Abstract
Drebrin is a major F-actin-binding protein in the brain. In the past two decades, many drebrin-binding proteins in addition to F-actin have been identified in several research fields including neuroscience, oncology, and immunology. Among the drebrin-binding proteins, there are various kinds of proteins including scaffold proteins, nuclear proteins, phosphatases, microtubule-binding proteins, G-actin-binding proteins, gap junction proteins, chemokine receptors, and cell-adhesion-related proteins. The interaction between drebrin and its binding partners seems to play important roles in higher brain functions, because drebrin is involved in the pathogenesis of some neurological diseases with cognitive defects. In this chapter, we will first review the interaction of Homer and spikar with drebrin, particularly focusing on spine morphogenesis and synaptic function. Homer contributes to spine morphogenesis by cooperating with shank and activated Cdc42 small GTPase, suggesting a novel signaling pathway comprising Homer, drebrin, shank, and Cdc42 for spine morphogenesis. Drebrin sequesters spikar in the cytoplasm and stabilizes it in dendritic spines, leading to spine formation. Finally, we will introduce some other drebrin-binding proteins including end-binding protein 3 (EB3), profilin, progranulin, and phosphatase and tensin homologue (PTEN). These proteins are involved in Alzheimer’s disease and cancer. Therefore, further studies on drebrin and its binding proteins will be of great importance to elucidate the pathologies of various diseases and may contribute to their medical treatment and diagnostics development.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Drebrin is a typical F-actin-binding protein in the dendritic spines in the brain of vertebrates. Additionally, flies and nematodes express a drebrin-like protein, although they contain a characteristic SH3 domain in the C-terminal region (Butkevich et al. 2015). They may be considered partly homologues to drebrin and mouse actin-binding protein 1 (mAbp1/SH3P7) (Lappalainen et al. 1998; Yamazaki et al. 2001). Therefore, the interaction between drebrin and its binding partners seems critical for higher brain functions such as learning and memory, which are observed in vertebrates. We elucidated various drebrin characters as an F-actin-binding protein between the 1980s and the 1990s (Shirao and Obata 1985; Ishikawa et al. 1994); however, there have only been a few reports about drebrin-binding proteins, even after the discovery that drebrin is a component of the actin cytoskeleton. In 1996, we found myosin and gelsolin in the drebrin-bound actin complex using co-immunoprecipitation analysis with anti-drebrin antibody (mAb M2F6) (Hayashi et al. 1996). Soon after, Mammoto et al. have found that the G-actin-binding protein, profilin , binds to drebrin (Mammoto et al. 1998). In the twenty-first century, the identification reports of novel drebrin-binding proteins including Homer and spikar rapidly increased (Fig. 14.1). Homer is involved in various brain functions including synaptic functions, neuronal development, neurological diseases, and behavior (for review, see Szumlinski et al. 2006; Foa and Gasperini 2009; Pouliquin and Dulhunty 2009; Luo et al. 2012). Spikar is a transcription cofactor that is also involved in dendritic spine formation. We identified spikar as a drebrin-binding protein (Yamazaki et al. 2014), which was registered in the DDBJ/EMBL/GenBank databases in 2001. In this chapter, we first focus on Homer and spikar and introduce their function in synapse formation. In the latter half, we introduce other drebrin-binding proteins reported in the past two decades and discuss the relationship and implications of their interactions regarding brain functions and disorders.
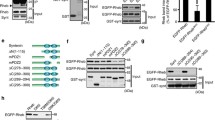
Schematic representation of drebrin and the binding sites of drebrin-binding proteins. Spikar binds to the ADF-H domain in the N-terminal region. Homer binds to the conserved sequence, PPXXF, in the C-terminal region. Profilin and afadin bind to the proline-rich region. CXCR4 binds to the N-terminal region (1–271). PTEN regulates the phosphorylation of Ser 659 of rat drebrin A (Ser 647 in the human homologue) by activity-dependent phosphatase activity. The binding sites on drebrin of EB3, connexin 43, progranulin, GAS8, SK1, and Arp3 are unknown. ADF-H ADF homology domain, AB1 Actin-binding region 1, AB2 Actin-binding region 2, In2 Ins 2 (drebrin A-specific sequence), P Proline-rich region
2 Homer
Homer protein family members are encoded by three genes (Homer1, 2 and 3), and many splicing variants have been identified (Shiraishi-Yamaguchi and Furuichi 2007). The subtypes are also called Vesl, PSD-Zip45, and Cupidin. Each Homer gene expresses long- and short-type Homers; the long types are Homer1b–d, Homer2a and b, and Homer3a and b, and the short types are Homer1a (Vesl-1s), Homer2c and d, and Homer3c and d. Short-type Homers lack self-assembly region, which contains coiled-coil domain and leucine zipper motif. Homer1a is a short-type protein that was originally identified as an immediate early gene induced by seizures in rat hippocampus (Kato et al. 1997; Brakeman et al. 1997). Homer1a protein was also isolated as a metabotropic glutamate receptor (mGluR) -binding protein during the same period (Brakeman et al. 1997). Since then, several binding partners of Homer have been identified. It has been elucidated that the Ena/VASP homology 1 (EVH1) domain of Homer recognizes a proline-rich consensus sequence (PPXXF or FPPPP) in several proteins including mGluR, Shank, inositol trisphosphate receptor (IP3R) , transient receptor potential canonical (TRPC) channels , dynamin3, and drebrin (Xiao et al. 1998; Tu et al. 1999, 1998; Yuan et al. 2003; Gray et al. 2003; Shiraishi et al. 2003). Homer forms a mesh-like matrix structure with shank, which causes spine enlargement and spine recruitment of several proteins including IP3R, postsynaptic density (PSD)-95, F-actin, guanylate kinase-associated protein (GKAP), and glutamate receptors (Sala et al. 2001; Hayashi et al. 2009).
The drebrin E and drebrin A isoforms, both of which have two PPXXF motifs in their C-terminal region, have been shown to bind to Homer2b (Shiraishi-Yamaguchi et al. 2009). Double mutation of the PPXXF motifs causes loss of the Homer2b-binding activity of drebrin, although it is not clear which motif is crucial for the drebrin-Homer2b binding (Shiraishi-Yamaguchi et al. 2009). The long subtype Homer proteins form an intermolecular disulfide bond between cysteine residues of C-terminal coiled-coil domain by oxidation, resulting in dimerization (Nepliouev et al. 2011). Disruption of the disulfide bond with a reducing reagent such as dithiothreitol (DTT) attenuates the binding of Homer and drebrin (Nepliouev et al. 2011). Another assay using Homer1a showed that either of the drebrin mutants, F543A or F621A, could disrupt the interaction between drebrin and Homer1a (Nepliouev et al. 2011), indicating that both of the PPXXF motifs of drebrin bind to Homer. Thus, it has been suggested that drebrin can bind to the long subtype Homer dimers more efficiently than to the short subtype Homer. Because the EVH1 domains of Homer are highly conserved between Homer family proteins, drebrin is supposed to be able to bind to Homer3 as well. Conversely, the brain-specific short-type drebrin isoform, s-drebrin A, contains actin-binding regions but no PPXXF motif (Jin et al. 2002), suggesting that s-drebrin A regulates the actin cytoskeleton in the spine without being implicated in a Homer-protein complex (Fig. 14.1).
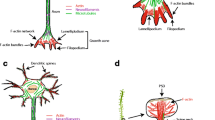
What is the physiological role of the interactions between drebrin and Homer? Although Homer can bind to several PSD proteins, it localizes not only at the PSD but also in the cytoplasmic area of dendritic spines (Shiraishi et al. 2004), where it seems to interact with IP3R on the endoplasmic reticulum (ER). Drebrin is concentrated in the cytoplasmic area of dendritic spines rather than in the PSD area (Kobayashi et al. 2007). Thus, drebrin may interact with Homer in the center of the spines. Drebrin attenuates the interaction between actin and myosin V (Ishikawa et al. 2007), and myosin Va, concentrated at the tip of the ER tubule, plays a pivotal role in the ER targeting. Therefore, it has been suggested that drebrin is involved in the localization of the Homer-IP3R-ER complex at the center of dendritic spines (Fig. 14.2).
