Abstract
The heat shock response is a fundamental mechanism to adapt against various proteotoxic stresses in all living organisms. This response is characterized by the induction of heat shock proteins (HSPs) and regulated mainly at the level of transcription by heat shock factor (HSF). Vertebrate cells possess four HSF genes, which are located in the conserved syntenic regions among species. The amino acid sequences of the DNA-binding domain (DBD) and oligomerization domain (HR-A/B) located in the N-terminal region are highly conserved. The DBD interacts with genomic DNA, and HR-A/B is required for the formation of an HSF trimer that binds to DNA with high affinity. The HR-A/B is flanked by two nuclear localization signals. There are some activation or regulatory domains in the C-terminal region. Among HSF family members, HSF1 is a master regulator of the expression of HSP gene in mammalian cells, while that of non-HSP genes is also regulated by HSF2, HSF3, and HSF4. Furthermore, the HSF family members cooperatively or competitively regulate the expression of some common targets.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Amino acid
- Consensus sequence
- DNA binding
- Evolution
- Hydrophobic heptad repeat
- Structure
- Transcriptional activity
1 Introduction
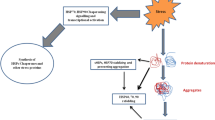
When cells are exposed to elevated temperatures, cellular proteins are denatured and aggregated. Cells have the ability to adapt to this proteotoxic stress by inducing the expression of heat shock proteins (HSPs) or molecular chaperones, which facilitate protein folding and suppress protein aggregation. This response is called the heat shock response and is a universal mechanism to maintain protein homeostasis and to protect cell from protein-damaging insults (Lindquist 1986). This response is well conserved from bacteria to human. In eukaryotes, the heat shock response is regulated mainly at the level of transcription by heat shock factor (HSF). HSF stays as an inert monomer in normal growth condition. In response to heat shock, HSF is quickly converted to a trimer that binds to the heat shock response element (HSE) located in the promoters of heat shock genes encoding HSPs including HSP70. The consensus sequence of the HSE is at least three inverted repeats of a pentanucleotide nGAAn (Fernandes et al. 1994). As a result, the expression of HSPs is rapidly induced.
HSF is required not only for the heat shock response but also cell growth and differentiation and normal lifespan in yeast, C. elegans, and Drosophila (Hsu et al. 2003; Morano et al. 1999; Morley and Morimoto 2004). There is a single HSF in these invertebrates, whereas four HSF genes (HSF1 to HSF4) exist in higher animals (Nakai et al. 1997; Nakai and Morimoto 1993; Rabindran et al. 1991; Schuetz et al. 1991). In mammals, HSF1 is required for the heat shock response, whereas HSF3 is required for this response in birds (McMillan et al. 1998; Tanabe et al. 1998). Both mammal HSF1 and chicken HSF3 are necessary for acquisition of the thermotolerance, which is correlated with the induction of HSPs. In addition, members of HSF family are involved in various developmental processes including gametogenesis and neurogenesis and maintenance of sensory organs (Chang et al. 2006; Kallio et al. 2002; Takaki et al. 2006; Wang et al. 2004). Thus, the HSF gene has duplicated and acquired multifaceted functions during evolution. In this review, we describe the structure of HSF family members; e.g., the DNA-binding domain, oligomerization domain, and activation domain. We also review the transcriptional activity, biochemical characters, and target sequences of HSF family members and discuss about evolution of the HSF gene family.
2 Structure of the HSF Family Members
The sizes and amino acid sequences of eukaryotic HSF family members are different, except for some conserved domains (Fig. 2.1) (Clos et al. 1993). Especially, amino acid sequences of the DNA-binding domain (DBD) and oligomerization domain (HR-A/B, hydrophobic heptad repeat-A/B) are highly conserved in eukaryotic species (Nakai et al. 1997; Nakai and Morimoto 1993; Sarge et al. 1991). In addition, there are several short sequences including the nuclear localization signal, which are conserved in all or parts of the HSF family members. The location and sequences of the transcriptional activation domain are different in the HSF family members.
Structures of the HSF family members. Diagrammatic representation of structures of the HSF family members. The percentage identities between amino acid sequences in regions of human HSF1 and those corresponding regions in other HSFs are established using the computer program GENETYX-WIN. The number of amino acids of each HSF is shown at the N-terminus. DBD DNA-binding domain; HR hydrophobic heptad repeat; DHR downstream of HR-C; h human; m mouse; c chicken; Dm Drosophila melanogaster; Ce C. elegans; Sc Saccharomyces cerevisiae. hHSF1 (an isoform hHSF1α) (Rabindran et al. 1991); hHSF2 (hHSF2α) (Schuetz et al. 1991); hHSF4 (hHSF4b) (Nakai et al. 1997); mHSF1 (mHSF1α) and mHSF2 (mHSF2α) (Sarge et al. 1993); mHSF3 (mHSF3a) (Fujimoto et al. 2010); mHSF4 (mHSF4b) (Tanabe et al. 1999); cHSF1, cHSF2, and cHSF3 (Nakai and Morimoto 1993); cHSF4 (cHSF4b) (Fujimoto et al. 2010); DmHSF (Clos et al. 1990); CeHSF1 (Swiss-P accession no Q9XW45); ScHSF (Wiederrecht et al. 1988). The HR-C is not conserved in HSF4, CeHSF1, and ScHSF
2.1 DNA-Binding Domain
The DBD is located near the N-terminus of all HSFs. Amino acid sequence of the DBD of human HSF1 (hHSF1) is 46 % identical to that of yeast HSF (ScHSF) (Fig. 2.1). The crystal structure and NMR solution structure of the DBD in ScHSF show that HSF belongs to winged helix-turn-helix DNA-binding proteins containing mammalian ETS and HNF-3/forkhead proteins (Gajiwala and Burley 2000) and consisted of three helical bundles (H1, H2, and H3) and four-stranded, antiparallel β-sheets (Fig. 2.2) (Damberger et al. 1995; Harrison et al. 1994; Vuister et al. 1994a, b). The central α-helix H3 interacts with the major groove of DNA. In contrast, the α-helix H1 and wing (loop) region are exposed outside and interact with regulatory factors such as RPA1 and ATF1 (Fujimoto et al. 2012; Takii et al. 2015). The wing is not necessary for the trimer formation of HSF but is required for full activity of HSF (Cicero et al. 2001; Littlefield and Nelson 1999).
