Abstract
The Ischemic Heart Disease (IHD) is considered a clinical condition characterized by myocardial ischemia causing an imbalance between myocardial blood supply and demand, leading to morbidity and mortality across the worldwide. Prompt diagnostic and prognostic represents key factors for the treatment and reduction of the mortality rate. Therefore, one of the newest frontiers in cardiovascular research is related to non-coding RNAs (ncRNAs), which prompted a huge interest in exploring ncRNAs candidates for utilization as potential therapeutic targets for diagnostic and prognostic and/or biomarkers in IHD. However, there are undoubtedly many more functional ncRNAs yet to be discovered and characterized. Here we will discuss our current knowledge and we will provide insight on the roles and effects elicited by some ncRNAs related to IHD.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Ischemic Heart Disease (IHD) is the leading death cause in the western countries, which happens when the heart became unable to pump blood properly due to myocardial damage provoked by ischemia. Ischemia is mainly caused by the interruption of heart blood flow, which leads to heart infarcts [1,2,3]. During short ischemia and despite of the decrease in oxygen supply, there is a reversible loss of cardiac contractile function. However, when ischemia is sustained for a prolonged period, there is an irreversible cardiac muscle damage resulting in adverse cardiac remodeling [1]. Remodeling is primarily achieved by myocardial fibrosis resulting in decreased cardiac function, impairment of cardiac conduction system and at last arrhythmia. Actually, prompt and rapid myocardial reperfusion reduces significantly myocardial infarct size and improves clinical outcome [4]. Paradoxically, the subsequent reperfusion also activates various injury responses and tissue lesions. This phenomenon is known as Ischemia and Reperfusion (I/R) injury [4]. The absence of oxygen and nutrients during ischemia causes metabolic and biochemical changes. Furthermore, reperfusion provokes calcium overload, oxidative stress, mitochondrial dysfunction and activation of apoptotic and autophagy pathways, which worsen the cardiac remodeling [5,6,7].
Current therapeutic strategies applied in the treatment of myocardial infarction have effectively lowered early mortality from IHD. However, significant number of myocardial infarcted patients still suffers from the adverse left ventricular remodeling and further heart failure progression. For this reason, a better understanding of the pathophysiology of IHD and novel therapeutic strategies to provide more effective monitoring of disease progression are eagerly needed. Non-coding RNAs (ncRNAs) represent one of the increasing areas in the cardiovascular research field [8, 9]. There are different types of ncRNAs according to their sequences length: silencing RNA (siRNAs), small nucleolar RNAs (snoRNAs), microRNAs (miRNAs), long non-coding RNAs (lncRNAs), and the latest Piwi-interacting RNA (pi-RNAs) [10, 11].
There are increasing amount of studies of ncRNAs in recent years explained by the amount of biological functions and pathologies where ncRNAs seem involved [12,13,14]. In heart, ncRNAs regulate a plethora of cellular processes, including cardiomyocyte apoptosis, necrosis and fibrosis [15, 16]. They have been related to different Cardiovascular Diseases (CVD) processes such as atherosclerosis, I/R injury and myocardial infarction. ncRNAs are also examined as sensitive biomarkers for IHD that will allow early prognostic of patients with high risks of post-infarction remodeling and malfunction of the left ventricle [9, 17]. Currently, an extensive list of cardiovascular ncRNAs as well as mRNA targets have been reported. In this review, we will discuss the most relevant ncRNAs involved in I/R and cardioprotection.
2 Classification, Synthesis and Regulation of Non-coding RNAs
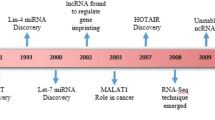
Until 1970s, central dogma of molecular biology established RNA as an intermediate in the process of protein translation from genes encoded in DNA [18]. Later on, this idea was challenged due to new discoveries of RNA molecules and the publication of the result of the International Human Genome Sequencing Consortium [19,20,21]. This consortium stated that approximately 98% of human genome contained non-protein coding sequences. Initially, these non-coding sequences were qualified as “DNA junk”. With the recent emergence of high-throughput technologies and the establishment of new consortiums, like the Encyclopedia of DNA Elements (ENCODE), transcripts generated from these DNA were re-valued and given the importance that they deserved [22, 23]. Nowadays, these transcripts are named ncRNAs [24]. ncRNAs are divided into short non-coding (sncRNAs; <200 nt), including miRNAs, piRNAs, siRNAs as well as snoRNAs and lncRNA (200 nt–100 kb) [10, 11].
2.1 Long Non-coding RNAs
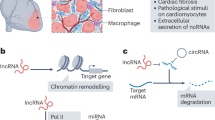
LncRNAs covers a heterogeneous group involved principally in the regulation of transcription at different levels. LncRNAs are transcripts which own a range of nucleotide from 200 nt to over 100 kb [10, 25]. Currently, 392 human lncRNAs are registered and published in the HUGO-Gene Nomenclature Committee (HGNC) (https://www.genenames.org/cgi-bin/genefamilies/set/788) and NONCODE collection of lncRNAs includes a number of 96,308 human lncRNAs gene loci and 172,216 human lncRNAs transcripts (http://www.noncode.org/analysis.php) [11].
LncRNAs can be grouped attending to different criteria, such as their sequence, structure, function, metabolism or interaction with genes and other DNA elements [26]. Nevertheless, a single and acceptable classification remains needed [27]. The most used classification is based on their localization with regard to protein-coding genes. Thus, there are sense lncRNAs, antisense lncRNAs, intronic lncRNAs, intergenic lncRNAs and enhancer lncRNAs [28]. Briefly, sense lncRNAs are located within exons; antisense lncRNAs are synthesized from the antisense DNA strand of protein exons; intronic lncRNAs are produced from protein intron; intergenic lncRNAs are positioned between protein-coding genes; and enhancer lncRNAs, transcripts from enhancer regions of protein-coding genes which can be mono or bidirectional [25, 29,30,31]. Conversely, in reference to their mechanism of action, lncRNAs can act as: signals enabling transcription control like a transcription factor, decoys that bind with effector to prevent their access and action, guides to ribonucleoprotein/chromatin complexes to locate target genes, scaffolds to generate a ribonucleoprotein complex acting as an adapter, and enhancers to build loops that connect enhancer and promoters regions [28, 32]. Furthermore, lncRNAs can also act as a regulator of alternative splicing in three ways. Concisely, lncRNAs interact directly with splicing factors, create a RNA-RNA complex with other pre-mRNA and/or interfere with chromatin remodeling [33]. Likewise, lncRNAs modulate post-transcriptional expression through translation control or altering mRNA stability [34, 35].
In relation to others ncRNAs, lncRNAs can be precursor of sncRNAs such as siRNAs or can control expression and action of miRNAs [32]. Interestingly, circular lncRNAs have been described as “sponges” able to sequestrate miRNAs [36, 37].
