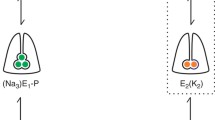Abstract
All animals are characterized by steep gradients of Na+ and K+ across the plasma membrane, and in spite of their highly similar chemical properties, the ions can be distinguished by numerous channels and transporters. The gradients are generated by the Na+,K+-ATPase, or sodium pump, which pumps out Na+ and takes up K+ at the expense of the chemical energy from ATP. Because the membrane is more permeable to K+ than to Na+, the uneven ion distribution causes a transmembrane voltage difference, and this membrane potential forms the basis for the action potential and for much of the neuronal signaling in general. The potential energy stored in the concentration gradients is also used to drive a large number of the secondary transporters responsible for transmembrane carriage of solutes ranging from sugars, amino acids, and neurotransmitters to inorganic ions such as chloride, inorganic phosphate, and bicarbonate. Furthermore, Na+ and K+ themselves are important enzymatic cofactors that typically lower the energy barrier of substrate binding.
In this chapter, we describe the roles of Na+ and K+ in the animal cell with emphasis on the creation and usage of the steep gradients across the membrane. More than 50 years of Na+,K+-ATPase research has revealed many details of the molecular machinery and offered insights into how the pump is regulated by post-translational modifications and specific drugs.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- action potential
- ion gradients
- membrane potential
- Na+ and K+ homeostasis
- ouabain
- palytoxin
- renal tubular system
- secondary transporters
- sodium pump
- voltage-gated channels
- Please cite as: Met. Ions Life Sci. 12 (2013) 41–67
1 Introduction
Studies of the ionic gradients in animals have been fundamental for the development of modern day molecular biology. Already in the 18th century, Luigi Galvani recognized that electrical currents can activate the muscles in a frog leg [1], and in 1902 Ernest Overton found that muscles lose their ability to contract after incubation in an isotonic solution of sucrose, but regain excitability upon addition of sodium. At this time it was already recognized that muscles leak some of their potassium content when stimulated, and after discovering the requirement of extracellular sodium, Ernest Overton very cleverly noted that some counter mechanism that restores these ion gradients must exist [2]. During the Second World War, blood stored cold for transfusions was found to gradually lose potassium, a process that could be reverted by the addition of glucose to restore cellular ATP [3,4]. In 1953, the constant regeneration of the sodium and potassium gradients were found to be sensitive to the cardiotonic glycoside strophanthin [5], and in 1957 the ratio of the energy dependent transport was found to be two K+ for three Na+ [6]. In 1957, Jens Christian Skou finally demonstrated that the Na+/K+ exchange is due to a Mg2+-dependent ATPase, now recognized as the Na+,K+-ATPase or simply the sodium pump [7], a discovery awarded the Nobel prize in 1997.
The sodium pump is responsible for the steep ionic gradients across the plasma membrane with high concentrations of intracellular potassium and extracellular sodium (∼140 mM) and low concentrations of intracellular sodium and extracellular potassium (∼10 mM). A fascinating hypothesis is that the high intracellular potassium concentration is a remnant of the very first proto-cells that evolved on Earth [8]. These cells had neither ion-tight membranes nor membrane pumps, so their intracellular environment would resemble that of the surroundings. The early, most basic cellular machineries thus evolved under ionic conditions similar to those in the primordial pond, but as life spread to different environments, cells evolved sophisticated pumps and transporters that allowed the maintenance of an intracellular solution similar to that found where life originated [8]. This theory also explains why it is a common trait across the kingdoms of life to have an intracellular concentration of potassium much higher than that of sodium.
Life thrives on Earth under exceedingly varying ionic conditions; even in the Dead Sea with NaCl concentrations approaching saturation, archaea, bacteria, and fungi have been described. Organisms that tolerate extreme salt concentration (>2.5 M NaCl) are known as halophilic microorganisms, and they rely on halophilic proteins to cope with the unusual ionic concentrations [9]. To maintain the osmotic balance, some of the microbes synthesize small molecules, e.g., glycerol and amino acids [10]. Others build up high cytosolic levels of K+ [11] and have evolved highly acidic halophilic proteins with an increased number of small hydrophobic residues to favor a negative surface [12].
2 Sodium and Potassium as Enzymatic Cofactors
2.1 Ionic Properties
Sodium and potassium both belong to the group 1 (previously named IA) metals and have similar chemical properties with a relatively small ionic radius and a single valence electron in the s-orbital that is easily lost. In solution, Na+ interacts with three or four water molecules, while the slightly larger K+ optimally coordinates four or five water molecules, but less strongly [13]. An illustrative example of how this difference is utilized in biological systems is the K+-channel, which is highly selective for K+ over Na+ even though Na+ is the smaller ion (Figure 1). The selectivity is achieved by the pore having carbonyl oxygen atoms perfectly substituting for the waters that coordinate free K+ [14] (Figure 1).
The structure of Na+ and K+ selective channels. At the top, the topology within the membrane of a single subunit of a tetrameric bacterial Nav is shown. In mammals, Nav channels are expressed as a single polypeptide with four repeats of structurally similar domains. The selectivity filter is green with the central glutamate shown in stick, and the voltage sensor is purple with arginines shown in sticks. In the middle, an extracellular view of the sodium channel in surface Figure 1 (continued) representation. Ions pass through the central pore, and the four voltage-sensing domains are on the rim of the channel ring. At the bottom, the selectivity filter of the sodium channel is overlayed with that of the potassium channel Kv1.2 (grey). For both channels, only two of the four subunits are shown. In Kv1.2, the potassium-coordinating oxygens are indicated with sticks, and the potassium ions are indicated with red spheres. Within the selectivity filter, there are four potassium binding sites of which 1 and 3 or 2 and 4 will be occupied in a conducting channel. The Nav selectivity filter is shorter with a central glutamate compensating sodium dehydration. The figure was made with pymol (www.pymol.org) using PDB IDs 3RVY (Arcobacter butzleri Nav) and 2A79 (Rattus norvegicus Kv1.2).
When bound within proteins, Na+ and K+ are mostly coordinated by oxygen atoms from amino acid side chains or the backbone, but cases of cation-π interactions with Tyr, Phe or Trp have also been reported (Figure 2a) [15].
Coordination of Na+ and K+ in enzymes. (a) Tagatose-1,6-bisphosphate aldolase coordinates a Na+ (yellow sphere) with four oxygen ligands (dashed green lines) and a π interaction (dashed red line) with a tyrosine. PGH: Phosphoglycolohydroxamic acid. PDB code 1GVF. (b) Diol dehydratase uses five enzyme oxygens to coordinate K+ which is then used to capture the substrate propanediol (PGO) via coordination to the two hydroxyl oxygen atoms. PDB code 1DIO. Figure 2 (continued) (c) Pyruvate kinase coordinates K+ (red sphere) with three oxygen ligands from protein residues and one from the substrate PGA, 2-phosphoglycolic acid (dashed green lines). There is also a Mn2+ ion (green sphere) in the active site. PDB code 1A3W. (d) Dialkylglycine decarboxylase coordinates K+ with five oxygens from amino acids and a single water molecule (dashed green lines). The ion is not directly coordinated by the substrate pyridoxal-5’-phosphate (PLP), but the potassium ion is required for the structure of the active site. PDB code 1DKA. (e) Thrombin coordinates Na+ (yellow sphere) with two oxygen ligands from protein residues and four water molecules (dashed green lines). PDB code 1SG8.
The cellular Na+ and K+ have their lead roles in transmembrane transport and signalling rather than as enzyme regulators. Nonetheless, there are examples of them serving as important facilitators of substrate binding by lowering energy barriers, and there is a strong correlation between the localization of an enzyme and its preferred monovalent ion; intracellular enzymes generally use K+, extracellular enzymes use Na+ [16].
