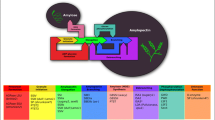
Abstract
Plants have developed two distinct starch biosynthetic systems composed of over 30 kinds of enzymatic reaction network in photosynthetic and non-photosynthetic cells. Higher plants have also evolved a process in which cells can accumulate huge amounts of starch as granules inside the amyloplast of the reserve organs. Primarily the coordinated expression of several sets of distinct isozymes with specific enzymatic properties enables plant cells to synthesize starch with distinct fine structure and form the starch granules with specific semicrystalline structure, granular morphology, and physicochemical properties in plastids. This chapter overviews the current status of our understanding of metabolic regulation of starch biosynthesis in reserve organs by focusing on functions of individual isozymes examined by numerous in vivo and in vitro studies that have been performed to reveal how individual isozymes contribute to the synthesis of the reserve starch. The results raised the high possibility that at least some isozymes have multiple functions in starch biosynthesis under different conditions depending on the presence of various glucans and interaction/association with other enzymes. The features of starch biosynthetic process in plant tissues are also discussed with emphasis on the biochemical mechanism(s) underlying the coordinate actions among various enzymes.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Amylopectin
- Amyloplast
- Assimilatory starch
- Chloroplast
- Cluster structure
- Isoamylase
- Phosphorylase
- Pullulanase
- Reserve starch
- Starch branching enzyme
- Starch debranching enzyme
- Starch synthase
1 Introduction
Starch is stored in plastids of essentially all plant tissues and consumed as both energy and carbon source when required. During photosynthesis plants accumulate storage compounds mainly in the form of starch and sucrose from CO2 and water. CO2 is first incorporated into the phosphorylated intermediates in the Calvin-Benson cycle. Starch and sucrose are synthesized from fructose 6-P (F6P) and triose-P such as dihydroxyacetone-P or 3-phosphoglycerate (PGA), the Calvin-Benson cycle intermediates, in the chloroplast and cytosol, respectively. The phosphorylated intermediates are also used for the biosynthesis of other compounds such as proteins, lipids, nucleic acids, and other polysaccharides. ATP and NAD(P)H needed for these compounds including starch and sucrose are basically provided from the photophosphorylation system and the light-dependent electron transport system of photosynthesis, respectively. Thus, the balance (ratio) of inorganic phosphate (Pi)/total P (inorganic plus organic P), ATP/ADP, and NAD(P)H/NAD(P)+ in each cellular compartment, such as chloroplast and cytosol, is fundamentally important to maintain high rates of photosynthesis and homeostasis of whole cells. The synthesis of starch and sucrose plays an important role in photosynthetic CO2-fixation activity by returning Pi to the photophosphorylation system and the Calvin-Benson cycle. Since sucrose is synthesized in the cytosol, the released Pi is returned to chloroplasts by the triose-P/Pi translocator (TPT) located in the chloroplast inner envelope when carrying triose-P/PGA into the cytoplasm, as shown in Fig. 5.1a. The photosynthetic reactions must run very quickly to support all the energy-requiring physiological and biochemical events in plants under the light condition. Therefore, the carbon flow from the Calvin-Benson cycle to the starch and sucrose synthesis forms the primary metabolic process in plant cells.
Schematic representation of the starch biosynthetic pathway and the related metabolism in photosynthetic and non-photosynthetic tissues. (a) The synthesis of assimilatory starch in the leaf. (b) The synthesis of reserve starch in storage tissues. (c) The starch synthetic pathway from G1P. Abbreviations: BT1 Brittle-1 protein (ADPglucose transporter), TPT triose phosphate/Pi translocator, GPT G6P/Pi translocator, G1PT putative G1P transporter (Fettke et al. 2010b)
As shown in Fig. 5.1c, the first step for the starch biosynthesis in green plants is the formation of ADPglucose, a glucose donor, from glucose 1-phosphate (G1P) and ATP as catalyzed by ADPglucose pyrophosphorylase (AGPase, EC 2.7.7.27) according to the following equation:
This step needs ATP and releases inorganic pyrophosphate (PPi), which is quickly decomposed into two molar equivalents of Pi by a high activity of inorganic pyrophosphatase in the stroma. Thus, the AGPase reaction actually proceeds in the ADPglucose synthetic direction, although the reaction is reversible in nature. The synthesis of starch is supported by chain elongation by transferring glucose moieties derived from ADPglucose to the nonreducing end of a glucan primer according to the equation catalyzed by starch synthase:
where G n and G n + 1 denote glucans having number of glucose residues with n and n + 1, respectively.
The amylopectin chains are elongated by a soluble form of starch synthase (SS, EC 2.4.1.21), whereas the amylose chain elongation is catalyzed by a starch granule-bound form of starch synthase, GBSS (Fig. 5.1c).
The α-1,6-glucosidic linkage is then introduced by starch branching enzyme (BE, EC 2.4.1.18). Finally, the fine structure of amylopectin is achieved via the trimming action (removal of unnecessary branches) of starch debranching enzymes (DBE).
In this way, amylopectin is basically synthesized by coordinate actions of three classes of enzymes, SS, BE, and DBE, whereas amylose is mainly synthesized by GBSS. However, the regulation of the starch synthesis is complex because each class of the enzymes has multiple isozymes with distinct properties.
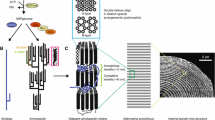
The amylopectin and amylose molecules are densely packed into the starch granules having a high specific gravity up to 1.6 (Rundle et al. 1944), and the fine structure of amylopectin is responsible for the semicrystalline nature of starch granules. It is evident that if normal amylopectin fine structure is disturbed, the morphology and physicochemical properties of starch granules are altered or sometimes the granular structure is lost. The distinct structure of amylopectin is frequently described as “cluster structure,” “bimodal chain-length distribution structure,” or “asymmetric branching structure” (Hizukuri 1986; Thompson 2000; Pérez and Bertoft 2010, also see Chap. 1 in this volume). This means that the branches of amylopectin molecules are formed in at least two regions referred to as the amorphous lamella and the crystalline lamella, keeping a constant length of a single cluster (about 9 nm) in all plant sources (Jenkins et al. 1993, also see Chap. 1 in this volume) (Fig. 5.2).
Schematic representation of the cluster structure of amylopectin and the synthesis of cluster chains and branches in cereal endosperm via concerted elongation reactions by SSI, SSIIa, and SSIIIa isozymes and by branching reactions by BEI and BEIIb isozymes. The basic structure of amylopectin cluster composed of the repeated amorphous lamellae and the crystalline lamellae. The figure also presents the lengths of amylopectin chains determined by SS isozymes. SSI, SSIIa, and SSIIIa play distinct roles in the synthesis of very short chains, short and intermediate A- and B1-chains, and long B1- and B2-3-chains, respectively, and that branches localized in the amorphous lamellae and the crystalline lamellae and/or the interregion between both lamellae are mainly formed by BEI and BEIIb, respectively. The figure is drawn based on the results with rice plants and enzymes (Nakamura 2002, 2014)
Starch biosynthesis in reserve organs is characterized by a high rate of starch production from sucrose and an efficient formation of the starch granules including amylopectin and amylose. At the initial developmental stage of reserve organs, the initial starch synthetic process is critical, while during the maturing stages when starch is synthesized at the maximum rate, the primary starch biosynthetic process which can be referred to as the starch amplification process (or reproduction process) must be predominantly operating. This chapter mainly deals with the amplification process where the homogeneous glucan molecules are very rapidly reproduced to increase the starch amount, whereas the initiation process is described in Chap. 9 (also see D’Hulst and Merida 2012).
2 Comparison of Starch Biosynthetic Pathway Between Photosynthetic and Non-photosynthetic Tissues
Plants have developed the energy storage system in which a great amount of starch granules can be accumulated in reserve tissue. These starch granules are usually referred to as the reserve starch while those synthesized in chloroplasts of photosynthetic cells are called the assimilatory starch. The physiological role of both types of starch is different, i.e., the reserve starch is stored for the long period and used for the next generation while the assimilatory starch is temporarily accumulated during the day but is consumed during the following night period to support the biological activities in plant tissues. The importance of the different functions of both starches seems to be supported by the structural differences between them.
The whole metabolic pathway and events in the starch biosynthetic process as well as other related metabolic pathways in higher plants are different between assimilatory starch produced in chloroplasts (Fig. 5.1a) and reserve starch produced in amyloplasts (Fig. 5.1b). First, the pathway of the synthesis of ADPglucose, a glucose donor for starch synthesis by serving the substrate for SS and GBSS, totally differs between chloroplasts and amyloplasts. In chloroplasts, F6P, a member of the Calvin-Benson cycle, is first converted to glucose 6-P (G6P) and then to G1P by a plastidial glucose-phosphate isomerase (PGI) and phosphoglucomutase (PGM), respectively. Subsequently, G1P is metabolized to ADPglucose by AGPase (Fig. 5.1a). In amyloplasts, however, the starting metabolite for starch synthesis is sucrose which is translocated to reserve tissues from leaves through phloem (Fig. 5.1b). Sucrose enters the cytosol via the sucrose transporter, and the first step for sucrose degradation is catalyzed by sucrose synthase to form UDPglucose and fructose. UDPglucose is then converted to G1P by UDPglucose pyrophosphorylase (UGPase). In cereals, G1P is mainly used as substrate for ADPglucose synthesizing reaction catalyzed by a cytosolic AGPase (Denyer et al. 1996; see also Chap. 11), and ADPglucose is then transported into amyloplasts via the ADPglucose transporter, sometimes referred to as the Brittle1 protein (Sullivan 1995; Shannon et al. 1998). In other reserve organs such as potato tubers and pea embryo, most of the G1P are transferred to G6P, which is imported into amyloplasts by the G6P translocator. G6P is metabolized to ADPglucose via G1P by both plastidial PGM and AGPase activities in amyloplasts. It is also possible that G1P is partially directly moved into amyloplasts by the putative G1P transporter (Fettke et al. 2010b). The translocated sucrose is also, at least partly, degraded by cellulose-bound invertase into glucose and fructose. After both compounds are transported into the cytosol by the hexose transporter, they are converted to G6P and F6P by hexokinase. F6P is converted into G6P by cytosolic PGI and, subsequently, to G1P to form ADPglucose by the actions of PGM and AGPase in the cereal cytoplasm or directly enters into amyloplasts in other species. During the early developmental stage of the cereal endosperm, it is considered that G6P moves into amyloplasts to some extent to be metabolized to ADPglucose by plastidial PGM and AGPase.
Second, the supply of ATP is different between chloroplast and amyloplast although ATP is essential for the AGPase reaction in both. In chloroplasts ATP is supplied from the photophosphorylation system functioning during photosynthesis. On the other hand, in non-photosynthetic cells, ATP is provided from the oxidative phosphorylation system operating in mitochondria during dark respiration (Fig. 5.1b). The generated ATP is provided to cytosolic AGPase in the cytoplasm or plastidial AGPase after the import into amyloplasts via the nucleoside triphosphate transporter.
Third, the types of translocators/transporters functioning in the non-photosynthetic cells are different from those in photosynthetic cells. In non-photosynthetic cells, many metabolic processes localized in different cellular compartments and organelles play distinct roles in starch synthesis. Thus, many translocators/transporters are needed to provide enzymes with metabolites and cofactors required for these actions and activities, as explained above. In contrast, chloroplasts in photosynthetic cells contain almost all of enzymes and cofactors for the whole starch biosynthetic processes. Thus, these processes can be independently operative during photosynthesis. More precisely, chloroplasts are specialized to produce two major photosynthates, starch and sucrose, and triose-P and/or PGA, the major primary CO2-fixation products, are imported into the cytosol by the triose-P translocator (TPT) to produce sucrose in the cytosol (Fig. 5.1a). To facilitate the cytosolic sucrose synthesis, TPT transports inorganic phosphate (Pi) from cytosol into chloroplasts in exchange for triose-P and/or PGA as the counterpart. It is noted that translocators/transporters in non-photosynthetic reserve cells are different from those in photosynthetic cells and must have been developed during the process of plant evolution (Weber and Linka 2011).
Fourth, the regulatory mechanism for enzymatic actions and activities is different between reserve and assimilatory starch syntheses. In chloroplasts, starch synthesis is closely related to photosynthesis; the regulatory properties of AGPase and possibly other starch synthetic enzymes too, which are present in photosynthetic cells, are different from those in non-photosynthetic cells (see in details in Chap. 11).
Fifth, starch structure including the fine structure of amylopectin and starch granular structure differs between reserve and assimilatory starches.
Sixth, isozymes are differently expressed in both tissues, in addition to different levels of enzymatic activities between tissues (Nakamura et al. 1989). For example, some isozymes such as SSIIa, GBSSI, BEIIb, and ISA1 are almost exclusively or preferentially expressed in developing rice endosperm while others such as SSIIb, GBSSII, BEIIa, and ISA3 in the leaf (Ohdan et al. 2005), as presented in Fig. 5.3.
Transcript levels of genes encoding starch metabolism in endosperm and leaf of rice (Data are from Ohdan et al. 2005)
3 Functions and Properties of Enzymes Involved in the Biosynthesis of Reserve Starch
The contribution of each isozyme of starch synthetic enzymes to the synthesis of amylopectin and amylose has been investigated by using a variety of plant sources and species, purified enzyme preparations, mutants, and transformants generated in at least the last three decades. The accumulated results provide us with the most reliable and invaluable fundamental information and materials. First, the presence and the relative activities or protein amounts of multiple isozymes and isoforms have been demonstrated. Second, their enzymatic properties such as reactivity toward various glucans including branched glucans (amylopectin and glycogen), amylose, and malto-oligosaccharides (MOS); effects of chemicals, metabolites, and temperature on activities; and association with other enzymes or starch granules have been elucidated. Third, the genes encoding each isozyme of starch synthetic enzymes were isolated, and structural features of individual proteins and their expression patterns in various tissues and the developmental stages were determined. Fourth, mutants defective in the target genes were isolated and used to characterize the functions of the genes. Fifth, the transgenic lines in which the target gene functions were modified were prepared, and various starch-related phenotypes were analyzed. The use of the transformants was of great interest to prove the function of the gene by detecting whether the mutated phenotype reverted to the wild-type one and to test the possibility of how and to what extent structural and functional properties of starch granules can be modified by bioengineering of the gene. Sixth, in vitro studies with the purified preparations of the target isozymes provided direct information on their enzymatic properties, functions, and enzymatic reaction mechanisms. Since in in vitro experiments the influences of other functionally overlapping and/or interacting isozymes can be neglected, properties of the target isozymes can be directly examined. In addition, the reaction conditions including the use of the appropriate primer/substrate glucans can be selected. Kinetic parameters and effects of cofactors on the activity can be determined. Seventh, the currently developed “omics” analyses enabled us to have the whole scope of the roles of all enzymes operating in the same tissues under various physiological conditions (Smith et al. 2004; Ohdan et al. 2005). By this approach, we can scrutinize the contribution of the target gene/enzyme to starch biosynthesis, which might fluctuate by tissue-, developmental stage-, and diurnal-specific manners. The different phenomena occurring between plant species can be also compared at the molecular level.
All these results serve as invaluable bases of most complexities and variations in various aspects of regulation of starch biosynthesis.
3.1 The Biosynthesis of Amylopectin
3.1.1 Soluble Starch Synthase (SS)
SS elongates the preexisting α-1,4-glucosidic chains of amylopectin by adding glucose moiety from ADPglucose to the nonreducing end of the chains. This reaction is the only step to increase the number of glucose residues of the starch because no change occurs in glucose amount of the starch after the BE and DBE reactions. In amylopectin molecules, the length or degree of glucose polymerization (DP) of side chains should be strictly controlled, because otherwise double helices (Kainuma and French 1972) composed of adjacent chains having similar lengths are not uniformly arranged. How can this structure be achieved? The rate of starch biosynthesis is very rapid and the newly formed amylopectin clusters are immediately packed into the starch granules. Therefore, chain length should be controlled simultaneously as they are formed because trimming of chain length of individual chains afterward is actually impossible once they are formed. The other basic requirement for side chains of amylopectin is that their nonreducing part should be left non-branched so that the external segments are long enough to form double helices with DP ≧ 10 (Gidley and Bulpin 1987). During the synthesis of amylopectin side chains by SS, the chain branching reactions catalyzed by BE are restrained until the chains reach the length of DP 12, the minimal chain length required for BE reaction (Guan et al. 1997; Nakamura et al. 2010; Sawada et al. 2014), or longer. To satisfy all these requirements, plants possess multiple SS isozymes with distinct chain elongation properties for reserve starch biosynthesis.