2.2 Small Non-coding RNAs
2.2.1 miRNAs
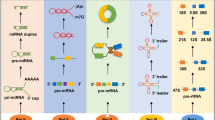
Huge number of studies made special attention to miRNAs within the ncRNAs, due to their high stability and the possibility to quantify easily in biological fluids. Nowadays, more than 2600 human mature miRNAs are known (http://www.mirbase.org/cgi-bin/query.pl?terms=hsa). miRNAs are molecules of sncRNAs (18–25 nt) vastly conserved, which participate in genetic regulation [22, 38, 39]. Generally, transcription of miRNA to primary-miRNAs (pri-miRNAs) is carried out by RNA polymerase II [40, 41]. There are two pathways to complete miRNAs biogenesis: canonical, the most typical pathway, and non-canonical. The pri-miRNA is next endonucleolytically cleaved by the nuclear microprocessor complex formed by the RNase III enzyme Drosha and the DiGeorge critical region 8 (DGCR8) protein Exportin 5/RanGTP complex is the responsible to transport the pre-miRNA to the cytoplasm and then other RNase III, Dicer, cleaves the terminal loop and generates a mature miRNA duplex. Once associated with the Argonaute (AGO) family of proteins, this duplex of RNA removes the passenger strand. Hence, AGO with mature miRNA guide strand conforming miRNA-Induced Silencing Complex (miRISC) [41, 42]. Non-canonical pathways are less characterized; they do not use one of the RNases (DROSHA or DICER) to reach the miRISC construction and use alternative ways and molecules. Interestingly, through this pathway “mirtrons”, pre-miRNAs created from introns of mRNA during splicing are generated [43, 44]. Generally, miRNAs have a guide role within RISC in RNA silencing 3′UTR level, whereas, other seed matches region have been described [45].
2.2.2 siRNAs
siRNAs are 19–24 nt widely used in gene silencing studies, including therapeutic purposes [46,47,48]. This kind of sncRNAs were characterized due to their highly stable double strand of RNA and a perfect complementarity with the target mRNA [49]. siRNAs are transcribed by RNase III, and the rest of the biogenesis is roughly similar to miRNAs biogenesis. Thus, siRNAs conducted the silencing post-transcriptional process through RISC complex too. Interestingly, in the same way as miRNAs, there are evidences suggesting that siRNAs actively participate in epigenetic modifications [50, 51]. Therefore, siRNAs are considered as valuable experimental tools [52]. However, their clinical use still remain limited because of the low efficacy of their delivery to tissue [53].
2.2.3 piRNAs
piRNAs are special sncRNAs with a 26–31 nt of length able to bind with a kind of argonaute proteins, Piwi. The association of Piwi and Piwi-like proteins with piRNAs generates a complex which participates in gene’s expression at epigenetic and post-transcriptional level, mainly in germline and gonadal somatic cells [54,55,56]. piRNAs are single strand sncRNAs with 2′-O-methylation at the 3′end [56]. According to the meiotic phase where piRNAs acts, there are two different sub-clusters called pachytene and pre-pachytene piRNA cluster. Currently, scientists hypothesize, but still with no consensus, about the piRNAs biogenesis. Two main ways comprise primary and secondary amplification cycle, known as “ping-pong cycle” [57]. What is clear is that piRNAs biology is more complex than other ncRNAs, and more studies are necessary to elucidate these emerging tools of genes’ expression.
3 Non-coding RNAs in Ischemia and Reperfusion
Relevant advances have been made in determining the role of ncRNAs in cellular process associated with ischemia. Earlier, most research focus on the role of miRNAs in ischemic responses. Only in recent years and thanks to the advances in OMICs technologies (genomics, transcriptomics, proteomics, metabolomics, and beyond), there is an increasing interest on studying the others ncRNAs. Here, we will highlight role of miRNAs and lncRNAs in responses to I/R, as well as in strategies of cardioprotection.
3.1 Role of miRNAs
miRNAs control many processes in the infarcted heart, such as cardiomyocyte cell death and proliferation, neovascularization and progenitor-cell-mediated repair [25, 58]. Initial attentions have been given to describe the role of miRNA in cellular processes associated with ischemia and/or revascularization in patients undergoing percutaneous coronary intervention and in vitro and in vivo, using animal models of Acute Myocardial Infarcts (AMI) [59, 60]. During the AMI, miRNAs can be up- or down-regulated, having either a pathological or protective role because they are involved in genes’ regulation, inflammation, stress responses, angiogenesis or apoptosis [3, 61,62,63]. Independent reports suggested a protective role of miRNAs, whereas others demonstrated deleterious effects of miRNAs dysregulation in AMI and I/R [3, 64,65,66,67,68].
Since the number of miRNAs related to ischemia and/or reperfusion is substantially increasing, here we will describe the role of only few of them on cell-death and survival.
3.1.1 miRNAs and Cardiomyocytes Survival
Earlier studies highlighted the role of miRNAs in cardiomyocytes survival and the regulation of apoptosis, necrosis or autophagy after considerable duration of ischemia. Gain- and loss-of-function studies were conducted in vivo and in vitro to demonstrate that miRNAs may promote or impair cardiomyocyte survival by regulation of caspases, Bcl-2 family or p53, among other apoptotic signaling pathways [58, 69].
Plenty of miRNAs families have been related to anti-apoptosis effects and cell survival during cardiac I/R as illustrated in Fig. 15.1, such as miR-1, miR-21, miR-24, miR-125, miR-133 or miR-98 [3, 70,71,72,73,74,75,76,77,78] as illustrated in Fig. 15.1. Using experimental model of AMI and I/R previous studies indicated that miR-21 has anti-apoptotic action regulating the called Programmed Cell Death 4 (PDCD4) [70], PTEN/Akt signaling pathway [71] or via Akt and the Bcl2/Bax pathway [72]. miR-1 and miR-133a mimics also attenuate apoptosis by the inhibition of caspase 9. In contrast pre-treatment of rat hearts with antimiR-133a increases caspase-9 and the apoptosis rate induced by I/R. Other reports demonstrated that miR-24 and miR-214 suppress cardiomyocyte apoptosis by Bim-1 repression, and attenuate infarct size in mouse model of AMI [74, 75]. A recent report suggested that miR-214 mimic suppresses the expression and translocation of Bim1 from cytosol to mitochondria and induces Bad phosphorylation, involving PTEN suppression in H9c2 cardiac cell line under I/R [75]. Similarly, miR-93 inhibits cardiomyocyte I/R-mediated apoptosis by targeting PI3K/Akt/PTEN signaling in H9c2 [79]. miR-98 also attenuates apoptosis-induced by I/R in H9c2, by inhibiting of Bcl-2, Bax and caspase-3 among others apoptotic genes [77]. Recently, we demonstrated that the transfection of neonatal cardiac myocyte with miR-125a-3p mimics inhibits the expression of BRCA1 [3].
