Abstract
Comparative studies of closely related species or populations in contrasting environments can potentially provide insights into adaptive mechanisms. We review the phylogeography and population diversity of brackish water species derived from marine species, Ulva prolifera Müller (Ulvophyceae) and Pyropia tenera (Kjellman) N. Kikuchi, M. Miyata, M.S. Hwang & H.G. Choi (Bangiophyceae). Brackish U. prolifera and marine Ulva linza L. have been resolved as closely related species based on phylogenetic analysis of moleclar markers, with U. linza apparently parental to U. prolifera. Our analyses of 5S rDNA spacer region in samples from an Ulvalean bloom in Qingdao, China, indicate that the species appear to be derived from Japanese U. prolifera. Hybridization tests suggest that U. linza and the Qingdao bloom samples are probably distinct species, but gene flow is possible between them. The threatened brackish water species, P. tenera, is morphologically and phylogenetically related to the coastal species, Pyropia yezoensis (Ueda) M.S. Hwang & H.G. Choi. One form, P. yezoensis Ueda f. narawaensis Miura (new combination “Pyropia yezoensis f. narawaensis” has not yet been proposed), is the largest aquaculture source of “Nori” in Japan. Hybridization between these species has been reported, especially between male P. tenera and female P. yezoensis. Sequences of the nrITS1 region and rbcL gene, and PCR-RFLP (ARP4 gene) analyses suggested that P. tenera is distributed across 15 prefectures from Kyushu to Tohoku in Japan; but is restricted to estuarine and brackish water habitats. Based on SSR analysis on the genetically identified P. tenera samples, we conclude that this species spread from Kyushu to Tohoku through the Kanto region.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Macroalgae comprise three major groups: Ulvophyceae (green seaweeds), Phaeophyceae (brown seaweeds) and Rhodophyceae (red seaweeds), which are found on seashores all over the world (van den Hoek et al. 1995). Comparative studies of related species in different environments can potentially elucidate mechanisms of adaptation . In addition to marine species, each group includes brackish and freshwater species, e.g., Dichotomosiphon tuberosus (Braum) Ernst (Bryopsidales, Ulvophyceae), Heribaudiella fluviatilis (Areschoug) Svedelius (Sphacelariales, Phaeophyceae) and Caloglossa continua (Okamura) King et Puttock (Ceramiales, Rhodophyceae) (Guiry et al. 2015). Molecular phylogenetic studies have revealed that many of these brackish/freshwater species evolved from marine ancestors (Lam and Zechman 2006; Krayesky et al. 2012; Silberfeld et al. 2014). Brackish water species have acquired molecular, functional and physiological characteristics required for adaptation to low-salinity conditions. However, many questions about the biological evolution of brackish and freshwater algae are yet to be resolved. Which genes are responsible for this new environmental adaptation? When and where did the brackish/freshwater species separate from marine ancestral species? What are their distributions relative to environmental gradients?
Phylogeography and population genetics are among the most powerful tools used to understand the provenance, genetic structure, distributional expansion and population diversity of organisms (e.g., dinoflagellates, Nagai et al. 2009; crayfish, Koizumi et al. 2012; humans, Jinam et al. 2012; butterflies, Jeratthitikul et al. 2013; hares, Nunome et al. 2014a, b). Land plants have been particularly well studied (van Inghelandt et al. 2010; Kaya et al. 2013; Xu et al. 2014). For instance, the probable distributions of woody plants at the Last Glacial Maximum (LGM, 0.026–0.019 Ma) has been estimated with palaeodistribution modeling (Sakaguchi et al. 2012), endangered plants were interpreted using population diversity and conservation units (Matsuda and Setoguchi 2012), and extinction risks for wild pollinated conifer populations were results from Simple Sequence Repeat (SSR) marker analyses (Iwasaki et al. 2013). These research techniques are also useful for examining the evolutionary history of seaweeds and phylogenetically related brackish and freshwater macroalgae (Perevra et al. 2009). Phylogeographic research can provide important clues to help understand molecular evolution, as it relates to environmental adaptation through all three aquatic environments (freshwater, brackish water, and seawater).
In this chapter, we review the phylogeography and population diversity of brackish water species derived from marine species in two case studies: Ulva prolifera Müller (Ulvophyceae) and Pyropia tenera (Kjellman) N. Kikuchi, M. Miyata, M.S. Hwang & H.G. Choi (Bangiophyceae).
2 Ulva (Ulvophyceae, Chlorophyta)
The genus Ulva is one of the most common seaweeds (Canter-Lund and Lund 1995; van den Hoek et al. 1995; Shimada et al. 2007). New species from the genus have been described year after year (Horimoto et al. 2011; Masakiyo and Shimada 2014; Matsumoto and Shimada 2015). This genus includes brackish species (U. prolifera) (Hiraoka and Shimada 2004) and freshwater species ( Ulva limnetica Ichihara et Shimada) (Ichihara et al. 2009a). Due to the euryhaline nature of Ulva, low-salinity tolerance and adaptation can be found, as reported in many studies (Reed and Russell 1979; Martins et al. 1999; Ohno et al. 1999; Kamer and Fong 2000; McAvoy and Klung 2005; Ichihara et al. 2009b, 2011, 2013).