3.1.1.1 Multiple SS Isozymes
Green plants have four types of SS isozymes, the SSI, SSII, SSIII, and SSIV types, while it is likely that some higher plants such as rice and potato have in addition the SSV type (see review by Fujita and Nakamura 2012). Since early 1970s, many reports have shown the occurrence of multiple SS isoforms in reserve organs and leaves of rice (Tanaka and Akazawa 1971), maize (Ozbun et al. 1971b; Tanaka and Akazawa 1971), potato (Hawker et al. 1972), and spinach (Ozbun et al. 1971a, 1972; Hawker et al. 1974), which were separated by DEAE column chromatography and distinguished from each other based on reactivities toward the glucan type and chemicals such as citrate, suggesting the involvement of coordinate actions of multiple SS isozymes in starch biosynthesis. Later, genome analysis indicated that higher plants have many SS genes, e.g., a total of nine genes (one SSI, three SSII, two SSIII, two SSIV, and one SSV) in rice plants (Hirose and Terao 2004; Ohdan et al. 2005).
3.1.1.2 Structures and Basic Enzymatic Properties
SS as well as GBSS belongs to the CAZy glucosyltransferase family 5 (GT5). Green plants have at least four types of SS and one type of GBSS, which are structurally and functionally distinct. These five enzymes have commonly a core sequence with an approximate size of 60 kDa and include two conserved regions referred to as the GT5 and GT1 domains which are separated by a linker region. In addition, they have the N-terminal extensions with different sizes – those of GBSS and SSIII/SSIV being the shortest and longest, respectively. The GT5 domain contains the highly conserved motif KXGGL considered to be responsible for the catalytic activity, while the N-terminal extension is likely to regulate catalytic activity, affecting binding affinity of SS for carbohydrates (Gao et al. 1998; Cao et al. 1999; Li et al. 2000; Leterrier et al. 2008; Schwarte et al. 2013).
The crystal structure of barley SSI with an omission of the N-terminal extension was revealed by Cuesta-Seijo et al. (2013), following the analysis of rice GBSSI (Momma and Fujimoto 2012). Structural similarities among plant SS were verified by the presence of the three distinct domains, i.e., the N-terminal domain, the central catalytic domain, and the C-terminal domain (Leterrier et al. 2008; Schwarte et al. 2013).
It is generally known that plant SS isozymes exhibit high activities toward various branched glucans including amylopectin and glycogen while the activities for linear MOS and amylose are very low (Edwards et al. 1999a; Damager et al. 2001; Imparl-Radosevich et al. 2003; Senoura et al. 2007; Nakamura et al. 2014). It is also well known that SSI and SSII are markedly activated by high concentrations (≧0.3 M) of citrate (Ozbun et al. 1971b; Hawker et al. 1974; Boyer and Preiss 1979; Pollock and Preiss 1980; Senoura et al. 2007; Nakamura et al. 2014) (for other properties of SS, see review by Fujita and Nakamura 2012).
3.1.1.3 Chain Preferences of SS Isozymes
The contributions of individual SS isozymes to starch synthesis, the fine structure of amylopectin, and physicochemical properties of starch granules have been extensively examined using SS mutants and transformants generated from a number of plant species (see review by Fujita and Nakamura 2012) and by in vitro studies with purified enzyme preparations and selected glucan primers.
Although SSI accounted for about 60–70 % of the SS activity in wild-type cereal endosperm and Arabidopsis leaves, the single ss1 mutation was likely to cause only a slight change in the starch-related phenotypes. It is, however, noted that amylopectin from rice ss1 mutant lines with no or varied levels of residual SSI activities in the endosperm had consistently reduced chains of DP 8–12 and enriched chains of DP 6–7 and DP 16–19, whereas changes in the proportion of long chains of DP ≧ 21 were very low or unchanged (Fujita et al. 2006) (Fig. 5.4b). The similar impact of suppression of SSI expression on amylopectin chain profile was reported in wheat endosperm (McMaugh et al. 2014). These results suggest that SSI plays a distinct role in the synthesis of chains of DP 8–12 from A-chains of DP 6–7 and the external segments of B-chains emerging from the nearest branch point. Delvallé et al. (2005) also found that the mutation of Arabidopsis SSI gene markedly reduced chains of DP 8–12 but increased chains of DP 17–20. These results are consistent with the view that SSI is involved in the synthesis of short external segments of amylopectin chains.
Changes in structure of amylopectin of rice mutants defective in starch synthase (SS) isozymes. (a) Amylopectin in endosperm from wild-type japonica rice cultivar Nipponbare (blue bars), as compared to phytoglycogen from sugary-1 mutant line (EM41) of rice (brown bars). (b) Amylopectin from ss1 mutant (Fujita et al. 2006). (c) Amylopectin from ss2a mutant (Nakamura et al. 2005a). (d) Amylopectin from ss3a mutant (Fujita et al. 2007)
It is well known that defective SSII gene function has profound effects on starch synthesis and amylopectin structure in reserve organs of many plant species. Craig et al. (1998) reported that mutations of SSII gene (the rugosus5 locus) of pea dramatically reduced the amount of intermediate chains and increased the amount of short chains. Later, Edwards et al. (1999b) and Lloyd et al. (1999) extensively analyzed the effects of antisense inhibition of either SSII or SSIII and both activity levels in potato tubers on the amylopectin chain lengths. The consequence of SSII inhibition was distinct because short chains of DP 6–13 greatly decreased while intermediate chains of DP 15–25 markedly increased. A series of studies by Umemoto et al. (1999, 2002) and Nakamura et al. (2002, 2005a) characterized the contribution of rice SSIIa to the fine structure of amylopectin. They showed that most of japonica-type rice varieties had enriched chains of DP 6–10 and depleted chains of DP about 12–24, compared with typical indica-type varieties although no significant difference was found in the proportion of B2- and B3-chains (Umemoto et al. 1999; Nakamura et al. 2002) (Fig. 5.4c) and that the japonica-type varieties had an inactive mutated ss2a gene while the indica-varieties had an active SSIIa gene (Nakamura et al. 2005a). The same trends in chain profiles for amylopectin were observed in the ss2a mutants from wheat (the sgp-1 mutant, Yamamori et al. 2000) and barley (the sex6 mutant, Morell et al. 2003). Similar results were reported in maize sugary-2 mutant endosperm (Takeda and Preiss 1993; Zhang et al. 2004), sweet potato cv. Quick sweet tuberous root (Katayama et al. 2002; Kitahara et al. 2005), and Arabidopsis leaf (Zhang et al. 2008). All these results support the conclusion that SSII(a) plays a distinct role in the synthesis of intermediate chains of amylopectin, and this role cannot be effectively supplemented by other SS isoforms (Fig. 5.5).
Pattern of changes in the cluster structure of amylopectin caused by different levels of key enzymes in starch biosynthesis. The fine structure of amylopectin is altered by different levels of SSIIa, BEIIb, and ISA1 activities in rice endosperm, keeping the distinct structural pattern of the cluster depending on the individual enzyme, and this is referred to as “cluster-world” (Nakamura et al. 2009; Nakamura 2014)
It is known that the dull1 mutation of maize caused by defective SSIIIa gene had a modified granular morphology and physicochemical properties (Mangelsdorf 1947; Davis et al. 1955; Gao et al. 1998, see also reviews by Shannon and Garwood 1984 and Shannon et al. 2007). Fontaine et al. (1993) and Ral et al. (2006) proposed a specific role of SSIII in the synthesis of intermediate and long chains of amylopectin in Chlamydomonas reinhardtii based on the chain-length analysis of the ss3 mutant, sta-3. Loss of SSIII in potato tuber resulted in changes in amylopectin and starch granule structure, and data suggested that SSIII is involved in the synthesis of long chains although the extent of difference in chain-length profile was too small to clearly evaluate the function of SSIII (Edwards et al. 1999b; Lloyd et al. 1999). Fujita et al. (2007) analyzed in details the amylopectin chain profiles of amylopectin in rice endosperm and found that the ss3a mutant had fewer chains of DP 6–9, DP 16–19, and DP 33–55 and more chains of DP 10–15 and DP 20–25 than the wild-type (Fig. 5.4d). They proposed a major role of SSIIIa in synthesizing long chains (B2- and B3-chains), judging from the observation that the increased pattern of long chains in the ss3a mutant was specific and the increase of very short chains (DP 6–9) was considered to be caused by the elevated level of SSI due to the pleiotropic effect of ss3a mutation. Recently, Lin et al. (2012) reported the fine structure of amylopectin in the maize dull1 mutants, similar to the rice ss3a mutants. The barley ss3a mutant (amo1) endosperm contained amylopectin with less short chains of DP 9–10 and more intermediate chains of DP 15–24 while no significant change was detected in long chains (Li et al. 2011).
The ss3(a) mutation frequently produced marked starch-related phenotypic changes such as the starch content, starch granule morphology, and the starch physicochemical properties, despite the absence of noticeable change in chain-length pattern. The results suggest that SSIII(a) plays an additional integral role in starch granule biosynthesis.
Although SSIV accounted for only a minor portion of the total SS activity, its function in starch biosynthesis was likely to be specific. Roldán et al. (2007) revealed that loss of SSIV reduced the number of starch granules per chloroplast in Arabidopsis leaves, suggesting that SSIV plays an essential part in the starch granule initiation. The specific role of SSIV in Arabidopsis was verified by further studies (Szydlowsky et al. 2009; Crumpton-Taylor et al. 2013). The observation that the ss4/ss3 double mutants completely lost starch granules suggests that the role of SSIV can be partially supported by SSIII, possibly due to some structural and functional similarities (Szydlowsky et al. 2009; 2011). Although studies on the contribution of SSIV to the starch biosynthesis in cereal endosperm are limited, recently Toyosawa et al. (2015) found that a single mutation of the SSIVa or SSIVb gene had minor impacts on starch phenotype in rice endosperm, whereas the ss4b/ss3a double mutants altered the polygonal starch granules into the spherical granules, although the number of granules per amyloplast and the starch content per seed were only slightly affected. The results suggest the possibility that the impact of SSIV depends on plant species (see details in Chaps. 6, 9, and 10).
Despite numerous in vivo studies on SS, only a few in vitro studies have been reported. Commuri and Keeling (2001) examined the chain-length specificities of maize SSI and determined the relationship between dissociation constant (Kd) of SSI for the external chain length of synthesized glycogen and the SSI activity. The catalytic activity evidently decreased with the increase in the average DP values of external chains and almost dropped to zero when DP exceeds about 11, these DP values being in accord with the increase of glucan affinity. They concluded that the activity of SSI decreases with the chain length of external chains due to its very high affinity for the longer chains. Recently, Nakamura et al. (2014) directly measured the chain length of the products formed by in vitro enzymatic reactions between rice SSI, SSIIa, or SSIIIa and amylopectin. It is noted that SSI specifically elongated very short chains of DP 6 and 7 of A-chains and external segments of B-chains to mainly the length of DP 8 in these chains, although they could be further elongated by the prolonged incubation. Analysis of the SSIIa and SSIIIa reaction products was consistent with in vivo results, suggesting that SSIIa and SSIIIa preferentially synthesize intermediate chains and intermediate/long chains, respectively.
3.1.1.4 The Roles of SS Isozymes in Amylopectin Biosynthesis
Undoubtedly, concerted reactions of individual SS isozymes are essential for construction of the organized structure of amylopectin with uniform size and shape of unit clusters. The chain-length distribution of amylopectin in reserve starch granules forms two distinct peaks: a large short- and intermediate-chain peak and a small long-chain peak with most abundant chains of approximately DP 10–14 and DP 35–45, respectively, sharply in contrast with the pattern of phytoglycogen or glycogen lacking the second peak (Fig. 5.4a). The lengths of the former intra-cluster chains (A- and B1-chains) and those of the latter inter-cluster chains (B2- and B3-chains) must be strictly controlled, otherwise the shape of the bimodal distribution would be distorted and the ratio of the two peaks must be changed. However, it is well known that these features are primarily conserved during development of the storage organ of each plant species. The chain length in itself is the result of the SS elongation reaction. BE indirectly affects the length by introducing branched linkages, which results in the reactivation of the long external chains to SS by decreasing the length of the external segment because each SS isozyme very specifically and sensitively recognizes the length of the external segment of substrate and product chain. The most likely explanation for requirement of multiple SS isozymes for the synthesis of amylopectin is that the branched chains are elongated by coordinate reactions of SSI, SSII, and SSIII isozymes with different chain-length preferences until the lengths reach the nonreducing end of the cluster.
Past biochemical, molecular, and genetic investigations supported the almost consistent view that properties of SSI, SSII, and SSIII are specific to forming very short chains of DP up to about 10, intermediate intra-cluster A- and B1-chains of DP up to about 24, and long inter- and intra-cluster B2-/B3-chains and B1-chains, respectively, of DP longer than about 20, respectively (Fujita and Nakamura 2012) (Fig. 5.2). The features of SS reaction properties are as follows: first, activities of SS are largely dependent on the chain length of the external segment of primer chains, being sharply in contrast with those of Pho and GBSS because both of them can synthesize very long chains such as amylose and amylopectin super-long chains. Second, SS shows a high activity toward branched glucan such as amylopectin and glycogen, whereas it has a very low activity toward linear glucan or MOS. In contrast, both Pho and GBSS are similarly reactive to either linear or branched glucans.
How can SS limit the length of amylopectin chains? Although no convincing evidence is available, several possibilities can be considered. First, as described above, Commuri and Keeling (2001) reported that catalytic ability of maize SSI decreased with the increase of external chain length in the range of DP 6–14 and then exponentially dropped to DP 21, due to the increase of its affinity for the longer chains. However, it remains to be resolved whether and to what extent the properties of SSII and SSIII are influenced by chain lengths. Second, the amylopectin chain increases the hydrophobic properties in aqueous medium with the increase in the chain length, as found in linear MOS, and this might inhibit the accessibility of SS to the nonreducing end of the primer chain. Third, since SS generally shows a higher catalytic action with branched glucans than linear glucans, as stated above, the actual substrate for SS might be the branched duplicate chains positioned in the same direction and having the similar length. As elongation of the single chain by SS progresses, the difference in chain length between adjacent chains will become larger, dramatically reducing the rate of chain elongation. All these properties might result in the cessation of further increase of the cluster size. It was found that rice SSIIa had a capacity for the synthesis of l-type amylopectin having longer A- and B1-chains in test tubes when it was incubated with s-type amylopectin having shorter cluster chains and prepared from a rice ss2a mutant japonica cultivar (Nakamura et al. 2005a) (Fig. 5.5), indicating that SSIIa is specialized to elongate intra-cluster chains by acting on short chains formed by BE reactions.
3.1.2 Starch Branching Enzyme (BE)
BE introduces the branch structure into α-glucan with α-1,6-glucosidic linkage by transferring the chain at its reducing end onto the acceptor chain. The transferred chain is derived from the donor chain by cleaving off its nonreducing part of the chain, and the remaining part of the chain is referred to as the residual chain (Borovsky et al. 1979) (Fig. 5.6). When BE uses an acceptor chain different from the donor chain, the reaction is referred to as the inter-chain BE reaction, whereas in the intra-chain reaction, BE utilizes the same single chain as both acceptor and donor chains. In both cases, the total number of chains increases while that of glucose residues is unchanged during the BE reaction.
Although glucose amount in the glucan during the BE reaction is unchanged, it plays an essential role in starch biosynthesis in terms of not only the glucan fine structure but also eventually in the catalytic activity of SS and the rate of starch production in plant tissues. The amylopectin cluster structure enables the macromolecule to form the semicrystalline granular structure so that plant cells can store the inert glucans for a long period. The branched structure also has numerous chains, serving for SS reaction as acceptors at their nonreducing ends, and this supports a high rate of starch production. The formation of a branch in the middle of linear long chain also affects the functionality of the chain. Because the linear α-1,4-chain segment increases its hydrophobic nature with the increase of DP, the long non-branched chain segment becomes insensitive to SS isozymes (Commuri and Keeling 2001). Thus, the branch formation by BE results in reactivation of long chains by accepting the transferred chains because SS can again act on them at their external segments.