Schematic diagram of presumable drebrin-complex and the signaling mechanism. Amyloid-beta recruits PTEN into dendritic spine, and then PTEN dephosphorylates drebrin. Ca2+ influx through NMDA receptor induces drebrin exodus that is the translocation of drebrin-F-actin complex from spine to dendrite (Mizui et al. 2014). In contrast, AMPA-receptor activity stabilizes drebrin within spines. Biou et al. (2008) suggested that drebrin remodels F-actin in the downstream of Ras signaling. Drebrin-EB3 complex plays a role as an adaptor between F-actin and microtubules
Drebrin is known to be involved in spine morphogenesis and formation (Takahashi et al. 2003; Ivanov et al. 2009). Homer2 interacts with activated Cdc42 , and a Homer2 mutant lacking the Cdc42-binding region impaired synapse formation and function (Shiraishi-Yamaguchi et al. 2009). Cdc42 and its regulators such as guanine nucleotide exchange factors (GEFs) play important roles in spine morphogenesis and formation (Moon and Zheng 2003; Chen et al. 2012). The Cdc42-binding region of Homer is separate from the EVH1 domain, which may enable drebrin association with the Homer-Cdc42 complex. These data raise the possibility of a novel signaling pathway for spine morphogenesis involving drebrin and activated small G proteins.
3 Spikar/ZMYND8
Spikar (also called KIAA1125, CTCL, tumor antigen Se14-3, ZMYND8, Prkcbp1, or RACK7) was first isolated as a drebrin-binding protein by a yeast two-hybrid screening of a rat brain library (Yamazaki et al. 2014). Previously, this gene was reported as a protein kinase C-binding protein (Fossey et al. 2000). Many splice variants of spikar are found in various database sites including NCBI. We have classified spikar into three major isoforms: spikar A, spikar B, and spikar delta-C (Fig. 14.3). Spikar A is similar to spikar B but lacks 20 residues in the N-terminal and 28 residues in the C-terminal region. Spikar delta-C lacks the C-terminal region of spikar A/B and contains a specific sequence in its C-terminal. There are no reports about the function of spikar delta-C so far.
Spikar contains a PHD domain, a bromodomain, a PWWP domain, a MYND domain, and a nuclear-receptor recognition motif (LSYLL) (Fig. 14.3). The PHD domain consists of two zinc fingers, and many PHD-containing proteins associate with chromatin and regulate its activities (Bienz 2006; Baker et al. 2008). The bromodomain associates with acetylated lysine residues found in the C-terminal region of histones (Sanchez and Zhou 2009). Bromodomain -containing proteins are associated with obesity, inflammation, cancer, and neurological disorders (Padmanabhan et al. 2016; Sanchez and Zhou 2009; Belkina and Denis 2012) and have become new therapeutic targets for these diseases. The nuclear-receptor recognition motif of spikar in the bromodomain binds to nuclear receptors including thyroid hormones and steroid receptors (Savkur and Burris 2004). The PWWP domain is named after a conserved Pro-Trp-Trp-Pro motif; however, the first three amino acids are not strictly conserved (Stec et al. 2000; Qin and Min 2014). The PWWP motif of spikar is Pro-Phe-Trp-Pro. The PWWP domain has a DNA- and histone-binding ability (Qiu et al. 2002; Wu et al. 2011). Like the PHD domain, the MYND domain contains a zinc finger motif, which can fold two zinc atoms (Lutterbach et al. 1998). The most studied MYND domain-containing proteins are the SMYD family proteins, which consist of SMYD1–5 and function as a lysine methyltransferase (Spellmon et al. 2015). Taken together, spikar is likely a nuclear protein.
BS69 is likely to be a spikar family protein, because it contains a PHD domain, a bromodomain, a PWWP domain, and a MYND domain in the same order within the molecule. BS69 is an adenovirus E1A-binding protein that interacts with the nuclear-receptor corepressor, N-CoR, and functions as a transcription corepressor (Masselink and Bernards 2000). However, we found that spikar functions as a coactivator of thyroid hormone receptor, glucocorticoid receptor, and estrogen receptor (Yamazaki et al. 2014). Interestingly, it has recently been reported that spikar acts as a transcription corepressor of histone H3 lysine 4 (H3K4) demethylase (JARID1D) (Li et al. 2016). Moreover, spikar is associated with the nucleosome remodeling and deacetylase (NuRD) complex in the nucleus, where it plays a role as a DNA damage response (DDR) factor, recruiting the NuRD complex to damaged chromatin, to repress transcription and promote double strand break repair by homologous recombination (Kloet et al. 2015; Gong et al. 2015; Adhikary et al. 2016).
The drebrin-binding region of spikar was assessed by a yeast two-hybrid assay and a GST pull-down assay, which showed that the N-terminal region of spikar interacts with the ADF-H domain of drebrin (Yamazaki et al. 2014). The ADF-H domain is found in all drebrin isoforms. In addition, the drebrin-binding region of spikar is found in spikar A, spikar B, and spikar delta-C. In the drebrin-binding region of spikar, there are nucleosome-associating domains, suggesting that drebrin may interfere with the association between spikar and the nucleosome.
Spikar is ubiquitously expressed in various tissues and concentrated in the cellular nuclei, although the expression level is higher in the spleen and thymus (Yamazaki et al. 2014). Spikar contains a nuclear localization signal (NLS, KKKKK) in the N-terminal region and is localized in the nucleus, but a spikar mutant (mNLS-spikar) with a mutated NLS (KTTKK) does not localize in the nucleus. In the rat brain, spikar is highly expressed in the cerebellum, although it is expressed throughout the brain (Yamazaki et al. 2014). In cultured astrocytes, however, spikar was hardly detected by western blotting. However, immunocytochemical studies and observation of GFP-spikar localization indicated that spikar is concentrated in the nucleus in HEK293 cells and cultured neurons (Yamazaki et al. 2014). Interestingly, in neurons, spikar is found in the cytoplasm in addition to the nucleus. Particularly, spikar is localized in dendritic spines , where drebrin is concentrated (Yamazaki et al. 2014). Furthermore, spikar tends to localize in drebrin-rich spines (Yamazaki et al. 2014). These findings suggest that spikar is localized in dendritic spines in a drebrin-dependent manner. In contrast, the localization of drebrin in dendritic spines was unaffected in spikar-knockdown neurons (Yamazaki et al. 2014). In addition to the postsynaptic localization, our subcellular fractionation data suggest that spikar is localized in presynapses associated with synaptic vesicles (Fig. 14.4).
Schematic diagram illustrating the anchoring of spikar in dendritic spines and its function in the nucleus. AMPA-receptor activity stabilizes drebrin in dendritic spines. Drebrin sequesters spikar in dendritic spines, and then spikar accumulates in drebrin-rich spines. Spikar function depends on drebrin. mNLS-spikar concentrates in dendritic spines, suggesting that spikar transports into the nucleus from dendritic spines with the importin complex. In the nucleus, spikar is involved in the regulation of gene transcription. Inset immunoblot image shows the subcellular distribution of spikar. The protein extract (20 μg protein) from each fraction was analyzed by western blotting with antibodies against spikar, drebrin, PSD-95, synaptophysin, and histone H3. The fractions were as follows: H homogenate, P1 cell nuclei and debris, P2 synaptosomal fraction, S2 non-synaptosomal fraction, S3 cytosolic fraction, P3 microsomal fraction, LP1 synaptosomal plasma-membrane fraction, LP2 synaptic vesicle fraction
Knockdown of spikar using shRNA resulted in decreased spine density without alteration of spine morphology (Yamazaki et al. 2014). We conducted rescue experiments using wild-type spikar and mNLS-spikar, which showed that not only wild-type spikar but also mNLS-spikar were able to recover the spine density (Yamazaki et al. 2014). Importantly, the fact that cytoplasmic spikar could rescue the spine density indicates that spikar plays multifunctional roles in both the nucleus and cytoplasm. In HEK293 cells, GFP-spikar was localized in the nucleus and rarely observed in the cytoplasm. In contrast, in HEK293 cells overexpressing drebrin, GFP-spikar tended to localize in the cytoplasm in addition to the nucleus (Yamazaki et al. 2014). Moreover, in drebrin-knockdown neurons, the spine localization of spikar was weak (Yamazaki et al. 2014). These observations indicate that the cytoplasmic localization of spikar, in particular the spine localization in neurons, depends on drebrin. Live-cell imaging of spikar-knockdown cultured neurons showed that spikar plays a role in spine maintenance and de novo spine sprouting (Yamazaki et al. 2014). Conversely, overexpression of mNLS-spikar increased the spine and filopodium number. Furthermore, drebrin knockdown abolished the effect of the mNLS-spikar overexpression. These data suggest that the localization and the function of spikar depend on drebrin (Yamazaki et al. 2014). Because drebrin accumulates in filopodia from an early stage of neuronal development (Takahashi et al. 2003), we propose that drebrin sequesters and stabilizes spikar in dendritic spines and contributes to spine formation and maintenance (Fig. 14.4). This idea is consistent with the fact that the spine localization of drebrin does not depend on spikar.