Structure of the DNA-binding domain in human HSF1. (a) Sequences of the DBD in hHSF1. Boxes indicate α-helices and arrows indicate β-sheets. (b) A model to predict the structure of the DBD in hHSF1 by comparison with that in Drosophila HSF1 using SWISS-MODEL (http://swissmodel.expasy.org/). Each absolute structure is indicated by rainbow color. It shows a winged helix-turn-helix motif, which consisted of H2 (orange) and H3 (red). The α-helix H3 interacts with the major groove of DNA. The α-helix H1 and wing (loop) region are exposed outside
2.2 Oligomerization Domain
The HR-A/B is connected to the DBD by a flexible linker of 10–20 amino acids (Flick et al. 1994). It contains two α-helices, HR-A and HR-B, which consist of a repeating pattern of seven amino acids. Especially, amino acids at positions a and d of the heptad repeat, a-b-c-d-e-f-g, are hydrophobic amino acids such as leucine (L), isoleucine (I), and valine (V) (Fig. 2.3a). HSF uniquely forms a trimer that binds to DNA with a high affinity through the hydrophobic interaction between three HR-A/B domains located in parallel when HSF is activated (Clos et al. 1990; Peteranderl et al. 1999) (Fig. 2.3a). The electrostatic interactions also occur primarily between positions e and g, which are often charged residues such as arginine (R), lysine (K), aspartic acid (D), and glutamic acid (E) (Fig. 2.3b) (Creighton 1993). The HR-A/B is required for the formation of not only a trimer but also a dimer (Nakai et al. 1995; Sistonen et al. 1994). Therefore, the HR-A/B is also called the trimerization domain or oligomerization domain. The linker domain modulates the trimerization of HSF (Liu and Thiele 1996). There is another hydrophobic heptad repeat HR-C near the C-terminus of HSF. The HR-C inhibits the oligomerization of the HR-A/B by forming an intramolecular coiled coil with the HR-A/B and keeps HSF as a monomer that cannot bind to DNA in control conditions. Therefore, the point mutation of hydrophobic amino acids in the HR-C of hHSF1 results in the formation of a DNA-binding trimer (Rabindran et al. 1993; Zuo et al. 1995). Because HSF in yeasts and HSF4 lack the HR-C, they exist mostly as DNA-binding trimers in control conditions (Giardina et al. 1995; Jakobsen and Pelham 1988; Nakai et al. 1997; Sorger and Nelson 1989; Sorger and Pelham 1988; Wiederrecht et al. 1988). Another hydrophobic heptad repeat DHR exists downstream of the HR-C in most vertebrate HSFs (Nakai et al. 1997), but its function is unknown yet.
Trimer formation through interaction between the hydrophobic heptad repeats. (a) Amino acid sequences of the HR-A/B and HR-C in hHSF1. The open and solid squares indicate amino acids at positions a and d, respectively, in the repeating seven amino acids. Hydrophobic amino acids are marked by red and charged amino acids by green. (b). Trimer formation of the HR-A/B. The HR-A/B forms a trimer through the interaction between hydrophobic amino acids at positions a and d (solid lines) in the HR-A/B. The electrostatic interactions also occur primarily between positions e and g (dotted lines), which are often charged residues
2.3 Nuclear Localization Signal
The HR-A/B domains in the HSF family members are surrounded by two putative nuclear localization signals (NLSs), NLS1 and NLS2 (Fig. 2.4). These sequences fit the consensus for a bipartite NLS, which consisted of two clusters of basic amino acids, separated by a spacer of about ten amino acids (Sheldon and Kingston 1993). Human HSF2 (hHSF2) partly accumulates to the nucleus upon heat shock (Shinkawa et al. 2011), but a mutated hHSF2 lacking a cluster of the basic amino acids in the NLS1 or NLS2 does not translocate to the nucleus (Sheldon and Kingston 1993). Thus, both NLS1 and NLS2 are required for the nuclear translocation of HSF2. Human HSF1 (hHSF1) is a master regulator of the HSP expression during heat shock. The NLS2 of hHSF1 is required for the nuclear translocation, whereas the NLS1 is dispensable for that (Vujanac et al. 2005). Lysine amino acids in the NLSs are acetylated and related with the formation of nuclear stress bodies in response to heat shock (Raychaudhuri et al. 2014). Chicken HSF3 (cHSF3) is a master regulator of the HSP expression in birds, and the NLS2, but not the NLS1, is required and sufficient for the nuclear translocation of cHSF3 upon heat shock (Nakai and Ishikawa 2000). Drosophila HSF also has a bipartite NLS located downstream of the HR-A/B which is required for its nuclear translocation in response to heat shock and during development (Fang et al. 2001; Orosz et al. 1996). In plants, HSFs have a nuclear export signal (NES) in the C-terminal activation domain (Kotak et al. 2004), but there is no report about the NES in vertebrate HSFs.
In addition to the NLS, there are at least seven conserved sequences, termed sites a to g in the regions X and Y, in vertebrate HSF members (Fujimoto et al. 2010; Nakai et al. 1997; Nakai and Morimoto 1993; Tanabe et al. 1999). The function of these conserved sequences should be revealed in the future.
2.4 Transcriptional Activation Domain
HSF from a budding yeast Saccharomyces cerevisiae possesses the N-terminal and C-terminal transcriptional activation domains (Sorger 1990). In contrast, HSF from another budding yeast Kluyveromyces lactis does not have the N-terminal activation domain but has the C-terminal activation domain, whose sequence is completely divergent from that of S. cerevisiae HSF (Jakobsen and Pelham 1991). Thus, the sequences of transcriptional activation domains in HSFs are not evolutionally conserved. However, transcriptional activity of the activation domain is repressed in unstressed condition by a conserved motif CE2, which is located near the activation domain.
Mammalian HSF1 is functionally analogous to yeast HSF and robustly induces the expression of HSPs during heat shock. It has a potent transcriptional activation domain (AD) in the C-terminus, which is divided into two regions (Green et al. 1995; Shi et al. 1995; Zuo et al. 1995) (Fig. 2.5a). The AD1 (a.a. 372–431) contains the HR-C and is predicted to form an α-helix, while AD2 (a.a. 432–529) is rich in proline (13 %) and glycine (8 %). Hydrophobic residues in the AD1 and AD2 are involved in the elongation step of the transcription processes, and acidic residues are in the initiation steps (Brown et al. 1998). The regulatory domain (RD, a.a. 221–310) located downstream of the HR-A/B represses the two activation domains in unstressed condition and confers heat shock inducibility of the transcriptional activity upon heat shock (Green et al. 1995; Newton et al. 1996).
