Abstract
Bacterial blight of rice, caused by the bacterial pathogen Xanthomonas oryzae pv. oryzae (Xoo) in rice, represents one of the most well-studied crop diseases and is also well-known as a model for studying host/microbe interaction. TALEs (transcription activator-like effectors), as a group of pathogenesis factors and once translocated into the host cells from pathogen, recognize and activate host genes to condition disease susceptibility and also trigger host resistance responses dependent on the nature of target genes in plants. TALEs and their target genes have become the foci of the molecular battles between Xoo and rice. The continuing battles have led to incredibly diverse virulence mechanisms in pathogen and counteracting defense mechanisms in rice. Extensive efforts have been made to understand the TALE biology, identify host target genes, and elucidate their interaction and resulting physiological relevance to rice blight and other crop diseases. This review aims to summarize how much we have learned about TALEs and their role in bacterial blight of rice, as well as associated susceptibility and resistance genes in the host. The review also intends to provide a prospect of engineering genetic resistance by applying precise genome editing of TALE-associated target genes in rice.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
19.1 Introduction
Rice (Oryza sativa L.), an important stable crop that provides food for more than half of the world’s population, suffers a variety of biotic and abiotic stresses, some of which cause severe loss to crop production and affect global food security (Oerke 2006). During the cropping season, rice diseases occur at different stages. In particular, bacterial blight of rice, caused by Xanthomonas oryzae pv. oryzae (Xoo), is one of the most devastating rice diseases (Mew et al. 1993). Bacterial blight of rice is a vascular disease, starting with the pathogen infection of rice through entering the wound and hydathodes of plant leaves, multiplying and colonizing the vascular bundle cells, and eventually resulting in gray to white opaque necrotic lesion, so-called leaf blight (Niño-Liu et al. 2006). There are no effective measures to chemically control this disease in the field; the most effective and economically friendly way to manage the disease is through genetic resistance that is conferred by dominant or recessive disease resistance genes, the collectively so-called R genes that have been recurrently bred into the rice cultivars of interest (Mew 1987). However, newly emerging virulent, pathogenic isolates or races of Xoo can overcome the integrated resistance gene or genes rapidly and render the time-consuming breeding efforts a failure (McDonald and Linde 2002). Therefore, a better understanding of the virulence mechanisms of pathogen and host response to the infection as well as coupling with the advent of biotechnologies and advanced breeding programs will provide promise to effectively control this disease.
The blight disease of rice, like most of other plant diseases, is the outcome of molecular interaction between the host rice and pathogen Xoo. One such interaction is bridged through a type III secretion system (T3SS) of bacterial origin which translocates the bacterial effector proteins into host cells (White and Yang 2009). The effectors function per se to manipulate the host transcriptional and physiological processes, which condition a state of disease susceptibility in host, while host counteracts the effectors by involving changes in target genes and generation of resistance genes to prevent disease from occurring. In turn, pathogen has evolved new mechanisms to adapt the host resistance, leading to newly emerging pathogen isolates. This coadaptation and arms race between rice and Xoo also fit a broad aspect of host/microbe interaction characteristic of a “zigzag” model (Jones and Dangl 2006).
Genes encoding transcription activator-like effectors (TALEs) are present in every sequenced Xoo genome in a large, varying number, ranging from low 9 in AXO1947 to high 19 in PXO99 (Bogdanove et al. 2011; Booher et al. 2015; Huguet-Tapia et al. 2016; Ochiai et al. 2005; Quibod et al. 2016; Salzberg et al. 2008; Yu et al. 2015). In contrast to forming a family of closely related TALEs, other T3SS effectors, so-called non-TAL effectors, are diverse and amount to about 20 in individual Xoo strains (www.xanthomonas.org). Those T3SS effectors collectively play important roles in determining the pathogenesis (resistance and susceptibility) of individual Xoo strains in interacting with their host rice plants carrying diverse genetic background (White and Yang 2009). Additionally, environmental factors (e.g., humidity and temperature) also influence host resistance (or susceptibility) and eventually the severity of symptom and yield loss of the crop. For instance, unlike in most cases where higher temperature comprises effectiveness of disease resistance genes, the rice R gene Xa7 becomes more effective in resistance against Xoo strains harboring the cognate avirulence TALE gene AvrXa7 (Webb et al. 2010).
Serendipitously, bacterial blight of rice has become an excellent model for studying host/microbe interaction due to its genetic amenability, richness of genetic and molecular resource, and a rich history of extensive research endeavor (Niño-Liu et al. 2006). Breakthroughs have been made in identifying key virulence factors, including TALEs that Xoo uses to hijack the host’s biological processes to facilitate the infection and disease development, and in discovering host counteracting strategies that make plants resistant. Discovery derived from this rice/Xoo system has spawned development of biotechnological platforms (e.g., TALEN genome editing) and testing strategies for resistance against this particularly devastating disease and maybe applicable to other crop diseases such as citrus canker and cotton blight wherein TALEs also play an important role in pathogenesis. Better understanding of the biological process of disease has facilitated the forming of new and effective disease control strategies with the advent of new biotechnologies. For example, the newly emerging genome editing technologies provide promise for precisely changing the host targets utilized by the pathogen virulence factors or enhancing the resistance process used by the host. The aims of this review article are to summarize the progress that has been made in the past decades in the rice/Xoo interaction mediated by TALEs and to highlight the significance of combining a basic understanding of disease biology with the application of biotech tools to effectively control crop diseases.
19.2 Unique Structural and Functional Domains of TALEs
TALEs have been found to exist in the bacteria of Xanthomonas and Ralstonia solanacearum, and they form the largest family of T3SS effector proteins. Hundreds of TALE homolog sequences are available at the NCBI (www.ncbi.nlm.nih.gov), where they are highly conserved at the nucleotide and amino acid levels. Each TALE possesses an N-terminal domain with an embedded type III secretion signal, central repetitive region of nearly identical repeats of 33–34 amino acids (CRR), and a C-terminal region with functional nuclear localization signals (NLSs) and a transcription activation domain (AD) (Boch et al. 2010) (Fig. 19.1a). TALEs are DNA-binding proteins that recognize and bind to specific sequences of target genes, namely, the effector binding elements (EBEs), in the host genome. They recognize the DNA sequence of the target following a TALE code that involves (1) one repeat of TALE corresponding to one nucleotide of EBE and (2) one type of repeat preferentially binding to one type of four nucleotides. The identity of each repeat is largely dictated by the composition of two amino acid residues at the positions 12 and 13, so named RVD (repeat variable di-residues), which are highly variable in relation to the amino acid residues at other positions (Boch et al. 2009; Moscou and Bogdanove 2009). For instance, the predominant four repeats represented by HD, NI, NG, and NN match to nucleotides C, A, T, and G, respectively. The EBE sites usually are located within the promoter regions of the target genes. After binding to EBE, the TAL effector transcriptionally activates the host target gene, leading to disease susceptibility of the host or triggering host resistance against the otherwise virulent bacteria based on the nature of the host target gene. Therefore, a combination of number of repeats and compositions of RVDs, and the polymorphism of repeat domains, govern the specificity of TALEs for their EBEs and consequentially functionality of each TALE corresponding to the genetic background of the host plants (Bogdanove et al. 2010).
