Abstract
The discovery of noncoding RNAs (ncRNAs) including short microRNAs, long ncRNAs and circular RNAs has broaden our knowledge about mammalian genomes and transcriptomes. A growing number of evidence on aberrantly regulated ncRNAs in cardiovascular diseases has indicated that ncRNAs are critical contributors to cardiovascular pathophysiology. Moreover, multiple recent studies have reported that ncRNAs can be detected in the bloodstream that differs between health subjects and diseased patients and some of them are remarkably stable. Although our knowledge about the origin and function of the circulating ncRNAs is still limited, these molecules have been regarded as promising noninvasive biomarker for risk stratification, diagnosis and prognosis of various cardiovascular diseases. In this chapter, we have described biological characteristics of circulating ncRNAs and discussed current trends and future prospects for the usage of circulating ncRNAs as biomarkers for common cardiovascular diseases.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Background
Only less than 3% of the human genome encodes messenger RNAs (mRNAs) that are encoded and participate in protein biosynthesis [1]. On the other hand, there are much more non-coding RNAs (ncRNAs) in the genome, most of which have undetermined functions. NcRNAs can be divided into basic ncRNAs and regulatory ncRNAs. Regulatory ncRNA can be further divided into microRNA (miRNA), long non-coding RNA (lncRNA), circular RNA (circRNA), piwi-interacting RNA (piRNA) and small interfering RNA (siRNA) [2]. Not surprisingly, regulatory ncRNAs have been found to be critical players in pathogenesis of human diseases, including cardiovascular diseases (CVDs). Numerous studies have demonstrated that miRNAs play a key role in driving gene expression changes in multiple cardiovascular pathological processes including cardiac hypertrophy, fibrosis, ischemia injury and heart failure [3, 4]. Whereas the role of lncRNAs and circRNAs are less understood compared to miRNAs, to date only a few candidates have been investigated in detail in cardiovascular system. Circulating ncRNAs have been have recently emerged as promising non-invasive biomarkers because their tissue- and time-specific expression pattern in CVDs and their ability to circulate in the bloodstream in a relative stable extracellular form [5]. Here, we present a brief introduction on miRNAs, lncRNAs and circRNAs, summarize the recent discovery of these ncRNAs in related to biomarker potential for common type of CVDs and, finally, discuss the current limitations and future prospects in developing ncRNAs as CVD biomarkers.
1.1 MicroRNAs
MiRNAs are abundant family of small ncRNAs in human genomes, containing more than 2000 different loci for miRNA generation [6], it is estimated that over 30% of the cellular transcriptome is orchestrated by miRNAs [7]. MiRNA is a short 18-22 nt ncRNA produced by transcription of specific genomic locus and specialized RNA endonuclease treatment [8, 9]. MiRNA transcription is conducted by RNA Pol II and is controlled by RNA Pol II-associated transcription factors and epigenetic regulators [10,11,12,13]. The primary miRNA (pri-miRNA) then go through several steps to become mature miRNA, with the help of nuclear RNase III Drosha [14,15,16]. Mature miRNAs can negatively regulate expression of target genes by binding to the 3’-UTR region of mRNAs and recruiting specific silencing proteins that form RNA-induced silencing complexes (RISC) [7]. In healthy condition, miRNAs act as modulators to steady protein levels that maintain physiological homeostasis. The regulatory activity of miRNAs depends on the abundance of their targets, so the same miRNA may have different regulatory function in different cell types [17]. While during pathological process, ectopic or aberrant expression of a particular miRNA in its original tissue can result in deregulation of its target transcripts and imbalanced physical functions. MiRNAs have been found present in variety of extracellular human body fluids including plasma, serum, saliva and urine [18,19,20,21]. Blood circulating miRNAs are most studied and found that the majority of circulating miRNAs in human blood are related with a protein named Argonaute 2 (Ago2) [22]. Ago2 is the effector component of the miRNA-induced silencing complex (RISC), it can directly bind miRNAs and drive mRNA suppression [23, 24]. Therefore it has been speculated that the majority of circulating miRNAs may be products released from dead cells that remain in extracellular space because of the high stability of Ago2-miRNA complex [5], which makes them potential indicators for various pathological conditions.
1.2 Long Non-coding RNAs
Long non-coding RNAs, also known as long ncRNAs or lncRNAs, are non-coding RNA transcripts that are longer than 200 nt, similar to protein-coding genes but lacking evident ORFs [25,26,27]. LncRNAs represent the majority of the ncRNAs, to date more than 58,000 lncRNAs has been classified [28]. However, only a few of them has been characterized with structure, function and impact in physical or pathological process. LncRNAs are produced with RNA polymerase II, which can be antisense, interleaved or overlapping with protein-coding genes, those sequences of lncRNA that do not overlap protein-coding genes term long intervening/intergenic noncoding RNAs (lincRNAs) [29]. LncRNAs are suggested very relevant players in the regulation of cellular functions because evidence shows they can interact with genomic DNA and RNA as a flexible molecular scaffold to recruit chromatin-modifying enzymes and transcription factors and to guide their transportation to the correct functional localization [30]. In addition, lncRNAs can act as guide molecules for DNA methyltransferase and histone modifier such as polycomb repressive complex PRC2 and histone H3 lysine 9 (H3K9) methyltransferases, which lead to repressive heterochromatin and the resultant transcriptional repression [31,32,33]. LncRNAs have also been reported to control the activity of other ncRNAs, particularly miRNAs, as decoys or sponges that can absorb miRNAs from their mRNA targets (and thus act as competing endogenous RNAs or ceRNAs) [34]. Loss of function experiments have provided evidence for the functional importance of lncRNAs in regulation of gene expression patterns that control cellular pluripotency, differentiation and survival [35]. Because of the enormous potential of lncRNAs to regulate gene expression, there is a growing interest in the potential roles of these RNAs in disease pathogenesis. In fact, numerous studies have shown a correlation between lncRNA dysregulation, with changes in gene expression and pathogenesis. Moreover, some studies have also suggested potential role of lncRNA in gene regulation outside the cell and between different cells. Recent researches have demonstrated that it is possible to detect the presence of lncRNAs in human body fluids, indicating the possible connection between circulating lncRNA concentration and disease initiation and development, makes lncRNAs potential novel diagnostic and prognostic tools [36]. However, on the other hand, most lncRNAs rapidly evolve at sequence and expression levels, it has been suggested that tissue-specific and possible three-dimensional structures of lncRNA are only conserved among closely related species.
