Abstract
Proteolysis is arguably the most important of all post-translational modifications in that it is effectively irreversible, and leads to the partial or complete breakdown of proteins. Cells have adapted various mechanisms to control proteolysis as despite its importance, its destructive nature requires tight regulation. Evidence now clearly shows that the loss of proteolytic regulation is a key component in the progression of various pathophysiologies from metabolic disorders to cancer. This chapter discusses the central role for cellular compartmentalization of proteolysis to ensure homeostasis and highlights examples of how the loss of this regulation can promote disease.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
3.1 Introduction
Compartmentalization of proteolysis is essential for homeostasis in all tissues and organs, not only at the level of communication of cells with their environment, but also at the sub-cellular level (Fig. 3.1). Within the context of intracellular compartmentalization, it is important to consider the definition of organelles as sub-cellular regions limited by biological membranes that form distinct compartments featuring specific biochemical environments (Fig. 3.2).
Schematic representation of compartmentalized proteolysis. Proteases perform their tasks at different locations within cells or tissues. Functions of distinct proteolytic enzymes differ depending on the position they fill in the complex network of proteases and where those meet their natural substrates. Compartmentalization occurs at the level of tissues consisting of different cell types (a) and at the sub-cellular level (b). Regulation of proteolysis in space and time is possible by means of different distributions of proteases, their activators, their co-factors, their inhibitors and their substrates (c). Thus, compartmentalization of proteolysis is influenced by signalling events and thereby also depends on the cross-talk of cells with each other and their environment. Bolt signifies a signal from the environment
Schematic representation of compartmentalization based on biochemical features of cellular organelles. Peptidases are faced with strikingly different biochemical conditions for substrate interaction and proteolytic cleavage depending on whether extra- or intracellular proteolysis takes place. The latter processes depend on where within the cellular compartments proteases engage in substrate cleavage; a number of examples of different biochemical conditions of proteolytic processing are depicted including oxidizing versus reducing, and acidic versus basic or neutral conditions. The ionic strength of certain compartments, e.g. the secretory granules, or the presence of metal ions are further well known factors contributing to the regulation of the activities of proteolytic enzymes
Compartmentalization, in a cellular context, distinguishes hydrophilic sub-cellular regions of defined properties that are separated from one another by hydrophobic membranes (Palade 1964, 1966; Clegg 1991; Lipowsky 1995). Proteolysis, the catalytic hydrolysis (and thus cleavage) of peptide bonds, was traditionally thought to require hydrophilic environments (Gerlt and Babbitt 2001). Lipid-enriched, hydrophobic conditions were not considered conducive to proteolytic catalysis where a molecule of water plays a central role (Chaps. 1 and 2; Brown et al. 2000). However, we have learned during the past decades that such (strict) proposals are too narrow and no longer fully valid (Weihofen and Martoglio 2003; Wolfe and Kopan 2004; Ha 2007). Undisputedly, evolution has found a huge variety of solutions to the demands of our body in modifying proteins—be they folded or unfolded, monomeric or multimeric—by protease-mediated hydrolysis that takes place in all possible environments including even the lipid bilayer itself (Chaps. 4 and 7–15; Fritz et al. 1973; Steiner et al. 1980; Fricker 1988; Travis and Fritz; 1991; Stubbs and Bode 1994; Bode et al. 1997; Chapman et al. 1997; Khan and James 1998; Galivan et al. 2000; Davies 2001; Chambers and Laurent 2002; Hooper 2002; Seiki 2002; Potempa et al. 2003; Schechter 2005; Arolas et al. 2007; Barrett and Rawlings 2007; Brix et al. 2008; Cobbe et al. 2009; Ovaere et al. 2009; Schweitzer and Naumann 2010).
Proteases were first defined by their presumably predominant functions as protein degrading enzymes. However, it has been conclusively shown that many proteases undertake a range of functions—and not simply as mere destructive enzymes. Although some proteases are essential in protein turnover, ensuring removal of unwanted or damaged proteins and producing pools of amino acids for de novo protein biosynthesis (Bonifacino and Weissman 1998; Reed 2003; Ciechanover and Iwai 2004), discrete cleavage or post-translational processing of target proteins is just as important. These processing events can promote protein folding, maturation, or achieve initiation, termination or modulation of protein functionality (Dobson 2003; Ciechanover 2005; Collins and Tansey 2006). They also engage in peptide processing, for example in antigen presentation, in the cellular defence of intruders and pathogens (Fineschi and Miller 1997; Nakagawa et al. 1998; Villadangos et al. 1999; Riese and Chapman 2000; Trombetta and Mellman 2005; Herget and Tampe 2007; Watts 2012). Additionally, it is worthy to note that some proteases can also have non-proteolytic roles, a topic that requires additional attention as we attempt to grasp the complexity of the world of proteolytic enzymes beyond their active sites (Chaps. 2, 5, 6, and 8; Reudelhuber et al. 1998; Blasi and Carmeliet 2002; Friedl and Wolf 2003; Yu et al. 2006; Lamkanfi et al. 2007; Uddin et al. 2008; Strongin 2010; Bhat and Greer 2011; Kwak et al. 2011).
Proteases can act on substrates in a number of ways: in a simple one-to-one fashion, or to cut specific bonds in complex substrates consisting of several proteins, or to sever high molecular mass proteins containing multiple domains. More than one protease may cleave a specific peptide bond in any one substrate at a time, further adding to the complexity of proteolytic control. It may be of further relevance to note that proteases can act as single molecules or as oligomers (e.g. the 26S proteasome). Increasingly, we are finding evidence for the need for proteases to complex with activators or co-factors (Fig. 3.3). Although these co-factors can be other proteins, they may also comprise lipids (invadolysin, secretases, SPPases) (McHugh et al. 2004; Cobbe et al. 2009; Di Cara et al. 2013), carbohydrates (cathepsin K, ADAMs, ADAM-TSs) or even nucleic acids (truncated cathepsin V, separases) (Chaps. 1 and 2; Bode and Huber 1992, 2000; Uhlmann 2003; Ong et al. 2007; Sun et al. 2009). Proteases may position themselves in a particular cellular context by interacting with scaffolding factors, for example in apoptosis (Chap. 8; Seiki 2002; Schweitzer and Naumann 2010; Kersse et al. 2011), or they may traffic along traditional and novel, sometimes unexpected, transport pathways to reach the location required for proteolytic activity (e.g. endo-lysosomal enzymes, “alternative” secretion of some convertases) (Chap. 4; Linke et al. 2002a; Buth et al. 2004, 2007; Brix and Jordans 2005; Sloane et al. 2006; Brix et al. 2008; Sameni et al. 2009) (Figs. 3.2, 3.4, and 3.5). Proteolytic activity may be controlled by physical compartmentalization of the protease as well as the presence of specific endogenous inhibitor(s).
Schematic representation of compartmentalization by lipid membranes, scaffolding factors, and complex formation. Proteases act as single molecules, in complexes, or in sequential modes. Proteolytic enzymes may be soluble, equipped with transmembrane domains, or interact with receptors and other scaffolding factors eventually initiating, terminating, or otherwise affecting the extent of substrate cleavage. Some proteases form multimeric complexes with other macromolecules such as other proteins, glycoproteins or proteoglycans in the extra- and pericellular space. In contrast, the formation of macromolecular assemblies within the cytosol results in substrate processing in e.g. the proteasome which provides a cleavage chamber with specific conditions, thus, separating the protease-substrate interaction from the surrounding cytosol without using the principle of compartmentalization by lipid membranes. Most proteolytic processes require hydrophilic conditions whereas others are enabled to take place near or at the level of lipid membranes for cleaving signal peptides or to carry out proteolysis within transmembrane domains
Schematic representation of different principles of compartmentalization. (a) Proteolytic enzymes of mammalian cells are often synthesized as inactive precursors, the zymogen forms (1), that are matured by proteolytic processing or other means at the sub-cellular location where the active protease is required to function (2). In contrast, compartments equipped with inactivating factors such as endogenous protease inhibitors provide conditions that restrict proteolytic cleavages (3). Shuttling between the different compartments is realized by directed protein transport processes ensuring protease trafficking to the desired location for meeting with their substrates at a given time (left panels). In some cases, individual compartments house different forms of the same proteolytic enzymes which are regulated in their activity by interaction with activating or inhibiting factors to initiate or terminate proteolytic substrate cleavage (right panel). (b) Proteases belong to distinct protein families (for classification see, http://merops.sanger.ac.uk) which are sorted into different (left) or the same sub-cellular compartments of mammalian cells (right)
Compartmentalization of cathepsins in mammalian cells. Confocal fluorescence micrographs of cryosections prepared from mouse tissue (a–c) and of cultured human cells (d and e) after immunolabelling with antibodies specific for the aspartic protease cathepsin D or the cysteine peptidases cathepsins B, K, L, or V, and fluorophore-conjugated secondary antibodies (green). Nuclei were counter-stained with Draq 5 (red in a, c, and d; blue in e) and tissue structures viewed in phase contrast (b); overlays of the single channels are displayed in false colours. Although all cathepsins depicted in this figure belong to the so-called “endo-lysosomal proteases”, their distribution patterns differ dramatically between the tissues and individual cell types. (a) Cathepsin L is detectable within vesicles gathering in the peri-nuclear regions of fibroblasts (F) and macrophages (Mø) of the lamina propria as well as of enteroendocrine cells (En) of the mucosal epithelium in mouse intestine (arrows). While this staining pattern is expected from the classical view on this member of the papain-like peptidase family, cathepsin L is almost absent from M-cells (M) and even secreted from the enterocytes (E) and goblet cells (G) of the mucosa (double headed arrows), from where it reaches the intestinal lumen and re-associates with the apical plasma membrane of the intestine epithelial cells (open arrowheads). These distribution patterns point to different functions that cathepsin L fulfils in the various cell types found in the intestinal mucosa. (b and c) Cathepsins D and K belong to different families of enzymes since they act as aspartic or cysteine peptidases, respectively. The epithelial cells of the thyroid gland sort both proteases into vesicles (arrows) that differ in size and are destined to either the apical (b) or the basolateral poles of thyroid epithelial cells depending on their physiological states (c). Both of these cathepsins together with a number of other related peptidases cleave the prohormone thyroglobulin by endo- and exopeptidatic modes, thereby liberating the thyroid hormones 3′,3,5-triiodothyronine T3 and thyroxine T4. The precise cleavage patterns of the substrate depend on the position and activity of its processing enzyme in the various compartments of the endocytic pathway of thyrocytes and within the extracellular lumen of thyroid follicles. Thus, transport of proteases and substrate cleavage appear compartmentalized and tightly regulated in the polarized epithelial cells of the thyroid gland. (d and e) Human keratinocytes exhibit vesicular staining of the cysteine cathepsin B (arrows in d) whereas the cysteine cathepsin V is additionally and predominantly detected in an ER-like reticular pattern throughout the cytosol of HaCaT cells (circles in e). These localizations point to different maturation stages of the two cysteine cathepsins resulting from distinct transport routes under steady state conditions. Sub-compartmentalization is even more complex because molecular variants of cathepsin B and, even more prominent, cathepsin V variants lacking the signal peptides are detectable in the nuclei of proliferating keratinocytes (arrowheads in e). N denotes nuclei, and N* indicates an apoptotic cell in which the subcellular compartments appear condensed and structurally altered in comparison to the normal appearance of organelles in this keratinocyte cell line. For further details see references (Brix and Herzog 1994; Brix et al. 1996, 1997; Lemansky et al. 1998; Tepel et al. 2000; Linke et al. 2002a, b; Friedrichs et al. 2003; Buth et al. 2004, 2007; Mayer et al. 2006, 2008, 2009; Jordans et al. 2009; Vreemann et al. 2009; Tedelind et al. 2010, 2011; Arampatzidou et al. 2011a, 2012; Dauth et al. 2011b, 2012; Haugen et al. 2013; reviewed in Brix et al. 2001, 2008, 2011; Brix 2005; Brix and Jordans 2005; Arampatzidou et al. 2011b; Dauth et al. 2011a)
In this chapter, we highlight some of the facets of compartmentalization of proteolysis—without claiming to be comprehensive in discussing the unlimited variations on the theme of proteolytic activity. We will start by discussing the seemingly most obvious aspect of compartmentalization of proteolysis, that is, the action of proteases in different cellular compartments.
