Abstract
Hepatitis B virus (HBV) is a major human pathogen lacking a reliable curative therapy. Current therapeutics target the viral reverse transcriptase/DNA polymerase to inhibit viral replication but generally fail to resolve chronic HBV infections. Due to the limited coding potential of the HBV genome, alternative approaches for the treatment of chronic infections are desperately needed. An alternative approach to the development of antiviral therapeutics is to target cellular gene products that are critical to the viral life cycle. As transcription of the viral genome is an essential step in the viral life cycle, the selective inhibition of viral RNA synthesis is a possible approach for the development of additional therapeutic modalities that might be used in combination with currently available therapies. To address this possibility, a molecular understanding of the relationship between viral transcription and replication is required. The first step is to identify the transcription factors that are the most critical in controlling the levels of HBV RNA synthesis and to determine their in vivo role in viral biosynthesis. Mapping studies in cell culture utilizing reporter gene constructs permitted the identification of both ubiquitous and liver-enriched transcription factors capable of modulating transcription from the four HBV promoters. However, it was challenging to determine their relative importance for viral biosynthesis in the available human hepatoma replication systems. This technical limitation was addressed, in part, by the development of non-hepatoma HBV replication systems where viral biosynthesis was dependent on complementation with exogenously expressed transcription factors. These systems revealed the importance of specific nuclear receptors and hepatocyte nuclear factor 3 (HNF3)/forkhead box A (FoxA) transcription factors for HBV biosynthesis. Furthermore, using the HBV transgenic mouse model of chronic viral infection, the importance of various nuclear receptors and FoxA isoforms could be established in vivo. The availability of this combination of systems now permits a rational approach toward the development of selective host transcription factor inhibitors. This might permit the development of a new class of therapeutics to aid in the treatment and resolution of chronic HBV infections, which currently affects approximately 1 in 30 individuals worldwide and kills up to a million people annually.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
Hepatitis B virus (HBV) infects man and great apes [1,2,3,4,5,6,7,8,9,10,11]. Viral tropism is restricted to the hepatocytes within the liver of the host [12,13,14,15,16,17]. HBV biosynthesis within the liver is noncytopathic [17,18,19]. However, the cellular immune response to HBV antigens synthesized during infection and presented at the cell surface of these hepatocytes in the context of human leukocyte antigens (HLA) results in cell death by T-cell-mediated cytotoxicity, compensating liver regeneration and associated fibrosis [18, 19]. In long-term chronic carriers where these processes have occurred for many years, cirrhosis and end-stage liver diseases can occur [18, 19]. Furthermore, chronic HBV carriers are at much greater risk of developing hepatocellular carcinoma (HCC) [18,19,20,21]. Liver cirrhosis and HCC are associated with significant morbidity and mortality [22]. It is estimated that approximately one in three individuals in the world will be infected with HBV in their lifetime, resulting in about 1 in 30 individuals currently being chronic carriers [23, 24]. This translates into approximately 248 million chronic HBV carriers worldwide today and an associated yearly mortality due to HBV-associated disease of about 600,000 individuals [22,23,24]. Therefore, HBV is a major public health concern, which currently lacks any therapies capable of efficiently resolving chronic infection [25, 26]. Current therapies are limited to type 1 interferons and nucleoside analog drugs, which modulate the immune response and inhibit the HBV reverse transcriptase/DNA polymerase, respectively [25, 26]. As these long-term therapies are generally used to limit disease progression [25, 26], there is an urgent need for additional therapeutic modalities capable of resolving chronic HBV infections within a limited treatment time period.
2 Transcription of the HBV Genome
The cloning and sequencing of HBV genomic DNA identified four open reading frames within the viral 3.2kbp genome [27,28,29,30]. Here, the sequence coordinates of the HBVayw subtype (genotype D [27, 31]) will be used, but the overall genome organization is essentially identical for all replication-competent viral genomes despite modest nucleotide and amino-acid differences among the various genotypes (subtypes) [27,28,29,30]. The core or nucleocapsid open reading frame encodes the hepatitis B early and core antigens, HBeAg and HBcAg, respectively (Fig. 3.1) [1, 32]. HBeAg is synthesized from the first translation initiation codon of the nucleocapsid 212 amino-acid open reading frame [33,34,35,36,37]. The first 19 amino-terminal hydrophobic signal sequence residues are cleaved by the signal peptidase as the precore sequence is translocated into the endoplasmic reticulum [33, 36,37,38,39]. Subsequently, the 34 carboxyl-terminal arginine-rich nuclear localization sequence residues are cleaved from the HBeAg precursor by a furin protease in the Golgi apparatus [33, 40,41,42]. This results in the secretion of a 36 kDa HBeAg protein comprising a dimer of the 159 amino-acid polypeptide generated as a result of the amino- and carboxyl-terminal cleavage events of the product of the complete nucleocapsid open reading frame [43, 44]. The 21 kDa HBcAg polypeptide is synthesized from the second in-frame translation initiation codon of the nucleocapsid open reading frame, which can assemble to generate the viral capsid comprising 120 dimers [33, 45,46,47,48].
Organization of the HBV genome. The circular HBV genome (subtype ayw) is 3182 nucleotides in length. The position of nucleotide coordinates 800 (0.8), 1600 (1.6), 2400 (2.4), and 3182 (3.2/0.0) are indicated. (A) The viral open reading frames (ORFs) are represented by black arrows. Orientation is N terminal to C-terminal for the PS (presurface), S (surface), X (X-gene), PC (precore), C (core), and P (polymerase) ORFs. The direction of transcription (>) from (B) the large surface antigen promoter (PSp), (C) the major surface antigen promoter (Sp), (D) the enhancer 1/X gene promoter (Enh1/Xp), and (E) the enhancer 2/core or nucleocapsid promoter (Enh2/Cp) is shown. Abundant 3.5-kb and 2.1-kb HBV transcripts are indicated by the solid green and blue arrows and the relatively rare 2.4-kb and 0.7-kb transcripts are indicated by the broken brown and purple arrows, respectively. The four transcripts terminate at the single polyadenylation site located around nucleotide coordinate 1940
The surface antigen open reading frame encodes the viral envelope proteins (Fig. 3.1) [1, 27, 33]. There are three in-frame translation initiation codons within this open reading frame, which are translated to produce the large, middle, and major surface antigen proteins, HBsAg [1, 27, 33]. The large surface antigen protein, p39/gp43, includes the 108 amino acid preS1, 55 amino acid preS2, and 226 amino acid major surface antigen domains, whereas the middle surface antigen protein, gp33/gp36, includes only the pres2 and major surface antigen domains [33, 49, 50]. The major surface antigen, p25/gp28, is translated from the third initiation codon and encodes the carboxyl-terminal 226 amino acids of the surface antigen open reading frame [33, 51,52,53]. All three HBsAg translation products are partially glycosylated at asparagine 146 of the major surface antigen open reading frame, whereas asparagine 4 of the pres2 domain present in the middle surface antigen polypeptide is completely glycosylated [54,55,56,57]. This gives rise to the six different forms of the HBsAg polypeptide present in the virus particles [58].
The HBV viral genome encodes two additional open reading frames. The HBV reverse transcriptase/DNA polymerase open reading frame encodes a 94 kDa polypeptide with three major domains (Fig. 3.1) [27]. The amino-terminal domain of this open reading frame encodes the terminal protein, which serves as the primer for HBV minus-strand DNA synthesis [59,60,61,62,63]. The middle domain encodes the reverse transcriptase/DNA polymerase activity, while the carboxyl-terminal domain encodes for the RNaseH activity responsible for the degradation of the viral pregenomic RNA during the process of minus-strand DNA synthesis [64,65,66,67,68,69,70,71]. The smallest open reading frame in the viral genome codes for a 154 amino-acid polypeptide, HBxAg (Fig. 3.1) [27]. The 17 kDa X-gene open reading frame encodes a protein that is essential for productive viral infection in vivo and has been ascribed a large variety of functions when assayed under various conditions [72,73,74,75]. Currently, it is unclear which, if any, of these functions explains the requirement for this protein for productive infection in vivo.
Analysis of the HBV viral transcripts during natural infection of humans and chimpanzees has been modest due to the limited availability of liver samples. However, two predominant viral transcripts of 3.5 kb and 2.1 kb have been detected during natural infection (Fig. 3.1) [12,13,14,15,16]. Furthermore, analysis of viral transcripts present in cells transfected with HBV genomic DNA and HBV transgenic mice has permitted a more detailed analysis of the transcripts derived from viral genomes. In addition to the major transcripts, two additional unspliced HBV RNAs of 2.4 and 0.7 kb have been routinely described in a variety of systems that can support viral biosynthesis (Fig. 3.1) [76,77,78,79,80,81,82,83,84,85,86,87,88,89]. The 3.5 kb HBV transcripts identified by RNA filter hybridization analysis represent two distinct transcripts, the precore and pregenomic RNAs, as determined by 5′-end mapping studies, which differ by approximately 36 nucleotides (Fig. 3.2) [76,77,78, 90]. The 3.5 kb HBV precore RNA initiates at a cluster of sites centered at approximately nucleotide coordinate 1785 and its translation from the initiation codon at nucleotide 1816 results in the synthesis of HBeAg [76,77,78, 90]. The 3.5 kb HBV pregenomic or core RNA initiates at a cluster of sites centered at approximately nucleotide coordinate 1821 and its translation from the initiation codon at nucleotide 1903 results in the synthesis of HBcAg [76,77,78, 90]. The 3.5 kb HBV pregenomic RNA is also translated from an internal initiation codon at nucleotide 2309, which results in the synthesis of the viral reverse transcriptase/DNA polymerase polypeptide although this presumably occurs at a much lower frequency than translation of the HBcAg polypeptide [65, 91, 92]. In this manner, the structural HBcAg is synthesized at a level much greater than the viral polymerase, which supports efficient viral biosynthesis. Furthermore, the HBV polymerase recognizes the RNA stem/loop/bulge structure, epsilon (ε), at the 5′-end of the 3.5 kb pregenomic RNA as it is being translated from the ribosome and forms a ribonucleoprotein complex, which is encapsidated by HBcAg to generate immature core particles [69, 93,94,95]. Within these immature core particles, the viral polymerase reverse transcribes the 3.5 kb pregenomic RNA to generate the mature core particle containing the 3.2 kb relaxed circular HBV DNA genome [93, 94]. Mature core particles can bind to envelope antigen, HBsAg, located within the membrane of the endoplasmic reticulum and subsequently bud into the lumen to be secreted from the hepatocytes by transit through the Golgi apparatus [96,97,98,99,100,101]. Alternatively, mature capsids can cycle viral genomes back into the nucleus to amplify and/or replenish the pool of HBV covalently closed circular DNA (HBV cccDNA) that represents the template for transcription by the host RNA polymerase II [79, 102].