1.3 Circular RNAs (CircRNAs)
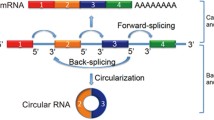
CircRNAs are a new class of endogenous noncoding RNAs and a field with much research activity, although the existence of circulating transcripts have been discovered for more than 20 years [37]. They are characterized by a covalently closed loop structure formed by back-splicing event that inversely connect exon boundaries [38]. These circular molecules have long been regarded as the artifacts of aberrant splicing or prerogative of several types of virus [39,40,41]. However, recent studies by using specific computational algorithm to identify circular molecules have demonstrated that in many cells the production of circRNA is not as rare as previously believed [42,43,44]. A growing number of evidence indicates that circRNAs are abundant, conserved, and stably accumulated in cells, and the expression pattern of circRNAs is highly dependent on cell type and species [45, 46]. Besides, circular RNAs are highly resistant to exonuclease RNase R, which makes them much more stable than to linear RNAs [47, 48], that explain their relative high evaluation conservation. The regulatory functions of circRNA remain to be further explored. Scientists have suggested several putative mechanisms of gene regulation by circular RNAs. (1) miRNA sponge: competitive endogenous RNA hypothesis is currently the most intensively studied and well accepted mechanism on regulatory activity of circRNAs on gene expression. CircRNA molecules contain lots of miRNA response elements (MREs) that allow them to competitively bind to miRNAs, causing suppression of the functional miRNA molecules and subsequent elevation of target miRNAs [47, 49]. (2) Interaction with RNA binding proteins (RBPs): strong direct interaction between circRNAs and their target RBPs enable gene regulation by competing with linear splicing [41]. (3) Regulation of parental gene transcription: some intronic circRNAs enhance the transcription of their hosting gene, probably by modulating RNA polymerase II in cis [50]. (4) Protein translation: some recent studies have demonstrated the potential of circRNA for direct protein translation, such as circ-ZNF609, circMbl3 and circ-SHPRH [51,52,53]. Furthermore, computational analysis of human transcriptomes sequencing has revealed the universal existence of circRNAs with coding potential [54, 55]. Due to their emerging role as regulators of gene expression, circRNAs are considered as important players in disease development. In addition, the stability of these circular molecule allow them to be easily identified and quantified in body fluid, which makes them high promising diagnostic biomarkers [46].
2 Circulating Non-coding RNA as Biomarkers for Cardiovascular Diseases
2.1 Myocardial Infarction
Myocardial infarction (MI) is the leading cause of death worldwide and is characterized by ischemia-induced localized heart tissue damage that induces cardiac remodeling and may progress to chronic heart failure. Appropriate therapies are required to reduce the mortality and thus a rapid diagnosis with high sensitivity and specificity is critical. MI is characterized by cell death and hypoxic stress, resulting in the release of various cardiac-specific proteins into the circulation. Classic MI biomarkers include serum concentrations of cardiac troponin (cardiac troponin T and I) and creatine kinase MB (CK-MB) [56]. Other than traditional protein markers, myocardium also releases ncRNAs into the bloodstream once injured. Numerous studies have described that single or a group of miRNAs in circulation can act as potential biomarker for cardiac injury including MI [57, 58]. MiR-1, which is abundantly expressed in cardiac and skeletal muscle and crucial in muscle differentiation and cardiac development, is firstly suggested as a circulating miRNA biomarker for acute MI [59,60,61]. Another high muscle-expressed miRNA miR-133, which is a crucial regulator of muscle development and pathophysiological alterations, has also been suggested to be a diagnostic biomarker for acute MI without prognostic potential on future left ventricular remodeling after MI [61, 62]. Cardiac-specific miRNAs miR-499 and miR-208a/b expressed by cardiac myosin genes have been suggested as biomarkers for myocardial damage and infarct severity [63]. In addition, results from patient and animal models showed a positive correlation between muscle and myocardial circulating miRNAs and acute MI with T-segment elevation (STEMI). The circulating levels of miRNAs including miR-1, miR-499-5p, miR-133a and miR-133b and followed the same pattern as rising of cardiac troponin T level and left ventricular ejection fraction (LVEF) in STEMI patients. Therefore, these miRNAs were regarded to be related to the extent of myocardial damage and necrosis after infarction [60]. It is worth mention that circulating non-muscle miRNA levels in patients with STEMI, such as liver miR-122-5p or pancreas-specific miR-375, showed an opposite pattern of muscle and cardiac-specific miRNAs, which was down-regulated in STEMI group of patients. These results were not consistent with the results observed in animal models of cardiogenic shock, in which plasma levels of liver-specific miR-122 showed a massive increase after external cardiac intervention and could indicate the time of infarction [64]. Additionally, a recent study has shown that plasma miR-122 levels measured less than 8 h after infarction demonstrated the same pattern of increase as that in animal models and miR-122-5p/133b ratio can act as a prognostic biomarker for successful stratification of STEMI patients [65]. Level of miR-133b in MI was measured in infarct-related artery (IRA) occlusion, without ST-segment elevation. Patients with closed IRA were found with higher levels of miR-133a, miR-133b then patients with patented IRA, but there was no difference in troponin T levels. These resulted suggested that elevated circulating miRNAs reveal the degree of IRA in MI and may indicate patients requiring urgent coronary revascularization [66].
