Abstract
Pseudomonas aeruginosa is a ubiquitous and opportunistic human pathogen that represents a critical problem to the clinician due to the increased number of resistant strains isolated from hospital settings. In addition, there is a great variety of pathologies associated with this versatile Gram-negative bacterium. P. aeruginosa cells are able to produce an incredible arsenal of virulence factors, especially secreted molecules that act singly or together to ensure the establishment, maintenance, and persistence of a successful infection in susceptible hosts. In this context, pseudomonal proteases’ roles are highlighted due to their ability to cleave key host proteinaceous substrates as well as to modulate several biological processes, for example, escaping and modulating the host immune responses in the bacterial own favor. Proteases secreted by P. aeruginosa include elastase A (LasA), elastase B (LasB), alkaline protease (AP), protease IV (PIV), Pseudomonas small protease (PASP), large protease A (LepA), MucD, and P. aeruginosa aminopeptidase (PAAP). In the present review, we discuss the role of each of these relevant proteases produced by P. aeruginosa taking into consideration their main biological functions in the bacterium–host interaction that favors the establishment of the infectious process.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Pseudomonads are bacteria well known for their metabolic versatility and widespread spatiotemporal distribution [1]. One of the most important species of pseudomonads is, with no doubt, Pseudomonas aeruginosa, which is a fascinating ubiquitous Gram-negative bacterium with rod shape measuring 0.5–0.8 µm × 1.5–3.0 µm (Fig. 1a) [1, 2]. P. aeruginosa presents the following metabolic features: non-fermentative, catalase positive, oxidase positive, ammonia producer, and usually aerobic, but it also can grow in an anaerobic environment if nitrate, citrate, and arginine are available [3]. The production of 2-aminoacetophenone by the bacterial cells generates the fruity grape-like odor that is characteristic of this pseudomonad species. On blood agar plates, colonies of P. aeruginosa often display beta-hemolysis and a greenish metallic sheen due to the production of pigments [2]. The characteristic that most distinguishes P. aeruginosa from the other pseudomonads, and from the other species of Gram-negative non-fermenting bacteria, is its ability to produce pyocyanin, a blue-green phenazine pigment that gives the green color to the bacterial colony (Fig. 1b) and also to the pus observed in P. aeruginosa-infected tissues. This pigment and several others, such as pyochelin (purple-cyan), pyoverdin (yellow, green and fluorescent), pyomelanin (light-brown), and pyorubin (red-brown), are secondary metabolites of P. aeruginosa, which play an important role in bacterial nutrition, such as iron acquisition and pathogenesis [2, 3]. Almost all P. aeruginosa strains are motile due to the presence of a single polar flagellum that facilitates the locomotion and colonization of a wide range of environmental niches [2]. This microorganism can grow within the temperature range from 4 to 42 °C in terrestrial (soil) and aquatic habitats (polluted, salt, and freshwater) as well as on the surface of animate hosts (insects, plants, animals, and humans) and inanimate surroundings, mainly in the hospital environment (distilled water, disinfectants, sinks, medical devices, and equipment), being an important causative agent of nosocomial infections, particularly in intensive care units (ICUs) [1,2,3,4]. One of the interesting characteristics of P. aeruginosa is its pan-genome, which presents a larger genetic repertoire than the human genome. This intriguing feature explains the broad metabolic capabilities of P. aeruginosa and its distribution and adaptability in diverse environments [5].
P. aeruginosa is one of the most important bacterial species for public health considerations due to its high resistance to different classes of antibiotics and its capability to cause serious health care-associated as well as nosocomial infections [6, 7]. Results reported from an International Nosocomial Infection Control Consortium (INICC) surveillance study, performed between 2007 and 2012, in Latin America, Asia, Africa, and Europe, in which prospective data were collected from 605,310 patients hospitalized in 503 ICUs, displayed frequencies of 42.8% of Pseudomonas isolates resistant to amikacin and 42.4% to imipenem [8]. In the USA, an estimated 51,000 health care-associated P. aeruginosa infections occur each year, in which more than 6,000 (13%) of these are multidrug-resistant and 400 deaths per year are attributed to these infections [9]. The analyses based on data extracted from the Public Health England (PHE) voluntary surveillance database in the period 2008–2012 showed that 92% of Pseudomonas spp. isolates identified from bacteremia in 3,457 reports were P. aeruginosa [10]. In Brazil, the National Health Surveillance Agency (ANVISA), through the National Monitoring Microbial Resistance Network Health Services (RM Network), published a report that shows the main etiologic agents and the resistance phenotypes responsible for causing primary bloodstream infections associated with the use of central venous catheter in adult patients interned at ICUs from Brazilian hospitals between January and December 2013. According to that study, 18,233 notifications were reported, of which 1,850 (10.1%) were caused by P. aeruginosa, being the fifth pathogen most often reported as the etiologic agent. The resistance rate to the carbapenems reached 37.4% (692 P. aeruginosa isolates) [11]. Additionally, the Infectious Diseases Society of America has highlighted P. aeruginosa as part of a faction of antibiotic-resistant bacteria, called ‘the ESKAPE pathogens’—Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter spp., capable of ‘escaping’ the bactericidal action of antibiotics and mutually representing new paradigms in pathogenesis, transmission, and resistance [12].
P. aeruginosa is extensively resistant to multiple drugs and is increasingly resistant to most available antibiotics, being a great emergency problem in the hospital settings worldwide [13]. Interestingly, P. aeruginosa has evolved over time in its ability to find new ways to be resistant to different classes of chemical compounds as well as to build strategies to exchange genetic materials, allowing that other bacteria also become drug-resistant [5]. Generally, resistance usually occurs due to a combination of factors acting synergistically: (i) P. aeruginosa is intrinsically resistant to antimicrobial agents due to its outer membrane/cell envelope composition that reduces the permeability of several drugs; and (ii) P. aeruginosa expresses a powerful repertoire of resistance mechanisms that can be developed through mutations in the genomic content that regulates resistance genes, and also acquired from other organisms via plasmids, transposons, or bacteriophages [14].