Nucleotide sequence of the HBV genomic DNA (subtype ayw) showing the location of the transcription factors binding to the enhancer 1/X-gene promoter, enhancer 2/core promoter region, the intragenic core gene sequence, the large surface antigen promoter, and the major surface antigen promoter [27]. The nucleotide coordinates are derived from the GenBank database (ID: V01460). The orientation of the direct repeat sequences homologous to the consensus nuclear receptor–binding site direct repeat sequence 5′-AGGTCA-3′ are indicated with arrows. The underlined sequences in the enhancer 1/X-gene promoter region indicate the location of the CCAAT/enhancer-binding protein-binding sites (C/EBP) [139], the p53 tumor suppressor gene product–binding site (p53) [140], the interferon regulatory factor–binding site (IRF) [141], the nuclear factor 1–binding sites (NF1) [142, 143], the forkhead box protein A/hepatocyte nuclear factor 3–binding sites (FOXA/HNF3) [144, 145], the hepatocyte nuclear factor 4–binding site (HNF4) [127], the retinoid X receptor plus the peroxisome proliferator-activated receptor heterodimer–binding site (RXR:PPAR) [127, 128, 146], the COUPTF-binding site (COUPTF) [120, 127], the RFX1-binding site (RFX1) [127, 147, 148], the activator protein 1–binding site (AP1) [143], the cyclic AMP response element–binding protein-binding site (CREB) [149], and the activating transcription factor 2–binding site (ATF2) [149]. The underlined sequences in the enhancer 2/core promoter region represent the RFX1-binding site (RFX1) [108], the Sp1-binding sites (Sp1) [109], the CCAAT/enhancer-binding protein-binding site (C/EBP) [110, 111], the retinoid X receptor plus the farnesoid X receptor heterodimer–binding site (RXR:FXR) [115,116,117,118], the liver receptor homolog 1/fetoprotein transcription factor–binding sites (LRH1/FTF) [112,113,114], the hepatic leukemia factor–binding site (HLF) [113], the E4BP4-binding site (E4BP4) [119], the hepatocyte nuclear factor 4–binding sites (HNF4) [120, 121], the forkhead box protein A/hepatocyte nuclear factor 3–binding sites (FOXA/HNF3) [122], the retinoid X receptor plus the peroxisome proliferator-activated receptor heterodimer–binding site (RXR:PPAR) [120], the COUPTF binding site (COUPTF) [120, 123, 124], the estrogen-related receptor (ERR) [117, 125], and the TATA-box-binding protein (TBP) site [126]. The underlined sequences in the intragenic core gene region spanning nucleotide coordinates 2110 to 2200 sequence indicate the location of the Sp1-binding sites (Sp1), the forkhead box protein A/hepatocyte nuclear factor 3–binding site (FOXA/HNF3), and the hepatocyte nuclear factor 4–binding site (HNF4). The underlined sequences in the large surface antigen promoter region indicate the location of the hepatocyte nuclear factor 1–binding sites (HNF1) [129, 130], the forkhead box protein A/hepatocyte nuclear factor 3–binding site (FOXA/HNF3) [131], the Sp1-binding sites (Sp1) [132], and the TATA-box-binding protein (TBP) site [133]. The underlined sequences in the major surface antigen promoter region indicate the location of the forkhead box protein A/hepatocyte nuclear factor 3–binding sites (FOXA/HNF3) [134], the nuclear factor 1–binding site (NF1) [135, 136], the Sp1-binding sites (Sp1) [137], and the nuclear factor Y–binding site (NF-Y) [138]. The approximate positions of the major transcription start sites are indicated by solid circles plus arrows indicating the direction of transcription. The transcription polyadenylation signal sequence, 5′-UAUAAA-3′, and the sights of polyadenylation for the viral RNAs are indicated with open and closed boxes, respectively. The protein translation initiation codons for the seven HBV polypeptides are indicated with solid triangles
The 2.1 kb HBV transcripts identified by RNA filter hybridization analysis appear to initiate synthesis at a cluster of locations positioned between nucleotide coordinates 3156 and 8, as determined by 5′-end mapping studies (Fig. 3.2) [76,77,78, 103]. As a consequence of the heterogeneous nature of the transcription start sites and their proximity to the preS2 initiation codon at nucleotide coordinate 3174, the 2.1 kb HBV surface antigen RNA is translated to a rather modest degree from the preS2 initiation codon at nucleotide coordinate 3174 to produce limited amounts of the middle HBsAg polypeptide and is robustly translated from the initiation codon at nucleotide coordinate 157 to produce large quantities of the major surface antigen protein [76,77,78, 103]. The minor 2.4 kb HBV presurface RNA initiates at a cluster of sites centered at approximately nucleotide coordinate 2809 and its translation from the initiation codon at nucleotide 2850 results in the synthesis of a limited amount of the large surface antigen polypeptide [76,77,78]. Consequently the large, middle, and major HBsAg polypeptides are synthesized at appropriate ratios to support the synthesis of virus particles, which require the large surface antigen polypeptide, plus orders of magnitude more subviral particles, which are present in the sera of infected individuals [58]. The 0.7 kb HBV X-gene RNA, which has been observed in some cell culture systems, HBV transgenic mice, and infected liver tissues, appears to initiate at multiple sites spanning nucleotide coordinates 1157 and 1340 and its translation from the initiation codon at nucleotide 1376 could result in the synthesis of the HBV X-gene polypeptide [89, 104,105,106]. The X-gene-encoded protein product has not been convincingly demonstrated in natural infection although antibodies to this polypeptide have been detected in the sera of chronic HBV carriers [89, 107]. Therefore, it is not apparent if the HBV X-gene polypeptide is encoded by its own transcript during natural infection, translated from one or more of the larger HBV RNAs by internal ribosome entry, or translated from a minor spliced HBV transcript. For all of the HBV transcripts, polyadenylation of the viral RNAs occurs between nucleotide coordinates 1936 and 1943, mediated in part by the nonconventional polyA recognition sequence, 5′-UAUAAA-3′, located between nucleotide coordinates 1918 and 1923 [85, 103].
3 Cis-Acting Transcriptional Regulatory Sequence Elements and Trans-Acting DNA-Binding Proteins
The cloning of the HBV genome and the mapping of the transcripts suggested that there were likely to be four transcriptional regulatory regions controlling viral RNA synthesis. With the extensive use of reporter gene constructs and transfection analysis utilizing both hepatoma and nonheptoma cell lines, the cis-acting transcriptional regulatory sequence elements within the viral genome were mapped in detail by deletion and mutational analysis (Fig. 3.2). Sequences of 70–240 nucleotides located upstream of the transcription start sites for each of the viral transcripts were shown to correspond to the enhancer/promoter regions governing the levels of RNA synthesis. The 200 base pair region located between nucleotide coordinates 1600 and 1800 bound both ubiquitous (RFX1, SP1, COUPTF, ERR, and TBP) and liver-enriched (C/EBP, LRH1, HNF4, RXR, FXR, FOXA and PPAR) transcription factors, which contributed to the level of the nucleocapsid or core promoter activity that directs the expression of the HBV 3.5 kb precore and pregenomic RNAs encoding the HBeAg and HBcAg polypeptides [108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128]. The 70 base pair region located between nucleotide coordinates 2720 and 2790 bound both ubiquitous (SP1 and TBP) and liver-enriched (FOXA and HNF1) transcription factors, which contribute to the level of the presurface antigen promoter activity that directs the expression of the HBV 2.4 kb presurface antigen RNA encoding the large surface antigen [129,130,131,132,133]. The 240 base pair region located between nucleotide coordinates 2910 and 3150 bound both ubiquitous (NF1, SP1 and NF-Y) and liver-enriched (FOXA) transcription factors, which contributed to the level of the surface antigen promoter activity that directs the expression of the HBV 2.1 kb surface antigen RNAs encoding the middle and major surface antigens [134,135,136,137,138]. Similarly, the 220 base pair region located between nucleotide coordinates 1020 and 1240 bound both ubiquitous (p53, IRF, NF1, COUPTF, RFX1, AP1, CREB, and ATF2) and liver-enriched (C/EBP, FOXA, HNF4, RXR and PPAR) transcription factors, which contributed to the level of the X-gene promoter activity that may direct the expression of the HBV 0.7 kb X-gene RNAs encoding the X-gene polypeptide [120, 127, 139,140,141,142,143,144,145,146,147,148,149]. Furthermore, the X-gene and core promoter regions can act as enhancer sequences, enhancer 1 and 2, respectively, under certain circumstances leading to increased transcription from the other HBV promoters [104, 112, 113, 145, 146, 150,151,152,153,154,155,156,157,158,159,160,161,162,163,164,165,166,167,168]. The enhancer function of the X-gene and core promoter regions may be important for the coordinated liver-specific expression of the HBV transcripts. Similarly, the contribution of individual transcription factors to multiple HBV promoter activities may also control the coordinate expression of the various transcripts at levels appropriate for viral biosynthesis. For example, FOXA binds to and regulates expression from all four HBV promoters to various extents [131]. However, the large surface antigen promoter is considerably weaker than the other HBV promoters due, in part, to its limited number of transcription factor–binding sites [166]. This ensures that limiting amounts of the large surface antigen are synthesized and hence prevents the inhibition of viral secretion due to the overproduction of surface antigen tubules that can limit viral envelope secretion [33, 49, 169].
The mapping of the cis-acting promoter sequences permitted the identification of regulatory sequence elements that were transcriptionally active only in hepatoma cells and not in nonhepatoma cells [87, 90, 105, 136, 150, 151, 158, 159, 164, 166, 167, 170,171,172,173,174,175,176,177,178,179,180]. These regulatory elements bound liver-enriched transcription factors, whereas the promoter regulatory elements that were transcriptionally active in both cell types bound more ubiquitously expressed transcription factors. Combinations of DNaseI footprinting and electrophoretic mobility shift assays (EMSAs) using cell extracts and purified factors permitted the identification of many of the trans-acting transcription factors binding to the HBV promoter regulatory sequence elements [109, 120,121,122, 124, 129,130,131,132,133,134, 136, 137, 150, 157, 161, 165, 170, 177, 178, 181,182,183,184,185,186]. Functional validation of the roles of the identified DNA binding proteins in governing the activities of the HBV promoter was evaluated using transfection analysis of wild-type and mutant HBV reporter gene constructs in the presence of exogenously expressed transcription factors [109, 120, 121, 124, 129, 131, 133, 137, 157, 178, 183, 184, 186, 187]. These studies led to a relatively comprehensive map of the HBV enhances/promoters and the functional importance of the transcription factors that bind to these regulatory sequence elements (Fig. 3.2). Despite generating a relatively comprehensive map of the cis-acting regulatory sequence elements governing viral RNA synthesis and the trans-acting factors that bound to them, it remained unclear what the relative importance of the various identified transcription factors might be for HBV biosynthesis, either in cell culture or in vivo.
4 Role of Liver-Enriched Transcription Factors in HBV Transcription, Replication, and Tissue Tropism
The mapping of transcription factor–binding sites to the viral promoters permitted their role in controlling HBV RNA synthesis to be evaluated in the context of viral replication. A significant limitation of these studies arose, because robust viral replication could only be observed in a very limited number of hepatoma cell lines where all the necessary factors for HBV biosynthesis were present [76, 80, 82, 188]. This meant that the effects of exogenously expressed transcription factors on viral transcription were typically rather modest [115, 189]. Furthermore, it was challenging to map these modest effects to specific transcription factor–binding sites by mutational analysis, in part, because of the redundancies in the transcriptional regulation of HBV RNA synthesis and the potential effects of mutations on the viral-coding capacity or cis-acting sequences involved in the regulation of viral replication. The use of short interfering RNA (siRNA) technologies to reduce specific transcription factor abundances in hepatoma cells also has limited utility because of the functional redundancies in the DNA-binding proteins regulating HBV biosynthesis. For these reasons, it became necessary to develop additional approaches to study the effects of specific transcription factors on HBV RNA synthesis, and consequently viral replication.