Circulating miRNAs have also been used to predict individual risk for future fatal acute MI in healthy individuals [67]. The HUNT study examined 112 healthy subjects and identified 10 plasma miRNAs that were differentially expressed between lethal cases and controls. The best miRNA expression model for prediction of future fatal MI consists of miR-106a, miR-424, let-7 g, miR-144 and miR-660 levels, which provided a correct risk assessment of 77.6% (74.1% and 81.8% for men and woman respectively). Other circulating miRNAs such as miR-34a, miR-192 and miR-194 have also been shown to be good predictors of risk assessment for heart failure after MI [68]. These miRNAs are expressed in a p53-dependent manner, linking them to other miRNAs that have been described as driving factors for CVDs [69]. Collectively, multiple studies have confirmed the ideas that circulating miRNA may serve as sensitive and specific biomarkers for MI, the combination of miRNA with cardiac troponin might be accurate diagnostic and prognostic tool for patients.
Recently, some circulating lncRNAs and circRNAs have also been explored as potential biomarkers of acute MI. CDR1 antisense (CDR1AS) and cyclic zinc finger antisense 1 (ZFAS1) showed significant differential expression between acute MI patients and healthy subjects; and similar changes in circulating CDR1AS and ZFAS1 were also consistently observed in the mouse models. Thereby researchers suggested changes in circulating CDR1AS and ZFAS1 could independently predict acute MI [70]. Another lncRNA urothelial carcinoma associated 1 (UCA1) was studied as well, which was found to be expressed in bladder and lung cancer and suggested as a predictive biomarker. UCA1 is specifically expressed in the heart of healthy adult individuals; while plasma UCA1 levels are reduced in the early state of patients with acute MI and increased on day 3 post-MI. The level of UCA1 circulating was also found negatively correlated with the expression of miR-177 [71].
2.2 Coronary Artery Disease
Coronary Artery Disease (CAD) is caused by the formation of atherosclerotic plaques, resulting in structural remodeling of the arterial wall, activation of endothelial cells and inflammatory cells may eventually lead to myocardial ischemia [72]. Activation of endothelial cells is critical for atherosclerosis; it is a potential source to seek new biomarkers for early diagnosis and identification of instable plaque, which eventually allow risk stratification of patients. It has been suggested that miRNAs associated with cellular components formed by atherogenesis are deregulated in CAD [73]. However, the cyclic signature data of CAD miRNAs are not consistent. Endothelial cells (miR-17, miR-92a and miR-126), inflammation (miR-155) and smooth muscle cell-associated (miR-145) miRNA were found to decrease in the circulation of CAD patients, while plasma myocardium and muscle miRNA (miR- 133a, miR-208a and miR-499) were increased. It was believed that miRNA may be cleared from the bloodstream by ingestion of atherosclerotic lesions or vasculature, and that enhanced release and elevation of miRNA may reflect myocardial damage [74]. In contrast, the miRNA signature of miR-126 and miR-17/92a cluster was up-regulated together with miR-451, miR-106b/25 cluster and miR-21/590-5p family in vulnerable CAD and was suggested as a novel biomarker [75]. Consistent with previous results, miR-1, miR-133a/b, miR-122, miR-126 and miR-199a have been reported elevated in the circulation of stable and unstable angina patients and miR-92a and miR-486 were associated with high-density lipoprotein components identified as potential circulating biomarkers for coronary plaque [76,77,78]. In addition, the severity of CAD for patients with hyperlipidemia was found associated with increased plasma levels of lipid metabolism-related miR-122 and miR-370 [79]. MiRNA signatures for risk assessment in patients with symptomatic obstructive CAD and chest pain were explored. MiR-134, miR-2861 and miR-3135b were associated with coronary artery calcification and were altered in patients with obstructive CAD [80]. A prognostic analysis evaluated circulating vascular and endothelial miRNAs in patients with CAD and found expression level of miR-126 and miR-199a contained in microvesicles but not freely circulating miRNA could predict the occurrence of cardiovascular events in patients with stable CAD [81]. Collectively, blood miRNAs have the potential to improve CAD diagnosis and prognosis, whereas, replication and validation of these findings in large independent cohorts are still required.
Recently, lncRNAs have get attention as CAD biomarkers. Microarray-based screening of plasma in CAD patients identified a transcript called CoroMarker as a marker for stability, sensitivity, and specificity of the CAD [82, 83]. This lncRNA is present in extracellular vesicles and circulating monocytes in peripheral blood. The same group reported another lncRNA LncPPARδ, which was elevated in circulating peripheral blood mononuclear cells, as another CAD biomarker in combination with other risk factors [84]. The combined use of circRNAs and miRNAs as biomarker for carotid plaque rupture was also investigated and found the ratio of serum circR-284/has-miR-221 was significantly increased in acutely symptomatic patients with carotid disease. This combination demonstrated favorable characteristics to be a prognostic biomarker of plaque rapture and stroke [85].