In contrast, other reports revealed that miRNAs might promote pro-apoptotic action and cell death in ischemic condition as shown in Fig. 15.1. Actually, significant upregulation of miR-15 was observed in the infarcted zone of porcine and mice cardiac tissue in response to ischemic injury, which was associated with cell-death through Bcl-2 activation [80, 81]. Similarly, miR-195 that is related to the miR-15 family, contributes to apoptosis by its downregulation of NAD-dependent protein deacetylase sirtuin-1 (SIRT1) [82]. Another study described that the inhibition of miR-143 with antagomir prevents its pro-apoptotic effects by caspase-3 inhibition and LDH release [83]. miR-29a and miR-29c also negatively regulate cardiac cell survival under I/R, because they increase the expression of Mcl-1, an anti-apoptotic Bcl-2 family member [84]. Indeed, miR-29a or miR-29c downregulation with antagomiRs significantly reduce myocardial infarct size and apoptosis in hearts subjected to I/R injury [84]. Likewise, significant upregulation of miR-34 family (miR-34a, miR-34b, and miR-34c), was observed after AMI. miR-34 family are regulated by cellular tumor antigen p53, and contribute to cardiomyocyte cell death. In fact, the inhibition of the three miR-34 in vivo using antimiRs or antagomiRs improves cardiomyocyte survival after AMI and preserves cardiac contractile function [85, 86].
Altogether, these independent studies demonstrated that gain- and loss-of-functions of some miRNAs may play a pivital role in AMI-mediated cardiac malfunction and cell death.
3.1.2 miRNAs and Cardioprotection
Compelling evidence confirmed the important role of miRNAs under different strategies of cardioprotection as summarized in Fig. 15.2. In fact, several miRNAs are released in patients with AMI after coronary reperfusion with percutaneous coronary intervention, namely miR-1, miR-133a, miR-133b, and miR-499-5p [87]. Interestingly, a protocol using remote ischemic-preconditioning to attenuate myocardial I/R injury releases up to 26 miRNAs in blood sample of anaesthetized patients undergoing coronary artery bypass surgery [88].
Moreover, anesthetic and pain drugs have confirmed cardioprotective effects [11, 89, 90], involving miRNAs activation (90). For example, isoflurane protects mouse and rat hearts from I/R injury by miR-21 activation, involving Akt/nitric oxide synthase pathway [91] and PDCD4 respectively [92]. Fentanyl, a synthetic opiates, also reduces injury evoked by Hypoxia/Reoxygenation (H/R), a simulated in vitro protocol of I/R, through the inhibition of miR-145-5p and Bcl-2 Interacting Protein 3 (BNIP3) [93]. Others studies demonstrated that the administration of the δ-opioid receptor agonist in rats, under normoxic conditions, increases cardiac expression of miR-107-3p, miR-141-3p, and miR-350-5p, while it rises miR-7a/b, miR-107-3p, miR-200b-5p, miR-376a-3p, and miR134-5p levels under hypoxic conditions [94]. Although, the exact contribution of these miRNAs and their targets to cardioprotection have not been examined. Nevertheless, another study demonstrated that the activation of δ-opioid receptor mediated-cardioprotection modifies the expression of 39 miRNAs, while it decreases cell death and LDH levels in isolated cardiomyocytes subjected to H/R (93). This study demonstrated an upregulation of miR-7a-5p which inhibits I/R-induced apoptosis by negatively regulating the expression of PARP (Peroxisome Proliferator-Activated Receptor) [95], and miR-107-3p that regulates HIF-1β stimulation od endothelial progenitor cells differentiation [95]. Other group suggested that miR-133b-5p has a preponderant role in morphine signaling and cardioprotection by targeting Fas gene [96]. Recently, the upregulation of miR-133b-5p was demonstrated to contribute to preconditioning mediated cardioprotection in cardiomyocytes, associated with inhibition of caspase-8 and caspase-3 apoptotic signaling [97].
Others stimuli also changes the expression of miRNA under I/R. For example, pioglitazone, an agonist of PPAR-gamma, protects against myocardial I/R injury by miR-29a and miR-29c downregulation [84]. Recently, we showed that the addition of urocortin at the onset of reperfusion protects the heart from I/R injuries and dysregulates the expression of several miRNAs, such as miR-125a-3p, miR-139 and miR-324-3p [3]. We demonstrated that mimics of miR-125a-3p, miR-324-3p and miR-139-3p modify the expression of genes involved in cell death and apoptosis (BRCA1, BIM, STAT2), in cAMP and Ca2+ signaling (PDE4a, CASQ1), in cell stress (NFAT5, XBP1, MAP 3K12) and in metabolism (CPT2, FoxO1, MTRF1, TAZ). Interestingly, a recent study described that circadian rhythm is involved in ischemia preconditioning through the upregulation of the light elicited-circadian rhythm protein Period 2 (Per2). This study identified miR-21 as cardioprotective downstream target of Per2 [98].
Recently, a study belonging to the discussions of the European Union-CARDIOPROTECTION COST Action, confirmed that the concentration of numerous ncRNAs molecules is altered by ischemia, I/R, conditioning stimuli and medications to conclude that miR-21 and miR-125b are highly relevant for cardioprotection [99].
3.2 Role of LncRNAs in Ischemia and Reperfusion
3.2.1 LncRNAs and Cardiomyocytes Survival
LncRNAs play different roles in cellular physiology. Concretely, they participate in immune responses, chemotaxis, cell death and/or in the production of Reactive Oxygen Species (ROS) in I/R [100,101,102,103]. Actually, an aberrant expression of lncRNAs was observed at early stages of reperfusion in a mouse model of I/R, where the microarray analysis of sample taken from the infracted zone shows differential expression of 151 lncRNAs as compare to sham [100]. Using quantitative-PCR the upregulation of five lncRNAs was confirmed in the infarcted zone, namely; uc007prv.1, AK080112, ENSMUST00000170410, AK156124 and ENSMUST00000166777. Using gene ontology and pathways analyses, authors revealed several target genes for theses lncRNAs, related with immune responses, cytokine activity, NOD-like receptor and chemokine signaling pathways, which have been linked to I/R injury [100].