As a major opportunistic pathogen for humans, P. aeruginosa causes a plenty variety of acute and chronic infections and presents significant levels of morbidity and mortality [15, 16]. P. aeruginosa typically infects through airways, wounds, urinary tract, ear canal, via ocular and implanted medical devices (e.g., catheters or ventilators). Thereby, it is the main cause of eschars, conjunctivitis, keratitis, corneal ulcer, osteomyelitis, otitis, urinary infections, surgical site infections, bloodstream infections in ICUs and hospital-acquired pneumonia in immunocompromised individuals, mainly in patients with severe burn wounds, AIDS, lung cancer, chronic obstructive pulmonary disease, bronchiectasis, and cystic fibrosis [16,17,18].
It is known that Gram-negative bacteria are common causes of a huge diversity of infections including, intra-abdominal infections (IAIs), urinary tract infections (UTIs), ventilator-associated pneumonia (VAP), and bacteremia [19]. In particular, P. aeruginosa is one of the most important pathogens in the hospital setting, being responsible for 27% of all pathogens and 70% of all Gram-negative bacteria causing health care-associated infections in the USA, and it is the most common Gram-negative organism causing VAP and the second most common organism causing catheter-associated UTIs [7, 19]. The Centers for Disease Control and Prevention found that P. aeruginosa totalized 7.1% of health care-associated infection in the USA in 2011, being the second most common cause of pneumonia in hospital settings and the third most common Gram-negative bacterium to cause bloodstream infections [20]. P. aeruginosa is also a major cause of concern in the cystic fibrosis setting, being the most common pathogen isolated from cystic fibrosis sputum, and approximately 70% of adult cystic fibrosis patients are chronically colonized by this microorganism [21, 22].
The pathogenic potential of P. aeruginosa is not only due to its metabolic/genetic versatility and both intrinsic and acquired antibiotic resistance. Its ability to form biofilm and to produce an arsenal of virulence attributes, including cell-associated determinants (e.g., lipopolysaccharide, pili, and flagellum) and soluble secreted factors (e.g., extracellular polysaccharides, exotoxins, pigments, and proteases), is very important for the survival and adaptation of this pathogen in distinct environments [17, 22, 23].
2 Pseudomonas aeruginosa: Establishing and Maintaining an Infection
In order to establish an infection, P. aeruginosa cells count on a suite of virulence factors (Fig. 2) [17, 24]. These factors act together not only causing injuries on the host epithelial cell lining but also inducing dysfunctions in physiology and function, such as host cell shape, membrane permeability, and protein synthesis, as well as manipulating/overcoming host defenses, down-modulating the immune responses and preventing P. aeruginosa endocytosis and obstructing clearance mechanisms, thereby allowing this microbe to persist in cells/tissues and to establish an infection in the host [25, 26]. The virulence of P. aeruginosa is mediated by multiple mechanisms, but the major contributor is the production of extracellular proteases. In general, these enzymes regulate multiple cellular and physiological processes and are essential to the success of the infection. They degrade a wide array of host proteins, impairing host defenses and destroying physical barriers that normally prevent attachment and penetration of the bacteria [26,27,28].
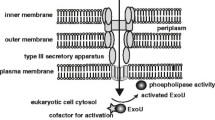
Virulence factors expressed/produced by P. aeruginosa cells: (i) lipopolysaccharide (LPS) that induces cytokine production, (ii) pili that help bacterial adherence to the respiratory epithelial cells, (iii) flagellum that participates in mobility, adherence, and internalization events, (iv) extracellularly released molecules like proteases (responsible for the cleavage of key host proteins), exotoxin A (inhibition of host protein synthesis), exoenzyme S (induces cytotoxic effect), exoenzyme U (antiphagocytic effect), phospholipase C (cleavage of membrane phospholipids), pigments (many biological effects, like pyocyanin that induces free radicals in host cells), rhamnolipids (detergent action), soluble lectins (inhibition of beating of lung cells), and alginate (phagocytosis inhibition, antifungal action, and host immune responses)
3 Proteolytic Enzymes Produced by Pseudomonas aeruginosa
P. aeruginosa is able to extracellularly release different kinds of proteases (Fig. 3), which together are responsible for invasion and destruction of host tissues. Because of the relevant roles played by proteases on the physiopathology of P. aeruginosa, it has been shown that the majority of environmental and clinical strains of P. aeruginosa exhibited proteolytic activity, particularly elastase activity [29,30,31]. According to Stover and co-workers [32], approximately 3% of the whole P. aeruginosa genome is composed by open reading frames that encode proteases [32]. Thus, the high genomic variability allows the bacterium to adapt its virulence arsenal machinery to support the variations of environment conditions, and for that, protease production in P. aeruginosa can vary greatly (Fig. 4) [32].
Production of extracellular proteases in clinical isolates of P. aeruginosa recovered from different anatomical sites. The proteolytic profiles were characterized by sodium dodecyl sulfate-containing polyacrylamide gel electrophoresis (SDS-PAGE) containing 0.1% gelatin as the protein substrate. Profile I—118 + 50 kDa; Profile II—118 + 90 + 50 kDa; Profile III—90 + 50 kDa; Profile IV—118 + 80 + 50 kDa; Profile V—90 + 80 kDa; Profile VI—118 kDa; Profile VII—90 kDa, and Profile VIII—36 kDa
The expression of extracellular proteolytic enzymes in P. aeruginosa is directly influenced by environmental factors and changes in the physicochemical properties of culture medium (e.g., nutrients, temperature, pH, and aeration), which significantly modulate the production of these crucial virulence factors [26, 33]. In addition, the amount of protease produced depends on the cell cycle moment (e.g., lag, exponential, or stationary growth phase) and on the growing lifestyle (e.g., planktonic or biofilm). For instance, the total protease production (Fig. 5a) as well as the specific elastase secretion increases along the first 48 h of in vitro cultivation of P. aeruginosa planktonic cells (Fig. 5b). Further, according to Hastie and co-workers [34], after 85 h of bacterial growth, the elastase production dropped off.