Several studies have shown a relationship between spikar and cancer. For example, spikar expression is upregulated in squamous cell cancer and cervical intraepithelial neoplasia (Bierkens et al. 2013). In colorectal cancer, a high mutation frequency was found in the spikar gene (Park et al. 2002). Additionally, spikar plays a role in promoting tumor angiogenesis (Kuroyanagi et al. 2014). Because drebrin is highly expressed in some cancer cells and is involved in tumor formation and invasion (Vaskova et al. 2011; Terakawa et al. 2013; Lin et al. 2014; Mizutani et al. 2014; Xu et al. 2015), the interaction between drebrin and spikar may be critical for cancer growth. Investigation of the role of the interaction between drebrin and spikar in cancer cells may provide new insight into the mechanism of cancer migration and growth.
4 EB3
EB3 was isolated as a molecule that binds to the C-terminal region of APCL, which is a brain-specific homologue of the tumor suppressor gene APC, by yeast two-hybrid screening. The name is based on its significant homology to the microtubule-binding protein, EB1 (Su et al. 1995). EB1 and EB3 are expressed ubiquitously in many tissues, although EB3 is particularly highly expressed in the brain and skeletal muscle (Nakagawa et al. 2000). EB3 and EB1 bind to the plus-tip of microtubules and regulate its dynamics (Straube and Merdes 2007). Drebrin was found in a protein complex that was pulled down with GST-EB3 from the growth cone cytosol but not with GST-EB1 (Geraldo et al. 2008). This indicates that drebrin specifically binds to EB3 but not to EB1. The binding site for drebrin is in the middle region of EB3, between the microtubule-biding region and the coiled-coil region. This middle region is absent from the EB1 sequence (Geraldo et al. 2008). In growth cones, drebrin interacts with EB3 at the tips of microtubules in the proximal region of the filopodia. Drebrin knockdown or overexpression of dominant negative EB3 causes an inhibition of neurite elongation. These findings indicate that the drebrin-EB3 interaction is critical for growth cone mobility by regulating the association between actin-filaments and microtubules.
Recently, we and others have shown that CDK5 phosphorylates drebrin at Ser 142 (pan drebrin) and Ser 342 (drebrin A) (Tanabe et al. 2014; Worth et al. 2013). Gordon-Weeks and colleagues proposed a model in which the phosphorylation at Ser 142 causes a conformational change of drebrin, consequently exposing the coiled-coil domain and EB3 binding region to the outside of the molecule (see Fig. 4.3 in Chap. 4) (Worth et al. 2013). It has been suggested that phosphorylated drebrin can more easily interact with EB3 than nonphosphorylated drebrin. Additionally, the phosphorylation-induced conformational change of drebrin enables bundling of two discrete actin-filaments. However, at present, this model needs to be verified by more studies.
While it has previously been believed that microtubules are not present in dendritic spines, imaging technology using fluorescent proteins and the RNAi technique have revealed that dynamic microtubules can transiently protrude into dendritic spines and regulate their development (Gu et al. 2008). The protrusion of microtubules into spines depends on Ca2+ influx through synaptic NMDA receptors (Merriam et al. 2013). Conversely, excessive Ca2+ influx induced by 50 μM glutamate removes EB3 from the growing microtubule plus-ends in dendrites (Kapitein et al. 2011). In addition, we have reported that the Ca2+ influx through NMDA receptors induced drebrin exodus from dendritic spines (Mizui et al. 2014). Therefore, drebrin may function as an adaptor that links the EB3 complex of microtubule plus-ends to the actin cytoskeleton in dendritic spines and enables a transient protrusion of microtubules into dendritic spines (Fig. 14.2). It is consistent with a recent report that drebrin knockdown decreased spine invasion of microtubules (Merriam et al. 2013).
The interaction of EB3 and drebrin has also been suggested in non-neuronal cells. Drebrin E is localized in the apical domain of columnar epithelial cells (Bazellières et al. 2012). Drebrin knockdown influences the apicobasal elongation during cell development, a phenotype that is observed in EB3-knockdown cells (Bazellières et al. 2012). The drebrin-EB3 complex regulates the cooperation between the actomyosin apical network and the apico-lateral microtubule network. Disruption of this interaction results in destabilization of the actin-based terminal web.
5 Profilin
Profilin is a classical actin-binding protein, which was isolated as an inhibitor molecule of actin polymerization in 1977 (Carlsson et al. 1977). Profilin sequesters G-actin, which results in inhibition of actin polymerization (Carlsson et al. 1977; Mockrin and Korn 1980). However, profilin also has an actin polymerization function (Tilney et al. 1983; Pring et al. 1992; Pantaloni and Carlier 1993). The interaction between the profilin I isoform and drebrin was first identified by affinity column chromatography with GST-profilin from rat brain cytosol (Mammoto et al. 1998) and was confirmed by a membrane overlay assay . Because profilin I can be purified with poly-proline agarose beads (Tanaka and Shibata 1985), it is likely to bind to the proline-rich region of drebrin (Fig. 14.1). Profilin II is specifically expressed in the brain (Honore et al. 1993), although both profilin I and profilin II are found in the brain. The interaction between profilin II and drebrin has not been reported, yet; however, drebrin is expected to bind to profilin II, because profilin II binds to several ligands (poly-proline, actin, phosphoinositides) that profilin I binds to (Lambrechts et al. 2000). Profilin II accumulates in dendritic spines after induction of long-term potentiation (LTP) in vitro and in vivo (Ackermann and Matus 2003). In addition, it has been indicated that emotional excitation such as fear conditioning drives profilin into dendritic spines in rat amygdala (Lamprecht et al. 2006). Similarly, drebrin immunoreactivity increased in the middle molecular layer of the hippocampus concomitantly with enhancement in F-actin content after in vivo LTP induction (Fukazawa et al. 2003). These observations suggest that the interaction between drebrin and profilin is enhanced after LTP induction.
6 Progranulin
Progranulin, also known as granulin-epithelin precursor, proepithelin, acrogranin, and GP88/PC-cell-derived growth factor, is a ubiquitously expressed, secreted protein throughout the body, which was originally identified as a growth factor-like molecule (Shoyab et al. 1990; Anakwe and Gerton 1990; Plowman et al. 1992; Bhandari et al. 1993; Zanocco-Marani et al. 1999; Zhu et al. 2002; Bateman and Bennett 2009). Progranulin is a multifunctional protein that is involved in several types of physiological phenomena including inflammation, cell proliferation, wound healing, development, and tumorigenesis (Konopka et al. 2014; Bateman and Bennett 2009). The interaction between progranulin and drebrin has recently been discovered in the course of cancer research. Drebrin was found in the eluted proteins of a pull-down assay with recombinant progranulin and protein extracts of 5637 bladder cancer cells (Xu et al. 2015). Drebrin modulated the progranulin-induced actin cytoskeleton remodeling and motility of cancer cells. Drebrin-depletion in UMUC-3 bladder cancer cells inhibited tumor formation in implanted mice in vivo (Xu et al. 2015).