2.3 Cardiomyopathy
Cardiomyopathies are a group of heart diseases characterized by morphological and functional abnormalities in the myocardium. When they originate from myocardial dysfunction or changes in the body, they can be classified as primary or intrinsic cardiomyopathy; and when their pathogenic factors are external factors for the heart, they can be classified as secondary or extrinsic cardiomyopathy [72]. Intrinsic cardiomyopathy can be obtained with a genetic basis or in response to stress on the myocardium. Inherited cardiomyopathies are the most common form of the disease, hence genetic testing is the most common diagnosis. Also, patients with cardiomyopathy often receive a series of biochemical tests to detect biomarkers that can assist diagnose [86]. Several therapies for cancer and other diseases can cause serious side effects that affect cardiovascular health. Cardiotoxicity and damage impede heart function and cause high blood pressure, apoptosis, arrhythmia, fibrosis, and finally heart failure. Therefore, minimizing or preventing these side effects by early monitoring of drug-induced cardiotoxicity and damage would be very significant for treatment strategies [87]. A few studies have evaluated ncRNA plasma levels in drug-induced cardiomyopathies. An in vivo study evaluating isoproterenol-induced cardiactoxic model in rats reported an increase in serum miR-208 level in a time-dependent manner and was associated with traditional myocardial injury cardiac troponin I [88]. Other animal studies support the response of miR-208 to isoproterenol, metaproterenol, allylamine and mitoxantrone [89,90,91]. On the other hand, miR-208 did not respond to a single administration of doxorubicin, and doxorubicin treatment induced other muscle and heart-specific miRNAs. In the chemotherapy treatment of doxorubicin, circulating miR-208a was not detected in the bloodstream of breast cancer patients [92]. These differences may be due to species-specificity, time- or dose-dependent effects, or indicate that different drugs may cause different circulating miRNA patterns. Therefore, other miRNAs should be considered to be biomarkers of drug-induced cardiotoxicity. Zhao et al. determined whether detectable levels of specific miRNAs are released into the circulation for bevacizumab-induced cardiotoxicity. They identified two cancer-associated miRNAs (miR-579 and miR-1254) that were specifically elevated in the circulation of bevacizumab-induced cardiotoxic patients and distinguish this patient group from AMI patients. MiR-1254 also showed strong correlation to the clinical diagnosis of bevacizumab-induced cardiotoxicity [93].
3 Prospects and Challenges
We have accumulated a great deal about the association of circulating miRNAs with various types of human heart diseases and injuries. Other circulating ncRNAs species as lncRNAs and circRNAs are also promising biomarkers of CVDs, however their physiological or pathological roles in the context of CVDs remain largely unknown. The study of circulating lncRNAs asa cardiac biomarkers is not as advanced as miRNA, partially because of the general assumption that lncRNAs are unstable in body fluids and the findings that lncRNAs are not conserved among species as miRNAs. However, recent data indicate that most lncRNAs are stable in neuroblastoma cell lines, although this study does not address the problem of extracellular stability [94]. Additionally, due to the emerging role of circRNA as regulators of gene expression, circRNA is likely to be an important player in the initiation and progress of diseases including CVDs. Research on the circular and stable RNA molecules has just begun and is a completely new field of research that will help to better understand the pathogenesis of CVDs. Regulatory networks occur in complex organisms and the surprising stability of these molecules suggests that circRNAs have great potential to be developed as CVD biomarkers.
Nevertheless, currently the use of ncRNA is still limited by (1) insufficient knowledge about the origin and function of ncRNAs especially for lncRNA and circRNAs; (2) diversity of RNA extraction and ncRNA detection methods without a standard protocol; (3) lack of consistency and standardization in different studies on same type of CVDs; (4) a relatively small patient cohort to date [95]. Still, some circulating ncRNAs appear to have stronger diagnostic and prognostic value than conventional biomarkers, not only because of their tissue and disease-specific expression patterns, but also because of the high physicochemical properties and their high stability in circulation system [96,97,98]. Whether circulating ncRNAs represent attractive diagnostic and prognostic biomarkers required future studies of large cohorts with standardized protocol for processing body fluid and RNA procreation and consistent analysis method.
References
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov JP, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C, Stange-Thomann Y, Stojanovic N, Subramanian A, Wyman D, Rogers J, Sulston J, Ainscough R, Beck S, Bentley D, Burton J, Clee C, Carter N, Coulson A, Deadman R, Deloukas P, Dunham A, Dunham I, Durbin R, French L, Grafham D, Gregory S, Hubbard T, Humphray S, Hunt A, Jones M, Lloyd C, McMurray A, Matthews L, Mercer S, Milne S, Mullikin JC, Mungall A, Plumb R, Ross M, Shownkeen R, Sims S, Waterston RH, Wilson RK, Hillier LW, McPherson JD, Marra MA, Mardis ER, Fulton LA, Chinwalla AT, Pepin KH, Gish WR, Chissoe SL, Wendl MC, Delehaunty KD, Miner TL, Delehaunty A, Kramer JB, Cook LL, Fulton RS, Johnson DL, Minx PJ, Clifton SW, Hawkins T, Branscomb E, Predki P, Richardson P, Wenning S, Slezak T, Doggett N, Cheng JF, Olsen A, Lucas S, Elkin C, Uberbacher E, Frazier M, Gibbs RA, Muzny DM, Scherer SE, Bouck JB, Sodergren EJ, Worley KC, Rives CM, Gorrell JH, Metzker ML, Naylor SL, Kucherlapati RS, Nelson DL, Weinstock GM, Sakaki Y, Fujiyama A, Hattori M, Yada T, Toyoda A, Itoh T, Kawagoe C, Watanabe H, Totoki Y, Taylor T, Weissenbach J, Heilig R, Saurin W, Artiguenave F, Brottier P, Bruls T, Pelletier E, Robert C, Wincker P, Smith DR, Doucette-Stamm L, Rubenfield M, Weinstock K, Lee HM, Dubois J, Rosenthal A, Platzer M, Nyakatura G, Taudien S, Rump A, Yang H, Yu J, Wang J, Huang G, Gu J, Hood L, Rowen L, Madan A, Qin S, Davis RW, Federspiel NA, Abola AP, Proctor MJ, Myers RM, Schmutz J, Dickson M, Grimwood J, Cox DR, Olson MV, Kaul R, Raymond C, Shimizu N, Kawasaki K, Minoshima S, Evans GA, Athanasiou M, Schultz R, Roe BA, Chen F, Pan H, Ramser J, Lehrach H, Reinhardt R, McCombie WR, de la Bastide M, Dedhia N, Blocker H, Hornischer K, Nordsiek G, Agarwala R, Aravind L, Bailey JA, Bateman A, Batzoglou S, Birney E, Bork P, Brown DG, Burge CB, Cerutti L, Chen HC, Church D, Clamp M, Copley RR, Doerks T, Eddy SR, Eichler EE, Furey TS, Galagan J, Gilbert JG, Harmon C, Hayashizaki Y, Haussler D, Hermjakob H, Hokamp K, Jang W, Johnson LS, Jones TA, Kasif S, Kaspryzk A, Kennedy S, Kent WJ, Kitts P, Koonin EV, Korf I, Kulp D, Lancet D, Lowe TM, McLysaght A, Mikkelsen T, Moran JV, Mulder N, Pollara VJ, Ponting CP, Schuler G, Schultz J, Slater G, Smit AF, Stupka E, Szustakowki J, Thierry-Mieg D, Thierry-Mieg J, Wagner L, Wallis J, Wheeler R, Williams A, Wolf YI, Wolfe KH, Yang SP, Yeh RF, Collins F, Guyer MS, Peterson J, Felsenfeld A, Wetterstrand KA, Patrinos A, Morgan MJ, de Jong P, Catanese JJ, Osoegawa K, Shizuya H, Choi S, Chen YJ, Szustakowki J, International Human Genome Sequencing C. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921.