Transcriptional activation domains of human HSFs. (a) Localization of activation domains in human HSF1, HSF2, and HSF4 and chicken HSF3. Red bars indicate transcriptional activation domains (ADs), and blue bars indicate repression domains of the activation domains. Human HSF1 possesses two activation domains in the C-terminus (AD1, a.a. 372–431; AD2, a.a. 432–529) (Green et al. 1995). The AD1 is predicted to form an α-helix, while AD2 is rich in proline and glycine. The regulatory domain (RD, a.a. 221–310) represses the two activation domains (Green et al. 1995). In human HSF2, two activation domains (a.a. 282–386 and a.a. 472–536) and three negative regulatory domains (a.a. 199–238, a.a. 389–411, and a.a. 428–445) are present (Yoshima et al. 1998b). Human HSF4 also has an activation domain (a.a. 296–395) and a negative regulatory domain (a.a. 200–295) (Nakai et al. 1997). Chicken HSF3 has at least a strong activation domain (a.a. 261–363) (Tanabe et al. 1997). (b) Human HSF1, hHSF2, and HSF4 have different potential to activate transcription. Reporter analysis in human cells treated without (control) and with heat shock (HS) (42 °C for 1 h and then recovery at 37 °C for 6 h) (Nakai et al. 1997; Yoshima et al. 1998a)
In contrast to the strong potential of mammalian HSF1 to activate transcription upon heat shock, the potential of HSF2 and HSF4 is weak in both unstressed and heat-shocked conditions (Tanabe et al. 1999; Yoshima et al. 1998a) (Fig. 2.5b). Nevertheless, HSF2 and HSF4 have a potential to activate the reporter gene constitutively, and HSF4 induces the gene expression in response to proteotoxic stresses including heat shock. In human HSF2, two activation domains (a.a. 282–386 and a.a. 472–536) and three negative regulatory domains (a.a. 199–238, a.a. 389–411, and a.a. 428–445) are present (Yoshima et al. 1998b). Human HSF4 also has an activation domain (a.a. 296–395) and a negative regulatory domain (a.a. 200–295) (Nakai et al. 1997). Chicken HSF3, like human HSF1, has a strong potential to activate transcription upon heat shock and possesses at least a strong activation domain (a.a. 261–363) (Tanabe et al. 1997).
3 Vertebrate HSF Gene Family
In contrast to invertebrate cells, vertebrate cells have four HSF genes. The human, mouse, and chicken genome sequences have become available (International Chicken Genome Sequencing Consortium 2004; International Human Genome Sequencing Consortium 2001; Mouse Genome Sequencing Consortium 2002); therefore, it is possible to compare the syntenic regions, where the same genes occur in a similar order along the chromosomes of different organisms (Koonin et al. 2000). The orthologues of each HSF gene are located in the same syntenic regions, which are derived from the same ancestral genomic region (Fujimoto et al. 2010) (Fig. 2.6). For example, HSF2 is flanked by the SERINCI gene in human, mouse, and chicken orthologous segments. HSF4 was located in a region between the TRADD-FBXL8 and NoL3 in these genomes, and HSF3 was located between the Vsig4 and HEPH. The exon-intron structures of each HSF gene are also evolutionally conserved in three species (Fujimoto et al. 2010).
Syntenic regions containing the vertebrate HSF genes. The location of each segment is as follows: HSF1, human Chr. 8 q24.3 and mouse Chr. 15 D3; HSF2, human Chr. 6 q22.31, mouse Chr. 10 B4, and chicken Chr. 3 63.95–63.98 Mb; HSF3, human Chr. X q12, mouse Chr. X B4, and chicken 0.252–0.265 Mb; HSF4, human Chr. 16 q22.1, mouse Chr. 8 D3, and chicken 2.44–2.45 Mb. A genomic sequence corresponding to chicken HSF1 cDNA has not yet been identified. Arrows indicate the 5′ to 3′ orientation of each gene. Colored boxes indicate the HSF genes, which are flanked by genes shown as white boxes. Human HSF3 is a pseudogene. Note that size markers differ in human, mouse, and chicken HSF3 loci
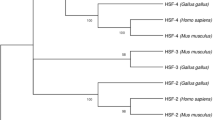
Four HSF family members are expressed in mouse and chicken cells, whereas human HSF3 is not expressed since human HSF3 is a pseudogene (Fujimoto et al. 2010). Because all of human, mouse, and chicken HSF genes have been molecularly cloned and functionally characterized, phylogenetic tree is generated from predicted amino acid sequences of the HSF family members in these endotherms (Fig. 2.7). It shows that the sequence of each HSF (HSF1, HSF2, HSF3, or HSF4) in a species is more related with those of orthologous ones in other species than those of paralogous ones. Furthermore, the sequences of HSF1 orthologues are more related with those of HSF4 orthologues than those of HSF2 or HSF3 orthologues, whereas those of HSF2 orthologues are more related with those of HSF3 orthologues. It has been suggested that two rounds of whole-genome duplication have occurred in vertebrate ancestral cells more than 440 million years (Myr) ago (Holland et al. 1994; Ohno 1970; Putnam et al. 2008), which resulted in polyploidization. Thereafter, avian and mammalian cells evolved differently from an ancestral cell 310 Myr ago, and human and mouse cells did 75 Myr ago. Taken together, four HSF genes may have been generated through the two rounds of whole-genome duplication, and thereafter the sequences and functions have been conserved or diverged during vertebrate evolution (Semon and Wolfe 2007). For example, mammal HSF1 is required for the induction of HSPs during heat shock, whereas HSF3, but not HSF1, is required for this response in birds (McMillan et al. 1998; Tanabe et al. 1998). In contrast, HSF4 is dominantly expressed in the lenses of the human, mouse, and chicken eyes (Bu et al. 2002; Fujimoto et al. 2004; Fujimoto et al. 2010). The phylogenetic tree also shows that the relatedness of chicken HSF3 with mammalian HSF3 is much weaker than that of chicken HSF1 with mammalian HSF1, that of chicken HSF2 with mammalian HSF2, or even that of chicken HSF4 with mammalian HSF4. These estimations indicate that nucleic acid sequences of HSF3 diverged most quickly during evolution, whereas those of HSF1 and HSF2 were similarly conserved. The divergence rates of the HSF4 sequence might also be relatively high.