Recent studies demonstrated that lncRNAs might interact with miRNAs to modulate cell death. For instance, lncRNA Necrosis-Related Factor (NRF) inhibits the expression of miR-873 which blocks RIPK1 and RIPK3, involved in I/R-induced myocardial necrosis [104]. Meanwhile, lncRNA FTX regulates cardiomyocyte apoptosis in I/R animal models, through modulation of the Bcl2l2 expression, which is mediated by miR-29b-1-5 [105]. A recent research demonstrated that the upregulation of lncRNA RMRP exacerbates H/R injury by downregulation of miR-206 and subsequently upregulation of ATG3 in H9c2. In contrast, suppression of RMPR improves cardiac function and inhibited apoptosis after H/R [106]. Other studies also demonstrated the contribution of lncRNAs to apoptosis (Fig. 15.1). For instance, UCA1 stimulates p27 protein and caspase3 in I/R rat model [107] as well as ROR, which aggravates H/R-induced myocardial injury through the stimulation of ROS production and apoptosis in H9c2 cells [101]. Indeed, ROR increases the expression of Bax, cytochrome C, Smac/Diablo, cleaved-caspase-3 and cleaved-caspase-9 expressions, but it also decreases Bcl-2 expression in H9c2 under H/R [101]. Finally, lncRNA E230034O05Rik is considered as effective modulator of autophagy since it repressed autolysosome formation under H/R in in H9c2 [108]. In fact, silencing of this lncRNA markedly decreased autophagy and increased H9c2 myocytes viability during H/R [108].
3.2.2 LncRNAs and Cardioprotection
The role of lncRNAs in cardioprotection has been barely explored. Nevertheless, recent studies suggested that they could be valuable therapeutic target in myocardial I/R (Fig. 15.2). For example, knockdown of lncRNA AK12348 prevents I/R-induced LDH release and inhibits PARP and caspase-3 [109]. Similarly, MALAT1 was suggested as a key mediator of cardioprotective effects of fentanyl against I/R injury. MALAT1 inhibition prevents LDH release and apoptosis, involving miR-145 and BNIP3 axis [93]. Furthermore, suppression of the lncRNA LINC00652 restores sevoflurane-induced cardioprotection. Moreover, its silencing reduces I/R injury and alleviates inflammatory damage by targeting the receptor of Glucagon-Like Peptide-1 (GLP-1), a protein with known anti-oxidative effect on various tissues [110].
4 ncRNAs as Biomarkers for Ischemic CVD
Great interest has arisen toward the potential use of ncRNAs as promising novel biomarkers for the diagnosis and/or prognosis of CVD. Researchers made special emphasis on miRNAs because of their high stability against circulating RNases, their easy detection in human samples obtained through minimal invasive methods (e.g. serum), their specific expression pattern in the disease and their long expectance and easy quantification by quantitative PCR [39, 111,112,113,114]. Precisely, it has been demonstrated that ncRNAs participate actively in the pathophysiology of ischemic CVD [60, 113, 115, 116].
Increasing data are confirming that changes in the expression of miRNAs were observed in plasma of patients with AMI [117,118,119]. Recently, a pilot study examined and compared the expression profile of circulating miRNAs in patients with normal coronary artery, unstable angina and with ST elevation myocardial infarct (STEMI). This study identified 38 miRNAs whose expression level is consistently changed in unstable angina and STEMI patients compared with control patients. This fact indicates dynamic changes of miRNAs expression with the pathogenesis and progression of coronary artery disease [119]. Bioinformatic analysis suggests that target genes of these miRNAs are involved in various biological processes including angiogenesis, inflammation, proliferation, migration and apoptosis [119]. Another study suggested that miRNA-1254 fairly predicts ventricular remodeling at 6 months after STEMI [120]. Similarly, the analysis of circulating miRNAs in patients with an acute coronary syndrome determined that 3 miRNAs (miR-26b-5p, miR-320a and miR-660-5p) are significantly and differentially expressed in patients with Major Cardiovascular Event (MACE), defined as cardiac death or recurrent myocardial infarction, within 1 year of follow-up. These data suggest that these three miRNAs may reflect the activation of molecular pathways that will improve the clinical outcome after STEMI [121].
In the case of lncRNAs, information is limited comparing to miRNAs, however the number of lncRNAs associated with diagnosis and/or prognosis of ischemic CVD keep increasing [122,123,124]. Precisely, changes in the expression of MYHEART, HIF1A-AS2, KCNQ1OT, MALAT1, LIPCAR and UCA1 were proposed as warning sign for the diagnosis of STEMI [125, 126]. Actually, LIPCAR is considered a potential biomarker of STEMI, which could predict the severity and progression of coronary artery disease [126]. Interestingly, another study analyzed the expression of lncRNAs in peripheral blood mononuclear cells to evaluate their role as diagnostic biomarkers to differentiate between STEMI and non-STEMI patients. This study identified 58 lncRNAs and confirmed by qRT-PCR that ENST00000508020.2, LNC_002011, LNC_000303, LNC_000898, ENST00000573866.2 and ENST00000562710.1 are abnormally expressed in mononuclear cells of STEMI compared with non-STEMI participants [127]. Future studies will be helpful to understand whether lncRNAs may serve as a potential noninvasive diagnostic for AMI.
5 Conclusions and Perspectives
ncRNAs have emerged in recent years as key factors in a multitude of pathways across several diseases including CVD and specially IHD. Clinicians are in pursuit of a reliable ncRNAs marker similar to the widely used cardiac troponin, to evaluate the extent of AMI injury. Whereas, it remains challenging to understand which of them are important and how their implication and effect is completely achieved.
We recommend to take in consideration:
-
New bioinformatic tools to predict the ncRNAs-IHD associated. Certainly, they will be helpful in identifying biological functions of ncRNAs in disease prevention, diagnosis and management.
-
Specific ncRNAs-disease associated (and/or -disease severity associated), to be used as novel diagnosis biomarkers.
-
For cardioprotection purpose; specific and reliable method to deliver possible therapeutic ncRNAs to the heart during ischemia, or during primary percutaneous coronary intervention to modulate gene’s expression in the infarcted heart.
-
It can be envisaged that deep understanding of ncRNAs identification, characterization and regulation in cardiovascular health and disease will yield novel therapeutic interventions tailored to the development of patients’ disease.
Abbreviations
- AGO:
-
Argonaute
- AMI:
-
acute myocardial infarct
- BNIP3:
-
Bcl2 interacting protein 3
- CVD:
-
Cardiovascular Disease
- DGCR8:
-
DiGeorge critical Region 8
- ENCODE:
-
Encyclopedia of DNA elements
- GLP-1:
-
glucagon like peptide 1
- HGNC:
-
Hugo Gene Nomenclature Committee
- H/R:
-
hypoxia/reoxygenation
- IHD:
-
Ischemia Heart Disease
- lncRNAs:
-
long non-coding RNAs
- miRISC:
-
miRNA-induced silencing complex
- miRNAs:
-
microRNAs
- ncRNAs:
-
non-coding RNAs
- NRF:
-
necrosis related factor
- PARP:
-
peroxisome proliferator activated receptor
- PDCD4:
-
programme cell death 4
- piRNAs:
-
piwi-interacting RNAs
- pri-miRNA:
-
primary-miRNAs
- ROS:
-
reactive oxygen species
- siRNA:
-
silencing RNAs
- SIRT1:
-
sirtuin-1
- snoRNAs:
-
small nucleolar RNAs
- STEMI:
-
ST Elevation myocardial infarct
References
Pfeffer MA, Braunwald E. Ventricular remodeling after myocardial infarction. Experimental observations and clinical implications. Circulation. 1990;81(4):1161–72.