The best-known important characteristic of progranulin in neurology is its implication in frontotemporal dementia (FTD) and Alzheimer’s disease (AD) . Mutations in the progranulin gene are found in familial FTD patients (Baker et al. 2006), and the mutations are risk factors for AD (Carecchio et al. 2009; Perry et al. 2013). Progranulin knockdown in cultured neurons results in a decrease in dendritic spine density and an increase in filopodium-like protrusion (Tapia et al. 2011). Interestingly, drebrin is decreased in the brain of patients with Alzheimer’s disease (Harigaya et al. 1996) and Down syndrome (Shim and Lubec 2002). In addition, the loss of drebrin function by an antisense oligonucleotide treatment caused a spine density decrease (Takahashi et al. 2006). Therefore, drebrin may be involved in a progranulin-related signal transduction pathway for spine formation and development.
7 PTEN
Phosphatase and tensin homologue (PTEN), which is a tumor suppressor, functions as a phosphatase for phosphatidylinositol (3,4,5)-trisphosphate (PIP3) and a protein phosphatase (Song et al. 2012). Drebrin was found in PTEN-immunoprecipitates from rat brain and liver by mass spectrometry analysis (Kreis et al. 2013). PTEN dephosphorylates drebrin at Ser 647 (Ser 659 in rat drebrin A) in an activity-dependent manner (Kreis et al. 2013). The Ser residue is conserved among chicken, human, rabbit, mouse, rat, and xenopus (Kreis et al. 2013). It has recently been reported that PTEN is essential for Alzheimer’s disease pathology (Knafo et al. 2016). As mentioned above, drebrin is decreased in Alzheimer’s disease. Furthermore, amyloid-β oligomer induced PTEN recruitment into dendritic spines (Knafo et al. 2016) and drebrin decreased in dendritic spines (Lacor et al. 2007; Ishizuka et al. 2014). Moreover, PTEN is known to be essential for NMDA-receptor-dependent long-term depression (Arendt et al. 2014). It has been reported that overactivation of NMDA receptor induces calpain-mediated proteolysis of drebrin (Chimura et al. 2015). Taken together, the dephosphorylation of drebrin at Ser 659 by PTEN may be involved in the pathology of Alzheimer’s disease, possibly through drebrin degradation.
8 Ras
Ras is a small G protein family, which consists of three major members: H-Ras, N-Ras, and K-Ras (Ellis et al. 1981; Shimizu et al. 1983). They were first identified as oncogenes, and its signal transduction pathways including extracellular and intracellular signals have been studied in detail (Malumbres and Barbacid 2003). In addition to tumor formation driven by Ras mutation, Ras proteins are engaged on several vital phenomena such as cell proliferation and differentiation (Feramisco et al. 1984; Hagag et al. 1986). In synapses, Ras is involved in neurotransmitter release and synaptic plasticity such as LTP and LTD (Brambilla et al. 1997; Jovanovic et al. 2000; Komiyama et al. 2002; Kim et al. 2003; Hou and Klann 2004; Schenk et al. 2005; Qin et al. 2005; Li et al. 2006; Rumbaugh et al. 2006; Banko et al. 2006). The activity of Ras is regulated by exchanging GTP binding (active form) or GDP binding (inactive form), and this conversion is mediated by their upstream regulators GAP/GEF (Shih et al. 1980; Calés et al. 1988; Adari et al. 1988; West et al. 1990; Crechet et al. 1990; Huang et al. 1990). Among a number of downstream signaling pathway of Ras, MAPK and PI3K cascades are assumed to be associated with synaptic function such as memory formation (Atkins et al. 1998; Blum et al. 1999; Chen et al. 2005; Peineau et al. 2007; Kim et al. 2011; Choi et al. 2014).
It is not clear whether drebrin forms a protein complex with Ras. However, Biou et al. (2008) have reported that Ras modulates drebrin function in dendritic spines . The overexpression of constitutively active Ras resulted in a similar phenotype as the drebrin overexpression in cultured neurons, and the effect on dendritic spine morphology was dependent on drebrin (Biou et al. 2008). Additionally, dominant negative Ras abolished the phenotype of drebrin overexpression (Biou et al. 2008). The downstream effectors of Ras play significant roles in spine growth and morphogenesis. For instance, PI3K regulates the activity of Rho family small G proteins, which are closely associated with dendritic spine morphogenesis (Sjolander et al. 1991; Rodriguez-Viciana et al. 1994; Nakayama et al. 2000; Penzes et al. 2001; Tashiro and Yuste 2008). Moreover, constitutively active Ras binds to Rac GEF Tiam1, and the interaction is essential for Rac1 activation (Yamauchi et al. 2005). Taken together, these findings suggest that Ras activation is involved in drebrin-mediated dendritic spine morphogenesis.
9 Other Drebrin-Binding Proteins
In addition to the drebrin-binding proteins described above, there are various other studies about drebrin-related proteins, which form protein-complexes with drebrin (Fig. 14.1). Rufy3, GAS8, SK1, Arp3, and myosin IIB were found in immunoprecipitated complexes with drebrin (Wei et al. 2014; Zhao et al. 2009; Yagoub et al. 2014; Li et al. 2011; Cheng et al. 2000); however, so far it is unknown whether drebrin directly binds to these proteins. Nonetheless, it is known that drebrin binds to connexin 43 (Butkevich et al. 2004), which is a well-known component of gap junctions in glial cells (Rash et al. 2000), and stabilizes gap junctions to the submembrane actin cytoskeleton (Butkevich et al. 2004). Afadin is a cell-adhesion-related protein, which binds to the poly-proline region of drebrin with its PR1-2 region (Rehm et al. 2013). CXCR4 , a chemokine receptor, binds to the N-terminal region (1–271) of drebrin (Perez-Martinez et al. 2010). In T lymphocytes, drebrin recruits CXCR4 to the immune synapses upon super-antigen stimulation (Perez-Martinez et al. 2010). The relationships between drebrin and connexin 43, afadin, and CXCR4 are reviewed in detail in other chapters of this book.
10 Conclusion
In this chapter, we reviewed drebrin-binding proteins with regard to neuronal functions. Identification of novel drebrin-binding proteins has increased across various disciplines in recent years, although the implications of their association with drebrin have not been fully determined. In some cases, such as connexin 43, drebrin acts like a stabilizer for binding partners on the actin cytoskeleton, and its knockdown leads to alteration of their subcellular localization. Based on these observations, drebrin may also function as a scaffold protein tethering its binding partners to the actin cytoskeleton in the cytoplasmic area of dendritic spines. We suppose that this interaction is regulated by synaptic activity. Because drebrin is involved in the mechanism of neurological diseases, further studies on the drebrin interactome will be of great importance to determining the pathologies of various diseases and may contribute to their medical treatment and diagnostics development.
