Abstract
Helicobacter pylori strains from different geographic areas exhibit clear phylogeographical differentiation. The genotype of the virulence genes is useful as a tool to track human migration utilizing the high genetic diversity and frequent recombination between different H. pylori strains. Using combinations of the virulence genes, five major groups have been defined according to geographical associations. Multilocus sequence typing (MLST) analysis using seven housekeeping genes also are widely used markers for genomic diversity. It was revealed that seven modern population types of H. pylori which derived from six ancestral populations provide more detailed information on human migration than does the analysis of human genetics. Although approaches by MLST and virulence factors are effective, these methods focus on a small number of genes and may miss information conveyed by the rest of the genome. Genome-wide analyses using DNA microarray or whole-genome sequencing technology give a broad view on the genome of H. pylori. In particular, next-generation sequencers, which can read DNA sequences in less time and at lower costs than Sanger sequencing, enabled us to efficiently investigate not only the evolution of H. pylori, but also novel virulence factors and genomic changes related to drug resistance.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Helicobacter pylori
- Asia
- Virulence factors
- Multilocus sequence typing
- Next-generation sequencer
- Migration
1 Introduction
More than half of all humans are infected with Helicobacter pylori, a gram-negative spiral bacterium whose ecological niche is the human stomach which is also linked to severe gastritis-associated diseases, including peptic ulcer and gastric cancer [1]. H. pylori strains from different geographical areas show clear phylogeographic features; these features enabled us to assume the migration of human populations by phylogeographic analyses of H. pylori. In addition, the genetic diversity within H. pylori is greater than within most other bacteria [2] and about 50-fold greater than that of the human population [3]. Moreover, frequent recombination between different H. pylori strains [4] leads to only partial linkage disequilibrium between polymorphic loci, which provide additional information for population genetic analysis [5].
Several virulence factors of H. pylori have been demonstrated to be predictors of gastric atrophy, intestinal metaplasia, and severe clinical outcomes [6–11]. Currently, two most extensively studied virulence factors of H. pylori, cagA and vacA, are used as markers for genomic diversity within distinct populations [12]. In addition, multilocus sequence typing (MLST) analysis, which uses seven housekeeping genes, is also useful to predict the history of human migrations [2, 5, 13–15]. MLST was proposed in 1998 as a tool for the epidemiological study of bacteria [16]. Recently, the genomic diversity within H. pylori populations was examined by employing the MLST method using seven housekeeping genes (atpA, efp, mutY, ppa, trpC, ureI, and yphC) [5, 13, 14]. MLST analysis is reported to give more detailed information about human population structure than the method using human microsatellite or mitochondrial DNA [15]. Moreover recently the whole-genome sequencing technology is another powerful tool to study the evolution and pathogenicity of H. pylori. In this chapter we describe the current knowledge about the usefulness of virulence factors and housekeeping genes for elucidating the history of human migration and overview on the utilization of genome-wide information for advanced studies.
2 Migration out of Africa to the Pacific
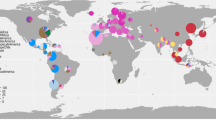
It was believed that H. pylori was already established in human stomachs at least 100,000 years ago [17] from an unknown source. It is most likely transmitted from large felines which contained H. acinonychis to San peoples (hpAfrica2; very distinct and has only been isolated in South Africa) and then widespread throughout Africa (hpAfrica1 and hpNEAfrica) [18]. hpAfrica1 divided into two subpopulations, hspWAfrica (West Africans, South Africans, and Afro-Americans) and hspSAfrica (South Africans). On the other hand, hpNEAfrica is predominant in isolates from Northeast Africa [19]. H. pylori is predicted to have spread from East Africa over the same time period as anatomically modern humans (~58,000 years ago) and mirrors the human pattern of increased genetic distance and decreased diversity with distance from Africa (Fig. 2.1) [13, 14]. Using MLST the modern populations derived from six ancestral populations (Table 2.1) which were designated ancestral European 1 (AE1), ancestral European 2 (AE2), ancestral East Asia, ancestral Africa 1, ancestral Africa 2 [5], and ancestral Sahul [14]. These ancestrals recently derived to seven population types based on geographical associations: hpEurope, hpEastAsia, hpAfrica1, hpAfrica2, hpAsia2, hpNEAfrica, and hpSahul (Fig. 2.2) [5, 13, 14].
