in typing the paper.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Bordron, A., Devauchelle-Pensec, V., Le Dantec, C., Capdeville, A., Brooks, W.H., Renaudineau, Y. (2020). Epigenetics in Primary Sjögren’s Syndrome. In: Chang, C., Lu, Q. (eds) Epigenetics in Allergy and Autoimmunity. Advances in Experimental Medicine and Biology, vol 1253. Springer, Singapore. https://doi.org/10.1007/978-981-15-3449-2_11
Download citation
DOI: https://doi.org/10.1007/978-981-15-3449-2_11
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-3448-5
Online ISBN: 978-981-15-3449-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)