The phylogenetic tree of HSF family members. Phylogenetic tree generated in CLUSTAL X (Thompson et al. 1997), using amino acid sequences of human (h), mouse (m), and chicken (c) HSF family members as well as Drosophila melanogaster HSF (DmHSF), C. elegans HSF1 (CeHSF1), and Saccharomyces cerevisiae HSF (ScHSF) (Fujimoto et al. 2010; Inouye et al. 2003). The number represents bootstrap values (1000 bootstrap replicates were performed). The unrooted tree was drawn using program TreeView (Page 1996). The bar represents 0.05 substitutions per site
In contrast to the HSF family members in the endotherms, genes of the HSF members in ectotherms, such as frog, lizard, and fish, have been molecularly cloned and functionally characterized only partially (Hilgarth et al. 2004; Råbergh et al. 2000; Stump et al. 1995; Swan et al. 2012; Yeh et al. 2006; Zatsepina et al. 2000) (Fig. 2.8). Analysis using the gene database (http://www.ensembl.org/index.html) suggests that HSF1, HSF2, and HSF4 genes are evolutionally conserved in all vertebrate species. In contrast, HSF3 is conserved in rodents, avians, reptiles, and amphibians, while HSF3 seems to be lost in some mammalian and fish species. In plants, 252 HSF family members, which have similar structures and features, are identified from nine plant species (Scharf et al. 2012). It is necessary to examine the function of each HSF member in ectotherms in the future.
Members of vertebrate HSF gene family. (a). Evolution of the vertebrates. Phylogenetic tree among main groups of the vertebrates is shown (Patterson 2001). Two rounds (2R) of whole-genome duplication (WGD) may have occurred in vertebrate ancestral cells more than 440 million years (Myr) ago. (b) HSF genes in vertebrate species. HSF genes, whose products are registered in NCBI database (http://www.ncbi.nlm.nih.gov/), are shown. 1 HSF1; 2 HSF2; 3 HSF3; 4 HSF4. HSF3 protein is not registered in chimpanzee, zebra fish, medaka, and coelacanth (x). Human HSF3 is a pseudogene (white circle). Cloned cDNAs are zebra fish HSF1 (Hsu and Yeh 2002; Råbergh et al. 2000), HSF2 (Yeh et al. 2006), and HSF4 (Swan et al. 2012); frog HSF1 (Mercier et al. 1997) and HSF2 (Hilgarth et al. 2004); chicken HSF1, HSF2, HSF3 (Nakai and Morimoto 1993), and HSF4 (Fujimoto et al. 2010); mouse HSF1, HSF2 (Sarge et al. 1991), HSF3 (Fujimoto et al. 2010), and HSF4 (Tanabe et al. 1999); and human HSF1 (Rabindran et al. 1991), HSF2 (Schuetz et al. 1991), HSF3 (Fujimoto et al. 2010), and HSF4 (Nakai et al. 1997). The genes for shark HSF3 (XP_007891631), frog HSF3 (NP_001039053), turtle HSF3 (XP_006136197), lizard HSF3 (XP_008118582), alligator HSF3 (XP_006258300), and zebra finch HSF3 (XP_004177297.1) are located on the same syntenic regions in the genomes of these species
4 Potential to Activate HSP and Non-HSP Genes During Heat Shock
The heat shock response is characterized by the induction of HSP gene expression during heat shock. After the identification of multiple HSF genes in vertebrate genomes (Fujimoto et al. 2010; Nakai et al. 1997; Nakai and Morimoto 1993; Rabindran et al. 1991; Sarge et al. 1991; Schuetz et al. 1991), researchers have investigated which HSF activates the HSP genes. Biochemical analyses showed that human and mouse HSF1, but not HSF2, acquires DNA-binding activity and translocated to the nucleus in response to heat shock, suggesting that HSF1 is involved in the induction of HSP genes (Baler et al. 1993; Sarge et al. 1993). Subsequently, disruption of HSF genes in mouse embryonic fibroblasts (MEFs) demonstrated that mouse HSF1 is required for the HSP induction during heat shock (McMillan et al. 1998; Zhang et al. 2002), while HSF2 is not (McMillan et al. 2002). Overexpression of human HSF1 restored the induction of HSPs during heat shock (Inouye et al. 2003). HSF1 is also responsible for the induction of the HSP expression in response to other proteotoxic stresses including a blockade of proteasome and an incorporation of amino acid analogues. Thus, HSF1 is a master regulator of the HSP expression in mammalian cells. HSF2 modulates the expression of HSPs to some extent, probably through the direct interaction with HSF1 (Ostling et al. 2007).
In addition to the expression of HSP genes, the expression of many non-HSP genes is induced during heat shock in mammalian cells (Trinklein et al. 2004). HSF1 plays a major role in the induction of these non-HSP genes, but HSF2, HSF3, and HSF4 are also involved in the induced expression of some non-HSP genes in MEF cells (Fujimoto et al. 2008; Fujimoto et al. 2010; Shinkawa et al. 2011).
Avian HSF genes have functionally diverged during evolution. Both HSF1 and HSF3 acquired the DNA-binding activity during heat shock in chicken cells (Nakai et al. 1995). Unexpectedly, disruption of chicken HSF3 in chicken B-lymphocyte DT40 cells resulted in a severe reduction in the induction of HSP70 expression during heat shock, and the expression of HSP110, HSP90α, HSP90β, and HSP40 was not induced at all (Tanabe et al. 1998). The disruption of chicken HSF1 had no effect on the induction of HSP expression during heat shock in DT40 cells, and overexpression of chicken HSF1 did not restore the induction of HSPs during heat shock in HSF1-null MEF cells (Inouye et al. 2003). These results demonstrate that HSF3, but not HSF1, is a master regulator of the HSP expression in chicken cells.
5 Oligomeric State
HSFs are defined by its ability to bind to the heat shock response element (HSE) that is composed of three inverted repeats of an nGGAn pentanucleotide. Therefore, activated HSFs form homotrimers that bind to the HSE with high affinity, through the interaction between the HR-A/B domains (Fig. 2.9). In budding yeast Saccharomyces cerevisiae, HSF is constitutively a DNA-binding trimer because it does not have the HR-C domain that inhibits oligomerization of the HR-A/B (Sorger and Pelham 1988; Wiederrecht et al. 1988). It acquires elevated potential to activate transcription via phosphorylation upon heat shock (Sorger 1990). In contrast, HSF from fission yeast Schizosaccharomyces pombe or Drosophila melanogaster stays an inert monomer in unstressed condition and is converted to a DNA-binding trimer in heat shock condition (Clos et al. 1990; Gallo et al. 1993). Thus, a monomer-to-trimer transition of HSF is a fundamental mechanism of HSF activation in response to heat shock (Rabindran et al. 1993).