Thum T. Facts and updates about cardiovascular non-coding RNAs in heart failure. ESC Heart Failure. 2015;2(3):108–11.
Papait R, Kunderfranco P, Stirparo GG, Latronico MV, Condorelli G. Long noncoding RNA: a new player of heart failure? J Cardiovasc Transl Res. 2013;6(6):876–83.
Thum T, Condorelli G. Long noncoding RNAs and microRNAs in cardiovascular pathophysiology. Circ Res. 2015;116(4):751–62.
Turchinovich A, Weiz L, Langheinz A, Burwinkel B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 2011;39(16):7223–33.
Kozomara A, Griffiths-Jones S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014;42(Database issue):D68–73.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136(2):215–33.
Zhuo Y, Gao G, Shi JA, Zhou X, Wang X. miRNAs: biogenesis, origin and evolution, functions on virus-host interaction. Cell Physiol Biochem. 2013;32(3):499–510.
Lee Y, Jeon K, Lee JT, Kim S, Kim VN. MicroRNA maturation: stepwise processing and subcellular localization. Eur Mol Biol Organ J. 2002;21(17):4663–70.
Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet. 2010;11(9):597–610.
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, Kim VN. MicroRNA genes are transcribed by RNA polymerase II. Eur Mol Biol Organ J. 2004;23(20):4051–60.
Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol. 2009;10(2):126–39.
Davis-Dusenbery BN, Hata A. Mechanisms of control of microRNA biogenesis. Biochem Med. 2010;148(4):381–92.
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, Lee J, Provost P, Radmark O, Kim S, Kim VN. The nuclear RNase III Drosha initiates microRNA processing. Nature. 2003;425(6956):415–9.
Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ. Processing of primary microRNAs by the Microprocessor complex. Nature. 2004;432(7014):231–5.
Han J, Lee Y, Yeom KH, Nam JW, Heo I, Rhee JK, Sohn SY, Cho Y, Zhang BT, Kim VN. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell. 2006;125(5):887–901.
Lu J, Clark AG. Impact of microRNA regulation on variation in human gene expression. Genome Res. 2012;22(7):1243–54.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O’Briant KC, Allen A, Lin DW, Urban N, Drescher CW, Knudsen BS, Stirewalt DL, Gentleman R, Vessella RL, Nelson PS, Martin DB, Tewari M. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 2008;105(30):10513–8.
Zubakov D, Boersma AW, Choi Y, van Kuijk PF, Wiemer EA, Kayser M. MicroRNA markers for forensic body fluid identification obtained from microarray screening and quantitative RT-PCR confirmation. Int J Leg Med. 2010;124(3):217–26.
Gupta SK, Bang C, Thum T. Circulating microRNAs as biomarkers and potential paracrine mediators of cardiovascular disease. Circ Cardiovasc Genet. 2010;3(5):484–8.
Wang L, Lv Y, Li G, Xiao J. MicroRNAs in heart and circulation during physical exercise. J Sport Health Sci. 2018;7(4):433–41.
Arroyo JD, Chevillet JR, Kroh EM, Ruf IK, Pritchard CC, Gibson DF, Mitchell PS, Bennett CF, Pogosova-Agadjanyan EL, Stirewalt DL, Tait JF, Tewari M. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc Natl Acad Sci U S A. 2011;108(12):5003–8.
Song JJ, Liu J, Tolia NH, Schneiderman J, Smith SK, Martienssen RA, Hannon GJ, Joshua-Tor L. The crystal structure of the Argonaute2 PAZ domain reveals an RNA binding motif in RNAi effector complexes. Nat Struct Biol. 2003;10(12):1026–32.
Ma JB, Ye K, Patel DJ. Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature. 2004;429(6989):318–22.
Dey BK, Mueller AC, Dutta A. Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription. 2014;5(4):e944014.
Rinn JL, Chang HY. Genome regulation by long noncoding RNAs. Annu Rev Biochem. 2012;81:145–66.
Anderson DM, Anderson KM, Chang CL, Makarewich CA, Nelson BR, McAnally JR, Kasaragod P, Shelton JM, Liou J, Bassel-Duby R, Olson EN. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell. 2015;160(4):595–606.
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, Barrette TR, Prensner JR, Evans JR, Zhao S, Poliakov A, Cao X, Dhanasekaran SM, Wu YM, Robinson DR, Beer DG, Feng FY, Iyer HK, Chinnaiyan AM. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47(3):199–208.
Ransohoff JD, Wei Y, Khavari PA. The functions and unique features of long intergenic non-coding RNA. Nat Rev Mol Cell Biol. 2018;19(3):143–57.
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, Li R, West RB, van de Vijver MJ, Sukumar S, Chang HY. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010;464(7291):1071–6.
Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science. 2008;322(5902):750–6.
Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R, Fraser P. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science. 2008;322(5908):1717–20.