TALEs are structurally unique. (a). Schematic structure of TALE. TALE consists of three typical regions: N-terminus, central repetitive region (CRR), and C-terminus with nuclear localization signal (NLS) and transcription activation domain (AD). The CRR contains a varying number of 34-amino acid repeats with the two residues at the 12th and 13th positions defined as RVDs to represent individual repeats. (b). Representative and important TALEs that play important roles in the pathogenesis of Xanthomonas bacteria. The CRRs of individual TALEs are presented as the RVDs. A single letter is used for each amino acid, and asterisk (*) represents a missing amino acid residue at position 13
The N-terminal T3SS secretion signal is required for translocation of TALE into host cells and is therefore indispensable for function of TALE (Buttner and Bonas 2002; Szurek et al. 2002). The C-terminal NLSs are functional for directing TALEs to the nuclei of eukaryotic cells and are also required for virulence and avirulence activities of examined TALEs, including TALEs involved in bacterial blight of rice (PthA, AvrBs3, AvrXa7, AvrXa27) (Gu et al. 2005; Van den Ackerveken et al. 1996; Yang et al. 2000; Yang and Gabriel 1995). Additionally, a short sequence of acidic residues at the end of the C-terminus of TALEs is characteristic of eukaryotic transcription activation domains (Szurek et al. 2001; Zhu et al. 1998, 1999). Similar to NLSs, the activation domains of the TALEs AvrXa10 and AvrXa27 are required for their ability to trigger resistance responses in Xa10- and Xa27-containing rice varieties, respectively, while that of AvrXa7 is required for its ability to trigger resistance in Xa7 plants and condition a state of disease in otherwise susceptible rice (Gu et al. 2005; Yang et al. 2000; Zhu et al. 1999).
19.3 TALEs as Virulence Factors Condition a State of Host Susceptibility
The virulence function of TALEs was first reported in citrus canker pathogen Xanthomonas citri that carried PthA (pathogenicity A) based on the finding that its presence in the strain 3213 was associated with hypertrophy symptoms at infection sites in citrus leaves (Swarup et al. 1991). The rupture of the canker-like lesions was speculated to promote the release and spread of bacterial cells, providing ecological fitness to the pathogen; deletion of the pthA gene deprived the pathogen of the ability to cause canker-like lesion (Swarup et al. 1991, 1992). More PthA-like TALEs (e.g., PthA4, PthC, PthAW, and PthA*) of virulence from different strains of X. citri have been identified that cause canker-like lesions in citrus (Al-Saadi et al. 2007). Similarly, AvrBs3, initially identified for its ability to trigger resistance responses in Bs3-containing pepper plants and found to exist in certain isolates of X. campestris pv. vesicatoria, induces host cell hypertrophy in susceptible pepper plants lacking Bs3 and provides benefits to bacterial virulence in the field condition (Marois et al. 2002; Wichmann and Bergelson 2004). Some pathogenic strains of Xanthomonas bacteria in cassava and cotton blight diseases also benefit from TALE genes for increased virulence. All sequenced X. axonopodis pv. manihotis (Xam) strains possess TALE genes (Bart et al. 2012); it has been found that several TALEs (TALE1Xam, TAL14 Xam668, TAL20 Xam668) contribute to the strain virulence (ability to multiply and move systemically through the vascular system) in causing cassava bacterial blight (Castiblanco et al. 2013; Cohn et al. 2014, 2016). In cotton blight, caused by X. campestris pv. malvacearum, Avrb6 enhances disease symptoms of water-soaking and necrosis of the infected tissue and increases the bacterial release from the infected sites in susceptible cotton plants (Yang et al. 1994, 1996).
Remarkably, pathogenic X. oryzae pv. oryzae depends on critical TALEs to condition a state of disease susceptibility including in planta bacterial growth and spread of bacterial cells for colonization of vascular tissue in susceptible rice (Streubel et al. 2013; Sugio et al. 2007; Vera Cruz et al. 2000; Yang and White 2004; Yang et al. 2000; Yu et al. 2011). All sequenced or examined Xoo strains contain multiple copies of TALE sequences which are diverse in terms of their amount of the central repeats and composition of RVDs. The TALE sequences are predicted to express intact (or full-length) TALEs or variants of truncated TALEs (Grau et al. 2016). Six naturally occurring TALEs (AvrXa7, PthXo1, PthXo2, PthXo3, TalC, and Tal5, so-called major TALEs) have been found to be the essential virulence factors of Xoo in causing blight disease. When the major TALE genes were inactivated, the resultant mutant Xoo strains acted like nonpathogenic bacteria, lacking the ability to spread and colonize the vascular tissue of infected rice leaves, a unique feature for rice/Xoo pathosystem (Bai et al. 2000; Yang and White 2004; Yu et al. 2011). In addition, some other TALEs (e.g., PthXo6, PthXo7) moderately contribute to virulence so that some Xoo strains cause blight in susceptible rice (Bai et al. 2000; Sugio et al. 2007). Those TALEs are mainly distinguishable by the number of central repeats and composition of the RVDs (Fig. 19.1b). Their roles in virulence contribution also depend on the identity of their host target genes (see below).