HBV does not replicate in nonhepatoma cells, presumably because these cells lack the specific transcription factors necessary to support the synthesis of the 3.5 kb RNA from the viral core promoter [189, 190]. The suggestion was supported by the observation that viral 3.5 kb pregenomic RNA synthesis driven by the cytomegalovirus (CMV) immediate early promoter was sufficient to support robust HBV replication in nonhepatoma cells [191]. These findings suggested that complementation of HBV genomic DNA with the appropriate liver-enriched transcription factors in nonhepatoma cells represented an approach to identifying the roles of specific transcription factors in the synthesis of HBV 3.5 kb pregenomic RNA and hence viral replication [189]. Indeed, this approach identified nuclear receptors as the sole class of transcription factors capable of robustly activating viral RNA and DNA syntheses in nonhepatoma cells [117, 189]. This approach indicated that HNF4, RXR, PPAR, FXR, and LRH1 represented liver-enriched nuclear receptors capable of supporting viral biosynthesis in nonhepatoma cells and hence likely contributed in a significant manner to the hepatocyte-specific tropism of HBV [117, 189]. The suggestion that HBV tropism is determined at the level of HBV 3.5 kb pregenomic RNA transcription is strongly supported by the tissue-specific expression pattern observed in the HBV transgenic mouse model of chronic viral infection [17]. In this model, viral transcription and biosynthesis are largely restricted to tissues where these transcription factors are highly expressed with lower levels of transcription being observed in tissues where these transcription factors are expressed at more modest levels [17, 192].
The development of the nonhepatoma cell system for the analysis of the transcriptional regulation of HBV biosynthesis identified nuclear receptors as the only transcription factors capable of supporting viral biosynthesis in this system [117, 189]. Furthermore, most of the activity was shown to map to the proximal nuclear receptor binding site located within the core promoter region [117, 189]. However, it was unclear what the role of the other liver-enriched transcription factors known to bind to the viral promoters might be in governing HBV transcription and replication. To date, none of the other liver-enriched transcription factors, except FoxA/HNF3, have been shown to modulate HBV biosynthesis in nonhepatoma cells [189]. In the nonhepatoma cell viral biosynthesis system, FoxA/HNF3 has been shown to antagonize nuclear receptor–mediated HBV transcription and replication [189, 191]. FoxA-/HNF3-mediated reduction in viral biosynthesis involves both HBeAg-mediated inhibition of HBV biosynthesis, possibly by reducing the efficiency of capsid assembly, plus inhibition of RNA elongation presumably by interfering with RNA polymerase II transcription through the viral promoters located within the DNA regions encoding the HBV 3.5 kb pregenomic RNA [191].
5 Redundant Functions for Nuclear Receptors in HBV Biosynthesis
The use of the nonhepatoma cell system permitted the identification of multiple nuclear receptors capable of supporting HBV biosynthesis due to their ability to bind to the viral nucleocapsid promoter and direct the expression of the HBV 3.5 kb pregenomic RNA [117, 189]. These observations may, in part, explain the difficulties in determining which transcription factors contribute most to HBV transcription in hepatoma cells and hepatocytes in vivo [118, 125, 193,194,195]. Differentiated hepatoma cells express a variety of liver-enriched transcription factors and support HBV transcription and replication [76, 80, 82, 188]. Consequently, ectopic expression of liver-enriched transcription factors can only enhance HBV transcription to a modest extent [115, 189]. Furthermore, reduction or elimination of any specific transcription factor involved in HBV RNA synthesis also only has a very modest effect as there are additional transcription factors capable of substituting for the loss of any particular transcription factor [196]. This situation has also been observed in vivo where individual nuclear receptor-null HBV transgenic mice have displayed only modest perturbations in HBV biosynthesis. Specifically, the PPARα-null HBV transgenic mice displayed wild-type levels of HBV RNA and DNA under control conditions but failed to show enhanced biosynthesis when challenged with PPARα agonists [192, 193]. In contrast, liver-specific HNF4-null HBV transgenic mice displayed a complete loss of viral biosynthesis, indicating that this nuclear receptor was a major determinant of the developmental expression of HBV RNA [195]. However, early neonatal loss of HNF4 expression affects the abundance of additional nuclear receptors (and liver-enriched transcription factors), which are potentially critical for robust HBV RNA synthesis [197], making it unclear the degree to which HNF4 plays a direct or indirect role in the developmental regulation of HBV expression [195].
FXR has also been implicated in the regulation of HBV biosynthesis [115, 117, 118]. However, treatment of HBV transgenic mice with bile acids has only a very limited effect on viral biosynthesis [196]. This effect was not dependent upon inhibition by the corepressor, small heterodimer partner (SHP), as SHP-null HBV transgenic mice have a similar phenotype to their wild-type controls whether or not they were fed a diet supplemented with bile acids [196]. The redundant function of multiple nuclear receptors may explain these observations [117, 196]. In the nonhepatoma cell system, HNF4 and FXR are both capable of independently activating HBV biosynthesis [117]. In the presence of HNF4, FXR can only modestly modulate HBV biosynthesis accounting for the in vivo observations [196]. Therefore, the development of the nonhepatoma cell–based HBV replication system has permitted the reconstitution of viral biosynthesis and the demonstration of the redundant mechanisms, which probably operate in vivo to govern the level of viral transcription under different physiologically relevant conditions [196, 198].
6 Regulation of HBV Biosynthesis by Transcriptional Coactivators and Corepressors
The development of the nonhepatoma cell HBV replication system permitted a more detailed investigation of the potential roles of coactivator and corepressor proteins in the regulation of HBV transcription and replication [118, 125, 194, 199,200,201]. These studies demonstrated that the coactivators, PGC1α, CBP, SRC1, and PRMT1, and the corepressor, SHP, were all capable of modulating HBV transcription to some degree depending on cellular context [118, 125, 194, 199, 200]. Furthermore, the observation that PGC1α and SHP could modulate the nuclear receptor–dependent HBV biosynthesis in nonhepatoma cells further indica ted the potential importance of various nuclear receptors in the transcriptional regulation of viral biosynthesis [118, 125, 194, 199, 200].
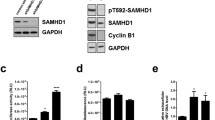
Examination of PGC1α-dependent HBV biosynthesis in the nonhepatoma cell line, HEK293T, in the absence of exogenously expressed nuclear receptors revealed two major aspects of the transcriptional regulation of viral DNA synthesis [200]. First, PGC1α acted as an adapter molecule for the recruitment of additional coactivators in the absence of exogenously expressed nuclear receptors in this particular cell line [200]. This indicated that the endogenous coactivators present in HEK293T cells that were unrelated to the PGC1 family of coactivators were unable to activate HBV 3.5 kb RNA synthesis independently of PGC1α [200]. Therefore, the recruitment of additional coactivators was PGC1-dependant and mutational analysis suggested that PGC1 was recruited to the HBV nucleocapsid promoter, at least in part, through endogenous nuclear receptors present in HEK293T cells [200]. In addition to serving as an adaptor molecule for the recruitment of additional coactivator proteins, PGC1α enhanced HBV transcription in HEK293T cells such that these cells could now support robust viral replication [200]. Detailed analysis of the mechanism governing this observation demonstrated that the concentration of HBcAg passed a critical threshold necessary for core dimers to cooperatively form viral capsids (Fig. 3.3) [200, 202, 203]. Therefore, this cell culture system demonstrated compelling evidence that very modest changes in HBV 3.5 kb pregenomic RNA synthesis that led to less than a two-fold increase in HBcAg were, nonetheless, associated with a dramatic increase in viral DNA synthesis [200]. This finding showed the absence of a linear relationship between core protein synthesis and capsid-associated HBV biosynthesis, which is a critical observation that should be considered when evaluating the transcriptional regulation of viral replication.
The HBV replication cycle showing the intracellular pathway for the synthesis and secretion of HBV, HBsAg, and HBeAg polypeptides. (a) Lower levels of the HBV pregenomic 3.5 kb RNA preclude cytoplasmic dimer oligomerization, immature capsid formation, and hence HBV DNA synthesis. (b) Modestly higher levels of the HBV pregenomic 3.5 kb RNA permit cytoplasmic dimer oligomerization, immature capsid formation, and hence HBV DNA synthesis
The observation that the activities of coactivators and corepressors, which were shown to modulate HBV biosynthesis, are responsive to changes in metabolic cellular states led to the suggestion that viral transcription and replication might be modulated by the physiological state of the cell [201]. Indeed, based on these types of observations, the term “metabolovirus” was suggested to describe the potential relationship between HBV biosynthesis and the metabolic state of the cell [204]. This suggestion is supported by the observations that PGC1α activity is enhanced in vivo by fasting [198, 205,206,207,208] and SHP activity is induced by bile acids [196, 209,210,211], demonstrating a direct relationship between metabolic challenges and coactivator and corepressor activities. However, despite these observations, there is very limited evidence linking metabolic perturbations to changes in specific coactivator- or corepressor-mediated changes in HBV biosynthesis in either hepatoma cell lines or animal models of chronic viral infection [116, 196, 198, 201]. This may reflect the lack of importance of this form of regulation of HBV biosynthesis or the presence of multiple compensating mechanisms that maintain the homeostatic regulation of viral RNA and DNA as the relative abundances of coactivators and corepressor activities change in response to altering physiological conditions.
7 Transcriptional Regulation of HBV Replication In Vivo
As HBV animal infection models are essentially limited to man, chimpanzees, and tree shrews [3, 4, 7, 212,213,214,215], a detailed understanding of the transcriptional regulation of HBV replication in vivo has been very challenging. None of the available models of HBV infection are suitable to investigate the in vivo relevance of the transcriptional regulation of viral biosynthesis revealed in cell culture analysis. The small animal models of hepadnavirus infection including the woodchuck hepatitis virus (WHV) and the duck hepatitis B virus (DHBV) are also not useful models to understand the transcriptional regulation of HBV biosynthesis in vivo as the transcription of both WHV and DHBV is regulated in a distinct manner from HBV [216,217,218]. For all these reasons, the HBV transgenic mouse model of chronic HBV infection represents the most relevant and tractable small animal model for the study of the transcriptional regulation of HBV biosynthesis in vivo [17, 219]. In this model, a single replication competent HBV genome comprising 1.3 copies of the HBVayw DNA sequence has been incorporated into the mouse germline [17]. Consequently, every cell in the HBV transgenic mouse carries the viral transgene, which obviates the species barrier associated with viral infection. Furthermore, the HBV transgene is highly transcribed only in the tissues expressing the liver-enriched transcription factors identified in cell culture studies to control viral RNA synthesis [17, 192, 220]. Therefore, it appears that the HBV transgenic mouse model is a system that probably reflects quite closely the transcriptional regulation of HBV biosynthesis observed in the liver during natural infection. Furthermore, in the absence of any alternative in vivo model system for the investigation of the transcriptional regulation of HBV biosynthesis, it is appropriate to utilize this system to support findings in cell culture. As many of the observations in cell culture have been validated in the HBV transgenic mouse model, it is reasonable to assume that they probably reflect, in part, the situation in natural infection under certain circumstances.
A concern regarding the HBV transgenic mouse model was the absence of nuclear HBV cccDNA and hence the possibility that aspects of the viral life cycle in addition to infection were absent from this system [17]. The alternative explanation for the absence of HBV cccDNA was that cycling of capsids back to the nucleus was not preferred as a result of the high level of surface antigen expression and the large surface antigen in particular that is essential for capsid envelopment and virion secretion through the endoplasmic reticulum to the Golgi apparatus secretion system [221]. Based on this assumption, the HNF1α-null HBV transgenic mouse was created and analyzed [79, 222]. HNF1α regulates the level of expression from the large surface antigen promoter, so loss of HNF1α should be associated with a reduction in the level of HBV 2.4 kb RNA and hence translation of the large surface antigen polypeptide [129, 223]. This was predicted to lead to a reduction in virion production and the recycling of newly synthesized capsids to the nuclei [102, 224]. Interestingly, intracellular viral replication intermediates increased within the livers of HNF1α-null HBV transgenic mice despite very limited effects on HBV RNA synthesis [79]. Most interestingly, HBV cccDNA was readily apparent in these mice, demonstrating that recycling of capsids occurs in this HBV transgenic mouse model of chronic infection [79]. Furthermore, enhanced levels of viral replication were observed despite very limited changes in HBV transcription, supporting the contention that small changes in viral RNAs can be associated with large effects on DNA replication intermediates [79, 200].