Pandey RR, Mondal T, Mohammad F, Enroth S, Redrup L, Komorowski J, Nagano T, Mancini-Dinardo D, Kanduri C. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell. 2008;32(2):232–46.
Wang Y, Hou J, He D, Sun M, Zhang P, Yu Y, Chen Y. The emerging function and mechanism of ceRNAs in Cancer. Trends Genet. 2016;32(4):211–24.
Chen ZH, Wang WT, Huang W, Fang K, Sun YM, Liu SR, Luo XQ, Chen YQ. The lnc RNA HOTAIRM1 regulates the degradation of PML-RARA oncoprotein and myeloid cell differentiation by enhancing the autophagy pathway. Cell Death Differ. 2017;24(2):212–24.
Huang X, Yuan T, Tschannen M, Sun Z, Jacob H, Du M, Liang M, Dittmar RL, Liu Y, Liang M, Kohli M, Thibodeau SN, Boardman L, Wang L. Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genomics. 2013;14:319.
Nigro JM, Cho KR, Fearon ER, Kern SE, Ruppert JM, Oliner JD, Kinzler KW, Vogelstein B. Scrambled exons. Cell. 1991;64(3):607–13.
Lasda E, Parker R. Circular RNAs: diversity of form and function. RNA. 2014;20(12):1829–42.
Kos A, Dijkema R, Arnberg AC, van der Meide PH, Schellekens H. The hepatitis delta (delta) virus possesses a circular RNA. Nature. 1986;323(6088):558–60.
Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci U S A. 1976;73(11):3852–6.
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S. circ RNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56(1):55–66.
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012;7(2):e30733.
Wu Q, Wang Y, Cao M, Pantaleo V, Burgyan J, Li WX, Ding SW. Homology-independent discovery of replicating pathogenic circular RNAs by deep sequencing and a new computational algorithm. Proc Natl Acad Sci U S A. 2012;109(10):3938–43.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, Marzluff WF, Sharpless NE. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19(2):141–57.
Danan M, Schwartz S, Edelheit S, Sorek R. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res. 2012;40(7):3131–42.
Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z, Sharpless NE. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLos Genet. 2010;6(12):e1001233.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, Kocks C, le Noble F, Rajewsky N. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333–8.
Enuka Y, Lauriola M, Feldman ME, Sas-Chen A, Ulitsky I, Yarden Y. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 2016;44(3):1370–83.
Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505(7483):344–52.
Zhang Y, Zhang XO, Chen T, Xiang JF, Yin QF, Xing YH, Zhu S, Yang L, Chen LL. Circular intronic long noncoding RNAs. Mol Cell. 2013;51(6):792–806.
Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O, Fatica A, Santini T, Andronache A, Wade M, Laneve P, Rajewsky N, Bozzoni I. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol Cell. 2017;66(1):22–37.. e29
Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E, Perez-Hernandez D, Ramberger E, Shenzis S, Samson M, Dittmar G, Landthaler M, Chekulaeva M, Rajewsky N, Kadener S. Translation of CircRNAs. Mol Cell. 2017;66(1):9–21.. e27
Begum S, Yiu A, Stebbing J, Castellano L. Novel tumour suppressive protein encoded by circular RNA, circ-SHPRH, in glioblastomas. Oncogene. 2018;37(30):4055.
Abe N, Matsumoto K, Nishihara M, Nakano Y, Shibata A, Maruyama H, Shuto S, Matsuda A, Yoshida M, Ito Y, Abe H. Rolling circle translation of circular RNA in living human cells. Sci Rep. 2015;5:16435.
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, Jin Y, Yang Y, Chen LL, Wang Y, Wong CC, Xiao X, Wang Z. Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res. 2017;27(5):626–41.
Hachey BJ, Kontos MC, Newby LK, Christenson RH, Peacock WF, Brewer KC, McCord J. Trends in use of biomarker protocols for the evaluation of possible myocardial infarction. J Am Heart Assoc. 2017;6(9)
Goldberg L, Tirosh-Wagner T, Vardi A, Abbas H, Pillar N, Shomron N, Nevo-Caspi Y, Paret G. Circulating microRNAs: a potential biomarker for cardiac damage, inflammatory response, and left ventricular function recovery in pediatric viral myocarditis. J Cardiovasc Transl Res. 2018;11(4):319–28.
Deddens JC, Vrijsen KR, Colijn JM, Oerlemans MI, Metz CH, van der Vlist EJ, Nolte-’t Hoen EN, den Ouden K, Jansen Of Lorkeers SJ, van der Spoel TI, Koudstaal S, Arkesteijn GJ, Wauben MH, van Laake LW, Doevendans PA, Chamuleau SA, Sluijter JP. Circulating extracellular vesicles contain miRNAs and are released as early biomarkers for cardiac injury. J Cardiovasc Transl Res. 2016;9(4):291–301.
Ai J, Zhang R, Li Y, Pu J, Lu Y, Jiao J, Li K, Yu B, Li Z, Wang R, Wang L, Li Q, Wang N, Shan H, Li Z, Yang B. Circulating microRNA-1 as a potential novel biomarker for acute myocardial infarction. Biochem Biophys Res Commun. 2010;391(1):73–7.
D’Alessandra Y, Devanna P, Limana F, Straino S, Di Carlo A, Brambilla PG, Rubino M, Carena MC, Spazzafumo L, De Simone M, Micheli B, Biglioli P, Achilli F, Martelli F, Maggiolini S, Marenzi G, Pompilio G, Capogrossi MC. Circulating microRNAs are new and sensitive biomarkers of myocardial infarction. Eur Heart J. 2010;31(22):2765–73.
Kuwabara Y, Ono K, Horie T, Nishi H, Nagao K, Kinoshita M, Watanabe S, Baba O, Kojima Y, Shizuta S, Imai M, Tamura T, Kita T, Kimura T. Increased microRNA-1 and microRNA-133a levels in serum of patients with cardiovascular disease indicate myocardial damage. Circ Cardiovasc Genet. 2011;4(4):446–54.