By a southern coastal route, the ancestors of modern humans passed from India to the Southeast and Australasia [20] during their first “out of Africa” migration, which subsequently resulted in the Asian lineages (hpAsia2). Recently hpAsia2 strains have been isolated in South, Southeast, and Central Asia [19]. Most strains in India initially belonged to hpAsia2 [13], whereas some strains belonged to hpEurope [21]. However H. pylori in the Indian population is more heterogeneous in origin, reflecting perhaps both earlier common ancestry and recent imports. It is notable that hpAsia2 strains from Ladakh Indians and Malaysian Indians can be divided into two subpopulations, hspLadakh and hspIndia [22]. From mainland Asia the route extended along the Pleistocene landmass, known as Sundaland (i.e., the Malay Peninsula, Sumatra, Java, Borneo, and Bali), that was joined to the Asian mainland as a result of low sea levels during the last ice age (12,000–43,000 years ago). Low sea levels also meant that Australia, New Guinea, and Tasmania were connected in a continent called Sahul, separated from Sundaland by a few narrow deep-sea channels [14]. Recently hpSahul strains are isolated from aborigines of Australia and highlanders of New Guinea [19].
Subsequent migrations of ancestors of the African hpNEAfrica and/or the Asian hpAsia2 populations resulted in the admixed hpEurope population which then became the predominant population of extant H. pylori in Europe, the Middle East, and Western Asia. The modern humans settled in Europe about 30,000–40,000 years ago, probably entering via two routes: from Turkey along the Danube corridor into Eastern Europe and along the Mediterranean coast [20]. hpEurope includes almost all H. pylori strains isolated from ethnic Europeans, including people from countries colonized by Europeans. The hpEurope can be divided into AE1 and AE2. AE1 originated in Central Asia, because it shares phylogenetic signals with isolates from Estonia, Finland, and Ladakh in India. It is not clear which population arrived first, but AE1 has a higher frequency in Northern Europe, while AE2 is more common in southern Europe. MLST analyses from Iran also provided evidence that H. pylori strains from Iran are similar to other isolates from Western Eurasia and can be placed in the previously described hpEurope population [23].
Human migrations in Southeast Asia have also been clarified on the basis of MLST analyses from Cambodia [24]. Cambodian strains have been classified in two groups, hpEurope and hspEAsia, which have resulted from three ancient human migrations: (1) from India, introducing hpEurope into Southeast Asia; (2) from China, carrying hspEAsia; and (3) from Southern China into Thailand carrying hpAsia2 [20, 24]. Their findings also support two recent migrations within the last 200 years: (1) from the Chinese to Thailand and Malaysia spreading hspEAsia strains and (2) from Indians to Malaysia carrying hpAsia2 and hpEurope [20, 24]. In concordance with this study, H. pylori isolates from Malaysia are classified as hpEastAsia, hpAsia2, or hpEurope. A new subpopulation within hpAsia2, hspIndia, may reflect as the Malaysian Indians mainly came from South India.
hpEastAsia is common in H. pylori isolates from East Asia. hpEastAsia also includes subpopulations, i.e., hspMaori (Polynesians, Melanesians, and native Taiwanese), hspAmerind (Amerindians), and hspEAsia (East Asians). Approximately 12,000 years ago, H. pylori (hspAmerind) accompanied humans when they crossed the Bering Strait from Asia to the Americas [12]. Our previous data showed that four strains isolated from the Ainu ethnic group, living in Hokkaido, a northern island of Japan, belong to the hspAmerind population [25]. Japanese aboriginal people, known as Jomon people, are thought to have migrated to the northern or southern area such as Hokkaido and Okinawa because of the immigration of the Yayoi people from the Korean Peninsula [26]. Finally around 5000 years ago, H. pyllori (hspMaori) accompanied several subgroup of the Austronesia language family spread from Taiwan through the Pacific [14] included several islands in east Indonesia [27] into Melanesia and Polynesia.
3 Virulence Factors for Tracking Human Migration
The relationships between MLST and virulence factors were reported [28, 29]. The phylogeny of most cag pathogenicity island (PAI) genes, an approximately 40-kilobase pair region that is thought to have been incorporated into the H. pylori genome by horizontal transfer from an unknown source [30], was similar to that of MLST, indicating that cag PAI was probably acquired only once by H. pylori, and its genetic diversity reflects the isolation by distance which has shaped this bacterial species since modern humans migrated out of Africa [29]. The cagA gene which encodes a highly immunogenic protein (CagA) is located at one end of the cag PAI. The cag PAI encodes a type IV secretion system, through which CagA is delivered into host cells [31–33]. After delivery into gastric epithelial cells, CagA is tyrosine phosphorylated at Glu-Pro-Ile-Tyr-Ala (EPIYA) motifs located in the 3′ region of the cagA gene [34]. Supporting that H. pylori mirrors the human pattern of increased genetic distance and decreased diversity, our group has reported that the structure of the 3′ region of the cagA gene varies between strains from East Asian and Western countries [9, 12, 35, 36]. In East Asian strains, two types of repeats are found: 57 bp repeats followed by 162 bp repeats (East Asian-type cagA). Western strains have similar 57 bp repeats; however, they are followed by a repeat region consisting of 102 bp repeats, which are completely different from those of East Asian strains (Western-type cagA). Previous reports also show that the structure of the 5′ region of the cagA gene varies between strains from East Asian and Western countries [12, 37]. East Asian-type cagA is only observed in H. pylori isolates from the East Asian population, whereas Western-type cagA is widely distributed among isolates from European, South and Central Asian, North and South American, and African populations [12]. Almost all H. pylori isolates from East Asia possess the cagA gene, whereas approximately 20–40 % of isolates from Europe and Africa are cagA negative. Thailand is at the cultural crossroads between East and South Asia and, indeed, approximately half of the strains in Thailand possess East Asian-type cagA, whereas others possess Western-type cagA [38]. Interestingly Western-type cagA detected in strains from Okinawa (J-Western-type cagA) formed a different cluster compared to the original Western-type cagA [39]. The pre-EPIYA region of cagA also shows geographic divergence [40].