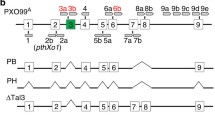
Surprisingly, there are two classes of TALE variants that function as the virulence factors of Xoo to interfere with the resistance in rice that contains the R gene Xa1 that encodes an NLR (nucleotide-binding domain, leucine-rich repeat) protein and recognizes most, if not all, full-length TALEs as its cognate avirulence effectors (Ji et al. 2016; Read et al. 2016). Those TALE variants were previously annotated as pseudogenes due to the mutations of premature stop codons or large deletions at their 3′ ends of coding sequences but were found to be expressed as the C-terminally truncated proteins, so named iTALEs (interfering TALEs). As the prototypes characterized from PXO99A, Tal3a with a premature stop codon encodes a TALE with a C-terminal truncation of 103 amino acids (the last NLS and AD deleted), while Tal3b carries a large deletion within its 3′ end results in a predicted protein with 229 amino acids deleted (knockout of three NLSs and AD) and 10 amino acids added (including a single, new NSL) due to a frame shift. Both genes contain two identical deletions (129 and 45 bp) within their 5′ regions compared to the intact TALE genes. Tal3a and Tal3b contain identical amino termini, distinguishable CRRs, and distinct C-termini. Deletion of the clustered Tal3a and Tal3b genes resulted in a mutant of PXO99A that triggered typical HR (hypersensitive reaction)-associated resistance in Xa1-containing rice. Furthermore, such 2 types of TALE variants are prevalent among 29 out 36 worldwide Xoo strains and all 9 sequenced X. oryzae pv. oryzicola (Xoc) strains (Ji et al. 2016). In another study, one such TALE variant, Tal2h named as truncTALE, from Xoc strain BLS256, was found to suppress resistance conferred by the Xo1 locus (an unknown but most likely NLR R gene) in the heirloom rice variety Carolina Gold (Read et al. 2016). The molecular mechanisms for Xa1-mediated recognition and activation of resistance to TALEs as well as how iTALEs or truncTALEs to interfere with such Xa1 resistance remain to be determined.
19.4 TALEs as Avirulence Factors Elicit Resistance Responses in Rice Against Xoo Infection
Some TALE genes, like all other microbial avirulence genes, were identified as the genetic determinants for eliciting resistance responses in host plants that contain the cognate resistance (R) gene. The resistance responses include activation of defense genes and hypersensitive reaction (HR). AvrBs3, the prototypic member of TALEs, was cloned from X. campestris pv. vesicatoria on its ability to trigger resistance responses in the R gene Bs3-containing pepper (Bonas et al. 1991; Minsavage et al. 1990). TALE genes avrXa10 and avrXa7 were later cloned from the Xoo strain PXO86 based on their homology to avrBs3 and their abilities to elicit resistance responses in rice varieties containing the R gene Xa10 and Xa7, respectively (Hopkins et al. 1992). avrXa7, characterized in the Korean, Japanese, and Philippine Xoo strains, is an effector gene with a dual function of virulence and avirulence depending on the genetic context of the host rice (Ochiai et al. 2000; Yang and White 2004). Similarly, TALEs of avirulence, AvrXa27 and AvrXa23, were cloned from PXO99A based on their relationships with the respective R genes Xa27 and Xa23 (Gu et al. 2005; Wang et al. 2014). Those four TALEs are also distinguishable by their number of central repeats and composition of RVDs (Fig. 19.1b), which both collectively determine the resistance specificity of the pathogen races and host cultivars. The resistance response is mediated in a gene-for-gene manner, i.e., one avirulence gene corresponds to one cognate R gene in Xoo and rice, respectively. On the other hand, most, if not all, TALEs function as resistance elicitors (avirulence factors) in rice that possess Xa1 or Xa1-like NLR R gene(s). The specificity of race/cultivar is not determined by the identity of TALEs but instead by the presence of iTALEs or truncTALEs (Ji et al. 2016; Read et al. 2016; Triplett et al. 2016).
19.5 TAL Effectors Induce Disease Susceptibility (S) Genes for Xoo Strain Virulence
Typical TALEs contain the characteristic eukaryotic transcription activator domains (NLS and AD), which tempt the initial hypothesis of host gene activation for the function of TALEs. The early work revealed the genes differentially activated by AvrBs3 in pepper using the cDNA-amplified fragment length polymorphism technique (Marois et al. 2002). The association of host gene induction and virulence functionality of pathogen TALEs came with the identification of Os8N3 (a member of MtN3 gene family) induced by its cognate PthXo1 in bacterial blight of rice (Yang et al. 2006). In that work, the rice microarrays were used to identify the differentially expressed genes from rice leaf samples treated with PXO99A and its mutant lacking pthXo1. RNAi-mediated gene silencing of Os8N3 in susceptible rice varieties or natural mutations in the Os8N3 promoter of rice IRBB13 (a recessive R gene, xa13, containing rice) rendered either plant resistant to PXO99A that relied on pthXo1 for virulence (Yang et al. 2006). The MtN3 gene family including Os8N3 was later found to encode a family of sugar transporters, so-called SWEETs, and Os8N3 or Xa13 (a dominant allele of xa13) was, therefore, renamed OsSWEET11 (Chen et al. 2010). Other SWEETs were also found to be specifically induced by other major TALEs of virulence. For example, OsSWEET13 is specifically induced by PthXo2 (Zhou et al. 2015); OsSWEE14 is induced by multiple TALEs, AvrXa7, PthXo3, TalC, and Tal5 (Antony et al. 2010; Streubel et al. 2013; Yu et al. 2011) (Fig. 19.2). Other two clade-III SWEETs in rice have been shown to also be the susceptibility genes to Xoo when induced with Xoo strains containing the artificial or designer TALEs (Li et al. 2013; Streubel et al. 2013).
Major TALEs and their SWEET targets in rice blight. (a). The TALE code in terms of nucleotides of target DNA and corresponding TALE repeats as illustrated in RVDs. (b). Symbols used for important components in the promoters of SWEETs. (c–e). Schematic gene structures of three SWEETs and their associated, cognate TALEs (presented as RVDs)
The relationship between TALEs and SWEET genes for disease susceptibility has been expanded from the originally described rice/Xoo system to other plant diseases such as cassava/X. axonopodis pv. manihotis (Cohn et al. 2014) and cotton/X. citri pv. malvacearum (Cox et al. 2017). The prevailing hypothesis for SWEETs to condition disease susceptibility is that elevated membrane-bound SWEETs render the sugar, especially sucrose, the direct product of photosynthesis in plant, leaking into the extracellular space wherein bacteria use carbohydrate to grow (Chen et al. 2010). Alternatively, leaking sugar out of plant cells depletes the cells of nutrient, and starvation of host cells results in compromised host immunity. Either hypothesis needs to be experimentally validated. TALEs of moderate virulence in Xoo also induce the cognate host target genes of non-SWEET to condition disease susceptibility. For example, PthXo6 and PthXo7 from PXO99A activate two host transcription factor genes, respectively, one for bZIP transcription factor and another for the general transcription factor TFIIA small subunit in chromosome 1 in rice (Sugio et al. 2007). Additionally, TALE of X. citri spp. citri plays an important role in citrus canker by targeting the host canker susceptibility gene lateral organ boundaries CsLOB1, also a transcription factor (Hu et al. 2014).