2.2 Sodium- and Potassium-Dependent Enzymes
The chemical properties of Na+ and K+ are utilized by several enzymes. In diol and glycerol dehydratases, K+ is coordinated by five enzyme oxygens and directly interacts with two oxygens from the substrate (Figure 2b). In accordance with the tight substrate-ion interaction, these enzymes are functionally dependent on K+, and in the apo state, the ion remains bound with two water molecules substituting for substrate [17]. Glycerol dehydrogenase is gaining increased interest from the industry because of its crucial role in the production of 1,3-propanediol from fermentation of glycerol. 1,3-propanediol has a potential use in the fabrication of, e.g., polyesters, cosmetics, and lubricants, and glycerol is readily available, since it is a by-product of biodiesel extraction [18].
The phosphoryl transfer enzymes, e.g., pyruvate kinase, also depend on K+ for optimal substrate binding. Together with K+, the enzyme binds a divalent cation, usually Mg2+, and the ions are located at the substrate binding site where they create an optimal docking pocket for phosphoenolpyruvate (Figure 2c). Pyruvate kinases catalyse the final reaction of glycolysis where phosphoenolpyruvate and ADP are converted into pyruvate and ATP. The phosphate part of the substrate couples electrostatically to R49 and K240, but is also stabilized by the positively charged cations. When the phosphate group from the substrate is transferred to ADP, the enzyme uses R49, K240, Mn2+, and K+ as activators [16]. Pyruvate kinases can use Na+ as a substitute for K+, but only with an 8% activity, even though the structures of K+ and Na+ bound enzyme are indistinguishable, emphasizing the importance of their different electrostatic contributions [19].
Dialkylglycine decarboxylase is an example of an enzyme that requires K+, but not in the direct coordination of the substrate. The enzyme catalyzes two coupled half-reactions: the decarboxylation of 2,2-dialkylglycine to CO2 and dialkyl ketone, and the formation of L-alanine from pyruvate. The enzyme is a tetrameric homodimer of dimers with two alkali metal ion binding sites, one at the surface where Na+ binds, and one near the active site where various alkali metals can bind, but the activity is only optimal with K+ [20]. When K+ is bound at the site close to the active site, it is coordinated by five oxygens from amino acids and a single water molecule (Figure 2d), while a Na+ bound at the same site lacks the interactions with T303 and S80. The smaller ionic radius of Na+ therefore leads to a steric clash between S80 and Y301, which reduces enzyme activity.
As many other serine proteases [16], the clotting protease thrombin requires Na+ for activation. Na+ binds in the vicinity of the substrate-binding pocket where it is coordinated by backbone oxygens of K224 and R221 together with four water molecules (Figure 2e). From this location the positive charge of the ion orients D189 for correct engagement with the substrate, and the catalytic S195 is affected through a network of buried water molecules [21].
3 The Membrane Potential
The plasma membrane is practically impermeable to charged solutes such as K+, Na+, and Cl–, but it contains a multitude of protein channels and transporters that allow specific ions to pass under certain conditions. In general, the membrane is most permeable to K+ because of the “two-pore domain” potassium channels that allow a K+ leak through the plasma membrane [14,22,23]. This outward flow of positively charged ions polarizes the membrane, making the inside negative with respect to the outside, until a resting potential is reached, where the chemical driving force of the ion gradient is exactly counterbalanced by the force of the electric field. The equilibrium between the chemical energy associated with movement of ions in a concentration gradient and the electrical energy associated with movement of a charged particle in an electric field is described by the Nernst equation:
where E is the membrane potential required to counterbalance the driving force of the chemical concentration gradient between [X1] and [X2]. R, T. z, and F are the gas constant, the temperature, the valence of the charged particle, and the Faraday number, respectively. If R and F are constant and T is room temperature, the Nernst equation simplifies to:
with E in mV.
The Nernst equation shows that the equilibrium potential of an ion depends on the concentration ratio of its chemical gradient and on its valence. The equilibrium potentials for K+ and Na+ differ between cell types, but they are typically around –80 to –100 mV and +40 to +60 mV, respectively. In addition to K+, the membrane of most cells is also slightly permeable to Na+, and in some instances also to Cl–. The relationship between the membrane potential and the chemical gradients of these ions, when their respective permeabilities are considered, is described in a modified Nernst equation known as the Goldman equation:
where Pion is the membrane permeability to that ion. The Goldman equation predicts that the resting membrane potential of a cell depends on the ionic gradients and on the ionic permeabilities, and since the plasma membrane is more permeable to K+, the resting potential is close to the equilibrium potential of K+, resulting in a membrane potential of –50 to –90 mV in most cells.
3.1 The Action Potential
One of the most illustrative examples of how the cells can utilize the steep gradients of sodium and potassium is the action potential. In neuronal networks, fast intercellular communication occurs via electrical signals made possible by the action of the Na+,K+-ATPase, and the majority of ATP hydrolyzed in the brain is used to fuel the sodium pump.
Neurons are not directly connected to each other, but communicate over structures know as synapses. A synapse has two terminals separated by the synaptic cleft; the transmitting cell signals via the pre-synaptic terminal, and the target cell receives the signal at the post-synaptic terminal. The typical neuron receives the majority of electrical inputs via synapses localized at subcellular domains called spines. Neuronal spines are confined compartments connected to the dendritic shaft via a narrow spine neck. Due to diffusion limitations through the neck, spines are relatively isolated chemically, allowing ion gradients to exist between the spine head and the dendritic shaft [24]. A synapse can be either inhibitory (hyperpolarizing the membrane potential) or excitatory (depolarizing the membrane potential). Inhibitory synapses increase Cl– permeability, causing an influx of negative charge, while excitatory synapses typically increase Na+ permeability to allow an inward flow of Na+, and the Na+ concentration within a spine may rise dramatically during periods of intense activity to levels around 100 mM [25].
The approximately –70 mV resting potential of a neuron is constantly influenced by excitatory and inhibitory signals from other neurons. If the membrane potential becomes depolarized to a certain threshold, an action potential is initiated (Figure 3). The threshold potential is defined by voltage-sensitive channels that are closed at the resting potential, but open when the membrane becomes sufficiently depolarized. Voltage-gated Na+ channels (Nav) open first, so the initial phase is dominated by increased Na+ permeability and thereby further depolarization towards ENa (cf. Figure 3), initiating a positive feedback loop where Na+ permeability increases progressively. The time spent near ENa is limited by two other voltage-dependent events, namely inactivation of Nav and opening of voltage-gated K+ channels (Kv) that let K+ flow out of the cell. The re-polarization phase of an action potential may drive the membrane voltage below the resting potential because the increased K+ permeability does not return to the resting state immediately (Figure 3).
An action potential is initiated when the threshold voltage is reached. This causes a rapid increase in the Na+ permeability because Navs open with fast kinetics and in a positive feedback loop. Before the voltage reaches the equilibrium potential for Na+ the Navs are inactivated in a voltage-dependent manner immediately reducing the Na+ permeability. Kvs open and close with slower kinetics than Navs, and therefore allow the voltage to depolarize before the increased K+ permeability hyperpolarizes the membrane to near the equilibrium potential of K+.
The phenomenon where the membrane hyperpolarizes with respect to the resting potential is known as after-hyperpolarization and is functionally implicated in the regulation of action potential firing patterns. Recently, two independent studies have linked the after-hyperpolarization that follows a period of high frequency stimulation to the activity of the Na+,K+-ATPase. The calyces of Held are large pre-synaptic terminals that fire with high precision at 1 kHz. In rats, large quantities of the sodium pump α3 isoform in the terminals cause hyperpolarization after a period of high activity, which may function to increase firing reliability [26]. A similar discovery was made in Drosophila melanogaster larval motor neurons, where Na+,K+-ATPase activity mediated long lasting hyperpolarization following a train of action potentials, and this was speculated to be involved in a novel short term cellular memory mechanism [27].