Thiele H, Akin I, Sandri M. PCI strategies in patients with acute myocardial infarction and cardiogenic shock. N Engl J Med. 2017;377(25):2419–32.
Díaz I, Calderón-Sánchez E, del Toro R, Ávila-Médina J, de Rojas-de Pedro ES, Domínguez-Rodríguez A, Rosado JA, Hmadcha A, Ordóñez A, Smani T. miR-125a, miR-139 and miR-324 contribute to Urocortin protection against myocardial ischemia-reperfusion injury. Sci Rep. 2017;7:8898.
Hausenloy DJ, Garcia-Dorado D, Bøtker HE, et al. Novel targets and future strategies for acute cardioprotection: position paper of the European Society of Cardiology Working Group on Cellular Biology of the Heart. Cardiovasc Res. 2017;113(6):564–85.
Manuscript A, Diseases N. NIH Public. Access. 2014;36:1–17.
Ruiz-Meana M, García-Dorado D. Fisiopatología del daño miocárdico por isquemia-reperfusión: nuevas oportunidades terapéuticas en el infarto agudo de miocardio. Rev Esp Cardiol. 2009;62(2):199–209.
Dhalla NS, Saini HK, Tappia PS, Sethi R, Mengi SA, Gupta SK. Potential role and mechanisms of subcellular remodeling in cardiac dysfunction due to ischemic heart disease. J Cardiovasc Med (Hagerstown). 2007;8(4):238–50.
Giral H, Landmesser U, Kratzer A. Into the wild: GWAS exploration of non-coding RNAs. Front Cardiovasc Med. 2018;5:181.
Hobuß L, Bär C, Thum T. Long non-coding RNAs: at the heart of cardiac dysfunction? Front Physiol. 2019;10:30.
Bhat SA, Ahmad SM, Mumtaz PT, Malik AA, Dar MA, Urwat U, Shah RA, Ganai NA. Long non-coding RNAs: mechanism of action and functional utility. Non-coding RNA Res. 2016;1(1):43–50.
Melo Z, Ishida C, de la Paz Goldaraz M, Rojo R, Echavarria R. Novel roles of non-coding RNAs in opioid signaling and cardioprotection. Non-coding RNA. 2018;4(3):22.
Yang L, Cai Y, Zhang D, Sun J, Xu C, Zhao W, Jiang W, Pan C. miR-195/miR-497 regulate CD274 expression of immune regulatory ligands in triple-negative breast Cancer. J Breast Cancer. 2018;21(4):371.
Heo MJ, Yun J, Kim SG. Role of non-coding RNAs in liver disease progression to hepatocellular carcinoma. Arch Pharm Res. 2019;42(1):48–62.
Chen J-B, Zhu Y-W, Guo X, et al. Microarray expression profiles analysis revealed lncRNA OXCT1-AS1 promoted bladder cancer cell aggressiveness via miR-455-5p/JAK1 signaling. J Cell Physiol. 2019;234(8):13592–601.
Guo Y, Luo F, Liu Q, Xu D. Regulatory non-coding RNAs in acute myocardial infarction. J Cell Mol Med. 2017;21(5):1013–23.
Das A, Samidurai A, Salloum FN. Deciphering non-coding RNAs in cardiovascular health and disease. Front Cardiovasc Med. 2018;5:73.
Wang S-S, Wu L-J, Li J-J-H, Xiao H-B, He Y, Yan Y-X. A meta-analysis of dysregulated miRNAs in coronary heart disease. Life Sci. 2018;215:170–81.
CRICK F. Central dogma of molecular biology. Nature. 1970;227:561–3.
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE, Horvitz HR, Ruvkun G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature. 2000;403(6772):901–6.
Human Genome Sequencing Consortium I. Finishing the euchromatic sequence of the human genome. Nature. 2004;431(7011):931–45.
Mattick JS. The state of long non-coding RNA biology. Non-coding RNA. 2018;4(3):17.
He L, Hannon GJ. Erratum: MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Niu D-K, Jiang L. Can ENCODE tell us how much junk DNA we carry in our genome? Biochem Biophys Res Commun. 2013;430(4):1340–3.
Brosnan CA, Voinnet O. The long and the short of noncoding RNAs. Curr Opin Cell Biol. 2009;21(3):416–25.
Boon RA. Non-coding RNAs in cardiovascular health and disease. Front Cardiovasc Med. 2018;5:71.
Bolha L, Ravnik-Glavač M, Glavač D. Long noncoding RNAs as biomarkers in Cancer. Dis Markers. 2017;2017:1–14.
St Laurent G, Wahlestedt C, Kapranov P. The landscape of long noncoding RNA classification. Trends Genet. 2015;31(5):239–51.
Dhanoa JK, Sethi RS, Verma R, Arora JS, Mukhopadhyay CS. Long non-coding RNA: its evolutionary relics and biological implications in mammals: a review. J Anim Sci Technol. 2018;60:25.
Ma L, Bajic VB, Zhang Z. On the classification of long non-coding RNAs. RNA Biol. 2013;10(6):925–33.
Hangauer MJ, Vaughn IW, McManus MT. Pervasive transcription of the human genome produces thousands of previously unidentified Long intergenic noncoding RNAs. PLoS Genet. 2013;9(6):e1003569.
Kim T-K, Hemberg M, Gray JM. Enhancer RNAs: a class of long noncoding RNAs synthesized at enhancers. Cold Spring Harb Perspect Biol. 2015;7(1):a018622.
Goyal N, Kesharwani D, Datta M. Lnc-ing non-coding RNAs with metabolism and diabetes: roles of lncRNAs. Cell Mol Life Sci. 2018;75(10):1827–37.
Romero-Barrios N, Legascue MF, Benhamed M, Ariel F, Crespi M. Survey and summary splicing regulation by long noncoding RNAs. Nucleic Acids Res. 2018;46(5):2169–84.
Geisler S, Coller J. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol. 2013;14(11):699–712.
Gong C, Maquat LE. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature. 2011;470(7333):284–8.
Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333–8.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495(7441):384–8.
Pasquinelli AE. MicroRNAs: heralds of the noncoding RNA revolution. RNA. 2015;21(4):709–10.
De Gonzalo-Calvo D, Iglesias-Gutié Rrez E, Llorente-Corté V. Epigenetic biomarkers and cardiovascular disease: circulating microRNAs. Rev Esp Cardiol (Engl Ed). 2017;70(9):763–9.
Lee Y, Kim M, Han J, Yeom K-H, Lee S, Baek SH, Kim VN. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004;23(20):4051–60.