3.1.2.1 Multiple BE Isozymes
Numerous past investigations reported that green plants have two types of BE isozymes, BEI and BEII, in various species such as maize (Boyer and Preiss 1978; Dang and Boyer 1988), rice (Nakamura et al. 1992a; Yamanouchi and Nakamura 1992; Mizuno et al. 1993), wheat (Morell et al. 1997; Rahman et al. 2001), barley (Sun et al. 1998), pea (Smith 1988; Burton et al. 1995), kidney bean (Nozaki et al. 2001; Hamada et al. 2001), potato (Larsson et al. 1996), spinach (Hawker et al. 1974), popular (Han et al. 2007b), and apple (Han et al. 2007a). BEII is known to be further functionally and structurally differentiated into BEIIa and BEIIb, encoded by different genes in cereals such as rice (Yamanouchi and Nakamura 1992; Mizuno et al. 1993), maize (Guan and Preiss 1993; Fisher et al. 1996), barley (Sun et al. 1998), wheat (Morell et al. 1997), and sorghum (Mutisya et al. 2003), whereas generally no such differentiation of BEIIb-type is found in dicot plants. Generally, BEIIb is specifically or mainly expressed in cereal endosperm, playing a distinct role in determining the fine structure of amylopectin while BEIIa is ubiquitously expressed in every tissue (see reviews by Nakamura 2002, 2014; Tetlow 2012). Thus, monocots have totally three functional BE isozymes in the endosperm and two isozymes in the other tissues such as leaf. On the other hand, most dicot tissues are likely to have only two BE isozymes. Although some dicots such as Arabidopsis and apple have multiple BEII-type isozymes (Han et al. 2007a), their function and structure were very similar and it is unlikely that they play different roles in amylopectin biosynthesis. The location and the clustering of the α-1,6-linkages is fundamentally important to the construction of the amylopectin structure, such that the formation of branches in amylopectin molecules by coordinated actions of BE isozymes must be precisely controlled. Considering this requirement, the limited number of BE isozymes expressed in plant tissues is rather surprising compared with many SS isozymes. How can only two or three isozymes suffice such requirements? The point might be further strengthened if one considers that plant BEs must have a common origin as that of glycogen BEs from eukaryotes such as animals and fungi, but not the bacterial BE origin (Patron and Keeling 2005; Deschamps et al. 2008).
The different expressions of multiple BE genes were examined in various plant species during development of reserve organs. It was generally seen that in both monocots and dicots, BEII-type gene(s) was (were) expressed from the early developmental stage of the reserve organ, while BEI-type gene was highly expressed at the latter developmental stage in both monocot seeds such as rice (Mizuno et al. 1993; Hirose and Terao 2004; Ohdan et al. 2005), maize (Gao et al. 1996), wheat (Morell et al. 1997), and barley (Sun et al. 1998) and dicot seeds such as pea (Burton et al. 1995) and kidney bean (Hamada et al. 2001). In cereal endosperm, BEIIa was found to be expressed from the very early stage, followed by a marked expression of BEIIb (Hirose and Terao 2004; Ohdan et al. 2005). The physiological significance of these expression patterns remains largely to be resolved in the future.
3.1.2.2 Structures and Basic Enzymatic Properties
BEs are members of the CAZy glycoside hydrolase family 13 (GH13), which is also referred to as the α-amylase family. Eukaryote BEs are further classified as subfamily 8, while almost all of bacterial BEs are members of subfamily 9 (Stam et al. 2006). Some thermophilic bacteria have other types of BEs belonging to the GH57 family (Murakami et al. 2006; Palomo et al. 2009). The GH13 BEs have three major domains, the A domain or the central (β/α)-barrel catalytic domain, the N-terminal domain, and the C-terminal domain, although the A domain sequence is highly conserved, whereas other two domains show a high variability, and this structural features were directly revealed by the X-ray crystallographic analysis of rice BEI (Noguchi et al. 2011; Chaen et al. 2012, also see review by Tetlow 2012 and many references therein).
Enzymatic properties of BEs have been elucidated. These include reactivity toward various glucans including branched glucans (amylopectin and glycogen), amylose, and malto-oligosaccharides (MOS) (Tetlow 2012); chemicals such as cyclodextrins (Vikso-Nielsen and Blennow 1998), phosphorylated metabolites (Morell et al. 1997), and citrate (Hamada et al. 2001, 2002); temperature dependence of activities (Takeda et al. 1993; Hamada et al. 2002; Ohdan et al. 2011; Sawada et al. 2013); and association with starch granules (Rahman et al. 1995; Mu-Forster et al. 1996; Hamada et al. 2001, 2002; Grimaud et al. 2008; Liu et al. 2012b; Abe et al. 2014). For other properties of BE, see review by Tetlow (2012).
3.1.2.3 Chain Preferences of BE Isozymes
The contributions of individual BE isozymes to the fine structure of amylopectin have been studied by analyzing changes in amylopectin structure in be mutants and transformants generated from a number of plant species such as maize (Hedman and Boyer 1982; Blauth et al. 2001, 2002; Yao et al. 2004, also see review by Shannon and Garwood 1984), rice (Nishi et al. 2001; Nakamura 2002; Satoh et al. 2003; Tanaka et al. 2004; Butardo et al. 2011), wheat (Regina et al. 2006; Sestili et al. 2010), barley (Regina et al. 2010; Carciofi et al. 2012), and pea (Smith 1988; Bhattacharyya et al. 1990). For other be mutants, see Tetlow (2012).
The impacts of these mutations were apparent when BEII-type forms were lacking while be1 mutants showed no or only a slight morphological and starch-related phenotypic changes. One most remarkable example is the pea mutant line called rugosus (r) which was used by Mendel as his experimental material and had a shriveled seed with a reduced starch synthesis caused by a loss of BEII-type (or A-type) isozyme (Smith 1988; Bhattacharyya et al. 1990). The other well-known example is the maize be2b mutant referred to as the amylose extender (ae) which caused a dramatic increase in amylose content and an alteration of amylopectin structure with longer chain length, which resulted in the commercially useful functional properties of the endosperm starch being resistant to digestion and gelatinization (Shannon et al. 2007; Li et al. 2008). The results suggest the specific role of BEII in starch biosynthesis in dicot reserve organs and BEIIb in monocot seeds, and their roles cannot be significantly supported by BEI and BEI plus BEIIa, respectively. Conversely, the role of BEI can be mostly complemented by BE isozyme(s). However, attention should be paid to the observations that in wheat and barley endosperms, the impact of BEIIa deficiency was much more similar to that of BEIIb deficiency of maize and rice (Yao et al. 2004; Nishi et al. 2001; Tanaka et al. 2004; Butardo et al. 2011), whereas suppression of BEIIb expression in wheat and barley resulted only in minor changes in starch phenotypes (Regina et al. 2006; Regina et al. 2010) (see Sect. 5.5.2).
Figure 5.7b shows that loss of BEIIb activity in rice endosperm caused a marked increase in long chains of DP ≧ 35 (B2- and B3-chains) and intermediate chains of DP 15–30 (A-chains plus B1-chains) and a great decrease in short chains of DP 6–12 in amylopectin (Nishi et al. 2001). The results support the notion that BEIIb forms short chains in the middle of the cluster instead of the root of the cluster, and therefore in its absence the number of chains per cluster (the ratio of A-chains plus B1-chains/B2-chains plus B3-chains) is significantly reduced (Figs. 5.2, 5.4, and 5.7). By contrast, the be1 mutant of rice had amylopectin with more chains of DP 6–12 and slightly less chains of DP 16–25 and DP ≧ 37 (Fig. 5.7a), suggesting that BEI plays a part in the synthesis of intermediate and long chains (Satoh et al. 2003). The situation is likely common among cereal endosperm (Yao et al. 2004; Regina et al. 2010; Butardo et al. 2011; Liu et al. 2012a), although the possibility that BEI also affects the position of branches of amylopectin in maize endosperm cannot be ruled out (Xia et al. 2011).
Changes in the amylopectin structure of mutants defective in starch branching enzyme (BE) isozymes of rice. Changes in the fine structure of amylopectin in rice endosperm caused by be1 (Satoh et al. 2003) (a) and be2b (Nishi et al. 2001) (b) mutations. The data shown are differences after subtraction of chain-length distribution of amylopectin of the wild-type from that of the mutant
The contribution of each BE isozyme to the amylopectin structure has been analyzed more precisely and in details by in vitro studies. It was firstly reported that maize BEII preferentially transferred very short chains of DP 6 to about 12 with most preferred chains of DP 6 for BEIIa and DP 7 and 6 for BEIIb, while BEI formed most effectively intermediate chains of DP about 10–12 (Takeda et al. 1993; Guan et al. 1997). Subsequently, BEs from other plant sources such as rice (Mizuno et al. 2001; Nakamura et al. 2010), kidney bean (Hamada et al. 2001; Nozaki et al. 2001), potato (Rydberg et al. 2001), green alga Chlorella (Sawada et al. 2009), and cyanobacterium Synechococcus (Sawada et al. 2013, 2014) were shown to have similar chain-length preferences as maize BE isozymes (see review by Tetlow 2012) and consistent with the enzymatic properties suggested by in vivo studies.
As an example of these in vitro studies, results with three rice BE isozymes are summarized (Nakamura et al. 2010), as shown in Fig. 5.8. First, when branched glucan such as amylopectin is used as substrate, the minimum chain lengths of substrate chains and the transferred chains for BE reaction were DP 12 and 6, respectively. In contrast, the minimum chain length of amylose required for branching was about DP 48 for BEI and DP ≻ 100 for BEIIa and BEIIb. Second, the most preferable chain lengths for branching of amylopectin by BEI, BEIIa, and BEIIb were DP 10–12, DP 6, and DP 7 and 6, respectively. Third, BEI could attack the internal chain segments of amylopectin chains but to a lesser extent than the external chain segments, while BEIIa and BEIIb could not. These results are basically consistent with those of corresponding BE isozymes from other plant species as described above, although precise comparison with BEs from various sources is difficult because these studies used different substrate, reaction conditions, and methods for measuring chain profiles.
Chain preference of starch branching enzyme (BE) isozymes analyzed by in vitro studies. The chain-length distribution of the reaction product of rice BE with ae-amylopectin used as substrate is shown. (a) The BEIIb reaction product. (b) The BEIIa reaction product. (c) The BEI reaction product (Data are from Sawada et al. 2014)
Recently, Sawada et al. (2013, 2014) analyzed and compared the chain preference of a total of ten BEs from various organisms including rice (OsBEI, OsBEIIa, and OsBEIIb), Chlorella kessleri (ChlBE), red alga Porphyridium purpureum (PopBE1 and PopBE2), Synechococcus elongatus (ScoBE), Escherichia coli BE (EcoBE), human (HosBE), and yeast (SacBE). It was found that EcoBE had the same enzymatic properties as OsBEI, while ScoBE and ChlBE had BEIIb-type properties. On the other hand, HosBE, SacBE, and two PopBEs exhibited the OsBEIIa-type properties. Analysis of chain-length profile of phosphorylase-limit dextrins (Φ-LD) of the BE reaction products revealed that EcoBE, ScoBE, PopBE1, and PopBE2 preferred A-chains as acceptors, while OsBEIIb used B-chains more frequently than A-chains. The preference of BE for acceptor chain seemed to reflect on the molecular structure of α-glucans in terms of the ratio of A-chains to B-chains in at least some organisms such as algae. The results showed for the first time the preference of some BEs to the acceptor chains and the location of branches. This information provides new insights into the mechanism for the synthesis of amylopectin cluster structure.
It is stressed that features of the chain preference of BE were maintained irrespective of the glucan substrate for the in vitro BE reaction, i.e., amylopectin or amylose (Guan et al. 1997; Nakamura et al. 2010), the duration of enzymatic reaction (Guan et al. 1997; Nakamura et al. 2010), and the reaction temperature (Nakamura et al. 2010; Ohdan et al. 2011), indicating that the specific chain preference of BE isozymes is consistent regardless of the structure of glucan substrate. Thus, the specificities of individual BEs for chain length of the glucans can be applicable to their roles under any physiological conditions.
3.1.2.4 The Roles of BE Isozymes in Amylopectin Biosynthesis in Cereal Endosperm
The contribution of BEIIb and BEI to the fine structure of amylopectin sharply contrasts from each other, and this is basically observed in any cereal endosperm. As seen in rice endosperm as a representative example, the view is supported by the different chain-length distribution pattern of amylopectin between be2b and be1 mutants (Fig. 5.7). The observation that the extent of change in chain-length profile was much higher in be2b mutant than be1 mutant indicates that the function of BEIIb is specific and this cannot be complemented by other BE isozymes while that of BEI can significantly be supported by other BEs. No phenotypic change in the amylopectin structure was detected in the be2a mutant, suggesting that the role of BEIIa is not specific and might merely be to support BEIIb and BEI. Because leaf be2a mutant amylopectin had less short chains compared to long chains, BEIIa might play an important part in forming short amylopectin chains in the leaf where BEIIb is not expressed. The specific role of BEIIb in rice endosperm was supported by the results with transgenic lines in which BEIIb gene was introduced into the be2b mutant (Tanaka et al. 2004). The extent of amylopectin chain pattern alteration among transformants was depending on the expressed level of BEIIb; remarkably most of the starch was replaced by water-soluble polysaccharides (WSP) when it was highly overexpressed (Fig. 5.5; see also Sect. 5.4.3.2).
3.1.3 Starch Debranching Enzyme (DBE)
Plants have two types of α-glucan debranching enzymes: isoamylase (ISA, EC 3.2.1.68) and pullulanase (PUL, EC 3.2.1.41). Both ISA and PUL directly debranch the α-1,6-glucosidic linkage, differing from DBE from heterotrophic eukaryotes such as animals and yeasts which removes the branched chain by the consecutive 4-α-glucanotransferase (EC 2.4.1.25) and amylo-1,6-glucosidase (EC 3.2.1.33) reactions (see reviews by Lee and Whelan 1971 and Nakamura 1996).
3.1.3.1 Involvement of DBE in Amylopectin Biosynthesis
One of the most striking biochemical events in the starch biosynthesis in plants is the fact that DBE is essential for the synthesis of cluster structure of amylopectin. Although some information suggesting the involvement of DBE in amylopectin synthesis had been reported through the analysis of phytoglycogen-producing sugary-1 mutants from maize (Pan and Nelson 1984) and rice (Nakamura et al. 1992b), direct evidence that ISA1 plays a crucial role in amylopectin biosynthesis and in its absence amylopectin is replaced by phytoglycogen devoid of cluster structure (Figs. 5.4a and 5.5) was reported by James et al. (1995) using the gene-tagging method. Since then the essential roles of DBEs in amylopectin synthesis have been verified by extensive investigations of a variety of mutants and transformants from maize (James et al. 1995; Beatty et al. 1999; Dinges et al. 2001; Dinges et al. 2003; Kubo et al. 2010; Lin et al. 2012, 2013), rice (Nakamura et al. 1996, 1997; Kubo et al. 1999, 2005; Fujita et al. 2003, 2009; Utsumi et al. 2011), potato (Hussain et al. 2003; Bustos et al. 2004), barley (Burton et al. 2002), Arabidopsis (Zeeman et al. 1998; Dellate et al. 2005; Wattebled et al. 2005, 2008; Streb et al. 2008; Pfister et al. 2014; Streb and Zeeman 2014), Chlamydomonas (Mouille et al. 1996; Colleoni et al. 1999a, 1999b; Dauvillée et al. 2000, 2001; Sim et al. 2014), and Cyanobacterium sp. CLg1 (Cenci et al. 2013) (also see review by Hennen-Bierwagen et al. 2012), although in this chapter experimental results with maize and rice endosperm are mainly described.
Green plants usually have four DBEs: three ISA isozymes, ISA1, ISA2, and ISA3, and one PUL (Ball et al. 2011). The essential role of ISA1 was proven by the molecular studies showing that the sugary-1 phenotype reverted to wild-type one when normal ISA1 gene was introduced into the sugary-1 mutant of rice (Kubo et al. 2005; Utsumi et al. 2011) (Fig. 5.9). Biochemical studies indicated that ISA2 functioned in a form of heteromeric complex with ISA1, although ISA2 itself lacked the catalytic site as DBE due to replacement and/or deletion of essential catalytic residues conserved in ISA1 and ISA3 (Hussain et al. 2003; Bustos et al. 2004). ISA3 likely plays a part in degrading branched glucans in the leaf (Hennen-Bierwagen et al. 2012), but not in the glucan synthesis in the endosperm (Yun et al. 2011). It is interesting that both ISA1 homomer and ISA1-ISA2 heteromer were present in maize endosperm (Kubo et al. 2010) and leaf (Lin et al. 2013), rice endosperm (Utsumi and Nakamura 2006; Utsumi et al. 2011), and Chlamydomonas (Sim et al. 2014). In contrast, only the ISA1-ISA2 heteromer existed in potato tuber (Hussain et al. 2003) as in Arabidopsis leaves (Dellate et al. 2005). It was found that the ISA1 homomer, but not the heteromer, was only the functional form in rice endosperm (Utsumi et al. 2011) while in maize endosperm the homomer played a major role in amylopectin synthesis but the heteromer could also support at least partly the normal starch synthesis (Kubo et al. 2010). These differences in the ISA functional form(s) between rice and maize were supported by the results indicating that the presence of the heteromer only caused by overexpression of ISA2 gene resulted in abnormal starch production in rice endosperm (Utsumi et al. 2011). It is noted that in the rice leaf the heteromer was the functional form for normal starch biosynthesis and that the heteromer was more resistant to high temperature at 40 °C than the homomer (Utsumi et al. 2011). In maize leaves, both homomer and heteromer existed (Lin et al. 2013). One likely reason for this phenomenon in rice is that the functional form of ISA should be more resistant in the leaf than in the endosperm presumably because intracellular environment due to diurnal and physiological conditions fluctuates more frequently and severely in the leaf than in the endosperm.