The crucial feature of the action potential is the ability of Nav and Kv to sense changes in the membrane potential and respond accordingly. The molecular architecture of the channels is comparable, but while Kv is a homotetramer of subunits with six transmembrane helices (TMs), Nav is a single polypeptide with four similar 6TM domains (Figure 1). The fourth TM helix of a subunit or domain contains several positively charged residues that respond to changes in the membrane potential by moving within the membrane, making TM4 the voltage-sensing domain. Selectivity between K+ and Na+ is obtained by a filter towards the extracellular side, where dehydrated ions are optimally coordinated. Since this compensation is structurally tight, only ions of an exact size fit perfectly and are efficiently allowed to pass.
4 Na+ and K+ Gradients and Secondary Transporters
The potential energy stored in the Na+ gradient is also used to drive a multitude of secondary transporters that translocate solute molecules across the membrane against their concentration gradients. By coupling transport to the Na+ gradient, solute flux can occur at rates 105 times faster than passive diffusion and against concentration gradients up to 106 [28,29]. The Na+ gradient is therefore essential for household cellular tasks like uptake of nutrients as well as for specialized uptake, e.g., in the nervous system. Several atomic structures of members from the different transporter families have now been solved, so we are starting to glimpse the mechanism for how the transporters can be energized by sodium co-transport as described in the following.
4.1 Uptake of Amino Acids, Sugars, and Transmitters
Communication between neurons occurs when an action potential from a transmitting cell reaches a presynaptic terminal and triggers release of neurotransmitters that bind receptors in the post synaptic terminal of the receiving cell. Termination of the signal is facilitated by fast re-uptake of released neurotransmitters by neuronal and glial cells. The majority of transport proteins involved in neurotransmitter re-uptake utilizes energy from the electrochemical Na+ gradient and belong to the family of neurotransmitter sodium symporters (NSSs), which includes GABA, glycine, noradrenaline, serotonin, and dopamine transporters. Malfunctions of these transport systems are linked to disorders like depression, schizophrenia, epilepsy, and Parkinson’s disease [30]. Norepinephrine and serotonin transporters are targets for several antidepressants, including the rather non-selective tricyclic agents and more modern selective serotonin re-uptake inhibitors (SSRIs), as well as psychostimulants like cocaine and amphetamine that also target dopamine transporters [30].
The dominant excitatory neurotransmitter of the central nervous system is the amino acid glutamate, and its re-uptake is carried out by a family of transporters, excitatory amino acid transporters (EAATs), that are structurally distinct from the NSSs, but also couple to the ionic gradients with co-import of three Na+ and a H+ and export of a K+ [31].
Uptake of ions and nutrients is carried out by polarized epithelial cells with an apical membrane that faces the lumen and a basolateral membrane oriented toward the tissue fluid. For instance, in kidney, colon, and small intestine, Na+,K+-ATPases are found exclusively in the basolateral membrane, where they vigorously keep the intracellular Na+ concentration low, while the counter imported K+ rapidly diffuses out of the cells through inwardly rectifying K+ channels, either basolaterally (e.g., Kir4.1/5.1) [32] or apically (e.g., ROMK/Kir1.1) [33].
Nutrient uptake in the small intestine and re-uptake in the kidney are largely carried out by transporters that are structurally and mechanistically similar to the NSSs. Following digestion, peptides and amino acids are absorbed by enterocytes lining the small intestine [34], and in the kidney, the proximal tubule is responsible for the re-absorption of approximately 95–99% of amino acids. Rather than having specific transporters for each amino acid, more general and partly overlapping systems handle uptake of neutral, cationic, and anionic groups. Such redundancy provides backup in case of mutational inactivation of a transport system [35].
In the small intestine and the proximal tubules of the kidneys, sodium-dependent glucose transporters (SGLTs) situated in the apical membrane drive simultaneous uptake of Na+ and glucose from the lumen at the expense of energy stored in the electrochemical Na+ gradient [36], and in the basolateral membrane, facilitative transporters (GLUTs) use the diffusion gradient of glucose to transport sugars into the blood.
X-ray crystal structures of Na+-coupled transporters have revealed common structural architecture, even between transporters with no identifiable sequence conservation. For example, the structures of sugar, amino acid, and the NSSs are evolutionary conserved from bacteria to higher eukaryotes with a characteristic core of two sets of 5TM helices that are interwoven in a 10TM domain with the subdomains related by a pseudo-two-fold axis of symmetry along the plane of the membrane. Structural information about NSSs comes to a large part from studies of a bacterial leucine transporter (LeuT) for which structures of several conformations in the catalytic cycle have been solved, and this has provided unprecedented insight into the molecular details of how the NSS family members work [37,38].
In LeuT, the two subdomains with an inverted topology in the plane of the membrane are comprised by TM1–TM5 and TM6–TM10. The transporter operates in an alternating access fashion, where the open to the outside conformation first binds two Na+ and then the substrate leucine. Upon substrate binding, the access tunnel to the extracellular solution closes, and the transporter transiently adopts an occluded conformation, where Na+ and leucine are shielded from both internal and external solutions, before an internal gate opens to allow the two Na+ and leucine to diffuse into the cell. The transitions from outward over occluded to inward open states and back again require large scale conformational changes, and the energy to drive this process comes solely from the electrochemical Na+ gradient. In some NSSs, the charge transferred during the translocation cycle is partly compensated by co-import of Cl–, and in Cl–-independent transporters, H+ antiport presumably has a similar role [39]. The inverted topology has suggested a ‘rocking-bundle’ theory, where symmetrical movement of the two subdomains during the cycle provides alternating access to either the cytoplasm or the extracellular space [40].
4.2 Co-transporters in Cell Volume and Ion Balances
4.2.1 Cell Volume
The volume of a mammalian cell is relentlessly confronted by changes in extracellular tonicity, transmembrane transfer of osmotically active substances and construction or withdrawal of intracellular osmoles [41]. Since alterations of cell volume can interfere with the structural integrity and the intracellular environment of cells, complex mechanisms continuously operate to maintain the required influx or efflux of water.
Most cells are water-permeable because of specialized water channels known as aquaporins that selectively allow water molecules to cross in single file while charged species, including protons, are impermeable. Since the water flow passively follows the net movement of ions, cell volume regulation is mediated primarily by ion transport systems that affect the overall osmolarity of the intracellular solution.
The gradients of Na+ and K+ drive import and export of Cl– via cation-chloride co-transporters (CCCs), including K+-coupled Cl– exporters (KCCs), Na+-coupled Cl– importers (NCCs), and Na+-coupled K+ and Cl– importers (NKCCs) [42]. Two isoforms of NKCC have been described, NKCC1, which is found in almost all mammalian cells, and NKCC2, which is restricted to the apical membrane of epithelial cells of the thick ascending loop of the kidney [43]. Recent data suggest that while NKCC2 is a dedicated ion transporter, NKCC1 may additionally co-import water along with ions, even against the osmotic gradient [44].
Cell swelling activates a process known as regulatory volume decrease, where the activation of KCCs [45] relieves pressure by exporting K+ and Cl–. Conversely, shrinkage results in activation of regulatory volume increase by Na+/H+ exchangers, NKCCs, and Na+ channels. A specific mechanism in cell volume is regulation of transporters by phosphorylation to activate NKCC1 and inhibit KCC [46]. The consensus within the field is that Cl–/volume sensitive kinases regulate the phosphorylation state of KCC and NKCC1 and hence control the fluxes of water and Cl– [47].