Ulva prolifera (Fig. 14.1a), which was originally described from Nebbelund, Lolland Island, Denmark (Müller 1778), has branched tubular thalli and is commonly distributed in estuarine and brackish waters (Burrow 1991; Shimada et al. 2008; Hiraoka et al. 2003, 2011). This species is of major importance to the Japanese fisheries industry, as it is a food for human consumption and has a regulated IQ (import quota) under Japanese domestic law (Kawashima et al. 2014). Due to an increasing demand, the supply of U. prolifera is also supplemented by land-based culture techniques in tanks using seawater (Hiraoka and Oka 2008). This species and Ulva linza L. are closely related based on phylogenetic studies (Shimada et al. 2003).
Cultured thalli of molecular-identified Ulva prolifera (a) and Ulva linza (b) under 20 °C, 16:8 h LD, 15–25 μE m−2 s−1 photon flux with PES medium (Provasoli 1968)
Ulva linza (Fig. 14.1b) is distributed in marine habitats, and possesses unbranched distromatic folious thalli without a stipe or margin (Brodie et al. 2007). In freshwater conditions, the cell viability of U. linza dropped to approximately 20 % after 7 days, while that of U. prolifera remained at almost 100 % (Ichihara et al. 2013). These physiological differences seem to be the source of their distinct distributions, with U. prolifera inhabiting estuaries with fluctuations in salinity, while U. linza occurs in marine environments (Ohno et al. 1999; Hiraoka and Shimada 2004). Ulva linza has higher intraspecific diversity than U. prolifera, suggesting that the latter brackish species is derived from marine U. linza (Shimada et al. 2008).
The “green tide” phenomenon occurs under conditions when an extensive biomass of free-floating green algae accumulates on shallow beaches. Free-floating Ulva causes green tide at several locations around the world (Fletcher 1996; Ohno 1999). In June 2008, it was reported worldwide that a vast algal bloom had occurred in Qingdao, China. This massive green-tide covered about 600 km2 along the coast of Qingdao, the host city for the Olympic sailing regatta. An army of more than 10,000 recruits was deployed for a period of one week to help remove the algal bloom. Qingdao city reported that the total mass of the bloom was about 10,000 ton wet weight (BBC NEWS 2008).
Based on satellite imaging data, Liu et al. (2009) proposed the possibility that nonlocal sources were responsible for the Qingdao bloom. According to these authors, on the 15th of May 2008, some small green patches covering a total surface area of about 80 km2 were observed near the coasts of Yancheng and Lianyungang, Jiangsu province. Soon, they grew rapidly and moved into the middle of the Yellow Sea. Finally, a large quantity of biomass formed the famous green tide along the coasts of Qingdao.
The world’s-largest super floating macroalgal bloom of U. prolifera has lasted eight years so far, reoccurring every summer in the Yellow Sea (Xing et al. 2014). The inter-annual variability in human-induced nutrient pollution from 2001 to 2012 was assessed, and a significant increase in nutrient uptake was found in the macroalgal bloom phase (2007–2012). Annual in situ nutrient concentrations increased rapidly from 2000 to 2011 in the Jiangsu Shoal, which was the origin of the drifting macroalgae (Xing et al. 2014).
Shimada et al. (2010) carried out a phylogeographic study of U. prolifera using samples from Japan and the Qingdao bloom. Figures 14.2 and 14.3 show the geographical distribution of sequence types of the 5S rDNA spacer region and the statistical parsimony network of U. prolifera, respectively. Type-A (red circle) individuals are widely distributed along the Pacific coast of Japan from Okinawa to the Kanto region. The blue lineage, derived from type-A in Kanto, radiated to the northern part of Honshu, and spread to the Sea of Japan through the Tsugaru channel (type-N: blue circle). The Qingdao bloom samples (type-QB: yellow circle) were derived from the blue lineage. There are 54 indels between type-QB and type-N. These results indicate that the individuals causing the Qingdao bloom might have recently derived from the Japanese U. prolifera, but are not the same taxon as the latter.
Geographical distributions of sequence types of the 5S rDNA spacer region. Combinations of color and pattern represent different types and are same as Fig. 14.3
Statistical parsimony network of 5S rDNA spacer region. Indels were treated as missing data. Each line connecting sequence types corresponds to one base mutation. A white circle in the parsimony network represents a missing sequence type. The size of circles corresponds to the frequency of each type
Hiraoka et al. (2011) experimentally hybridized individuals of U. linza, Japanese U. prolifera and the Qingdao bloom strain. The sexually reproducing Qingdao strains were successfully crossed with U. prolifera with no evidence of a reproductive barrier, but could not be crossed with U. linza due to gamete incompatibility. However, the results of U. prolifera × U. linza showed an unusual mating activity: male gametes of U. prolifera cannot fuse with female gametes of U. linza, but female gametes of U. prolifera successfully fuse with male gametes of U. linza. The hybrid zygotes can normally develop into sporophytes, and the F1 sporophyte produces zoospores via meiosis. Additionally, the zoospores grow into F1 gametophytes which can produce normal gametes. These results indicate that U. linza and the Qingdao bloom samples are probably different species, but gene flow is nonetheless still possible between them through the Japanese U. prolifera.