Changes in starch granular structure of mutants defective in isoamylase1 (ISA1) of rice and complementation of the mutant phenotype by introducing normal ISA1 gene from wheat. Granular morphology (a) and thermal properties (b) of α-glucans in rice endosperms of wild-type, an isa1 (sugary-1) mutant line EM914, and OsISA1 transformants (Data are from Kubo et al. 2005)
In the sugary-1 mutant endosperm, the activity of PUL was concomitantly reduced in maize (Pan and Nelson 1984; Dinges et al. 2003) and rice (Nakamura et al. 1996), and the reduced level of PUL was almost related with the severity of sugary-1 mutation or the extent of decreased ISA activity in rice endosperm (Nakamura et al. 1997), suggesting that PUL together with ISA plays a part in amylopectin biosynthesis in cereal endosperm. Later, Fujita et al. (2009) provided evidence that the role of PUL was to supplement the role of ISA when absent, but in its presence the PUL’s role was only limited, if any, in developing rice endosperm, basically consistent with previous results with maize (Beatty et al. 1999; Dinges et al. 2003) and rice endosperm (Kubo et al. 1999) endosperm.
3.1.3.2 Multiple DBE Isozymes, Structure, Basic Properties, and Functions
Both ISA and PUL are also members of the α-amylase family, the GH13, like BEs. The structural features of ISA (Katsuya et al. 1998; Sim et al. 2014) and PUL (Vester-Christensen et al. 2010) were revealed.
The ISA protein was first purified to homogeneity from developing endosperm of rice and shown to be in a form of ISA1 homomultimer (Fujita et al. 1999). However, Hussain et al. (2003) discovered that potato tuber had three ISA isozymes, ISA1, ISA2, and ISA3, and confirmed that the ISA1-ISA2 heteromer was responsible for the ISA activity contributing to the starch synthesis in potato tuber. The results are consistent with the previous report by Ishizaki et al. (1983) that the finally purified ISA activity preparation from potato tuber included two protein components with the same protein amount. Three distinct ISA genes and one PUL gene have been identified in all green plants so far examined (Hussain et al. 2003; Rahman et al. 2003; Deschamps et al. 2008; Hennen-Bierwagen et al. 2012).
The enzymatic properties of ISA have been examined in several sources. Utsumi and Nakamura (2006) showed that both ISA1 homomer and ISA1-ISA2 heteromer purified from developing rice endosperm had a broad chain-length preference like Pseudomonas amyloderamosa ISA. In contrast, the ISA from E. coli, which is sometimes referred to as GlgX protein and can be classified as an ISA subclass, was shown to be specific for hydrolysis of only very short chains of DP 2–4 (Dauvillée et al. 2005). Later, Takashima et al. (2007) proved that the ISA1-ISA2 heteromer purified from developing kidney bean had a broad chain-length preference, while the purified ISA3 only debranched the DP 2–4 chains. Comparative analyses by Kobayashi et al. (unpublished) also indicated that ISA-type DBE could be classified into three types, i.e., type 1, the rice ISA1 and P. amyloderamosa ISA that could act on any chains having various DP values; type 2, rice ISA3 and ISAs from cyanobacteria Cyanothece sp. ATCC51142 and Synechococcus elongatus PCC7942 that had the capacity for hydrolysis of short chains of DP up to about 13; and type 3, E. coli ISA that could only hydrolyze the DP 2–4 chains.
Information on the crystal structure of plant DBEs is available for Chlamydomonas ISA1 (CrISA1) (Sim et al. 2014) and barley PUL (Vester-Christensen et al. 2010). It is interesting that the homodimeric CrISA1 structure had a unique elongated structure with monomers connected end to end, and the distinct dimeric structure was commonly conserved in ISA1 homomer and ISA1-ISA2 heteromer at least in green plants (Dauvillée et al. 2000; Sim et al. 2014).
The molar mechanism as to how DBE is involved in amylopectin synthesis is not clear. This is because it is difficult to reveal the mechanism at molecular level under the present situation where nobody has succeeded to establish the reconstituted system for amylopectin biosynthesis in test tubes. Several ideas have been provided to explain why DBE is needed to synthesize the cluster structure of amylopectin. One most likely idea is that the role of DBE is to remove the unnecessary branches which are accidentally and rarely formed at the improper position on the cluster chains by BE (Nakamura 2002). The reason for the hypothesis is that cleavage of wrongly formed branches by DBE is indispensable to the cluster structure, because, otherwise, the presence of such branches will interfere with the formation of a bundle of double helices. This topic is further discussed below (Sect. 5.4.2).
3.1.4 Granule-Bound Starch Synthase (GBSS)
There are a few reports indicating the involvement of GBSS in amylopectin synthesis such as Chlamydomonas (Delrue et al. 1992; Maddelein et al. 1994). Rice endosperm frequently had the extra-long chains of DP > 100, which is referred to as super-long chains (SLC) (Takeda et al. 1987a; Hanashiro et al. 2005). Substantial evidence was provided that GBSS was responsible for the synthesis of the long chains (Takeda et al. 1987a; Inouchi et al. 2005; Hanashiro et al. 2008). The SLC was also detected in amylopectin from many dicot and monocot plants (see Chapter 1). It is known that the amount of SLC in amylopectin greatly affected the quality of cereal starch, which was derived from changes in physicochemical properties such as the setback value measured by rapid visco analyzer (Horibata et al. 2004; Inouchi et al. 2005).
3.2 The Biosynthesis of Amylose
3.2.1 GBSS and BE
Mutants having amylose-free starch called waxy mutants have been isolated from many plant species (Collins and Kempton 1911; Ikeno 1914; Jacobsen et al. 1989, see reviews by Shannon and Garwood 1984 and Shannon et al. 2007), and it is evident that GBSS was responsible for the synthesis of amylose (Nelson and Rines 1962; Murata et al. 1965; Visser et al. 1991).
There are two types of GBSS referred to as GBSSI and GBSSII or GBSSIb encoded by two different genes and expressed in reserve tissues such as cereal endosperm, potato tuber, and pea embryo and other tissues such as leaf, pericarp, cereal embryo, and aleurone layers (Fujita and Taira 1998; Nakamura et al. 1998; Tomlinson et al. 1998; Dian et al. 2003). Although assimilatory starch had a lower amylose content than reserve starch (Tomlinson et al. 1997, 1998), it is unknown if this was caused by the difference in enzyme features between GBSSI and GBSSII.
Denyer et al. (1999b) showed that GBSSI from pea embryo synthesized amylose in a processive manner by adding more than one glucose residue for each enzyme-glucan primer while SSII elongated MOS in a distributive manner, by adding only one glucose residue to the primer chain during association of the enzyme-primer complex. They also found that GBSSI had a much higher affinity for amylopectin as an effector than as a substrate, suggesting the critical feature of GBSSI for the synthesis of amylose (Denyer et al. 1999a).
It is known that some amylose molecules from a limited number of botanical sources so far analyzed had a branched structure with a less frequency of branches than amylopectin (Hizukuri et al. 1981; Takeda et al. 1987b, see also Chapter 2). Hanashiro et al. (2013) analyzed the features of branched chains in amylose molecules. Since short side chains of amylose were not highly ordered in amylose unlike in amylopectin, it is difficult at present to assess which BE isozyme is involved in the synthesis of amylose branches and what the donor chains are for providing amylose with branched chains.
4 Regulation and Features of the Biosynthetic Process of Reserve Starch
During their biological evolutions, green plants have developed a metabolic system in which the homogeneous structure of amylopectin molecules having the semicrystalline properties are efficiently reproduced at a high rate and immediately stored in the form of granule inside plastids. However, the fine structure of amylopectin changes in response to different expression levels of key isozymes involved in amylopectin synthesis, keeping a distinct pattern of change depending on each isozyme (Fig. 5.5), as described above. Thus, under a fixed physiological condition, the distinct amylopectin fine structure is basically determined by the balance between chain elongation reaction catalyzed by SS and chain branching reaction by BE, whereas DBE merely removes the improper branches to trim the cluster structure. How can only two kinds of reactions determine the skeleton of the highly organized cluster structure? One possible scenario is that plants have evolved the way by refining several combinations of interactions between different classes of starch synthetic enzymes and the same class of isozymes. How do plants coordinate the distinct and overlapping functions of individual isozymes during the synthesis of amylopectin?
4.1 Functional Interactions Between Multiple Enzymes
So far examined, no isoforms of SS, BE, and DBE show the sigmoidal curve in their activity changes in response to varied substrate and/or cofactor concentrations, a typical behavior for the regulatory enzymes like AGPase (see Chap. 11). Instead, their catalytic activities are controlled in a different way, and their activities greatly vary depending on the types of glucans such as amylopectin, amylose, maltodextrins, and MOS. More actually, reaction products formed by some enzymes affect the actions of other enzymes by providing suitable substrates for their catalysis. One of such examples is the observation that the length of branched chains formed by rice BEIIb was almost exclusively DP 7 or 6 (Nakamura et al. 2010) and the major substrate chains for rice SSI and its product were chains of DP 6–7 and DP 8, respectively (Nakamura et al. 2014). The DP 8 chain could then be elongated to form the intermediate intra-cluster chains by SSIIa (Nakamura et al. 2005a). The results suggest the close functional interactions among BEIIb, SSI, and SSIIa during amylopectin biosynthesis in rice endosperm and that these interactions between other combinations of enzymes play important roles in amylopectin biosynthesis in reserve tissues.
A number of results provided evidences for several kinds of physical association between enzymes while other results showed that some enzymes catalytically interacted with other enzymes/isozymes during the synthesis of amylopectin.
4.1.1 Catalytic Interactions
Recently, Nakamura et al. (2012, 2014) presented evidence that rice SSI and plastidial phosphorylase (Pho1, EC 2.4.1.1) could synthesize glucans in the absence of added glucan primer when either BE isozyme was present (see Chap. 9 in details). These SSI-BE and Pho1-BE interactions were established by stimulating mutual catalytic activities and increasing affinities for substrate glucans of the counterpart enzymes. The results might be some examples of the close catalytic interactions among some different enzymes in starch-synthesizing tissues. It is stressed that these interactions were realized in the presence of branched glucans or dextrins (Nakamura et al. 2012).
4.1.2 Protein-Protein Interactions
Tetlow et al. (2004, 2008) showed the protein-protein complex among starch synthetic enzymes in amyloplasts of developing wheat endosperm. It is noteworthy that the protein phosphorylation accelerated the formation of the protein multimer composed of BEIIb, BEI, and Pho1 as well as the BEIIb catalytic activity. The different combinations of starch synthetic enzyme complexes were also verified in developing maize endosperm, and the protein phosphorylation was likely to stimulate the functional roles of the enzymes in starch biosynthesis (Hennen-Bierwagen et al. 2008, 2009; Liu et al. 2012a, b, also see review by Emes and Tetlow 2012). These results strongly suggest that the heteromeric protein complexes are essentially involved in starch biosynthesis, and this topic is described in details in Chap. 8.
4.2 Models for the Starch Biosynthetic Process
During the course of evaluation of key enzymes, several models for explaining the functions of these enzymes have been proposed.
Ball et al. (1996) proposed a model by incorporating the newly established specific role of DBE (see Sect. 5.3.1.3) in the amylopectin synthesis by trimming the shape of its cluster structure, and therefore this model is referred to as “the glucan-trimming model” (Fig. 5.10a). They claim that the basic frame of amylopectin is constructed by combined actions of SS for chain elongation and BE for chain branching. Excess branches formed during BE reactions should be instantly cleared off by the action of DBE (mainly ISA), because otherwise these irregularly formed branches prevent the glucan crystallization.
Models for the synthesis of amylopectin. (a) The glucan-trimming model (Ball et al. 1996). (b) The glucan crystallization model (Myers et al. 2000). (c) The modified crystallization model (Hennen-Bierwagen et al. 2012). (d) The two-step branching and improper branches clearing model (Nakamura 2002) with a slight modification
Myers et al. (2000) stressed a concept that the crystallization process plays a distinct role during the biogenesis of amylopectin crystal (Fig. 5.10b). During the crystallization process, the removal of pre-amylopectin from the aqueous phase by DBE reaction is needed to make the mature amylopectin inaccessible to further enzymatic modification. This model explains the involvement of nonenzymatic chemical (or spontaneous) behavior of glucan chains after modification by various enzymes in the normal amylopectin biosynthetic process.
Nakamura (2002) proposed a newer model named “the two-step branching and improper branch clearing model” to explain the contribution of each isozyme during amylopectin biosynthesis in cereal endosperm (Fig. 5.10d), although his model was based on the biochemical analyses of SS, BE, and DBE isozymes in rice endosperm. This model emphasized that although interacting reactions between SS and BE isozymes play major roles in the synthesis of amylopectin structure, two different kinds of combinations between SS and BE isozymes play essential parts in the synthesis of normal cluster structure in cereal endosperm and that the role of ISA is to remove a few unnecessary branches that are spontaneously formed by BE reactions at the less-clustered improper positions. It is conceived that a small amount of such wrongly positioned branches is produced at regions remote from where most of the branches are correctly and densely synthesized, and these wrongly and dispersedly formed branches can be removed by ISA more easily than those normally and densely formed branches. This model hypothesizes that the first branches at the root of the cluster are mainly synthesized by BEI, eventually forming the amorphous lamellae, and the second branches are preferentially synthesized by BEIIb. As a result, most of the second branches are considered to be present at the border between the amorphous lamellae and the crystalline lamellae (Fig. 5.2). The first branched chains with the intermediate length synthesized by BEI are mainly elongated by SSIIIa and to a lesser extent by SSIIa and SSI, while the second branched chains with the short chain length formed by BEIIb are sequentially elongated by SSI and SSIIa, as described above (Sect. 5.4.1.1).
Recently, the Myers’ group demonstrated a modified model that explains in more details their ideas (Hennen-Bierwagen et al. 2012) (Fig. 5.10c). They postulated equilibrium between crystalline and non-crystalline glucans and claimed that ISA1-ISA2 plays a crucial role in transferring the non-crystalline disordered glucans into the crystalline-competent glucans by eliminating some branches that hinder the spontaneous crystallization.
Until now, no consensual model applicable for many plant species and tissues has been reported yet. A new model incorporating novel experimental results and ideas is waiting to be proposed.
4.3 Features of the Biosynthetic Process of Reserve Starch
4.3.1 Starch Biosynthesis and Its Degradation
For the most part the starch synthetic process is distinctly segregated from the degradation process. Classes and/or types of enzymes involved are almost different in both processes from each other, and especially in reserve organs, the timing of operation of each process is distinctive. Nevertheless, in several aspects, both processes are considered to be closely related. There are some direct and indirect evidences for the involvement of other enzymes besides SS, BE, and DBE in the starch biosynthesis in reserve tissues. At least some hydrolase, phosphorylase (Pho), and transglucosidase-type enzymes, which have been considered to play major roles in starch degradation, are highly expressed in tissues exhibiting vigorous starch production.