3.2 Sweet or Savoury—Salty or Tart: Biochemical Conditions of the Cleaving Environment
Evolution has given us proteases that are able to hydrolyze peptide bonds in a plethora of diverse biochemical conditions (Figs. 3.1 and 3.2). For example, calpain activity depends on free calcium concentrations ranging from 10−9 M in resting cells to 10−7 M in activated cells (Chap. 12; Vanlangenakker et al. 2008; Sorimachi et al. 2010), while other proteases like matrix metalloproteinases (MMPs) exhibit maximal activity in an extracellular environment characterized by 10−3 M free calcium. Furthermore, pH-values ranging from basic to highly acidic, and redox conditions from oxidizing (in the compartments of the secretory pathway and extracellular space) to reducing conditions (within endo-lysosomal compartments) are crucial determinants of protein folding and proteolytic activity (Chap. 2; Barrett and Kirschke 1981; Kirschke et al. 1995; McGrath 1999; Reinheckel et al. 2000; Pillay et al. 2002; Jordans et al. 2009; Zhou et al. 2010; Scott and Gruenberg 2011). Therefore, compartmentalization and complex formation not only generate efficient environments for proteolytic cleavage, but may modulate the local biochemical environment around an active protease.
One exquisite exemplification of this is in the proposed mechanism of Regulated Intramembrane Proteolysis (RIP) where proteases, like signal peptidases, sheddases, or the Alzheimer’s Precursor Protein (APP)-cleaving secretases, acquire the ability to cleave near or within amphipathic, α-helical transmembrane domains (Chaps. 9 and 10; Annaert and De Strooper 1999; De Strooper and Annaert 2000; Steiner and Haass 2000; Urban et al. 2001; Urban and Freeman 2002; Ehrmann and Clausen 2004; Parkin et al. 2004; Selkoe and Wolfe 2007; Freeman 2008; Murphy 2009; Lichtenthaler et al. 2011). The proteases involved in RIP, represented by soluble and peripheral membrane proteins, are proposed to form channel-like cleaving environments by arranging into multimeric complexes with other proteins. In contrast, turnover of transmembrane proteins in general, which includes extensive cleavage of hydrophobic transmembrane domains in addition to more straight-forward degradation of the hydrophilic luminal and cytosolic protein domains, is enabled by their extraction from within lipid-rich environments (membranes) through recruitment by the ESCRT (Endosomal Sorting Complex Required for Transport) machinery (Gu et al. 1997; Gruenberg 2001; Hicke and Dunn 2003; Gruenberg and Stenmark 2004; Piper and Katzmann 2007; van Meer and de Kroon 2011). A variety of hydrolytic enzymes present in the endocytic compartments, then degrade the transmembrane proteins in addition to their other substrates, such as membrane lipids (Chap. 4; Kirschke et al. 1995; McGrath 1999; Brix 2005).
Cleavage of peptide bonds in the extracellular milieu (Fig. 3.1) requires enzymes that thrive in carbohydrate- or glycosaminoglycan-rich environments; conditions of elevated negative charge and, hence, salt-‘enriching’ conditions (Jacques 1979; Bergers and Coussens 2000; Blobel 2000a; Rosenblum and Kozarich 2003; Stamenkovic 2003; Strongin 2010; Sato and Takino 2010; Zogg and Brandstetter 2011). Sequential cleavages mediated by cascades of proteases acting on one or several protein substrates (perhaps including the proteases themselves) in a tightly regulated manner seem to explain how compartmentalization is achieved even in the peri-cellular environment (Chaps. 13–15; Chapman et al. 1994; Owen and Campbell 1995; Holmbeck et al. 1999; Sternlicht and Werb 2001; Seiki 2002; Ellis 2003; Mott and Werb 2004; Sounni and Noel 2005; Munshi and Stack 2006), where lipid membranes are not the limiting structure of the reaction compartment, but rather serve as an organizing structural support. The solution of cells to such problems of focalized proteolytic events in the peri-cellular surrounding is often achieved by arranging for protease delivery to the cleavage environment in a sequential and spatially regulated manner (Basbaum and Werb 1996; Murphy and Gavrilovic 1999; Blobel 2000a; Hooper et al. 2001; Itoh 2006; Owen 2008; Vreemann et al. 2009; Friedl and Wolf 2009; Murphy and Nagase 2011; Pagano and Reboud-Ravaux 2011). Upon initiation via appropriate signals, fine-tuning the sequence of proteolysis occurs via interaction with partners comprising not only substrates but also other proteases, membrane receptors, and protease inhibitors (Chaps. 2 and 15; Brix et al. 2001; Schenk and Quaranta 2003; Dano et al. 2005; Jordans et al. 2009; Sun et al. 2009; Das et al. 2010) (Fig. 3.3).
Moreover, the chromatin environment of the nucleus also presents demanding conditions, characterized by extraordinarily long charged polymers such as DNA and RNA (negatively charged), and histones (positively charged). In addition, nucleic acids can provide scaffolding platforms for proteolysis (Ong et al. 2007) where the proteolytic enzymes feature positively-charged molecular surfaces to facilitate interaction with DNA as exemplified by separases to cleave the cohesin “glue” holding sister chromatids together (Heck 1997; Lamond and Earnshaw 1998; Nasmyth et al. 2000; Chapman 2004; Goulet et al. 2004; Goulet and Nepveu 2004; Ruchaud et al. 2007; Hudson et al. 2009; He et al. 2009; Sun et al. 2009; Yanagida 2009; Cauwe and Opdenakker 2010).
These examples clearly illustrate that proteases are dependent on the properties of the cleavage environment and that proteases may react or be modulated by rapid and transient alterations of the biochemical conditions during signalling events (Squier 2006; Salvesen and Dixit 1997). However, proteases themselves may be regulated by proteolytic cleavage and by other potentially reversible post-translational modifications including acetylation, methylation, glycosylation, hydrocarbonylation (farnesylation, geranylation, myristoylation), oxidation, phosphorylation, sulfation, SUMOylation, or ubiquitylation (Chap. 5; Doucet et al. 2008). These findings have profound implications on our understanding of the role of proteolysis as, above all, proteases are the mediators of possibly the most important post-translational modification of proteins—cleavage! This is because proteolytic action is essentially non-reversible and therefore frequently represents ‘points of no return’ as exemplified by caspase activation during apoptosis (Chap. 8).
Thus, eukaryotic cells have devised an almost unlimited repertoire of proteases; including soluble enzymes, transmembrane and peripheral-membrane proteins, macromolecular assemblies in the cytosol (proteasomes) or in the extracellular environment (meprins, MMPs, plasmin/uPA/uPAR). These proteases have the ability to interact with a likewise versatile group of substrates ranging from dipeptides to supramolecular protein assemblies that can be modified by endo- or exo-peptidic cleavages (Chaps. 1, 2 and 5). The diversity of proteolytic enzymes is represented by some hundreds of molecules classified on the basis of the catalytic mechanism of action as aspartic, cysteine, glutamic, metallo, serine, or threonine proteases (Chap. 1; Barrett et al. 2001, 2003; Barrett 2004; Rawlings et al. 2004; Rawlings 2010). In addition, a number of peptidases have not yet been classified into any of the known enzyme families.
3.3 The Architectural Art of Compartmentalization of Proteolysis: Specified Rooms for Proteolytic Processing
Like a room plan that is defined when constructing a new house, the eukaryotic cell hosts a variety of compartments with specific, well-defined conditions for specific [proteolytic] activities.
The nucleus, like an office or a creativity centre, can be considered a compartment of planning where proteolytic processing of proteins is a rare event needed for reading and distribution of the blueprint dictating when proliferating cells progress through the cell cycle, or pause in G0-phase in order to differentiate (Nasmyth et al. 2000; Pellman and Christman 2001; Goulet and Nepveu 2004). Therefore, histones, transcription factors, and structural proteins connecting sister chromatids are the main substrates known for nuclear proteases (Heck 1997; Goulet et al. 2004, 2007, 2008; Ong et al. 2007; Tedelind et al. 2010; Haugen et al. 2013). While more substrates will undoubtedly be identified in the future, it appears that protease activity in the nucleus of interphase cells is occasional and highly regulated. During mitosis, the nuclear envelope is disassembled (as a result of reversible phosphorylation cascades), and proteolysis follows the principles of the cytoplasm. Upon initiation of apoptosis by specific intrinsic or extrinsic factors, however, the nuclear envelope is also rearranged (with cleavage of nuclear lamins this time), eventually leading to programmed cell death (Wyllie et al. 1980).
The kitchen where meals are created can be compared with the centres of de novo protein biosynthesis such as the rough endoplasmic reticulum (rER), and the semi-autonomous organelles, mitochondria and plastids. Proteases of these compartments—perhaps with additional exceptions in the cytoplasm (see below)—are selective in their choice of substrates, i.e. they cut off short targeting sequences to ensure directionality of delivery of newly synthesized proteins (ER, mitochondria, plastids) and, together with chaperones, they sample and inspect protein domains for proper folding (cytosol, ER, mitochondria, plastids). The proteases of the compartments of de novo protein biosynthesis are involved in maturation of newly synthesized proteins, and therefore enable limited proteolysis that is domain-driven, sequence-targeted and restrictive—rarely involving “arbitrary” cleavage along the length of the polypeptide chain. Only in conditions of over-load or disease, resulting in massive misfolding of newly synthesized proteins, are the proteases of these compartments ‘up-regulated’ for enhanced protein processing. However, the cell will first take all measures possible to attempt enhanced folding or refolding assisted by chaperones that become up-regulated in the so-called Unfolded Protein Response (UPR). The misfolded proteins, however, like the kitchen garbage, are delivered to other compartments for storage and disposal in aggresomes and inclusion bodies (Kopito 2000; Singhvi and Garriga 2009). The improperly folded proteins will eventually be destined for degradation by proteasomes. This process, when involving the ER, is also known as ER-associated degradation (ERAD) during which newly synthesized but misfolded proteins of the ER lumen are retro-translocated for degradation in cytoplasmic and ER-associated proteasomes (Wiertz et al. 1996a, b; Meusser et al. 2005; Romisch 2005).
The post-ER compartments of the secretory pathway including the intermediate compartment, Golgi apparatus, trans-Golgi network (TGN) and secretory vesicles are crucially important for modification, further maturation, sorting and packaging of proteins (Farquhar and Palade 1981, 1998; Varki 1993; Rothman and Wieland 1996; Kaiser et al. 2002). Like in a living room, molecules involved in post-translational modifications of pro-proteins (such as glycosylation, phosphorylation, sulfation, or proteolytic processing) gather to perform the critical cellular actions in preparation for extracellular secretion of proteins that frequently takes place in response to communication of cells with their extracellular environment (Docherty and Steiner 1982; Zhou et al. 1999). Well-studied proteases known for their significance as proprotein-processing enzymes are furin and other proprotein-convertases (PCs) (Seidah et al. 1991; Garten et al. 1994; Schaller and Ryan 1996; Nakayama 1997; Steiner 1998; Seidah and Prat 2002; Thomas 2002; Rockwell and Thorner 2004; Henrich et al. 2005; Scamuffa et al. 2006; Creemers and Khatib 2008; Seidah 2011). The PCs interact in a transient and selective manner with a huge variety of soluble proteins thereby processing the pro-forms of secretory proteins that include ECM constituents as well as peptide or protein hormones (Steiner 1969; Steiner et al. 1980; Seidah and Chretien 1999; Fu et al. 2008).
The major sorting steps take place at the level of the TGN, a compartment that resembles endo-lysosomes with respect to acidity, and where cargo loading and packaging into secretory vesicles or secretory granules and other Golgi-derived vesicles occurs. Thus, proteins en-route to endo-lysosomes including the zymogen [inactive] forms of proteases such as the asparaginyl endopeptidase (AEP/legumain) or the cathepsins are also transported along the secretory route to the TGN. From here, they reach their next sub-cellular destinations in a non-processed and proteolytically inactive proform (Chap. 4; Mort and Buttle 1997; Schaschke et al. 1998; Khan and James 1998; Linke et al. 2002a, b; Mach 2002; Wiederanders et al. 2003; Burden et al. 2007; Buttle 2007; Brix et al. 2008). Some endo-lysosomal proteases, like cathepsins B and L, are believed to become sorted into and activated in secretory vesicles of pancreatic cells and in neurosecretory granules of pituitary cells (Tooze et al. 1991; Halangk et al. 2000; Hook et al. 2004, 2008; Meister et al. 2010; Funkelstein et al. 2010); these examples, however, are specific to certain cell types. Secretory granules and secretory vesicles of most eukaryotic cells may be considered as compartments where proteases that are almost ready to function (like chymase, tryptase, perforins) are stored together with a number of other secretory macromolecules in a highly concentrated and compacted fashion. Only very-limited proteolysis is taking place in these granules and vesicles, if at all (Gomez-Lazaro et al. 2010), and they may thus be considered analogous to hallways and corridors of houses that are used by both inhabitants and visitors simply for by-passing and through-traffic and possibly where coats are removed.