Ha M, Kim VN. Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol. 2014;15(8):509–24.
Kawamata T, Tomari Y. Making RISC. Trends Biochem Sci. 2010;35(7):368–76.
Denli AM, Tops BBJ, Plasterk RHA, Ketting RF, Hannon GJ. Processing of primary microRNAs by the microprocessor complex. Nature. 2004;432(7014):231–5.
O’Brien J, Hayder H, Zayed Y, Peng C. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol (Lausanne). 2018;9:402.
Xu W, San Lucas A, Wang Z, Liu Y. Identifying microRNA targets in different gene regions. BMC Bioinformatics. 2014;15:S4.
Karimizadeh E, Motamed N, Mahmoudi M, Jafarinejad-Farsangi S, Jamshidi A, Faridani H, Gharibdoost F. Attenuation of fibrosis with selective inhibition of c-Abl by siRNA in systemic sclerosis dermal fibroblasts. Arch Dermatol Res. 2015;307(2):135–42.
Suzuki K, Yokoyama J, Kawauchi Y, et al. Phase 1 clinical study of siRNA targeting carbohydrate sulphotransferase 15 in Crohn’s disease patients with active mucosal lesions. J Crohns Colitis. 2017;11(2):221–8.
Fitzgerald K, White S, Borodovsky A, et al. A highly durable RNAi therapeutic inhibitor of PCSK9. N Engl J Med. 2017;376(1):41–51.
Elbashir SM, Harborth J, Lendeckel W, Yalcin A, Weber K, Tuschl T. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature. 2001;411(6836):494–8.
Holoch D, Moazed D. RNA-mediated epigenetic regulation of gene expression. Nat Rev Genet. 2015;16(2):71–84.
Wei J-W, Huang K, Yang C, Kang C-S. Non-coding RNAs as regulators in epigenetics. Oncol Rep. 2017;37(1):3–9.
Morris KV, Mattick JS. The rise of regulatory RNA. Nat Rev Genet. 2014;15(6):423–37.
Dana H, Chalbatani GM, Mahmoodzadeh H, et al. Molecular mechanisms and biological functions of siRNA. Int J Biomed Sci. 2017;13(2):48–57.
Cox DN, Chao A, Baker J, Chang L, Qiao D, Lin H. A novel class of evolutionarily conserved genes defined by piwi are essential for stem cell self-renewal. Genes Dev. 1998;12:3715–27.
Kuramochi-Miyagawa S, Kimura T, Ijiri TW, et al. Mili, a mammalian member of piwi family gene, is essential for spermatogenesis. Development. 2004;131:839–49.
Weng W, Li H, Goel A. Piwi-interacting RNAs (piRNAs) and cancer: emerging biological concepts and potential clinical implications. Biochim Biophys Acta Rev Cancer. 2018; https://doi.org/10.1016/J.BBCAN.2018.12.005.
Czech B, Hannon GJ. One loop to rule them all: the ping-pong cycle and piRNA-guided silencing. Trends Biochem Sci. 2016;41:324–37.
Boon RA, Dimmeler S. MicroRNAs in myocardial infarction. Nat Rev Cardiol. 2015;12:135–42.
Eltzschig HK, Eckle T. Ischemia and reperfusion—from mechanism to translation. Nat Med. 2011;17:1391–401.
Su Q, Ye Z, Sun Y, Yang H, Li L. Relationship between circulating miRNA-30e and no-reflow phenomenon in S.TEMI patients undergoing primary coronary intervention. Scand J Clin Lab Investig. 2018;78:318–24.
Arif M, Pandey R, Alam P, Jiang S, Sadayappan S, Paul A, Ahmed RPH. MicroRNA-210-mediated proliferation, survival, and angiogenesis promote cardiac repair post myocardial infarction in rodents. J Mol Med. 2017;95:1369–85.
Yang T, Cao C, Yang J, Liu T, Lei XG, Zhang Z, Xu S. miR-200a-5p regulates myocardial necroptosis induced by Se deficiency via targeting RNF11. Redox Biol. 2018;15:159–69.
Shao H, Yang L, Wang L, Tang B, Wang J, Li Q. MicroRNA-34a protects myocardial cells against ischemia–reperfusion injury through inhibiting autophagy via regulating TNFα expression. Biochem Cell Biol. 2018;96:349–54.
Fan ZX, Yang J. The role of micrornas in regulating myocardial ischemia reperfusion injury. Saudi Med J. 2015;36:787–93.
Lorenzen JM, Batkai S, Thum T. Regulation of cardiac and renal ischemia-reperfusion injury by microRNAs. Free Radic Biol Med. 2013;64:78–84.
Condorelli G, Latronico MVG, Cavarretta E. microRNAs in cardiovascular diseases. J Am Coll Cardiol. 2014;63:2177–87.
Weiss JB, Eisenhardt SU, Stark GB, Bode C, Moser M, Grundmann S. MicroRNAs in ischemia-reperfusion injury. Am J Cardiovasc Dis. 2012;2:237–47.
Zhu H, Fan GC. Role of microRNAs in the reperfused myocardium towards post-infarct remodelling. Cardiovasc Res. 2012;94:284–92.
Sun T, Dong Y-H, Du W, Shi C-Y, Wang K, Tariq M-A, Wang J-X, Li P-F. The role of MicroRNAs in myocardial infarction: from molecular mechanism to clinical application. Int J Mol Sci. 2017;18(4):e745.
Cheng Y, Zhu P, Yang J, Liu X, Dong S, Wang X, Chun B, Zhuang J, Zhang C. Ischaemic preconditioning-regulated miR-21 protects heart against ischaemia/reperfusion injury via anti-apoptosis through its target PDCD4. Cardiovasc Res. 2010;87:431–9.
Tu Y, Wan L, Fan Y, Wang K, Bu L, Huang T, Cheng Z, Shen B. Ischemic postconditioning-mediated miRNA-21 protects against cardiac ischemia/reperfusion injury via PTEN/Akt pathway. PLoS One. 2013;8:e75872.
Ma N, Bai J, Zhang W, Luo H, Zhang X, Liu D, Qiao C. Trimetazidine protects against cardiac ischemia/reperfusion injury via effects on cardiac miRNA-21 expression, Akt and the Bcl-2/Bax pathway. Mol Med Rep. 2016;14(5):4216–22.
Xu X, Kriegel AJ, Jiao X, Liu H, Bai X, Olson J, Liang M, Ding X. miR-21 in ischemia/reperfusion injury: a double-edged sword? Physiol Genomics. 2014;46(21):789–97.
Qian L, Van Laake LW, Huang Y, Liu S, Wendland MF, Srivastava D. miR-24 inhibits apoptosis and represses Bim in mouse cardiomyocytes. J Exp Med. 2011;208(3):549–60.