4.2.2 Inhibitory or Excitatory Action of γ-Aminobutyric Acid
In neurons, KCC2 and NKCC1 play important roles in the tuning of the chloride gradient. During development, the expression of NKCC1 decreases, and that of KCC2 increases, effectively resulting in a reversion of the chloride gradient from being highest intracellularly to being lowest. The effects of neurotransmitters like GABA and glycine that open chloride channels are therefore reversed during development from excitatory during pre-natal development to inhibitory in adulthood.
4.2.3 Re-uptake of Inorganic Solutes
CCCs play a pivotal role in the reabsorption of salts from renal corpuscle filtrate in the renal tubules. Nephrons, the functional units of kidneys, include a tubular system that starts with the proximal convoluted tubule which is followed by the loop of Henle that contains a descending and an ascending limp that continues to the distal convoluted tubule and finally the collecting duct. Figure 4 gives an overview of some specialized epithelial cell types along the renal tubules and highlights some of the transport systems that are involved in the reabsorption of inorganic solutes and that are associated with the gradients of Na+ and K+.
The renal tubular system contains several specialized epithelial cell types that expresses different transporters and utilize the cell polarity with an apical and a basolateral membrane to reabsorb a huge amount of solutes from the tubular filtrate. The Na,K-ATPase in the basolateral membrane creates chemical and electrical gradients of Na+ and K+ that drive numerous secondary transporters, some of which are exemplified in this figure.
Phosphate is the most abundant anion in the human body and is implicated in numerous biological functions as bone mineralization, signaling, nucleotide and energy metabolism, and preservation of membrane integrity [48]. In the kidney, re-uptake of inorganic phosphate (Pi) depends on Na+-coupled symporters (NaPi) that are located in the apical membrane of proximal tubule epithelial cells [49].
Sodium-coupled bicarbonate transporters (NCBTs) are important regulators of intra- and extracellular pH, and exist as both electrogenic (NBCe1 and NCBe2) and electroneutral (NBCn1, NBCn2, and NDCBE) forms. In the proximal tubule of the kidney nephrons, NCBe1 is localized in the basolateral membrane where it couples with apically localized Na+-coupled H+ exporters (NHEs) and aquaporins to drive the reabsorption of approximately 80% of the filtered HCO\( {}_{3}^{-} \)which is of crucial importance for maintenance of, e.g., blood pH. The remaining 20% is reabsorbed in more distal parts of the renal tubules so virtually no HCO\( {}_{3}^{-} \)escapes with the urine. HCO\( {}_{3}^{-} \)that in the filtrate is titrated to CO2 and H2O by H+ that is secreted by NHEs. Both CO2 and H2O then enter the proximal tubule epithelium at the apical membrane via aquaporins and are converted into HCO\( {}_{3}^{-} \)by the enzyme carbonic anhydrase before it is exported by the basolateral NCBe1 together with Na+ in a 3:1 ratio [50]. Since NCBe1 is co-exporting Na+ against the electrochemical gradient, the driving force of this transporter is partly the membrane potential [51].
The proximal convoluted tubule is connected to the distal convoluted tubule via the loop of Henle which encompasses a descending and an ascending loop that dives into the medulla of the kidney where the interstitial fluid has a high concentration of solutes. Since water permeability in the descending limp is high, massive amount of water is drawn out of the tubular fluid and into the interstitium, while solutes remain due to a low population of transporters in the descending limp epithelium. The fluid that reaches the ascending limp is consequently highly concentrated and, because the epithelia that line this part of the tubular system does not express aquaporins, only salts are re-absorbed via, e.g., NKCCs and NHEs. In the distal convoluted tubule NCCs are responsible for the reabsorption of approximately 5% of the renal corpuscle filtrate [52].
Electroneutral cation-chloride co-transporters are targets for some of the most prescribed drugs in the world, and are used to treat hypertension and heart failure because they inhibit salt re-uptake, and hence increase water output through urination. NCC represents the major target site for thiazide-type diuretics that are used to treat hypertension, extracellular fluid volume overload, and renal stones. Furosemide is an inhibitor of NKCC and is primarily used in the treatment of hypertension and edema, but has also been the center of some controversy due to its usage on race horses. Fast or intense exercise often causes pulmonary hemorrhage in race horses due to the combination of capillary and alveolar pressure increments. The hemorrhage in the lungs is often seen after a race as bleeding from the nostrils, and the condition results in poor athletic performance. Pre-race administration of furosemide is known to increase performance because it decreases the incidence and severity of exercise-induced pulmonary hemorrhage [53].
5 Homeostasis of the Na+ and K+ Gradients by the Na+,K+-ATPase
5.1 The Subunits of the Na+,K+-ATPase
The Na+,K+-ATPase is an oligomeric protein that, as a minimum, contains an α-subunit with ten transmembrane spanning helixes and a 1TM type II glycosylated β-subunit [54]. All principal ion pump activity is managed by the α-subunit, which contains the translocation channel with ion-binding sites, the ATP hydrolysis core, and the target site for the specific Na+,K+-ATPase inhibitors cardiotonic glycosides. The β-subunit associates with the α-subunit in the endoplasmic reticulum (ER) and is required for stability and sorting to the plasma membrane [55–57]. In addition to the essential α- and β-subunits, the Na+,K+-ATPase may also associate with a 1TM regulatory subunit of the FXYD family of membrane proteins (Figure 5) [58].
Structural overview of the sodium pump in the membrane. In the α-subunit, the transmembrane part is brown, the phosphorylation (P) domain is light blue, the actuator (A) domain is light brown, and the nucleotide (N) domain is red. The β-subunit is in purple and the γ-subunit is in green. The two occluded K+ and the cytoplasmic regulatory K+ are indicated with red spheres. The phosphorylation is mimicked by magnesium fluoride shown as green and light blue spheres. PDB code 2ZXE.
Animal cells depend strongly on the large gradients of Na+ and K+ that Na+,K+-ATPases create and maintain, and the great diversity of cell types within the animal kingdom call for an assortment of various sodium pump subtypes. To fulfill the different requirements, multiple isoforms of the Na+,K+-ATPase subunits have evolved, in mammals there are four different types of α-, three of β-, and seven of FXYD-subunits. Since all isoform combinations yield functional pumps, the potential number of different Na+,K+-ATPases in mammals is 84.
The subunit isoforms of the Na+,K+-ATPase display very distinct expression profiles depending on cell type, tissue and developmental state. The α1 isoform is ubiquitously expressed, α2 is found predominantly in muscle, adipose, heart and lung tissues, and in glial cells [59], the α3 isoform is mostly found in neurons [60], but has also been detected in other tissues, including the developing rat cardiac ventricle [61] and erythroid cells [62]. The most restricted Na+,K+-ATPase isoform is α4, which is only expressed in sperm cells [63,64]. Both β- and FXYD- subunits are viewed as regulators of the principal α-subunit, and their complex expression profiles further illustrate the fine-tuning of pump function required for different cell types [65].
Diverse subunit compositions yield Na+,K+-ATPases optimized to operate under different cellular conditions. The α3 isoform for example is associated with low affinity for intracellular Na+ and high affinity for ATP [65]. In essence, this gives a pump that remains relatively dormant under basal conditions where the level of intracellular Na+ is approximately 10 mM, but in neurons, α3 accumulates in dendritic spines, and the concentration of Na+ increases dramatically during high neuronal activity and will not be limiting. The high affinity for intracellular ATP further ensures that α3 remains active even when energy is low. Another adaptive technique is seen for the α2-subunit isoform, which is the isoform most sensitive to the membrane potential, being inhibited by hyperpolarization.