Microsatellite/SSR markers have been developed for many organisms due to their unique advantages such as being codominant, highly polymorphic, well distributed throughout the genome, and their simplicity of application for analysis and consistency (Varshney et al. 2002; Tanaka et al. 2011). Recently, Expressed Sequence Tag (EST) -SSR markers were developed for U. prolifera from the south Yellow Sea (Zhang et al. 2014). The analysis of genetic variation using the SSR markers indicated that most individuals collected at the same site clustered together in a tree produced by the unweighted pair-group mean analysis method (UPGMA). The clusters in the tree showed some consistency with the geographical origins of the samples. In addition, 12 of 13 free-floating samples were grouped as a single clade separated from the attached samples. These free-floating samples were collected from different sites at different dates. These results indicate that the free-floating masses of U. prolifera may share the same origin and have been dispersed along coastal areas by wind and ocean currents (Zhang et al. 2014).
Ulva linza and U. prolifera, including the Qingdao blooms, is an ideal species-complex to study speciation and environmental adaptation to low (from U. linza to Japanese U. prolifera) and high (from Japanese U. prolifera to the Qingdao bloom samples) salinities. Kawashima et al. (2013) reported that the 5S rDNA spacer analysis of Japanese and Chinese U. prolifera commercial products detected the potential for intercrossing between U. linza and U. prolifera. Therefore, hybrids of U. linza and U. prolifera are likely to appear in nature. We are now analyzing gene flow between field populations in both species to gain an accurate understanding of their taxonomic status. RNA-seq analysis is being conducted using next generation sequencing (NGS) to elucidate the molecular evolution of low-salinity adaptation in U. prolifera, by comparing the gene expression levels of U. prolifera and U. linza cultured in seawater, brackish water and freshwater conditions. NGS allows us to study gene expression in nonmodel organisms without previous genomic resources (e.g., species-specific primers), and RNA-seq is a very promising application for the study of environmental adaptation (Ekblom and Galindo 2011). RAD -seq (Restriction Site Associated DNA Sequence) from NGS analysis is also useful in studies of speciation, population genetics and phylogeography (Davey and Biaxter 2010; Wagner et al. 2013). NGS analyses will undoubtedly provide a better understanding into algal problems such as green tides . These studies also present new insights into speciation and environmental adaptation, which is the major driver of biological evolution.
3 Pyropia (Bangiophyceae, Rhodophyta)
Pyropia is a foliose red algal genus, consisting of 60 species (Guiry et al. 2015). It is currently recognized as a distinct genus from the closely related genera Porphyra , Boreophyllum , Miuraea , Lysithea , Fuscifolium , and Wildemania (Sutherland et al. 2011). “Nori” (laver), used in sushi, is an edible seaweed product made using thalli of Pyropia species obtained from the field and/or aquaculture, including Pyropia yezoensis (Ueda) M.S. Hwang & H.G. Choi, Pyropia haitanensis (T.J. Chang & B.F. Zheng) N. Kikuchi, M. Miyata, Pyropia pseudolinearis (Ueda) N. Kikuchi, M. Miyata, M.S. Hwang & H.G. Choi and others (Park et al. 2003; Touhata et al. 2013; Xia et al. 2013).
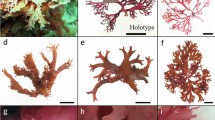
Pyropia yezoensis was originally described from Hokkaido, Japan (Ueda 1932) (Fig. 14.4a). Its type locality is “Muroran?” based on the label of the holotype specimen (Sutherland et al. 2011). One form of this species, P. yezoensis Ueda f. narawaensis Miura (new combination “Pyropia yezoensis f. narawaensis” has not yet been proposed), is the largest aquaculture source of laver in Japan (Miura 1988). Recently, Nakamura et al. (2013) determined a symbiont-free genome sequence of this species using NGS of purified protoplasts. The P. yezoensis genome could represent a model genome for examining red algal life history, and will provide insights into Nori aquaculture in the near future (Nakamura et al. 2013).
Cultured young thalli of molecular-identified samples of Pyropia yezoensis (a) and Pyropia tenera (b) under 15 °C, 8:16 h LD, 15–25 μE m−2 s−1 photon flux with PES medium (Provasoli 1968)
Pyropia tenera (Kjellman) N. Kikuchi, M. Miyata, M.S. Hwang & H.G. Choi was described in Japan based on dried Nori sheets (Kjellman 1897) (Fig. 14.4b). Although this species was used for Nori cultivation from the Edo period (17 century) to the 1960s, P. yezoensis is better than P. tenera in color, gloss and tolerance of strong waves and windy conditions, and thus almost all cultivators now use P. yezoensis in Japanese aquaculture (Miura 1988; Notoya 2004). Pyropia tenera has been categorized as an endangered species (CR + EN) by the Ministry of the Environment in Japan (Environment Agency of Japan 2000). The habitat of this species thought to be limited primarily to the vicinity of the mouths of large and small rivers (brackish water), and only eight localities of P. tenera in Japan were known in 2002 (Kikuchi et al. 2002).
This threatened species, P. tenera, is morphologically and phylogenetically related to P. yezoensis (Yoshida 1998; Sutherland et al. 2011). Hybridization between P. tenera and P. yezoensis has been reported, especially between male P. tenera and female P. yezoensis (Niwa et al. 2009). The reverse combination (hybridization between female P. tenera and male P. yezoensis) also exists, but does not lead to normal development, and thus it is thought that there are few occurrences in the field (Niwa et al. 2010; Niwa and Sakamoto 2010). Although hybrids between female P. tenera and male P. yezoensis showed P. tenera-type in both nrITS (nuclear encoded Internal Transcribed Spacer) and rbcL (ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit) because of concerted evolution, all four types of individuals (1: pure P. tenera, 2: pure P. yezoensis , 3: hybrids between male P. tenera and female P. yezoensis, and 4: hybrids between female P. tenera and male P. yezoensis) can be clearly distinguished by a combination of Niwa’s methods using the direct sequence of nrITS and rbcL, and PCR-RFLP (Restriction Fragment Length Polymorphism) of the ARP4 gene (Niwa et al. 2009, 2010; Niwa and Sakamoto 2010).