Oligomeric states of mammalian HSFs. HSF1 remains mostly as an inert monomer in unstressed condition and is converted to a DNA-binding trimer. The activated HSF1 binds to the HSE located in the promoters of HSP and non-HSP genes. HSF2 stays as a dimer and a trimer and is converted to a trimer in response to proteasome inhibition or mild heat shock. HSF4 exists as a DNA-binding trimer constitutively because it lacks the HR-C. HSF3 may be a trimer in unstressed mouse cells. Mouse HSF3 may exist in trimer in unstressed condition
Oligomeric status of vertebrate HSFs involves a monomer, dimer, and trimer (Fig. 2.9). Mammalian HSF1, like Drosophila HSF, mostly stays as an inert monomer in unstressed condition and is converted to a DNA-binding trimer (Baler et al. 1993; Sarge et al. 1993). Mammalian HSF2 stays as both an inert dimer and a DNA-binding trimer in unstressed condition, and the latter increased during the treatment with cells with proteasome inhibitors such as MG132, lactacystin, and hemin (Sistonen et al. 1994; Mathew et al. 1998). Although HSF2 is unstable in severe heat shock conditions such as 42 °C (Sistonen et al. 1994), it is stable and acquires a DNA-binding activity through trimerization in febrile-range, mild heat shock conditions such as 40 °C (Shinkawa et al. 2011). Mammalian HSF4 lacks the HR-A/B domain and therefore forms a DNA-binding trimer constitutively in the lens and other tissues including the brain and lung (Fujimoto et al. 2004; Tanabe et al. 1999). HSF3 may exist as a trimer in unstressed mouse cells, because sequences of the heptad repeats of hydrophobic amino acids in the HR-C domain are not well conserved (Fujimoto et al. 2010). It is worth noticing that chicken HSF3, a master regulator of HSP expression, stays mostly as an inert dimer in control condition and is converted to a DNA-binding trimer upon heat shock (Nakai et al. 1995). Therefore, the dimer-to-trimer transition of HSF2 and HSF3 is also one of HSF activation mechanisms in vertebrates.
6 Recognition Sequences
Extensive studies have revealed that HSP genes in Drosophila and other eukaryotic species have a 14 bp conserved promoter element, CnnGAAnnTTCnnG (n is any nucleic acid), termed the heat shock response element (HSE) (Pelham 1985). Furthermore, inverted repeats of a 5 bp sequence nGAAn are shown to be key features of the HSE (Xiao and Lis 1988; Amin et al. 1998). In fact, Drosophila HSF trimer stably binds to the HSE composed of three contiguous inverted 5 bp units (Perisic et al. 1989; Xiao et al. 1991). Analysis of crystal structure of the DBD of yeast HSF reveals that the second G and ninth C of 5′-nGAAnnTTCn-3′ in major groove of DNA are essential for the interaction with the α-helix H3 of the winged helix-turn-helix motif (Littlefield and Nelson 1999).
Recognition sequences of mammalian HSFs have been analyzed using in vitro random oligonucleotide selection. As is expected from the fact that sequences of the DBD in all of eukaryotic HSFs are highly conserved, both mouse HSF1 and HSF2 recognize inverted repeats of a pentameric consensus nGAAn (Kroeger and Morimoto 1994) (Fig. 2.10). Each HSF has some preference for the nucleotides flanking the core GAA motif. In contrast, human HSF4 uniquely recognizes inverted repeats of an ambiguous nGnnn sequence in vitro (Fujimoto et al. 2008). Furthermore, most of the HSF4 binding region contains none or only one GAA sequence even in vivo. The numbers of consensus pentamers per binding sites are high (four to five pentamers) in only HSF1-selected oligonucleotide, suggesting that cooperative binding between trimers could affect the binding of HSF1 (Bonner et al. 1994; Kroeger and Morimoto 1994).
In vitro consensus sequence of mammalian HSF binding sites. Consensus recognition sequences for mammalian HSF1, HSF2, and HSF4 binding in vitro were generated with WebLogo (http://weblogo.berkeley.edu/) by using DNA sequences of HSF1, HSF2 and HSF4 binding fragments (Fujimoto et al. 2008; Kroeger and Morimoto 1994). The height of each letter represents the relative frequency of nucleotides at different positions in the consensus
7 Cooperativity and Competition Between Distinct HSFs
Vertebrate HSF family members typically form a homotrimer when they are activated and bind to the HSEs in the promoters of target genes. Cooperativity of HSF1 and HSF2 has been shown in spermatogenesis (Wang et al. 2004). Spermatogenesis is normal in HSF1-null mice (Izu et al. 2004) and is partially impaired in HSF2-null mice (Kallio et al. 2002; Wang et al. 2003). These male mice are still fertile, although the number of sperm is markedly reduced in HSF2-null mice. In marked contrast, spermatogenesis is completely blocked in mice lacking both HSF1 and HSF2, and the double-null mice are infertile (Wang et al. 2004). Analysis of target genes shows that the expression of some genes, including HSP70-2, HSC70t, and acrosin, is significantly reduced only in double-null testis. Thus, HSF1 and HSF2 cooperatively upregulate target genes and support spermatogenesis. In addition, both factors are required for the constitutive and inducible expression of CRYAB in mouse embryonic fibroblasts (Shinkawa et al. 2011), and HSF2 enhances the HSF1-mediated inducible expression of HSP70 (Östling et al. 2007). Mechanisms of these cooperativities are unclear yet. HSF1 and HSF2 trimers may simultaneously occupy the target gene promoters, or HSF1 trimer may interact with HSF2 trimer. Alternatively, HSF1 and HSF2 may form heterotrimers (Sandqvist et al. 2009).
On the contrary, there is evidence that HSF1 and HSF4 compete with each other. In lens fiber cells of the mouse lens, both HSF1 and HSF4 cooperatively upregulate the expression of γ-crystallin genes. In contrast, in lens epithelial cells, HSF4 downregulates the expression of FGF genes, while HSF1 upregulates it (Fujimoto et al. 2004). Therefore, increased proliferation and premature differentiation due to the elevated expression of FGFs in the HSF4-null lens are alleviated in the lens lacking both HSF4 and HSF1. Furthermore, HSF1 and HSF4 have opposing effects on the expression of LIF gene in the mouse olfactory epithelium (Takaki et al. 2006). Thus, HSF4 competes with HSF1 for the expression of genes such as cytokine genes.
In chicken B-lymphocyte DT40 cells, disruption of HSF1 or HSF3 gene does not affect the expression of HSP90α (Nakai and Ishikawa 2001). However, disruption of both genes results in the marked reduction of the constitutive expression of HSP90α and blockade of cell cycle progression at elevated temperature. Thus, avian HSF1 and HSF3 redundantly regulate the expression of HSP90α, which is required for cell cycle transition under stress condition (Morano et al. 1999; Zarzov et al. 1997).