The dining room, designed for consumption of meals, serves as a reference point for the compartments of the endocytic pathway, i.e. endosomes and lysosomes (Gruenberg and Howell 1989; Schmid 1993; Gruenberg and Maxfield 1995; Mukherjee et al. 1997; Gu and Gruenberg 1999; Gagescu et al. 2000; Nichols and Lippincott-Schwartz 2001; Conner and Schmid 2003). A number of entry points exist for proteins reaching endo-lysosomes (Rechsteiner 1990; Seglen et al. 1996; Felberbaum-Corti et al. 2003; Lin et al. 2004; Klionsky 2007; Mizushima et al. 2008; Arias and Cuervo 2011; Chen and Klionsky 2011). Endo-lysosomes are multi-faceted and interchangeable sub-cellular compartments that perform numerous functions above their main catabolic tasks in the break-down of proteins and other internalized molecules (Brix 2005; Brix et al. 2008). Restricted and limited proteolysis in early endocytic compartments is extremely important in antigen processing for subsequent MHC class II dependent presentation (Riese and Chapman 2000). The early endosome is considered a ligand-unloading and a sorting station (not only for polarized epithelial cells), and it has receiving and distribution functions resembling those of the TGN in many respects (Howell et al. 1989; Sachse et al. 2002). Sorting may result in subsequent trafficking of proteins like receptors back to the plasma membrane for recycling, or further transport to the later endocytic compartments for transient storage or degradation (and thus, down-regulation) (Katzmann et al. 2002; Lin et al. 2004). The late endosome, however, is clearly the dining table where incoming proteins are taken apart and often broken down to the level of single amino acids. The menu (sequence) and the selection of substrates available on the dinner table will need one or many proteases to proteolytically process any given protein substrate (Tedelind et al. 2011). Thus, early, or possibly even all compartments along the endocytic pathway may be utilised—and, like the arrangement of our dinner tables, the number and types of cutlery will differ depending on the food served. The remnants of proteolytic processing will be stored in poorly accessible residual bodies often accumulating in lysosomes and in vacuoles [plant], which we may therefore compare with the trash bags of our house.
Peroxisomes of animal cells, glyoxysomes of plant cells, and the smooth ER of all eukaryotic cells are the major compartments for detoxification (Platta and Erdmann 2007) in which, in part, extremely oxidative environments enable beta-oxidation which facilitates protein degradation. The composition of peroxisomes and glyoxysomes does not appear to contain proteases. Yet, occasionally, there have been reports of proteolytic enzymes like insulin-degrading enzyme (IDE) that are involved in the degradation of oxidized proteins within peroxisomes (Authier et al. 1996; Morita et al. 2000). However, these compartments are certainly not considered protein-processing compartments of major impact on cellular functions, so we may compare them to the trash compactor or garbage disposal of a house that dispose of contaminating and potentially toxic components.
3.4 Proteases Facing the Extracellular Space
The outside deck/terrace of eukaryotic cells is represented by the plasma membrane with a variety of cellular appendages like cilia, flagella, or microvilli reaching into the extracellular surroundings (Kenny and Maroux 1982). Like the plasma membrane itself, these appendages can carry a multitude of proteolytic enzymes anchored as transmembrane proteins or, if soluble, by binding to cell surface receptors (Chaps. 7, 9 and 11; Bode et al. 1996; Rosenblum and Kozarich 2003; Mentlein 2004; List et al. 2006; Bugge et al. 2007; Turner and Nalivaeva 2007; Lopez-Otin and Bond 2008; Szabo and Bugge 2008; Sohail et al. 2008; Sterchi et al. 2008; Choi et al. 2009; Ramsay et al. 2009; Yao 2010; Hendrickx et al. 2011). Such co-ordination of proteolytic activity in the peri-cellular environment is well known for important contributions in protein breakdown and peptide processing, in particular, at the apical plasma membrane domains of enterocytes (Chap. 11; Kenny and Maroux 1982; Bank et al. 2008; Matteucci and Giampietro 2009; Yu et al. 2010; Arampatzidou et al. 2011a, 2012). A classic example of this is in the most vital event of fertilization, where proteases are secreted from within the sperm head acrosome for degradation and modification of the protective vitellin layer surrounding the oocyte (Blobel 2000b). The oocyte is eventually reached by the sperm for subsequent fusion and zygote formation through explosive, actin polymerization-driven extrusion of microvillus-like extensions.
In addition to outward-looking appendages of cells, invaginations of the plasma membrane also play a role in the dynamics of proteolysis. Some invaginations form during internalization of molecules destined to reach the compartments of the endocytic pathway. Others, like caveolae (little flask-like caves), seem to be less well connected to the cellular interior, but provide a specific biochemical environment in which protein processing is facilitated (Simionescu et al. 1972; Hajjar and Acharya 2000; Predescu et al. 2001; Kim and Hajjar 2002; Pelkmans and Helenius 2002; Navarro et al. 2004). Thus, plasma membrane indentations—caveolae and other non-caveolin coated microdomains—can provide areas of concentrated cell surface receptors for restricting and focalizing proteases to specified sub-domains of the cell surface (Owen and Campbell 1995; Mai et al. 2000; Cavallo-Medved and Sloane 2003; Gumy et al. 2010).
Invadopodia may be analogous to caveolae in their protease concentrating roles but are constructed so as to extend finger-like into the extracellular matrix (Chen 1996; Murphy and Gavrilovic 1999; Linder 2007; O’Brien and O’Connor 2008; Frittoli et al. 2011). Invadopodia have become prominently known for their distinctive composition and functional duality for being involved in both cell adhesion and extracellular matrix degradation through associated proteases. Hence, delicate and rapidly interchangeable cellular extensions that protrude and communicate with the surrounding environment like an easily remodelled patio may also contribute in multiple ways to peri- and extra-cellular proteolytic processes (Woodward et al. 2007; Brix et al. 2008; Korkmaz et al. 2008; Pham 2008; Stetler-Stevenson 2008; Sato and Takino 2010). The proteases found in such exposed positions are usually characterized by extended extracellular domains which are often heavily glycosylated to promote stability against proteolytic attacks, and like their soluble relatives, provide many platforms and binding sites for interaction with other macromolecules or regulatory factors (Gahmberg and Tolvanen 1996; Manon-Jensen et al. 2010; Cawston and Young 2010).
The extracellular space itself, the garden, also houses a myriad of proteases that are derived from the many secretions of different cells in a tissue (Chaps. 6, 13–15; Andrews 2000; Overall and Blobel 2007). Proteolytic enzymes in the extracellular space may either be secreted in active form or remain latent until activation triggers other proteolytic enzymes to convert them into active proteases that often interact with each other to cleave protein substrates by sequential modes (Brix and Herzog 1994; Brix et al. 1996, 2001; Tepel et al. 2000; Linke et al. 2002a; Mayer et al. 2009; Vreemann et al. 2009; Tedelind et al. 2011). The extracellular space can therefore be considered a rich reservoir of proteolytic enzymes that are not only crucial for tissue remodelling but, in part, also contribute to organization of extracellular matrix components, a function that is crucial during morphogenesis in embryonic development (using ADAMs [A Disintegrin And Metalloprotease], ADAM-TSs) (Chap. 9; White 2003; Noel et al. 2004; Apte 2004; Edwards et al. 2008; Kveiborg et al. 2008; Reiss and Saftig 2009; van Goor et al. 2009; Dikic and Schmidt 2010; Urban 2010; Saftig and Reiss 2011).
3.5 Democracy as an Answer for Radical Decision-Making Processes
Radical decision-making during peptide bond cleavage may be seen as the hallmark of proteolytic activitiy, yet proteases can hardly be thought of as dictators. Often proteases act in proteolytic networks in which it is not the individual enzyme that counts (Chaps. 2 and 5). Collaboration, including finely-tuned and highly-regulated actions amongst several proteases, is instrumental in paving the way to success. Clearly, cleaving substrates at the right time and place, and with the desired pace and the required specificity is crucial (Liu et al. 1999; Kidd et al. 2001; Rao 2003; Baruch et al. 2004; Joyce and Hanahan 2004; Blum et al. 2005; Brix and Jordans 2005; Carlson and Cravatt 2007; Gocheva and Joyce 2007; Victor and Sloane 2007; Blum 2008; Brix et al. 2008; Jedeszko et al. 2008; Paulick and Bogyo 2008). We shall discuss these aspects of sub-compartmentalization by looking in more detail at the specialists. These include the endo-lysosomal proteases, which come in a most astounding array and act, not exclusively but optimally in endosomes and lysosomes of mammalian cells (Chap. 4; Kirschke et al. 1995; Chapman et al. 1997; McGrath 1999; Turk et al. 2001; Brix 2005; Mohamed and Sloane 2006; Brix et al. 2008; Reiser et al. 2010) (Fig. 3.5).
The compartments of the endocytic pathway contains a wide range of protein processing and degrading enzymes including the cysteine peptidase AEP/legumain that engages in proteolytic maturation of the proforms of other endo-lysosomal proteases (Barrett and Rawlings 2001; Ishidoh and Kominami 2002; Watts et al. 2005; Haugen et al. 2013), aspartic cathepsins D and E (Barrett 1979; Yamamoto 1995; Dunn et al. 1998; Tatnell et al. 1998; Rochefort et al. 2000; Tsukuba et al. 2000; Dash et al. 2003; Nakanishi 2003; Liaudet-Coopman et al. 2006; Zaidi et al. 2008; Hook et al. 2008; Nicotra et al. 2010), the serine cathepsins A and G (Travis 1988; Hiraiwa 1999; Pham 2006; Caughey 2007; Korkmaz et al. 2008; Meyer-Hoffert 2009; Kessenbrock et al. 2011), and the most versatile group of the cysteine cathepsins B, C, F, H, K, L, O, S, V, W, and X/Z in man (Chap. 6; Kirschke et al. 1995; McGrath 1999; Turk et al. 2001; Greenbaum et al. 2002; Brix 2005; Sloane et al. 2005; Mohamed and Sloane 2006; Gocheva and Joyce 2007; Brix et al. 2008; Reiser et al. 2010). Rodents express even more (placental) cysteine cathepsins (Sol-Church et al. 2002) and protozoa like Tetrahymena are clearly very extreme examples with dozens of these proteolytic enzymes. The cathepsins that can be found in endo-lysosomes of every eukaryotic cell are the cathepsins B, D, G, H, and L (Turk et al. 2000; Reinheckel et al. 2001; Vasiljeva et al. 2007; Brix et al. 2008; Reiser et al. 2010), while other cathepsins may be present only in certain cell types that are facing very specific challenges, e.g. osteoclasts express high levels of cathepsin K to facilitate bone matrix turnover (Bromme and Okamoto 1995; Saftig et al. 1998; Tepel et al. 2000; Lecaille et al. 2003; Desmarais et al. 2009; Podgorski 2009; Rachner et al. 2011). Nonetheless, all of these proteolytic enzymes can process their substrates by limited or by full proteolysis (Brix and Herzog 1994; Brix et al. 1996, 2001; Friedrichs et al. 2003; Dauth et al. 2011a, b).