We first determined the distribution of P. tenera and the intercross of P. tenera-yezoensis in 165 samples identified morphologically as P. tenera collected at 46 localities in 17 prefectures around Japan (Ohnishi et al. 2013). Sequences of the nrITS1 region and rbcL gene, and PCR-RFLP (ARP4 gene) indicated that P. tenera occurred at 38 localities in 15 prefectures (Fig. 14.5). Pyropia tenera was restricted to estuarine and brackish water habitats, mainly in the Kyushu region (western part of Japan) and in the Tohoku region, but not in the Sea of Japan. Nemalionopsis tortuosa Yoneda et Yagi (Thoreales, Rhodophyta), designated as an endangered species (CR + EN), is also found in many localities around Kyushu (Shimada et al. 2012). Since several threatened algal species have been reported from Kyushu (Environment Agency of Japan 2000), it is one of the most important regions for the conservation of macroalgae in Japan.
Hybrids between male P. tenera and female P. yezoensis were detected at 11 localities in 8 prefectures, mostly located in Tokai, and Kanto to Tohoku regions (Fig. 14.5). Hybridization was restricted to P. tenera’s brackish water habitat. The original distribution of P. yezoensis is thought to have been from Kanto to Hokkaido (Niwa et al. 2009), thus it is reasonable that the hybrids are most common in the eastern to northern parts of Honshu. As these species are cultivated on the Pacific coast of Japan from Kyushu to Tohoku region (Miura 1988), hybrids in the Nagasaki Prefecture may be the result of crosses between wild P. tenera and cultivated P. yezoensis. Crossing between the two species apparently occurs relatively easily.
Next, from SSR analysis of the molecular-identified P. tenera samples, we determined genotypes of three microsatellite markers (Pye13, Pye41, Pye53 of Niwa et al. 2010). Three genetic groups (G1, G2 and G3) appeared in neighbor-joining trees of inter-population genetic distances (Nei and Tajima 1983) computed with POPULATION 1.2.31 (http://bioinformatics.org/~tryphon/populations/) (Fig. 14.6). G1 comprised Kumamoto and Nagasaki populations (Kyushu region) with a high bootstrap value (85 %). Populations of G2 (51 % bootstrap value) were mainly distributed along the Pacific coast from the Kanto to Tohoku region. G3 was supported by a bootstrap value of only 46 %, and the populations in this cluster were geographically scattered (Fig. 14.6).
Population tree of Japanese Pyropia tenera constructed by neighbor-joining method with D A data from three SSR markers. Modified from Ohnishi et al. (2013)
The cluster analysis generated using STRUCTURE (Pritchard et al. 2000) and Structure Harvester v. 2.3.1 (Earl and von Holdt 2012) revealed four genetically differentiated clusters in the Japanese P. tenera (Fig. 14.7). The clustering patterns for K = 4 were highly consistent for 10 independent runs. The result was consistent with the previous POPULATION analysis. Cluster 1 (yellow) was present in only Kumamoto and Nagasaki populations, which is consistent with the G1 POPULATION analysis. Cluster 2 (red) appeared mostly in G2 populations. Clusters 3 (green) and 4 (blue) were detected in almost all populations, indicating that these lineages in these clusters might be ancestral. G3 populations (Fukuoka, Aichi, Ohita, Hiroshima, and Kanagawa populations) overlapped at least 50 % with clusters 3 and 4 and were absent cluster 1.
Best clustering result (K = 4 clusters) for the 14 populations of the Japanese Pyropia tenera based on the STRUCTURE method. a Cluster distribution in each prefecture, b bar plot showing each individual. Each individual is represented by a vertical line partitioned into colored segments; the segment length is proportional to the individual’s estimated K cluster membership coefficient. The sampled populations are separated by black vertical lines. c Neighbor-joining tree of the four clusters. Modified from Ohnishi et al. (2013)
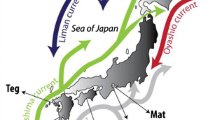
The populations of Kumamoto Prefecture had higher levels of genetic diversity (Allele Richness (AR): 1.542, H E (expected heterozygosity): 0.508, F IS (inbreeding coefficient of an individual with respect to the local subpopulation): 0.556) and Nagasaki (AR: 1.615, H E: 0.582, F IS: 0.575) than the average of all the other populations (AR: 1.444, H E: 0.278, F IS: 0.239). This large genetic diversity may indicate a long stable history during which diversity has accumulated. However, the high F IS indicated inbreeding or subpopulations. On the other hand, the populations in the region extending from Kanto to Tohoku region consisted largely of cluster 2 (red) and had moderately low levels of genetic diversity (AR: 1.291–1.389, H E: 0.167–0.323) (Table 6 in Ohnishi et al. 2013). This result and the direction of the Kuroshio Current , flowing from the southern to the northern Pacific coast of Japan, indicate that P. tenera probably spread from Kyushu to Tohoku through the Kanto region.