Protease detection in P. aeruginosa. a Total extracellular protease production was analyzed by the degradation of casein (1%) incorporated into Luria Bertani agar medium up to 48 h at 37 °C. b The elastase activity was measured in the cell-free culture supernatant obtained from P. aeruginosa cells grown in tryptic soy broth up to 48 h at 37 °C, using the fluorogenic peptide substrate Abz-Ala-Gly-Leu-Ala-p-Nitro-Benzyl-Amide. Results were expressed as fluorescence arbitrary units (FAU). In parallel, the number of bacterial cells along each time point was evaluated by plating cells onto agar medium and expressed as colony-forming units (CFU)
3.1 Elastase B
One of the best proteases characterized in Pseudomonas is elastase B (LasB), also known as pseudolysin. This 33-kDa enzyme belongs to the M4 thermolysin-like family of neutral, Zn-dependent metallo-endopeptidases (Fig. 6). This enzyme is encoded by lasB gene as a pre-pro-protein, containing at the N-terminal region a signal peptide of 23 amino acids that transport the enzyme through the inner membrane to periplasmic place by bacterial secretory system [35].
Elastase of P. aeruginosa is a typical zinc-metalloprotease. The purified elastase B is able to cleave the fluorogenic peptide substrate Abz-Ala-Gly-Leu-Ala-p-Nitro-Benzyl-Amide along the time. Conversely, 1,10-phenanthroline (1,10-Phen), a metalloprotease inhibitor, at 10 μM was able to block the substrate cleavage. FAU, fluorescence arbitrary units
The first and the most studied substrate of elastase B is bovine and human elastin [36,37,38]. Some reports correlate the elastinolytic activity of elastase B to Pseudomonas infections in cystic fibrosis patients [39,40,41,42,43]. Histological studies have detected altered elastin fibers in lung alveoli of cystic fibrosis patients on autopsy, indicating a probable elastase activity on cystic fibrosis lung [39]. In addition, the elastase activity is associated with vascular inflammation during P. aeruginosa infection, since the disorganization of elastin fiber in vascular tissue caused by protease degradation was observed [44]. Previously, our group analyzed the production of virulence attributes in 96 clinical strains of P. aeruginosa recovered from patients attended at hospitals located in three States of Brazil (Espírito Santo, Minas Gerais, and Rio de Janeiro), and it was shown that all bacterial strains exhibited a homogeneous elastase activity, with an average of 1069.28 ± 213.95 fluorogenic arbitrary units (FAU) with no correlation with the original anatomical site of isolation [16]. On the other hand, P. aeruginosa strains recovered from trachea, urinary tract, and wounds of patients attended at University Medical Center/Texas Tech Health Sciences Center were able to produce different amounts of elastase [45]. Woods and co-workers [46] showed that Canadian P. aeruginosa strains isolated from acute lung infections showed the highest production of elastase (0.053 ± 0.021 mg/ml) compared with elastase activity of strains isolated from burns, wounds, cystic fibrosis lung, and blood.
LasB is also able to cleave other host extracellular matrix proteins, such as collagen type III and IV. Interestingly, after subcutaneous injection of purified elastase B into mice, an intense degradation of basement membranes was observed, and elastase B was responsible for severe hemorrhage and tissue damage [47]. Several studies have demonstrated that LasB-associated epithelial disruption is mediated by the attack to intracellular tight junctions and cytoskeleton reorganization via inhibition of protein kinase C and activation of EGFR, ERK1/2 and NFκB, urokinase, and protease-activated receptor 2 (PAR-2) [48,49,50,51,52,53]. Elastase B can also interfere with the host bacterial clearance by degrading several components of innate and adaptive immune defenses, including tumor necrosis factor-α (TNF-α), interferon-γ (IFN-γ) and interleukin-2 (IL-2), monocyte chemotactic protein-1 (MCP-1), and epithelial neutrophil activating protein-78 (ENA-78) [52,53,54,55,56,57]. In addition, it was shown that elastase B was efficient in the inactivation of key components of the complement system such as fluid-phase and cell-bound C1 and C3 and fluid-phase C5, C8, and C9 [44]. This multifunctional enzyme is also able to cleave surfactant protein A and D (SP-A and SP-D), also known as collectin. SP-A and SP-D are synthesized by alveolar type II epithelial cells and are responsible for the recognition and binding to oligosaccharides present on the cell surface of many bacteria to be phagocytized by host macrophages [58]. Previously, Meyer and co-workers [59] have reported that a decrease on the SP-A and SP-D levels in bronchoalveolar lavage (BAL) was observed in the lung of cystic fibrosis individuals. Also, SP-D knockout mice were more sensible to P. aeruginosa corneal infections when compared to wild-type animals, and only the wild-type mice recovered completely of the infection [60]. Based on this, elastase B was suggested to be responsible for the SP-D degradation in the eye [25, 26]. Furthermore, pseudomonal elastase can interact with host adaptive immune system by degrading immunoglobulins [61,62,63]. Bainbrigde and Flick [61] showed that elastase B was able to cleave IgG molecules recovered from cystic fibrosis patients and the degradation products bound to IgG-receptors of human neutrophils, thereby inhibiting the opsonization of bacterial invaders. Lomholt and Kilian [63] reported the IgA degradation in tears from patients infected with P. aeruginosa. They also observed that isogenic mutants of P. aeruginosa knockout to either elastase or alkaline protease were not able to completely inhibit the IgA degradation, indicating that several proteases were working in concert to cleave IgA.