References
Ackermann M, Matus A (2003) Activity-induced targeting of profilin and stabilization of dendritic spine morphology. Nat Neurosci 6:1194–1200
Adari H, Lowy DR, Willumsen BM, Der CJ, McCormick F (1988) Guanosine triphosphatase activating protein (GAP) interacts with the p21 ras effector binding domain. Science 240:518–521
Adhikary S, Sanyal S, Basu M, Sengupta I, Sen S, Srivastava DK, Roy S, Das C (2016) Selective recognition of H3.1K36 dimethylation/H4K16 acetylation facilitates the regulation of all-trans-retinoic acid (ATRA)-responsive genes by putative chromatin reader ZMYND8. J Biol Chem 291:2664–2681
Anakwe OO, Gerton GL (1990) Acrosome biogenesis begins during meiosis: evidence from the synthesis and distribution of an acrosomal glycoprotein, acrogranin, during guinea pig spermatogenesis. Biol Reprod 42:317–328
Arendt KL, Benoist M, Lario A, Draffin JE, Munoz M, Esteban JA (2014) PTEN counteracts PIP3 upregulation in spines during NMDA-receptor-dependent long-term depression. J Cell Sci 127:5253–5260
Atkins CM, Selcher JC, Petraitis JJ, Trzaskos JM, Sweatt JD (1998) The MAPK cascade is required for mammalian associative learning. Nat Neurosci 1:602–609
Baker M et al (2006) Mutations in progranulin cause tau-negative frontotemporal dementia linked to chromosome 17. Nature 442:916–919
Baker LA, Allis CD, Wang GG (2008) PHD fingers in human diseases: disorders arising from misinterpreting epigenetic marks. Mutat Res 647:3–12
Banko JL, Hou L, Poulin F, Sonenberg N, Klann E (2006) Regulation of eukaryotic initiation factor 4E by converging signaling pathways during metabotropic glutamate receptor-dependent long-term depression. J Neurosci 26:2167–2173
Bateman A, Bennett HP (2009) The granulin gene family: from cancer to dementia. Bioessays 31:1245–1254
Bazellières E, Massey-Harroche D, Barthelemy-Requin M, Richard F, Arsanto JP, Le Bivic A (2012) Apico-basal elongation requires a drebrin-E-EB3 complex in columnar human epithelial cells. J Cell Sci 125:919–931
Belkina AC, Denis GV (2012) BET domain co-regulators in obesity, inflammation and cancer. Nat Rev Cancer 12:465–477
Bhandari V, Giaid A, Bateman A (1993) The complementary deoxyribonucleic acid sequence, tissue distribution, and cellular localization of the rat granulin precursor. Endocrinology 133:2682–2689
Bienz M (2006) The PHD finger, a nuclear protein-interaction domain. Trends Biochem Sci 31:35–40
Bierkens M, Krijgsman O, Wilting SM, Bosch L, Jaspers A, Meijer GA, Meijer CJ, Snijders PJ, Ylstra B, Steenbergen RD (2013) Focal aberrations indicate EYA2 and hsa-miR-375 as oncogene and tumor suppressor in cervical carcinogenesis. Genes Chromosomes Cancer 52:56–68
Biou V, Brinkhaus H, Malenka RC, Matus A (2008) Interactions between drebrin and Ras regulate dendritic spine plasticity. Eur J Neurosci 27(11):2847–2859
Blum S, Moore AN, Adams F, Dash PK (1999) A mitogen-activated protein kinase cascade in the CA1/CA2 subfield of the dorsal hippocampus is essential for long-term spatial memory. J Neurosci 19:3535–3544
Brakeman PR, Lanahan AA, O'Brien R, Roche K, Barnes CA, Huganir RL, Worley PF (1997) Homer: a protein that selectively binds metabotropic glutamate receptors. Nature 386:284–288
Brambilla R, Gnesutta N, Minichiello L, White G, Roylance AJ, Herron CE, Ramsey M, Wolfer DP, Cestari V, Rossi-Arnaud C, Grant SG, Chapman PF, Lipp HP, Sturani E, Klein R (1997) A role for the Ras signalling pathway in synaptic transmission and long-term memory. Nature 390:281–286
Butkevich E, Hulsmann S, Wenzel D, Shirao T, Duden R, Majoul I (2004) Drebrin is a novel connexin-43 binding partner that links gap junctions to the submembrane cytoskeleton. Curr Biol 14:650–658
Butkevich E, Bodensiek K, Fakhri N, von Roden K, Schaap IA, Majoul I, Schmidt CF, Klopfenstein DR (2015) Drebrin-like protein DBN-1 is a sarcomere component that stabilizes actin filaments during muscle contraction. Nat Commun 6:7523
Calés C, Hancock JF, Marshall CJ, Hall A (1988) The cytoplasmic protein GAP is implicated as the target for regulation by the ras gene product. Nature 332:548–551
Carecchio M, Fenoglio C, De Riz M, Guidi I, Comi C, Cortini F, Venturelli E, Restelli I, Cantoni C, Bresolin N, Monaco F, Scarpini E, Galimberti D (2009) Progranulin plasma levels as potential biomarker for the identification of GRN deletion carriers. A case with atypical onset as clinical amnestic mild cognitive impairment converted to Alzheimer's disease. J Neurol Sci 287:291–293
Carlsson L, Nystrom LE, Sundkvist I, Markey F, Lindberg U (1977) Actin polymerizability is influenced by profilin, a low molecular weight protein in non-muscle cells. J Mol Biol 115:465–483
Chen X, Garelick MG, Wang H, Lil V, Athos J, Storm DR (2005) PI3 kinase signaling is required for retrieval and extinction of contextual memory. Nat Neurosci 8:925–931
Chen C, Wirth A, Ponimaskin E (2012) Cdc42: an important regulator of neuronal morphology. Int J Biochem Cell Biol 44:447–451
Cheng XT, Hayashi K, Shirao T (2000) Non-muscle myosin IIB-like immunoreactivity is present at the drebrin-binding cytoskeleton in neurons. Neurosci Res 36:167–173
Chimura T, Launey T, Yoshida N (2015) Calpain-mediated degradation of drebrin by excitotoxicity in vitro and in vivo. PLoS One 10:e0125119
Choi JH, Park P, Baek GC, Sim SE, Kang SJ, Lee Y, Ahn SH, Lim CS, Lee YS, Collingridge GL, Kaang BK (2014) Effects of PI3Kgamma overexpression in the hippocampus on synaptic plasticity and spatial learning. Mol Brain 7:78
Crechet JB, Poullet P, Mistou MY, Parmeggiani A, Camonis J, Boy-Marcotte E, Damak F, Jacquet M (1990) Enhancement of the GDP-GTP exchange of RAS proteins by the carboxyl-terminal domain of SCD25. Science 248:866–868
Ellis RW, Defeo D, Shih TY, Gonda MA, Young HA, Tsuchida N, Lowy DR, Scolnick EM (1981) The p21 src genes of Harvey and Kirsten sarcoma viruses originate from divergent members of a family of normal vertebrate genes. Nature 292:506–511
Feramisco JR, Gross M, Kamata T, Rosenberg M, Sweet RW (1984) Microinjection of the oncogene form of the human H-ras (T-24) protein results in rapid proliferation of quiescent cells. Cell 38:109–117
Foa L, Gasperini R (2009) Developmental roles for Homer: more than just a pretty scaffold. J Neurochem 108:1–10
Fossey SC, Kuroda S, Price JA, Pendleton JK, Freedman BI, Bowden DW (2000) Identification and characterization of PRKCBP1, a candidate RACK-like protein. Mamm Genome 11:919–925
Fukazawa Y, Saitoh Y, Ozawa F, Ohta Y, Mizuno K, Inokuchi K (2003) Hippocampal LTP is accompanied by enhanced F-actin content within the dendritic spine that is essential for late LTP maintenance in vivo. Neuron 38:447–460
Geraldo S, Khanzada UK, Parsons M, Chilton JK, Gordon-Weeks PR (2008) Targeting of the F-actin-binding protein drebrin by the microtubule plus-tip protein EB3 is required for neuritogenesis. Nat Cell Biol 10:1181–1189
Gong F, Chiu LY, Cox B, Aymard F, Clouaire T, Leung JW, Cammarata M, Perez M, Agarwal P, Brodbelt JS, Legube G, Miller KM (2015) Screen identifies bromodomain protein ZMYND8 in chromatin recognition of transcription-associated DNA damage that promotes homologous recombination. Genes Dev 29:197–211