In spite of the high expression levels of Pho1 in reserve tissues from many plant species in accordance with the increase in their starch production, the in vivo function of Pho1 is still a matter of controversy. The Pho1 reaction is freely reversible and Pho1 itself can catalyze either the glucan synthetic or the degradable direction. However, the mutations at the Pho1 locus caused a severe reduction of starch biosynthesis in Chlamydomonas (Dauvillée et al. 2006) and rice endosperm (Satoh et al. 2008), strongly suggesting that Pho1 is involved in starch biosynthesis. Several reports have supported this idea. First, Fettke et al. (2010a, 2012) showed that G1P was efficiently and directly incorporated into reserve starch granules in potato tuber cells, suggesting the G1P pathway functions in parallel with the authentic AGPase pathway in starch biosynthesis. Second, Hwang et al. (2010) proved that rice Pho1 could elongate MOS to be used by SS in the glucan synthetic direction even under physiological conditions of high Pi/G1P concentrations. Third, Nakamura et al. (2012) showed the catalytic interaction in glucan synthesis between rice Pho1 and BE in the omission of added glucan primer, as described above. All these results support the view that Pho1 plays a critical role in starch biosynthesis as well as starch degradation.
It was reported that plastidial disproportionating enzyme (DPE1, EC 2.4.1.25) was involved in amylopectin biosynthesis in Chlamydomonas reinhardtii (Colleoni et al. 1999a, 1999b; Wattebled et al. 2003), although the contradictory results that DPE1 had no direct role in starch biosynthesis in Arabidopsis leaves were shown (Critchley et al. 2001).
The potential involvement of the chloroplastic α-amylase (EC 3.2.1.1) in the synthesis of phytoglycogen in Arabidopsis leaves under the condition where all DBE activities were lacking was reported by Streb et al. (2008). The authors hypothesized that SS and BE can basically synthesize starch-type glucans while DBE is indirectly involved in the synthesis (Zeeman et al. 2010). They explained that the role of DBE is to facilitate the crystallization of glucans by removing wrongly positioned branches because otherwise the glucan-degrading enzymes such as α-amylase can easily attack these untrimmed glucans. As a result, in the absence of DBE, the glucans unusually have very short chains of DP ≤ 5, which results in the formation of WSP.
It should be taken into account that a significant amount of MOS is liberated during the process of trimming of amylopectin cluster by DBE, and this MOS should be recycled into the synthetic process. Under physiological condition, most of its glucose residue must be recovered for the biosynthetic process to maintain the high yield of starch production from sucrose. In fact, the level of MOS is usually low in in vivo conditions, although the metabolic process involved in the fate of MOS during starch synthesis has not been fully examined. Besides the four major classes of starch synthetic enzymes (AGPase, SS, BE, and DBE), several types of enzymes presumably play distinct roles in this process. Initially, MOS can be easily degraded until it becomes the length of DP4 by Pho1 to form G1P. In addition, glucose and maltose can be generated by DPE1, α-amylase, and β-amylase (EC 3.2.1.2) from MOS, and this glucose will be converted to G1P by hexokinase and PGM. G1P will be again used by AGPase to form ADPglucose which enters the starch synthetic process via the ADPglucose pathway. The role of DPE1 is specific because it disproportionates the original MOS ranging from much smaller including glucose and maltose to larger MOS, which eventually makes MOS more easily metabolized by hexokinase for glucose and Pho1 and amylases for larger MOS.
All these results lead us to a conclusion that Pho1, DPE, and amylases have dual functions in both the synthesis and degradation of starch granules under various physiological and genetic conditions.
4.3.2 Factors Causing Damage to the Cluster Structure
The basic cluster structure of amylopectin is highly conserved even in the mutants whose chain-length profiles are profoundly modified by lesion of any SS, BE, and DBE isozymes. So far examined, six factors are known to be able to change the reserve glucans devoid of the cluster structure, although most of these factors were already stated above (Sect. 5.3.1). First, loss of ISA activity caused phytoglycogen production instead of amylopectin. Second, overexpression of ISA2 gene also resulted in phytoglycogen synthesis by bereaving the ISA1 homomer in rice endosperm because all the ISA1 proteins formed the ISA1-ISA2 heteromer that had no or little trimming activity for the synthesis of normal amylopectin at least in rice and maize endosperm. Third, excess of BEIIb activity inhibited the formation of organized branches and resulted in the synthesis of WSP in rice endosperm (Tanaka et al. 2004) (Fig. 5.11). Fourth, Lin et al. (2012) showed that when activities of both SSIII and ISA1 or ISA2 were defective in maize, phytoglycogen was produced in the endosperm. The close correlation between ISA and SS activities in starch/WSP accumulation was also found in Arabidopsis leaves (Pfister et al. 2014). Fifth, the mutation of rice Sugary-2 gene led to WSP synthesis especially at early stage of the endosperm development (Nakagami et al. unpublished) (Fig. 5.11), although the gene was recently mapped on the rice genome position different from any DBE genes (Nakamura T et al. unpublished). Sixth, α-amylase removed the capacity of Arabidopsis for the synthesis of starch-type glucan in the leaf when its ISA activity was lost (Streb et al. 2008), as described above.
Mutants and transformants producing water-soluble polysaccharide-(WSP)-type glucans in the rice endosperm. (a) Cross-section of rice kernels: wild-type (WT), a sugary-1 (isa1) line, an ISA2 overexpressed (ISA2-OX) line, a sugary-2 (isa2) line. (b) Seed morphology of rice transformants with different expression levels of BEIIb in which the normal rice BEIIb gene was introduced into a be2b mutant line (EM10) (see details by Tanaka et al. 2004). Note that the BEIIb overexpressed line (BEIIb-OX, #1-1), the mature seed became shriveled due to accumulation of water-soluble polysaccharides. (c) Scanning electron micrographs of glucan granular morphology in endosperm of the ISA2 overexpressed line (Utsumi et al. 2011), a sugary-2 mutant line (Nakagami et al. unpublished), and the BEIIb overexpressed line (Tanaka et al. 2004) of rice (See text in details)
In summary, the activity of ISA is likely to be the primary factor in the prevention of phytoglycogen production. In addition, balance of activities between multiple enzymes such as between ISA1 and ISA2, SSIII and ISA, BEIIb and SS, and DBE and α-amylase might be important for the synthesis of starch-type glucans, although comprehensive and more precise biochemical analyses are needed to draw the conclusion.
4.3.3 Minimum Sets of Enzymes Required for Construction of the Cluster Structure
It is known that in addition to green plants, some primitive algae including cyanobacteria, grey algae, and red algae could produce starch-like granules, although their amylopectin had less organized cluster structure than higher plant amylopectin and hence it is usually referred to as semi-amylopectin (Nakamura et al. 2005b; Deschamps et al. 2008; Shimonaga et al. 2008; Sawada et al. 2014, also see review by Colleoni and Suzuki 2012 and Chap. 4). Compared with a marked differentiation of starch synthetic enzymes into multiple isozymes in green plants, no significant differentiation in the glucan-metabolizing enzyme genes was found in the starch-synthesizing primitive algae. Some semi-amylopectin producing cyanobacteria such as Crocosphaera watsonii had three SS gene, three BE genes, and one ISA gene (Patron and Keeling 2005; Deschamps et al. 2008; Ball et al. 2011). Cyanidioschyzon merolae had one SS gene, one BE gene, and two ISA genes, and another unicellular red alga Porphyridium purpureum having one SS gene, two BE genes, and one ISA gene (Coppin et al. 2005; Patron and Keeling 2005; Deschamps et al. 2008; Ball et al. 2011; Bhattacharya et al. 2013) could also synthesize semi-amylopectin (Hirabaru et al. 2010; Shimonaga et al. 2006, 2008). It is also known that many mutant and transformant lines prepared from higher plants having a single major SS isozyme, BE isozyme, or ISA1 out of corresponding multiple isozymes could also produce starch-like glucans (Tetlow 2012; Fujita and Nakamura 2012; Hennen-Bierwagen et al. 2012).
All these results indicate that the presence of multiple isozymes in each class of starch synthetic enzymes is not an absolute requirement for the synthesis of starch. Rather, it is likely that multiple isozymes enable plant tissues to efficiently produce starch under varying physiological and environmental conditions by giving them differentiation in capacities for production of various types of tissue- and species-specific starches with refined structural features.
4.3.4 Reactivity and Binding Affinity of Starch Biosynthetic Enzymes Toward Glucans
During the starch synthesis, the fine structure of glucan molecule changes in every step of consecutive reactions involved in numerous starch synthetic enzymes. How can each isozyme act correctly on the exact glucan substrate by selecting it among various structures of glucans simultaneously formed by many other enzymes? Biochemical in vitro studies showed that SS and BE recognized the length of external segments of A- and B-chains (Guan et al. 1997; Commuri and Keeling 2001; Nakamura et al. 2010, 2014; Sawada et al. 2014), although some BE isozymes such as BEI could also attack the internal segments of B2- and B3-chains to some extent (Nakamura et al. 2010; Sawada et al. 2014). It was also established that SS and BE were much more preferentially reactive toward branched glucans than linear MOS and dextrins/glucans (Nakamura et al. 2010, 2014 and many references therein), suggesting that the actual glucan substrates for SS and BE during in vivo starch synthesis are the dual parallel chains or the double helical chains, respectively, of the branched glucans/dextrins, as proposed by Borovsky et al. (1979) for BE. Recently, it was found that SSI from rice (Nakamura et al. 2014) and SS from Arabidopsis (Brust et al. 2014) could efficiently synthesize glucans by interacting closely with BE isozymes, possibly forming the protein complex, in the absence of added primer as found in the interaction between Pho1 and BE (Nakamura et al. 2012). It is noted that these interactions were mediated by branched glucans, but not by linear glucans (Nakamura et al. 2012; Brust et al. 2014). These results strongly suggest that at least SSI, BEs, and Pho1 can closely interact with their counterparts including many types of glucans varied under different physiological conditions.
5 Future Perspectives for Research on the Biosynthesis of Reserve Starch
5.1 Major Subjects Unresolved
Despite huge amounts of biochemical, molecular, genetic, physicochemical, and structural studies as well as theoretical analyses and proposals for long time, we have to admit that there are still a lot of important subjects related to starch biosynthesis remaining unresolved.
First, how is the cluster structure constructed, especially how the length of single cluster is made constant? Concomitant with analytical studies on the enzymatic reaction mechanisms, the minimum requirements of the glucan fine structure needed for construction of the cluster should be defined (see Chap. 1).
Second, although currently the most conventional method for determination of the fine structure of glucans is to measure their chain-length distributions by HPLC or capillary electrophoresis, the method only measures the proportion of component chains of the glucan, but neither method can give any information on the position of branches. However, it is much more important to know the localized positions of branched chains and the lengths of internal segments of chains than to simply know the chain-length distribution of the glucan.
Third, how are the protein-protein multimeric structures related to their specific functions? Further analytical data are needed to clarify the mechanism for the actions of multimeric structure and to reveal the extent of involvement of enzyme interactions in starch biosynthesis.
Fourth, in which way do the enzymes interact with glucans? Starch biosynthetic enzymes are known to closely interact with environmental glucans in different ways, and some of them such as GBSS, BEIIb, and SSI were tightly and lightly bound to starch granules at different proportions while others such as DBEs were almost exclusively soluble in the stroma compartment. However, since enzymes were usually non-covalently associated with carbohydrates, it was difficult to elucidate the mechanism on how the bindings of enzymes to glucans influenced their functions/activities. The mechanism on how the carbohydrate-binding module (CBM) present in GH13 family enzymes (Janeček et al. 2011) and SSIII (Schwarte et al. 2013) relates to their enzymatic actions awaits resolution.
Fifth, how are enzymes related with glucan granule morphology? There have been some reports showing that some enzymes were even present in specific plastid membrane spaces and compartments, e.g., SSIIIa, SSIVb, GBSSI, and ISA3 (Gámez-Arjona et al. 2011; Yun et al. 2011; Kawagoe 2013; Toyosawa et al. 2015). It is likely that they play important roles in the starch granule initiation, development, and morphology, as well as in the conventional starch biosynthetic process, although the mechanisms as to how they are involved in these events remain to be elucidated.
5.2 Diversity of Regulation of Starch Biosynthesis Among Plant Species and Pleiotropic Effects
Notably it is known that relative activities of major SS isozymes greatly differed among plant species and this affected the species-specific structural and functional features of starch (Edwards et al. 1999b; Fujita and Nakamura 2012). For example, SSI was a major component of the total SS activity in the endosperm of maize (Cao et al. 1999), rice (Fujita et al. 2006), and Arabidopsis leaf (Delvallé et al. 2005; Zhang et al. 2005), but not in pea embryo (Craig et al. 1998) and potato tuber (Marshall et al. 1996) where SSII and SSIII, respectively, accounted for the bulk activity of SS. The results might support the variation on the mode of amylopectin chains synthesized through concerted actions of different combinations of SS isozymes and/or other synthetic enzymes such as BE and DBE depending on the plant species and tissues.
BEII is differentiated into distinct BEIIa and BEIIb isozymes in cereals, but not in dicots. In maize and rice endosperm, the specific chain preference of BEIIb was likely to be one of the strongest determinants of starch properties, and this role could not be supplemented by BEIIa (Nishi et al. 2001; Klucinec and Thompson 2002; Yao et al. 2004; Butardo et al. 2011). In contrast, the implication of BEIIa was more significant than BEIIb in wheat and barley endosperm (Regina et al. 2005), while loss of both BEIIa and BEIIb showed much more profound effects on starch structure than that of only BEIIa (Regina et al. 2006, 2010; Sestili et al. 2010). The results suggest that in wheat and barley, the functions of BEIIa and BEIIb largely overlap, whereas in rice and maize, BEIIb has a distinct role in starch biosynthesis. It is likely that the difference in contribution of BEIIa and BEIIb was not merely due to the relative activity of BEIIa to BEIIb because in rice endosperm their activities were at the same level (Yamanouchi and Nakamura 1992) and the distinct role of BEIIb was evident even if it had lower activity levels compared to the BEIIa (Tanaka et al. 2004). In addition, the silencing of the BEIIa gene profoundly influenced the amylose content and starch granule morphology in wheat and barley (Regina et al. 2006, 2010), while the be2b mutation had a minor impact on amylose content in rice (Nishi et al. 2001; Butardo et al. 2011). Remarkably, Carciofi et al. (2012) succeeded to produce amylose-only starch in barley endosperm by silencing all the BE genes. The impact of BEIIb/IIa gene on amylose content in maize was more similar to wheat and barley than rice because maize ae starch contained high amylose content (Li et al. 2008). Starch phenotypes of be2b (ae) mutants were considerably different between maize and rice (Banks et al. 1974; Baba et al. 1982; Nishi et al. 2001; Li et al. 2008). These results indicate that BEIIb exhibited a great variation in terms of its distinct function and expression level in cereal endosperm among species.
The roles of ISAs, their presence and composition, and their contributions to starch biosynthesis were dramatically different depending on plant species and tissues (Hennen-Bierwagen et al. 2012; Facon et al. 2013). One example is that the complete loss of ISA1 gene caused most cells to synthesize both starch-type glucans and WSP in many plant sources, while starch was completely replaced by WSP in rice endosperm (Nakamura et al. 1997) and Chlamydomonas (Mouille et al. 1996), although the exact reason remains to be clarified.
Several reports showed that multiple forms of some enzymes such as SSI (Baba et al. 1993) and BEIIa (Yamanouchi and Nakamura 1992) in rice endosperm, BEI in wheat endosperm (Båga et al. 2000), and BEII from kidney bean (Hamada et al. 2002) were generated from a single gene by transcriptional or posttranscriptional regulation and/or modification. However, the physiological significances of multiple forms of the same enzymes were unclear.
There are ample evidences showing the pleiotropic effects of losses of some enzymes on the enhancement or reduction on the levels of other enzymes (Sestili et al. 2010 and references therein). These suggest the presence of some factors which integrate the expression and amounts of numerous genes and proteins involved in starch metabolism and the close interrelation via protein-protein and protein-glucan interactions. The effects were possibly caused by many reasons at various levels. Recently, the specific and complex relations were found in protein levels and presence (e.g., binding to starch granules) among SSI, SSIIIa, BEI, and BEIIb in rice endosperm using various SSI-BE (Abe et al. 2014) and SSIIIa-BE (Asai et al. 2014) double mutants (see Chap. 10 in details), although some of them might have been caused by effects on association of protein complex among them in maize endosperm reported by Liu et al. (2012a, b). The molecular mechanism and physiological significance for some pleiotropic effects will be elucidated in the future.
5.3 New Approaches and Techniques Required
A number of new approaches will help us to understand the regulatory mechanisms for starch biosynthesis. First, a new practical method to determine the branch position in glucan chains should be established. This would give new insights into the possible roles of enzymes in determining the distribution and location of branches in amylopectin (Xia et al. 2011). Hizukuri (1996) proposed the enzymatic method for determination of the average span length and the average number of α-1,4-linkages between the nearest branch linkages, by calculating the number of glucosyl stubs of the B-chains treated with a sequential hydrolysis by β-amylase, isoamylase, and β-amylase. Usui et al. (2009) established a method for measuring directly the α-1,6-glucosidic linkages and the α-1,4-linkages along with MOS by negative ion Q-TOF MS/MS spectrometry method. However, a new standard method to determine exactly the position of α-1,6-linkages of glucans needs to be developed.