As analysis in nonhepatoma cells indicated that nuclear receptors were major determinants of viral tropism, because they were critical for HBV 3.5 kb pregenomic RNA synthesis [189], it was of interest to determine their role in the in vivo regulation of HBV biosynthesis. Initially, PPARα-null HBV transgenic mice were characterized [193]. These mice displayed no major effect on HBV biosynthesis, indicating that PPARα did not contribute to viral transcription and replication under normal physiological conditions [193]. However, activation of PPARα by the agonists, clofibric acid and Wy-14,643, enhanced HBV biosynthesis in the liver of wild type but not PPARα-null HBV transgenic mice [193]. This finding demonstrated that activated PPARα can enhance the basal level of HBV biosynthesis observed in HBV transgenic mice [193]. As plasticizers and some drugs used to treat hypertriglyceridemia can activate PPARα, it seems possible that exposure to these compounds might affect viral loads and disease state of chronic HBV carriers due to their effects on viral biosynthesis [225, 226]. Furthermore, it was noted that the effect of PPARα activation in the HBV transgenic mouse activated viral DNA synthesis considerably more than RNA synthesis, suggesting that modest increases in transcription in vivo may be associated with much larger increase in viral replication as also recently observed in cell culture [193, 200].
Liver-specific HNF4α-null HBV transgenic mice died by postnatal day 15 [227]. The absence of HNF4α expression in the livers of these mice was associated with a dramatic loss in the increase in HBV biosynthesis observed during early neonatal development [227]. As HNF4α is a major contributor to the liver-specific transcriptional network that defines the hepatocyte phenotype [197], it is not clear if the effect of HNF4α on HBV biosynthesis is direct or indirect. However, the in vivo loss of HBV RNA and DNA synthesis associated with the absence of HNF4α expression is consistent with a direct role for this nuclear receptor in viral biosynthesis, as observed in the nonhepatoma replication system [189]. Furthermore, the observed increase in the developmental expression of HNF4α correlates with a similar developmental increase in HBV biosynthesis, supporting its potentially direct role in viral transcriptional regulation in vivo [189, 197]. However, the developmental expression HNF4α in the liver also supports the expression of additional transcription factors including LRH1, RXRα, FXRα, and FoxA2, which are also important regulators of HBV transcription and replication [189, 197]. If any of these transcription factors are critical determinants of viral biosynthesis, the effects of HNF4α on HBV RNA and DNA synthesis in vivo might be indirect rather than direct [189, 197].
Analysis of the liver-enriched transcription factors capable of complementing HBV transcription in nonhepatoma cells indicated that only nuclear receptors could independently activate HBV biosynthesis [117, 189]. This raised the interesting issue of the role of the other liver-enriched transcription factors in HBV biosynthesis in this system. Only FoxA/HNF3 modulated nuclear receptor-mediated biosynthesis in this system [189]. Indeed, it appeared that FoxA mediated its effects by preferentially reducing the expression of the HBV 3.5 kb pregenomic RNA at the level of transcriptional elongation, presumably due to its binding to the presurface, surface, X-gene and nucleocapsid promoters that are intragenic with respect to the transcription of the pregenomic RNA [191]. To address the in vivo relevance of these observations, HBV biosynthesis was determined in the liver-specific FoxA2/HNF3β-overexpressing HBV transgenic mouse [228]. As observed in the nonhepatoma cells, overexpression of FoxA2/HNF3β in the liver of the HBV transgenic mouse resulted in a dramatic reduction in HBV biosynthesis [228]. In this case, a large decrease in HBV replication was associated with a more modest reduction in viral transcription [228]. This observation suggests that the viral biosynthesis in the HBV transgenic mouse is positioned such than the small changes in HBV RNA synthesis result in limited effects on core polypeptide synthesis, which, due to the cooperative nature of capsid assembly, have a dramatic effect on capsid-dependent reverse transcription of pregenomic RNA in a manner similar to that recently reported in cell culture [200].
Since FoxA/HNF3 overexpression in the HBV transgenic mouse was associated with the loss of viral replication, it was of interest to determine the in vivo effect of the loss of FoxA/HNF3 on HBV biosynthesis [198, 220]. The FoxA3/HNF3γ-null HBV transgenic mouse displayed a very limited phenotype, suggesting that the other FoxA/HNF3 isoforms in the liver were either compensating for the loss FoxA3/HNF3γ or FoxA3/HNF3γ was relatively unimportant for HBV biosynthesis [198]. Consequently, a FoxA/HNF3-deficient HBV transgenic mouse expressing only a single FoxA3/HNF3γ allele was generated and characterized [220]. This mouse was viable and displayed no overt phenotype despite biliary epithelial cell proliferation, stellate cell activation, and bridging fibrosis within the liver [220, 229]. However, HBV transcription and replication were essentially absent within the liver [220]. Indeed, the HBV transgene had been permanently transcriptionally silenced due to DNA methylation of its non-CpG island sequences [220]. This observation indicated that the pioneer transcription factor, FoxA/HNF3, was essential for the demethylation of the HBV transgene during liver development and this may account, in part, for the observed increase in HBV biosynthesis during postnatal liver maturation [220, 227]. Further studies are required to determine when FoxA/HNF3 marks the HBV genome for demethylation during liver development and whether this process involves active demethylation by ten-eleven translocation (TET) methylcytosine dioxygenase-mediated oxidation of the 5-methylcytosine residues or passive demethylation involving DNA methyltransferase (DNMT) inhibition in the presence of chromosome replication [230]. Regardless of the mechanism of action of FoxA/HNF3, these observations suggest that targeting FoxA/HNF3 at the appropriate stage of liver development might lead to permanent DNA methylation and inactivation of HBV cccDNA as a transcriptional template necessary for viral biosynthesis and hence might represent a therapeutic target for the resolution of neonatal (and possibly adult) chronic infections.
8 Conclusions
HBV is a significant human pathogen responsible for approximately 600,000 deaths annually [22,23,24]. Current therapies are not curative and nucleoside-analog drugs target a single viral protein, the HBV reverse transcriptase/DNA polymerase, leading to the selection of drug-resistant variants [231, 232]. Additional therapeutic targets are urgently needed to address this unmet need. Unfortunately, due to the small size of the viral genome and hence limited coding capacity, there are only a very limited number of HBV proteins that might serve as potential additional targets for the development of antiviral therapeutics. The HBV core antigen is a potential target and compounds affecting capsid assembly and/or function have been identified, but, to date, they have not been developed into therapeutic modalities [233,234,235,236,237,238].
Given the challenges with the development of antiviral therapeutics targeting viral proteins, an alternative approach is to target cellular gene products that are vital for the viral life cycle but are dispensable at some level for host viability. In this regard, our current understanding of the transcriptional regulation of HBV biosynthesis offers some cellular therapeutic targets that might potentially be exploited for the development of antiviral compounds. Nuclear receptors are ligand-dependent transcription factors governing the level of HBV 3.5 kb pregenomic RNA synthesis. Antagonists targeting HNF4α, PPARα, FXRα, or LRH1 could potentially lead to a reduction in HBV biosynthesis especially if viral transcription is reduced to a level where HBcAg dimers are expressed below the level required to support capsid assembly [200]. The limitations of nuclear receptors as antiviral targets include the functional redundancy resulting from multiple nuclear receptors governing HBV 3.5 kb pregenomic RNA synthesis and the possible undesirable effects on host metabolic function associated with their reduced activities, which might induce cellular toxicity. Targeting FoxA transcription factors at the appropriate developmental stage might be more challenging but potentially more therapeutically beneficial. Transient inhibition of FoxA activity during early neonatal development could potentially lead to the DNA methylation of viral genomes transmitted from mother to child at birth. This could lead to the transcriptional inactivation of the HBV cccDNA, which effectively and permanently terminates viral biosynthesis with the functional eradication of the viral replication intermediate that is refractory to current therapeutic modalities. The major challenge with this approach is the difficulty in effectively targeting FoxA while limiting any possible long-term negative effects on normal cellular and tissue physiology. Regardless of these challenges, the study of the transcriptional regulation of HBV biosynthesis has revealed several interesting aspects of both HBV and liver developmental biology while indicating a number of potential approaches to the development of novel therapeutic modalities targeting host gene products. Going forward, it is hoped that combinations of current and future therapies might result in effective treatments, leading to the resolution of chronic HBV infections and ultimately the worldwide eradication of this devastating human pathogen.