Most strains isolated from East Asia have a 39-bp deletion, but this deletion was absent in most strains from Western countries. On the other hand, an 18-bp deletion was common in Vietnamese strains. In addition, we found that the frequencies of the EPIYT and ESIYT motifs are relatively high among the sequences of the Okinawa strains [41]. Amerindian-type cagA from part of Machiguenga-speaking residents of the Shimaa village in the remote Peruvian Amazon (AM-I) also contained ESIYT motifs, which supports the possibility that these populations share the same origin [42]. A recent study revealed the recombination processes of cagA [43]. Interestingly, the left half of the EPIYA-D segment of East Asian-type cagA was derived from the Western-type EPIYA, with the Amerindian-type EPIYA as intermediate, through rearrangement of specific sequences within the gene. J-Western type EPIYA is phylogenetically located between the Western-type EPIYA and Amerindian-type EPIYA. This finding suggests that the original H. pylori strain had a Western-type cagA sequence. Subsequently, they evolved to the J-Western-type cagA, to the Amerindian-type cagA, and then to the East Asian-type cagA.
The right end of the cag PAI has been divided into five subtypes according to deletion, insertion, and substitution motifs [44]. Type I is most common in isolates from ethnic European groups and from Africa, type II is predominant in those from East Asia, and type III is predominant in isolates from South Asia [12, 44]. Type IV is very rare and, therefore, has not been assigned to a specific geographical area. Type V is found in a few strains from Calcutta, India [12, 44]. Interestingly, our report showed that type V was present in 10 % of isolates from patients of Thailand, and the ratio was especially high in strains obtained from ethnic Thai (21 %) [38]. The presence of this genotype in Thailand suggests that it migrated to the east of Calcutta. Overall, these data might show that transmission of specific genotypes remains conserved within ethnic groups for at least several generations.
On the other hand, the overall topology of the vacA tree did not always match with that of MLST [28]. Furthermore, rooting the vacA tree with out-group sequences from the closely related H. acinonychis revealed that the ancestry of vacA is different from the African origin. VacA is a vacuolating cytotoxin that induces cytoplasmic vacuoles in various eukaryotic cells. Unlike the case of the cagA gene, all H. pylori strains carry a functional vacA gene. However, there is considerable variation in vacuolating activities among strains [45, 46], primarily as a result of differences in the vacA gene structure at the signal region (s1 and s2) and the middle region (m1 and m2) [47]. Interestingly, the vacA s1 genotype is closely correlated with the presence of the cagA gene [8, 48, 49]. The vacA gene may comprise any combination of signal and middle-region types, although the s2/m1 combination is rare [47, 50]. All East Asian H. pylori strains are of the vacA s1 type [8, 12]. Within East Asian countries, the m1 type is predominant in Japan and Korea, whereas the prevalence of m2 types gradually increases in southern parts of East Asia (Vietnam, Hong Kong) [12]. The vacA s1 type is subdivided into s1a, s1b, and s1c [37, 47], and the m1 type is subdivided into m1a, m1b, and m1c [51]. The vacA s1c and m1b types are typical of H. pylori from East Asia (i.e., more than 95 % of s1 and m1) and the s1a and m1c types are common in South Asia (i.e., approximately 85 % of s1 and nearly 100 % of m1) [12, 52]. The vacA m1c genotype is also found in strains from Central Asia (ethnic Kazakhs) [12]. The m1a type is typical of Africans and ethnic Europeans (i.e., nearly 100 % of m1) [12, 37]. Both the s1a and s1b types are common in ethnic European strains, and s1b types are especially common in strains from the Iberian Peninsula and Latin America (i.e., approximately 85 % of s1) [37, 53]. The s1b type is also predominant in Africa (i.e., approximately 90 % of s1) [50, 53]. The H. pylori genotypes circulating among ethnic groups (Blacks, White Hispanics, Whites, and Vietnamese) living in the same region (Houston, Texas, USA) [54] have been examined by our group. According to ethnicity genotypes, the most common were the following: Blacks, s1b-m1; Hispanics, s1b-m1; Whites, s1a-m2; and Vietnamese, s1c-m2, which completely overlap with the predominant genotypes of Africa, the Iberian Peninsula, Northern and Eastern Europe, and Vietnam, respectively. In Thailand, the predominant vacA genotypes among s1-m1 strains are s1a-m1c in ethnic Thai people and s1b-m1b in ethnic Chinese people, which are the same as the predominant genotypes of South Asia and East Asia, respectively [38].