Substrates reach the endocytic compartments not only by internalization from the extracellular space, but also by direct entry from the cytosol via chaperone-mediated or “classic/conventional” autophagy (Dice 2007). The outcome of substrate proteolysis by endo-lysosomal proteases depends on whether individual or multiple proteases act on the substrate with endo- and/or exo-peptidase modes of cleavage (Chaps. 1, 2, 6, 15; Barrett and Kirschke 1981; Kirschke et al. 1995; McGrath 1999; Tepel et al. 2000; Jordans et al. 2009). Moreover, proteolytic processing in endocytic compartments is aided by increasingly reductive conditions that support protein unfolding (Pillay et al. 2002; Jordans et al. 2009; Scott and Gruenberg 2011). In general, the hydrolytic enzymes of endo-lysosomes will team up so that almost all protein substrates with their many other, non-proteolytic post-translational modifications can be handled very efficiently. Interestingly enough, endo-lysosomal proteolytic enzymes are sometimes selective in their substrate choice, whereas others are less selective and cleave almost any protein substrate. The pH optimum of substrate cleavage by endo-lysosomal proteolytic enzymes spans an astonishingly wide range from neutral pH-values for cathepsin S down to the most acidic pH-values for cathepsin D (Takahashi and Tang 1981; Kirschke and Wiederanders 1994). Hence, protein processing and degradation by endo-lysosomal proteases is already initiated in the peri-cellular vicinity of specialized cells that secrete these enzymes in a regulated or non-regulated manner, and, in general, the activity of endo-lysosomal proteases is available in all compartments of the endocytic pathway (Brix and Herzog 1994; Brix et al. 2008). An example of this kind can be found in the thyroid gland, where a huge protein substrate (thyroglobulin) and its covalently cross-linked supramolecular assemblies are handled by the cathepsins B, K, L, and S acting in a temporal- and spatially-regulated, sequential manner for proteolytic liberation of iodinated thyronines from within the polypeptide chain (Brix et al. 1996, 2001; Friedrichs et al. 2003; Jordans et al. 2009; Dauth et al. 2011a).
The cocktail of proteases that is present in one or another sub-compartment of the endocytic pathway will determine the extent of protein substrate cleavages. Numerous questions therefore arise: what is an endosome and what is a lysosome? What governs or triggers transport of endo-lysosomal proteases into the interchangeable compartments of the endocytic pathway? And what are the targeting sequences and which mechanisms explain why some compartments retain fewer, while others contain more, of these proteolytic enzymes? The answers to these questions are partially derived from the study of mannose 6-phosphorylation of the pro-forms of endo-lysosomal proteases (Kornfeld 1992; Peters and von Figura 1994; Bresciani and von Figura 1996). This post-translational modification is maintained in the compartments of the secretory pathway, but partially also persists in the extracellular space. The well known cation-dependent and -independent mannose 6-phosphate receptors can deal with sorting at the level of the TGN, or with re-internalization should pro-forms become secreted (Peters et al. 1990; Hille et al. 1992; Koster et al. 1993; McIntyre et al. 1994; Pohlmann et al. 1995). However, this elegant molecular mechanism of endo-lysosomal enzyme trafficking is not realized in all cell types (Ludwig et al. 1994; Pohlmann et al. 1995; Tanaka et al. 2000). Therefore, the answers to these questions will not be universal, but vary from cell type to cell type (Linke et al. 2002a, b; Brix 2005; Brix and Jordans 2005; Brix et al. 2008; Tedelind et al. 2011).
3.6 Debating Clubs: Proteases and Their Inhibitors
Proteolysis is a result of balancing proteolytic and anti-proteolytic factors (Chaps. 6 and 15; Turk and Bode 1991; Basbaum and Werb 1996; Turk et al. 1997; Matrisian 1999; Coussens et al. 2002; Abrahamson et al. 2003; Turk et al. 2003; Kaiserman et al. 2006; Mohamed and Sloane 2006; Lopez-Otin and Matrisian 2007; Vasiljeva et al. 2007; Scott and Taggart 2010; Reiser et al. 2010). While this notion may have been formulated early on in the examination of proteolysis, it still remains an apt proposal. However, the temporal and spatial regulation of proteolysis must be considered as equally important to the balance with anti-proteolytic factors, if not more. Hence, a well orchestrated debating event between substrates, proteases, and their inhibitors is required for a given cellular process to dictate the final outcome of proteolytic cleavage. So far, many more investigations exist on the proposal of this house: “how and when and where to start proteolysis”, rather than on: “how and when and where to halt the event”. However, signalling may intervene (Mackie et al. 2002; Ichihara et al. 2006; Mockaitis and Estelle 2008; Murphy et al. 2009; Smith and Marshall 2010) and continued proteolysis may be required in some instances. Control is essential as if proteolysis is exceeded, it may lead to severe disease and eventual death. Therefore, inhibitors of proteases, that typically act intracellularly, are highly concentrated in extracellular fluids for prevention of premature proteolysis around a cell or tissue should the proteases happen to escape from cells in unplanned events (Travis and Salvesen 1983; Turk and Bode 1991; Nagase et al. 1996; Bode et al. 1999; Deveraux et al. 1999; Silverman et al. 2001; Salvesen and Duckett 2002; Abrahamson et al. 2003; Whisstock and Bottomley 2008; Drag and Salvesen 2010). Thus, termination of proteolytic cleavage is as important as its initiation and the Master of Ceremony that determines the sequence and types of proteolytic events taking place must consider protease inhibitors as essential factors in the compartmentalization of proteolysis.
Again, endo-lysosomal enzymes represent good examples for illustrating such control mechanisms by their endogenous inhibitors. Endo-lysosomal enzymes can become extremely dangerous for other cellular compartments or even for the entire cell if released into the extracellular space in an uncontrolled fashion. Therefore, endo-lysosomal proteolytic enzymes must be transported in a defined manner and premature activation is further shielded by potent 60–100 residue inhibitory N-terminal pro-peptide domains as in the case of cysteine cathepsin proteases (Chap. 4; Mach et al. 1994; Mach 2002; Wiederanders et al. 2003). These domains have two functions; firstly, they adopt their own clearly defined secondary structure which acts as a folding chaperone to facilitate the larger catalytic domain to fold. Secondly, the pro-peptide binds tightly across the active site of the catalytic domain in reverse orientation to normal substrates, thus acting as competitive inhibitors. This tight binding is pH dependent and as the inactive zymogen species is transported to the lysosomes, the propeptide will dissociate from the catalytic domain in the acidified lysosomal lumen, leaving the active site exposed and active and results in the irreversible proteolytic removal of the propeptide domain leaving the mature active protease domain (Pungercar et al. 2009; Linke et al. 2002a, b). However, despite this understanding of the role of the pro-peptide, the recent studies highlighting the presence of cysteine cathepsins in the nucleus, a consequence of downstream translational initiation (Chapman 2004; Goulet and Nepveu 2004; Goulet et al. 2004; Tedelind et al. 2010), highlights that these control mechanisms for the pro-peptide require further supplementation by the presence of inhibitors such as Stefin B in the nucleus (Ong et al. 2007; Ceru et al. 2010).
Cystatins and stefins are the most powerful counter-players of some of the endo-lysosomal proteases (Turk and Bode 1991; Turk et al. 1997, 2003; Abrahamson et al. 2003). Some families of these protease inhibitors reside within the cytosol whereas others are secreted into the extracellular space for abundant presence in extracellular fluids of tissues and body organs. Hence, the cysteine peptidase inhibitors are separated as safe-guarding molecules from the endo-lysosomal cysteine proteases by a membrane, the endo-lysosomal membrane and/or the plasma membrane. Very similar strategies are realized for serpins that inhibit both serine- and cysteine peptidases and which are abundantly present in the cyto- and nucleoplasm of eukaryotic cells (Silverman et al. 2001; Whisstock and Bottomley 2006, 2008). The aspartic peptidases of endo-lysosomes are kept in control by another group of cross-class inhibitors, the thyropins, which also interfere with cysteine peptidase activities (Chap. 2; Lenarcic and Bevec 1998; Novinec et al. 2006). Another regulatory mechanism of keeping the team of endo-lysosomal proteases well within their borders is reflected by the relatively strict pH-requirements for both proteolytic activity and stability of these enzymes, thereby explaining why many members of this large group of proteases can be kept in check, simply by compartmentalization to specific cellular regions of defined biochemical conditions (Mort et al. 1984; Mort and Buttle 1997; Jordans et al. 2009).
MMPs, meprins/BMPs, ADAMs and ADAM-TSs are best known for their proteolytic activities exerted in the extra- and peri-cellular space (Chaps. 7, 9, 11, 13–15; Brinckerhoff and Matrisian 2002; Coussens et al. 2002; Norman et al. 2003; Edwards et al. 2008; Reiss and Saftig 2009; Rosenberg 2009; Tallant et al. 2010). These enzymes are either soluble or transmembrane proteins. However, many act in sequence thereby establishing proteolytic cascades in which one metalloprotease activates the next (Chakraborti et al. 2003). These cascades of proteolytic activation often take place at the cell surface with the plasma membrane and its receptors serving as scaffolding support. Another excellent example of a similar kind is provided by the urokinase-type plasminogen activator (uPA) that interacts with the plasminogen activator receptor (uPAR), a transmembrane protein of the plasma membrane (Magdolen et al. 2000; Blasi and Carmeliet 2002; Behrendt 2004; Mondino and Blasi 2004; Shi and Stack 2007). Moreover, in a number of proteolytic processes that involve MMPs and PAs (Plasminogen Activators), the endogenous protease inhibitors TIMPs (tissue inhibitors of MMPs) and PAIs (PA inhibitors) become part of the proteolytic cascade and therefore fine-tune and regulate the peri-cellular activities of these proteolytic enzymes (Bode and Renatus 1997; Bode and Maskos 2001, 2003; Stetler-Stevenson 2008; Brew and Nagase 2010). Final termination of such proteolytic events that can involve multiple members of the same and also of related and unrelated protease families is considered to occur by down-regulation (Jiang et al. 2001), i.e. degradation by endo-lysosomal pathways.
3.7 Speed-Dating of Proteolytic Enzymes and Their Substrates
So far we have not described one of the most intriguing examples of sub-compartmentalization of proteolysis and of processivity of proteolytic cleavage—the processing of linear polypeptides by the very well organized protease-machines of the cytosol, the proteasomes (Groll and Clausen 2003).
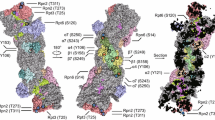
The cytoplasm, as the largest sub-compartment of eukaryotic cells, bears a variety of complex protein-processing assemblies (De Mot et al. 1999; Brandstetter et al. 2001; Rosenblum and Kozarich 2003). The best understood of such cytoplasmic protease assemblies is the 26S proteasome that can function in cleaving a variety of different peptide-bonds, and which can even be considered a protein processing machine when acting in conjunction with 19S cap-structures that comprise energy-dependent unfolding factors (Wolf and Hilt 2004; Vierstra 2009; Stadtmueller and Hill 2011). The structure of the yeast proteasome has been determined in atomic detail to a resolution of 2.4 Å by X-ray crystallography which also discovered that the proteasome belongs to a new class of proteases, the threonine peptidases (Groll et al. 1997). These astonishingly versatile, and, in cellular immune response reactions, highly adaptable (Groettrup et al. 1996, 2001a, b, 2010; van den Eynde and Morel 2001; Kloetzel and Ossendorp 2004; Rivett and Hearn 2004; Driscoll and Dechowdhury 2010) protease complexes provide a hydrolytic environment with unique properties in that four heptameric rings assemble to form a cleavage chamber (Groll and Clausen 2003; Stadtmueller and Hill 2011) that concentrates, guides and shields protease substrates from surrounding influences without being enclosed by a lipid membrane. The fully functional proteasome is already the 20S assembly which is complemented by two 19S regulatory caps, one at each end to prevent inappropriate degradation in the larger 26S proteasome.
The precise localization of proteasomes is still a matter of debate (Rivett 1998; Brooks et al. 2000a, b; Wojcik and DeMartino 2003). While their cytoplasmic location is undisputed, it is less clear under which circumstances—and for which reasons, proteasomes may be detected in the nucleus. Proteasomes are often associated with the cytosolic leaflets of the ER membrane or the nuclear envelope where they engage in ERAD, the ER-associated degradation of misfolded protein intermediates that are destined for destruction in order to protect the cell from an overload and accumulation of unfolded proteins (Wiertz et al. 1996a, b). Likewise, proteasomes are detectable as constituents of the so-called aggresomes, non-membrane enclosed regions of the cytoplasm of cells that harbour inclusions of misfolded proteins resulting from protein over-expression (Kopito 2000). In all these latter cases, protein substrates may or may not require ubiquitinylation as a proteasome-targeting signal (Pickart 1997; Schwartz and Hochstrasser 2003; Ciechanover 2003; Tai and Schuman 2008).