References
Seeger C, Mason WS (2015) Molecular biology of hepatitis B virus infection. Virology 479–480:672–686
Vaudin M, Wolstenholme AJ, Tsiquay KN, Zuckerman AJ, Harrison TJ (1988) The complete nucleotide sequence of the genome of a hepatitis B virus isolated from a naturally infected chimpanzee. J Gen Virol 69:1383–1389
Lichter EA (1969) Chimpanzee antibodies to Australia antigen. Nature 224:810–811
Maynard JE, Hartwell WV, Berquist KR (1971) Hepatitis-associated antigen in chimpanzees. J Infect Dis 123:660–664
Hirschman RJ, Shulman R, Barker LF, Smith KO (1969) Virus-like particles in sera of patients with infectious and serum hepatitis. JAMA 208:1667–1670
Barker LF, Maynard JE, Purcell RH, Hoofnagle JH, Berquist KR, London WT (1975) Viral hepatitis, type B, in experimental animals. Am J Med Sci 270:189–194
Maynard JE, Berquist KR, Krushak DH, Purcell RH (1972) Experimental infection of chimpanzees with the virus of hepatitis B. Nature 237:514–515
Acs G, Sells MA, Purcell RH, Price P, Engle R, Shapiro M et al (1987) Hepatitis B virus produced by transfected HepG2 cells causes hepatitis in chimpanzees. Proc Natl Acad Sci USA 84:4641–4644
Sureau C, Eichberg JW, Hubbard GB, Romet-Lemonne JL, Essex M (1988) A molecularly cloned hepatitis B virus produced in vitro is infectious in a chimpanzee. J Virol 62:3064–3067
Thornton SM, Walker S, Zuckerman JN (2001) Management of hepatitis B virus infections in two gibbons and a western lowland gorilla in a zoological collection. Vet Rec 149(4):113–115
Warren KS, Heeney JL, Swan RA, Heriyanto, Verschoor EJ (1999) A new group of hepadnaviruses naturally infecting orangutans (Pongo pygmaeus). J Virol 73(9):7860–7865
Cattaneo R, Will H, Hernandez N, Schaller H (1983) Signals regulating hepatitis B surface antigen transcription. Nature 305:336–338
Cattaneo R, Will H, Schaller H (1984) Hepatitis B virus transcription in the infected liver. EMBO J 3:2191–2196
Yokosuka O, Omata M, Imazeki F, Ito Y, Okuda K (1986) Hepatitis B virus RNA transcripts and DNA in chronic liver disease. N Engl J Med 315:1187–1192
Imazeki F, Yaginuma K, Omata M, Okuda K, Kobayashi M, Koike K (1987) RNA transcripts of hepatitis B virus in hepatocellular carcinoma. Hepatology 7:753–757
T-S S, Lui W-Y, Lin L-H, Han S-H, P’eng F-K (1989) Analysis of hepatitis B virus transcripts in infected human livers. Hepatology 9:180–185
Guidotti LG, Matzke B, Schaller H, Chisari FV (1995) High-level hepatitis B virus replication in transgenic mice. J Virol 69:6158–6169
Ganem D (1982) Persistent infection of humans with hepatitis B virus: mechanisms and consequences. Rev Infect Dis 4:1026–1047
Chisari FV, Ferrari C (1995) Hepatitis B virus Immunopathogenesis. Annu Rev Immunol 13(1):29–60
Beasley RP (1988) Hepatitis B virus: the major etiology of hepatocellular carcinoma. Cancer 61:1942–1956
Beasley RP, Hwang L-Y, Lin C-C, Chien C-S (1981) Hepatocellular carcinoma and hepatitis B virus – a prospective study of 22707 men in Taiwan. Lancet 2:1129–1133
Goldstein ST, Zhou F, Hadler SC, Bell BP, Mast EE, Margolis HS (2005) A mathematical model to estimate global hepatitis B disease burden and vaccination impact. Int J Epidemiol 34(6):1329–1339
Ott JJ, Stevens GA, Groeger J, Wiersma ST (2012) Global epidemiology of hepatitis B virus infection: new estimates of age-specific HBsAg seroprevalence and endemicity. Vaccine 30(12):2212–2219
Schweitzer A, Horn J, Mikolajczyk RT, Krause G, Ott JJ (2015) Estimations of worldwide prevalence of chronic hepatitis B virus infection: a systematic review of data published between 1965 and 2013. Lancet 386(10003):1546–1555
Terrault NA, Bzowej NH, Chang K-M, Hwang JP, Jonas MM, Murad MH (2016) AASLD guidelines for treatment of chronic hepatitis B. Hepatology 63(1):261–283
Wong DK-H, Seto W-K, Fung J, Ip P, Huang F-Y, Lai C-L et al (2013) Reduction of hepatitis B surface antigen and covalently closed circular DNA by nucleos(t)ide analogues of different potency. Clin Gastroenterol Hepatol 11(8):1004–10.e1
Galibert F, Mandart E, Fitoussi F, Tiollais P, Charnay P (1979) Nucleotide sequence of the hepatitis B virus genome (subtype ayw) cloned in E. coli. Nature 281:646–650
Valenzuela P, Quiroga M, Zaldivar J, Gray P, Rutter WJ (1980) The nucleotide sequence of the hepatitis B viral genome and the identification of the major viral genes. In: Fields B, Jaenisch R, Fox CF (eds) Animal virus genetics. Academic, New York, pp 57–70
Ono Y, Onda H, Sasada R, Igarashi K, Sugino Y, Nishioka K (1983) The complete nucleotide sequences of the cloned hepatitis B virus DNA: subtype adr and adw. Nucleic Acids Res 11:1747–1757
Okamoto H, Imai M, Shimozaki M, Hoshi Y, Iizuka H, Gotanda T et al (1986) Nucleotide sequence of a cloned hepatitis B virus genome, subtype ayr: Comparison with genomes of the other three subtypes. J Gen Virol 67:2305–2314
Kramvis A, Kew M, François G (2005) Hepatitis B virus genotypes. Vaccine 23(19):2409–2423
Lamontagne RJ, Bagga S, Bouchard MJ (2016) Hepatitis B virus molecular biology and pathogenesis. Hepatoma Res 2:163–186
McLachlan A, Milich DR, Raney AK, Riggs MG, Hughes JL, Sorge J et al (1987) Expression of hepatitis B virus surface and core antigens: influences of pre-s and precore sequences. J Virol 61:683–692
Ou JH, Laub O, Rutter WJ (1986) Hepatitis B virus gene function: the precore region targets the core antigen to cellular membranes and causes the secretion of the e antigen. Proc Natl Acad Sci USA 83:1578–1582
Roossinck MJ, Jameel S, Loukin SH, Siddiqui A (1986) Expression of hepatitis B viral core region in mammalian cells. Mol Cell Biol 6:1393–1400
Bruss V, Gerlich WH (1988) Formation of transmembranous hepatitis B e-antigen by cotranslational in vitro processing of the viral precore protein. Virology 163:268–275
Garcia PD, Ou J-H, Rutter WJ, Walter P (1988) Targeting of the hepatitis B virus precore protein to the endoplasmic reticulum membrane: after signal peptide cleavage translocation can be aborted and the product released into the cytoplasm. J Cell Biol 106:1093–1104
Standring DN, Ou J-H, Masiarz FR, Rutter WJ (1988) A signal peptide encoded within the precore region of hepatitis B virus directs the secretion of heterogeneous population of e antigens in Xenopus oocytes. Proc Natl Acad Sci USA 85:8405–8409
Takahashi K, Kishimoto S, Ohori K, Yoshizawa H, Machida A, Ohnuma H et al (1991) Molecular heterogeneity of e antigen polypeptides in sera from carriers of hepatitis B virus. J Immunol 147:3156–3160
Takahashi K, Machida A, Funatsu G, Nomura M, Usuda S, Aoyagi S et al (1983) Immunochemical structure of hepatitis B e antigen in the serum. J Immunol 130:2903–2907
Messageot F, Salhi S, Eon P, Rossignol JM (2003) Proteolytic processing of the hepatitis B virus e antigen precursor – cleavage at two furin consensus sequences. J Biol Chem 278(2):891–895
Ito K, Kim KH, Lok AS-F, Tong S (2009) Characterization of genotype-specific carboxyl-terminal cleavage sites of hepatitis B virus e antigen precursor and identification of furin as the candidate enzyme. J Virol 83(8):3507–3517
DiMattia Michael A, Watts Norman R, Stahl Stephen J, Grimes Jonathan M, Steven Alasdair C, Stuart David I et al (2013) Antigenic switching of hepatitis B virus by alternative dimerization of the capsid protein. Structure 21(1):133–142
Watts NR, Vethanayagam JG, Ferns RB, Tedder RS, Harris A, Stahl SJ et al (2010) Molecular basis for the high degree of antigenic cross-reactivity between hepatitis B virus capsids (HBcAg) and dimeric capsid-related protein (HBeAg): insights into the enigmatic nature of the e-antigen. J Mol Biol 398(4):530–541
Conway JF, Cheng N, Zlotnick A, Wingfield PT, Stahl SJ, Steven AC (1997) Visualization of a 4-helix bundle in the hepatitis B virus capsid by cryo-electron microscopy. Nature 386:91–94
Wynne SA, Crowther RA, Leslie AGW (1999) The crystal structure of the human hepatitis B virus capsid. Mol Cell 3(6):771–780
Roossinck MJ, Siddiqui A (1987) In vivo phosphorylation and protein analysis of hepatitis B virus core antigen. J Virol 61:955–961
Weimer T, Salfeld J, Will H (1987) Expression of the hepatitis B virus core gene in vitro and in vivo. J Virol 61:3109–3113
Persing DH, Varmus HE, Ganem D (1986) Inhibition of secretion of hepatitis B surface antigen by a related presurface polypeptide. Science 234:1388–1391
Cheng K-C, Moss B (1987) Selective synthesis and secretion of particles composed of the hepatitis B virus middle surface protein directed by a recombinant vaccinia virus: induction of antibodies to pre-S and S epitopes. J Virol 61:1286–1290
Crowley CW, Liu CC, Levinson AD (1983) Plasmid-directed synthesis of hepatitis B surface antigen in monkey cells. Mol Cell Biol 3(1):44–55
Hsiung N, Fitts R, Wilson S, Milne A, Hamer DH (1984) Efficient production of hepatitis B surface antigen using a bovine papilloma virus-metallothionein vector. J Mol App Gen 2:497–506
Moriarty AM, Hoyer BH, Shih JW-K, Gerin JL, Hamer DH (1981) Expression of the hepatitis B virus surface antigen gene in cell culture by using a simian virus 40 vector. Proc Natl Acad Sci USA 78:2606–2610
Schmitt S, Glebe D, Alving K, Tolle TK, Linder M, Geyer H et al (1999) Analysis of the pre-S2 N- and O-linked glycans of the M surface protein from human hepatitis B virus. J Biol Chem 274(17):11945–11957
Schmitt S, Glebe D, Tolle TK, Lochnit G, Linder D, Geyer R et al (2004) Structure of pre-S2 N- and O-linked glycans in surface proteins from different genotypes of hepatitis B virus. J Gen Virol 85(7):2045–2053
Mehta A, Lu XY, Block TM, Blumberg BS, Dwek RA (1997) Hepatitis B virus (HBV) envelope glycoproteins vary drastically in their sensitivity to glycan processing: evidence that alteration of a single N-linked glycosylation site can regulate HBV secretion. Proc Natl Acad Sci USA 94:1822–1827
Machida A, Kishimoto S, Ohnuma H, Miyamoto H, baba K, Oad K et al (1982) A glycopeptide containing 15 amino acid residues derived from hepatitis B surface antigen particles: demonstration of immunogenicity to raise anti-HBs in mice. Mol Immunol 19:1087–1093
Heermann KH, Goldmann U, Schwartz W, Seyffarth T, Baumgarten H, Gerlich WH (1984) Large surface proteins of hepatitis B virus containing the pre-s sequence. J Virol 52:396–402
Bartenschlager R, Schaller H (1988) The amino-terminal domain of the hepadnaviral P-gene encodes the terminal protein (genome-linked protein) believed to prime reverse transcription. EMBO J 7:4185–4192
Zoulim F, Seeger C (1994) Reverse transcription in hepatitis B viruses is primed by a tyrosine residue of the polymerase. J Virol 68:6–13
Wang G-H, Seeger C (1992) The reverse transcriptase of hepatitis B virus acts as a protein primer for viral DNA synthesis. Cell 71:663–670