Second, methods to establish a reconstituted system to synthesize glucans with the cluster structure by in vitro enzymatic reactions are desirable. So far, no attempts have been successful to synthesize glucans having the cluster structure by in vitro reactions, whereas it is easy to synthesize glycogen-type glucans in test tubes in the presence of SS and BE. Since the use of readily available glucans composed of a mixture of chains with different lengths does not lead to a clear conclusion regarding characteristics of the enzyme, the choice of appropriate glucan primers with known structural features and lengths of component chains which are chemically and/or enzymatically prepared and the appropriate combination of candidate enzymes might be essentially important.
Third, factors which potentially integrate the starch biosynthetic process in plant tissues (She et al. 2010; Wang et al. 2013; Peng et al. 2014) need to be fully characterized. The use of transgenic lines carrying various types of constructs might provide important insights into the subject, because multiple candidate gene functions can be simultaneously modified by using, for example, the RNAi technology.
References
Abe N, Asai H, Yago H et al (2014) Relationships between starch synthase I and branching enzyme isozymes determined using double mutant rice lines. BMC Plant Biol 14:80
Asai H, Abe N, Matsushima R, et al. (2014) Deficiencies in both starch synthase (SS) IIIa and branching enzyme IIb lead to a significant increase in amylose in SSIIa-inactive japonica rice seeds. J Exp Bot 65:5497–5507
Baba T, Arai Y, Yamamoto T et al (1982) Some structural features of amylomaize starch. Phytochemistry 21:2291–2296
Baba T, Nishihara M, Mizuno K et al (1993) Identification, cDNA cloning, and gene expression of soluble starch synthase in rice (Oryza sativa L.) immature seeds. Plant Physiol 103:565–573
Båga M, Nair RB, Repellin A et al (2000) Isolation of a cDNA encoding a granule-bound 152-kilodalton starch-branching enzyme in wheat. Plant Physiol 124:253–263
Ball S, Guan H-P, James MG et al (1996) From glycogen to amylopectin: a model for the biogenesis of the plant starch granule. Cell 86:349–352
Ball S, Colleoni C, Cenci U et al (2011) The evolution of glycogen and starch metabolism in eukaryotes gives molecular clues to understand the establishment of plastid endosymbiosis. J Exp Bot 62:1775–1801
Banks W, Greenwood CT, Muir DD (1974) Studies on starches of high amylose content. Part 17. A review of current concepts. Starch 26:289–300
Beatty MK, Rahman A, Cao H et al (1999) Purification and molecular genetic characterization of ZPU1, a pullulanase-type starch-debranching enzyme from maize. Plant Physiol 119:255–265
Bhattacharya D, Price DD, Chan CX et al (2013) Genome of the red alga Porphyridium purpureum. Nat Commun 4:2931
Bhattacharyya MK, Smith AM, Ellis THN et al (1990) The wrinkled-seed character of pea described by Mendel is caused by a transposon-like insertion in a gene encoding starch-branching enzyme. Cell 60:115–122
Blauth SL, Yao Y, Klucinec JD et al (2001) Identification of Mutator insertional mutants of starch-branching enzyme 2a in corn. Plant Physiol 125:1396–1405
Blauth SL, Kim KN, Klucinec J et al (2002) Identification of Mutator insertional mutants of starch-branching enzyme 1 (sbe1) in Zea mays L. Plant Mol Biol 48:287–297
Borovsky D, Smith EC, Whelan WJ et al (1979) The mechanism of Q-enzyme action and its influence on the structure of amylopectin. Arch Biochem Biophys 198:627–631
Boyer CD, Preiss J (1978) Multiple forms of (1–4)-α-D-glucan, (1–4)-α-D-glucan-6-glycosyl transferase from developing Zea mays L. kernels. Carbohydr Res 61:321–334
Boyer CD, Preiss J (1979) Properties of citrate-stimulated starch synthesis catalyzed by starch synthase I of developing maize kernels. Plant Physiol 64:1039–1042
Brust H, Lehman T, D’Hulst C et al (2014) Analysis of the functional interaction of Arabidopsis starch synthase and branching enzyme isoforms reveals that the cooperative action of SSI and BEs results in glucans with polymodal chain length distribution similar to amylopectin. PLoS One 9:e102364
Burton R, Bewley JD, Smith AM et al (1995) Starch branching enzymes belonging to distinct enzyme families are differentially expressed during pea embryo development. Plant J 7:3–15
Burton RA, Jenner H, Carrangis L et al (2002) Starch granule initiation and growth are altered in barley mutants that lack isoamylase activity. Plant J 31:97–112
Bustos R, Fahy B, Hylton CM et al (2004) Starch granule initiation is controlled by a heteromultimeric isoamylase in potato tubers. Proc Natl Acad Sci U S A 101:2215–2220
Butardo VM, Fitzgerald MA, Bird AR et al (2011) Impact of down-regulation of starch branching enzyme IIb in rice by artificial microRNA- and hairpin RNA-mediated RNA silencing. J Exp Bot 62:4927–4941
Cao H, Imparl-Radosevich J, Guan H et al (1999) Identification of the soluble starch synthase activities of maize endosperm. Plant Physiol 120:205–215
Carciofi M, Blennow A, Jensen SL et al (2012) Concerted suppression of all starch branching enzyme genes in barley produces amylase-only starch granules. BMC Plant Biol 12:223
Cenci U, Chabi M, Ducatez M et al (2013) Convergent evolution of polysaccharide debranching defines a common mechanism for starch accumulation in cyanobacteria and plants. Plant Cell 25:3961–3975
Chaen K, Noguchi J, Omori T (2012) Crystal structure of the rice branching enzyme I (BEI) in complex with maltopentaose. Biochem Biophys Res Commun 424:508–511
Colleoni C, Suzuki E (2012) Storage polysaccharide metabolism in cyanobacteria. In: Tetlow I (ed) Starch: origins, structure and metabolism, vol 5, Essential reviews in experimental biology. The Society for Experimental Biology, London, pp 217–253
Colleoni C, Dauvillee D, Mouille G et al (1999a) Genetic and biochemical evidence for the involvement of α-1,4 glucanotransferases in amylopectin synthesis. Plant Physiol 120:993–1004
Colleoni C, Dauvillee D, Mouille G et al (1999b) Biochemical characterization of the Chlamydomonas reinhardtii α-1,4 glucanotransferase supports a direct function in amylopectin biosynthesis. Plant Physiol 120:1005–1014
Collins GN, Kempton JF (1911) Inheritance of waxy endosperm in hybrids of Chinese maize. Proc IV Int Genet Congr (Paris) 347–356
Commuri PD, Keeling PL (2001) Chain-length specificities of maize starch synthase I enzyme: studies of glucan affinity and catalytic properties. Plant J 25:475–486
Coppin A, Varre JS, Lienard L et al (2005) Evolution of plant-like crystalline storage polysaccharide in the protozoan parasite Toxoplasma gondii argues for a red alga ancestry. J Mol Evol 60:257–267
Craig J, Lloyd JR, Tomlinson K et al (1998) Mutations in the gene encoding starch synthase II profoundly alter amylopectin structure in pea embryos. Plant Cell 10:413–426
Critchley JH, Zeeman S, Takaha T et al (2001) A critical role for disproportionating enzyme in starch breakdown is revealed by a knock-out mutation in Arabidopsis. Plant J 26:89–100
Crumpton-Taylor M, Pike M, Lu K et al (2013) Starch synthase 4 is essential for coordination of starch granule formation with chloroplast division during Arabidopsis leaf expansion. New Phytol 200:1064–1075
Cuesta-Seijo JA, Nielsen MM, Marri L et al (2013) Structure of starch synthase I from barley: insight into regulatory mechanisms of starch synthase activity. Acta Crystallogr D Biol Crystallogr D69:1013–1025
D’Hulst C, Merida A (2012) Once upon a time: inception of the understanding of starch initiation in plants. In: Tetlow I (ed) Starch: origins, structure and metabolism. , vol 5, Essential reviews in experimental biology. The Society for Experimental Biology, London, pp 55–76
Damager I, Denyer K, Motawia MS et al (2001) The action of starch synthase II on 6’-α-maltotoriosyl-maltohexaose comprising the branch point amylopectin. Eur J Biochem 268:4878–4884
Dang PL, Boyer CD (1988) Maize leaf and kernel starch synthases and starch branching enzymes. Phytochemistry 27:1255–1259
Dauvillée D, Mestre V, Colleoni C et al (2000) The debranching enzyme complex missing in glycogen accumulating mutants of Chlamydomonas reinhardtii displays an isoamylase-type specificity. Plant Sci 157:145–156
Dauvillée D, Colleoni C, Mouille G et al (2001) Biochemical characterization of wild-type and mutant isoamylases of Chlamydomonas reinhardtii supports a function of the multimeric enzyme organization in amylopectin maturation. Plant Physiol 125:1723–1731
Dauvillée D, Kinderf IS, Li Z et al (2005) Role of the Escherichia coli glgX gene in glycogen metabolism. J Bacteriol 187:1465–1473
Dauvillée D, Chochois V, Steup M et al (2006) Plastidial phosphorylase is required for normal starch synthesis in Chlamydomonas reinhardtii. Plant J 48:274–285
Davis JH, Kramer HH, Whistler RL (1955) Expression of the gene du in the endosperm of maize. Agron J 47:232–235
Dellate T, Trevisan M, Parker ML et al (2005) Arabidopsis mutants Atisa1 and Atisa2 have identical phenotypes and lack the multimeric isoamylase, which influences the branch point distribution of amylopectin during starch synthesis. Plant J 41:815–830
Delrue B, Fontaine T, Routier F et al (1992) Waxy Chlamydomonas reinhardtii: monocellular algal mutants defective in amylose biosynthesis and granule-bound starch synthase activity accumulate a structurally modified amylopectin. J Bacteriol 174:3612–3620
Delvallé D, Dumez S, Wattebled F et al (2005) Soluble starch synthase I: a major determinant for the synthesis of amylopectin in Arabidopsis thaliana leaves. Plant J 43:398–412
Denyer K, Dunlap F, Thornbjørnsen T et al (1996) The major form of ADP-glucose pyrophosphorylase in maize (Zea mays L.) endosperm is extra-plastidial. Plant Physiol 112:779–785
Denyer K, Waite D, Edwards A et al (1999a) Interaction with amylopectin influences the ability of granule-bound starch synthase I to elongate malto-oligosaccharides. Biochem J 342:647–653
Denyer K, Waite D, Motawia S et al (1999b) Granule-bound starch synthase I in isolated starch granules elongates malto-oligosaccharides processively. Biochem J 340:183–191
Deschamps P, Colleoni C, Nakamura Y et al (2008) Metabolic symbiosis and the birth of the plant kingdom. Mol Biol Evol 25:536–548
Dian W, Jiang H, Chen Q et al (2003) Cloning and characterization of the granule-bound starch synthase II gene in rice: gene expression is regulated by the nitrogen level, sugar and circadian rhythm. Planta 218:261–268
Dinges JR, Colleoni C, Myers AM et al (2001) Molecular structure of three mutations at the maize sugary1 locus and their allele-specific phenotypic effects. Plant Physiol 125:1406–1418
Dinges JR, Colleoni C, James MG et al (2003) Mutational analysis of the pullulanase-type debranching enzyme of maize indicates multiple functions in starch metabolism. Plant Cell 15:666–680
Edwards A, Borthakur A, Bornemann S et al (1999a) Specificity of starch synthase isoforms from potato. Eur J Biochem 266:724–736
Edwards A, Fulton DC, Hylton CM et al (1999b) A combined reduction in activity of starch synthases II and III of potato has novel effects on the starch of tubers. Plant J 17:251–261
Emes MJ, Tetlow IJ (2012) The role of heteromeric protein complexes in starch synthesis. In: Tetlow I (ed) Starch: origins, structure and metabolism, vol 5, Essential reviews in experimental biology. The Society for Experimental Biology, London, pp 255–278
Facon M, Lin Q, Azzaz AM et al (2013) Distinct functional properties of isoamylase-type starch debranching enzymes in monocot and dicot leaves. Plant Physiol 163:1363–1375
Fettke J, Albrecht T, Hejazi M et al (2010a) Glucose 1-phosphate is efficiently taken up by potato (Solanum tuberosum) tuber parenchyma cells and converted to reserve starch granules. New Phytol 185:663–675
Fettke J, Malinova I, Albrecht T et al (2010b) Glucose-1-phosphate transport into protoplasts and chloroplasts from leaves of Arabidopsis. Plant Physiol 155:1723–1734
Fettke J, Leifels L, Brust H et al (2012) Two carbon fluxes to reserve starch in potato (Solanum tuberosum L.) tuber cells are closely interconnected but differently modulated by temperature. J Exp Bot 63:3011–3029
Fisher DK, Gao M, Kim K (1996) Allelic analysis of the maize amylose-extender locus suggests that independent genes encode starch-branching enzymes IIa and IIb. Plant Physiol 110:611–619
Fontaine T, D-Hulst C, Maddelein ML et al (1993) Toward an understanding of the biogenesis of the starch granules. Evidence that Chlamydomonas starch synthase II controls the synthesis of intermediate size glucans of amylopectin. J Biol Chem 268:16223–16230
Fujita N, Nakamura Y (2012) Distinct and overlapping functions of starch synthase isoforms. In: Tetlow I (ed) Starch: origins, structure and metabolism, vol 5, Essential reviews in experimental biology. The Society for Experimental Biology, London, pp 115–140
Fujita N, Taira T (1998) A 56 kDa protein is a novel granule-bound starch synthase existing in the pericarps, aleurone layers, and embryos of immature seeds of diploid wheat (Triticum monococcum L.). Planta 207:125–132
Fujita N, Kubo A, Francisco PB Jr et al (1999) Purification, characterization, and cDNA structure of isoamylase from developing endosperm of rice. Planta 208:283–293
Fujita N, Kubo A, Suh DS et al (2003) Antisense inhibition of isoamylase alters the structure of amylopectin and the physiological properties of starch in rice endosperm. Plant Cell Physiol 44:607–618
Fujita N, Yoshida M, Asakura N et al (2006) Function and characterization of starch synthase I using mutants in rice. Plant Physiol 140:1070–1084
Fujita N, Yoshida M, Kondo T et al (2007) Characterization of SSIIIa-deficient mutants of rice: the function of SSIIIa and pleiotropic effects by SSIIIa deficiency in the rice endosperm. Plant Physiol 144:2009–2023
Fujita N, Toyosawa Y, Utsumi Y et al (2009) Characterization of pullulanase (PUL)-deficient mutants of rice (Oryza sativa L.) and the function of PUL on starch biosynthesis in the developing rice endosperm. J Exp Bot 60:1009–1023
Gámez-Arjona FM, Li J, Raynaud S et al (2011) Enhancing the expression of starch synthase class IV results in increased levels of both transitory and long-term storage starch. Plant Biotechnol J 9:1049–1060
Gao M, Fisher DK, Kim K et al (1996) Evolutionary conservation and expression patterns of maize starch branching enzyme I and IIb genes suggest isoform specialization. Plant Mol Biol 30:11223–11232
Gao M, Wanat J, Stinard PS et al (1998) Characterization of dull1, a maize gene coding for a novel starch synthase. Plant Cell 10:399–412
Gidley MJ, Bulpin PV (1987) Crystallization of malto-oligosaccharides as models of the crystalline forms of starch. Carbohydr Res 161:291–300
Grimaud F, Rogniaux H, James MG et al (2008) Proteome and phosphoproteome analysis of starch granule-associated proteins from normal maize and mutants affected in starch biosynthesis. J Exp Bot 59:3395–3406
Guan H, Preiss J (1993) Differentiation of the properties of the branching isozymes from maize (Zea mays). Plant Physiol 102:1269–1273
Guan H, Li P, Imparl-Radosevich J et al (1997) Comparing the properties of Escherichia coli branching enzyme and maize branching enzyme. Arch Biochem Biophys 342:92–98
Hamada S, Nozaki K, Ito H et al (2001) Two starch-branching-enzyme isoforms occur in different fractions of developing seeds of kidney bean. Biochem J 359:23–34