By combining the cagA, cag right-end junction, and vacA genotypes of more than 1000 H. pylori strains collected from East Asia, Southeast Asia, South Asia, Central Asia, Europe, Africa, North America, and South America [12, 38], four major groups (East Asia type, South/Central Asia type, Iberian/Africa type, and Europe type) can be defined according to geographical associations (Table 2.2). In these groups, cagA-negative and/or vacA m2 genotypes are not taken into account, but we can predict the geographical origins of each group using available genotypes (i.e., strains with cagA negative, but vacA s1a-m1a is predicted to be of the Europe type). Overall, the genotype of the virulence genes is important, not only as a tool to track human migration but also for epidemiological studies of H. pylori-related gastroduodenal diseases, especially in areas where multiple genotypes coexist (e.g., virulent East Asian type and less virulent South/Central Asian type in Thailand).
The genotypes of the virulence genes have provided important information about human migration to the Americas. The Americas were populated by humans of East Asian ancestry approximately 15,000 years ago. Over the last 500 years, Europeans and Africans have come to the Americas, leading to an increasing Mestizo (mixture of Amerindian and European ancestry) population. Our group has discovered that approximately 25 % of the H. pylori isolates from Native Colombians and Native Alaskans possess novel vacA and/or cagA structures that are similar, but not identical, to structures from East Asia (i.e., vacA s1c-m1b-like, East Asian-like cagA) [12]. Native Venezuelans are also reported to have a high frequency of the vacA s1c genotype [55]. These data confirm that H. pylori accompanied humans when they crossed the Bering Strait from Asia to the New World. Importantly, none of the H. pylori strains from Mestizo populations possess East Asian-like genotypes. Sequence analysis of H. pylori genomes has shown that East Asian-like Amerindian strains are the least genetically diverse, probably because of a genetic bottleneck, whereas European strains are the most diverse among Amerindian, European, African, and East Asian strains [56]. If diversity is important for the success of H. pylori colonization, the East Asian-like Amerindian strains may lack the needed diversity to compete with the diverse H. pylori population brought to the New World by non-Amerindian hosts and has therefore disappeared.
4 Genome-Wide for Evolutionary Study
Analysis of MLST data and virulence factors revealed much information about the pathogenicity and genealogy of H. pylori; however, these approaches focus on a small number of genes and may miss information conveyed by the rest of the genome. Genome-wide analyses using DNA microarray or whole-genome sequencing technology give a broad view on the genome of H. pylori.
Microarray analysis provides comprehensive information about gene contents of different strains and helps identify strain-specific genes as well as core genes shared by multiple strains. Salama et al. examined the genomic content of 15 clinical isolates using a whole-genome DNA microarray and defined 1281 genes as functional core genes [57]. They identified candidates of virulence genes on the basis of coinheritance with the cag PAI. A similar approach was used to elucidate the genomic diversity of isolates obtained from clinical patients in China [58]. The whole-genome sequencing technology is another powerful tool to study the evolution and pathogenicity of H. pylori.
Since the first release of the whole genome of strain 26,695 [59], the sequences of more than 20 genomes were determined by Sanger sequencing or the massively parallel sequencing technology. Accumulation of whole-genome data enables extensive sequence analyses of H. pylori strains. About 1200 core genes were identified by comparison of peptic ulcer strain P12 and six other H. pylori genomes, which were in agreement with preceding studies [60]. The authors found that the P12 genome contains three plasticity zones and that one of them is capable of self-excision and horizontal transfer by conjugation. Their result suggests that conjugative transfer of genomic islands may contribute to the genetic diversity of H. pylori. McClain et al. compared genome sequences of an isolate obtained from a patient with gastric cancer (strain 98–10) and an isolate from a patient with gastric ulcer (strain B128) [61]. Strain 98–10 was found to be closely related to East Asian strains, while strain B128 was related to European strains. They determined strain-specific genes of strain 98–10 as candidate genes associated with gastric cancer. East Asian strains are known for their stronger carcinogenicity compared to strains of other areas. Kawai et al. investigated the evolution of East Asian strains using 20 whole genomes of Japanese, Korean, Amerindian, European, and West African strains [62]. Phylogenetic analysis revealed a greater divergence between the East Asian strains and the European strains in genes related to virulence factors, outer membrane proteins, and lipopolysaccharide synthesis enzymes. They examined positively selected amino acid changes and mapped the identified residues on CagA, VacA, HomC, SotB, and MiaA proteins.