Clark DN, Flanagan JM, Hu J (2017) Mapping of functional subdomains in the terminal protein domain of hepatitis B virus polymerase. J Virol 91(3):e01785-16
Weber M, Bronsema V, Bartos H, Bosserhoff A, Bartenschlager R, Schaller H (1994) Hepadnavirus P protein utilizes a tyrosine residue in the TP domain to prime reverse transcription. J Virol 68:2994–2999
Bavand M, Feitelson M, Laub O (1989) The hepatitis B virus-associated reverse transcriptase is encoded by the viral pol gene. J Virol 63:1019–1021
Chang L-J, Pryciak P, Ganem D, Varmus HE (1989) Biosynthesis of the reverse transcriptase of hepatitis B viruses involves de novo translational initiation not ribosomal frameshifting. Nature 337:364–368
Wang G-H, Zoulim F, Leber EH, Kitson J, Seeger C (1994) Role of RNA in enzymatic activity of the reverse transcriptase of hepatitis B viruses. J Virol 68:8437–8442
Lanford RE, Notvall L, Lee H, Beames B (1997) Transcomplementation of nucleotide priming and reverse transcription between independently expressed TP and RT domains of the hepatitis B virus reverse transcriptase. J Virol 71:2996–3004
Chen Y, Marion PL (1996) Amino acids essential for RNase H activity of hepadnaviruses are also required for efficient elongation of minus-strand viral DNA. J Virol 70:6151–6156
Radziwill G, Tucker W, Schaller H (1990) Mutational analysis of the hepatitis B virus P gene product: domain structure and RNase activity. J Virol 64:613–620
Wei X, Peterson DL (1996) Expression, purification, and characterization of an active RNase H domain of the hepatitis B viral polymerase. J Biol Chem 271:32617–32622
Ko C, Shin Y-C, Park W-J, Kim S, Kim J, Ryu W-S (2014) Residues Arg703, Asp777, and Arg781 of the RNase H domain of hepatitis B virus polymerase are critical for viral DNA synthesis. J Virol 88(1):154–163
Zoulim F, Saputelli J, Seeger C (1994) Woodchuck hepatitis virus X protein is required for viral infection in vivo. J Virol 68:2026–2030
Chen H-S, Kaneko S, Girones R, Anderson RW, Hornbuckle WE, Tennant BC et al (1993) The woodchuck hepatitis virus X gene is important for establishment of virus infection in woodchucks. J Virol 67:1218–1226
Slagle BL, Bouchard MJ (2016) Hepatitis B virus X and regulation of viral gene expression. Cold Spring Harb Perspect Med 6(3):a021402
Slagle BL, Andrisani OM, Bouchard MJ, Lee CGL, Ou JHJ, Siddiqui A (2015) Technical standards for hepatitis B virus X protein (HBx) research. Hepatology 61(4):1416–1424
Yaginuma K, Shirakata Y, Kobayashi M, Koike K (1987) Hepatitis B virus (HBV) particles are produced in a cell culture system by transient expression of transfected HBV DNA. Proc Natl Acad Sci USA 84:2678–2682
Sells MA, Zelent AZ, Shvartsman M, Acs G (1988) Replicative intermediates of hepatitis B virus in HepG2 cells that produce infectious virions. J Virol 62:2836–2844
Farza H, Hadchouel M, Scotto J, Tiollais P, Babinet C, Pourcel C (1988) Replication and gene expression of hepatitis B virus in a transgenic mouse that contains the complete viral genome. J Virol 62:4144–4152
Raney AK, Eggers CM, Kline EF, Guidotti LG, Pontoglio M, Yaniv M et al (2001) Nuclear covalently closed circular viral genomic DNA in the liver of hepatocyte nuclear factor 1α-null hepatitis B virus transgenic mice. J Virol 75(6):2900–2911
Sureau C, Romet-Lemonne J-L, Mullins JI, Essex M (1986) Production of hepatitis B virus by a differentiated human hepatoma cell line after transfection with cloned circular HBV DNA. Cell 47:37–47
Tsurimoto T, Fujiyama A, Matsubara K (1987) Stable expression and replication of hepatitis B virus genome in an integrated state in a human hepatoma cell line transfected with the cloned viral DNA. Proc Natl Acad Sci USA 84:444–448
Chang C, K-S J, C-P H, Lo SJ, T-S S, Ting L-P et al (1987) Production of hepatitis B virus in vitro by transient expression of cloned HBV DNA in a hepatoma cell line. EMBO J 6:675–680
Kaneko S, Miller RH (1988) X-region-specific transcript in mammalian hepatitis B virus-infected liver. J Virol 62:3979–3984
Araki K, Miyazaki J-I, Hino O, Tomita N, Chisaka O, Matsubara K et al (1989) Expression and replication of hepatitis B virus genome in transgenic mice. Proc Natl Acad Sci USA 86:207–211
Simonsen CC, Levinson AD (1983) Analysis of processing and polyadenylation signals of the hepatitis B virus surface antigen gene by using simian virus 40-hepatitis B virus chimeric plasmids. Mol Cell Biol 3:2250–2258
Gough NM (1983) Core and e antigen synthesis in rodent cells transformed with hepatitis B virus DNA is associated with greater than genome length viral messenger RNAs. J Mol Biol 165:683–699
Siddiqui A, Jameel S, Mapoles J (1986) Transcriptional control elements of hepatitis B surface antigen gene. Proc Natl Acad Sci USA 83:566–570
Saito I, Oya Y, Shimojo H (1986) Novel RNA family structure of hepatitis B virus expressed in human cells, using a helper-free adenovirus vector. J Virol 58:554–560
Siddiqui A, Jameel S, Mapoles J (1987) Expression of the hepatitis B virus X gene in mammalian cells. Proc Natl Acad Sci USA 84:2513–2517
Honigwachs J, Faktor O, Dikstein R, Shaul Y, Laub O (1989) Liver-specific expression of hepatitis B virus is determined by the combined action of the core gene promoter and the enhancer. J Virol 63:919–924
Roychoudhury S, Shih C (1990) Cis rescue of a mutated reverse transcriptase gene of human hepatitis B virus by creation of an internal ATG. J Virol 64:1063–1069
Schlicht H-J, Radziwill G, Schaller H (1989) Synthesis and encapsidation of duck hepatitis B virus reverse transcriptase do not require formation of core-polymerase fusion proteins. Cell 56:85–92
Summers J, Mason WS (1982) Replication of the genome of a hepatitis B-like virus by reverse transcription of an RNA intermediate. Cell 29:403–415
Will H, Reiser W, Weimer T, Pfaff E, Buscher M, Sprengle R et al (1987) Replication strategy of human hepatitis B virus. J Virol 61:904–911
Hirsch RC, Lavine JE, Chang L-J, Varmus HE, Ganem D (1990) Polymerase gene products of hepatitis B viruses are required for genomic RNA packaging as well as for reverse transcription. Nature 344:552–555
Stein O, Fainaru M, Stein Y (1972) Visualization of virus-like particles in endoplasmic reticulum of hepatocytes of Australia antigen carriers. Lab Invest 26:262–269
Huang S-N, Groh V (1973) Immunoagglutination electron microscopic study on virus-like particles and Australia antigen in liver tissue. Lab Invest 29:353–366
Yamada G, Nakane PK (1977) Hepatitis B core and surface antigens in liver tissue. Lab Invest 36:649–659
Yamada G, Sakamoto Y, Mizuno M, Nishihara T, Kobayashi T, Takahashi T et al (1982) Electron and immunoelectron microscopic study of Dane particle formation in chronic hepatitis B virus infection. Gastroenterology 83:348–356
Gerber MA, Sells MA, Chen M-L, Thung SN, Tabibzadeh SS, Hood A et al (1988) Morphologic, immunohistochemical, and ultrastructural studies of the production of hepatitis B virus in vitro. Lab Invest 59:173–180
Kamimura T, Yoshikawa A, Ichida F, Sasaki H (1981) Electron microscopic studies of Dane particles in hepatocytes with special reference to intracellular development of Dane particles and their relation with HBeAg in serum. Hepatology 1:392–397
Tuttleman JS, Pourcel C, Summers J (1986) Formation of the pool of covalently closed circular viral DNA in hepadnavirus-infected cells. Cell 47:451–460
Standring DN, Rutter WJ, Varmus HE, Ganem D (1984) Transcription of the hepatitis B surface antigen gene in cultured murine cells initiates within the presurface region. J Virol 50:563–571
Hu K-Q, Siddiqui A (1991) Regulation of the hepatitis B virus gene expression by the enhancer element I. Virology 181:721–726
Treinin M, Laub O (1987) Identification of a promoter element located upstream from the hepatitis B virus X gene. Mol Cell Biol 7:545–548
Tokusumi Y, Ma Y, Song X, Jacobson RH, Takada S (2007) The new core promoter element XCPE1 (X core promoter element 1) directs activator-, mediator-, and TATA-binding protein-dependent but TFIID-independent RNA polymerase II transcription from TATA-less promoters. Mol Cell Biol 27(5):1844–1858
Moriarty AM, Alexander H, Lerner RA, Thornton GB (1985) Antibodies to peptides detect new hepatitis B antigen: serological correlation with hepatocellular carcinoma. Science 227:429–433
Buckwold VE, Chen M, Ou JH (1997) Interaction of transcription factors RFX1 and MIBP1 with the gamma motif of the negative regulatory element of the hepatitis B virus core promoter. Virology 227:515–518
Zhang P, Raney AK, McLachlan A (1993) Characterization of functional Sp1 transcription factor binding sites in the hepatitis B virus nucleocapsid promoter. J Virol 67:1472–1481
Yuh C-H, Ting L-P (1991) C/EBP-like proteins binding to the functional box-α and box-β of the second enhancer of hepatitis B virus. Mol Cell Biol 11:5044–5052
Choi BH, Park GT, Rho HM (1999) Interaction of hepatitis B viral X protein and CCAAT/enhancer-binding protein α synergistically activates the hepatitis B viral enhancer II pregenomic promoter. J Biol Chem 274(5):2858–2865
Li M, Xie YH, Kong YY, Wu X, Zhu L, Wang Y (1998) Cloning and characterization of a novel human hepatocyte transcription factor, hB1F, which finds and activates enhancer II of hepatitis B virus. J Biol Chem 273(44):29022–29031
Ishida H, Ueda K, Ohkawa K, Kanazawa Y, Hosui A, Nakanishi F et al (2000) Identification of multiple transcription factors, HLF, FTF, and E4BP4, controlling hepatitis B virus enhancer II. J Virol 74(3):1241–1251
Gilbert S, Galarneau L, Lamontagne A, Roy S, Bélanger L (2000) The hepatitis B virus core promoter is strongly activated by the liver nuclear receptor fetoprotein transcription factor or by ectopically expressed steroidogenic factor 1. J Virol 74(11):5032–5039
Ramiere C, Scholtes C, Diaz O, Icard V, Perrin-Cocon L, Trabaud MA et al (2008) Transactivation of the hepatitis B virus core promoter by the nuclear receptor FXRα. J Virol 82(21):10832–10840
Reese VC, Oropeza CE, McLachlan A (2013) Independent activation of hepatitis B virus biosynthesis by retinoids, peroxisome proliferators, and bile acids. J Virol 87(2):991–997
Reese VC, Ondracek CR, Rushing CN, Li L, Oropeza CE, McLachlan A (2011) Multiple nuclear receptors may regulate hepatitis B virus biosynthesis during development. Int J Biochem Cell Biol 43(2):230–237
Ondracek CR, Reese VC, Rushing CN, Oropeza CE, McLachlan A (2009) Distinct regulation of hepatitis B virus biosynthesis by peroxisome proliferator-activated receptor γ coactivator 1α and small heterodimer partner in human hepatoma cell lines. J Virol 83(23):12545–12551
Lai CK, Ting LP (1999) Transcriptional repression of human hepatitis B virus genes by a bZIP family member, E4BP4. J Virol 73(4):3197–3209