8 Future Perspectives
In this chapter, we described mainly the structure and function of four HSF genes in vertebrates. The crystal structure and NMR solution structure of the DBD in yeast HSF revealed the precise structure in the DBD. It is proposed that an inert HSF1 monomer is folded for the HR-A/B and HR-C to interact with each other in unstressed condition and is converted to three-bundle, rod-shaped structure in response to heat shock. The structure of full-length HSF1 or domains other than the DBD is not analyzed yet, and these analyses should be done in the future. Among domains of HSFs, amino acid sequences of the DBD and oligomerization domains are highly conserved, and functions of these domains are well known. On the other hand, roles of the regions X and Y and the DHR are little known. There are several conserved sequences termed a to g sites in these regions, whose functions should be revealed.
In mammalian cells, HSF1 is required for the heat shock response, while roles of other HSF family members had hardly been known until recently. It is now known that not only HSF1 but also HSF2, HSF3, and HSF4 regulate proteostasis capacity within a cell to adapt to proteotoxic stresses. However, target genes and activation mechanisms of each HSF are not well known at present. In addition, mechanisms of cooperative and competitive regulation by the HSF family members are still unclear and should be clarified in the future.
References
Amin J, Anathan J, Voellmy R (1998) Key features of heat shock regulatory elements. Mol Cell Biol 8:3761–3769
Baler R, Dahl G, Voellmy R (1993) Activation of human heat shock genes is accompanied by oligomerization, modification, and rapid translocation of heat shock transcription factor HSF1. Mol Cell Biol 13:2486–2496
Bonner JJ, Ballou C, Fackenthal DL (1994) Interactions between DNA-bound trimers of the yeast heat shock factor. Mol Cell Biol 14:501–508
Brown SA, Weirich CS, Newton EM et al (1998) Transcriptional activation domains stimulate initiation and elongation at different times and via different residues. EMBO J 17:3146–3154
Bu L, Jin Y, Shi Y et al (2002) Mutant DNA-binding domain of HSF4 is associated with autosomal dominant lamellar and Marner cataract. Nat Genet 31:276–278
Chang Y, Ostling P, Akerfelt M et al (2006) Role of heat-shock factor 2 in cerebral cortex formation and as a regulator of p35 expression. Genes Dev 20:836–847
Cicero MP, Hubl ST, Harrison CJ et al (2001) The wing in yeast heat shock transcription factor (HSF) DNA-binding domain is required for full activity. Nucleic Acids Res 29:1715–1723
Clos J, Westwood JT, Becker PB et al (1990) Molecular cloning and expression of a hexameric Drosophila heat shock factor subject to negative regulation. Cell 63:1085–1097
Clos J, Rabindran S, Wisniewski J et al (1993) Induction temperature of human heat shock factor is reprogrammed in a Drosophila cell environment. Nature 364:252–255
Creighton TE (1993) Proteins: structures and molecular properties. W H Freeman & Company, New York
Damberger FF, Pelton JG, Liu C et al (1995) Refined solution structure and dynamics of the DNA-binding domain of the heat shock factor from Kluyveromyces lactis. J Mol Biol 254:704–719
Fang X, Chen T, Tran K, Parker CS (2001) Developmental regulation of the heat shock response by nuclear transport factor karyopherin-alpha3. Development 128(17):3349–3358
Fernandes M, O’Breien T, Lis JT (1994) Structure and regulation of heat shock gene promoters. In: Morimoto RI, Tissieres A, Georgopoulos C (eds) The biology of heat shock proteins and molecular chaperones. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp 375–393
Flick KE, Gonzalez L Jr, Harrison CJ et al (1994) Yeast heat shock transcription factor contains a flexible linker between the DNA-binding and trimerization domains. Implications for DNA binding by trimeric proteins. J Biol Chem 269:12475–12481
Fujimoto M, Izu H, Seki K et al (2004) HSF4 is required for normal cell growth and differentiation during mouse lens development. EMBO J 23:4297–4306
Fujimoto M, Oshima K, Shinkawa T et al (2008) Analysis of HSF4 binding regions reveals its necessity for gene regulation during development and heat shock response in mouse lenses. J Biol Chem 283:29961–29970
Fujimoto M, Hayashida N, Katoh T et al (2010) A novel mouse HSF3 has the potential to activate non-classical heat shock genes during heat shock. Mol Biol Cell 20:106–116
Fujimoto M, Takaki E, Takii R et al (2012) RPA assists HSF1 access to nucleosomal DNA by recruiting histone chaperone FACT. Mol Cell 48:182–194
Gajiwala KS, Burley SK (2000) Winged helix proteins. Curr Opin Struct Biol 10:110–116
Gallo GJ, Prentice H, Kingston RE (1993) Heat shock factor is required for growth at normal temperature in the fission yeast Schizosaccharomyces pombe. Mol Cell Biol 13:749–761
Giardina C, Lis JT (1995) Dynamic protein-DNA architecture of a yeast heat shock promoter. Mol Cell Biol 15:2737–2744
Green M, Schuetz TJ, Sullivan EK et al (1995) A heat shock-responsive domain of human HSF1 that regulates transcription activation domain function. Mol Cell Biol 15:3354–3362
Harrison CJ, Bohm AA, Nelson HC (1994) Crystal structure of the DNA binding domain of the heat shock transcription factor. Science 263:224–227
Hilgarth RS, Murphy LA, O’Connor CM et al (2004) Identification of Xenopus heat shock transcription factor-2: conserved role of sumoylation in regulating deoxyribonucleic acid-binding activity of heat shock transcription factor-2 proteins. Cell Stress Chaperones 9:214–220
Holland PW, Garcia-Fernàndez J, Williams NA et al (1994) Gene duplications and the origins of vertebrate development. Development 1994(Suppl):125–133, PMID: 7579513
Hsu T, Yeh FL (2002) Differential regulation of spontaneous and heat-induced HSP 70 expression in developing zebrafish (Danio rerio). J Exp Zool 293:349–359
Hsu AL, Murphy CT, Kenyon C (2003) Regulation of aging and age-related disease by DAF-16 and heat-shock factor. Science 300:1142–1145
Inouye S, Katsuki K, Izu H et al (2003) Activation of heat shock genes is not necessary for protection by heat shock transcription factor 1 against cell death due to a single exposure to high temperatures. Mol Cell Biol 23:5882–5895
International Chicken Genome Sequencing Consortium (2004) Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 432:695–716