Currently, we took advantage of next-generation sequencers to read genomic sequences of more than 40 H. pylori strains mainly from Asian populations and attempted de novo assembly (unpublished observation). Although we cannot determine the whole genomes yet, we could construct a substantial size of contigs and predicted 1200–1500 genes for each strain. Using these data, we determined orthologous genes among our samples and strains whose whole genomes were released into public databases. A phylogenetic tree constructed by concatenated sequences of the orthologous genes showed more reliable results than a phylogenetic tree constructed by using MLST data. Compared with the tree based on MLST data, the tree constructed by using concatenated genes showed better branching with higher bootstrap values between hpEurope and hpAsia2, as well as between hspEAsia and hspAmerind. Data obtained by using the massively parallel sequencing technology provide valuable information on the genealogy of H. pylori strains, as well as on candidates of drug resistance genes and new virulence factors.
5 Conclusion
H. pylori typing is very useful as a tool for tracking human migrations. Serial studies of large numbers of H. pylori strains from all over the world, including strains isolated from aboriginal populations, have shown that MLST analysis of H. pylori sequences provides more detailed information on human migrations than does human genetic analysis. However, there are still a number of untapped areas in the world, including a number of isolated aboriginal populations in Siberia, Mongolia, China, Indonesia, and Japan (Ainu tribe), and it will be interesting to study H. pylori strains isolated from these different groups. To date, subcategorization of East Asian strains (hspEAsia) has not been possible because of high homology among East Asian strains. However, the genotyping of virulence genes has shown that the vacA middle region can be useful for distinguishing strains of the northern parts of East Asian countries from those of the south. Genome-wide analyses using DNA microarray or whole-genome sequencing technology give a broad view on the genome of H. pylori. These methods may complete the weakness of MLST and virulence factors. In particular, next-generation sequencers enabled us to efficiently investigate not only the evolution of H. pylori, but also novel virulence factors and genomic changes related to drug resistance.
Abbreviations
- cagA :
-
Cytotoxin-associated gene
- MLST:
-
Multilocus sequence typing
References
Suerbaum S, Michetti P. Helicobacter pylori infection. N Engl J Med. 2002;347(15):1175–86.
Achtman M, Azuma T, Berg DE, Ito Y, Morelli G, Pan ZJ, Suerbaum S, Thompson SA, van der Ende A, van Doorn LJ. Recombination and clonal groupings within Helicobacter pylori from different geographical regions. Mol Microbiol. 1999;32(3):459–70.
Li WH, Sadler LA. Low nucleotide diversity in man. Genetics. 1991;129(2):513–23.
Suerbaum S, Josenhans C. Helicobacter pylori evolution and phenotypic diversification in a changing host. Nat Rev Microbiol. 2007;5(6):441–52.
Falush D, Wirth T, Linz B, Pritchard JK, Stephens M, Kidd M, Blaser MJ, Graham DY, Vacher S, Perez-Perez GI, et al. Traces of human migrations in Helicobacter pylori populations. Science. 2003;299(5612):1582–5.
Yamaoka Y. Mechanisms of disease: Helicobacter pylori virulence factors. Nat Rev Gastroenterol Hepatol. 2010;7(11):629–41.
Yamaoka Y, Ojo O, Fujimoto S, Odenbreit S, Haas R, Gutierrez O, El-Zimaity HM, Reddy R, Arnqvist A, Graham DY. Helicobacter pylori outer membrane proteins and gastroduodenal disease. Gut. 2006;55(6):775–81.
Yamaoka Y, Kodama T, Kita M, Imanishi J, Kashima K, Graham DY. Relationship of vacA genotypes of Helicobacter pylori to cagA status, cytotoxin production, and clinical outcome. Helicobacter. 1998;3(4):241–53.
Yamaoka Y, El-Zimaity HM, Gutierrez O, Figura N, Kim JG, Kodama T, Kashima K, Graham DY. Relationship between the cagA 3' repeat region of Helicobacter pylori, gastric histology, and susceptibility to low pH. Gastroenterology. 1999;117(2):342–9.
Yamaoka Y. Roles of Helicobacter pylori BabA in gastroduodenal pathogenesis. World J Gastroenterol. 2008;14(27):4265–72.