Recently, P. tenera has been rediscovered at the Tamagawa River (Kanagawa Prefecture, Kanto Region), on the border between Tokyo and Kanagawa, near Haneda International Airport, (Kikuchi and Niwa 2006). In Tokyo Bay, the mouth of the Tamagawa River, P. tenera was farmed from the seventeenth century until the 1950s (Okamura 1909; Miyashita 2003). However, since the 1950s, aquaculture of P. tenera has not been carried out in the vicinity of Tokyo Bay due to increased landfill and deterioration of farming environments (Kawasaki City Museum 1995). The samples from the Tamagawa River site often showed ancestral clusters 3 and 4 in our SSR analysis. These results indicate that the populations of P. tenera at Tamagawa River may include remnant descendants from the Edo Period. Also, a population-specific allele has been observed in this area (allele 229 of pye53) (Table 7 in Ohnishi et al. 2013). In recent years, however, the number of individuals in this region has declined, increasing the risk of extinction (Kikuchi and Niwa 2006). In order to protect this area-specific genetic diversity , specific conservation activities are a high priority.
There are a few reports about differences between the two species: P. tenera blades are softer in texture and considered better in taste and flavor (Ueda 1932), and it easier to infect with a virus than P. yezoensis (Tatyana et al. 2012). However, it is very difficult to identify P. tenera, P. yezoensis and their hybrids by gross morphology alone (Niwa et al. 2005; Kikuchi 2014). By using molecular-identified samples, researchers can better investigate differences in their gross morphology and color, growth rate, adaptation to marine/brackish water habitats, taste and flavor, and resistance to viral infections. It is hoped that such studies can contribute to elucidating the mechanisms of morphological/physiological evolution of seaweeds.
References
BBC NEWS. 2008. http://news.bbc.co.uk/2/hi/asia-pacific/7486814.stm.
Brodie J, Maggs CA, John DM. Green seaweeds of Britain and Ireland. London: British Phycological Society; 2007. p. 242.
Burrow EM (1991) Seaweeds of the British Isles, vol 2: chlorophyta. London: British Museum (Natural History), p. 238.
Canter-Lund H, Lund JWG. Freshwater algae. Their microscopic world explored. Bristol: Biopress; 1995.
Davey JW, Biaxter ML. RADseq: next-generation population genetics. Brief Funct Genom. 2010;9:416–23.
Earl D, von Holdt B. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Res. 2012;4:359–61.
Ekblom R, Galindo J. Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity. 2011;107:1–15.
Environment Agency of Japan. In: Threatened wildlife of Japan. Red data book, 2nd ed. 2000.
Fletcher RT. The occurrence of “green tide”. In: Schramm W, Nienhuis PH, editors. Marine benthic vegetation—recent changes and the effects of eutrophication. Berlin: Springer; 1996. p. 7–43.
Guiry M, Guiry MD, Guiry GM. AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. 2015. http://www.algaebase.org; Accessed 21 Jan 2015.
Hiraoka M, Oka N. Tank cultivation of Ulva prolifera in deep seawater using a new “germling cluster” method. J Appl Phycol. 2008;20:97–102.
Hiraoka M, Shimada S. Biology of a special green laver, Ulva prolifera from the Shimanto River. Aquabiology. 2004;26:508–15.
Hiraoka M, Dan A, Shimada S, Hagihira M, Migita M, Ohno M. Different life histories of Enteromorpha prolifera (Ulvales, Chlorophyta) from four rivers on Shikoku Island, Japan. Phycologia. 2003;42:275–84.
Hiraoka M, Ichihara K, Zhu W, Ma J, Shimada S. Culture and hybridization experiments on an Ulva clade including the Qingdao strains blooming in the Yellow sea. PLoS ONE. 2011;6(5):e19371.
Horimoto R, Masakiyo Y, Ichihara K, Shimada S. Enteromorpha-like Ulva (Ulvophyceae, Chlorophyta) growing in the Todoroki river, Ishigaki island, Japan, with special reference to Ulva meridionalis Horimoto et Shimada sp. nov. Bull Nat Sci Mus. 2011;37:155–67.
Ichihara K, Arai S, Uchimura M, Fay EJ, Ebata H, Hiraoka M, Shimada S. New species of freshwater Ulva, Ulva limnetica (Ulvales, Ulvophyceae) from Ryukyu archipelago, Japan. Phycol Res. 2009a;57:94–103.
Ichihara K, Arai S, Shimada S. cDNA cloning of a lectin-like gene preferentially expressed in freshwater from the macroalga Ulva limnetica (Ulvales, Chlorophyta). Phycol Res. 2009b;57:104–10.
Ichihara K, Mineur F, Shimada S. Isolation and temporal expression analysis of freshwater-induced genes in Ulva limnetica (Ulvales, Chlorophyta). J Phycol. 2011;47:584–90.
Ichihara K, Miyaji K, Shimada S. Comparing the low-salinity tolerance of Ulva species distributed in different environments. Phycol Res. 2013;61:52–7.
Iwasaki T, Sase T, Takeda S, Ohsawa TA, Ozaki K, Tani N, Ikeda H, Suzuki M, Endo R, Tohei K, Watano Y. Extensive selfing in an endangered population of Pinus parviflora var. parviflora (Pinaceae) in the Boso hills, Japan. Tree Genet Genomes. 2013;9:693–705.