International Human Genome Sequencing Consortium (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Izu H, Inouye S, Fujimoto M et al (2004) HSF1 is involved in quality control mechanisms in male germ cells. Biol Reprod 70:18–24
Jakobsen BK, Pelham HR (1988) Constitutive binding of yeast heat shock factor to DNA in vivo. Mol Cell Biol 8:5040–5042
Jakobsen BK, Pelham HR (1991) A conserved heptapeptide restrains the activity of the yeast heat shock transcription factor. EMBO J 10:369–375
Kallio M, Chang Y, Manuel M et al (2002) Brain abnormalities, defective meiotic chromosome synapsis and female subfertility in HSF2 null mice. EMBO J 21:2591–2601
Koonin EV, Aravind L, Kondrashov AS (2000) The impact of comparative genomics on our understanding of evolution. Cell 101:573–576
Kotak S, Port M, Ganguli A et al (2004) Characterization of C-terminal domains of Arabidopsis heat stress transcription factors (Hsfs) and identification of a new signature combination of plant class a Hsfs with AHA and NES motifs essential for activator function and intracellular localization. Plant J 39:98–112
Kroeger PE, Morimoto RI (1994) Selection of new HSF1 and HSF2 DNA-binding sites reveals difference in trimer cooperativity. Mol Cell Biol 14:7592–7603
Lindquist S (1986) The heat-shock response. Ann Rev Biochem 55:1151–1191
Littlefield O, Nelson HC (1999) A new use for the ‘wing’ of the ‘winged’ helix-turn-helix motif in the HSF-DNA cocrystal. Nat Struct Biol 6:464–470
Liu XD, Thiele DJ (1996) Oxidative stress induced heat shock factor phosphorylation and HSF-dependent activation of yeast metallothionein gene transcription. Genes Dev 10:592–603
Mathew A, Mathur SK, Morimoto RI (1998) Heat shock response and protein degradation: regulation of HSF2 by the ubiquitin-proteasome pathway. Mol Cell Biol 18:5091–5098
McMillan DR, Xiao X, Shao L et al (1998) Targeted disruption of heat shock transcription factor 1 abolishes thermotolerance and protection against heat-inducible apoptosis. J Biol Chem 273:7523–7528
McMillan DR, Christians E, Forster M et al (2002) Heat shock transcription factor 2 is not essential for embryonic development, fertility, or adult cognitive and psychomotor function in mice. Mol Cell Biol 22:8005–8014
Mercier PA, Foksa J, Ovsenek N et al (1997) Xenopus heat shock factor 1 is a nuclear protein before heat stress. J Biol Chem 272:14147–14151
Morano KA, Santoro N, Koch KA et al (1999) A trans-activation domain in yeast heat shock transcription factor is essential for cell cycle progression during stress. Mol Cell Biol 19:402–411
Morley JF, Morimoto RI (2004) Regulation of longevity in Caenorhabditis elegans by heat shock factor and molecular chaperones. Mol Biol Cell 15:657–664
Mouse Genome Sequencing Consortium (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420:520–562
Nakai A, Ishikawa T (2000) A nuclear localization signal is essential for stress-induced dimer-to-trimer transition of heat shock transcription factor 3. J Biol Chem 275:34665–34671
Nakai A, Ishikawa T (2001) Cell cycle transition under stress conditions controlled by vertebrate heat shock factors. EMBO J 20:2885–2895
Nakai A, Morimoto RI (1993) Characterization of a novel chicken heat shock transcription factor, heat shock factor 3, suggests a new regulatory pathway. Mol Cell Biol 13:1983–1997
Nakai A, Kawazoe Y, Tanabe M et al (1995) The DNA-binding properties of two heat shock factors, HSF1 and HSF3 are induced in the avian erythroblast cell line HD6. Mol Cell Biol 15:5268–5278
Nakai A, Tanabe M, Kawazoe Y et al (1997) HSF4, a new member of the human heat shock factor family which lacks properties of a transcriptional activator. Mol Cell Biol 17:469–481
Newton EM, Knauf U, Green M et al (1996) The regulatory domain of human heat shock factor 1 is sufficient to sense heat stress. Mol Cell Biol 16:839–846
Ohno S (1970) Evolution by gene duplication. Springer, New York
Orosz A, Wisniewski J, Wu C (1996) Regulation of Drosophila heat shock factor trimerization : global sequence requirements and independence of nuclear localization. Mol Cell Biol 16:7018–7030
Östling P, Björk JK, Roos-Mattjus P et al (2007) Heat shock factor 2 (HSF2) contributes to inducible expression of hsp genes through interplay with HSF1. J Biol Chem 282:7077–7086
Page RD (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Patterson C (2001) Evolution (Translated by Mawatari S, Uehara M, Isono N), 2nd edn, p 181
Pelham HR (1985) Activation of heat-shock genes in eukaryotes. Trends Genet 1:31–35
Perisic O, Xiao H, Lis JT (1989) Stable binding of Drosophila heat shock factor to head-to-head and tail-to tail repeats of a conserved 5bp recognition unit. Cell 59:797–806
Peteranderl R, Rabenstein M, Shin YK et al (1999) Biochemical and biophysical characterization of the trimerization domain from the heat shock transcription factor. Biochemistry 38:3559–3569
Putnam NH, Butts T, Ferrier DE et al (2008) The amphioxus genome and the evolution of the chordate karyotype. Nature 453:1064–1071
Råbergh CM, Airaksinen S, Soitamo A et al (2000) Tissue-specific expression of zebrafish (Danio rerio) heat shock factor 1 mRNAs in response to heat stress. J Exp Biol 203:1817–1824
Rabindran SK, Giorgi G, Clos J et al (1991) Molecular cloning and expression of a human heat shock factor, HSF1. Proc Natl Acad Sci U S A 88:6906–6910
Rabindran SK, Haroun RI, Clos J et al (1993) Regulation of heat shock factor trimer formation: role of a conserved leucine zipper. Science 259:230–234
Raychaudhuri S, Loew C, Körner R et al (2014) Interplay of acetyltransferase EP300 and the proteasome system in regulating heat shock transcription factor 1. Cell 156:975–985
Sandqvist A, Björk JK, Åkerfelt M et al (2009) Heterotrimerization of heat-shock factors 1 and 2 provides a transcriptional switch in response to distinct stimuli. Mol Biol Cell 20:1340–1347
Sarge KD, Zimarino V, Holm K et al (1991) Cloning and characterization of two mouse heat shock factors with distinct inducible and constitutive DNA-binding ability. Genes Dev 5:1902–1911
Sarge KD, Murphy SP, Morimoto RI (1993) Activation of heat shock gene transcription by heat shock factor 1 involves oligomerization, acquisition of DNA-binding activity, and nuclear localization and can occur in the absence of stress. Mol Cell Biol 13:1392–1407