The proteasome is thus a beautiful example where evolution has devised a complex protein assembly employing a variety of proteolytic activities to cleave substrates in a more or less specific fashion. The result may be release of peptides of defined length, suitable to enter the ER lumen via TAP-transporter for MHC class I dependent antigen presentation of virus-derived peptides (Abele and Tampe 2006, 2009; Hansen and Bouvier 2009), or tailor-made for further destruction by other proteolytic enzymes of the cytosol that feature amino- or carboxypeptidase activities (Gomis-Ruth 2008; Gomis-Ruth et al. 2012) in order to replenish the cellular pool of free amino acids.
3.8 How to Organise Proteolytic Actions in Busy Times Like Rapid Cell Cycle Progression, Remodelling of Cellular Components, or Cell Death
The cytoplasm is already considered as a compartment of de novo protein biosynthesis. However, it is also the cellular compartment of protein maintenance and turnover in response to signalling or mechanical damage of cytoskeletal proteins (Chap. 12; Goldberg and Dice 1974; Dice and Walker 1979; Dice 1987, 1990; Olson and Dice 1989; Rechsteiner and Hill 2005). It is also well understood that cell cycle progression depends on cyclic degradation and reformation of cyclins. Moreover, the cytoplasm contains many scaffolding factors, proteins and ribonucleoproteins, that have the ability to build and organize larger assemblies of macromolecules, such as the apoptosome that initiates a sequence of dramatic proteolytic events eventually leading to cell death in a tightly regulated and programmed manner (Chap. 8; Tschopp et al. 2003; Vanlangenakker et al. 2008; Declercq et al. 2009; Pop and Salvesen 2009; Vandenabeele et al. 2010; Krysko et al. 2011). Hence, while the cytoplasm may be the largest compartment of eukaryotic cells, it has established unique strategies to sub-compartmentalize proteolytic actions to certain required areas. As calcium waves emerging from the sperm entry point specify polarity of the zygote, and thus the body axis at the earliest stage of development, the cytoplasm of eukaryotic cells has found intriguing and elegant solutions to the primary question of this chapter, compartmentalization of proteolysis.
Besides proteasomes and a number of other proteolytic enzymes that are important contributors to guarantee the pool of free amino acids, other proteases are found abundantly in the cytosol: the procaspases (Earnshaw et al. 1999; Riedl and Shi 2004; He et al. 2009; Pop and Salvesen 2009; Drag and Salvesen 2010) and the calpains (Sorimachi et al. 1997, 2010; Reverter et al. 2001; Storr et al. 2011). Procaspases are cysteine peptidases cleaving after aspartic acid upon their proteolytic activation in the initiation of apoptotic pathways leading to programmed cell death (Alnemri et al. 1996). The assembly of caspases in apoptosomes is triggered by a variety of intrinsic and extrinsic signalling pathways (Wyllie et al. 1980; Budihardjo et al. 1999; Song and Steller 1999; Salvesen 2002; Riedl and Salvesen 2007; Kersse et al. 2011). Again, a sub-compartmentalization of their proteolytic activities can be conceived from the formation of restricted, caspase-enriched cytoplasmic areas where scaffolding factors serve as the platforms for apoptosome formation, and, as in the case of proteasomes, without the help of lipid membranes. Calpains (also cysteine peptidases) represent yet another solution to restricting proteolytic activities, as they are dependent on cytosolic free calcium in the range of nanomolar concentrations that exist only transiently in response to activation of cells via e.g. protein kinase C-mediated signalling (Sato and Kawashima 2001). Calpains are known to be involved in cytoskeletal protein remodelling in skeletal muscle cells under conditions of excessive contractile activity that may cause micro-damage to the muscle fibers (Chap. 12; Koohmaraie 1992).
Besides the above described mechanisms of restricting proteolytic activity of cytoplasmic proteases to certain areas of the cytoplasm, a number of ‘safe-guarding’ protease inhibitors can be found in certain areas of the cytoplasm as well (Turk et al. 2003; Kaiserman et al. 2006) (see also above). First, and foremost, the serpins which act as cross-class inhibitors of both serine and cysteine peptidases are found (Blasi 1993; Smirnova et al. 1994; Brunner and Preissner 1994; Pappot et al. 1995; Bailey et al. 2006; Kaiserman et al. 2006; Izuhara et al. 2008). Secondly, the cystatins are localised not only in all body fluids and in endocytic compartments, but other members of this inhibitor family are also present in the cytoplasm (Travis and Salvesen 1983; Turk and Bode 1991; Bode et al. 1999; Deveraux et al. 1999; Silverman et al. 2001; Abrahamson et al. 2003; Whisstock and Bottomley 2008). Cystatins are believed to protect cells from sudden death by inhibiting cysteine peptidases which not only reside and act normally in the cytoplasm, but which may become enriched in the cytosol if endo-lysosomes suffer membrane leakage or rupture (Chap. 8; Nixon and Cataldo 1993; McNeil and Steinhardt 1997; Gerasimenko et al. 2001; Degli Esposti 2008; Ivanova et al. 2008; Turk and Turk 2009). Thus, eukaryotic cells can further transiently modulate proteolytic activity through the expression and localisation of cognate inhibitors.
3.9 Themes Emerging from the Discovery of the New Kids on the Block
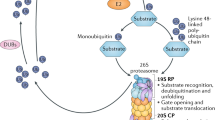
The role for selected proteolysis in ubiquitin biology is an area that is fast developing. Ubiquitin (and indeed related peptides such as SUMO, ISG15) is post-translationally added to proteins by E3 ubiquitin ligase complexes (Pickart and Eddins 2004). This modification of a specific protein can have a broad range of effects, from simple targeting of a protein for degradation, to modification of its activity in a manner not dissimilar from the role of phosphorylation throughout biological networks. Ubiquitination forms a key signaling role in processes such as cell cycle regulation (Song and Rape 2008), enzyme activation/inactivation (Adhikari et al. 2007) and DNA repair (Kennedy and D’Andrea 2005).
Ubiquitin is a 76 residue peptide, the free C-terminus of which is conjugated onto lysine side chain amino groups on target proteins forming a pseudo-peptide bond (Pickart 2001). However, the form of this ubiquitination can vary substantially from a simple mono-ubiquitination to addition of a number of ubiquitins (poly-ubiquitination). This post-translational modification is further complicated in that ubiquitin itself has a number of lysine residues and a branched poly-ubiquitin can be created through these points—each of which has its own distinctive conformation and role (K48 branched polymers labels proteins for degeneration and K63 polymers activate target proteins) (Pickart and Fushman 2004).
Key to the role of ubiquitination signaling is the fact that it is reversible and removal of ubiquitin is coordinated by proteases called deubiquitinating enzymes (DUBs). To date, almost 100 DUB proteases have been identified, belonging to cysteine and metalloprotease families (Reyes-Turcu et al. 2009). However, it is interesting to note that no endogenous inhibitors of these proteases have been identified to date and control of their activity appears to use other mechanisms including transcriptional control, substrate recognition and physical compartmentalization (Zhao et al. 2008).
In addition to these mechanisms of control, within the cytoplasm where the majority of these proteases are located, a major source of control is the presence of binding partners and adaptor proteins. Indeed, many of the DUB proteases contain other domains in addition to their catalytic core to facilitate binding to other proteins, and in particular scaffold proteins where it is thought that the DUB forms part of a multi-protein complex (often with the cognate E3 ubiquitin ligase complex), coordinating its activity and substrate specificity (Marfany and Denuc 2008). In this regard, it is perhaps particularly striking that invadolysin, a metalloprotease coordinating mitotic progression, has recently been found to interact genetically with non-stop, a DUB (MM Heck, personal communication).
3.10 And Now, We Retire to the Study
We conclude that we have only glimpsed the tip of the ice-berg. Many more intriguing features of how proteolysis is compartmentalized in the realm of the cell are to be discovered, understood and set into context before we will fully grasp the complexity of proteolytic enzymes and their compartmentalization principles. Faulty localization, delayed or too early, too fast or too slow, extensive or non-productive interactions of proteases, substrates, inhibitors and other factors regulating proteolysis will result in dramatic effects to the cell—and likely in diseases detrimental to the functioning of an organism. Understanding the regulation of intra- and extra-cellular proteolysis will be an important step to assess the physiology of eu- and prokaryotic cells. In turn, this understanding will be crucial to clarifying the routes leading to disease, and ultimately, shed light on potential therapeutic pathways.
References
Abele R, Tampe R (2006) Modulation of the antigen transport machinery TAP by friends and enemies. FEBS Lett 580(4):1156–1163
Abele R, Tampe R (2009) Peptide trafficking and translocation across membranes in cellular signaling and self-defense strategies. Curr Opin Cell Biol 21(4):508–515
Abrahamson M, Alvarez-Fernandez M, Nathanson CM (2003) Cystatins. Biochem Soc Symp 70:179–199
Adhikari A, Xu M, Chen ZJ (2007) Ubiquitin-mediated activation of TAK1 and IKK. Oncogene 26(22):3214–3226
Alnemri ES, Livingston DJ, Nicholson DW, Salvesen G, Thornberry NA, Wong WW, Yuan J (1996) Human ICE/CED-3 protease nomenclature. Cell 87(2):171
Andrews NW (2000) Regulated secretion of conventional lysosomes. Trends Cell Biol 10(8):316–321
Annaert W, De Strooper B (1999) Presenilins: molecular switches between proteolysis and signal transduction. Trends Neurosci 22(10):439–443
Apte SS (2004) A disintegrin-like and metalloprotease (reprolysin type) with thrombospondin type 1 motifs: the ADAMTS family. Int J Biochem Cell Biol 36(6):981–985
Arampatzidou M, Mayer K, Iolyeva ME, Gebre Asrat S, Ravichandran M, Günther R, Schüle T, Reinheckel T, Brix K (2011a) Studies of intestinal morphology and cathepsin B expressing in a transgenic mouse aiming at intestine-specific expression of CathB-EGFP. Biol Chem 392:983–993
Arampatzidou M, Rehders M, Dauth S, Yu DMT, Tedelind S, Brix K (2011b) Imaging of protease functions – current guide to spotting cysteine cathepsins in classical and novel scenes of action in mammalian epithelial cells and tissues. Ital J Anat Embryol 116(1):1–19
Arampatzidou M, Schütte A, Hansson GC, Saftig P, Brix K (2012) Effects of cathepsin K deficiency on intercellular junction proteins, luminal mucus layers, and extracellular matrix constituents in the mouse colon. Biol Chem 393:1391–1403