Bauters C, Kumarswamy R, Holzmann A, Bretthauer J, Anker SD, Pinet F, Thum T. Circulating miR-133a and miR-423-5p fail as biomarkers for left ventricular remodeling after myocardial infarction. Int J Cardiol. 2013;168(3):1837–40.
Wang GK, Zhu JQ, Zhang JT, Li Q, Li Y, He J, Qin YW, Jing Q. Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur Heart J. 2010;31(6):659–66.
Andersson P, Gidlof O, Braun OO, Gotberg M, van der Pals J, Olde B, Erlinge D. Plasma levels of liver-specific miR-122 is massively increased in a porcine cardiogenic shock model and attenuated by hypothermia. Shock. 2012;37(2):234–8.
Cortez-Dias N, Costa MC, Carrilho-Ferreira P, Silva D, Jorge C, Calisto C, Pessoa T, Robalo Martins S, de Sousa JC, da Silva PC, Fiuza M, Diogo AN, Pinto FJ, Enguita FJ. Circulating miR-122-5p/miR-133b ratio is a specific early prognostic biomarker in acute myocardial infarction. Circ J. 2016;80(10):2183–91.
Gacon J, Kablak-Ziembicka A, Stepien E, Enguita FJ, Karch I, Derlaga B, Zmudka K, Przewlocki T. Decision-making microRNAs (miR-124, -133a/b, −34a and −134) in patients with occluded target vessel in acute coronary syndrome. Kardiologia Polska. 2016;74(3):280–8.
Bye A, Rosjo H, Nauman J, Silva GJ, Follestad T, Omland T, Wisloff U. Circulating microRNAs predict future fatal myocardial infarction in healthy individuals – The HUNT study. J Mol Cell Cardiol. 2016;97:162–8.
Matsumoto S, Sakata Y, Suna S, Nakatani D, Usami M, Hara M, Kitamura T, Hamasaki T, Nanto S, Kawahara Y, Komuro I. Circulating p53-responsive microRNAs are predictive indicators of heart failure after acute myocardial infarction. Circ Res. 2013;113(3):322–6.
Evans S, Mann DL. Circulating p53-responsive microRNAs as predictive biomarkers in heart failure after acute myocardial infarction: the long and arduous road from scientific discovery to clinical utility. Circ Res. 2013;113(3):242–4.
Zhang Y, Sun L, Xuan L, Pan Z, Li K, Liu S, Huang Y, Zhao X, Huang L, Wang Z, Hou Y, Li J, Tian Y, Yu J, Han H, Liu Y, Gao F, Zhang Y, Wang S, Du Z, Lu Y, Yang B. Reciprocal changes of circulating long non-coding RNAs ZFAS1 and CDR1AS predict acute myocardial infarction. Sci Rep. 2016;6:22384.
Mentz RJ, O’Connor CM. Pathophysiology and clinical evaluation of acute heart failure. Nat Rev Cardiol. 2016;13(1):28–35.
Maron BJ, Towbin JA, Thiene G, Antzelevitch C, Corrado D, Arnett D, Moss AJ, Seidman CE, Young JB, American Heart A, Council on Clinical Cardiology HF, Transplantation C, Quality of C, Outcomes R, Functional G, Translational Biology Interdisciplinary Working G, Council on E, Prevention. Contemporary definitions and classification of the cardiomyopathies: an American heart association scientific statement from the council on clinical cardiology, heart failure and transplantation committee; quality of care and outcomes research and functional genomics and translational biology interdisciplinary working groups; and council on epidemiology and prevention. Circulation. 2006;113(14):1807–16.
Lu M, Yuan S, Li S, Li L, Liu M, Wan S. The exosome-derived biomarker in atherosclerosis and its clinical application. J Cardiovasc Transl Res. 2019;12(1):68–74.
Fichtlscherer S, De Rosa S, Fox H, Schwietz T, Fischer A, Liebetrau C, Weber M, Hamm CW, Roxe T, Muller-Ardogan M, Bonauer A, Zeiher AM, Dimmeler S. Circulating microRNAs in patients with coronary artery disease. Circ Res. 2010;107(5):677–84.
Ren J, Zhang J, Xu N, Han G, Geng Q, Song J, Li S, Zhao J, Chen H. Signature of circulating microRNAs as potential biomarkers in vulnerable coronary artery disease. PLoS One. 2013;8(12):e80738.
D’Alessandra Y, Carena MC, Spazzafumo L, Martinelli F, Bassetti B, Devanna P, Rubino M, Marenzi G, Colombo GI, Achilli F, Maggiolini S, Capogrossi MC, Pompilio G. Diagnostic potential of plasmatic MicroRNA signatures in stable and unstable angina. PLoS One. 2013;8(11):e80345.
Niculescu LS, Simionescu N, Sanda GM, Carnuta MG, Stancu CS, Popescu AC, Popescu MR, Vlad A, Dimulescu DR, Simionescu M, Sima AV. MiR-486 and miR-92a identified in circulating HDL discriminate between stable and vulnerable coronary artery disease patients. PLoS One. 2015;10(10):e0140958.
Al-Muhtaresh HA, Salem AH, Al-Kafaji G. Upregulation of circulating cardiomyocyte-enriched miR-1 and miR-133 associate with the risk of coronary artery disease in Type 2 diabetes patients and serve as potential biomarkers. J Cardiovasc Transl Res. 2019;12(4):347–57.
Gao W, He HW, Wang ZM, Zhao H, Lian XQ, Wang YS, Zhu J, Yan JJ, Zhang DG, Yang ZJ, Wang LS. Plasma levels of lipometabolism-related miR-122 and miR-370 are increased in patients with hyperlipidemia and associated with coronary artery disease. Lipids Health Dis. 2012;11:55.