Gray NW, Fourgeaud L, Huang B, Chen J, Cao H, Oswald BJ, Hemar A, McNiven MA (2003) Dynamin 3 is a component of the postsynapse, where it interacts with mGluR5 and Homer. Curr Biol 13:510–515
Gu J, Firestein BL, Zheng JQ (2008) Microtubules in dendritic spine development. J Neurosci 28:12120–12124
Hagag N, Halegoua S, Viola M (1986) Inhibition of growth factor-induced differentiation of PC12 cells by microinjection of antibody to ras p21. Nature 319:680–682
Harigaya Y, Shoji M, Shirao T, Hirai S (1996) Disappearance of actin-binding protein, drebrin, from hippocampal synapses in Alzheimer's disease. J Neurosci Res 43:87–92
Hayashi K, Ishikawa R, Ye LH, He XL, Takata K, Kohama K, Shirao T (1996) Modulatory role of drebrin on the cytoskeleton within dendritic spines in the rat cerebral cortex. J Neurosci 16:7161–7170
Hayashi MK, Tang C, Verpelli C, Narayanan R, Stearns MH, Xu RM, Li H, Sala C, Hayashi Y (2009) The postsynaptic density proteins Homer and Shank form a polymeric network structure. Cell 137:159–171
Honore B, Madsen P, Andersen AH, Leffers H (1993) Cloning and expression of a novel human profilin variant, profilin II. FEBS Lett 330:151–155
Hou L, Klann E (2004) Activation of the phosphoinositide 3-kinase-Akt-mammalian target of rapamycin signaling pathway is required for metabotropic glutamate receptor-dependent long-term depression. J Neurosci 24:6352–6361
Huang YK, Kung HF, Kamata T (1990) Purification of a factor capable of stimulating the guanine nucleotide exchange reaction of ras proteins and its effect on ras-related small molecular mass G proteins. Proc Natl Acad Sci U S A 87:8008–8012
Ishikawa R, Hayashi K, Shirao T, Xue Y, Takagi T, Sasaki Y, Kohama K (1994) Drebrin, a development-associated brain protein from rat embryo, causes the dissociation of tropomyosin from actin filaments. J Biol Chem 269:29928–29933
Ishikawa R, Katoh K, Takahashi A, Xie C, Oseki K, Watanabe M, Igarashi M, Nakamura A, Kohama K (2007) Drebrin attenuates the interaction between actin and myosin-V. Biochem Biophys Res Commun 359:398–401
Ishizuka Y, Shimizu H, Takagi E, Kato M, Yamagata H, Mikuni M, Shirao T (2014) Histone deacetylase mediates the decrease in drebrin cluster density induced by amyloid beta oligomers. Neurochem Int 76:114–121
Ivanov A, Esclapez M, Pellegrino C, Shirao T, Ferhat L (2009) Drebrin A regulates dendritic spine plasticity and synaptic function in mature cultured hippocampal neurons. J Cell Sci 122:524–534
Jin M, Tanaka S, Sekino Y, Ren Y, Yamazaki H, Kawai-Hirai R, Kojima N, Shirao T (2002) A novel, brain-specific mouse drebrin: cDNA cloning, chromosomal mapping, genomic structure, expression, and functional characterization. Genomics 79:686–692
Jovanovic JN, Czernik AJ, Fienberg AA, Greengard P, Sihra TS (2000) Synapsins as mediators of BDNF-enhanced neurotransmitter release. Nat Neurosci 3:323–329
Kapitein LC, Yau KW, Gouveia SM, van der Zwan WA, Wulf PS, Keijzer N, Demmers J, Jaworski J, Akhmanova A, Hoogenraad CC (2011) NMDA receptor activation suppresses microtubule growth and spine entry. J Neurosci 31(22):8194–8209
Kato A, Ozawa F, Saitoh Y, Hirai K, Inokuchi K (1997) Vesl, a gene encoding VASP/Ena family related protein, is upregulated during seizure, long-term potentiation and synaptogenesis. FEBS Lett 412:183–189
Kim JH, Lee HK, Takamiya K, Huganir RL (2003) The role of synaptic GTPase-activating protein in neuronal development and synaptic plasticity. J Neurosci 23:1119–1124
Kim JI et al (2011) PI3Kgamma is required for NMDA receptor-dependent long-term depression and behavioral flexibility. Nat Neurosci 14:1447–1454
Kloet SL, Baymaz HI, Makowski M, Groenewold V, Jansen PW, Berendsen M, Niazi H, Kops GJ, Vermeulen M (2015) Towards elucidating the stability, dynamics and architecture of the nucleosome remodeling and deacetylase complex by using quantitative interaction proteomics. FEBS J 282:1774–1785
Knafo S et al (2016) PTEN recruitment controls synaptic and cognitive function in Alzheimer’s models. Nat Neurosci 19:443–453
Kobayashi C, Aoki C, Kojima N, Yamazaki H, Shirao T (2007) Drebrin a content correlates with spine head size in the adult mouse cerebral cortex. J Comp Neurol 503:618–626
Komiyama NH, Watabe AM, Carlisle HJ, Porter K, Charlesworth P, Monti J, Strathdee DJ, O’Carroll CM, Martin SJ, Morris RG, O’Dell TJ, Grant SG (2002) SynGAP regulates ERK/MAPK signaling, synaptic plasticity, and learning in the complex with postsynaptic density 95 and NMDA receptor. J Neurosci 22:9721–9732
Konopka J, Richbourgh B, Liu C (2014) The role of PGRN in musculoskeletal development and disease. Front Biosci (Landmark Ed) 19:662–671
Kreis P, Hendricusdottir R, Kay L, Papageorgiou IE, van Diepen M, Mack T, Ryves J, Harwood A, Leslie NR, Kann O, Parsons M, Eickholt BJ (2013) Phosphorylation of the actin binding protein Drebrin at S647 is regulated by neuronal activity and PTEN. PLoS One 8:e71957
Kuroyanagi J, Shimada Y, Zhang B, Ariyoshi M, Umemoto N, Nishimura Y, Tanaka T (2014) Zinc finger MYND-type containing 8 promotes tumour angiogenesis via induction of vascular endothelial growth factor-A expression. FEBS Lett 588:3409–3416
Lacor PN, Buniel MC, Furlow PW, Clemente AS, Velasco PT, Wood M, Viola KL, Klein WL (2007) Abeta oligomer-induced aberrations in synapse composition, shape, and density provide a molecular basis for loss of connectivity in Alzheimer's disease. J Neurosci 27:796–807
Lambrechts A, Braun A, Jonckheere V, Aszodi A, Lanier LM, Robbens J, Van Colen I, Vandekerckhove J, Fassler R, Ampe C (2000) Profilin II is alternatively spliced, resulting in profilin isoforms that are differentially expressed and have distinct biochemical properties. Mol Cell Biol 20:8209–8219
Lamprecht R, Farb CR, Rodrigues SM, LeDoux JE (2006) Fear conditioning drives profilin into amygdala dendritic spines. Nat Neurosci 9:481–483
Lappalainen P, Kessels MM, Cope MJ, Drubin DG (1998) The ADF homology (ADF-H) domain: a highly exploited actin-binding module. Mol Biol Cell 9:1951–1959
Li S, Tian X, Hartley DM, Feig LA (2006) Distinct roles for Ras-guanine nucleotide-releasing factor 1 (Ras-GRF1) and Ras-GRF2 in the induction of long-term potentiation and long-term depression. J Neurosci 26:1721–1729
Li MW, Xiao X, Mruk DD, Lam YL, Lee WM, Lui WY, Bonanomi M, Silvestrini B, Cheng CY (2011) Actin-binding protein drebrin E is involved in junction dynamics during spermatogenesis. Spermatogenesis 1:123–136
Li N, Li Y, Lv J, Zheng X, Wen H, Shen H, Zhu G, Chen TY, Dhar SS, Kan PY, Wang Z, Shiekhattar R, Shi X, Lan F, Chen K, Li W, Li H, Lee MG (2016) ZMYND8 reads the dual histone mark H3K4me1-H3K14ac to antagonize the expression of metastasis-linked genes. Mol Cell 63:470–484
Lin Q, Tan HT, Lim TK, Khoo A, Lim KH, Chung MC (2014) iTRAQ analysis of colorectal cancer cell lines suggests Drebrin (DBN1) is overexpressed during liver metastasis. Proteomics 14:1434–1443
Luo P, Li X, Fei Z, Poon W (2012) Scaffold protein Homer 1: implications for neurological diseases. Neurochem Int 61:731–738
Lutterbach B, Sun D, Schuetz J, Hiebert SW (1998) The MYND motif is required for repression of basal transcription from the multidrug resistance 1 promoter by the t(8;21) fusion protein. Mol Cell Biol 18:3604–3611