Hamada S, Ito H, Hiraga S et al (2002) Differential characteristics and subcellular localization of two starch-branching enzyme isoforms encoded by a single gene in Phaseolus vulgaris L. J Biol Chem 277:16538–16546
Han Y, Bendik E, Sun F et al (2007a) Genomic isolation of genes encoding starch branching enzyme II (SBEII) in apple: Towards characterization of evolutionary disparity in SbeII genes between monocots and eudicots. Planta 226:1265–1276
Han Y, Sun F, Rosales-Mendoza S et al (2007b) Three orthologs in rice, Arabidopsis, and Populus encoding starch branching enzymes (SBEs) are different from other SBE gene families in plants. Gene 401:123–130
Hanashiro I, Matsunaga J, Egashira T et al (2005) Structural characterization of long unit-chains of amylopectin. J Appl Glycosci 52:233–237
Hanashiro I, Ito K, Kuratomi Y et al (2008) Granule-bound starch synthase I is responsible for biosynthesis of extra-long unit chains of amylopectin in rice. Plant Cell Physiol 49:925–933
Hanashiro I, Sakaguchi I, Yamashita H (2013) Branched structures of rice amylose examined by differential fluorescence detection of side-chain distribution. J Appl Glycosci 60:79–85
Hawker JS, Ozbun JL, Preiss J (1972) Unprimed starch synthesis by soluble ADPglucose-starch glucosyltransferase from potato tubers. Phytochemistry 11:1287–1293
Hawker JS, Ozbun JL, Ozaki H et al (1974) Interaction of spinach leaf adenosine diphosphate glucose α-1,4-glucan α-4-glucosyl transferase and α-1,4-glucan, α-1,4-glucan-6-glucosyl transferase in synthesis of branched α-glucan. Arch Biochem Biophys 160:530–551
Hedman KD, Boyer CD (1982) Gene dosage at the amylose-extender locus of maize: effects on the levels of starch branching enzymes. Biochem Genet 20:483–492
Hennen-Bierwagen TA, Liu F, Marsh RS et al (2008) Starch biosynthetic enzymes from developing Zea mays endosperm associate in multisubunit complexes. Plant Physiol 146:1892–1908
Hennen-Bierwagen TA, Lin Q, Grimaud F et al (2009) Proteins from multiple metabolic pathways associate with starch biosynthetic enzymes in high molecular weight complexes: a model for regulation of carbon allocation in maize amyloplasts. Plant Physiol 149:1541–1559
Hennen-Bierwagen TA, James MG, Myers AM (2012) Involvement of debranching enzymes in starch biosynthesis. In: Tetlow I (ed) Starch: origins, structure and metabolism, vol 5, Essential reviews in experimental biology. The Society for Experimental Biology, London, pp 179–215
Hirabaru C, Izumo A, Fujiwara S et al (2010) The primitive rhodophyte Cyanidioschyzon merolae contains a semiamylopectin-type, but not an amylose-type α-glucan. Plant Cell Physiol 51:682–693
Hirose T, Terao T (2004) A comprehensive expression analysis of the starch synthase gene family in rice (Oryza sativa L.). Planta 220:9–16
Hizukuri S (1986) Polymodal distribution of the chain lengths of amylopectins, and its significance. Carbohydr Res 147:342–347
Hizukuri S (1996) Starch: analytical aspects. In: Eliasson AC (ed) Carbohydrates in food starch. Essential reviews in experimental biology, vol 5. Marcel Dekker Inc, New York/Basel/Hong Kong; The Society for Experimental Biology, London, pp 115–140
Hizukuri S, Takeda Y, Yasuda M et al (1981) Multi-branched nature of amylose and the action of debranching enzymes. Carbohydr Res 94:205–213
Horibata T, Nakamoto M, Fuwa H et al (2004) Structural and physicochemical characteristics of endosperm starches of rice cultivars recently bred in Japan. J Appl Glycosci 51:303–313
Hussain H, Mant A, Seale R et al (2003) Three isoforms of isoamylase contribute different catalytic properties for the debranching of potato glucans. Plant Cell 15:133–149
Hwang S, Nishi A, Satoh H et al (2010) Rice endosperm-specific plastidial α-glucan phosphorylase is important for synthesis of short-chain malto-oligosaccharides. Arch Biochem Biophys 495:82–92
Ikeno S (1914) Über die Bestäubung und die Bastardierung von Reis. Z Pflanzenzucht 2:495–503
Imparl-Radosevich JM, Gameon JR, McKean A et al (2003) Understanding catalytic properties and functions of maize starch synthase isozymes. J Appl Glycosci 50:177–182
Inouchi N, Hibiu H, Li T et al (2005) Structure and properties of endosperm starches from cultivated rice of Asia and other countries. J Appl Glycosci 52:239–246
Ishizaki Y, Taniguchi H, Maruyama Y et al (1983) Debranching enzymes of potato tubers (Solanum tuberosum L.). I. Purification and some properties of potato isoamylase. Agric Biol Chem 47:771–779
Jacobsen E, Hovenkamp-Hermelink JHM, Krijgheld HT et al (1989) Phenotypic and genotypic characterization of an amylase-free starch mutant of the potato. Euphytica 44:43–48
James MG, Robertson DS, Myers AM (1995) Characterization of the maize gene sugary1, a determinant of starch composition in kernels. Plant Cell 7:417–429
Janeček Š, Svensson B, MacGregor EA (2011) Structural and evolutional aspects of two families of non-catalytic domains present in starch and glycogen binding proteins from microbes, plants and animals. Enzym Microb Technol 49:429–440
Jenkins PJ, Cameron RE, Donald AM (1993) A universal feature in the starch granules from different botanical sources. Starch 45:417–420
Kainuma K, French D (1972) Naegeli amylodextrin and its relationship to starch granule structures. III Role of water in crystallization of B-starch. Biopolymers 11:2241–2250
Katayama K, Komae K, Kohyama K et al (2002) New sweet potato line having low gelatinization temperature and altered starch structure. Starch 54:51–57
Katsuya Y, Mezaki Y, Kubota M et al (1998) Three-dimensional structure of Pseudomonas isoamylase at 2.2 Ǻ resolution. J Mol Biol 281:885–897
Kawagoe Y (2013) The characteristic polyhedral, sharp-edged shape of compound-type starch granules in rice endosperm is achieved via the septum-like structure of the amyloplast. J Appl Glycosci 60:29–36
Kitahara K, Fukunaga S, Katayama K et al (2005) Physicochemical properties of sweet potato starches with different gelatinization temperatures. Starch 57:473–479
Klucinec JD, Thompson DB (2002) Structure of amylopectins from ae-containing maize starches. Cereal Chem 79:19–23
Kubo A, Fujita N, Harada K et al (1999) The starch-debranching enzymes isoamylase and pullulanase are both involved in amylopectin biosynthesis in rice endosperm. Plant Physiol 121:399–409
Kubo A, Rahman S, Utsumi Y et al (2005) Complementation of sugary-1 phenotype in rice endosperm with the wheat Isoamylase1 gene supports a direct role for isoamylase1 in amylopectin biosynthesis. Plant Physiol 137:43–56
Kubo A, Colleoni C, Dinges JR et al (2010) Functions of heteromeric and homomeric isoamylase-type starch debranching enzymes in developing maize endosperm. Plant Physiol 153:956–969
Larsson CT, Hofvander P, Khoshnoodi J et al (1996) Three isoforms of starch synthase and two isoforms of branching enzyme are present in potato tuber starch. Plant Sci 117:9–16
Lee EYC, Whelan WJ (1971) Glycogen and starch debranching enzymes. In: Boyer PD (ed) The enzymes, vol 5. Academic, New York, pp 191–234
Leterrier M, Holappa L, Broglie KE et al (2008) Cloning, characterization and comparative analysis of a starch synthase IV gene in wheat: functional and evolutional implications. BMC Plant Biol 8:98
Li Z, Mouille G, Kosar-Hashemi B et al (2000) The structure and expression of the wheat starch synthase III gene: motif in the expressed gene define the lineage of the starch synthase III gene family. Plant Physiol 123:613–624
Li L, Jiang H, Campbell M, et al. (2008) Characterization of maize amylose-extender (ae) mutant starches. Part I: Relationship between resistant starch contents and molecular structures. Carbohydr Polym 74:396–404
Li Z, Li D, Du X et al (2011) The barley amo1 locus is tightly linked to the starch synthase IIIa gene and negatively regulates expression of granule-bound starch synthetic genes. J Exp Bot 62:5217–5231
Lin Q, Huang B, Zhang M et al (2012) Functional interactions between starch synthase III and isoamylase-type starch-debranching enzyme in maize endosperm. Plant Physiol 158:679–692
Lin Q, Facon M, Putaux JL et al (2013) Function of isoamylase-type starch debranching enzymes ISA1 and ISA2 in the Zea mays leaf. New Phytol 200:1009–1021
Liu F, Ahmed Z, Lee EA et al (2012a) Allelic variants of the amylose-extender mutation of maize demonstrate phenotypic variation in starch structure resulting from modified protein-protein interactions. J Exp Bot 63:1167–1183
Liu F, Romanova N, Lee EA et al (2012b) Glucan affinity of starch synthase IIa determines binding of starch synthase I and starch-branching enzyme IIb to starch granules. Biochem J 448:373–387
Lloyd JR, Landschütze V, Kossman J (1999) Simultaneous antisense inhibition of two starch-synthase isoforms in potato tubers leads to accumulation of grossy modified amylopectin. Biochem J 338:515–521
Maddelein ML, Libessart N, Bellanger F et al (1994) Toward an understanding of the biogenesis of the starch granule. J Biol Chem 269:25150–25157
Mangelsdorf PC (1947) The inheritance of amylaceous sugary endosperm and its derivatives in maize. Genetics 32:448–458
Marshall J, Sidebottom C, Debet M et al (1996) Identification of the major starch synthase in the soluble fraction of potato tuber. Plant Cell 8:1121–1135
McMaugh SJ, Thistleton JL, Anschaw E et al (2014) Suppression of starch synthase I expression affects the granule morphology and granule size and fine structure of starch in wheat endosperm. J Exp Bot 65:2189–2201
Mizuno K, Kawasaki T, Shimada H et al (1993) Alteration of the structural properties of starch components by the lack of an isoform of starch branching enzyme in rice seeds. J Biol Chem 268:19084–19091
Mizuno K, Kobayashi E, Tachibana M et al (2001) Characterization of an isoform of rice starch branching enzyme, RBE4, in developing seeds. Plant Cell Physiol 42:349–357
Momma M, Fujimoto Z (2012) Interdomain disulfide bridge in the rice granule bound starch synthase I catalytic domain as elucidated by X-ray structural analysis. Biosci Biotechnol Biochem 76(8):1591–1595
Morell M, Blennow A, Kosar-Hashemi B (1997) Differential expression and properties of starch branching enzyme isoforms in developing wheat endosperm. Plant Physiol 113:201–208
Morell MK, Kosar-Hashemi B, Cmiel M et al (2003) Barley sex6 mutants lack starch synthase IIa activity and contain a starch with novel properties. Plant J 34:173–185
Mouille G, Maddelein M-L, Libessart N et al (1996) Preamylopectin processing: a mandatory step for starch biosynthesis in plants. Plant Cell 8:1353–1366
Mu-Forster C, Huang R, Powers JR et al (1996) Physical association of starch biosynthetic enzymes with starch granules of maize endosperm. Plant Physiol 111:821–829
Murakami T, Kanai T, Takata H et al (2006) A novel branching enzyme of the GH-57 family in the hyperthermophilic archaeon Thermococcus kodakaraensis KODI. J Bacteriol 188:5915–5924
Murata T, Sugiyama T, Akazawa T (1965) Enzymic mechanism of starch synthesis in glutinous rice grains. Biochem Biophys Res Commun 18:371–376
Mutisya J, Sathish P, Sun C et al (2003) Starch branching enzymes in sorghum (Sorghum bicolor) and barley (Hordeum vulgare): Comparative analyses of enzyme structure and gene expression. J Plant Physiol 160:921–930
Myers AM, Morell MK, James MG et al (2000) Recent progress toward understanding biosynthesis of the amylopectin crystal. Plant Physiol 122:989–997
Nakamura Y (1996) Some properties of starch debranching enzymes and their possible role in amylopectin biosynthesis. Plant Sci 121:1–18
Nakamura Y (2002) Towards a better understanding of the metabolic system for amylopectin biosynthesis plants: Rice endosperm as a model tissue. Plant Cell Physiol 43:718–725
Nakamura Y (2014) Mutagenesis and transformation of starch biosynthesis of rice and the production of novel starches. In: Tomlekova N, Kozgar I, Wani R (eds) Mutagenesis: exploring novel genes and pathways. Wageningen, Wageningen Academic, pp 251–278
Nakamura Y, Yuki K, Park SY et al (1989) Carbohydrate metabolism in the developing endosperm of rice grains. Plant Cell Physiol 30:833–839
Nakamura Y, Takeichi T, Kawaguchi K et al (1992a) Purification of two forms of starch branching enzyme (Q-enzyme) from developing rice endosperm. Physiol Plant 84:329–335
Nakamura Y, Umemoto T, Takahata Y et al (1992b) Characteristics and roles of key enzymes associated with starch biosynthesis in rice endosperm. Gamma Field Symp 31:25–44
Nakamura Y, Umemoto T, Takahata Y et al (1996) Changes in structure of starch and enzyme activities affected by sugary mutations in developing rice endosperm. Possible role of starch debranching enzyme (R-enzyme) in amylopectin biosynthesis. Physiol Plant 97:491–498
Nakamura Y, Kubo A, Shimamune T et al (1997) Correlation between activities of starch debranching enzyme (R-enzyme or pullulanase) and α-glucan structure in endosperms of sugary-1 mutants of rice. Plant J 12:143–153
Nakamura T, Vrinten P, Hayakawa K et al (1998) Characterization of a granule-bound starch synthase isoform found in the pericarp of wheat. Plant Physiol 118:451–459
Nakamura Y, Sakurai A, Inaba Y et al (2002) The fine structure of amylopectin in endosperm from Asian cultivated rice can be largely classified into two classes. Starch 54:117–131
Nakamura Y, Francisco PB Jr, Hosaka Y et al (2005a) Essential amino acids of starch synthase IIa differentiate amylopectin structure and starch quality between japonica and indica rice varieties. Plant Mol Biol 58:213–227
Nakamura Y, Takahashi J, Sakurai A et al (2005b) Some cyanobacteria synthesize semi-amylopectin type α-polyglucan instead of glycogen. Plant Cell Physiol 46:539–545
Nakamura Y, Fujita N, Utsumi Y et al (2009) Revealing the complex system of starch biosynthesis in higher plants using rice mutants and transformants. In: Shu Q (ed) Induced mutations in the genomics era. Food and Agriculture Organization of the United Nations, Rome, pp 165–167
Nakamura Y, Utsumi Y, Sawada T et al (2010) Characterization of the reactions of starch branching enzymes from rice endosperm. Plant Cell Physiol 51:776–794
Nakamura Y, Ono M, Utsumi Y et al (2012) Functional interaction between plastidial starch phosphorylase and starch branching enzymes from rice during the synthesis of branched maltodextrins. Plant Cell Physiol 53:869–878
Nakamura Y, Aihara S, Crofts N et al (2014) In vitro studies of enzymatic properties of starch synthases and interactions between starch synthase I and starch branching enzymes from rice. Plant Sci 224:1–8
Nelson OE, Rines HW (1962) The enzymatic deficiency in the waxy mutant of maize. Biochem Biophys Res Commun 9:297–300
Nishi A, Nakamura Y, Tanaka N et al (2001) Biochemical and genetic analysis of the effects of amylose-extender mutation in rice endosperm. Plant Physiol 127:459–472
Noguchi J, Chaen K, Vu NT et al (2011) Crystal structure of the branching enzyme I (BEI) from Oryza sativa L with implications for catalysis and substrate binding. Glycobiology 21:1108–1116
Nozaki K, Hamada S, Nakamori T et al (2001) Major isoforms of starch branching enzymes in premature seeds of kidney bean (Phaseolus vulgaris L.). Biosci Biotechnol Biochem 65:1141–1148
Ohdan T, Francisco PB Jr, Hosaka Y et al (2005) Expression profiling of genes involved in starch synthesis in sink and source organs of rice. J Exp Bot 56:3229–3244
Ohdan T, Sawada T, Nakamura Y (2011) Effects of temperature on starch branching enzyme properties of rice. J Appl Glycosci 58:19–26
Ozbun JL, Hawker JS, Preiss J (1971a) Multiple forms of α-1,4 glucan synthetase from spinach leaves. Biochem Biophys Res Commun 43:631–636