19.6 Host Resistance in Response to TALEs
To counteract the virulence strategy of pathogens that targets host genes of physiological importance through binding to their promoters and transcriptionally activating the gene expression, plants have evolved a “trapping mechanism” by luring some TALEs to the promoter of defense genes or R genes that have stronger defense activity than classic defense genes (e.g., pathogenesis-related PR genes) and are functionally equivalent to the dominant R genes. Those types of R genes are also called “executor” R genes (Bogdanove et al. 2010). The promoter element to which TALEs bind is also called EBE, the effector binding element. The first such case was identified for the pair of Xa27 and avrXa27 wherein rice variety IRBB27 harboring Xa27 recognizes the AvrXa27-secreting Xoo strains through trapping AvrXa27 to the promoter of Xa27 for its activation and subsequent resistance including hypersensitive responses at the infection sites (Gu et al. 2005). The nature of resistance/susceptibility of the variety and Xoo race is determined by the polymorphism where AvrXa27 could or could not match the EBE of Xa27 or xa27. Similarly, resistance genes Xa10 and Xa23, the closely related R gene family members, specifically lure the cognate TALEs AvrXa10 and AvrXa23 to their promoter EBEs for gene activation and resulting HR-associated resistance in specific varieties (Tian et al. 2014; Wang et al. 2014). Such pairing of TALEs of avirulence and executor R genes also exists beyond rice blight. The pepper R gene Bs3 (encoding a flavin monooxygenase) traps AvrBs3 for manifestation of resistance against Xanthomonas campestris pv. vesicatoria harboring the elicitor AvrBs3; similarly, Bs4C (encoding a unique R protein) again in pepper was found to be induced by TALE AvrBs4 for disease resistance; in both cases, the matchings between the EBEs and RVDs are required for gene induction and resistance, and polymorphisms between the resistant and susceptible alleles are primarily attributable to the outcome of diseases (Römer et al. 2007; Strauss et al. 2012).
Rice uses another resistance strategy to prevent its disease susceptibility genes from being induced by TALEs of virulence. Naturally occurring alleles with polymorphisms in EBE of OsSWEET11 (Os8N3 or Xa13) for PthXo1 are collectively named xa13; alterations of 1-bp substitution in rice variety BJ1 and 243-bp insertion in IRBB13 relative to the susceptible allele in IR24 are attributable to the mismatching between RVDs of PthXo1 and its cognate EBE and abolishment of xa13 induction by PthXo1 (Chu et al. 2006; Yang et al. 2006; Yuan et al. 2009). xa13 represents alleles carrying the promoter mutations instead of knockout mutations. It, presumably like Xa13, encodes a SWEET protein competent for normal physiological function of sucrose transport in the absence of bacterial infection. Another recessive R gene xa25 was found to be allelic to the bacterial blight susceptibility gene OsSWEET13 and was implicated as a host target of a major TALE of the Xoo strain PXO339 (Liu et al. 2011). It has been identified and experimentally validated later that xa25 is a nonresponsive allele of OsSWEET13 to another TALE, PthXo2, and that the EBE polymorphism in xa25 and OsSWEET13 determines the compatibility between respective rice varieties and Xoo strains that depend on PthXo2 alone for virulence (Zhou et al. 2015). The third recessive R gene due to loss of susceptibility to the Xoo TALEs AvrXa7 and Tal5 is xa41(t). The OsSWEET14 gene in Oryza barthii and O. glaberrima contains an 18-bp deletion just within the EBE for AvrXa7 and Tal5 relative to OsSWEET14 in Oryza sativa. The natural polymorphism or mutation in xa41(t) is responsible for non-response of OsSWEET14 to AvrXa7 and Tal5 for induction and, thus, responsible for the resistance to the specific Xoo strains (Hutin et al. 2015).
The third strategy deployed by rice for resistance against Xoo pathogens is to also interfere with S gene induction by TALEs not through alteration in EBEs of the S SWEET genes but instead through encoding a variant of general transcription factor TFIIA small subunit, the gene product of the recessive R gene xa5. xa5, being located on rice chromosome 5, encodes a protein with a valine to glutamic acid change (V39E) in TFIIAγ5 (or Xa5) (Iyer and McCouch 2004; Jiang et al. 2006). TFIIAγ5 is part of the transcription pre-initiation complex (Mediator) that is globally conserved across eukaryotic organisms (Allen and Taatjes 2015). Resistance to bacterial blight conferred by xa5 in IRBB5 line is broad spectrum against many strains of Xoo (Garris et al. 2003; Iyer and McCouch 2004; Mishra et al. 2013). It has been recently found that xa5 compromises the gene induction of OsSWEET13 and OsSWEET14 by their cognate TALEs PthXo2, PthXo3, and AvrXa7 (Huang et al. 2016). It has been hypothesized that reduced levels of OsSWEET13 and OsSWEET14 in the infected rice tissue are not sufficient enough to support bacterial multiplication. However, xa5 is not effective to resist to Xoo strains that contain PthXo1 for targeting OsSWEET11 for virulence. This is probably because the OsSWEET11 transcript as induced by PthXo1 in xa5 plants is exceptionally high, a level above the threshold for bacteria to be pathogenic (Huang et al. 2016). It has also been found that induction of Xa27 by TALE AvrXa27 is compromised and the resulting resistance is diminished in the xa5 background in bacterial blight (Gu et al. 2009). The hypothesis that TALEs directly interact with the plant Mediator for gene induction of S genes has been tested recently, and direct interaction of TFIIAγ5 with several TALEs has been demonstrated (Yuan et al. 2016).