Jeratthitikul E, Hara T, Yago M, Itoh T, Wang M, Usami S, Hikida S. Phylogeography of Fischer’s blue, Tongeia fischeri, in Japan: evidence for introgressive hybridization. Mol Phylogenet Evol. 2013;66:316–26.
Jinam T, Nishida N, Hirai M, Kawamura S, Oota H, Umetsu K, Kimura R, Ohashi J, Tajima A, Yamamoto T, Tanabe H, Mano S, Suto Y, Kaname T, Naritomi K, Yanagi K, Niikawa N, Omoto K, Tokunaga K, Saitou N. The history of human populations in the Japanese archipelago inferred from genome-wide SNP data with a special reference to the Ainu and the Ryukyuan populations. J Human Genet. 2012;57:787–95.
Kamer K, Fong P. A fluctuating salinity regime mitigates the negative effects of reduced salinity on the estuarine macroalga, Enteromorpha intestinalis (L.) link. J Exp Mar Biol Ecol. 2000;254:53–69.
Kawasaki City Museum. Sea and life. The period when Nori was foaming at Kawasaki. Kawasaki: Kawasaki City Museum; 1995.
Kawashima Y, Akasaki T, Matsumoto Y, Yamazaki Y, Shimada S. Species identification of imported and Japanese commercial green algal products, based on phylogenetic analyses using the nrITS2 and 5S rDNA spacer regions. Fish Sci. 2013;79:521–9.
Kawashima Y, Akasaki T, Matsumoto Y, Shimada S. Development of a rapid and accurate PCR-based detection method for commercially valuable green algal species. Fish Sci. 2014;80:859–67.
Kaya HB, Cetin O, Kaya H, Sahin M, Sefer F, Kahraman A, Tanyolac B. SNP discovery by Illumina-based transcriptome sequencing of the olive and the genetic characterization of Turkish olive genotypes revealed by AFLP, SSR and SNP markers. PLoS ONE. 2013;8(9):e73674.
Kikuchi N. Discrimination between Pyropia species and forma by morphological data. In: Fujiyoshi E, Tamaki M, Kobayashi M, Aritaki M, editors. Characteristics of Nori cultivars. Nagasaki: Seikai National Fisheries Research Institute; 2014.
Kikuchi N, Niwa K. Habitat and morphology of the endangered species Porphyra tenera (Bangiales, Rhodophyta) at the estuary of Tamagawa River in Tokyo Bay. Jap J Phycol. 2006;54:149–56.
Kikuchi M, Yoshida T, Yoshinaga K. Distribution of some endangered species of Porphyra. Ecosophia. 2002;9:112–7.
Kjellman FR. Japanska arter af slägtet Porphyra. Bihang Till K. Svenska Vet Akad Handlingar. 1897;23:1–34.
Koizumi I, Usio N, Kawai T, Azuma N, Masuda R. Loss of genetic diversity means loss of geological information: the endangered Japanese crayfish exhibits remarkable historical footprints. PLoS ONE. 2012;7(3):e33986.
Krayesky DM, Norris JN, West JA, Kamiya M, Viguerie M, Wysor BS, Fredericq S. Two new species of Caloglossa (Delesseriaceae, Rhodophyta) from the Americas, C. confusa and C. fluviatilis. Phycologia. 2012;51(5):513–30.
Lam DW, Zechman FW. Phylogenetic analyses of the Bryopsidales (Ulvophyceae, Chlorophyta) based on rubisco large subunit gene sequences. J Phycol. 2006;42:669–78.
Liu D, Keesing JK, Xing Q, Shi P. World’s largest macroalgal bloom caused by expansion of seaweed aquaculture in China. Mar Pollut Bull. 2009;58:888–95.
Martins I, Oliveria JM, Flindt MR, Marques JC. The effect of salinity on the growth rate of the macroalgae Enteromorpha intestinalis (Chlorophyta) in the Mondego estuary (West Portugal). Acta Oecol. 1999;20:259–65.
Masakiyo Y, Shimada S. Species diversity of the genus Ulva (Ulvophyceae, Chlorophyta) in Japanese waters, with special reference to Ulva tepida Masakiyo et S. Shimada sp. nov. Bull Nat Mus Nat Sci Ser B. 2014;40:1–13.
Matsuda J, Setoguchi H. Isolation and characterization of microsatellite loci in Asarum leucosepalum (Aristolochiaceae), an endangered plant endemic to Tokunoshima Island in the Ryukyu Archipelago. Conserv Genet Res. 2012;4:579–81.
Matsumoto M, Shimada S. Systematics of green algae resembling Ulva conglobata, with a description of Ulva adhaerens sp. nov. (Ulvales, Ulvophyceae). Eur J Phycol. 2015;50:100–11.
McAvoy KM, Klug JL. Positive and negative effects of riverine input on the estuarine green alga Ulva intestinalis (syn. Enteromorpha intestinalis) (Linnaeus). Hydrobiologia. 2005;545:1–9.
Miura A. Taxonomic studies of Porphyra species cultivated in Japan, referring to their transition to the cultivated variety. J Tokyo Univ Fish. 1988;75:311–25.
Miyashita A. Thing and human cultural history. Tokyo: Hosei University Press; 2003.
Müller OF (1778) Icones plantarum. Florae danicae, vol. 5, fasc 13. Copenhagem.