Ozbun JL, Hawker JS, Preiss J (1971b) Adenosine diphosphoglucose-starch glucosyl transferases from developing kernels of waxy maize. Plant Physiol 48:765–769
Ozbun JL, Hawker JS, Preiss J (1972) Soluble adenosine diphosphate glucose-arufa-1,4-glucan arufa-4-glucosyltransferases from spinach leaves. Biochem J 126:953–963
Palomo M, Kralj S, van der Maarel MJEC et al (2009) The unique branching pattern of Deinococcus glycogen branching enzymes are determined by their N-terminal domains. Appl Environ Microbiol 75:1355–1362
Pan D, Nelson NE (1984) A debranching enzyme deficiency in endosperms of sugary-1 mutants of maize. Plant Physiol 74:324–328
Patron NJ, Keeling PJ (2005) Common evolutionary origin of starch biosynthetic enzymes in green and red algae. J Phycol 41:1131–1141
Peng C, Wang Y, Liu F et al (2014) FLOURY ENDOSPERM6 encodes a CBM48 domain-containing protein involved in compound granule formation and starch synthesis in rice endosperm. Plant J 77:917–930
Pérez S, Bertoft E (2010) The molecular structures of starch components and their contribution to the architecture of starch granules: a comprehensive review. Starch 62:389–420
Pfister B, Lu K, Eicke S, et al (2014) Genetic evidence that chain length and branch point distributions are linked determinants of starch granule formation in Arabidopsis. Plant Physiol 165:1457–1474
Pollock C, Preiss J (1980) The citrate-stimulated starch synthase of starchy maize kernels: Purification and properties. Arch Biochem Biophys 204:578–588
Rahman S, Kosar-Hashemi B, Samuel MS et al (1995) The major proteins of wheat starch granules. Aust J Plant Physiol 22:793–803
Rahman S, Regina A, Li Z et al (2001) Comparison of starch-branching enzyme genes reveals evolutionary relationships among isoforms. Characterization of a gene for starch-branching enzyme IIa from the wheat D genome donor Aegilops tauschii. Plant Physiol 125:1314–1324
Rahman S, Nakamura Y, Li Z et al (2003) The sugary-type isoamylase gene from rice and Aegilops tauschii: characterization and comparison with maize and Arabidopsis. Genome 46:496–506
Ral JP, Colleoni C, Wattebled F et al (2006) Circadian clock regulation of starch metabolism establishes GBSSI as a major contributor to amylopectin synthesis in Chlamydomonas reinhardtii. Plant Physiol 142:305–317
Regina A, Kosar-Hashemi B, Li Z et al (2005) Starch branching enzyme IIb in wheat is expressed at low levels in the endosperm compared to other cereals and encoded at a non-syntenic locus. Planta 222:899–909
Regina A, Bird A, Topping D et al (2006) High-amylose wheat generated by RNA interference improves indices of large-bowel health in rats. Proc Natl Acad Sci U S A 103:3546–3551
Regina A, Kosar-Hashemi B, Ling S et al (2010) Control of starch branching in barley defined through differential RNAi suppression of starch branching enzyme IIa and IIb. J Exp Bot 61:1469–1482
Roldán L, Wattebled F, Lucas MM et al (2007) The phenotype of soluble starch synthase IV defective mutant of Arabidopsis thaliana suggests a novel function of elongation enzymes in the control of starch granule formation. Plant J 49:492–504
Rundle RE, Daasch L, French D (1944) The structure of the “B” modification of starch from film and fiber diffraction diagrams. J Am Chem Soc 66:130–134
Rydberg U, Andersson L, Andersson R et al (2001) Comparison of starch branching enzyme I and II from potato. Eur J Biochem 268:6140–6145
Satoh H, Nishi A, Yamashita K et al (2003) Starch-b ranching enzyme I-deficient mutation specifically affects the structure and properties of starch in rice endosperm. Plant Physiol 133:1111–1121
Satoh H, Shibahara K, Tokunaga T et al (2008) Mutation of the plastidial α-glucan phosphorylase gene in rice affects the synthesis of branched maltodextrins. Plant Cell 20:1833–1849
Sawada T, Francisco PB Jr, Aihara S et al (2009) Chlorella starch branching enzyme II (BEII) can complement the function of BEIIb in rice endosperm. Plant Cell Physiol 50:1062–1074
Sawada T, Nakagami T, Utsumi Y et al (2013) Characterization of starch and glycogen branching enzymes from various sources. J Appl Glycosci 60:69–78
Sawada T, Nakamura Y, Ohdan T et al (2014) Diversity of reaction characteristics of glucan branching enzymes and the fine structure of α-glucan from various sources. Arch Biochem Biophys 562:9–21
Schwarte S, Brust H, Steup M et al (2013) Intraspecific sequence variation and differential expression in starch synthase genes of Arabidopsis thaliana. BMC Res Notes 6:84
Senoura T, Isono N, Yoshikawa M et al (2007) Characterization of starch synthase I and II expressed in early developing seeds of kidney bean (Phaseolus vulgaris L.). Biosci Biotechnol Biochem 68:1949–1960
Sestili F, Janni M, Doherty A et al (2010) Increasing the amylose content of durum wheat through silencing of the SBEIIa genes. BMC Plant Biol 10:144
Shannon JC, Garwood DL (1984) Genetics and physiology of starch development. In: Whistler RL, BeMiller JN, Paschall EF (eds) Starch: chemistry and technology, 2nd edn. Academic, New York, pp 25–86
Shannon JC, Pein FM, Cao HP et al (1998) Brittle-1, an adenylate translocator, facilitates transfer of extraplastidial synthesized ADP-glucose into amyloplasts of maize endosperms. Plant Physiol 117:1235–1252
Shannon JC, Garwood DL, Boyer CD (2007) Genetics and physiology of starch development. In: BeMiller JN, Whistler RL (eds) Starch: chemistry and technology, 3rd edn. Academic, New York, pp 23–82
She K, Kusano H, Koizumi K et al (2010) A novel factor FLOURY ENDOSPERM2 is involved in regulation of rice grain size and starch quality. Plant Cell 22:3280–3294
Shimonaga T, Fujiwara S, Kaneko M et al (2006) Variation in storage α-polyglucans of red algae: Amylose and semi-amylopectin-types in Porphyridium and glycogen-type in Cyanidium. Mar Biotechnol 9:192–202
Shimonaga T, Konishi M, Oyama Y et al (2008) Variation in storage α-glucans of the Porphyridiales (Rhodophyta). Plant Cell Physiol 49:103–116
Sim L, Beeren SR, Findinier J, et al (2014) Crystal structure of the Chlamydomonas starch debranching enzyme isoamylase ISA1 reveals insights into the mechanism of branch trimming and complex assembly. J Biol Chem 289:22991–23003
Smith AM (1988) Major differences in isoforms of starch-branching enzyme between developing embryos of round- and wrinkled-seeded peas (Pisum sativum L.). Planta 175:270–279
Smith SM, Fulton DC, Chia T et al (2004) Diurnal changes in the transcriptome encoding enzymes of starch metabolism provide evidence for both transcriptional and posttranscriptional regulation of starch metabolism in Arabidopsis leaves. Plant Physiol 136:2687–2699
Stam MR, Danchin EGJ, Ranchured C et al (2006) Dividing the large glycoside hydrolase family 13 into subfamilies: towards improved functional annotations of amylase-related proteins. Protein Eng Des Sel 19:555–562
Streb S, Zeeman SC (2014) Replacement of the endogenous starch debranching enzymes ISA1 and ISA2 of Arabidopsis with the rice orthologs reveals a degree of functional conservation during starch synthesis. PLoS One 9:e92174
Streb S, Delatte T, Umhang M et al (2008) Starch granule biosynthesis in Arabidopsis is abolished by removal of all debranching enzymes but restored by the subsequent removal of an endoamylase. Plant Cell 20:3448–3466
Sullivan TD (1995) The maize brittle1 gene encodes amyloplast membrane polypeptides. Planta 196:477–484
Sun C, Sathish P, Ahlandsberg S et al (1998) The two genes encoding starch-branching enzymes IIa and IIb are differentially expressed in barley. Plant Physiol 118:37–49
Szydlowsky N, Ragel P, Raynaud S et al (2009) Starch granule initiation in Arabidopsis requires the presence of either class IV or class II starch synthases. Plant Cell 21:2443–2457
Szydlowsky N, Ragel P, Hennen-Bierwagen TA (2011) Integrated functions among multiple starch synthases determine both amylopectin chain length and branch linkage location in Arabidopsis leaf starch. J Exp Bot 62:4547–4559
Takashima Y, Senoura T, Yoshizaki T et al (2007) Differential chain-length specificities of two isoamylase-type starch debranching enzymes from developing seeds of kidney bean. Biosci Biotechnol Biochem 71:2308–2312
Takeda Y, Hizukuri S, Juliano BO (1987a) Structure of rice amylopectins with low and high affinities for iodine. Carbohydr Res 168:79–88
Takeda Y, Hizukuri S, Takeda C et al (1987b) Structures of branched molecules of amyloses of various botanical origins, and molar fractions of branched and unbranched molecules. Carbohydr Res 165:139–145
Takeda Y, Guan H, Preiss J (1993) Branching of amylose by the branching isoenzymes of maize endosperm. Carbohydr Res 240:253–263
Takeda Y, Preiss J (1993) Structures of B90 (sugary) and W64A (normal) maize starches. Carbohydr Res 240:265–275
Tanaka Y, Akazawa T (1971) Enzymic mechanism of starch synthesis in ripening rice grains VI. Isozymes of starch synthase. Plant Cell Physiol 12:493–505
Tanaka N, Fujita N, Nishi A et al (2004) The structure of starch can be manipulated by changing expression levels of starch branching enzyme IIb in rice endosperm. Plant Biotechnol J 2:507–516
Tetlow I (2012) Branching enzymes and their role in determining structural and functional properties of polyglucan. In: Tetlow I (ed) Starch: origins, structure and metabolism, vol 5, Essential reviews in experimental biology. The Society for Experimental Biology, London, pp 141–177
Tetlow IJ, Wait R, Lu Z et al (2004) Protein phosphorylation in amyloplasts regulates starch branching enzyme activity and protein-protein interactions. Plant Cell 16:694–708
Tetlow IJ, Beisel KG, Cameron S et al (2008) Analysis of protein complexes in amyloplasts reveals functional interactions among starch biosynthetic enzymes. Plant Physiol 146:1878–1891
Thompson DB (2000) On the non-random nature of amylopectin branching. Carbohydr Polym 43:223–239
Tomlinson KL, Lloyd JR, Smith AM (1997) Importance of isoforms of starch-branching enzyme in determining the structure of starch in pea leaves. Plant J 11:31–43
Tomlinson KL, Lloyd JR, Smith AM (1998) Major differences in isoform composition of starch synthase between leaves and embryos of pea (Pisum sativum L.). Planta 204:86–92
Toyosawa Y, Kawagoe Y, Matsushima R, et al. (2015) Deficiency of starch synthase IIIa and IVb leads to dramatic changes in starch granule morphology in rice endosperm (submitted)
Umemoto T, Nakamura Y, Satoh H et al (1999) Differences of amylopectin structure between two rice varieties in relation to the effects of temperature during grain-filling. Starch 51:58–62
Umemoto T, Yano M, Satoh H et al (2002) Mapping of a gene responsible for the difference in amylopectin structure between japonica-type and indica-type rice varieties. Theor Appl Genet 104:1–8
Usui T, Ogata M, Murata T et al (2009) Sequential analysis of α-glucooligosaccharides with α-(1–4) and α-(1–6) linkages by negative ion Q-TOF MS/MS spectrometry. J Carbohydr Chem 28:421–430
Utsumi Y, Nakamura Y (2006) Structural and enzymatic characterization of the isoamylase1 homo-oligomer and the isoamylase1-isoamylase2 hetero-oligomer from rice endosperm. Planta 225:75–87
Utsumi Y, Utsumi C, Sawada T et al (2011) Functional diversity of isoamylase oligomers: The ISA1 homo-oligomer is essential for amylopectin biosynthesis in rice endosperm. Plant Physiol 156:61–77
Vester-Christensen MB, Hachem MA, Svensson B et al (2010) Crystal structure of an essential enzyme in seed starch degradation: barley limit dextrinase in complex with cyclodextrins. J Mol Biol 403:739–750
Vikso-Nielsen A, Blennow A (1998) Isolation of starch branching enzyme I from potato using γ-cyclodextrin affinity chromatography. J Chromatogr 800:382–385
Visser RGF, Somhorst I, Kuipers GJ et al (1991) Inhibition of the expression of the gene for granule-bound starch synthase in potato by antisense constructs. Mol Gen Genet 225:289–296
Wang J, Xu H, Zhu Y et al (2013) OsbZIP58, a basic leucine zipper transcription factor, regulates starch biosynthesis in rice endosperm. J Exp Bot 64:3453–3466
Wattebled F, Ral JP, Dauvillée D et al (2003) STA11, a Chlamydomonas reinhardtii locus required for normal starch granule biogenesis, encodes disproportionating enzyme. Further evidence for a function of α-1,4 glucanotransferases during starch granule biosynthesis in green algae. Plant Physiol 132:137–145
Wattebled F, Dong Y, Dumez S et al (2005) Mutants of Arabidopsis lacking a chloroplastic isoamylase accumulates phytoglycogen and an abnormal form of amylopectin. Plant Physiol 138:184–195
Wattebled F, Planchot V, Dong Y et al (2008) Further evidence for the mandatory nature of polysaccharide debranching for the aggregation of semicrystalline starch and for overlapping functions of debranching enzymes in Arabidopsis leaves. Plant Physiol 148:1309–1323
Weber APM, Linka N (2011) Connecting the plastids: Transporters of the plastid envelope and their role in linking plastidial with cytosolic metabolism. Annu Rev Plant Biol 62:53–77
Xia H, Yandeau-Nelson M, Thompson DB et al (2011) Deficiency of maize starch-branching enzyme I results in altered starch fine structure, decreased digestibility and reduced coleoptiles growth during germination. BMC Plant Biol 11:95
Yamamori M, Fujita S, Hayakawa K et al (2000) Genetic elimination of a starch granule protein, SGP-1, of wheat generates an altered starch with apparent high amylose. Theor Appl Genet 101:21–29
Yamanouchi H, Nakamura Y (1992) Organ specificity of isoforms of starch branching enzyme (Q-enzyme) in rice. Plant Cell Physiol 33:985–991
Yao Y, Thompson DB, Guiltinan MJ (2004) Maize starch-branching enzyme isoforms and amylopectin structure. In the absence of starch-branching enzyme IIb, the further absence of starch-branching enzyme Ia leads to increased branching. Plant Physiol 136:3515–3523
Yun M, Umemoto T, Kawagoe Y (2011) Rice debranching enzyme isoamylase3 facilitates starch metabolism and affects plastid morphogenesis. Plant Cell Physiol 52:1068–1082
Zeeman SC, Umemoto T, Lue W et al (1998) A mutant of Arabidopsis lacking a chloroplastic isoamylase accumulates both starch and phytoglycogen. Plant Cell 10:1699–1711
Zeeman SC, Kossman J, Smith AM (2010) Starch: its metabolism, evolution, and biotechnological modification in plants. Annu Rev Plant Biol 61:209–234
Zhang X, Colleoni C, Ralushana V et al (2004) Molecular characterization demonstrates that the Zea mays gene sugary2 codes the starch synthase isoform SSIIa. Plant Mol Biol 54:865–879
Zhang X, Myers AM, James MG (2005) Mutations affecting starch synthase III in Arabidopsis alter leaf starch structure and increase the rate of starch synthesis. Plant Physiol 138:663–674
Zhang X, Szydlowski N, Delvallé D et al (2008) Overlapping functions of the starch synthase SSII and SSIII in amylopectin biosynthesis in Arabidopsis. BMC Plant Biol 8:96–113
Acknowledgments
The author thanks Dr. Perigio B. Francisco, Jr. for critical reading of the manuscript and polishing the English used in the paper. The author also thanks Dr. Takayuki Sawada for the assistance with some figures.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Japan
About this chapter
Cite this chapter
Nakamura, Y. (2015). Biosynthesis of Reserve Starch. In: Nakamura, Y. (eds) Starch. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55495-0_5
Download citation
DOI: https://doi.org/10.1007/978-4-431-55495-0_5
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55494-3
Online ISBN: 978-4-431-55495-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)