Arias E, Cuervo AM (2011) Chaperone-mediated autophagy in protein quality control. Curr Opin Cell Biol 23(2):184–189
Arolas JL, Vendrell J, Aviles FX, Fricker LD (2007) Metallocarboxypeptidases: emerging drug targets in biomedicine. Curr Pharm Des 13(4):349–366
Authier F, Posner BI, Bergeron JJ (1996) Insulin-degrading enzyme. Clin Invest Med 19(3):149–160
Bailey CM, Khalkhali-Ellis Z, Seftor EA, Hendrix MJ (2006) Biological functions of maspin. J Cell Physiol 209(3):617–624
Bank U, Bohr UR, Reinhold D, Lendeckel U, Ansorge S, Malfertheiner P, Tager M (2008) Inflammatory bowel diseases: multiple benefits from therapy with dipeptidyl- and alanyl-aminopeptidase inhibitors. Front Biosci 13:3699–3713
Barrett AJ (1979) Cathepsin D: the lysosomal aspartic proteinase. Ciba Found Symp 75:37–50
Barrett AJ (2004) Bioinformatics of proteases in the MEROPS database. Curr Opin Drug Discov Devel 7(3):334–341
Barrett AJ, Kirschke H (1981) Cathepsin B, Cathepsin H, and cathepsin L. Methods Enzymol 80(Pt C):535–561
Barrett AJ, Rawlings ND (2001) Evolutionary lines of cysteine peptidases. Biol Chem 382(5):727–733
Barrett AJ, Rawlings ND (2007) ‘Species’ of peptidases. Biol Chem 388(11):1151–1157
Barrett AJ, Rawlings ND, O’Brien EA (2001) The MEROPS database as a protease information system. J Struct Biol 134(2–3):95–102
Barrett AJ, Tolle DP, Rawlings ND (2003) Managing peptidases in the genomic era. Biol Chem 384(6):873–882
Baruch A, Jeffery DA, Bogyo M (2004) Enzyme activity–it’s all about image. Trends Cell Biol 14(1):29–35
Basbaum CB, Werb Z (1996) Focalized proteolysis: spatial and temporal regulation of extracellular matrix degradation at the cell surface. Curr Opin Cell Biol 8(5):731–738
Behrendt N (2004) The urokinase receptor (uPAR) and the uPAR-associated protein (uPARAP/Endo180): membrane proteins engaged in matrix turnover during tissue remodeling. Biol Chem 385(2):103–136
Bergers G, Coussens LM (2000) Extrinsic regulators of epithelial tumor progression: metalloproteinases. Curr Opin Genet Dev 10(1):120–127
Bhat KP, Greer SF (2011) Proteolytic and non-proteolytic roles of ubiquitin and the ubiquitin proteasome system in transcriptional regulation. Biochim Biophys Acta 1809(2):150–155
Blasi F (1993) Urokinase and urokinase receptor: a paracrine/autocrine system regulating cell migration and invasiveness. Bioessays 15(2):105–111
Blasi F, Carmeliet P (2002) uPAR: a versatile signalling orchestrator. Nat Rev Mol Cell Biol 3(12):932–943
Blobel CP (2000a) Remarkable roles of proteolysis on and beyond the cell surface. Curr Opin Cell Biol 12(5):606–612
Blobel CP (2000b) Functional processing of fertilin: evidence for a critical role of proteolysis in sperm maturation and activation. Rev Reprod 5(2):75–83
Blum G (2008) Use of fluorescent imaging to investigate pathological protease activity. Curr Opin Drug Discov Devel 11(5):708–716
Blum G, Mullins SR, Keren K, Fonovic M, Jedeszko C, Rice MJ, Sloane BF, Bogyo M (2005) Dynamic imaging of protease activity with fluorescently quenched activity-based probes. Nat Chem Biol 1(4):203–209
Bode W, Huber R (1992) Natural protein proteinase inhibitors and their interaction with proteinases. Eur J Biochem 204(2):433–451
Bode W, Huber R (2000) Structural basis of the endoproteinase-protein inhibitor interaction. Biochim Biophys Acta 1477(1–2):241–252
Bode W, Maskos K (2001) Structural studies on MMPs and TIMPs. Methods Mol Biol 151:45–77
Bode W, Maskos K (2003) Structural basis of the matrix metalloproteinases and their physiological inhibitors, the tissue inhibitors of metalloproteinases. Biol Chem 384(6):863–872
Bode W, Renatus M (1997) Tissue-type plasminogen activator: variants and crystal/solution structures demarcate structural determinants of function. Curr Opin Struct Biol 7(6):865–872
Bode W, Grams F, Reinemer P, Gomis-Ruth FX, Baumann U, McKay DB, Stocker W (1996) The metzincin-superfamily of zinc-peptidases. Adv Exp Med Biol 389:1–11
Bode W, Brandstetter H, Mather T, Stubbs MT (1997) Comparative analysis of haemostatic proteinases: structural aspects of thrombin, factor Xa, factor IXa and protein C. Thromb Haemost 78(1):501–511
Bode W, Fernandez-Catalan C, Nagase H, Maskos K (1999) Endoproteinase-protein inhibitor interactions. APMIS 107(1):3–10
Bonifacino JS, Weissman AM (1998) Ubiquitin and the control of protein fate in the secretory and endocytic pathways. Annu Rev Cell Dev Biol 14:19–57
Brandstetter H, Kim JS, Groll M, Huber R (2001) Crystal structure of the tricorn protease reveals a protein disassembly line. Nature 414(6862):466–470
Bresciani R, Von Figura K (1996) Dephosphorylation of the mannose-6-phosphate recognition marker is localized in later compartments of the endocytic route. Identification of purple acid phosphatase (uteroferrin) as the candidate phosphatase. Eur J Biochem 238(3):669–674
Brew K, Nagase H (2010) The tissue inhibitors of metalloproteinases (TIMPs): an ancient family with structural and functional diversity. Biochim Biophys Acta 1803(1):55–71
Brinckerhoff CE, Matrisian LM (2002) Matrix metalloproteinases: a tail of a frog that became a prince. Nat Rev Mol Cell Biol 3(3):207–214
Brix K (2005) Lysosomal proteases: revival of the sleeping beauty, Chap. 5. In: Saftig P (ed) Lysosomes. Landes Bioscience/Eurekah.com/Springer, Georgetown, TX
Brix K, Herzog V (1994) Extrathyroidal release of thyroid hormones from thyroglobulin by J774 mouse macrophages. J Clin Invest 93(4):1388–1396
Brix K, Jordans S (2005) Watching proteases in action. Nat Chem Biol 1(4):186–187
Brix K, Lemansky P, Herzog V (1996) Evidence for extracellularly acting cathepsins mediating thyroid hormone liberation in thyroid epithelial cells. Endocrinology 137(5):1963–1974
Brix K, Wirtz R, Herzog V (1997) Paracrine interaction between hepatocytes and macrophages after extrathyroidal proteolysis of thyroglobulin. Hepatology 26(5):1232–1240
Brix K, Linke M, Tepel C, Herzog V (2001) Cysteine proteinases mediate extracellular prohormone processing in the thyroid. Biol Chem 382(5):717–725
Brix K, Dunkhorst A, Mayer K, Jordans S (2008) Cysteine cathepsins: cellular roadmap to different functions. Biochimie 90(2):194–207
Brix K, Fuhrer D, Biebermann H (2011) Molecules important for thyroid hormone synthesis and action – known facts and future perspectives. Thyroid Res 4(Suppl 1):S9
Bromme D, Okamoto K (1995) Human cathepsin O2, a novel cysteine protease highly expressed in osteoclastomas and ovary molecular cloning, sequencing and tissue distribution. Biol Chem Hoppe Seyler 376(6):379–384
Brooks P, Fuertes G, Murray RZ, Bose S, Knecht E, Rechsteiner MC, Hendil KB, Tanaka K, Dyson J, Rivett J (2000a) Subcellular localization of proteasomes and their regulatory complexes in mammalian cells. Biochem J 346(Pt 1):155–161
Brooks P, Murray RZ, Mason GG, Hendil KB, Rivett AJ (2000b) Association of immunoproteasomes with the endoplasmic reticulum. Biochem J 352(Pt 3):611–615
Brown MS, Ye J, Rawson RB, Goldstein JL (2000) Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans. Cell 100(4):391–398
Brunner G, Preissner KT (1994) Pericellular enzymatic hydrolysis: implications for the regulation of cell proliferation in the vessel wall and the bone marrow. Blood Coagul Fibrinolysis 5(4):625–639
Budihardjo I, Oliver H, Lutter M, Luo X, Wang X (1999) Biochemical pathways of caspase activation during apoptosis. Annu Rev Cell Dev Biol 15:269–290
Bugge TH, List K, Szabo R (2007) Matriptase-dependent cell surface proteolysis in epithelial development and pathogenesis. Front Biosci 12:5060–5070
Burden RE, Snoddy P, Jefferies CA, Walker B, Scott CJ (2007) Inhibition of cathepsin L-like proteases by cathepsin V propeptide. Biol Chem 388(5):541–545
Buth H, Wolters B, Hartwig B, Meier-Bornheim R, Veith H, Hansen M, Sommerhoff CP, Schaschke N, Machleidt W, Fusenig NE, Boukamp P, Brix K (2004) HaCaT keratinocytes secrete lysosomal cysteine proteinases during migration. Eur J Cell Biol 83(11–12):781–795
Buth H, Luigi Buttigieg P, Ostafe R, Rehders M, Dannenmann SR, Schaschke N, Stark HJ, Boukamp P, Brix K (2007) Cathepsin B is essential for regeneration of scratch-wounded normal human epidermal keratinocytes. Eur J Cell Biol 86(11–12):747–761
Buttle DJ (2007) Factors controlling matrix turnover in health and disease. Biochem Soc Trans 35(Pt 4):643–646
Carlson EE, Cravatt BF (2007) Chemoselective probes for metabolite enrichment and profiling. Nat Methods 4(5):429–435
Caughey GH (2007) Mast cell tryptases and chymases in inflammation and host defense. Immunol Rev 217:141–154
Cauwe B, Opdenakker G (2010) Intracellular substrate cleavage: a novel dimension in the biochemistry, biology and pathology of matrix metalloproteinases. Crit Rev Biochem Mol Biol 45(5):351–423
Cavallo-Medved D, Sloane BF (2003) Cell-surface cathepsin B: understanding its functional significance. Curr Top Dev Biol 54:313–341
Cawston TE, Young DA (2010) Proteinases involved in matrix turnover during cartilage and bone breakdown. Cell Tissue Res 339(1):221–235
Ceru S, Konjar S, Maher K, Repnik U, Krizaj I, Bencina M, Renko M, Nepveu A, Zerovnik E, Turk B, Kopitar-Jerala N (2010) Stefin B interacts with histones and cathepsin L in the nucleus. J Biol Chem 285(13):10078–10086
Chakraborti S, Mandal M, Das S, Mandal A, Chakraborti T (2003) Regulation of matrix metalloproteinases: an overview. Mol Cell Biochem 253(1–2):269–285
Chambers RC, Laurent GJ (2002) Coagulation cascade proteases and tissue fibrosis. Biochem Soc Trans 30(2):194–200
Chapman HA (2004) Cathepsins as transcriptional activators? Dev Cell 6(5):610–611
Chapman HA Jr, Munger JS, Shi GP (1994) The role of thiol proteases in tissue injury and remodeling. Am J Respir Crit Care Med 150(6 Pt 2):S155–S159
Chapman HA, Riese RJ, Shi GP (1997) Emerging roles for cysteine proteases in human biology. Annu Rev Physiol 59:63–88
Chen WT (1996) Proteases associated with invadopodia, and their role in degradation of extracellular matrix. Enzyme Protein 49(1–3):59–71
Chen Y, Klionsky DJ (2011) The regulation of autophagy – unanswered questions. J Cell Sci 124(Pt 2):161–170
Choi SY, Bertram S, Glowacka I, Park YW, Pohlmann S (2009) Type II transmembrane serine proteases in cancer and viral infections. Trends Mol Med 15(7):303–312
Ciechanover A (2003) The ubiquitin proteolytic system and pathogenesis of human diseases: a novel platform for mechanism-based drug targeting. Biochem Soc Trans 31(2):474–481
Ciechanover A (2005) Proteolysis: from the lysosome to ubiquitin and the proteasome. Nat Rev Mol Cell Biol 6(1):79–87
Ciechanover A, Iwai K (2004) The ubiquitin system: from basic mechanisms to the patient bed. IUBMB Life 56(4):193–201
Clegg JS (1991) Metabolic organization and the ultrastructure of animal cells. Biochem Soc Trans 19(4):986–991
Cobbe N, Marshall KM, Gururaja Rao S, Chang CW, Di Cara F, Duca E, Vass S, Kassan A, Heck MM (2009) The conserved metalloprotease invadolysin localizes to the surface of lipid droplets. J Cell Sci 122(Pt 18):3414–3423