Malumbres M, Barbacid M (2003) RAS oncogenes: the first 30 years. Nat Rev Cancer 3:459–465
Mammoto A, Sasaki T, Asakura T, Hotta I, Imamura H, Takahashi K, Matsuura Y, Shirao T, Takai Y (1998) Interactions of drebrin and gephyrin with profilin. Biochem Biophys Res Commun 243:86–89
Masselink H, Bernards R (2000) The adenovirus E1A binding protein BS69 is a corepressor of transcription through recruitment of N-CoR. Oncogene 19:1538–1546
Merriam EB, Millette M, Lumbard DC, Saengsawang W, Fothergill T, Hu X, Ferhat L, Dent EW (2013) Synaptic regulation of microtubule dynamics in dendritic spines by calcium, F-actin, and drebrin. J Neurosci 33:16471–16482
Mizui T, Sekino Y, Yamazaki H, Ishizuka Y, Takahashi H, Kojima N, Kojima M, Shirao T, Sato M (2014) Myosin II ATPase activity mediates the long-term potentiation-induced exodus of stable F-actin bound by drebrin A from dendritic spines. PLoS One 9(1), e85367
Mizutani Y, Iwamoto I, Kanoh H, Seishima M, Nagata K (2014) Expression of drebrin, an actin binding protein, in basal cell carcinoma, trichoblastoma and trichoepithelioma. Histol Histopathol 29:757–766
Mockrin SC, Korn ED (1980) Acanthamoeba profilin interacts with G-actin to increase the rate of exchange of actin-bound adenosine 5′-triphosphate. Biochemistry 19:5359–5362
Moon SY, Zheng Y (2003) Rho GTPase-activating proteins in cell regulation. Trends Cell Biol 13:13–22
Nakagawa H, Koyama K, Murata Y, Morito M, Akiyama T, Nakamura Y (2000) EB3, a novel member of the EB1 family preferentially expressed in the central nervous system, binds to a CNS-specific APC homologue. Oncogene 19:210–216
Nakayama AY, Harms MB, Luo L (2000) Small GTPases Rac and Rho in the maintenance of dendritic spines and branches in hippocampal pyramidal neurons. J Neurosci 20:5329–5338
Nepliouev I, Zhang ZS, Stiber JA (2011) Effect of oxidative stress on homer scaffolding proteins. PLoS One 6:e26128
Padmanabhan B, Mathur S, Manjula R, Tripathi S (2016) Bromodomain and extra-terminal (BET) family proteins: new therapeutic targets in major diseases. J Biosci 41:295–311
Pantaloni D, Carlier MF (1993) How profilin promotes actin filament assembly in the presence of thymosin beta 4. Cell 75:1007–1014
Park J, Betel D, Gryfe R, Michalickova K, Di Nicola N, Gallinger S, Hogue CW, Redston M (2002) Mutation profiling of mismatch repair-deficient colorectal cancers using an in silico genome scan to identify coding microsatellites. Cancer Res 62:1284–1288
Peineau S, Taghibiglou C, Bradley C, Wong TP, Liu L, Lu J, Lo E, Wu D, Saule E, Bouschet T, Matthews P, Isaac JT, Bortolotto ZA, Wang YT, Collingridge GL (2007) LTP inhibits LTD in the hippocampus via regulation of GSK3beta. Neuron 53:703–717
Penzes P, Johnson RC, Sattler R, Zhang X, Huganir RL, Kambampati V, Mains RE, Eipper BA (2001) The neuronal Rho-GEF Kalirin-7 interacts with PDZ domain-containing proteins and regulates dendritic morphogenesis. Neuron 29:229–242
Perez-Martinez M, Gordon-Alonso M, Cabrero JR, Barrero-Villar M, Rey M, Mittelbrunn M, Lamana A, Morlino G, Calabia C, Yamazaki H, Shirao T, Vazquez J, Gonzalez-Amaro R, Veiga E, Sanchez-Madrid F (2010) F-actin-binding protein drebrin regulates CXCR4 recruitment to the immune synapse. J Cell Sci 123:1160–1170
Perry DC, Lehmann M, Yokoyama JS, Karydas A, Lee JJ, Coppola G, Grinberg LT, Geschwind D, Seeley WW, Miller BL, Rosen H, Rabinovici G (2013) Progranulin mutations as risk factors for Alzheimer disease. JAMA Neurol 70:774–778
Plowman GD, Green JM, Neubauer MG, Buckley SD, McDonald VL, Todaro GJ, Shoyab M (1992) The epithelin precursor encodes two proteins with opposing activities on epithelial cell growth. J Biol Chem 267:13073–13078
Pouliquin P, Dulhunty AF (2009) Homer and the ryanodine receptor. Eur Biophys J 39:91–102
Pring M, Weber A, Bubb MR (1992) Profilin-actin complexes directly elongate actin filaments at the barbed end. Biochemistry 31:1827–1836
Qin Y, Zhu Y, Baumgart JP, Stornetta RL, Seidenman K, Mack V, van Aelst L, Julius Zhu J (2005) State-dependent Ras signaling and AMPA receptor trafficking. Genes Dev 19(17):2000–2015
Qin S, Min J (2014) Structure and function of the nucleosome-binding PWWP domain. Trends Biochem Sci 39:536–547
Qiu C, Sawada K, Zhang X, Cheng X (2002) The PWWP domain of mammalian DNA methyltransferase Dnmt3b defines a new family of DNA-binding folds. Nat Struct Biol 9:217–224
Rash JE, Staines WA, Yasumura T, Patel D, Furman CS, Stelmack GL, Nagy JI (2000) Immunogold evidence that neuronal gap junctions in adult rat brain and spinal cord contain connexin-36 but not connexin-32 or connexin-43. Proc Natl Acad Sci U S A 97:7573–7578
Rehm K, Panzer L, van Vliet V, Genot E, Linder S (2013) Drebrin preserves endothelial integrity by stabilizing nectin at adherens junctions. J Cell Sci 126:3756–3769
Rodriguez-Viciana P, Warne PH, Dhand R, Vanhaesebroeck B, Gout I, Fry MJ, Waterfield MD, Downward J (1994) Phosphatidylinositol-3-OH kinase as a direct target of Ras. Nature 370:527–532
Rumbaugh G, Adams JP, Kim JH, Huganir RL (2006) SynGAP regulates synaptic strength and mitogen-activated protein kinases in cultured neurons. Proc Natl Acad Sci U S A 103:4344–4351
Sala C, Piech V, Wilson NR, Passafaro M, Liu G, Sheng M (2001) Regulation of dendritic spine morphology and synaptic function by Shank and Homer. Neuron 31:115–130
Sanchez R, Zhou MM (2009) The role of human bromodomains in chromatin biology and gene transcription. Curr Opin Drug Discov Devel 12:659–665
Savkur RS, Burris TP (2004) The coactivator LXXLL nuclear receptor recognition motif. J Pept Res 63:207–212
Schenk U, Menna E, Kim T, Passafaro M, Chang S, De Camilli P, Matteoli M (2005) A novel pathway for presynaptic mitogen-activated kinase activation via AMPA receptors. J Neurosci 25:1654–1663
Shih TY, Papageorge AG, Stokes PE, Weeks MO, Scolnick EM (1980) Guanine nucleotide-binding and autophosphorylating activities associated with the p21src protein of Harvey murine sarcoma virus. Nature 287:686–691
Shim KS, Lubec G (2002) Drebrin, a dendritic spine protein, is manifold decreased in brains of patients with Alzheimer's disease and Down syndrome. Neurosci Lett 324:209–212
Shimizu K, Goldfarb M, Perucho M, Wigler M (1983) Isolation and preliminary characterization of the transforming gene of a human neuroblastoma cell line. Proc Natl Acad Sci U S A 80:383–387
Shiraishi Y, Mizutani A, Mikoshiba K, Furuichi T (2003) Coincidence in dendritic clustering and synaptic targeting of homer proteins and NMDA receptor complex proteins NR2B and PSD95 during development of cultured hippocampal neurons. Mol Cell Neurosci 22:188–201
Shiraishi Y, Mizutani A, Yuasa S, Mikoshiba K, Furuichi T (2004) Differential expression of Homer family proteins in the developing mouse brain. J Comp Neurol 473:582–599
Shiraishi-Yamaguchi Y, Furuichi T (2007) The Homer family proteins. Genome Biol 8:206
Shiraishi-Yamaguchi Y, Sato Y, Sakai R, Mizutani A, Knopfel T, Mori N, Mikoshiba K, Furuichi T (2009) Interaction of Cupidin/Homer2 with two actin cytoskeletal regulators, Cdc42 small GTPase and Drebrin, in dendritic spines. BMC Neurosci 10:25