Scharf KD, Berberich T, Ebersberger I et al (2012) The plant heat stress transcription factor (Hsf) family: structure, function and evolution. Biochim Biophys Acta 1819:104–119
Schuetz TJ, Gallo GJ, Sheldon L et al (1991) Isolation of a cDNA for HSF2: evidence for two heat shock factor genes in humans. Proc Natl Acad Sci U S A 88:6911–6915
Sémon M, Wolfe KH (2007) Consequences of genome duplication. Curr Opin Genet Dev 17:505–512
Sheldon LA, Kingston RE (1993) Hydrophobic coiled-coil domains regulate the subcellular localization of human heat shock factor 2. Genes Dev 7:1549–1558
Shi Y, Kroeger PE, Morimoto RI (1995) The carboxyl-terminal transactivation domain of heat shock factor 1 is negatively regulated and stress responsive. Mol Cell Biol 15:4309–4318
Shinkawa T, Tan K, Fujimoto M et al (2011) Heat shock factor 2 is required for maintaining proteostasis against febrile-range thermal stress and polyglutamine aggregation. Mol Biol Cell 22:3571–3583
Sistonen L, Sarge KD, Morimoto RI (1994) Human heat shock factors 1 and 2 are differentially activated and can synergistically induce hsp70 gene transcription. Mol Cell Biol 14:2087–2099
Sorger PK (1990) Yeast heat shock factor contains separable transient and sustained response transcriptional activators. Cell 62:793–805
Sorger PK, Nelson HC (1989) Trimerization of a yeast transcriptional activator via a coiled-coil motif. Cell 59:807–813
Sorger PK, Pelham HR (1988) Yeast heat shock factor is an essential DNA-binding protein that exhibits temperature-dependent phosphorylation. Cell 54:855–864
Stump DG, Landsberger N, Wolffe AP (1995) The cDNA encoding Xenopus laevis heat-shock factor 1 (XHSF1): nucleotide and deduced amino-acid sequences, and properties of the encoded protein. Gene 160:207–211
Swan CL, Evans TG, Sylvain N et al (2012) Zebrafish HSF4: a novel protein that shares features of both HSF1 and HSF4 of mammals. Cell Stress Chaperones 17:623–637
Takaki E, Fujimoto M, Sugahara K et al (2006) Maintenance of olfactory neurogenesis requires HSF1, a major heat shock transcription factor in mice. J Biol Chem 281:4931–4937
Takii R, Fujimoto M, Tan K et al (2015) ATF1 modulates the heat shock response by regulating the stress-inducible heat shock factor 1 transcription complex. Mol Cell Biol 35:11–25
Tanabe M, Nakai A, Kawazoe Y et al (1997) Different thresholds in the responses of two heat shock transcription factors, HSF1 and HSF3. J Biol Chem 272:15389–15395
Tanabe M, Kawazoe Y, Takeda S et al (1998) Disruption of the HSF3 gene results in the severe reduction of heat shock gene expression and loss of thermotolerance. EMBO J 17:1750–1758
Tanabe M, Sasai N, Nagata K et al (1999) The mammalian HSF4 gene generates both an activator and a repressor of heat shock genes by alternative splicing. J Biol Chem 274:27845–27856
Thompson JD, Gibson TJ, Plewniak F et al (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Trinklein ND, Murray JI, Hartman SJ et al (2004) The role of heat shock transcription factor 1 in the genome-wide regulation of the mammalian heat shock response. Mol Biol Cell 15:1254–1261
Vuister GW, Kim SJ, Orosz A et al (1994a) Solution structure of the DNA-binding domain of Drosophila heat shock transcription factor. Nat Struct Biol 1:605–614
Vuister GW, Kim SJ, Wu C et al (1994b) NMR evidence for similarities between the DNA-binding regions of Drosophila melanogaster heat shock factor and the helix-turn-helix and HNF-3/forkhead families of transcription factors. Biochemistry 33:10–16
Vujanac M, Fenaroli A, Zimarino V (2005) Constitutive nuclear import and stress-regulated nucleocytoplasmic shuttling of mammalian heat-shock factor 1. Traffic 6:214–229
Wang G, Zhang J, Moskophidis D et al (2003) Targeted disruption of the heat shock transcription factor (hsf)-2 gene results in increased embryonic lethality, neuronal defects, and reduced spermatogenesis. Genesis 36:48–61
Wang G, Ying Z, Jin X et al (2004) Essential requirement for both hsf1 and hsf2 transcriptional activity in spermatogenesis and male fertility. Genesis 38:66–80
Wiederrecht G, Seto D, Parker CS (1988) Isolation of the gene encoding the S. cerevisiae heat shock transcription factor. Cell 54:841–853
Xiao H, Lis JT (1988) Germline transformation used to define key features of heat-shock response elements. Science 239:1139–1142
Xiao H, Perisic O, Lis JT (1991) Cooperative binding of Drosophila heat shock factor to arrays of a conserved 5 bp unit. Cell 64:585–593
Yeh FL, Hsu LY, Lin BA et al (2006) Cloning of zebrafish (Danio rerio) heat shock factor 2 (HSF2) and similar patterns of HSF2 and HSF1 mRNA expression in brain tissues. Biochimie 88:1983–1988
Yoshima T, Yura T, Yanagi H (1998a) Heat shock factor 1 mediates hemin-induced hsp70 gene transcription in K562 erythroleukemia cells. J Biol Chem 273:25466–25471
Yoshima T, Yura T, Yanagi H (1998b) Function of the C-terminal transactivation domain of human heat shock factor 2 is modulated by the adjacent negative regulatory segment. Nucleic Acids Res 26:2580–2585
Zarzov P, Boucherie H, Mann C (1997) A yeast heat shock transcription factor (Hsf1) mutant is defective in both Hsc82/Hsp82 synthesis and spindle pole body duplication. J Cell Sci 110:1879–1891
Zatsepina OG, Ulmasov KA, Beresten SF et al (2000) Thermotolerant desert lizards characteristically differ in terms of heat-shock system regulation. J Exp Biol 203:1017–1025
Zhang Y, Huang L, Zhang J et al (2002) Targeted disruption of hsf1 leads to lack of thermotolerance and defines tissue-specific regulation for stress-inducible Hsp molecular chaperones. J Cell Biochem 86:376–393
Zuo J, Rungger D, Voellmy R (1995) Multiple layers of regulation of human heat shock transcription factor 1. Mol Cell Biol 15:4319–4330
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Japan
About this chapter
Cite this chapter
Takii, R., Fujimoto, M. (2016). Structure and Function of the HSF Family Members. In: Nakai, A. (eds) Heat Shock Factor. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55852-1_2
Download citation
DOI: https://doi.org/10.1007/978-4-431-55852-1_2
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55850-7
Online ISBN: 978-4-431-55852-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)