5.2 The Structure of the Na+,K+-ATPase
The Na+,K+-ATPase belongs to a family of membrane proteins known as the P-type ATPases, where the defining term “P-type” comes from the notion that all members are phosphorylated and dephosphorylated at a conserved aspartate during the reaction cycle (Figure 6) [66]. All P-type ATPases have three intracellular domains that together constitute the engine of the pump, transforming energy in the form of ATP into mechanical energy that opens and closes the enzyme toward the cytoplasm and the extracellular solution in an alternating fashion. The three intracellular domains are named the actuator domain (A-domain), the nucleotide binding domain (N-domain) and the phosphorylation domain (P-domain) [67]. The transmembrane part of the Na+,K+-ATPase α-subunit contains three ion-binding sites within the lipid bilayer. Two of these sites are shared between Na+ and K+, while the third site binds only Na+. The C-terminal tail of the Na+,K+-ATPase is implanted within the transmembrane part of the pump in proximity to the exclusive Na+-binding site. On the extracellularly exposed part of the α-subunit, a cavity serves as an ion entrance/exit vestibule and as a binding site for the specific Na+,K+-ATPase inhibitors cardiotonic glycosides (cf. Figures 7 and 8 in Section 5.5.1).
The catalytic cycle of the sodium pump. The Na+,K+-ATPase exists in two principal conformations; E1 with high Na+ affinity and opened inwards, and E2 with high K+ affinity and opened outward. Each complete cycle results in the export of three Na+, import of two K+ and the hydrolysis of a single molecule of ATP. Recent speculations suggest that a cytoplasmic proton enters the pump and stabilizes it in the K+ bound state, and then returns to the cytoplasm when K+ is released [104].
Membrane view of a low-affinity Na+,K+-ATPase-ouabain complex. Indicated are ouabain, the occluded potassium ions, and the loop between TM1 and TM2, the M1-M2 loop. Surface representations of the subunits α (brown), β (purple), and γ (green). The figure was made with pymol using PDB ID 3A3Y (a shark Na,K-ATPase). In the high-affinity complex, ouabain most likely binds similarly, but helix movements are expected to close the binding site further.
Extracellular view of the structure shown in Figure 7 with similar color coding and with indication of the loop between TM1 and TM2, the M1-M2 loop. The two residues known to be important for ouabain binding are indicated.
The β-subunit is glycosylated and provides the majority of the extracellular architecture. It plays an important role in cell-cell adhesion because two opposing β-subunits can interact [68].
5.3 The Mechanism of the Na+,K+-ATPase
Since the discovery of the Na+,K+-ATPase, much has been learned about the cycle of conformational shifts required to translocate both Na+ and K+ against their respective chemical gradients, and for Na+ also against the electrical potential of the cell. The reaction scheme is named “The Post-Albers Model” after two of the pioneers in its formulation, and it is based on a cycle where the pump alternates between two major states; an E1 state open to the inside of the cell with high affinity for Na+, and an E2 state open to the outside with a high K+ affinity (Figure 6) [69,70]. From the combined biochemical, structural, and electrophysiological data, much of the mechanics driving the pump have been elucidated. High resolution crystal structures of both the Na+,K+-ATPase [71,72] and the closely related sarcoplasmic reticulum Ca2+-ATPase SERCA [73–79], the latter in several conformational states, have given astonishing insight into how the different domains of the pumps move as the cycle progresses.
The N-domain promotes the E2 to E1 conformational shift by binding of ATP. For the sodium pump, this means intracellular release of two K+ and binding of three Na+. The N-domain then phosphorylates the P-domain, whereby the three bound Na+ are occluded within the pump. Release of Na+ is presumably initiated by protonation of a membrane-embedded aspartate by an intracellular proton, and the opening of the structure to the external side is forced by a 90° rotation of the A-domain. In relation to the P-domain, the A-domain rotation situates it in a favorable position for acting as a phosphatase, and dephosphorylation induces occlusion of the two newly bound K+ (Figure 6).
The activity of a membrane-embedded molecular engine such as the Na+,K+-ATPase that catalyzes an electrogenic reaction will be influenced by the transmembrane potential. Energetically, the transport of positive charge against the electrical potential becomes increasingly expensive with hyperpolarization, and since the 3:2 stoichiometry is maintained at all investigated potentials, the pump’s turnover rate will cease if the electrochemical energetic cost of the ion transport approaches the energy of ATP hydrolysis [80].
5.4 Regulation of the Na+,K+-ATPase
The activity of the Na+,K+-ATPase can be regulated by cellular interaction partners and by posttranslational modifications that affect the pump’s kinetic properties or its cellular distribution.
5.4.1 Posttranslational Modifications
The best-studied posttranslational modifications are phosphorylations. In particular, phosphorylation of FXYD1 in heart and skeletal muscle increases pump activity, probably because unphosphorylated FXYD1 lowers sodium and potassium affinities, and phosphorylation releases the inhibitory effect [81]. PKA and PKC target the C-terminal part of FXYD1, and as indicated by its alternative name, phospholemman, it is phosphorylated to a large extent [82].
The α-subunit is also phosphorylated. The most conserved site is a serine in the cytoplasm/membrane interface, which is targeted by PKA. With the use of phosphospecific antibodies, the site has been shown to be phosphorylated in response to activation of a number of G-protein coupled receptors, and the phosphorylation generally leads to inhibition of the pump without affecting the level of pumps at the plasma membrane, suggesting direct inactivation [83]. The α isoforms can all be phosphorylated by PKA; for example, mouse studies showed that α2 phosphorylation levels respond to morphine administration [84]. Yet a detailed knowledge of the cellular pathways and roles of the phosphorylation remain uncertain.
Recently, the sodium pump has also been shown to be subjected to glutathionylation. In cardiac cells experiencing oxidative stress, a cysteine in the transmembrane part of β1 is modified by glutathione, which inhibits the pump [85], but FXYD proteins can reverse the inhibition [86]. During, e.g., heart failure, it may be an important pathophysiological consideration that oxidative stress inhibits the sodium pump, but since it is reversible, pump glutathionylation could in general serve a signaling role.
5.4.2 Cellular Interactions
A rapidly increasing number of protein interaction partners are being described for the sodium pump. In many cases, the interaction is believed to control pump activity by regulating, e.g., its subcellular localization [87,88], but intriguingly, there is also emerging evidence that the pump interacts with channels that functionally couple directly to the ion clearance, so far mainly sodium-selective channels.
The atypical Nax channels are expressed in glia cells in parts of the brain that control salt intake. The Nax channels open in response to high extracellular sodium levels, which leads to increased lactate production in the cell, and lactate functions as a signal to GABAergic neurons. The channel was shown to interact with the P-domains of α1 and α2, but not α3. The interaction was required for elevated sodium levels causing upregulated glucose intake and thereby lactate signalling. Tight coupling of Nax with the Na+,K+-ATPase secures efficient pump activation, since the substrate is directly loaded to the pump, but a direct physical interaction may serve additional functions [89].
The α1 sodium pump subunit was also shown to co-immunoprecipitate with AMPA receptor subunits GluR1 and 4 in lysates from rat cerebellum [90] and with GluR1 and 2 in rat cortical lysates [91]. The majority of excitatory neurotransmission in the CNS is attributable to the AMPA subgroup of the glutamate receptors. The channels are heterotetramers of the GluR1-4 subunits that allow influx of sodium upon activation and thus depend strongly on sodium pump activity. In cortical neurons, inactivation of the sodium pump by ouabain caused up to 80% of the AMPAR subunits to be internalized and subsequently degraded. Like the Nax channel, AMPAR function depends on the sodium gradient created by the Na,K-ATPase, and the interaction study further indicated that the sodium pump may play an active part at the synapse by determining the number of AMPARs and thereby the strength of the synapse [91].