Raney AK, Johnson JL, Palmer CNA, McLachlan A (1997) Members of the nuclear receptor superfamily regulate transcription from the hepatitis B virus nucleocapsid promoter. J Virol 71(2):1058–1071
Guo W, Chen M, Yen TSB, Ou J-H (1993) Hepatocyte-specific expression of the hepatitis B virus core promoter depends on both positive and negative regulation. Mol Cell Biol 13:443–448
Johnson JL, Raney AK, McLachlan A (1995) Characterization of a functional hepatocyte nuclear factor 3 binding site in the hepatitis B virus nucleocapsid promoter. Virology 208:147–158
Buckwold VE, Xu ZC, Yen TSB, Ou JH (1997) Effects of a frequent double-nucleotide basal core promoter mutation and its putative single-nucleotide precursor mutations on hepatitis B virus gene expression and replication. J Gen Virol 78:2055–2065
Yu XM, Mertz JE (1997) Differential regulation of the pre-C and pregenomic promoters of human hepatitis B virus by members of the nuclear receptor superfamily. J Virol 71:9366–9374
Ondracek CR, Rushing CN, Reese VC, Oropeza CE, McLachlan A (2009) Peroxisome proliferator-activated receptor γ coactivator 1α and small heterodimer partner differentially regulate nuclear receptor-dependent hepatitis B virus biosynthesis. J Virol 83(23):12535–12544
Chen I-H, Huang C-J, Ting L-P (1995) Overlapping initiator and TATA box functions in the basal core promoter of hepatitis B virus. J Virol 69:3647–3657
Garcia AD, Ostapchuk P, Hearing P (1993) Functional interaction of nuclear factors EF-C, HNF-4, and RXRα with hepatitis B virus enhancer I. J Virol 67:3940–3950
Huan B, Siddiqui A (1992) Retinoid X receptor RXRα binds to and trans-activates the hepatitis B virus enhancer. Proc Natl Acad Sci USA 89:9059–9063
Raney AK, Easton AJ, Milich DR, McLachlan A (1991) Promoter-specific transactivation of hepatitis B virus transcription by a glutamine- and proline-rich domain of hepatocyte nuclear factor 1. J Virol 65:5774–5781
Chang H-K, Wang B-Y, Yuh C-H, Wei C-L, Ting L-P (1989) A liver-specific nuclear factor interacts with the promoter region of the large surface protein gene of human hepatitis B virus. Mol Cell Biol 9:5189–5197
Raney AK, Zhang P, McLachlan A (1995) Regulation of transcription from the hepatitits B virus large surface antigen promoter by hepatocyte nuclear factor 3. J Virol 69:3265–3272
Raney AK, McLachlan A (1995) Characterization of the hepatitis B virus large surface antigen promoter Sp1 binding site. Virology 208:399–404
Raney AK, Easton AJ, McLachlan A (1994) Characterization of the minimal elements of the hepatitis B virus large surface antigen promoter. J Gen Virol 75:2671–2679
Raney AK, McLachlan A (1997) Characterization of the hepatitis B virus major surface antigen promoter hepatocyte nuclear factor 3 binding site. J Gen Virol 78:3029–3038
Santoro C, Mermod N, Andrews PC, Tjian R (1988) A family of human CCAAT-box-binding proteins active in transcription and DNA replication: cloning and expression of multiple cDNAs. Nature 334:218–224
Shaul Y, Ben Levy R, De Medina T (1986) High affinity binding site for nuclear factor I next to the hepatitis B virus S gene promoter. EMBO J 5:1967–1971
Raney AK, Le HB, McLachlan A (1992) Regulation of transcription from the hepatitis B virus major surface antigen promoter by the Sp1 transcription factor. J Virol 66:6912–6921
Lu CC, Yen TSB (1996) Activation of the hepatitis B virus S promoter by transcription factor NF-Y via a CCAAT element. Virology 225:387–394
Landschulz WH, Johnson PF, Adashi EY, Graves BJ, McKnight SL (1988) Isolation of a recombinant copy of the gene encoding C/EBP. Genes Dev 2:786–800
Ori A, Zauberman A, Doitsh G, Paran N, Oren M, Shaul Y (1998) p53 binds and represses the HBV enhancer: an adjacent enhancer element can reverse the transcription effect of p53. EMBO J 17(2):544–553
Nakao K, Nakata K, Yamashita M, Tamada Y, Hamasaki K, Ishikawa H et al (1999) p48 (ISGF-3gamma) is involved in interferon-α-induced suppression of hepatitis B virus enhancer-1 activity. J Biol Chem 274(40):28075–28078
Patel NU, Jameel S, Isom H, Siddiqui A (1989) Interactions between nuclear factors and the hepatitis B virus enhancer. J Virol 63:5293–5301
Ben-Levy R, Faktor O, Berger I, Shaul Y (1989) Cellular factors that interact with the hepatitis B virus enhancer. Mol Cell Biol 9:1804–1809
Chen M, Hieng S, Qian X, Costa R, Ou JH (1994) Regulation of hepatitis B virus ENI activity by hepatocyte-enriched transcription factor HNF3. Virology 205:127–132
Ori A, Shaul Y (1995) Hepatitis B virus enhancer binds and is activated by the hepatocyte nuclear factor 3. Virology 207:98–106
Huan B, Kosovsky MJ, Siddiqui A (1995) Retinoid X receptor α transactivates the hepatitis B virus enhancer 1 element by forming a heterodimeric complex with the peroxisome proliferator-activated receptor. J Virol 69:547–551
Siegrist CA, Durand B, Emery P, David E, Hearing P, Mach B et al (1993) RFX1 is identical to enhancer factor C and functions as a transactivator of the hepatitis B virus enhancer. Mol Cell Biol 13:6375–6384
Ostapchuk P, Scheirle G, Hearing P (1989) Binding of nuclear factor EF-C to a functional domain of the hepatitis B virus enhancer region. Mol Cell Biol 9:2787–2797
Maguire HF, Hoeffler JP, Siddiqui A (1991) HBV X protein alters the DNA binding specificity of CREB and ATF-2 by protein-protein interactions. Science 252:842–844
Zhang P, McLachlan A (1994) Differentiation-specific transcriptional regulation of the hepatitis B virus nucleocapsid gene in human hepatoma cell lines. Virology 202:430–440
Antonucci TK, Rutter WJ (1989) Hepatitis B virus (HBV) promoters are regulated by the HBV enhancer in a tissue-specific manner. J Virol 63:579–583
Lo W-Y, Ting L-P (1994) Repression of enhancer II activity by a negative regulatory element in the hepatitis B virus genome. J Virol 68:1758–1764
Doitsh G, Shaul Y (2004) Enhancer I predominance in hepatitis B virus gene expression. Mol Cell Biol 24(4):1799–1808
Faktor O, Budlovsky S, Ben Levy R, Shaul Y (1990) A single element within the hepatitis B virus enhancer binds multiple proteins and responds to multiple stimuli. J Virol 64:1861–1863
Shaul Y, Ben Levy R (1987) Multiple nuclear proteins in liver cells are bound to hepatitis B virus enhancer element and its upstream sequences. EMBO J 6:1913–1920
Shaul Y, Rutter WJ, Laub O (1985) A human hepatitis B viral enhancer element. EMBO J 4:427–430
Trujillo MA, Letovsky J, Maguire HF, Lopez-Cabrera M, Siddiqui A (1991) Functional analysis of a liver-specific enhancer of the hepatitis B virus. Proc Natl Acad Sci USA 88:3797–3801
Bulla GA, Siddiqui A (1988) The hepatitis B virus enhancer modulates transcription of the hepatitis B virus surface antigen gene from an internal location. J Virol 62:1437–1441
Jameel S, Siddiqui A (1986) The human hepatitis B virus enhancer requires trans-acting cellular factor(s) for activity. Mol Cell Biol 6:710–715
Guo W, Bell KD, Ou J-H (1991) Characterization of the hepatitis B virus EnhI enhancer and X promoter complex. J Virol 65:6686–6692
Yuh C-H, Ting L-P (1993) Differentiated liver cell specificity of the second enhancer of hepatitis B virus. J Virol 67:142–149
Yuh C-H, Ting L-P (1990) The genome of hepatitis B virus contains a second enhancer: cooperation of two elements within this enhancer is required for its function. J Virol 64:4281–4287
Wang Y, Chen P, Wu X, Sun A-L, Wang H, Zhu Y-A et al (1990) A new enhancer element, ENII, identified in the X gene of hepatitis B virus. J Virol 64:3977–3981
Vannice JL, Levinson AD (1988) Properties of the human hepatitis B virus enhancer: position effects and cell-type nonspecificity. J Virol 62:1305–1313
Raney AK, Milich DR, McLachlan A (1989) Characterization of hepatitis B virus major surface antigen gene transcriptional regulatory elements in differentiated hepatoma cell lines. J Virol 63:3919–3925
Raney AK, Milich DR, Easton AJ, McLachlan A (1990) Differentiation specific transcriptional regulation of the hepatitis B virus large surface antigen gene in human hepatoma cell lines. J Virol 64:2360–2368
Zhang P, Raney AK, McLachlan A (1992) Characterization of the hepatitis B virus X- and nucleocapsid gene transcriptional regulatory elements. Virology 191:31–41
Tognoni A, Cattaneo R, Serfling E, Schaffner W (1985) A novel expression selection approach allows precise mapping of the hepatitis B virus enhancer. Nucleic Acids Res 13:7457–7472
Ou J-H, Rutter WJ (1987) Regulation of secretion of the hepatitis B virus major surface antigen by the preS-1 protein. J Virol 61:782–786
Karpen S, Banerjee R, Zelent A, Price P, Acs G (1988) Identification of protein-binding sites in the hepatitis B virus enhancer and core promoter domains. Mol Cell Biol 8:5159–5165
Yee J-K (1989) A liver-specific enhancer in the core promoter region of human hepatitis B virus. Science 246:658–661
Raney AK, Milich DR, McLachlan A (1991) Complex regulation of transcription from the hepatitis B virus major surface antigen promoter in human hepatoma cell lines. J Virol 65:4805–4811
Chang HK, Chou CK, Chang C, Su TS, Hu C, Yoshida M et al (1987) The enhancer sequence of human hepatitis B virus can enhance the activity of its surface gene promoter. Nucleic Acids Res 15:2261–2268
De-Medina T, Faktor O, Shaul Y (1988) The S promoter of hepatitis B virus is regulated by positive and negative elements. Mol Cell Biol 8:2449–2455
Faktor O, De Medina T, Shaul Y (1988) Regulation of hepatitis B virus S gene promoter in transfected cell lines. Virology 162:362–368
Pourcel C, Louis A, Gervais M, Chenciner N, Dubois M-F, Tiollais P (1982) Transcription of the hepatitis B surface antigen gene in mouse cells transformed with cloned viral DNA. J Virol 42:100–105
Nakao K, Miyao Y, Ohe Y, Tamaoki T (1989) Involvement of an AFP1-binding site in cell-specific transcription of the pre-S1 region of the human hepatitis B virus surface antigen gene. Nucleic Acids Res 17:9833–9842
López-Cabrera M, Letovsky J, Hu K-Q, Siddiqui A (1990) Multiple liver-specific factors bind to the hepatitis B virus core/pregenomic promoter: trans-activation and repression by CCAAT/enhancer binding protein. Proc Natl Acad Sci USA 87:5069–5073
Waisman A, Aloni Y, Laub O (1990) In vitro regulation of human hepatitis B virus core gene transcription. Virology 177:737–744
Yaginuma K, Koike K (1989) Identification of a promoter region for 3.6-kilobase mRNA of hepatitis B virus and specific cellular binding protein. J Virol 63:2914–2920
Kosovsky MJ, Huan BF, Siddiqui A (1996) Purification and properties of rat liver nuclear proteins that interact with the hepatitis B virus enhancer 1. J Biol Chem 271:21859–21869
Zhou D-X, Yen TSB (1991) The ubiquitous transcription factor Oct-1 and the liver- specific factor HNF-1 are both required to activate transcription of a hepatitis B virus promoter. Mol Cell Biol 11:1353–1359