Liu W, Ling S, Sun W, Liu T, Li Y, Zhong G, Zhao D, Zhang P, Song J, Jin X, Xu Z, Song H, Li Q, Liu S, Chai M, Dai Q, He Y, Fan Z, Zhou YJ, Li Y. Circulating microRNAs correlated with the level of coronary artery calcification in symptomatic patients. Sci Rep. 2015;5:16099.
Jansen F, Yang X, Proebsting S, Hoelscher M, Przybilla D, Baumann K, Schmitz T, Dolf A, Endl E, Franklin BS, Sinning JM, Vasa-Nicotera M, Nickenig G, Werner N. MicroRNA expression in circulating microvesicles predicts cardiovascular events in patients with coronary artery disease. J Am Heart Assoc. 2014;3(6):e001249.
Cai Y, Yang Y, Chen X, Wu G, Zhang X, Liu Y, Yu J, Wang X, Fu J, Li C, Jose PA, Zeng C, Zhou L. Circulating ‘lncRNA OTTHUMT00000387022’ from monocytes as a novel biomarker for coronary artery disease. Cardiovasc Res. 2016;112(3):714–24.
Yang Y, Cai Y, Wu G, Chen X, Liu Y, Wang X, Yu J, Li C, Chen X, Jose PA, Zhou L, Zeng C. Plasma long non-coding RNA, CoroMarker, a novel biomarker for diagnosis of coronary artery disease. Clin Neurosci Res. 2015;129(8):675–85.
Cai Y, Yang Y, Chen X, He D, Zhang X, Wen X, Hu J, Fu C, Qiu D, Jose PA, Zeng C, Zhou L. Circulating “LncPPARdelta” from monocytes as a novel biomarker for coronary artery diseases. Medicine. 2016;95(6):e2360.
Tijsen AJ, Creemers EE, Moerland PD, de Windt LJ, van der Wal AC, Kok WE, Pinto YM. MiR423-5p as a circulating biomarker for heart failure. Circ Res. 2010;106(6):1035–9.
Coats CJ, Heywood WE, Mills K, Elliott PM. Current applications of biomarkers in cardiomyopathies. Expert Rev Cardiovasc Ther. 2015;13(7):825–37.
Sandhu H, Maddock H. Molecular basis of cancer-therapy-induced cardiotoxicity: introducing microRNA biomarkers for early assessment of subclinical myocardial injury. Clin Neurosci Res (Lond). 2014;126(6):377–400.
Ji X, Takahashi R, Hiura Y, Hirokawa G, Fukushima Y, Iwai N. Plasma miR-208 as a biomarker of myocardial injury. Clin Chem. 2009;55(11):1944–9.
Nishimura Y, Kondo C, Morikawa Y, Tonomura Y, Torii M, Yamate J, Uehara T. Plasma miR-208 as a useful biomarker for drug-induced cardiotoxicity in rats. J Appl Toxicol. 2015;35(2):173–80.
Calvano J, Achanzar W, Murphy B, DiPiero J, Hixson C, Parrula C, Burr H, Mangipudy R, Tirmenstein M. Evaluation of microRNAs-208 and 133a/b as differential biomarkers of acute cardiac and skeletal muscle toxicity in rats. Toxicol Appl Pharmacol. 2016;312:53–60.
Glineur SF, De Ron P, Hanon E, Valentin JP, Dremier S, Nogueira da Costa A. Paving the route to plasma miR-208a-3p as an acute cardiac injury biomarker: preclinical rat data supports its use in drug safety assessment. Toxicol Sci. 2016;149(1):89–97.
Oliveira-Carvalho V, Ferreira LR, Bocchi EA. Circulating mir-208a fails as a biomarker of doxorubicin-induced cardiotoxicity in breast cancer patients. J Appl Toxicol. 2015;35(9):1071–2.
Zhao Z, He J, Zhang J, Liu M, Yang S, Li N, Li X. Dysregulated miR1254 and miR579 for cardiotoxicity in patients treated with bevacizumab in colorectal cancer. Tumor Biol. 2014;35(6):5227–35.
Clark MB, Johnston RL, Inostroza-Ponta M, Fox AH, Fortini E, Moscato P, Dinger ME, Mattick JS. Genome-wide analysis of long noncoding RNA stability. Genome Res. 2012;22(5):885–98.
Moldovan L, Batte KE, Trgovcich J, Wisler J, Marsh CB, Piper M. Methodological challenges in utilizing miRNAs as circulating biomarkers. J Cell Mol Med. 2014;18(3):371–90.
Quinn JJ, Chang HY. Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet. 2016;17(1):47–62.
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X, Li Q, Li X, Wang W, Zhang Y, Wang J, Jiang X, Xiang Y, Xu C, Zheng P, Zhang J, Li R, Zhang H, Shang X, Gong T, Ning G, Wang J, Zen K, Zhang J, Zhang CY. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18(10):997–1006.
Arita T, Ichikawa D, Konishi H, Komatsu S, Shiozaki A, Shoda K, Kawaguchi T, Hirajima S, Nagata H, Kubota T, Fujiwara H, Okamoto K, Otsuji E. Circulating long non-coding RNAs in plasma of patients with gastric cancer. Anticancer Res. 2013;33(8):3185–93.
Acknowledgements
This work was supported by the grants from National Natural Science Foundation of China.
Competing Financial Interests
The authors declare no competing financial interests.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Zhao, C., Lv, Y., Duan, Y., Li, G., Zhang, Z. (2020). Circulating Non-coding RNAs and Cardiovascular Diseases. In: Xiao, J. (eds) Non-coding RNAs in Cardiovascular Diseases. Advances in Experimental Medicine and Biology, vol 1229. Springer, Singapore. https://doi.org/10.1007/978-981-15-1671-9_22
Download citation
DOI: https://doi.org/10.1007/978-981-15-1671-9_22
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1670-2
Online ISBN: 978-981-15-1671-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)