19.7 Concluding Remarks
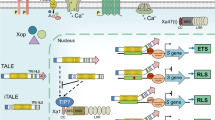
Through decades of research endeavor by scientists who had and have been working on bacterial blight of rice, significant progresses or breakthroughs have been made in the understanding of the biology of this disease. Achievements include revealing the key components of T3SS effectors (especially TALEs) and host target genes, and elucidating their interactions at the molecular level, which all play roles in bacterial infection, spread, and colonization in vascular tissue as well as host responses. In a simple model as illustrated in Fig. 19.3, the disease process starts with contact of Xoo with the plant cells and secretion and translocation of TALEs through T3SS into cytoplasmic space (scenario 1); TALEs go to nuclei and activate S genes for disease (scenario 2); plants change EBE sequences to avoid gene induction for loss of disease susceptibility (a resistance different from a dominant R gene-mediated resistance) (scenario 3); plants employ xa5 to reduce S gene induction by certain TALEs (scenario 4); plants also evolve executor R genes to trap TALEs to trigger resistance (scenario 5); and finally plants use the dominant NLR R genes to recognize TALEs and activate resistance independent of gene activation (scenario 6); in return Xoo mutates some TALEs to not only avoid NLR R protein recognition but also actively suppress resistance triggered by their progenitor TALEs (scenario 7). It is worthy to note that the genetically diverse Xoo populations produce several major TALEs capable of targeting multiple S SWEETs to overcome the usually single recessive resistance gene derived from the S SWEETs in rice.
Molecular interactions between TALEs and the cognate rice factors and the resulting bacterial blight disease. (a). In scenario 1, pathogen (e.g., Xoo) injects intact TALE and its variant iTALE into the cytoplasm (C) of host cell through a type III secretion system (T3SS); TALE and iTALE are localized to the cell nucleus (N) through their nuclear localization signals. In scenario 2, TALE targets the host disease susceptibility (S) gene through binding to its EBE (effector binding element) and transcriptionally activating the S gene, leading to a state of disease susceptibility, a compatible interaction between pathogen and host. In scenario 3, mutations (black dots) in EBE of the S gene in certain rice varieties prevent its activation by TALE, resulting in incompatible interaction between such a host and pathogen; the TALE-irresponsive allele of S gene becomes a genetically recessive resistance (r) gene. In scenario 4, rice with its TFIIA small subunit gene (TFIIAγ5) mutated (xa5) becomes incompatible to Xoo due to the inefficient activation of S gene by TALE. In scenario 5, rice use executor R gene to trap TALEs to trigger host resistance through gene activation. In scenario 6, rice also uses the NLR R gene to recognize TALEs for resistance independent of R gene activation. Finally, in scenario 7, pathogen uses truncated TALE variants to suppress TALE-triggered, NLR R gene-mediated resistance for compatible interaction between the host and pathogen. (b). Symbols used in Fig. 19.3a
The TALE targets (e.g., SWEETs) present a potential target for genetic manipulation of rice genome through biotechnologies for engineering broad resistance. Given the fact that the naturally occurring polymorphisms of S genes do not affect the normal physiological function of target genes, biotechnologies for such gene editing provide promise to mimic such polymorphisms for improved traits with possibly little negative effect on the overall performance of rice plants. The advanced genome-editing technologies enable precise, targeted genomic changes to specific loci. The editing-enabling technologies include zinc finger nucleases (ZFNs), TAL effector nucleases (TALENs), and most recently clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated (Cas) proteins (Weeks et al., 2016). The first TALEN-edited rice was generated by altering the overlapping EBEs of OsSWEET14 promoter for AvrXa7 and PthXo3 (Li et al. 2012). Through genetic transformation, TALEN reagents specifically targeting OsSWEET14 in rice susceptible to AvrXa7-dependent Xoo bacteria were used to generate plants with high mutagenesis frequency. Progeny plants homozygous for the EBE mutations and free of the TALEN transgene were highly resistant to Xoo strains that depended on AvrXa7 to induce OsSWEET14-associated rice blight (Li et al. 2012). A similar approach was used to mutate the EBEs in the promoter of OsSWEET14 targeted by TalC and Tal5, two TALEs prevalent among the African Xoo strains. Plants with respective mutations exhibited strong resistance to TalC- and Tal5-dependent strains that were pathogenic to the otherwise susceptible parental plant (Blanvillain-Baufumé et al. 2017). These studies provide the proof-of-concept successful cases in manipulation of a single SWEET gene for blight resistance. Multiplex genome-editing technologies based on CRISPR/Cas most likely enable simultaneous mutations in a single rice line of all five known EBEs in three S SWEETs (OsSWEET11, 13, and 14) targeted by all known TALEs. Such rice will be expected to confer the broadest resistance to most of, if not all, Xoo strains that are virulent to the otherwise susceptible parental rice.