In neuromuscular junctions, the Na+,K+-ATPase interacts with the nicotinic acetylcholine receptor (nAchR) at the postsynapse. The nAchRs are ligand-gated cation channels, so agonist binding leads to an influx of sodium, but also to potassium release. In mammals, the sodium pump α2 colocalizes and copurifies with nAchR, and the desensitized channel enhances pump activity [92]. In Caenorhabditis elegans, the Na+,K+-ATPase (EAT-6) is required for proper neuromuscular localization of the nAchR, again suggesting a critical role for the pump in synaptic organization [93].
It thus seems a recurring theme that the sodium pump associates with some of the channels that depend on it in a regulatory network. We are only now beginning to glimpse the higher-order complexes, but they will undoubtedly be in the focus for a fuller understanding of how the cell maintains and utilizes sodium and potassium gradients.
5.5 Na+,K+-ATPase Toxins
5.5.1 Cardiotonic Glycosides
Inhibitors of the Na+,K+-ATPase are produced by both plants and animals, and for centuries it has been recognized that low doses of the toxins can be used to treat cardiac insufficiency, hence their name, cardiotonic glycosides (Figures 7 and 8). Na+,K+-ATPase inhibition leads to higher intracellular sodium levels and, via the sodium-calcium exchangers, to higher calcium levels, which increases the force of myocardial contractions. However, the therapeutic index is quite narrow, and higher doses are lethal. Apart from accidental intake, human poisoning has mainly been fictional (e.g., in Agatha Cristie novels and a James Bond movie), but African hunters have used extracts with cardiotonic glycosides as arrow poison [94,95].
At least 14 different plant species and nine Bufo toads produce cardiotonic steroids. Furthermore, the larvae of the Monarch butterfly live solely on a plant that produces cardiotonic steroids, and both the larvae and the butterfly retain sufficient amounts of poison for it to serve as a defense against predators [96]. Similarly, two species of the leaf beetles Chrysochus eat plants that produce cardiotonic steroids, making the beetles toxic to predators [97]. Plants energize their membranes with a proton gradient and are therefore not affected by sodium pump inhibitors, but the animals that produce or retain the poison need to protect their own pumps. They avoid inhibition by having mutations in the extracellular part of their α subunits where the cardiotonic steroids otherwise bind (Figure 9).
Alignment of residues involved in ouabain binding. Arrows indicate residues known to affect ouabain affinity at the end of TM1 (corresponding to Gln118 in Figure 8) and at the beginning of TM2 (corresponding to Asn129 in Figure 8). The extracellular loop between TM1 and TM2, the M1-M2 loop, is indicated. Abbreviations used are Homo sapiens (Hs), Gallus gallus (Gg), Xenopus tropicalis (Xt), Rattus norvegicus (Rn), Dugesia japonica (Dj), Bufo marinus (Bm), Danaus plexippus (Dp), Chrysochus auratus (Ca), and Heterosigma akashiwo (Ha). In rodents, α1 is insensitive to ouabain. Cardiotonic glycosides are used for protection by Bufo toads. The monarch butterfly (Dp) and a leaf beetle (Ca) take up cardiotonic glycosides produced by the plants they feed on. Sequences of Na+,K+-ATPases from non-animal species, e.g., a marine algae (Ha), suggest that they are not sensitive to ouabain.
The cardiotonic glycosides inhibit the Na+,K+-ATPase by locking it in an E2P state, thus putting a break in the catalytic cycle. Surprisingly, the cardiotonic steroid ouabain is also being recognized to function as a hormone at very low concentrations (nM), causing intracellular signalling events to change gene expression and cell survival. There is no measurable effect on the Na+,K+-ATPase activity at these concentrations, but it is suggested that locking of a sub-portion of the pumps in the E2P state causes release of a Src kinase interacting with the pump, thus allowing it to start the signaling cascade via phosphorylations [98].
5.5.2 Palytoxin
While cardiotonic glycosides block the Na+,K+-ATPase, palytoxin transforms the pump into a cation channel with an open probability of around 90%. Compared to the normal pump, the palytoxin bound pump therefore allows sodium and potassium to flow in the opposite directions at speeds many thousandfold higher than pumping, so the ionic gradients are destroyed. Loss of the ionic gradients is extremely lethal, causing rapid heart failure upon intravenous exposure, and palytoxin is the second deadliest non-peptide molecule known with an LD50 in mice of less than 50 ng/kg [99].
Palytoxin was first isolated in 1971 from the Hawaiian coral Palythoa toxica. The molecule is huge with 64 stereogenic centers, and the mechanism of action remains for the most part mysterious, but somehow palytoxin recognizes the extracellular part of the pump and causes opening of both intracellular and extracellular ion pathways at the same time [99].
6 Pathophysiology of Na+,K+-ATPase Disturbance
Maintaining the sodium and potassium gradients is essential to all animal cells, so deleterious mutations in the gene for the ubiquitous α1 subunit are not compatible with life. For α2 and α3, haploinsufficiency is tolerated, but can cause the rare human diseases familial hemeplegic migraine 2 (FHM2 [100]) and rapid-onset dystonia parkinsonism (RDP [101]), respectively. FHM2 patients suffer from attacks of migraine with aura and hemiparesis, and the attacks can be accompanied by other symptoms such as ataxia, epilepsy/seizure, and loss of consciousness. In some FHM2 families, there are also reports of, e.g., weakened cognitive functions and psychiatric disorders [102].
Similar symptoms are described for patients with mutations in genes encoding the voltage-gated calcium channel CaV2.1 (causing FHM1) or the voltage-gated sodium channel NaV1.1 α1 subunit (causing FHM3), which suggests that the phenotype reflects an at least partly common mechanism where the ionic balances in the brain are misregulated.
RDP patients very abruptly (within hours to weeks) develop dystonia, where the muscles remain involuntarily contracted, causing twisting of the body, but there are also parkinsonian symptoms, e.g., slow movements and postural instability. The severity of the symptoms follows a rostro-caudal gradient with the face affected the most and the legs the least, and typical medications for parkinsonism have no effect. The onset is triggered by physical or emotional stress, often involving alcohol.
36 persons with RDP from ten families have been identified to have mutations in the ATP1A3 gene, while maybe an order of magnitude more have been identified with FHM2. For both syndromes, healthy family members carrying a disease-causing mutation have been identified, so there is incomplete penetration of the phenotype, and whether a mutation causes disease will depend on other genetic and environmental variations.
7 Concluding Remarks and Future Directions
The sodium and potassium gradients are absolutely required for animal life, since they form the basis for numerous essential cellular processes, and the sodium pump is required to regulate osmolarity, which is vital to the animal cells that, in contrast to, e.g., plant cells, are not protected by a cell wall. Although a Na+,K+-ATPase was probably not the primoridal P-type ATPase [103], it has been essential for metazoan evolution. The original Na+,K+-ATPase may have depended on a single subunit, and in, e.g., algae and archaea, genes predicted to encode the α- and not the β-subunit have been identified, but the β-subunit’s role in cell-cell interactions combined with the ionic gradients created by the α-subunit have likely been prerequisites for the earliest assemblies of polarized cells that evolved into animals.
The studies of sodium and potassium gradients began centuries ago, and the sodium pump has been known for more than fifty years. Nonetheless, there are still many unanswered questions regarding the mechanism, ion pathways and regulation, and the coming years will undoubtedly bring new perspectives on the sodium pump’s role in cellular networks.
