Abstract
The paper highlights the need of dimension reduction of voluminous gene expression microarray data for developing a robust classifier to predict patients with cancerous genes. The proposed algorithm builds a fuzzy rule based classifier with optimized rule set without much sacrificing classification accuracy. The gene expression matrix is first discretized using linguistic values. The importance factor of each gene is then evaluated representing the degree of presence of a unique linguistic value of the gene both in disease and nondisease classes. Initial fuzzy rule base consists higher ranking genes and gradually other genes are included in the rule base in order to achieve maximum classification accuracy. Thus optimum rule set is built with important genes for classification of test data set. The methodology proposed here has been successfully demonstrated for the lung cancer classification problem, which includes 97 smokers with lung cancer and 90 without lung cancer gene expression data. The results are promising even though maximum number of genes are removed from the original data.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Machine learning and data mining techniques have been successfully applied [1–4] for long in biomedical data analysis to extract knowledge [5–7]. Dimensionality reduction is very much relevant in bio-informatics research, particularly in the context of microarray data, characterized by relatively few samples in a high-dimensional gene (feature) space. Irrelevant genes (features) lead to insufficient classification accuracy and add extra difficulties in finding potentially useful knowledge [8, 9]. Gene selection becoming one of the main sub-fields in bio-informatics data mining [10–12]. In the context of classification, the main goal of gene selection is to search for an optimal gene subset that lead to improved classification performance. During the past decades, extensive research has been conducted from multidisciplinary fields including statistics, pattern recognition, machine learning and data mining [10, 13, 14].
Continuous valued attributes generate large rule set and most classifiers with such huge number of rules are unstable even for slight change of training data set. Fuzzy rule based systems have been successfully applied to various application areas such as control, decision making, classification and many more [15, 16]. Some of the machine learning based works which generate fuzzy if-then rules are demonstrated in [17–19]. A version of fuzzy-ID3 algorithm which induces fuzzy decision trees is proposed in [17]. While the main objective in designing fuzzy rule-based systems is maximization of performance, comprehensibility of the system has also been taken into account in some recent studies [20–26]. The comprehensibility of fuzzy rule-based systems is related to linguistic interpretability of each fuzzy set in the rule, separation of neighboring fuzzy sets and the number of fuzzy sets for each linguistic variable. Simplicity of fuzzy rule based systems (e.g. the number of input variables, the number of fuzzy if-then rules) and fuzzy reasoning power are the other important factors that determine comprehensibility of the system.
In this paper, fuzzy if-then rules are optimized using selected genes to design a robust comprehensible fuzzy rule base for gene pattern classification problem with continuous attribute values. In the proposed method, first continuous value of microarray gene expression data are discretized using linguistic values determined by the variance of each sample corresponding to each gene. The linguistic values are relevant for discriminating alternative phenotypes and represent activation level of the gene. With the linguistic value, gene expression matrix is rebuilt. As a next step, fuzzy logic has been applied on the linguistic gene expression matrix to evaluate importance of each gene. Based on the importance factor, genes are ranked representing their significance in sample classification. Finally, fuzzy rules are framed with higher ranking genes and gradually next higher ranking genes are included in the fuzzy rule base with an objective to achieve maximum classification accuracy. Thus optimized rule set is obtained to classify patients with cancerous genes using test microarray gene expression data set.
The rest of the paper is organized as follows. The proposed method is described in Sect. 2. Experimental results are presented in Sect. 3. Concluding remarks and suggestions for future work are given in Sect. 4.
2 Methodology
Classification is a supervised learning technique that aims at exploring proper class of given objects based on the similarity among the objects. In disease classification problem, it has been noticed that although there are thousands of genes for each observation, a few underlying genes may account for much of the data variation. For instance, many of the genes may not be relevant to the tumour metabolic process, so they are potentially noise features. Removing such noise features may help to obtain higher classification accuracy (better diagnosis), resulting identification of marker genes. Selected genes increase classification accuracy with less number of comparisons which is ultimate goal of gene research.