References
Allen BL, Taatjes DJ (2015) The mediator complex: a central integrator of transcription. Nat Rev Mol Cell Biol 16:155–166
Al-Saadi A, Reddy JD, Duan YP, Brunings AM, Yuan Q, Gabriel DW (2007) All five host-range variants of Xanthomonas citri carry one pthA homolog with 17.5 repeats that determines pathogenicity on citrus, but none determine host-range variation. Mol Plant-Microbe Interact 20:934–943
Antony G, Zhou J, Huang S, Li T, Liu B, White F, Yang B (2010) Rice xa13 recessive resistance to bacterial blight is defeated by induction of the disease susceptibility gene Os-11N3. Plant Cell 22:3864–3876
Bai J, Choi SH, Ponciano G, Leung H, Leach JE (2000) Xanthomonas oryzae pv. oryzae avirulence genes contribute differently and specifically to pathogen aggressiveness. Mol Plant-Microbe Interact 13:1322–1329
Bart R, Cohn M, Kassen A, McCallum EJ, Shybut M, Petriello A, Krasileva K, Dahlbeck D, Medina C, Alicai T, Kumar L, Moreira LM, Rodrigues Neto J, Verdier V, Santana MA, Kositcharoenkul N, Vanderschuren H, Gruissem W, Bernal A, Staskawicz BJ (2012) High-throughput genomic sequencing of cassava bacterial blight strains identifies conserved effectors to target for durable resistance. Proc Natl Acad Sci U S A 109:E1972–E1979
Blanvillain-Baufumé S, Reschke M, Solé M, Auguy F, Doucoure H, Szurek B, Meynard D, Portefaix M, Cunnac S, Guiderdoni E, Boch J, Koebnik R (2017) Targeted promoter editing for rice resistance to Xanthomonas oryzae pv. oryzae reveals differential activities for SWEET14-inducing TAL effectors. Plant Biotechnol J 15:306–317
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U (2009) Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326:1509–1512
Boch J, Bonas U, VanAlfen N, Bruening G, Leach J (2010) Xanthomonas AvrBs3 family-type III effectors: discovery and function. Annu Rev Phytopathol 48:419–436
Bogdanove A, Schornack S, Lahaye T (2010) TAL effectors: finding plant genes for disease and defense. Curr Opin Plant Biol 13:394–401
Bogdanove AJ, Koebnik R, Lu H, Furutani A, Angiuoli SV, Patil PB, Van Sluys MA, Ryan RP, Meyer DF, Han SW, Aparna G, Rajaram M, Delcher AL, Phillippy AM, Puiu D, Schatz MC, Shumway M, Sommer DD, Trapnell C, Benahmed F, Dimitrov G, Madupu R, Radune D, Sullivan S, Jha G, Ishihara H, Lee SW, Pandey A, Sharma V, Sriariyanun M, Szurek B, Vera-Cruz CM, Dorman KS, Ronald PC, Verdier V, Dow JM, Sonti RV, Tsuge S, Brendel VP, Rabinowicz PD, Leach JE, White FF, Salzberg SL (2011) Two new complete genome sequences offer insight into host and tissue specificity of plant pathogenic Xanthomonas spp. J Bacteriol 193:5450–5464
Bonas U, Schulte R, Fenselau S, Minsavage GV, Staskawicz BJ, Stall RE (1991) Isolation of a gene cluster from Xanthomonas campestris pv. vesicatoria that determines pathogenicity and the hypersensitive response on pepper and tomato. Mol Plant-Microbe Interact 4:81–88
Booher NJ, Carpenter SC, Sebra RP, Wang L, Salzberg SL, Leach JE, Bogdanove AJ (2015) Single molecule real-time sequencing of Xanthomonas oryzae genomes reveals a dynamic structure and complex TAL (transcription activator-like) effector gene relationships. Microb Genom 1:e000032
Buttner D, Bonas U (2002) Getting across – bacterial type III effector proteins on their way to the plant cell. EMBO J 21:5313–5322
Castiblanco LF, Gil J, Rojas A, Osorio D, Gutiérrez S, Muñoz-Bodnar A, Perez-Quintero AL, Koebnik R, Szurek B, López C, Restrepo S, Verdier V, Bernal AJ (2013) TALE1 from Xanthomonas axonopodis pv. manihotis acts as a transcriptional activator in plant cells and is important for pathogenicity in cassava plants. Mol Plant Pathol 14:84–95
Chen LQ, Hou BH, Lalonde S, Takanaga H, Hartung ML, Qu XQ, Guo WJ, Kim JG, Underwood W, Chaudhuri B, Chermak D, Antony G, White FF, Somerville SC, Mudgett MB, Frommer WB (2010) Sugar transporters for intercellular exchange and nutrition of pathogens. Nature 468:527–532
Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen JL, Zhang Q, Wang S (2006) Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev 20:1250–1255
Cohn M, Bart RS, Shybut M, Dahlbeck D, Gomez M, Morbitzer R, Hou BH, Frommer WB, Lahaye T, Staskawicz BJ (2014) Xanthomonas axonopodis virulence is promoted by a transcription activator-like effector-mediated induction of a SWEET sugar transporter in cassava. Mol Plant-Microbe Interact 27:1186–1198
Cohn M, Morbitzer R, Lahaye T, Staskawicz BJ (2016) Comparison of gene activation by two TAL effectors from Xanthomonas axonopodis pv. manihotis reveals candidate host susceptibility genes in cassava. Mol Plant Pathol 17:875–889
Cox KL, Meng F, Wilkins KE, Li F, Wang P, Booher NJ, Carpenter SCD, Chen LQ, Zheng H, Gao X, Zheng Y, Fei Z, Yu JZ, Isakeit T, Wheeler T, Frommer WB, He P, Bogdanove AJ, Shan L (2017) TAL effector driven induction of a SWEET gene confers susceptibility to bacterial blight of cotton. Nat Commun 8:15588
Garris AJ, McCouch SR, Kresovich S (2003) Population structure and its effect on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice (Oryza sativa L.) Genetics 165:759–769
Grau J, Reschke M, Erkes A, Streubel J, Morgan RD, Wilson GG, Koebnik R, Boch J (2016) AnnoTALE: bioinformatics tools for identification, annotation, and nomenclature of TALEs from Xanthomonas genomic sequences. Sci Rep 6:21077
Gu K, Yang B, Tian D, Wu L, Wang D, Sreekala C, Yang F, Chu Z, Wang GL, White FF, Yin Z (2005) R gene expression induced by a type-III effector triggers disease resistance in rice. Nature 435:1122–1125
Gu K, Tian D, Qiu C, Yin Z (2009) Transcription activator-like type III effector AvrXa27 depends on OsTFIIA gamma 5 for the activation of Xa27 transcription in rice that triggers disease resistance to Xanthomonas oryzae pv. oryzae. Mol Plant Pathol 10:829–835
Hopkins CM, White FF, Choi SH, Guo A, Leach JE (1992) Identification of a family of avirulence genes from Xanthomonas oryzae pv. oryzae. Mol Plant-Microbe Interact 5:451–459