Furthermore, elastase B plays a key role in the differentiation of pseudomonal biofilms. Tielen and co-workers [64] showed that strains that overexpress lasB gene were not able to form robust biofilms, and they observed the formation of few microcolonies after 72 h of contact with glass surface. Those authors also assigned that lasb-overexpressed strain shifted the composition of its extracellular polymeric substances, reducing the alginate content as well as enhancing the rhamnolipids concentration [64]. However, Yu and co-workers [65] demonstrated that elastase B is crucial for biofilm formation. They observed that ΔlasB mutant decreased the biofilm formation through down-regulation of rhamnolipids synthesis.
3.2 Elastase A
Another extracellular protease produced by P. aeruginosa is elastase A (LasA), a metalloprotease that belongs to the subgroup A of M23 family of staphylolytic or β-lytic zinc metallo-endopeptidases. LasA is codified as an elastase A pre-pro-protein with molecular mass of 40 kDa [66, 67]. After its synthesis in intracellular bacterial environment, LasA is secreted via type II secretion machinery and when it is secreted to the extracellular space, LasA is immediately converted to its mature and active form of 27 kDa due to the cleavage by other pseudomonal-secreted endopeptidases, such as LasB, LysC, and protease IV [68, 69].
Elastase A is also called as staphylolysin, because it is able to cleave the pentaglycine bonds in the peptidoglycan of Staphylococcus aureus [70]. As well, LasA degrades several glycine-rich synthetic peptides [71]. However, LasA exhibited a limited elastinolytic activity [72]. Kessler and co-workers [71] showed that LasA prefers cleaving Gly–Ala peptide bonds within the Gly-Gly-Ala sequences surrounded by apolar sequences. Such sequences are uncommon in elastin, resulting in low elastinolytic activity [26, 73]. Besides its own intrinsic elastinolytic activity, LasA enhances significantly the elastinolytic activity of other proteases, including LasB in P. aeruginosa, but also human leukocyte elastase and human neutrophil elastase [74, 75]. Moreover, LasA is responsible for inducing shedding of the host cell surface proteoglycan syndecan-1 (co-receptor proteins), which has been shown to be important for P. aeruginosa survival [25, 26].
3.3 Alkaline Protease
Another pseudomonal protein shown to be important for phagocytic evasion is alkaline protease (AprA), which is also known as aeruginolysin. Alkaline protease is a 50-kDa zinc-metalloprotease, member of subfamily B of the M10 peptidase family and metzincin superfamily. AprA, encoded by aprA gene, has a C-terminal secretion signal located within the last 50 amino acid residues necessary to be translocated and secreted by AprD, APrE, and AprF membrane proteins, which form the bacterial type I secretory machinery [35].
It was reported that alkaline protease is able to degrade a large number of host proteins, including fibronectin and laminin, important components of basal lamina and endothelium. Therefore, alkaline protease develops an important function in invasion and hemorrhagic tissue necrosis in infections caused by P. aeruginosa [76]. Furthermore, this protease was found in many isolates of P. aeruginosa recovered from different human anatomical sites with especial elevated expression in clinical isolates from eyes, gastrointestinal tract, and mucoid wounds exacerbated in cystic fibrosis patients [25, 61]. AprA is important to bacterial escape from the host immunological defenses, degrading complement proteins (C1q, C2, and C3) and cytokines (IFN-γ, TNF-α and IL-6) [76]. Also, alkaline protease and elastase B are able to inhibit chemotaxis of neutrophils and block efficiently the phagocytosis, which gives the pathogen an advantage in escaping from phagocyte cells that are one of the first lines of host defense mechanisms [25, 31, 77, 78]. Moreover, alkaline protease is able to inhibit flagellin recognition by TLR5 due to the degradation of free flagellin monomers, helping P. aeruginosa cells to avoid the immune detection [79]. This enzyme has also been shown to aid P. aeruginosa survival in iron limitations conditions during human infections by cleaving transferrin that increase the siderophore-mediated iron uptake [80]. Gupta and co-workers [81] also reported that treatment of mouse corneal tissue with alkaline protease (50 ng) increases the binding of P. aeruginosa to the epithelial surface.
3.4 Protease IV
P. aeruginosa secretes a serine-type protease designated as protease IV (PIV) or lysyl endopeptidase (PrpL), a 26-kDa protease belonging to the chymotrypsin family S1 that has been demonstrated to be an important virulence factor in the rabbit cornea, but is found in clinical isolates recovered from all the anatomical sites analyzed [35, 82]. Its catalytic domain is formed by the triad His72, Asp122, and Ser198. Moreover, it was demonstrated that the residue Ser197 adjacent to Ser198 is critical to the catalytic activity [83]. Protease IV is encoded by piv gene (PA4175), with a full length of 48 kDa, which is initially expressed in the cytoplasm in a pre-pro-enzyme form and then processed to the 26-kDa mature protease after its secretion into the extracellular milieu [83].