Abbreviations
- A-domain:
-
actuator domain
- ADP:
-
adenosine 5’-diphosphate
- AMPA:
-
2-amino-3-(5-methyl-3-oxo-1,2-oxazol-4-yl)propanoic acid
- ATP:
-
adenosine 5’-triphosphate
- CCCs:
-
cation-chloride co-transporters
- CNS:
-
central nervous system
- EAATs:
-
excitatory amino acid transporters
- ER:
-
endoplasmic reticulum
- FHM2:
-
familial hemeplegic migraine 2
- GABA:
-
γ-aminobutyric acid
- GLUTs:
-
glucose transporters
- KCCs:
-
K+-coupled Cl– exporters
- Kv :
-
voltage-gated K+ channels
- LD50 :
-
lethal dose, 50%
- LeuT:
-
leucine transporter
- nAchR:
-
nicotinic acetylcholine receptor
- NaPi:
-
Na+-coupled Pi symporter
- Nav :
-
voltage-gated Na+ channels
- Nax :
-
subfamily of voltage-gated sodium channels (formerly Nav2.1 in humans)
- NCBTs:
-
sodium-coupled bicarbonate transporters
- NCCs:
-
Na+-coupled Cl– importers
- N-domain:
-
nucleotide-binding domain
- NHEs:
-
Na+-coupled H+ exporters
- NKCCs:
-
Na+-coupled K+ and Cl– importers
- NSSs:
-
neurotransmitter sodium symporters
- P-domain:
-
phosphorylation domain
- Pi :
-
inorganic phosphate
- PKA:
-
protein kinase A
- PKC:
-
protein kinase C
- RDP:
-
rapid-onset dystonia parkinsonism
- SGLTs:
-
sodium-dependent glucose transporters
- SSRIs:
-
selective serotonin re-uptake inhibitors
- TMs:
-
transmembrane helices
References
C. McCaig, A. Rajnicek, B. Song, M. Zhao, Physiol. Rev. 2005, 85, 943–1021.
E. Overton, Pflügers Arch. 1902, 92, 346–386.
T. Danowski, J. Biol. Chem. 1941, 139, 693–705.
J. Harris, J. Biol. Chem. 1941, 141, 579–595.
H. Schatzmann, Helv. Physiol. Pharmacol. Acta 1953, 11, 346–400.
R. Post, P. Jolly, Biochim. Biophys. Acta 1957, 25, 118–146.
J. Skou, Biochim. Biophys. Acta 1957, 23, 394–795.
A. Mulkidjanian, A. Bychkov, D. Dibrova, M. Galperin, E. Koonin, Proc. Nat. Acad. Sci. USA 2012, 109, 30.
D. Madern, C. Ebel, G. Zaccai, Extremophiles: Life under Extreme Conditions 2000, 4, 91–99.
P. Yancey, J. Exper. Biol. 2005, 208, 2819–2849.
R. Vreeland, Crit. Rev. Microbiol. 1987, 14, 311–367.
S. Kennedy, W. Ng, S. Salzberg, L. Hood, S. DasSarma, Genome Res. 2001, 11, 1641–1691.
K. Collins, Biophys. J. 1997, 72, 65–141.
D. Doyle, J. Morais Cabral, R. Pfuetzner, A. Kuo, J. Gulbis, S. Cohen, B. Chait, R. MacKinnon, Science 1998, 280, 69–146.
D. Hall, C. Bond, G. Leonard, C. Watt, A. Berry, W. Hunter, J. Biol. Chem. 2002, 277, 22018–22042.
M. Page, E. Di Cera, Physiol. Rev. 2006, 86, 1049–1141.
N. Shibata, J. Masuda, T. Tobimatsu, T. Toraya, K. Suto, Y. Morimoto, N. Yasuoka, Structure 1999, 7, 997–2005.
E. Wilkens, A. Ringel, D. Hortig, T. Willke, K.-D. Vorlop, Appl. Microbiol. Biotechnol. 2012, 93, 1057–1120.
T. Larsen, M. Benning, I. Rayment, G. Reed, Biochemistry 1998, 37, 6247–6302.
M. Toney, E. Hohenester, J. Keller, J. Jansonius, J. Mol. Biol. 1995, 245, 151–230.
A. Pineda, C. Carrell, L. Bush, S. Prasad, S. Caccia, Z.-W. Chen, F. Mathews, E. Di Cera, J. Biol. Chem. 2004, 279, 31842–31895.
S. Brohawn, J. del Mármol, R. MacKinnon, Science 2012, 335, 436–477.
A. Miller, S. Long, Science 2012, 335, 432–438.
K. Svoboda, D. Tank, W. Denk, Science 1996, 272, 716–725.
C. Rose, A. Konnerth, J. Neurosci. 2001, 21, 4207–4221.
J. Kim, I. Sizov, M. Dobretsov, H. von Gersdorff, Nature Neuroscience 2007, 10, 196–401.
S. Pulver, L. Griffith, Nature Neuroscience 2010, 13, 53–62.
A. Chakrabarti, D. Deamer, Biochim. Biophys. Acta 1992, 1111, 171–178.
M. Roux, S. Supplisson, Neuron 2000, 25, 373–456.
M. Hahn, R. Blakely, Pharmacogenomics J. 2002, 2, 217–252.
N. Zerangue, M. Kavanaugh, Nature 1996, 383, 634–641.
S. Lachheb, F. Cluzeaud, M. Bens, M. Genete, H. Hibino, S. Lourdel, Y. Kurachi, A. Vandewalle, J. Teulon, M. Paulais, Am. J. Physiology. Renal Physiology 2008, 294, 407.