Hu Y, Zhang J, Jia H, Sosso D, Li T, Frommer WB, Yang B, White FF, Wang N, Jones JB (2014) Lateral organ boundaries 1 is a disease susceptibility gene for citrus bacterial canker disease. Proc Natl Acad Sci U S A 111:E521–E529
Huang S, Antony G, Li T, Liu B, Obasa K, Yang B, White FF (2016) The broadly effective recessive resistance gene xa5 of rice is a virulence effector-dependent quantitative trait for bacterial blight. Plant J 86:186–194
Huguet-Tapia JC, Peng Z, Yang B, Yin Z, Liu S, White FF (2016) Complete genome sequence of the African strain AXO1947 of Xanthomonas oryzae pv. oryzae. Genome Announc 4:e01730–e01715
Hutin M, Sabot F, Ghesquière A, Koebnik R, Szurek B (2015) A knowledge-based molecular screen uncovers a broad-spectrum OsSWEET14 resistance allele to bacterial blight from wild rice. Plant J 84:694–703
Iyer AS, McCouch SR (2004) The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol Plant-Microbe Interact 17:1348–1354
Ji Z, Ji C, Liu B, Zou L, Chen G, Yang B (2016) Interfering TAL effectors of Xanthomonas oryzae neutralize R-gene-mediated plant disease resistance. Nat Commun 7:13435
Jiang GH, Xia ZH, Zhou YL, Wan J, Li DY, Chen RS, Zhai WX, Zhu LH (2006) Testifying the rice bacterial blight resistance gene xa5 by genetic complementation and further analyzing xa5 (Xa5) in comparison with its homolog TFIIAgamma1. Mol Gen Genomics 275:354–366
Jones JD, Dangl JL (2006) The plant immune system. Nature 444:323–329
Li T, Liu B, Spalding MH, Weeks DP, Yang B (2012) High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat Biotechnol 30:390–392
Li T, Huang S, Zhou J, Yang B (2013) Designer TAL effectors induce disease susceptibility and resistance to Xanthomonas oryzae pv. oryzae in rice. Mol Plant 6:781–789
Liu Q, Yuan M, Zhou Y, Li X, Xiao J, Wang S (2011) A paralog of the MtN3/saliva family recessively confers race-specific resistance to Xanthomonas oryzae in rice. Plant Cell Environ 34:1958–1969
Marois E, Van den Ackerveken G, Bonas U (2002) The Xanthomonas type III effector protein AvrBs3 modulates plant gene expression and induces cell hypertrophy in the susceptible host. Mol Plant-Microbe Interact 15:637–646
McDonald B, Linde C (2002) Pathogen population genetics, evolutionary potential, and durable resistance. Annu Rev Phytopathol 40:349–379
Mew T (1987) Current status and future-prospects of research on bacterial-blight of rice. Annu Rev Phytopathol 25:359–382
Mew TW, Alvarez AM, Leach JE, Swing J (1993) Focus on bacterial blight of rice. Plant Dis 77:5–12
Minsavage GV, Dahlbeck D, Whalen MC, Kearney B, Bonas U, Staskawicz BJ, Stall RE (1990) Gene-for-gene relationships specifying disease resistance in Xanthomonas campestris pv. vesicatoria-pepper interactions. Mol Plant-Microbe Interact 3:41–47
Mishra D, Vishnupriya MR, Anil MG, Konda K, Raj Y, Sonti RV (2013) Pathotype and genetic diversity amongst Indian isolates of Xanthomonas oryzae pv. oryzae. PLoS One 8:e81996
Moscou MJ, Bogdanove AJ (2009) A simple cipher governs DNA recognition by TAL effectors. Science 326:1501
Niño-Liu DO, Ronald PC, Bogdanove AJ (2006) Xanthomonas oryzae pathovars: model pathogens of a model crop. Mol Plant Pathol 7:303–324
Ochiai H, Horino O, Miyajima K, Kaku H (2000) Genetic diversity of Xanthomonas oryzae pv. oryzae strains from Sri Lanka. Phytopathology 90:415–421
Ochiai H, Inoue Y, Takeya M, Sasaki A, Kaku H (2005) Genome sequence of Xanthomonas oryzae pv. oryzae suggests contribution of large numbers of effector genes and insertion sequences to its race diversity. Jpn Agric Res Q 39:275–287
Oerke EC (2006) Crop losses to pests. J Agric Sci 144:31–43
Quibod IL, Perez-Quintero A, Booher NJ, Dossa GS, Grande G, Szurek B, Vera Cruz C, Bogdanove AJ, Oliva R (2016) Effector diversification contributes to Xanthomonas oryzae pv. oryzae phenotypic adaptation in a semi-isolated environment. Sci Rep 6:34137
Read AC, Rinaldi FC, Hutin M, He YQ, Triplett LR, Bogdanove AJ (2016) Suppression of Xo1-mediated disease resistance in rice by a truncated, non-DNA-binding TAL effector of Xanthomonas oryzae. Front Plant Sci 7:1516
Römer P, Hahn S, Jordan T, Strauss T, Bonas U, Lahaye T (2007) Plant pathogen recognition mediated by promoter activation of the pepper Bs3 resistance gene. Science 318:645–648
Salzberg SL, Sommer DD, Schatz MC, Phillippy AM, Rabinowicz PD, Tsuge S, Furutani A, Ochiai H, Delcher AL, Kelley D, Madupu R, Puiu D, Radune D, Shumway M, Trapnell C, Aparna G, Jha G, Pandey A, Patil PB, Ishihara H, Meyer DF, Szurek B, Verdier V, Koebnik R, Dow JM, Ryan RP, Hirata H, Tsuyumu S, Won Lee S, Seo YS, Sriariyanum M, Ronald PC, Sonti RV, Van Sluys MA, Leach JE, White FF, Bogdanove AJ (2008) Genome sequence and rapid evolution of the rice pathogen Xanthomonas oryzae pv. oryzae PXO99A. BMC Genomics 9:204
Strauss T, van Poecke RM, Strauss A, Römer P, Minsavage GV, Singh S, Wolf C, Kim S, Lee HA, Yeom SI, Parniske M, Stall RE, Jones JB, Choi D, Prins M, Lahaye T (2012) RNA-seq pinpoints a Xanthomonas TAL-effector activated resistance gene in a large-crop genome. Proc Natl Acad Sci U S A 109:19480–19485
Streubel J, Pesce C, Hutin M, Koebnik R, Boch J, Szurek B (2013) Five phylogenetically close rice SWEET genes confer TAL effector-mediated susceptibility to Xanthomonas oryzae pv. oryzae. New Phytol 200:808–819
Sugio A, Yang B, Zhu T, White F (2007) Two type III effector genes of Xanthomonas oryzae pv. oryzae control the induction of the host genes OsTFIIA gamma 1 and OsTFX1 during bacterial blight of rice. Proc Natl Acad Sci USA 104:10720–10725
Swarup S, Feyter RD, Brlansky RH, Gabriel ADW (1991) A pathogenicity locus from Xanthomonas citri enables strains from several pathovars of X. campestris to elicit canker like lesions on citrus. Phytopathology 81:802–809