Collins GA, Tansey WP (2006) The proteasome: a utility tool for transcription? Curr Opin Genet Dev 16(2):197–202
Conner SD, Schmid SL (2003) Regulated portals of entry into the cell. Nature 422(6927):37–44
Coussens LM, Fingleton B, Matrisian LM (2002) Matrix metalloproteinase inhibitors and cancer: trials and tribulations. Science 295(5564):2387–2392
Creemers JW, Khatib AM (2008) Knock-out mouse models of proprotein convertases: unique functions or redundancy? Front Biosci 13:4960–4971
Dano K, Behrendt N, Hoyer-Hansen G, Johnsen M, Lund LR, Ploug M, Romer J (2005) Plasminogen activation and cancer. Thromb Haemost 93(4):676–681
Das R, Pluskota E, Plow EF (2010) Plasminogen and its receptors as regulators of cardiovascular inflammatory responses. Trends Cardiovasc Med 20(4):120–124
Dash C, Kulkarni A, Dunn B, Rao M (2003) Aspartic peptidase inhibitors: implications in drug development. Crit Rev Biochem Mol Biol 38(2):89–119
Dauth S, Arampatzidou M, Rehders M, Yu DMT, Tedelind S, Brix K (2011a) Thyroid cathepsin K – roles in physiology and thyroid disease. Clin Rev Bone Miner Metab 9:94–106
Dauth S, Sirbulescu RF, Jordans S, Rehders M, Avena L, Oswald J, Lerchl A, Saftig P, Brix K (2011b) Cathepsin K deficiency in mice induces structural and metabolic changes in the central nervous system that are associated with learning and memory deficits. BMC Neurosci 12(1):74
Dauth S, Schmidt MM, Rehders M, Dietz F, Kelm S, Dringen R, Brix K (2012) Characterization and metabolism of astroglia-rich primary cultures from cathepsin K-deficient mice. Biol Chem 393:959–970
Davies KJ (2001) Degradation of oxidized proteins by the 20S proteasome. Biochimie 83(3–4):301–310
De Mot R, Nagy I, Walz J, Baumeister W (1999) Proteasomes and other self-compartmentalizing proteases in prokaryotes. Trends Microbiol 7(2):88–92
De Strooper B, Annaert W (2000) Proteolytic processing and cell biological functions of the amyloid precursor protein. J Cell Sci 113(Pt 11):1857–1870
Declercq W, Vanden Berghe T, Vandenabeele P (2009) RIP kinases at the crossroads of cell death and survival. Cell 138(2):229–232
Degli Esposti M (2008) Organelle intermixing and membrane scrambling in cell death. Methods Enzymol 442:421–438
Desmarais S, Masse F, Percival MD (2009) Pharmacological inhibitors to identify roles of cathepsin K in cell-based studies: a comparison of available tools. Biol Chem 390(9):941–948
Deveraux QL, Stennicke HR, Salvesen GS, Reed JC (1999) Endogenous inhibitors of caspases. J Clin Immunol 19(6):388–398
Di Cara F, Duca E, Dunbar DR, Cagney G, Heck MM (2013) Invadolysin, a conserved lipid droplet-associated metalloprotease, is required for mitochondrial function in Drosophila. J Cell Sci, in press, doi:10.1242/jcs.133306
Dice JF (1987) Molecular determinants of protein half-lives in eukaryotic cells. FASEB J 1(5):349–357
Dice JF (1990) Peptide sequences that target cytosolic proteins for lysosomal proteolysis. Trends Biochem Sci 15(8):305–309
Dice JF (2007) Chaperone-mediated autophagy. Autophagy 3(4):295–299
Dice JF, Walker CD (1979) Protein degradation in metabolic and nutritional disorders. Ciba Found Symp 75:331–350
Dikic I, Schmidt MH (2010) Notch: implications of endogenous inhibitors for therapy. Bioessays 32(6):481–487
Dobson CM (2003) Protein folding and misfolding. Nature 426(6968):884–890
Docherty K, Steiner DF (1982) Post-translational proteolysis in polypeptide hormone biosynthesis. Annu Rev Physiol 44:625–638
Doucet A, Butler GS, Rodriguez D, Prudova A, Overall CM (2008) Metadegradomics: toward in vivo quantitative degradomics of proteolytic post-translational modifications of the cancer proteome. Mol Cell Proteomics 7(10):1925–1951
Drag M, Salvesen GS (2010) Emerging principles in protease-based drug discovery. Nat Rev Drug Discov 9(9):690–701
Driscoll JJ, Dechowdhury R (2010) Therapeutically targeting the SUMOylation, ubiquitination and proteasome pathways as a novel anticancer strategy. Target Oncol 5(4):281–289
Dunn BM, Oda K, Kay J, Rao-Naik C, Lowther WT, Beyer BM, Scarborough PE, Bukhtiyarova M (1998) Comparison of the specificity of the aspartic proteinases towards internally consistent sets of oligopeptide substrates. Adv Exp Med Biol 436:133–138
Earnshaw WC, Martins LM, Kaufmann SH (1999) Mammalian caspases: structure, activation, substrates, and functions during apoptosis. Annu Rev Biochem 68:383–424
Edwards DR, Handsley MM, Pennington CJ (2008) The ADAM metalloproteinases. Mol Aspects Med 29(5):258–289
Ehrmann M, Clausen T (2004) Proteolysis as a regulatory mechanism. Annu Rev Genet 38:709–724
Ellis V (2003) Plasminogen activation at the cell surface. Curr Top Dev Biol 54:263–312
Farquhar MG, Palade GE (1981) The Golgi apparatus (complex)-(1954-1981)-from artifact to center stage. J Cell Biol 91(3 Pt 2):77s–103s
Farquhar MG, Palade GE (1998) The Golgi apparatus: 100 years of progress and controversy. Trends Cell Biol 8(1):2–10
Felberbaum-Corti M, Van Der Goot FG, Gruenberg J (2003) Sliding doors: clathrin-coated pits or caveolae? Nat Cell Biol 5(5):382–384
Fineschi B, Miller J (1997) Endosomal proteases and antigen processing. Trends Biochem Sci 22(10):377–382
Freeman M (2008) Rhomboid proteases and their biological functions. Annu Rev Genet 42:191–210
Fricker LD (1988) Carboxypeptidase E. Annu Rev Physiol 50:309–321
Friedl P, Wolf K (2003) Proteolytic and non-proteolytic migration of tumour cells and leucocytes. Biochem Soc Symp 70:277–285
Friedl P, Wolf K (2009) Proteolytic interstitial cell migration: a five-step process. Cancer Metastasis Rev 28(1–2):129–135
Friedrichs B, Tepel C, Reinheckel T, Deussing J, von Figura K, Herzog V, Peters C, Saftig P, Brix K (2003) Thyroid functions of mouse cathepsins B, K, and L. J Clin Invest 111(11):1733–1745
Frittoli E, Palamidessi A, Disanza A, Scita G (2011) Secretory and endo/exocytic trafficking in invadopodia formation: the MT1-MMP paradigm. Eur J Cell Biol 90(2–3):108–114
Fritz H, Schiessler H, Schleuning WD (1973) Proteinases and proteinase inhibitors in the fertilization process: new concepts of control? Adv Biosci 10:271–286
Fu X, Parks WC, Heinecke JW (2008) Activation and silencing of matrix metalloproteinases. Semin Cell Dev Biol 19(1):2–13
Funkelstein L, Beinfeld M, Minokadeh A, Zadina J, Hook V (2010) Unique biological function of cathepsin L in secretory vesicles for biosynthesis of neuropeptides. Neuropeptides 44(6):457–466
Gagescu R, Gruenberg J, Smythe E (2000) Membrane dynamics in endocytosis: structure–function relationship. Traffic 1(1):84–88
Gahmberg CG, Tolvanen M (1996) Why mammalian cell surface proteins are glycoproteins. Trends Biochem Sci 21(8):308–311
Galivan J, Ryan TJ, Chave K, Rhee M, Yao R, Yin D (2000) Glutamyl hydrolase. Pharmacological role and enzymatic characterization. Pharmacol Ther 85(3):207–215
Garten W, Hallenberger S, Ortmann D, Schafer W, Vey M, Angliker H, Shaw E, Klenk HD (1994) Processing of viral glycoproteins by the subtilisin-like endoprotease furin and its inhibition by specific peptidylchloroalkylketones. Biochimie 76(3–4):217–225
Gerasimenko JV, Gerasimenko OV, Petersen OH (2001) Membrane repair: Ca(2+)-elicited lysosomal exocytosis. Curr Biol 11(23):R971–R974
Gerlt JA, Babbitt PC (2001) Divergent evolution of enzymatic function: mechanistically diverse superfamilies and functionally distinct suprafamilies. Annu Rev Biochem 70:209–246
Gocheva V, Joyce JA (2007) Cysteine cathepsins and the cutting edge of cancer invasion. Cell Cycle 6(1):60–64
Goldberg AL, Dice JF (1974) Intracellular protein degradation in mammalian and bacterial cells. Annu Rev Biochem 43:835–869
Gomez-Lazaro M, Rinn C, Aroso M, Amado F, Schrader M (2010) Proteomic analysis of zymogen granules. Expert Rev Proteomics 7(5):735–747
Gomis-Ruth FX (2008) Structure and mechanism of metallocarboxypeptidases. Crit Rev Biochem Mol Biol 43(5):319–345
Gomis-Ruth FX, Botelho TO, Bode W (2012) A standard orientation for metallopeptidases. Biochim Biophys Acta 1824(1):157–163
Goulet B, Nepveu A (2004) Complete and limited proteolysis in cell cycle progression. Cell Cycle 3(8):986–989
Goulet B, Baruch A, Moon NS, Poirier M, Sansregret LL, Erickson A, Bogyo M, Nepveu A (2004) A cathepsin L isoform that is devoid of a signal peptide localizes to the nucleus in S phase and processes the CDP/Cux transcription factor. Mol Cell 14(2):207–219
Goulet B, Sansregret L, Leduy L, Bogyo M, Weber E, Chauhan SS, Nepveu A (2007) Increased expression and activity of nuclear cathepsin L in cancer cells suggests a novel mechanism of cell transformation. Mol Cancer Res 5(9):899–907
Goulet B, Markovic Y, Leduy L, Nepveu A (2008) Proteolytic processing of cut homeobox 1 by neutrophil elastase in the MV4;11 myeloid leukemia cell line. Mol Cancer Res 6(4):644–653
Greenbaum D, Baruch A, Hayrapetian L, Darula Z, Burlingame A, Medzihradszky KF, Bogyo M (2002) Chemical approaches for functionally probing the proteome. Mol Cell Proteomics 1(1):60–68
Groettrup M, Soza A, Kuckelkorn U, Kloetzel PM (1996) Peptide antigen production by the proteasome: complexity provides efficiency. Immunol Today 17(9):429–435
Groettrup M, Khan S, Schwarz K, Schmidtke G (2001a) Interferon-gamma inducible exchanges of 20S proteasome active site subunits: why? Biochimie 83(3–4):367–372
Groettrup M, van den Broek M, Schwarz K, Macagno A, Khan S, de Giuli R, Schmidtke G (2001b) Structural plasticity of the proteasome and its function in antigen processing. Crit Rev Immunol 21(4):339–358
Groettrup M, Kirk CJ, Basler M (2010) Proteasomes in immune cells: more than peptide producers? Nat Rev Immunol 10(1):73–78
Groll M, Clausen T (2003) Molecular shredders: how proteasomes fulfill their role. Curr Opin Struct Biol 13(6):665–673
Groll M, Ditzel L, Lowe J, Stock D, Bochtler M, Bartunik HD, Huber R (1997) Structure of 20S proteasome from yeast at 2.4 A resolution. Nature 386(6624):463–471
Gruenberg J (2001) The endocytic pathway: a mosaic of domains. Nat Rev Mol Cell Biol 2(10):721–730
Gruenberg J, Howell KE (1989) Membrane traffic in endocytosis: insights from cell-free assays. Annu Rev Cell Biol 5:453–481
Gruenberg J, Maxfield FR (1995) Membrane transport in the endocytic pathway. Curr Opin Cell Biol 7(4):552–563
Gruenberg J, Stenmark H (2004) The biogenesis of multivesicular endosomes. Nat Rev Mol Cell Biol 5(4):317–323
Gu F, Gruenberg J (1999) Biogenesis of transport intermediates in the endocytic pathway. FEBS Lett 452(1–2):61–66
Gu F, Aniento F, Parton RG, Gruenberg J (1997) Functional dissection of COP-I subunits in the biogenesis of multivesicular endosomes. J Cell Biol 139(5):1183–1195
Gumy LF, Tan CL, Fawcett JW (2010) The role of local protein synthesis and degradation in axon regeneration. Exp Neurol 223(1):28–37
Ha Y (2007) Structural principles of intramembrane proteases. Curr Opin Struct Biol 17(4):405–411
Hajjar KA, Acharya SS (2000) Annexin II and regulation of cell surface fibrinolysis. Ann NY Acad Sci 902:265–271