PIV participates in the tissue invasion/damage processes and hemorrhagic events due to the cleavage of fibrinogen. It is well known that fibrinogen is required after vascular damage, but the degradation of fibrinogen by PIV leads to hemorrhage during P. aeruginosa infection [84]. PIV is also important to evade host immune defenses because it is able to degrade plasminogen, immunoglobulin, C1q and C3, and host antimicrobial peptide LL-37 [25, 68]. Furthermore, Malloy and co-workers [82] observed that PIV degrades the surfactant proteins, SP-A, SP-D, and SP-B, by a time- and dose-depended way in cell-free bronchoalveolar lavage fluid. Those authors reported that degradation of SPs by protease IV reduced the association among bacteria and alveolar macrophage. Interestingly, the incubation of pulmonary surfactant with pseudomonal protease IV reduced the ability of the surfactant to diminish the superficial tension within the lung [82]. Protease IV has been shown to be an iron-regulated protein, suggesting that its expression is regulated irrespective of quorum sensing system, which is distinct from other pseudomonal proteases [69]. Protease IV has also been correlated to ring abscess lesions present in pseudomonal keratitis [68]. Corroborating this finding, Engel and co-workers [85] showed that protease IV-deficient mutants exhibited lower ocular virulence in rabbits when intrastromally infected.
3.5 Pseudomonas Small Protease
P. aeruginosa small protease (PASP) is described as a 18.5-kDa secreted zinc-dependent leucine aminopeptidase. PASP gene has been found in a large number of P. aeruginosa clinical strains, but its higher expression is found during the ocular infection [86]. Previous reports showed that PASP is found only in the bacterial supernatant culture. According to Tang and co-workers [86], the sequence of PASP gene appears to have a signal peptide consistent with that needed for type II secretion system.
Direct inoculation of purified PASP into the rabbit cornea causes severe ocular pathology, including epithelial erosion and ulcer in stroma, edema, and neutrophil infiltration into the corneal stroma [87]. PASP has also been demonstrated to cleavage host proteins required for maintaining structure of cornea, such as collagens, fibrinogen (but not fibrin), complement C3, and antimicrobial peptide LL-37. Studies of PASP, coupled with those of PIV, strongly support the hypothesis that Pseudomonas proteases play a major role in keratitis [87].
3.6 Large Exoprotease A
Large exoprotease A (LepA) is an exoprotease with molecular mass of ~100 kDa produced by P. aeruginosa. LepA, as well as thrombin and trypsin, cleaves human protease-activated receptors (PARs) 1, 2, and 4 in order to activate the critical transcription factor NF-κB, which is associated with host inflammatory and immune responses [49, 88].
3.7 MucD
MucD was reported to be a serine endoprotease that is localized within the periplasmic space. Data suggest that MucD induced a significant reduction on the levels of IL-1β, neutrophil-chemoattractant chemokines KC, and macrophage-inflammatory protein-2 (MIP-2) in the early stages of bacterial infection as well as it inhibited the recruitment of polymorphonuclear (PMN) cells into the cornea. Furthermore, a decrease in PMN cells recruited to infection site favored the establishment of infection by P. aeruginosa. MucD may be secreted to the extracellular space, interfering with the biological functions of cytokines and chemokines, but further investigation is needed to understand the mechanisms underlying the role of MucD in keratitis [89, 90].
3.8 Aminopeptidase
The P. aeruginosa aminopeptidase (PAAP) or leucine aminopeptidase has been speculated as complementary enzyme to the activity of other endopeptidases. PAAP has an important function in bacterial physiology; it acts releasing free amino acids/small peptides from protein fragments produced by the others P. aeruginosa endopeptidases, thereby providing low molecular mass nutrients that can be taken up by the bacterium, which in turn may promote bacterial growth and proliferation [26].
4 Conclusions
P. aeruginosa is a metabolically versatile bacterium that can cause a wide range of severe opportunistic infections in hospitalized patients. To cause this huge variety of infections, P. aeruginosa has an arsenal of proteases that are involved in critical events of bacterial pathogenicity and virulence, which are important for survival in the host, tissue invasion, and evasion of host immune defenses. Therefore, this review has highlighted the importance of each pseudomonal protease in bacterial physiology and/or in infectious events. In this context, inhibitors able to block the proteases produced by P. aeruginosa cells would represent a new drug class quite promising to combat this widespread bacterial pathogen.
References
Tümmler B, Wiehlmann L, Klockgether J et al (2014) Advances in understanding Pseudomonas. F1000 Prime 6:9
Jayaseelan S, Ramaswamy D, Dharmaraj S (2014) Pyocyanin: production, applications, challenges and new insights. World J Microbiol Biotechnol 30:1159–1168
Vasil ML (1986) Pseudomonas aeruginosa: biology, mechanisms of virulence, epidemiology. J Pediatr 108:800–805
Wiehlmann L, Wagner G, Cramer N et al (2007) Population structure of Pseudomonas aeruginosa. PNAS 104:8101–8106