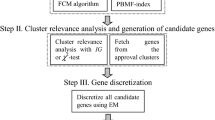
2.1 Optimized Fuzzy Rule Generation (OFRG) Algorithm
Feature selection process refers to choosing a subset of attributes from the set of original attributes. The purpose of feature selection is to identify the significant features, eliminate the irrelevant features and build a robust learning model.
The proposed gene selection algorithm evaluates fuzzy importance factor of each gene that signifies relevance of the gene in classifying the diseased patients using microarray gene expression data.
2.2 Terminologies for the Proposed OFRG Algorithm
Linguistic Gene Expression \( \left( {LGE_{(i,j)} } \right) \): To obtain linguistic gene expression value, first mean of each gene, say i for all samples are calculated, called gean_mean (i) . Difference between the \( j{\text{th}} \) sample value of \( i{\text{th}} \) gene and mean of the sample values of the corresponding gene (gean_mean (i)) as defined in (1) is used to assign the linguistic value at (\( LGE_{{(i,j){\text{th}}}} \)) position of the matrix. Thus the discretized gene expression matrix is constructed with the Linguistic gene expression value, which represent activity level of gene i for sample j.
Gene Importance Factor (i): Determines maximum number of unique Linguistic Gene Expression value for \( i{\text{th}} \) gene appear in the disease classes.
Unconstructive Impact Factor (i): Determines number of unique Linguistic Gene Expression value present in gean (i) for non-cancer classes. The Unconstructive Important Factor of gean (i) measures its contributions towards calculating the Gene Importance Factor, which leads to misclassification of data.
Fuzzy Importance Factor (FIF (i)): Distribution of gene expression data in two-dimension space reveals the fact that several genes are highly important compare to others in context of disease classification. Maximum number of samples with same Linguistic Gene Expression value in \( i{\text{th}} \) gene is used to evaluate importance of that gene. However, for gene (i) same linguistic value may appear in different non-cancer classes too (Unconstructive Impact Factor) which affects classification accuracy. Therefore, Fuzzy Importance Factor is proposed to isolate the Unconstructive Impact Factor from Gene Importance Factor using relation (2).
where k, m and n represent Gene Importance Factor, Unconstructive Impact Factor of \( i{\text{th}} \) gene and total number of samples in the gene expression matrix respectively. FIF (i) is zero when \( i{\text{th}} \) Gene Importance Factor is less/equal to the Unconstructive Impact Factor signifying that if Unconstructive Impact Factor is higher than the Gene Impact Factor then that gene has no effect to identify the proper class.

As the first step of the proposed method linguistic gene expression values are generated by applying Algorithm (1). Five different linguistic values are assigned to the samples of each gene by evaluating Eq. (1). Algorithm (2) is applied on discretized gene expression matrix to calculate fuzzy importance factor of each gene. Genes are clustered based on their similarity determined by Fuzzy Importance Factor. Finally, optimized fuzzy rule base is built with linguistic value of genes having maximum fuzzy importance factor as described in Algorithm 3.
3 Results and Comparisons
3.1 Data Set
Genes selected from microarray data sets of smokers with lung cancer and without lung cancer are compared. Database record series GSE4115 [27, 28] consisting of 22215 genes and each gene having 192 samples are considered for validating the scheme using Affymetrix genechip U133A.
3.2 Performance Measurements
OFRG algorithm has been applied on microarray data set to classify the samples by developing a fuzzy rule base classifier. Initial fuzzy rule base is built based on highest fuzzy importance factor and corresponding linguistic value are assigned for a specific disease class. Subsequently genes are included in the fuzzy rule base depending on their importance with an objective to improve classification accuracy.


A ten-fold cross-validation strategy is used by weka [29] classifier where BayesNet applied to measure the generalization accuracy of the Correlation feature selection (CFS) algorithm, shown in Table 1.