Jung SW, Sugimoto M, Graham DY. Yamaoka Y: homB status of Helicobacter pylori as a novel marker to distinguish gastric cancer from duodenal ulcer. J Clin Microbiol. 2009;47(10):3241–5.
Yamaoka Y, Orito E, Mizokami M, Gutierrez O, Saitou N, Kodama T, Osato MS, Kim JG, Ramirez FC, Mahachai V, et al. Helicobacter pylori in North and South America before Columbus. FEBS Lett. 2002;517(1–3):180–4.
Linz B, Balloux F, Moodley Y, Manica A, Liu H, Roumagnac P, Falush D, Stamer C, Prugnolle F, van der Merwe SW, et al. An African origin for the intimate association between humans and Helicobacter pylori. Nature. 2007;445(7130):915–8.
Moodley Y, Linz B, Yamaoka Y, Windsor HM, Breurec S, Wu JY, Maady A, Bernhoft S, Thiberge JM, Phuanukoonnon S, et al. The peopling of the Pacific from a bacterial perspective. Science. 2009;323(5913):527–30.
Wirth T, Wang X, Linz B, Novick RP, Lum JK, Blaser M, Morelli G, Falush D, Achtman M. Distinguishing human ethnic groups by means of sequences from Helicobacter pylori: lessons from Ladakh. Proc Natl Acad Sci U S A. 2004;101(14):4746–51.
Maiden MC, Bygraves JA, Feil E, Morelli G, Russell JE, Urwin R, Zhang Q, Zhou J, Zurth K, Caugant DA, et al. Multilocus sequence typing: a portable approach to the identification of clones within populations of pathogenic microorganisms. Proc Natl Acad Sci U S A. 1998;95(6):3140–5.
Covacci A, Telford JL, Del Giudice G, Parsonnet J, Rappuoli R. Helicobacter pylori virulence and genetic geography. Science. 1999;284(5418):1328–33.
Moodley Y, Linz B, Bond RP, Nieuwoudt M, Soodyall H, Schlebusch CM, Bernhoft S, Hale J, Suerbaum S, Mugisha L, et al. Age of the association between Helicobacter pylori and man. PLoS Pathog. 2012;8(5), e1002693.
Suzuki R, Shiota S, Yamaoka Y. Molecular epidemiology, population genetics, and pathogenic role of Helicobacter pylori. Infect Genet Evol. 2012;12(2):203–13.
Correa P, Piazuelo MB. Evolutionary history of the Helicobacter pylori genome: implications for gastric carcinogenesis. Gut Liver. 2012;6(1):21–8.
Devi SM, Ahmed I, Francalacci P, Hussain MA, Akhter Y, Alvi A, Sechi LA, Megraud F, Ahmed N. Ancestral European roots of Helicobacter pylori in India. BMC Genomics. 2007;8:184.
Tay CY, Mitchell H, Dong Q, Goh KL, Dawes IW, Lan R. Population structure of Helicobacter pylori among ethnic groups in Malaysia: recent acquisition of the bacterium by the Malay population. BMC Microbiol. 2009;9:126.
Latifi-Navid S, Ghorashi SA, Siavoshi F, Linz B, Massarrat S, Khegay T, Salmanian AH, Shayesteh AA, Masoodi M, Ghanadi K, et al. Ethnic and geographic differentiation of Helicobacter pylori within Iran. PLoS One. 2010;5(3), e9645.
Breurec S, Guillard B, Hem S, Brisse S, Dieye FB, Huerre M, Oung C, Raymond J, Tan TS, Thiberge JM, et al. Evolutionary history of Helicobacter pylori sequences reflect past human migrations in Southeast Asia. PLoS One. 2011;6(7), e22058.
Gressmann H, Linz B, Ghai R, Pleissner KP, Schlapbach R, Yamaoka Y, Kraft C, Suerbaum S, Meyer TF, Achtman M. Gain and loss of multiple genes during the evolution of Helicobacter pylori. PLoS Genet. 2005;1(4), e43.
Ishida T, Hinuma Y. The origin of Japanese HTLV-I. Nature. 1986;322(6079):504.
Miftahussurur M, Tuda J, Suzuki R, Kido Y, Kawamoto F, Matsuda M, Tantular IS, Pusarawati S, Nasronudin, Harijanto PN, et al. Extremely low Helicobacter pylori prevalence in North Sulawesi, Indonesia and identification of a Maori-tribe type strain: a cross sectional study. Gut Pathog. 2014;6(1):42.
Gangwer KA, Shaffer CL, Suerbaum S, Lacy DB, Cover TL, Bordenstein SR. Molecular evolution of the Helicobacter pylori vacuolating toxin gene vacA. J Bacteriol. 2010;192(23):6126–35.