Swarup S, Yang Y, Kingsley MT, Gabriel DW (1992) An Xanthomonas citri pathogenicity gene, pthA, pleiotropically encodes gratuitous avirulence on nonhosts. Mol Plant-Microbe Interact 5:204–213
Szurek B, Marois E, Bones U, Van den Ackerveken G (2001) Eukaryotic features of the Xanthomonas type III effector AvrBs3: protein domains involved in transcriptional activation and the interaction with nuclear import receptors from pepper. Plant J 26:523–534
Szurek B, Rossier O, Hause G, Bonas U (2002) Type III-dependent translocation of the Xanthomonas AvrBs3 protein into the plant cell. Mol Microbiol 46:13–23
Tian D, Wang J, Zeng X, Gu K, Qiu C, Yang X, Zhou Z, Goh M, Luo Y, Murata-Hori M, White F, Yin Z (2014) The rice TAL effector-dependent resistance protein XA10 triggers cell death and calcium depletion in the endoplasmic reticulum. Plant Cell 26:497–515
Triplett L, Cohen S, Heffelfinger C, Schmidt C, Huerta A, Tekete C, Verdier V, Bogdanove A, Leach J (2016) A resistance locus in the American heirloom rice variety Carolina gold select is triggered by TAL effectors with diverse predicted targets and is effective against African strains of Xanthomonas oryzae pv. oryzicola. Plant J 87:472–483
Van den Ackerveken G, Marois E, Bonas U (1996) Recognition of the bacterial avirulence protein AvrBs3 occurs inside the host plant cell. Cell 87:1307–1316
Vera Cruz CM, Bai J, Ona I, Leung H, Nelson RJ, Mew TW, Leach JE (2000) Predicting durability of a disease resistance gene based on an assessment of the fitness loss and epidemiological consequences of avirulence gene mutation. Proc Natl Acad Sci U S A 97:13500–13505
Wang C, Qin T, Yu H, Zhang X, Che J, Gao Y, Zheng C, Yang B, Zhao K (2014) The broad bacterial blight resistance of rice line CBB23 is triggered by a novel transcription activator-like (TAL) effector of Xanthomonas oryzae pv. oryzae. Mol Plant Pathol 15:333–341
Webb KM, Oña I, Bai J, Garrett KA, Mew T, Vera Cruz CM, Leach JE (2010) A benefit of high temperature: increased effectiveness of a rice bacterial blight disease resistance gene. New Phytol 185:568–576
Weeks D, Spalding M, Yang B (2016) Use of designer nucleases for targeted gene and genome editing in plants. Plant Biotechnol J 14:483–495
White F, Yang B (2009) Host and pathogen factors controlling the rice-Xanthomonas oryzae interaction. Plant Physiol 150:1677–1686
Wichmann G, Bergelson J (2004) Effector genes of Xanthomonas axonopodis pv. vesicatoria promote transmission and enhance other fitness traits in the field. Genetics 166:693–706
Yang Y, Gabriel D (1995) Xanthomonas avirulence/pathogenicity gene family encodes functional-plant nuclear targeting signals. Mol Plant-Microbe Interact 8:627–631
Yang B, White FF (2004) Diverse members of the AvrBs3/PthA family of type III effectors are major virulence determinants in bacterial blight disease of rice. Mol Plant-Microbe Interact 17:1192–1200
Yang Y, De Feyter R, Gabriel DW (1994) Host-specific symptoms and increased release of Xanthomonas citri and X. campestris pv. malvacearum from leaves are determined by the 102-bp tandem repeats of pthA and avrb6, respectively. Mol Plant-Microbe Interac 7:345–355
Yang Y, Yuan Q, Gabriel DW (1996) Watersoaking function(s) of XcmH1005 redundantly encoded by members of the Xanthomonas avr/pth gene family. Mol Plant-Microbe Interact 9:105–112
Yang B, Zhu W, Johnson L, White F (2000) The virulence factor AvrXa7 of Xanthomonas oryzae pv, oryzae is a type III secretion pathway-dependent nuclear-localized double-stranded DNA-binding protein. Proc Natl Acad Sci U S A 97:9807–9812
Yang B, Sugio A, White F (2006) Os8N3 is a host disease-susceptibility gene for bacterial blight of rice. Proc Natl Acad Sci U S A 103:10503–10508
Yu Y, Streubel J, Balzergue S, Champion A, Boch J, Koebnik R, Feng J, Verdier V, Szurek B (2011) Colonization of rice leaf blades by an African strain of Xanthomonas oryzae pv. oryzae depends on a new TAL effector that induces the rice nodulin-3 Os11N3 gene. Mol Plant-Microbe Interact 24:1102–1113
Yu YH, Lu Y, He YQ, Huang S, Tang JL (2015) Rapid and efficient genome-wide characterization of Xanthomonas TAL effector genes. Sci Rep 5:13162
Yuan M, Chu Z, Li X, Xu C, Wang S (2009) Pathogen-induced expressional loss of function is the key factor in race-specific bacterial resistance conferred by a recessive R gene xa13 in rice. Plant Cell Physiol 50:947–955
Yuan M, Ke Y, Huang R, Ma L, Yang Z, Chu Z, Xiao J, Li X, Wang S (2016) A host basal transcription factor is a key component for infection of rice by TALE-carrying bacteria. elife 5:e19605
Zhou J, Peng Z, Long J, Sosso D, Liu B, Eom J, Huang S, Liu S, Cruz C, Frommer W, White F, Yang B (2015) Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant J 82:632–643
Zhu W, Yang B, Chittoor JM, Johnson LB, White FF (1998) AvrXa10 contains an acidic transcriptional activation domain in the functionally conserved C terminus. Mol Plant-Microbe Interact 11:824–832
Zhu W, Yang B, Wills N, Johnson LB, White FF (1999) The C terminus of AvrXa10 can be replaced by the transcriptional activation domain of VP16 from the herpes simplex virus. Plant Cell 11:1665–1674
Acknowledgments
The authors gratefully acknowledge grant support from the Rural Development Administration (RDA) of the Republic of Korea (PJ012098 to S.P. and B.Y.) and the Bill and Melinda Gates Foundation (Grand Challenge Program, a subaward from Carnegie Institute for Science to BY).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Char, S.N., Park, S., Yang, B. (2018). Interaction of Rice and Xanthomonas TAL Effectors. In: Sasaki, T., Ashikari, M. (eds) Rice Genomics, Genetics and Breeding. Springer, Singapore. https://doi.org/10.1007/978-981-10-7461-5_19
Download citation
DOI: https://doi.org/10.1007/978-981-10-7461-5_19
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-7460-8
Online ISBN: 978-981-10-7461-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)