P. Welling, K. Ho, Am. J. Physiology. Renal Physiology 2009, 297, 63.
S. Adibi, S. Gray, E. Menden, Am. J. Clin. Nutrition 1967, 20, 24–57.
S. Bröer, Physiol. Rev. 2008, 88, 249–335.
E. Wright, D. Loo, B. Hirayama, Physiol. Rev. 2011, 91, 733–827.
H. Krishnamurthy, E. Gouaux, Nature 2012, 481, 469–543.
A. Yamashita, S. Singh, T. Kawate, Y. Jin, E. Gouaux, Nature 2005, 437, 215–238.
Y. Zhao, M. Quick, L. Shi, E. Mehler, H. Weinstein, J. Javitch, Nature Chem. Biol. 2010, 6, 109–125.
L. Forrest, R. Krämer, C. Ziegler, Biochim. Biophys. Acta 2011, 1807, 167–255.
F. Lang, G. Busch, H. Völkl, Cell. Physiol. Biochem.: Int. J. Exper. Cell. Physiol., Biochem., Pharmacol. 1998, 8, 1–46.
C. Lytle, J. Biol. Chem. 1997, 272, 15069–15146.
J. Russell, Physiol. Rev. 2000, 80, 211–287.
T. Zeuthen, N. Macaulay, J. Physiol. 2012, 590, 1139–1193.
P. Dunham, G. Stewart, J. Ellory, Proc. Nat. Acad. Sci. USA 1980, 77, 1711–1716.
G. Gamba, Physiol. Rev. 2005, 85, 423–516.
K. Kahle, J. Rinehart, A. Ring, I. Gimenez, G. Gamba, S. Hebert, R. Lifton, Physiology 2006, 21, 326–361.
A. Alizadeh Naderi, R. Reilly, Nature Rev. Nephrology 2010, 6, 657–722.
I. Forster, N. Hernando, J. Biber, H. Murer, Kidney Int. 2006, 70, 1548–1607.
W. Boron, J. Am. Soc. Nephrol.: JASN 2006, 17, 2368–2450.
I. Choi, H. Soo Yang, W. Boron, J. Physiol. 2007, 578, 131–173.
S. Hebert, D. Mount, G. Gamba, Pflügers Arch.: Eur. J. Physiol. 2004, 447, 580–673.
K. Hinchcliff, P. Morley, A. Guthrie, J. Am. Vet. Med. Assoc. 2009, 235, 76–158.
J. Kyte, J. Biol. Chem. 1971, 246, 4157–4222.
E. Cayanis, H. Bayley, I. Edelman, J. Biol. Chem. 1990, 265, 10829–10864.
K. Geering, FEBS Lett. 1991, 285, 189–282.
K. Geering, J. Kraehenbuhl, B. Rossier, J. Cell Biol. 1987, 105, 2613–2622.
G. Crambert, K. Geering, Science’s STKE: Signal Transduction Knowledge Environment 2003, 2003.
K. McGrail, J. Phillips, K. Sweadner, J. Neurosci. 1991, 11, 381–472.
P. Bøttger, Z. Tracz, A. Heuck, P. Nissen, M. Romero-Ramos, K. Lykke-Hartmann, J. Compar. Neurol. 2011, 519, 376-780.
P. Lucchesi, K. Sweadner, J. Biol. Chem. 1991, 266, 9327–9358.
J. F. Hoffman, Proc. Nat. Acad. Sci. USA 2002, 99.
J. Hlivko, S. Chakraborty, T. Hlivko, A. Sengupta, P. James, Mol. Reprod. Devel. 2006, 73, 101–116.
A. Woo, P. James, J. Lingrel, J. Membr. Biol. 1999, 169, 39–83.
G. Blanco, Seminars in Nephrology 2005, 25, 292–595.
P. L. Pedersen, E. Carafoli, Trends Biochem. Sci., 1987 , 12, 146–296.
M. Palmgren, P. Nissen, Annu. Rev. Biophys. 2011, 40, 243–309.
O. Vagin, L. Dada, E. Tokhtaeva, G. Sachs, Am. J. physiol. Cell Physiol. 2012.
R. Albers, Annu. Rev. Biochem. 1967, 36, 727–783.
R. Post, S. Kume, T. Tobin, B. Orcutt, A. Sen, J. Gen. Physiol. 1969, 54, 306–332.
J. Morth, B. Pedersen, M. Toustrup-Jensen, T. Sørensen, J. Petersen, J. Andersen, B. Vilsen, P. Nissen, Nature 2007, 450, 1043–1052.
T. Shinoda, H. Ogawa, F. Cornelius, C. Toyoshima, Nature 2009, 459, 446–496.
C. Olesen, M. Picard, A.-M. L. Winther, C. Gyrup, J. Morth, C. Oxvig, J. Møller, P. Nissen, Nature 2007, 450, 1036–1078.
C. Olesen, T. Sørensen, R. Nielsen, J. Møller, P. Nissen, Science 2004, 306, 2251–2256.
T. Sørensen, J. Clausen, A.-M. L. Jensen, B. Vilsen, J. Møller, J. Andersen, P. Nissen, J. Biol. Chem. 2004, 279, 46355–46363.
C. Toyoshima, T. Mizutani, Nature 2004, 430, 529–564.
C. Toyoshima, M. Nakasako, H. Nomura, H. Ogawa, Nature 2000, 405, 647–702.
C. Toyoshima, H. Nomura, Nature 2002, 418, 605–616.
C. Toyoshima, H. Nomura, T. Tsuda, Nature 2004, 432, 361–369.
R. Rakowski, D. Gadsby, P. De Weer, J. Gen. Physiol. 1989, 93, 903–944.
S. Despa, J. Bossuyt, F. Han, K. Ginsburg, L.-G. Jia, H. Kutchai, A. Tucker, D. Bers, Circulat. Res. 2005, 97, 252–261.
C. Palmer, B. Scott, L. Jones, J. Biol. Chem. 1991, 266, 11126–11156.
H. Poulsen, P. Morth, J. Egebjerg, P. Nissen, FEBS Lett. 2010, 584, 2589–2684.
Z.-Q. Wu, J. Chen, Z.-Q. Chi, J.-G. Liu, Mol. Pharmacol. 2007, 71, 519–549.
H. Rasmussen, E. Hamilton, C.-C. Liu, G. Figtree, Trends Cardiovasc. Med. 2010, 20, 85–175.
S. Bibert, C.-C. Liu, G. Figtree, A. Garcia, E. Hamilton, F. Marassi, K. Sweadner, F. Cornelius, K. Geering, H. Rasmussen, J. Biol. Chem. 2011, 286, 18562–18634.
D. Alves, G. Farr, P. Seo-Mayer, M. Caplan, Molecul. Biol. Cell 2010, 21, 4400–4408.
H. Blom, D. Rönnlund, L. Scott, Z. Spicarova, J. Widengren, A. Bondar, A. Aperia, H. Brismar, BMC Neuroscience 2011, 12, 16.
H. Shimizu, E. Watanabe, T. Hiyama, A. Nagakura, A. Fujikawa, H. Okado, Y. Yanagawa, K. Obata, M. Noda, Neuron 2007, 54, 59–131.
S. Santos, B. Manadas, C. Duarte, A. Carvalho, J. Proteome Res. 2010, 9, 1670–1752.
D. Zhang, Q. Hou, M. Wang, A. Lin, L. Jarzylo, A. Navis, A. Raissi, F. Liu, H.-Y. Man, J. Neuroscience 2009, 29, 4498–5009.
J. Heiny, V. Kravtsova, F. Mandel, T. Radzyukevich, B. Benziane, A. Prokofiev, S. Pedersen, A. Chibalin, I. Krivoi, J. Biol. Chem. 2010, 285, 28614–28640.
M. Doi, K. Iwasaki, Mol. Cell. Neurosci. 2008, 38, 548–606.
B. Cassels, J. Ethnopharmacol. 1985, 14, 273–354.
D. Watt, J. Simard, P. Mancuso, Comp. Biochem. Physiol. A, Comp. Physiol. 1982, 71, 375–457.
S. Zhan, C. Merlin, J. Boore, S. Reppert, Cell 2011, 147, 1171–1256.
E. Labeyrie, S. Dobler, Mol. Biol. Evolut. 2004, 21, 218–239.
Z. Li, Z. Xie, Pflügers Arch.: Eur. J. Physiol. 2009, 457, 635–679.
D. Hilgemann, Proc. Nat. Acad. Sci. USA 2003, 100, 386–394.
M. De Fusco, R. Marconi, L. Silvestri, L. Atorino, L. Rampoldi, L. Morgante, A. Ballabio, P. Aridon, G. Casari, Nature Genetics 2003, 33, 192–198.
P. de Carvalho Aguiar, K. Sweadner, J. Penniston, J. Zaremba, L. Liu, M. Caton, G. Linazasoro, M. Borg, M. Tijssen, S. Bressman, W. Dobyns, A. Brashear, L. Ozelius, Neuron 2004, 43, 169–244.
P. Bøttger, C. Doğanlı, K. Lykke-Hartmann, Neurosci. Biobehav. Rev. 2012, 36, 855–926.
K. Axelsen, M. Palmgren, J. Mol. Evolut. 1998, 46, 84–185.
H. Poulsen, H. Khandelia, J. Morth, M. Bublitz, O. Mouritsen, J. Egebjerg, P. Nissen, Nature 2010, 467, 99–201.
Acknowledgments
We are grateful to Poul Nissen for advice and support. MJC and HP were funded by the Danish National Research Center PUMPKIN and HP by The Lundbeck Foundation, The Carlsberg Foundation, and L’Oréal/UNESCO.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Clausen, M.J.V., Poulsen, H. (2013). Sodium/Potassium Homeostasis in the Cell. In: Banci, L. (eds) Metallomics and the Cell. Metal Ions in Life Sciences, vol 12. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-5561-1_3
Download citation
DOI: https://doi.org/10.1007/978-94-007-5561-1_3
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-007-5560-4
Online ISBN: 978-94-007-5561-1
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)