Olbermann P, Josenhans C, Moodley Y, Uhr M, Stamer C, Vauterin M, Suerbaum S, Achtman M, Linz B. A global overview of the genetic and functional diversity in the Helicobacter pylori cag pathogenicity island. PLoS Genet. 2010;6(8), e1001069.
Censini S, Lange C, Xiang Z, Crabtree JE, Ghiara P, Borodovsky M, Rappuoli R, Covacci A. cag, a pathogenicity island of Helicobacter pylori, encodes type I-specific and disease-associated virulence factors. Proc Natl Acad Sci U S A. 1996;93(25):14648–53.
Asahi M, Azuma T, Ito S, Ito Y, Suto H, Nagai Y, Tsubokawa M, Tohyama Y, Maeda S, Omata M, et al. Helicobacter pylori CagA protein can be tyrosine phosphorylated in gastric epithelial cells. J Exp Med. 2000;191(4):593–602.
Odenbreit S, Puls J, Sedlmaier B, Gerland E, Fischer W, Haas R. Translocation of Helicobacter pylori CagA into gastric epithelial cells by type IV secretion. Science. 2000;287(5457):1497–500.
Segal ED, Cha J, Lo J, Falkow S, Tompkins LS. Altered states: involvement of phosphorylated CagA in the induction of host cellular growth changes by Helicobacter pylori. Proc Natl Acad Sci U S A. 1999;96(25):14559–64.
Backert S, Selbach M. Role of type IV secretion in Helicobacter pylori pathogenesis. Cell Microbiol. 2008;10(8):1573–81.
Yamaoka Y, Kodama T, Kashima K, Graham DY, Sepulveda AR. Variants of the 3′ region of the cagA gene in Helicobacter pylori isolates from patients with different H. pylori-associated diseases. J Clin Microbiol. 1998;36(8):2258–63.
Yamaoka Y, Osato MS, Sepulveda AR, Gutierrez O, Figura N, Kim JG, Kodama T, Kashima K, Graham DY. Molecular epidemiology of Helicobacter pylori: separation of H. pylori from East Asian and non-Asian countries. Epidemiol Infect. 2000;124(1):91–6.
van Doorn LJ, Figueiredo C, Sanna R, Blaser MJ, Quint WG. Distinct variants of Helicobacter pylori cagA are associated with vacA subtypes. J Clin Microbiol. 1999;37(7):2306–11.
Vilaichone RK, Mahachai V, Tumwasorn S, Wu JY, Graham DY, Yamaoka Y. Molecular epidemiology and outcome of Helicobacter pylori infection in Thailand: a cultural cross roads. Helicobacter. 2004;9(5):453–9.
Truong BX, Mai VT, Tanaka H, le Ly T, Thong TM, Hai HH, Van Long D, Furumatsu K, Yoshida M, Kutsumi H, et al. Diverse characteristics of the CagA gene of Helicobacter pylori strains collected from patients from southern Vietnam with gastric cancer and peptic ulcer. J Clin Microbiol. 2009;47(12):4021–8.
Uchida T, Nguyen LT, Takayama A, Okimoto T, Kodama M, Murakami K, Matsuhisa T, Trinh TD, Ta L, Ho DQ, et al. Analysis of virulence factors of Helicobacter pylori isolated from a Vietnamese population. BMC Microbiol. 2009;9:175.
Xia Y, Yamaoka Y, Zhu Q, Matha I, Gao X. A comprehensive sequence and disease correlation analyses for the C-terminal region of CagA protein of Helicobacter pylori. PLoS One. 2009;4(11), e7736.
Suzuki M, Kiga K, Kersulyte D, Cok J, Hooper CC, Mimuro H, Sanada T, Suzuki S, Oyama M, Kozuka-Hata H, et al. Attenuated CagA oncoprotein in Helicobacter pylori from Amerindians in Peruvian Amazon. J Biol Chem. 2011;286(34):29964–72.
Furuta Y, Yahara K, Hatakeyama M, Kobayashi I. Evolution of cagA oncogene of Helicobacter pylori through recombination. PLoS One. 2011;6(8), e23499.
Kersulyte D, Mukhopadhyay AK, Velapatino B, Su W, Pan Z, Garcia C, Hernandez V, Valdez Y, Mistry RS, Gilman RH, et al. Differences in genotypes of Helicobacter pylori from different human populations. J Bacteriol. 2000;182(11):3210–8.
Cover TL, Blaser MJ. Purification and characterization of the vacuolating toxin from Helicobacter pylori. J Biol Chem. 1992;267(15):10570–5.
Leunk RD. Production of a cytotoxin by Helicobacter pylori. Rev Infect Dis. 1991;13 Suppl 8:S686–9.