Dikstein R, Faktor O, Shaul Y (1990) Hierarchic and cooperative binding of the rat liver nuclear protein C/EBP at the hepatitis B virus enhancer. Mol Cell Biol 10:4427–4430
López-Cabrera M, Letovsky J, Hu K-Q, Siddiqui A (1991) Transcriptional factor C/EBP binds to and transactivates the enhancer element II of the hepatitis B virus. Virology 183:825–829
Yuh C-H, Chang Y-L, Ting L-P (1992) Transcriptional regulation of precore and pregenomic RNAs of hepatitis B virus. J Virol 66:4073–4084
Li M, Xie YH, Wu X, Kong YY, Wang Y (1995) HNF3 binds and activates the second enhancer, ENII, of hepatitis B virus. Virology 214:371–378
Pei D, Shih C (1990) Transcriptional activation and repression by cellular DNA-binding protein C/EBP. J Virol 64:1517–1522
Sells MA, Chen M-L, Acs G (1987) Production of hepatitis B virus particles in HepG2 cells transfected with cloned hepatitis B virus DNA. Proc Natl Acad Sci USA 84:1005–1009
Tang H, McLachlan A (2001) Transcriptional regulation of hepatitis B virus by nuclear hormone receptors is a critical determinant of viral tropism. Proc Natl Acad Sci USA 98:1841–1846
Junker M, Galle P, Schaller H (1987) Expression and replication of the hepatitis B virus genome under foreign promoter control. Nucleic Acids Res 15:10117–10132
Tang H, McLachlan A (2002) Mechanisms of inhibition of nuclear hormone receptor dependent hepatitis B virus replication by hepatocyte nuclear factor 3β. J Virol 76:8572–8581
Raney AK, Kline EF, Tang H, McLachlan A (2001) Transcription and replication of a natural hepatitis B virus nucleocapsid promoter variant is reguated in vivo by peroxisome proliferators. Virology 289:239–251
Guidotti LG, Eggers CM, Raney AK, Chi SY, Peters JM, Gonzalez FJ et al (1999) In vivo regulation of hepatitis B virus replication by peroxisome proliferators. J Virol 73(12):10377–10386
Oropeza CE, Li L, McLachlan A (2008) Differential inhibition of nuclear hormone receptor dependent hepatitis B virus replication by small heterodimer partner. J Virol 82(8):3814–3821
Li L, Oropeza CE, Sainz B, Uprichard SL, Gonzalez FJ, McLachlan A (2009) Developmental regulation of hepatitis B virus biosynthesis by hepatocyte nuclear factor 4α. PLoS One 4(5):e5489
Reese VC, Moore DD, McLachlan A (2012) Limited effects of bile acids and small heterodimer partner on hepatitis B virus biosynthesis in vivo. J Virol 86(5):2760–2768
Kyrmizi I, Hatzis P, Katrakili N, Tronche F, Gonzalez FJ, Talianidis I (2006) Plasticity and expanding complexity of the hepatic transcription factor network during liver development. Genes Dev 20(16):2293–2305
Li L, Oropeza CE, Kaestner KH, McLachlan A (2009) Limited effects of fasting on hepatitis B virus (HBV) biosynthesis in HBV transgenic mice. J Virol 83(4):1682–1688
Ondracek CR, McLachlan A (2011) Role of peroxisome proliferator-activated receptor gamma coactivator 1α in AKT/PKB-mediated inhibition of hepatitis B virus biosynthesis. J Virol 85(22):11891–11900
Shalaby RE, Iram S, Çakal B, Oropeza CE, McLachlan A (2017) PGC1α transcriptional adaptor function governs hepatitis B virus replication by controlling HBcAg/p21 protein-mediated capsid formation. J Virol 91(20):e00790-17
Shlomai A, Paran N, Shaul Y (2006) PGC-1α controls hepatitis B virus through nutritional signals. Proc Natl Acad Sci USA 103(43):16003–16008
Seifer M, Zhou S, Standring DN (1993) A micromolar pool of antigenically distinct precursors is required to initiate cooperative assembly of hepatitis B virus capsids in Xenopus oocytes. J Virol 67:249–257
Porterfield JZ, Dhason MS, Loeb DD, Nassal M, Stray SJ, Zlotnick A (2010) Full-length hepatitis B virus core protein packages viral and heterologous RNA with similarly high levels of cooperativity. J Virol 84(14):7174–7184
Shlomai A, Shaul Y (2008) The “metabolovirus” model of hepatitis B virus suggests nutritional therapy as an effective anti-viral weapon. Med Hypotheses 71(1):53–57
Rhee J, Inoue Y, Yoon JC, Puigserver P, Fan ML, Gonzalez FJ et al (2003) Regulation of hepatic fasting response by PPARγ coactivator-1α (PGC-1): requirement for hepatocyte nuclear factor 4α in gluconeogenesis. Proc Natl Acad Sci USA 100(7):4012–4017
Puigserver P, Spiegelman BM (2003) Peroxisome proliferator-activated receptor-gamma coactivator 1α (PGC-1α): transcriptional coactivator and metabolic regulator. Endocr Rev 24(1):78–90
Lin J, Handschin C, Spiegelman BM (2005) Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab 1(6):361–370
Yoon JC, Puigserver P, Chen GX, Donovan J, Wu ZD, Rhee J et al (2001) Control of hepatic gluconeogenesis through the transcriptional coactivator PGC-1. Nature 413(6852):131–138
Wang L, Lee YK, Bundman D, Han Y, Thevananther S, Kim CS et al (2002) Redundant pathways for negative feedback regulation of bile acid production. Dev Cell 2(6):721–731
Goodwin B, Jones SA, Price RR, Watson MA, McKee DD, Moore LB et al (2000) A regulatory cascade of the nuclear receptors FXR, SHP-1, and LRH-1 represses bile acid biosynthesis. Mol Cell 6(3):517–526
Lu TT, Makishima M, Repa JJ, Schoonjans K, Kerr TA, Auwerx J et al (2000) Molecular basis for feedback regulation of bile acid synthesis by nuclear receptors. Mol Cell 6(3):507–515
Komiya Y, Katayama K, Yugi H, Mizui M, Matsukura H, Tomoguri T et al (2008) Minimum infectious dose of hepatitis B virus in chimpanzees and difference in the dynamics of viremia between genotype A and genotype C. Transfusion 48(2):286–294
Barker LF, Chisari FV, McGrath PP, Dalgard DW, Kirschstein RL, Almeida JD et al (1973) Transmission of type B viral hepatitis to chimpanzees. J Infect Dis 127:648–662
Walter E, Keist R, Niederöst B, Pult I, Blum HE (1996) Hepatitis B virus infection of tupaia hepatocytes in vitro and in vivo. Hepatology 24(1):1–5
Yan RQ, Su JJ, Huang DR, Gan YC, Yang C, Huang GH (1996) Human hepatitis B virus and hepatocellular carcinoma. I. Experimental infection of tree shrews with hepatitis B virus. J Cancer Res Clin Oncol 122(5):283–288
Tang H, McLachlan A (2002) Avian and mammalian hepadnaviruses have distinct transcription factor requirements for viral replication. J Virol 76(15):7468–7472
Summers J, Smolec JM, Snyder R (1978) A virus similar to human hepatitis B virus associated with hepatitis and hepatoma in woodchucks. Proc Natl Acad Sci USA 75:4533–4537
Mason WS, Seal G, Summers J (1980) Virus of Pekin ducks with structural and biological relatedness to human hepatitis B virus. J Virol 36:829–836
Chisari FV (1995) Hepatitis B virus transgenic mice: insights into the virus and the disease. Hepatology 22:1316–1325
McFadden VC, Shalaby RE, Iram S, Oropeza CE, Landolfi JA, Lyubimov AV et al (2017) Hepatic deficiency of the pioneer transcription factor FoxA restricts hepatitis B virus biosynthesis by the developmental regulation of viral DNA methylation. PLoS Pathog 13(2):e1006239
Bruss V, Ganem D (1991) The role of envelope proteins in hepatitis B virus assembly. Proc Natl Acad Sci USA 88:1059–1063
Anderson AL, Banks KE, Pontoglio M, Yaniv M, McLachlan A (2005) Alpha/beta interferon differentially modulates the clearance of cytoplasmic encapsidated replication intermediates and nuclear covalently closed circular hepatitis B virus (HBV) DNA from the livers of hepatocyte nuclear factor 1{alpha}-null HBV transgenic mice. J Virol 79(17):11045–11052
Chang HK, Wang BY, Yuh CH, Wei CL, Ting LP (1989) A liver-specific nuclear factor interacts with the promoter region of the large surface protein gene of human hepatitis B virus. Mol Cell Biol 9(11):5189–5197
Summers J, Smith PM, Horwich AL (1990) Hepadnavirus envelope proteins regulate covalently closed circular DNA amplification. J Virol 64:2819–2824
Peters JM, Shah YM, Gonzalez FJ (2012) The role of peroxisome proliferator-activated receptors in carcinogenesis and chemoprevention. Nat Rev Cancer 12:181
Peraza MA, Burdick AD, Marin HE, Gonzalez FJ, Peters JM (2006) The toxicology of ligands for peroxisome proliferator-activated receptors (PPAR). Toxicol Sci 90(2):269–295
Li L, Oropeza CE, Sainz B Jr, Uprichard SL, Gonzalez FJ, McLachlan A (2009) Developmental regulation of hepatitis B virus biosynthesis by hepatocyte nuclear factor 4α. PLoS One 4(5):e5489
Banks KE, Anderson AL, Tang H, Hughes DE, Costa RH, McLachlan A (2002) Hepatocyte nuclear factor 3β inhibits hepatitis B virus replication in vivo. J Virol 76:12974–12980
Li Z, White P, Tuteja G, Rubins N, Sackett S, Kaestner KH (2009) Foxa1 and Foxa2 regulate bile duct development in mice. J Clin Invest 119(6):1537–1545
Wu X, Zhang Y (2017) TET-mediated active DNA demethylation: mechanism, function and beyond. Nat Rev Genet 18:517
Gao S, Duan Z-P, Coffin CS (2015) Clinical relevance of hepatitis B virus variants. World J Hepatol 7(8):1086–1096
Glebe D, Geipel A (2014) Selected phenotypic assays used to evaluate antiviral resistance and viral fitness of hepatitis B virus and its variants. Intervirology 57(3–4):225–231
Li L, Chirapu SR, Finn MG, Zlotnick A (2013) Phase diagrams map the properties of antiviral agents directed against hepatitis B virus Core assembly. Antimicrob Agents Chemother 57(3):1505–1508
Venkatakrishnan B, Katen SP, Francis S, Chirapu S, Finn MG, Zlotnick A (2016) Hepatitis B virus capsids have diverse structural responses to small-molecule ligands bound to the heteroaryldihydropyrimidine pocket. J Virol 90(8):3994–4004
Deres K, Schröder CH, Paessens A, Goldmann S, Hacker HJ, Weber O et al (2003) Inhibition of hepatitis B virus replication by drug-induced depletion of nucleocapsids. Science 299(5608):893–896
Cho MH, Jeong H, Kim YS, Kim JW, Jung G (2014) 2-amino-N-(2,6-dichloropyridin-3-yl)acetamide derivatives as a novel class of HBV capsid assembly inhibitor. J Viral Hepat 21(12):843–852
Campagna MR, Liu F, Mao R, Mills C, Cai D, Guo F et al (2013) Sulfamoylbenzamide derivatives inhibit the assembly of hepatitis B virus nucleocapsids. J Virol 87(12):6931–6942
Wu S, Zhao Q, Zhang P, Kulp J, Hu L, Hwang N et al (2017) Discovery and mechanistic study of benzamide derivatives that modulate hepatitis B virus capsid assembly. J Virol 91(16):e00519-17
Acknowledgments
This work was supported by Public Health Service grant AI125401 from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Oropeza, C.E., Tarnow, G., Sridhar, A., Taha, T.Y., Shalaby, R.E., McLachlan, A. (2020). The Regulation of HBV Transcription and Replication. In: Tang, H. (eds) Hepatitis B Virus Infection. Advances in Experimental Medicine and Biology, vol 1179. Springer, Singapore. https://doi.org/10.1007/978-981-13-9151-4_3
Download citation
DOI: https://doi.org/10.1007/978-981-13-9151-4_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-9150-7
Online ISBN: 978-981-13-9151-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)