Shirao T, Obata K (1985) Two acidic proteins associated with brain development in chick embryo. J Neurochem 44:1210–1216
Shoyab M, McDonald VL, Byles C, Todaro GJ, Plowman GD (1990) Epithelins 1 and 2: isolation and characterization of two cysteine-rich growth-modulating proteins. Proc Natl Acad Sci U S A 87:7912–7916
Sjolander A, Yamamoto K, Huber BE, Lapetina EG (1991) Association of p21ras with phosphatidylinositol 3-kinase. Proc Natl Acad Sci U S A 88:7908–7912
Song MS, Salmena L, Pandolfi PP (2012) The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol 13:283–296
Spellmon N, Holcomb J, Trescott L, Sirinupong N, Yang Z (2015) Structure and function of SET and MYND domain-containing proteins. Int J Mol Sci 16:1406–1428
Stec I, Nagl SB, van Ommen GJ, den Dunnen JT (2000) The PWWP domain: a potential protein-protein interaction domain in nuclear proteins influencing differentiation? FEBS Lett 473:1–5
Straube A, Merdes A (2007) EB3 regulates microtubule dynamics at the cell cortex and is required for myoblast elongation and fusion. Curr Biol 17:1318–1325
Su LK, Burrell M, Hill DE, Gyuris J, Brent R, Wiltshire R, Trent J, Vogelstein B, Kinzler KW (1995) APC binds to the novel protein EB1. Cancer Res 55:2972–2977
Szumlinski KK, Kalivas PW, Worley PF (2006) Homer proteins: implications for neuropsychiatric disorders. Curr Opin Neurobiol 16:251–257
Takahashi H, Sekino Y, Tanaka S, Mizui T, Kishi S, Shirao T (2003) Drebrin-dependent actin clustering in dendritic filopodia governs synaptic targeting of postsynaptic density-95 and dendritic spine morphogenesis. J Neurosci 23:6586–6595
Takahashi H, Mizui T, Shirao T (2006) Down-regulation of drebrin A expression suppresses synaptic targeting of NMDA receptors in developing hippocampal neurones. J Neurochem 97(Suppl 1):110–115
Tanabe K, Yamazaki H, Inaguma Y, Asada A, Kimura T, Takahashi J, Taoka M, Ohshima T, Furuichi T, Isobe T, Nagata K, Shirao T, Hisanaga S (2014) Phosphorylation of drebrin by cyclin-dependent kinase 5 and its role in neuronal migration. PLoS One 9:e92291
Tanaka M, Shibata H (1985) Poly(L-proline)-binding proteins from chick embryos are a profilin and a profilactin. Eur J Biochem 151:291–297
Tapia L, Milnerwood A, Guo A, Mills F, Yoshida E, Vasuta C, Mackenzie IR, Raymond L, Cynader M, Jia W, Bamji SX (2011) Progranulin deficiency decreases gross neural connectivity but enhances transmission at individual synapses. J Neurosci 31:11126–11132
Tashiro A, Yuste R (2008) Role of Rho GTPases in the morphogenesis and motility of dendritic spines. Methods Enzymol 439:285–302
Terakawa Y, Agnihotri S, Golbourn B, Nadi M, Sabha N, Smith CA, Croul SE, Rutka JT (2013) The role of drebrin in glioma migration and invasion. Exp Cell Res 319:517–528
Tilney LG, Bonder EM, Coluccio LM, Mooseker MS (1983) Actin from Thyone sperm assembles on only one end of an actin filament: a behavior regulated by profilin. J Cell Biol 97:112–124
Tu JC, Xiao B, Yuan JP, Lanahan AA, Leoffert K, Li M, Linden DJ, Worley PF (1998) Homer binds a novel proline-rich motif and links group 1 metabotropic glutamate receptors with IP3 receptors. Neuron 21:717–726
Tu JC, Xiao B, Naisbitt S, Yuan JP, Petralia RS, Brakeman P, Doan A, Aakalu VK, Lanahan AA, Sheng M, Worley PF (1999) Coupling of mGluR/Homer and PSD-95 complexes by the Shank family of postsynaptic density proteins. Neuron 23:583–592
Vaskova M, Kovac M, Volna P, Angelisova P, Mejstrikova E, Zuna J, Brdicka T, Hrusak O (2011) High expression of cytoskeletal protein drebrin in TEL/AML1pos B-cell precursor acute lymphoblastic leukemia identified by a novel monoclonal antibody. Leuk Res 35:1111–1113
Wei Z, Sun M, Liu X, Zhang J, Jin Y (2014) Rufy3, a protein specifically expressed in neurons, interacts with actin-bundling protein Fascin to control the growth of axons. J Neurochem 130:678–692
West M, Kung HF, Kamata T (1990) A novel membrane factor stimulates guanine nucleotide exchange reaction of ras proteins. FEBS Lett 259:245–248
Worth DC, Daly CN, Geraldo S, Oozeer F, Gordon-Weeks PR (2013) Drebrin contains a cryptic F-actin-bundling activity regulated by Cdk5 phosphorylation. J Cell Biol 202:793–806
Wu H, Zeng H, Lam R, Tempel W, Amaya MF, Xu C, Dombrovski L, Qiu W, Wang Y, Min J (2011) Structural and histone binding ability characterizations of human PWWP domains. PLoS One 6:e18919
Xiao B, Tu JC, Petralia RS, Yuan JP, Doan A, Breder CD, Ruggiero A, Lanahan AA, Wenthold RJ, Worley PF (1998) Homer regulates the association of group 1 metabotropic glutamate receptors with multivalent complexes of homer-related, synaptic proteins. Neuron 21:707–716
Xu SQ, Buraschi S, Morcavallo A, Genua M, Shirao T, Peiper SC, Gomella LG, Birbe R, Belfiore A, Iozzo RV, Morrione A (2015) A novel role for drebrin in regulating progranulin bioactivity in bladder cancer. Oncotarget 6:10825–10839
Yagoub D, Wilkins MR, Lay AJ, Kaczorowski DC, Hatoum D, Bajan S, Hutvagner G, Lai JH, Wu W, Martiniello-Wilks R, Xia P, McGowan EM (2014) Sphingosine kinase 1 isoform-specific interactions in breast cancer. Mol Endocrinol 28:1899–1915
Yamauchi J, Miyamoto Y, Tanoue A, Shooter EM, Chan JR (2005) Ras activation of a Rac1 exchange factor, Tiam1, mediates neurotrophin-3-induced Schwann cell migration. Proc Natl Acad Sci U S A 102(41):14889–14894
Yamazaki H, Takahashi H, Aoki T, Shirao T (2001) Molecular cloning and dendritic localization of rat SH3P7. Eur J Neurosci 14:998–1008
Yamazaki H, Kojima N, Kato K, Hirose E, Iwasaki T, Mizui T, Takahashi H, Hanamura K, Roppongi RT, Koibuchi N, Sekino Y, Mori N, Shirao T (2014) Spikar, a novel drebrin-binding protein, regulates the formation and stabilization of dendritic spines. J Neurochem 128:507–522
Yuan JP, Kiselyov K, Shin DM, Chen J, Shcheynikov N, Kang SH, Dehoff MH, Schwarz MK, Seeburg PH, Muallem S, Worley PF (2003) Homer binds TRPC family channels and is required for gating of TRPC1 by IP3 receptors. Cell 114:777–789
Zanocco-Marani T, Bateman A, Romano G, Valentinis B, He ZH, Baserga R (1999) Biological activities and signaling pathways of the granulin/epithelin precursor. Cancer Res 59:5331–5340
Zhao L, Xu L, Zhou X, Zhu Q, Yang Z, Zhang C, Zhu X, Yu M, Zhang Y, Zhao X, Huang P (2009) Interaction of influenza virus NS1 protein with growth arrest-specific protein 8. Virol J 6:218
Zhu J, Nathan C, Jin W, Sim D, Ashcroft GS, Wahl SM, Lacomis L, Erdjument-Bromage H, Tempst P, Wright CD, Ding A (2002) Conversion of proepithelin to epithelins: roles of SLPI and elastase in host defense and wound repair. Cell 111:867–878
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Japan KK
About this chapter
Cite this chapter
Yamazaki, H., Shirao, T. (2017). Homer, Spikar, and Other Drebrin-Binding Proteins in the Brain. In: Shirao, T., Sekino, Y. (eds) Drebrin. Advances in Experimental Medicine and Biology, vol 1006. Springer, Tokyo. https://doi.org/10.1007/978-4-431-56550-5_14
Download citation
DOI: https://doi.org/10.1007/978-4-431-56550-5_14
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-56548-2
Online ISBN: 978-4-431-56550-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)