CFS [30] algorithm couples evaluation formula with an appropriate correlation measure and a heuristic search strategy. CFS was evaluated by experiments on artificial and natural data sets using three machine learning algorithms: a decision tree learner, an instance based learner, and Naive Bayes. Experiments on artificial data sets show that CFS quickly identifies and screens irrelevant, redundant and noisy features, and selects relevant features as long as their relevance does not strongly depend on other features. On natural domains, CFS typically eliminates well over half the features.
The OFRG algorithm extracts 4 (four) genes out of 22215 using GSE4115 [27, 28] data set showing more promising result than CFS algorithm (select 23 genes). Classification percentage comparison of CFS and OFRG algorithms are shown in Table 1 and Fig. 1, respectively. McNemar’s test using Chi squared and corrected for discontinuity comparison are shown in Table 2.
4 Conclusions
Dimension reduction is one of the main issues in microarray data classification and appropriate gene selection demonstrates promising outcome that enhance knowledge discovery and model interpretation.
Since biologists are often interested in identifying a collection of genes involved in a biological function or a pathway rather than individual genes, there has been considerable interest in recent years to develop statistical methods for identifying significant set of genes. This paper exploits structural information and proposes a three stage strategy for selecting significant set of genes and classifying disease and normal experimental conditions. Proposed OFRG algorithm selects optimized fuzzy rule base using maximum fuzzy importance factor of genes for disease sample which helps to identify samples in proper classes with less number of comparison. Moreover, variation of number of samples does not affect the classification accuracy of OFRG algorithm while it reduces in case of CFS algorithm.
References
Kononenko, I.: Inductive and bayesian learning in medical diagnosis. Appl. Artif. Intell. 7(4), 317–337 (1993)
Wolberg, W., Street, W.: Mangasarian ol. machine learning techniques to diagnose breast cancer from fine-needle aspirates. Cancer Lett. 77, 163–171 (1994)
Wolberg, W., Street, W.: Mangasarian ol. image analysis and machine learning applied to breast cancer diagnosis and prognosis. Anal. Quant. Cytol. Histol. 17(2), 77–87 (1995)
Kurgan, L., Cios, K., Tadeusiewicz, R., Ogiela, M., Goodenday, L.S.: Knowledge discovery approach to automated cardiac spect diagnosis. Artif. Intell. Med. 23(2), 149–169 (2001)
Antoniadis, A., Lambert-Lacroix, S., Leblanc, F.: Effective dimension reduction methods for tumor classification using gene expression data. Bioinformatics 19(5), 563–570 (2003)
Guyon, I., Weston, J., Barnhill, S., Vapnik, V.: Gene selection for cancer classification using support vector machines. Mach. Learn. 46, 389–422 (2002)
Yu, J., Ongarello, S., Fiedler, R., Chen, X., Toffolo, G., Cobelli, C., et al.: Ovarian cancer identification based on dimensionality reduction for high-throughput mass spectrometry data. Bioinformatics 21, 2200–2209 (2005)
Oh, I., Lee, J., Moon, B.: Hybrid genetic algorithms for feature selection. IEEE Trans. Pattern Anal. Mach. Intell. 26(11), 1424–1437 (2004)
Saeys, Y., Inza, I., Larranaga, P.: A review of feature selection techniques in bioinformatics. Bioinformatics 23(19), 2507–2517 (2007)
Liu, H., Motoda, H.: Feature extraction, construction and selection: a data mining perspective, 1st edn. Kluwer, Norwell (1998)
Conilione, P., Wang, D.: A comparative study on feature selection for E. coli promoter recognition. Int. J. Inf. Technol. 11, 54–66 (2005)