Wang X, Ha T, Hu Y, Lu C, Liu L, Zhang X, Kao R, Kalbfleisch J, Williams D, Li C. MicroRNA-214 protects against hypoxia/reoxygenation induced cell damage and myocardial ischemia/reperfusion injury via suppression of PTEN and Bim1 expression. Oncotarget. 2016;7(52):86926–36.
Tan H, Qi J, Fan B-Y, Zhang J, Su F-F, Wang H-T. MicroRNA-24-3p attenuates myocardial ischemia/reperfusion injury by suppressing RIPK1 expression in mice. Cell Physiol Biochem. 2018;51(1):46–62.
Zhai CL, Tang GM, Qian G, Hu HL, Wang SJ, Yin D, Zhang S. MicroRNA-98 attenuates cardiac ischemia-reperfusion injury through inhibiting DAPK1 expression. IUBMB Life. 2018;71(2):166–76.
He B, Xiao J, Ren A-J, Zhang Y-F, Zhang H, Chen M, Xie B, Gao X-G, Wang Y-W. Role of miR-1 and miR-133a in myocardial ischemic postconditioning. J Biomed Sci. 2011;18:22.
Ke Z-P, Xu P, Shi Y, Gao A-M. MicroRNA-93 inhibits ischemia-reperfusion induced cardiomyocyte apoptosis by targeting PTEN. Oncotarget. 2016;7(20):28796–805.
Hullinger TG, Montgomery RL, Seto AG, et al. Inhibition of miR-15 protects against cardiac ischemic injury. Circ Res. 2012;110(1):71–81.
Cimmino A, Calin GA, Fabbri M, et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci. 2005;102(39):13944–9.
Zhu H, Yang Y, Wang Y, Li J, Schiller PW, Peng T. MicroRNA-195 promotes palmitate-induced apoptosis in cardiomyocytes by down-regulating Sirt1. Cardiovasc Res. 2011;92(1):75–84.
Hong H, Tao T, Chen S, Liang C, Qiu Y, Zhou Y, Zhang R. MicroRNA-143 promotes cardiac ischemia-mediated mitochondrial impairment by the inhibition of protein kinase Cepsilon. Basic Res Cardiol. 2017;112(60):60.
Ye Y, Hu Z, Lin Y, Zhang C, Perez-Polo JR. Downregulation of microRNA-29 by antisense inhibitors and a PPAR-γ agonist protects against myocardial ischaemia–reperfusion injury. Cardiovasc Res. 2010;87(3):535–44.
Bernardo BC, Gao X-M, Winbanks CE, et al. Therapeutic inhibition of the miR-34 family attenuates pathological cardiac remodeling and improves heart function. Proc Natl Acad Sci U S A. 2012;109(43):17615–20.
Boon RA, Iekushi K, Lechner S, et al. MicroRNA-34a regulates cardiac ageing and function. Nature. 2013;495(7439):107–10.
D’Alessandra Y, Devanna P, Limana F, et al. Circulating microRNAs are new and sensitive biomarkers of myocardial infarction. Eur Heart J. 2010;31(22):2765–73.
Frey UH, Klaassen M, Ochsenfarth C, et al. Remote ischaemic preconditioning increases serum extracellular vesicle concentrations with altered micro-RNA signature in CABG patients. Acta Anaesthesiol Scand. 2018;63(4):483–92.
Lang X-E, Wang X, Jin J-H. Mechanisms of cardioprotection by isoflurane against I/R injury. Front Biosci (Landmark Ed). 2013;18:387–93.
Tanaka K, Kersten JR, Riess ML. Opioid-induced cardioprotection. Curr Pharm Des. 2014;20:5696–705.
Qiao S, Olson JM, Paterson M, et al. MicroRNA-21 mediates isoflurane-induced cardioprotection against ischemia-reperfusion injury via Akt/nitric oxide synthase/mitochondrial permeability transition pore pathway. Anesthesiology. 2015;123(4):786–98.
Olson JM, Yan Y, Bai X, Ge Z-D, Liang M, Kriegel AJ, Twaroski DM, Bosnjak ZJ. Up-regulation of microRNA-21 mediates isoflurane-induced protection of cardiomyocytes. Anesthesiology. 2015;122(4):795–805.
Zhao Z, Hao W, Meng Q, Du X, Lei S, Xia Z. Long non-coding RNA MALAT1 functions as a mediator in cardioprotective effects of fentanyl in myocardial ischemia-reperfusion injury. Cell Biol Int. 2017;41(1):62–70.
Zhi F, Xue L, Shao N, Deng D, Kang X, Chao D, Xu Y, Wang R, Yang Y, Xia Y. δ-opioid receptor activation and microRNA expression in the rat heart under prolonged hypoxia. Cell Physiol Biochem. 2016;39:1118–28.
Meng S, Cao J, Wang L, Zhou Q, Li Y, Shen C, Zhang X, Wang C. MicroRNA 107 partly inhibits endothelial progenitor cells differentiation via HIF-1β. PLoS One. 2012;7(7):e40323.
He S-F, Zhu H-J, Han Z-Y, Wu H, Jin S-Y, Irwin MG, Zhang Y. MicroRNA-133b-5p is involved in cardioprotection of morphine preconditioning in rat cardiomyocytes by targeting Fas. Can J Cardiol. 2016;32(8):996–1007.
Pan Y, Han Z, He S, Yang W, Cheng J, Zhang Y, Chen Z. miR-133b-5p contributes to hypoxic preconditioning-mediated cardioprotection by inhibiting the activation of caspase-8 and caspase-3 in cardiomyocytes. Mol Med Rep. 2018;17(5):7097–104.
Oyama Y, Bartman CM, Gile J, Eckle T. Circadian microRNAs in cardioprotection. Curr Pharm Des. 2017;23(25):3723–30.
Davidson SM, Andreadou I, Barile L, et al. Circulating blood cells and extracellular vesicles in acute cardioprotection. Cardiovasc Res. 2018;115(7):1156–66.
Liu Y, Li G, Lu H, Li W, Li X, Liu H, Li X, Li T, Yu B. Expression profiling and ontology analysis of long noncoding RNAs in post-ischemic heart and their implied roles in ischemia/reperfusion injury. Gene. 2014;543(1):15–21.
Zhang W, Li Y, Wang P. Long non-coding RNA-ROR aggravates myocardial ischemia/reperfusion injury. Braz J Med Biol Res. 2018;51(6):e6555.
Wu X, Zhu H, Zhu S, Hao M, Li Q. IncRNA expression character associated with ischemic reperfusion injury. Mol Med Rep. 2017;16:3745–52.
S yang Y, Tang L, hua ZS. Long noncoding RNAs: new players in ischaemia-reperfusion injury. Heart Lung Circ. 2018;27(3):322–32.