Ghodhbane H, Elaidi S, Sabatier JM et al (2015) Bacteriocins active against multi-resistant gram negative bacteria implicated in nosocomial infections. Infect Disord Drug Targets 15:2–12
Bartram J, Cotruvo J, Exner M, Fricker C, Glasmacher A (2003) WHO—World Health Organization. Heterotrophic plate counts and drinking-water safety. IWA Publishing, London. ISBN: 1 84339 025 6
Nielsen SL (2015) The incidence and prognosis of patients with bacteremia. Dan Med J 62:B5128
Rosenthal VD, Maki DG, Mehta Y, Leblebicioglu H et al (2014) International nosocomial Infection Control Consortium. International Nosocomial Infection Control Consortium (INICC) report, data summary of 43 countries for 2007–2012. Device-associated module. Am J Infect Control 42:942–956
Centers for Disease Control and Prevention. Healthcare-associated Infections (HAIs). Pseudomonas aeruginosa in Healthcare Settings. Available in: http://www.cdc.gov/hai/organisms/pseudomonas.html. Accessed on 17 June 2016
Public Health England (2012) Pseudomonas aeruginosa: guidance, data and analysis—voluntary surveillance of Pseudomonas spp. and Stenotrophomonas spp. causing bacteraemia in England, Wales and Northern Ireland. Available in: https://www.gov.uk/government/publications/pseudomonas-spp-and-stenotrophomonas-spp-voluntary-surveillance-2012. Accessed on 17 June 2016
ANVISA—Agência Nacional de Vigilância Sanitária. Boletim Informativo Segurança do Paciente e Qualidade em Serviços de Saúde—Ano V nº 09|Dezembro de 2014. Available in: http://portal.anvisa.gov.br. Accessed on 17 June 2016
Pendleton JN, Gorman SP, Gilmore BF (2013) Clinical relevance of the ESKAPE pathogens. Expert Rev Anti Infect Ther 11:297–308
Buhl M, Peter S, Willmann M (2013) Prevalence and risk factors associated with colonization and infection of extensively drug-resistant Pseudomonas aeruginosa: a systemic review. Expert Rev Anti-infect Ther 13:1159–1170
El Zowalaty ME, Al Thani AA, Webster TJ et al (2015) Pseudomonas aeruginosa: arsenal of resistance mechanisms, decades of changing resistance profiles, and future antimicrobial therapies. Future Microbiol 10:1683–1706
Bentzmann S, Plésiat P (2011) The Pseudomonas aeruginosa opportunistic pathogen and human infections. Environ Microbiol 13:1655–1665
Silva LV, Galdino ACM, Nunes APF et al (2014) Virulence attributes in Brazilian clinical isolates of Pseudomonas aeruginosa. Int J Med Microbiol 304:990–1000
Balasubramanian D, Schneper L, Kumari H, Mathee K (2013) A dynamic and intricate regulatory network determines Pseudomonas aeruginosa virulence. Nucleic Acids Res 41:1–20
Savoia D (2014) New perspectives in the management of Pseudomonas aeruginosa infections. Future Microbiol 9:917–928
Kaye KS, Pogue JM (2015) Infections caused by resistant Gram-negative bacteria: epidemiology and management. Pharmacotherapy 35:949–962
McCarthy K (2015) Pseudomonas aeruginosa: evolution of antimicrobial resistance and implications for therapy. Semin Respir Crit Care Med 36:44–55
Sousa AM, Pereira MO (2014) Pseudomonas aeruginosa diversification during infection development in cystic fibrosis lungs—a review. Pathogens 3:680–703
Oliver A, Mulet X, López-Causapé C, Juan C (2015) The increasing threat of Pseudomonas aeruginosa high-risk clones. Drug Resist Update 22:41–59
Kung VL, Ozer EA, Hauser AR (2010) The accessory genome of Pseudomonas aeruginosa. Microbiol Mol Biol 74:621–664
Crousilles A, Maunders E, Bartlett S, Fan C et al (2015) Which microbial factors really are important in Pseudomonas aeruginosa infections? Future Microbiol 10:1825–1836
Ballok AE, O’Toole GA (2013) Pouring salt on a wound: Pseudomonas aeruginosa virulence factors alter Na+ and Cl− flux in the lung. J Bacteriol 195:4013–4019
Kessler E, Safrin M (2014) Elastinolytic and proteolytic enzymes. In Pseudomonas methods and protocols. Methods Mol Biol 1149:135–169
McCarty SM, Cochrane CA, Clegg PD, Percival SL (2012) The role of endogenous and exogenous enzymes in chronic wounds: a focus on the implications of aberrant levels of both host and bacterial proteases in wound healing. Wound Repair Regen 20:125–136
Gellatly SL, Hancock REW (2013) Pseudomonas aeruginosa: new insights into pathogenesis and host defenses. Pathog Dis 67:159–173
Schmidtchen A, Wolff H, Hansson C (2001) Differential proteinase expression by Pseudomonas aeruginosa derived from chronic leg ulcers. Acta Derm Venereol 81:406–409
Tingpej P, Smith L, Rose B et al (2007) Phenotypic characterization of clonal and nonclonal Pseudomonas aeruginosa strains isolated from lungs of adults with cystic fibrosis. J Clin Microbiol 45:1697–1704
Thibodeau PH, Butterworth MB (2013) Proteases, cystic fibrosis and the epithelial sodium channel (ENaC). Cell Tissue Res 351:309–323
Stover CK, Pham XQ, Erwin AL et al (2000) Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature 406:959–964
Marquart ME, Dajcs JJ, Caballero AR et al (2005) Calcium and magnesium enhance the production of Pseudomonas aeruginosa protease IV, a corneal virulence factor. Med Microbiol Immunol 194:39–45
Hastie AT, Hingley ST, Kueppers F (1983) Protease production by Pseudomonas aeruginosa isolates from patients with cystic fibrosis. Infect Immun 40:506–513