Nagai S, Nishitani G, Sakamoto S, Sugaya T, Lee CK, Kim CH, Itakura S, Yamaguchi M. Genetic structuring and transfer of marine dinoflagellate Cochlodinium polykrikoides in Japanese and Korean coastal waters revealed by microsatellites. Mol Ecol. 2009;18:2337–52.
Nakamura Y, Sasaki N, Kobayashi M, Ojima N, Yasuike M, Shigenobu Y, Satomi M, Fukuma Y, Shiwaku K, Tsujimoto A, Kobayashi T, Nakayama I, Ito F, Nakajima K, Sano M, Wada T, Kuhara S, Inouye K, Gojobori T, Ikeo K. The first symbiont-free genome sequence of marine red alga, susabi-nori (Pyropia yezoensis). PLoS ONE. 2013;8(3):e57122.
Nei M, Tajima F. Maximum likelihood estimation of the number of nucleotide substitutions from restriction sites data. Genetics. 1983;105:207–17.
Niwa K, Sakamoto T. Allopolyploidy in natural and cultivated populations of Porphyra. J Phycol. 2010;46:1097–105.
Niwa K, Kikuchi N, Aruga Y. Morphological and molecular analysis of the endangered species Porphyra tenera (Bangiales, Rhodophyta). J Phycol. 2005;41:294–304.
Niwa K, Iida S, Kato A, Kawai H, Kikuchi N, Kobiyama A, Aruga Y. Genetic diversity and introgression in two cultivated species (Porphyra yezoensis and Porphyra tenera) and closely related wild species of Porphyra (Bangiales, Rhodophyta). J Phycol. 2009;45:493–502.
Niwa K, Kobiyama A, Sakamoto T. Interspecific hybridization in the haploid blade-forming marine crop Porphyra (Bangiales, Rhodophyta): occurrence of allodiploidy in surviving F1 gametophytic blades. J Phycol. 2010;46:693–702.
Notoya M. Porphyra. In: Ohno M, editor. Biology and technology of economic seaweeds. Tokyo: Uchida Rokakuho; 2004.
Nunome M, Kinoshita G, Tomozawa M, Torii H, Matsuki R, Yamada F, Matsuda Y, Suzuki H. Lack of association between winter coat colour and genetic population structure in the Japanese hare, Lepus brachyurus (Lagomorpha: Leporidae). Biol J Linn Soc. 2014a;111(4):761–76.
Nunome M, Kinoshida G, Motozawa M, Torii H, Matsuki R, Yamada F, Matsuda Y, Suzuki H. Lack of association between winter coat colour and genetic population structure in the Japanese hare, Lepus brachyurus (Lagomorpha: Leporidae). Biol J Linn Soc. 2014b;111:761–76.
Ohnishi M, Kikuchi N, Iwasaki T, Kawaguchi R, Shimada S. Population genomic structures of endangered species (CR + ER), Pyropia tenera (Bangiales, Rhodophyta). Jpn J Phycol. 2013;61:87–96.
Ohno M. Ulva and extensive biomass. In: Notoya M, editor. Utilization of Ulva and restoration of the environment. Tokyo: Seizando; 1999. p. 1–15.
Ohno M, Mizutani Y, Taino Takahashi I. Ecology of the edible green alga Enteromorpha prolifera in Shimanto River, Southern Japan. Bull Mar Sci Fish Kochi Univ. 1999;19:27–35.
Okamura K. Asakusa Nori (Porphyra tenera). Tokyo: Hakubunn-kan; 1909.
Park CS, Hwang EK, Sohn CH. A atable seeding method for Porphyra pseudolinearis Ueda (Rhodophyta): developing a new species for cultivation of Porphyra in Korea. Aquat Res. 2003;34:895–8.
Perevra RT, Bergström L, Kautsky L, Johannesson K. Rapid speciation in a newly opened postglacial marine environment, the Baltic Sea. BMC Evol Biol. 2009;9:70.
Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000;155:945–59.
Provasoli L. Media and prospects for the cultivation of marine algae. In: Watanabe A, Hattori A (eds) Cultures and collections of Algae. Proceedings of US Japan conference; September 1966, Hakone, Japan. Tokyo: Japanese Society of Plant Physiologists; 1968. pp 63–75.
Reed RH, Russell G. Adaptation to salinity stress in populations of Enteromorpha intestinalis (L.) link. Estu Coast Mar Sci. 1979;8:251–8.
Sakaguchi S, Qiu YX, Liu YH, Qi XS, Kim SH, Han J, Takeuchi Y, Worth JR, Yamasaki M, Sakurai S, Isagi Y. Climate oscillation during the quaternary associated with landscape heterogeneity promoted allopatric lineage divergence of a temperate tree Kalopanax septemlobus (Araliaceae) in East Asia. Mol Ecol. 2012;21(15):3823–38.
Shimada S, Hiraoka M, Nabata S, Iima M, Masuda M. Molecular phylogenetic analyses of the Japanese Ulva and Enteromorpha (Ulvales, Ulvophyceae), with special reference to the free-floating Ulva. Phycol Res. 2003;51:99–108.
Shimada S, Yokoyama N, Masuda M. Genus Ulva (Ulvophyceae, Chlorophyta) in Hokkaido, Japan. Jpn J Bot. 2007;82:190–204.