Degroeve, S., Baets, B., de Peer, Y., Rouzé, P.: Feature subset selection for splice site prediction. Bioinformatics 18(Suppl 2), 75–83 (2002)
Guyon, I., Elisseeff, A.: An introduction to variable and feature selection. J. Mach. Learn. Res. 3, 1157–1182 (2003)
Liu, H., Yu, L.: Toward integrated feature selection algorithms for classification and clustering. IEEE Trans. Knowl. Data Eng. 17(4), 491–502 (2005)
Kuncheva, L.: Fuzzy Classifier Design. Springer, Heidelberg (2000)
Leondes, C. (ed.): Fuzzy Theory Systems: Techniques and Applications, vol. 1–4. Academic Press, San Diego (1999)
Yuan, Y., Shaw, M.: Induction of fuzzy decision trees. Fuzzy Sets Syst. 25, 125–139 (1995)
Ichihashi, H., Shirai, T., Nagasaka, K., Miyoshi, T.: Neuro-fuzzy ID3: a method of inducing fuzzy decision trees with linear programming for maximizing entropy and an algebraic method for incremental learning. Fuzzy Sets Syst. 84, 1–19 (1996)
Yuan, Y., Zhuang, H.: A genetic algorithm for generating fuzzy classification rules. Fuzzy Sets Syst. 84, 1–19 (1996)
Castillo, L., Gonzalez, A., Perez, P.: Including a simplicity criterion in the selection of the best rule in a genetic fuzzy learning algorithm. Fuzzy Sets Syst. 120(2), 309–321 (2001)
Castro, J., Castro-Schez, J., Zurita, J.: Use of a fuzzy machine learning technique in the knowledge acquisition process. Fuzzy Sets Syst. 123(3), 307–320 (2001)
Jin, Y.: Fuzzy modeling of high-dimensional systems: complexity reduction and interpretability improvement. IEEE Trans. Fuzzy Syst. 8(2), 212–221 (2000)
de Oliveira, V.: Semantic constraints for membership function optimization. IEEE Trans. Syst. Man Cybern. Part A Syst. Hum. 29(1), 128–138 (1999)
Pedrycz, W., de Oliveira, V.: Optimization of fuzzy models. IEEE Trans. Systems Man Cybern. Part B Cybern. 26(4), 627–637 (1996)
Setnes, M., Babuska, R., Verbruggen, B.: Rule-based modeling: precision and transparency. IEEE Trans. Systems Man and Cybern. Part C Appl. Rev. 28(1), 165–169 (1998)
Setnes, M., Roubos, H.: GA-fuzzy based modeling and classification: complexity and performance. IEEE Trans. Fuzzy Syst. 8(5), 509–522 (2000)
Spira, A., Beane, J., Shah, V., Steiling, K., Liu, G., Schembri, F., Gilman, S., Dumas, Y., Calner, P., Sebastiani, P., Sridhar, S., Beamis, J., Lamb, C., Anderson, T., Gerry, N., Keane, J., Lenburg, M., Brody, J.: Airway epithelial gene expression in the diagnostic evaluation of smokers with suspect lung cancer. Nat. Med. 13(3), 361–366 (2007)
Gustafson, A., Soldi, R., Anderlind, C., Scholand, M., Qian, J., Zhang, X., Cooper, K., Walker, D., McWilliams, A., Liu, G., Szabo, E., Brody, J., Massion, P., Lenburg, M., Lam, S., Bild, A., Spira, A.: Airway PI3K pathway activation is an early and reversible event in lung cancer development. Sci. Transl. Med. 2(26), 26–25 (2010)
Hall, M., Frank, E., Holmes, G., Pfahringer, B., Reutemann, P., Witten, I.: The weka data mining software: an update. SIGKDD Explor. 11, 10–18 (2009)
Hall, M.: Correlation-based feature selection for machine learning. Thesis for the degree of Doctor of Philosophy (1999)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer India
About this paper
Cite this paper
Paul, A., Sil, J., Das Mukhopadhyay, C. (2014). Dimension Reduction of Gene Expression Data for Designing Optimized Rule Base Classifier. In: Biswas, G., Mukhopadhyay, S. (eds) Recent Advances in Information Technology. Advances in Intelligent Systems and Computing, vol 266. Springer, New Delhi. https://doi.org/10.1007/978-81-322-1856-2_15
Download citation
DOI: https://doi.org/10.1007/978-81-322-1856-2_15
Published:
Publisher Name: Springer, New Delhi
Print ISBN: 978-81-322-1855-5
Online ISBN: 978-81-322-1856-2
eBook Packages: EngineeringEngineering (R0)