Hoge R, Pelzer A, Rosenau F, Wilhelm S (2010) Weapons of a pathogen: proteases and their role in virulence of Pseudomonas aeruginosa. In: Méndez-Vilas A (ed) Current research, technology and education topics in applied microbiology and microbial biotechnology. Formatex Research Center, Badajoz, pp. 383–395
Hamdaoui A, Wund-Bisseret F, Bieth JG (1987) Fast solubilization of human lung elastin by Pseudomonas aeruginosa elastase. Am Rev Respir Dis 135:860–863
Saulnier JM, Curtil FM, Duclos MC, Wallach JM (1989) Elastolytic activity of Pseudomonas aeruginosa elastase. Biochim Biophys Acta 995:285–290
Yang J, Zhao HL, Ran YL et al (2015) Mechanistic insights into elastin degradation by pseudolysin, the major virulence factor of the opportunistic pathogen Pseudomonas aeruginosa. Sci Rep 9936
Bruce MC, Poncz L, Klinger JD et al (1985) Biochemical and pathologic evidence for proteolytic destruction of lung connective tissue in cystic fibrosis. Am Rev Respir Dis 132:529–535
Erickson DL, Endersby R, Kirkham A et al (2002) Pseudomonas aeruginosa quorum-sensing systems may control virulence factor expression in the lungs of patients with cystic fibrosis. Infect Immun 70:1783–1790
Kosorok MR, Zeng L, West SE et al (2001) Acceleration of lung disease in children with cystic fibrosis after Pseudomonas aeruginosa acquisition. Pediatr Pulmonol 32:277–287
Voynow JA, Fischer BM, Zheng S (2008) Proteases and cystic fibrosis. Int J Biochem Cell Biol 40:1238–1245
Van’t Wout EF, van Schadewijk A, van Boxtel R et al (2015) Virulence factors of Pseudomonas aeruginosa induce both the unfolded protein and integrated stress responses in airway epithelial cells. PLoS Pathog 11:e1004946
Schultz DR, Miller KD (1974) Elastase of Pseudomonas aeruginosa: inactivation of complement components and complement-derived chemotactic and phagocytic factors. Infect Immun 10:128–135
Hamood A, Griswold G, Colmer J (1996) Characterization of elastase-deficient clinical isolates of Pseudomonas aeruginosa. Infect Immun 64:3154–3160
Woods DE, Schaffer MS, Rabin HR et al (1988) Phenotypic comparison of Pseudomonas aeruginosa strains isolated from a variety of clinical sites. J Bacteriol 170:4309–4314
Komori Y, Nonogaki T, Nikai T (2001) Hemorrhagic activity and muscle damaging effect of Pseudomonas aeruginosa metalloproteinase (elastase). Toxicon 39:1327–1332
Bentzmann S, Polette M, Zahm JM et al (2000) Pseudomonas aeruginosa virulence factors delay airway epithelial wound repair by altering the actin cytoskeleton and inducing overactivation of epithelial matrix metalloproteinase-2. Lab Invest 80:209–219
Kida Y, Higashimoto Y, Inoue H et al (2008) A novel secreted protease from Pseudomonas aeruginosa activates NF-kappaB through protease-activated receptors. Cell Microbiol 10:491–504
Clark CA, Thomas LK, Azghani AO (2011) Inhibition of protein kinase C attenuates Pseudomonas aeruginosa elastase-induced epithelial barrier disruption. Am J Respir Cell Mol Biol 45:1263–1271
Cosgrove S, Chotirmall SH, Greene CM et al (2011) Pulmonary proteases in the cystic fibrosis lung induce interleukin 8 expression from bronchial epithelial cells via a heme/meprin/epidermal growth factor receptor/Toll-like receptor pathway. J Biol Chem 286:692–704
Kuang Z, Hao Y, Walling BE et al (2011) Pseudomonas aeruginosa elastase provides an escape from phagocytosis by degrading the pulmonary surfactant protein-A. PLoS ONE 6:e27091
Nomura K, Obata K, Keira T et al (2014) Pseudomonas aeruginosa elastase causes transient disruption of tight junctions and downregulation of PAR-2 in human nasal epithelial cells. Respir Res 18:15–21
Parmely M, Gale A, Clabaugh M et al (1990) Proteolytic inactivation of cytokines by Pseudomonas aeruginosa. Infect Immun 58:3009–3014
Horvat RT, Clabaugh M, Duval-Jobe C, Parmely MJ (1989) Inactivation of human gamma interferon by Pseudomonas aeruginosa proteases: elastase augments the effects of alkaline protease despite the presence of alpha 2-macroglobulin. Infect Immun 57:1668–1674
Theander TG, Kharazmi A, Pedersen BK et al (1988) Inhibition of human lymphocyte proliferation and cleavage of interleukin-2 by Pseudomonas aeruginosa proteases. Infect Immun 56:1673–1677
Leidal KG, Munson KL, Johnson MC et al (2003) Metalloproteases from Pseudomonas aeruginosa degrade human RANTES, MCP-1, and ENA-78. J Interferon Cytokine Res 23:307–318
Mariencheck WI, Alcorn JF, Palmer SM (2003) Pseudomonas aeruginosa elastase degrades surfactant proteins A and D. Am J Respir Cell Mol Biol 28:528–537
Meyer KC, Sharma R, Brown M et al (2000) Function and composition of pulmonary surfactant and surfactant-derived fatty acid profiles are altered in young adults with cystic fibrosis. Chest 118:164–174
McCormick CC, Hobden JA, Balzli CL et al (2007) Surfactant protein D in Pseudomonas aeruginosa keratitis. Ocular Immun Inflam 15:371–379
Bainbridge T, Fick RB (1989) Functional importance of cystic fibrosis immunoglobulin G fragments generated by Pseudomonas aeruginosa elastase. J Lab Clin Med 114:728–733
Heck LW, Alarcon PG, Kulhavy RM et al (1990) Degradation of IgA proteins by Pseudomonas aeruginosa elastase. J Immunol 144:2253–2257
Lomholt JA, Kilian M (2008) Degradation of uniquely glycosylated secretory immunoglobulin A in tears from patients with Pseudomonas aeruginosa keratitis. Invest Ophthalmol Vis Sci 49:1944–4939
Tielen P, Rosenau F, Wilhelm S et al (2010) Extracellular enzymes affect biofilm formation of mucoid Pseudomonas aeruginosa. Microbiology 156:2239–2252