Shimada S, Yokoyama N, Arai S, Hiraoka M. Phylogeography of the genus Ulva (Ulvophyceae, Chlorophyta), with special reference to the Japanese freshwater and brackish taxa. J Appl Phycol. 2008;20:979–89.
Shimada S, Nagano M, Hiraoka M, Ichihara K, Minerur F, Zhu W. Phylogeographic analysis of the genus Ulva (Ulvales, Chlorophyta), including bloom sample in Qingdao, China. Coast Mar Sci. 2010;34:117–22.
Shimada S, Ichihara K, Masakiyo Y, Iima M, Yoshida Y, Kumano S. Threatened species Nemalionopsis tortuosa (Thoreales, Rhodophyta) in Japan, new locality and current condition of its all reported habitats. Algal Res. 2012;5:9–16.
Silberfeld T, Rousseeau F, de Reviers B. An updated classification of brown algae (Ochrophyta, Phaeophyceae). Cryptog Algol. 2014;35(2):117–56.
Sutherland JE, Lindstrom SC, Nelson WA, Brodie J, Lynch MD, Hwang MS, Choi HG, Miyata M, Kikuchi N, Oliveira MC, Farr T, Neefus C, Mols-Mortensen A, Milstei D, Müller KM. A new look at an ancient order: generic revision of the Bangiales (Rhodophyta). J Phycol. 2011;47(5):1131–51.
Tanaka N, Demise T, Ishii M, Shji Y, Nakaoka M. Genetic structure and gene flow of eelgrass Zostera marina populations in Tokyo Bay, Japan: implications for their restoration. Mar Biol. 2011;158:871–82.
Tatyana A, Shim JB, Hwang MS, Kim GH. Host-parasiteinteractions and host species susceptibility of the marine oomycete parasite, Olpidiopsis sp., from Korea that infects red algae. J Appl Phycol. 2012;24:135–44.
Touhata K, Namikoshi A, Suzuki T, Iguchi J, Mizusawa N, Hara T, Imamura S, Yabu T, Yamashita Y, Yamashita M. Origin identification of dried seaweed product “Nori” by PCR–RFLP analysis of Pyropia yezoensis in the internal transcribed spacer ITS-1 region. Fish Sci. 2013;79:865–75.
Ueda S. Systematic study of the genus Porphyra in Japan. Suiko Kenkyu Kokoku. 1932;28:1–45.
van den Hoek C, Man DG, Jahns HM. Algae. An introduction to phycology. Cambridge: Cambridge University Press; 1995.
van Inghelandt D, Melchinger AE, Lebreton C, Stich B. Population structure and genetic diversity in a commercial maize breeding program assessed with SSR and SNP markers. Theor Appl Genet. 2010;120:1289–99.
Varshney RK, Thiel T, Stein N, Langridge P, Graner A. In silico analysis of frequency and distribution of microsatellites in ESTs of some cereal species. Cell Mol Biol Lett. 2002;7(2A):537–46.
Wagner CE, Keller I, Wittwer S, Selz OM, Mwaiko S, Greuter L, Sivasundar A, Seehausen O. Genome-wide RAD sequence data provide unprecedented resolution of species boundaries and relationships in the Lake Victoria cichlid adaptive radiation. Mol Ecol. 2013;22(3):787–98.
Xia C, Li B, Ji D, Chen C. Characterization of the global transcriptome for Pyropia haitanensis (Bangiales, Rhodophyta) and development of cSSR markers. BMC Genom. 2013;14:107.
Xing Q, Gao M, Gao X, Tosi L, Schmitt FG, Zhang Y, Shi P, Wei J, Luo Y. Progressive eutrophication behind the world-largest super floating macroalgal bloom in the Yellow Sea. Biogeosci Dis. 2014;11:7029–54.
Xu P, Xu SZ, Wu XH, Tao Y, Wang BG, Wang S, Qin DH, Lu ZF, Li GJ. Population genomic analyses from low-coverage RAD-Seq data: a case study on the non-model cucurbit bottle gourd. Plant J. 2014;77:430–42.
Yoshida T. Marine algae of Japan. Tokyo: Uchida Rokakuho; 1998.
Zhang L, Wang G, Liu C, Chi S, Liu T. Development and utility of EST-SSR markers in Ulva prolifera of the south Yellow Sea. Acta Oceanol Sin. 2014;33(10):105–13.
Acknowledgements
We are deeply indebted to Dr. Takaya Iwasaki of Tokyo Univ., for helpful discussions. We greatly appreciate the stimulating discussions, constructive criticism and valuable suggestions on an early version of the manuscript by Senior Lecturer Alecia Bellgrove and Casual Academic Vanessa Skrzypczyk of Marine Biologist at Deakin University Warrnamboolfor. We also thank the Japanese Society of Phycology for the permission of the reproduction of the figures.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Shimada, S., Ichihara, K., Masakiyo, Y., Kawaguchi, R., Kikuchi, N. (2016). Phylogeography of Macroalgal Species Distributed in Brackish Water: Ulva prolifera (Ulvophyceae) and Pyropia tenera (Bangiophyceae). In: Hu, ZM., Fraser, C. (eds) Seaweed Phylogeography. Springer, Dordrecht. https://doi.org/10.1007/978-94-017-7534-2_14
Download citation
DOI: https://doi.org/10.1007/978-94-017-7534-2_14
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-017-7532-8
Online ISBN: 978-94-017-7534-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)