Wang K, Liu F, Liu C-Y, et al. The long noncoding RNA NRF regulates programmed necrosis and myocardial injury during ischemia and reperfusion by targeting miR-873. Cell Death Differ. 2016;23(8):1394–405.
Long B, Li N, Xu X-X, Li X-X, Xu X-J, Guo D, Zhang D, Wu Z-H, Zhang S-Y. Long noncoding RNA FTX regulates cardiomyocyte apoptosis by targeting miR-29b-1-5p and Bcl2l2. Biochem Biophys Res Commun. 2018;495(1):312–8.
Kong F, Jin J, Lv X, Han Y, Liang X, Gao Y, Duan X. Long noncoding RNA RMRP upregulation aggravates myocardial ischemia-reperfusion injury by sponging miR-206 to target ATG3 expression. Biomed Pharmacother. 2019;109:716–25.
Liu Y, Zhou D, Li G, Ming X, Y feng T, Tian J, Lu H, Yu B. Long non coding RNA-UCA1 contributes to cardiomyocyte apoptosis by suppression of p27 expression. Cell Physiol Biochem. 2015;35(5):1986–98.
Huang Z, Ye B, Wang Z, Han J, Lin L, Shan P, Cai X, Huang W. Inhibition of LncRNA-HRIM increases cell viability by regulating autophagy levels during hypoxia/reoxygenation in myocytes. Cell Physiol Biochem. 2018;46(4):1341–51.
Zheng C, Wu Z, Tian L, et al. Long noncoding RNA AK123483 is involved in the regulation of myocardial ischaemia-reperfusion injury by targeting PARP and caspase-3. Heart Lung Circ. 2018;27(5):e51–8.
Zhang S-B, Liu T-J, Pu G-H, Li B-Y, Gao X-Z, Han X-L. Suppression of long non-coding RNA LINC00652 restores sevoflurane-induced cardioprotection against myocardial ischemia-reperfusion injury by targeting GLP-1R through the cAMP/PKA pathway in mice. Cell Physio Biochem. 2018;49(4):1476–91.
Xu Y, Huang R, Gu J, Jiang W. Identification of long non-coding RNAs as novel biomarker and potential therapeutic target for atrial fibrillation in old adults. Oncotarget. 2016;7(10):10803–11.
Backes C, Meese E, Keller A. Specific miRNA disease biomarkers in blood, serum and plasma: challenges and prospects. Mol Diagn Ther. 2016;20(6):509–18.
Vierek J, Thum T. Circulating noncoding RNAs as biomarkers of cardiovascular disease and injury. Circ Res. 2013;120(2):381–99.
Busch A, Eken SM, Maegdefessel L. Prospective and therapeutic screening value of non-coding RNA as biomarkers in cardiovascular disease. Ann Transl Med. 2016;4(12):236.
Liu X, Dong Y, Chen S, Zhang G, Zhang M, Gong Y, Li X. Circulating microRNA-146a and microRNA-21 predict left ventricular remodeling after ST-elevation myocardial infarction. Cardiology. 2015;132(4):233–41.
Vegter EL, Ovchinnikova ES, van Veldhuisen DJ, Jaarsma T, Berezikov E, van der Meer P, Voors AA. Low circulating microRNA levels in heart failure patients are associated with atherosclerotic disease and cardiovascular-related rehospitalizations. Clin Res Cardiol. 2017;106(8):598–609.
Wang G-K, Zhu J-Q, Zhang J-T, Li Q, Li Y, He J, Qin Y-W, Jing Q. Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur Heart J. 2010;31:659–66.
Gidlöf O, Andersson P, van der Pals J, Götberg M, Erlinge D. Cardiospecific microRNA plasma levels correlate with troponin and cardiac function in patients with ST elevation myocardial infarction, are selectively dependent on renal elimination, and can be detected in urine samples. Cardiology. 2011;118(4):217–26.
Zhong Z, Hou J, Zhang Q, Zhong W, Li B, Li C, Liu Z, Yang M, Zhao P. Circulating microRNA expression profiling and bioinformatics analysis of dysregulated microRNAs of patients with coronary artery disease. Medicine (Baltimore). 2018;97(27):e11428.
de Gonzalo-Calvo D, Cediel G, Bär C, et al. Circulating miR-1254 predicts ventricular remodeling in patients with ST-segment-elevation myocardial infarction: a cardiovascular magnetic resonance study. Sci Rep. 2018;8:15115.
Jakob P, Kacprowski T, Briand-Schumacher S, et al. Profiling and validation of circulating microRNAs for cardiovascular events in patients presenting with ST-segment elevation myocardial infarction. Eur Heart J. 2016;38(7):511–5.
Hu H, Wu J, Li D, Zhou J, Yu H, Ma L. Knockdown of lncRNA MALAT1 attenuates acute myocardial infarction through miR-320-Pten axis. Biomed Pharmacother. 2018;106:738–46.
Yan Y, Zhang B, Liu N, Qi C, Xiao Y, Tian X, Li T, Liu B. Circulating long noncoding RNA UCA1 as a novel biomarker of acute myocardial infarction. Biomed Res Int. 2016;2016:8079372.
Greco S, Zaccagnini G, Perfetti A, et al. Long noncoding RNA dysregulation in ischemic heart failure. J Transl Med. 2016;14(1):183.
Sun C, Jiang H, Sun Z, Gui Y, Xia H. Identification of long non-coding RNAs biomarkers for early diagnosis of myocardial infarction from the dysregulated coding-non-coding co-expression network. Oncotarget. 2016;7(45):73541–51.
Li MB, Wang L-FA, Yang X-CC, Xu LA, Li W-MDE, Xia KB, Zhang D-PC, Wu R-NC, Gan Corresponding Author T, Yang X-C. Circulating long noncoding RNA LIPCAR acts as a novel biomarker in patients with ST-segment elevation myocardial infarction. Med Sci Monit. 2018;24:5064–70.
Zhong Z, Hou J, Zhang Q, Li B, Li C, Liu Z, Yang M, Zhong W, Zhao P. Differential expression of circulating long non-coding RNAs in patients with acute myocardial infarction. Medicine (Baltimore). 2018;97(51):e13066.
Funding
This work was supported by Spanish Ministry of Economy and Competitiveness [BFU2016–74932-C2]; Institute of Carlos III [PI15/00203; PI16/00259; CB16/11/00431]; the Andalusia Government [PI-0313-2016]. This study was co-financed by FEDER Funds.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Smani, T. et al. (2020). Non-coding RNAs and Ischemic Cardiovascular Diseases. In: Xiao, J. (eds) Non-coding RNAs in Cardiovascular Diseases. Advances in Experimental Medicine and Biology, vol 1229. Springer, Singapore. https://doi.org/10.1007/978-981-15-1671-9_15
Download citation
DOI: https://doi.org/10.1007/978-981-15-1671-9_15
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1670-2
Online ISBN: 978-981-15-1671-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)