Yu H, He X, Xie W et al (2014) Elastase LasB of Pseudomonas aeruginosa promotes biofilm formation partly through rhamnolipid-mediated regulation. Can J Microbiol 60:227–235
Schad PA, Iglewski BH (1988) Nucleotide sequence and expression in Escherichia coli of the Pseudomonas aeruginosa lasA gene. J Bacteriol 170:2784–2789
Kessler E, Safrin M, Gustin JK et al (1998) Elastase and the LasA protease of Pseudomonas aeruginosa are secreted with their propeptides. J Biol Chem 273:30225–30231
Engel LS, Hill JM, Caballero AR (1998) Protease IV, a unique extracellular protease and virulence factor from Pseudomonas aeruginosa. J Biol Chem 273:16792–16797
Wilderman PJ, Vasil AI, Johnson Z (2001) Characterization of an endoprotease (prpl) encoded by a pvds-regulated gene in Pseudomonas aeruginosa. Infect Immun 69:5385–5394
Barequet IS, Bourla N, Pessach YN et al (2012) Staphylolysin is an effective therapeutic agent for Staphylococcus aureus experimental keratitis. Graefes Arch Clin Exp Ophthalmol 250:223–229
Kessler E, Safrin M, Abrams WR, Rosenbloom J, Ohman DE (1997) Inhibitors and specificity of Pseudomonas aeruginosa LasA. J Biol Chem 272:9884–9889
Kessler E, Safrin M, Blumberg S, Ohman DE (2004) A continuous spectrophotometric assay for Pseudomonas aeruginosa LasA prote- ase (staphylolysin) using a two-stage enzymatic reaction. Anal Biochem 328:225–232
Vessillier S, Delolme F, Bernillon J, Saulnier J, Wallach J (2001) Hydrolysis of glycine-containing elastin pentapeptides by LasA, a metalloelastase from Pseudomonas aeruginosa. Eur J Biochem 268:1049–1057
Peters JE, Galloway DR (1990) Purification and characterization of an active fragment of the LasA protein from Pseudomonas aeruginosa: enhancement of elastase activity. J Bacteriol 172:2236–2240
Peters JE, Park SJ, Darzins A et al (1992) Further studies on Pseudomonas aeruginosa LasA: analysis of specificity. Mol Microbiol 6:1155–1162
Laarman AJ, Bardoel BW, Ruyken M et al (2012) Pseudomonas aeruginosa alkaline protease blocks complement activation via the classical and lectin pathways. J Immunol 188:386–393
Kharazmi A, Hoiby N, Doring G, Valerius NH (1984) Pseudomonas aeruginosa exoproteases inhibit human neutrophil chemiluminescence. Infect Immun 44:587–591
Hong YQ, Ghebrehiwet B (1992) Effect of Pseudomonas aeruginosa elastase and alkaline protease on serum complement and isolated components C1q and C3. Clin Immunol Immunopathol 62:133–138
Bardoel BW, van Kessel KP, van Strijp JA, Milder FJ (2012) Inhibition of Pseudomonas aeruginosa virulence: characterization of the AprA-AprI interface and species selectivity. J Mol Biol 415:573–583
Kim SJ, Park RY, Kang SM (2006) Pseudomonas aeruginosa alkaline protease can facilitate siderophore-mediated iron-uptake via the proteolytic cleavage of transferrins. Biol Pharm Bull 29:2295–22300
Gupta SK, Masinick SA, Hobden JA et al (1996) Bacterial proteases and adherence of Pseudomonas aeruginosa to mouse cornea. Exp Eye Res 62:641–650
Malloy JL1, Veldhuizen RA, Thibodeaux BA et al (2005) Pseudomonas aeruginosa protease IV degrades surfactant proteins and inhibits surfactant host defense and biophysical functions. Am J Physiol Lung Cell Mol Physiol 288:409–418
Traidej M, Caballero AR, Marquart ME et al (2003) Molecular analysis of Pseudomonas aeruginosa protease IV expressed in Pseudomonas putida. Invest Ophthalmol Vis Sci 44:190–196
Matsumoto K (2004) Role of bacterial proteases in pseudomonal and serratial keratitis. Biol Chem 385:1007–1016
Engel LS, Hobden JA, Moreau JM et al (1997) Pseudomonas deficient in protease IV has significantly reduced corneal virulence. Invest Ophthalmol Vis Sci 38:1535–1542
Tang A, Marquart ME, Fratkin JD et al (2009) Properties of PASP: a Pseudomonas protease capable of mediating corneal erosions. Invest Ophthalmol Vis Sci 50:3794–3801
Tang A, Caballero AR, Marquart ME, O’callaghan RJ (2013) Pseudomonas aeruginosa small protease (PASP), a keratitis virulence factor. Invest Ophthalmol Vis Sci 54:2821–2828
Kida Y, Shimizu T, Kuwano K (2011) Cooperation between LepA and PlcH contributes to the in vivo virulence and growth of Pseudomonas aeruginosa in mice. Infect Immun 79:211–219
Mochizuki Y, Suzuki T, Oka N, Zhang Y et al (2014) Pseudomonas aeruginosa MucD protease mediates keratitis by inhibiting neutrophil recruitment and promoting bacterial survival. Invest Ophthalmol Vis Sci 55:240–246
Okuda J, Hayashi N, Tanabe S et al (2011) Degradation of interleukin 8 by the serine protease MucD of Pseudomonas aeruginosa. Infect Chemother 17:782–792
Acknowledgements
This study was supported by grants from the following Brazilian Agencies: Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), and Fundação de Amparo à Pesquisa no Estado do Rio de Janeiro (FAPERJ).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Galdino, A.C.M., Branquinha, M.H., Santos, A.L.S., Viganor, L. (2017). Pseudomonas aeruginosa and Its Arsenal of Proteases: Weapons to Battle the Host. In: Chakraborti, S., Dhalla, N. (eds) Pathophysiological Aspects of Proteases. Springer, Singapore. https://doi.org/10.1007/978-981-10-6141-7_16
Download citation
DOI: https://doi.org/10.1007/978-981-10-6141-7_16
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-6140-0
Online ISBN: 978-981-10-6141-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)