Atherton JC, Cao P, Peek Jr RM, Tummuru MK, Blaser MJ, Cover TL. Mosaicism in vacuolating cytotoxin alleles of Helicobacter pylori. Association of specific vacA types with cytotoxin production and peptic ulceration. J Biol Chem. 1995;270(30):17771–7.
Atherton JC, Peek Jr RM, Tham KT, Cover TL, Blaser MJ. Clinical and pathological importance of heterogeneity in vacA, the vacuolating cytotoxin gene of Helicobacter pylori. Gastroenterology. 1997;112(1):92–9.
Atherton JC. The pathogenesis of Helicobacter pylori-induced gastro-duodenal diseases. Annu Rev Pathol. 2006;1:63–96.
Letley DP, Lastovica A, Louw JA, Hawkey CJ, Atherton JC. Allelic diversity of the Helicobacter pylori vacuolating cytotoxin gene in South Africa: rarity of the vacA s1a genotype and natural occurrence of an s2/m1 allele. J Clin Microbiol. 1999;37(4):1203–5.
Mukhopadhyay AK, Kersulyte D, Jeong JY, Datta S, Ito Y, Chowdhury A, Chowdhury S, Santra A, Bhattacharya SK, Azuma T, et al. Distinctiveness of genotypes of Helicobacter pylori in Calcutta, India. J Bacteriol. 2000;182(11):3219–27.
Hisatsune J, Nakayama M, Isomoto H, Kurazono H, Mukaida N, Mukhopadhyay AK, Azuma T, Yamaoka Y, Sap J, Yamasaki E, et al. Molecular characterization of Helicobacter pylori VacA induction of IL-8 in U937 cells reveals a prominent role for p38MAPK in activating transcription factor-2, cAMP response element binding protein, and NF-kappaB activation. J Immunol. 2008;180(7):5017–27.
Sugimoto M, Yamaoka Y. The association of vacA genotype and Helicobacter pylori-related disease in Latin American and African populations. Clin Microbiol Infect. 2009;15(9):835–42.
Yamaoka Y, Malaty HM, Osato MS, Graham DY. Conservation of Helicobacter pylori genotypes in different ethnic groups in Houston, Texas. J Infect Dis. 2000;181(6):2083–6.
Ghose C, Perez-Perez GI, Dominguez-Bello MG, Pride DT, Bravi CM, Blaser MJ. East Asian genotypes of Helicobacter pylori strains in Amerindians provide evidence for its ancient human carriage. Proc Natl Acad Sci U S A. 2002;99(23):15107–11.
Dominguez-Bello MG, Perez ME, Bortolini MC, Salzano FM, Pericchi LR, Zambrano-Guzman O, Linz B. Amerindian Helicobacter pylori strains go extinct, as european strains expand their host range. PLoS One. 2008;3(10), e3307.
Salama N, Guillemin K, McDaniel TK, Sherlock G, Tompkins L, Falkow S. A whole-genome microarray reveals genetic diversity among Helicobacter pylori strains. Proc Natl Acad Sci U S A. 2000;97(26):14668–73.
Han YH, Liu WZ, Shi YZ, Lu LQ, Xiao S, Zhang QH, Zhao GP. Comparative genomics profiling of clinical isolates of Helicobacter pylori in Chinese populations using DNA microarray. J Microbiol. 2007;45(1):21–8.
Tomb JF, White O, Kerlavage AR, Clayton RA, Sutton GG, Fleischmann RD, Ketchum KA, Klenk HP, Gill S, Dougherty BA, et al. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature. 1997;388(6642):539–47.
Fischer W, Windhager L, Rohrer S, Zeiller M, Karnholz A, Hoffmann R, Zimmer R, Haas R. Strain-specific genes of Helicobacter pylori: genome evolution driven by a novel type IV secretion system and genomic island transfer. Nucleic Acids Res. 2010;38(18):6089–101.
McClain MS, Shaffer CL, Israel DA, Peek Jr RM, Cover TL. Genome sequence analysis of Helicobacter pylori strains associated with gastric ulceration and gastric cancer. BMC Genomics. 2009;10:3.
Kawai M, Furuta Y, Yahara K, Tsuru T, Oshima K, Handa N, Takahashi N, Yoshida M, Azuma T, Hattori M, et al. Evolution in an oncogenic bacterial species with extreme genome plasticity: Helicobacter pylori East Asian genomes. BMC Microbiol. 2011;11:104.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Japan
About this chapter
Cite this chapter
Miftahussurur, M., Yamaoka, Y. (2016). Human Migration. In: Suzuki, H., Warren, R., Marshall, B. (eds) Helicobacter pylori. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55705-0_2
Download citation
DOI: https://doi.org/10.1007/978-4-431-55705-0_2
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55704-3
Online ISBN: 978-4-431-55705-0
eBook Packages: MedicineMedicine (R0)