Halangk W, Lerch MM, Brandt-Nedelev B, Roth W, Ruthenbuerger M, Reinheckel T, Domschke W, Lippert H, Peters C, Deussing J (2000) Role of cathepsin B in intracellular trypsinogen activation and the onset of acute pancreatitis. J Clin Invest 106(6):773–781
Hansen TH, Bouvier M (2009) MHC class I antigen presentation: learning from viral evasion strategies. Nat Rev Immunol 9(7):503–513
Haugen MH, Johansen HT, Pettersen SJ, Solberg R, Brix K, Flatmark K, Maelandsmo GM (2013) Nuclear legumain activity in colorectal cancer. PLoS One 8(1):e52980
He B, Lu N, Zhou Z (2009) Cellular and nuclear degradation during apoptosis. Curr Opin Cell Biol 21(6):900–912
Heck MM (1997) Condensins, cohesins, and chromosome architecture: how to make and break a mitotic chromosome. Cell 91(1):5–8
Hendrickx AP, Budzik JM, Oh SY, Schneewind O (2011) Architects at the bacterial surface – sortases and the assembly of pili with isopeptide bonds. Nat Rev Microbiol 9(3):166–176
Henrich S, Lindberg I, Bode W, Than ME (2005) Proprotein convertase models based on the crystal structures of furin and kexin: explanation of their specificity. J Mol Biol 345(2):211–227
Herget M, Tampe R (2007) Intracellular peptide transporters in human–compartmentalization of the “peptidome”. Pflugers Arch 453(5):591–600
Hicke L, Dunn R (2003) Regulation of membrane protein transport by ubiquitin and ubiquitin-binding proteins. Annu Rev Cell Dev Biol 19:141–172
Hille A, Klumperman J, Geuze HJ, Peters C, Brodsky FM, von Figura K (1992) Lysosomal acid phosphatase is internalized via clathrin-coated pits. Eur J Cell Biol 59(1):106–115
Hiraiwa M (1999) Cathepsin A/protective protein: an unusual lysosomal multifunctional protein. Cell Mol Life Sci 56(11–12):894–907
Holmbeck K, Bianco P, Caterina J, Yamada S, Kromer M, Kuznetsov SA, Mankani M, Robey PG, Poole AR, Pidoux I, Ward JM, Birkedal-Hansen H (1999) MT1-MMP-deficient mice develop dwarfism, osteopenia, arthritis, and connective tissue disease due to inadequate collagen turnover. Cell 99(1):81–92
Hook V, Yasothornsrikul S, Greenbaum D, Medzihradszky KF, Troutner K, Toneff T, Bundey R, Logrinova A, Reinheckel T, Peters C, Bogyo M (2004) Cathepsin L and Arg/Lys aminopeptidase: a distinct prohormone processing pathway for the biosynthesis of peptide neurotransmitters and hormones. Biol Chem 385(6):473–480
Hook V, Schechter I, Demuth HU, Hook G (2008) Alternative pathways for production of beta-amyloid peptides of Alzheimer’s disease. Biol Chem 389(8):993–1006
Hooper NM (2002) Proteases: a primer. Essays Biochem 38:1–8
Hooper JD, Clements JA, Quigley JP, Antalis TM (2001) Type II transmembrane serine proteases. Insights into an emerging class of cell surface proteolytic enzymes. J Biol Chem 276(2):857–860
Howell KE, Devaney E, Gruenberg J (1989) Subcellular fractionation of tissue culture cells. Trends Biochem Sci 14(2):44–47
Hudson DF, Marshall KM, Earnshaw WC (2009) Condensin: architect of mitotic chromosomes. Chromosome Res 17(2):131–144
Ichihara A, Kaneshiro Y, Suzuki F (2006) Prorenin receptor blockers: effects on cardiovascular complications of diabetes and hypertension. Expert Opin Investig Drugs 15(10):1137–1139
Ishidoh K, Kominami E (2002) Processing and activation of lysosomal proteinases. Biol Chem 383(12):1827–1831
Itoh Y (2006) MT1-MMP: a key regulator of cell migration in tissue. IUBMB Life 58(10):589–596
Ivanova S, Repnik U, Bojic L, Petelin A, Turk V, Turk B (2008) Lysosomes in apoptosis. Methods Enzymol 442:183–199
Izuhara K, Ohta S, Kanaji S, Shiraishi H, Arima K (2008) Recent progress in understanding the diversity of the human ov-serpin/clade B serpin family. Cell Mol Life Sci 65(16):2541–2553
Jacques LB (1979) Heparin: an old drug with a new paradigm. Science 206(4418):528–533
Jedeszko C, Sameni M, Olive MB, Moin K, Sloane BF (2008) Visualizing protease activity in living cells: from two dimensions to four dimensions. Curr Protoc Cell Biol Chapter 4:Unit 4.20
Jiang A, Lehti K, Wang X, Weiss SJ, Keski-Oja J, Pei D (2001) Regulation of membrane-type matrix metalloproteinase 1 activity by dynamin-mediated endocytosis. Proc Natl Acad Sci U S A 98(24):13693–13698
Jordans S, Jenko-Kokalj S, Kuhl NM, Tedelind S, Sendt W, Bromme D, Turk D, Brix K (2009) Monitoring compartment-specific substrate cleavage by cathepsins B, K, L, and S at physiological pH and redox conditions. BMC Biochem 10:23
Joyce JA, Hanahan D (2004) Multiple roles for cysteine cathepsins in cancer. Cell Cycle 3(12):1516–1619
Kaiser CA, Chen EJ, Losko S (2002) Subcellular fractionation of secretory organelles. Methods Enzymol 351:325–338
Kaiserman D, Whisstock JC, Bird PI (2006) Mechanisms of serpin dysfunction in disease. Expert Rev Mol Med 8(31):1–19
Katzmann DJ, Odorizzi G, Emr SD (2002) Receptor downregulation and multivesicular-body sorting. Nat Rev Mol Cell Biol 3(12):893–905
Kennedy RD, D’Andrea AD (2005) The Fanconi Anemia/BRCA pathway: new faces in the crowd. Genes Dev 19(24):2925–2940
Kenny AJ, Maroux S (1982) Topology of microvillar membrance hydrolases of kidney and intestine. Physiol Rev 62(1):91–128
Kersse K, Verspurten J, Berghe TV, Vandenabeele P (2011) The death-fold superfamily of homotypic interaction motifs. Trends Biochem Sci 36(10):541–552
Kessenbrock K, Dau T, Jenne DE (2011) Tailor-made inflammation: how neutrophil serine proteases modulate the inflammatory response. J Mol Med 89(1):23–28
Khan AR, James MN (1998) Molecular mechanisms for the conversion of zymogens to active proteolytic enzymes. Protein Sci 7(4):815–836
Kidd D, Liu Y, Cravatt BF (2001) Profiling serine hydrolase activities in complex proteomes. Biochemistry 40(13):4005–4015
Kim J, Hajjar KA (2002) Annexin II: a plasminogen-plasminogen activator co-receptor. Front Biosci 7:d341–d348
Kirschke H, Wiederanders B (1994) Cathepsin S and related lysosomal endopeptidases. Methods Enzymol 244:500–511
Kirschke H, Barrett AJ, Rawlings ND (1995) Proteinases 1: lysosomal cysteine proteinases. Protein Profile 2(14):1581–1643
Klionsky DJ (2007) Autophagy: from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol 8(11):931–937
Kloetzel PM, Ossendorp F (2004) Proteasome and peptidase function in MHC-class-I-mediated antigen presentation. Curr Opin Immunol 16(1):76–81
Koohmaraie M (1992) The role of Ca(2+)-dependent proteases (calpains) in post mortem proteolysis and meat tenderness. Biochimie 74(3):239–245
Kopito RR (2000) Aggresomes, inclusion bodies and protein aggregation. Trends Cell Biol 10(12):524–530
Korkmaz B, Moreau T, Gauthier F (2008) Neutrophil elastase, proteinase 3 and cathepsin G: physicochemical properties, activity and physiopathological functions. Biochimie 90(2):227–242
Kornfeld S (1992) Structure and function of the mannose 6-phosphate/insulinlike growth factor II receptors. Annu Rev Biochem 61:307–330
Koster A, Saftig P, Matzner U, von Figura K, Peters C, Pohlmann R (1993) Targeted disruption of the M(r) 46,000 mannose 6-phosphate receptor gene in mice results in misrouting of lysosomal proteins. EMBO J 12(13):5219–5223
Krysko DV, Agostinis P, Krysko O, Garg AD, Bachert C, Lambrecht BN, Vandenabeele P (2011) Emerging role of damage-associated molecular patterns derived from mitochondria in inflammation. Trends Immunol 32(4):157–164
Kveiborg M, Albrechtsen R, Couchman JR, Wewer UM (2008) Cellular roles of ADAM12 in health and disease. Int J Biochem Cell Biol 40(9):1685–1702
Kwak J, Workman JL, Lee D (2011) The proteasome and its regulatory roles in gene expression. Biochim Biophys Acta 1809(2):88–96
Lamkanfi M, Festjens N, Declercq W, Vanden Berghe T, Vandenabeele P (2007) Caspases in cell survival, proliferation and differentiation. Cell Death Differ 14(1):44–55
Lamond AI, Earnshaw WC (1998) Structure and function in the nucleus. Science 280(5363):547–553
Lecaille F, Weidauer E, Juliano MA, Bromme D, Lalmanach G (2003) Probing cathepsin K activity with a selective substrate spanning its active site. Biochem J 375(Pt 2):307–312
Lemansky P, Brix K, Herzog V (1998) Iodination of mature cathepsin D in thyrocytes as an indicator for its transport to the cell surface. Eur J Cell Biol 76(1):53–62
Lenarcic B, Bevec T (1998) Thyropins–new structurally related proteinase inhibitors. Biol Chem 379(2):105–111
Liaudet-Coopman E, Beaujouin M, Derocq D, Garcia M, Glondu-Lassis M, Laurent-Matha V, Prebois C, Rochefort H, Vignon F (2006) Cathepsin D: newly discovered functions of a long-standing aspartic protease in cancer and apoptosis. Cancer Lett 237(2):167–179
Lichtenthaler SF, Haass C, Steiner H (2011) Regulated intramembrane proteolysis–lessons from amyloid precursor protein processing. J Neurochem 117(5):779–796
Lin SX, Mallet WG, Huang AY, Maxfield FR (2004) Endocytosed cation-independent mannose 6-phosphate receptor traffics via the endocytic recycling compartment en route to the trans-Golgi network and a subpopulation of late endosomes. Mol Biol Cell 15(2):721–733
Linder S (2007) The matrix corroded: podosomes and invadopodia in extracellular matrix degradation. Trends Cell Biol 17(3):107–117
Linke M, Herzog V, Brix K (2002a) Trafficking of lysosomal cathepsin B-green fluorescent protein to the surface of thyroid epithelial cells involves the endosomal/lysosomal compartment. J Cell Sci 115(Pt 24):4877–4889
Linke M, Jordans S, Mach L, Herzog V, Brix K (2002b) Thyroid stimulating hormone upregulates secretion of cathepsin B from thyroid epithelial cells. Biol Chem 383(5):773–784
Lipowsky R (1995) The morphology of lipid membranes. Curr Opin Struct Biol 5(4):531–540
List K, Bugge TH, Szabo R (2006) Matriptase: potent proteolysis on the cell surface. Mol Med 12(1–3):1–7
Liu H, Lazarus SC, Caughey GH, Fahy JV (1999) Neutrophil elastase and elastase-rich cystic fibrosis sputum degranulate human eosinophils in vitro. Am J Physiol 276(1 Pt 1):L28–L34
Lopez-Otin C, Bond JS (2008) Proteases: multifunctional enzymes in life and disease. J Biol Chem 283(45):30433–30437
Lopez-Otin C, Matrisian LM (2007) Emerging roles of proteases in tumour suppression. Nat Rev Cancer 7(10):800–808
Ludwig T, Munier-Lehmann H, Bauer U, Hollinshead M, Ovitt C, Lobel P, Hoflack B (1994) Differential sorting of lysosomal enzymes in mannose 6-phosphate receptor-deficient fibroblasts. EMBO J 13(15):3430–3437
Mach L (2002) Biosynthesis of lysosomal proteinases in health and disease. Biol Chem 383(5):751–756
Mach L, Mort JS, Glossl J (1994) Maturation of human procathepsin B. Proenzyme activation and proteolytic processing of the precursor to the mature proteinase, in vitro, are primarily unimolecular processes. J Biol Chem 269(17):13030–13035
Mackie EJ, Pagel CN, Smith R, de Niese MR, Song SJ, Pike RN (2002) Protease-activated receptors: a means of converting extracellular proteolysis into intracellular signals. IUBMB Life 53(6):277–281
Magdolen V, Arroyo de Prada N, Sperl S, Muehlenweg B, Luther T, Wilhelm OG, Magdolen U, Graeff H, Reuning U, Schmitt M (2000) Natural and synthetic inhibitors of the tumor-associated serine protease urokinase-type plasminogen activator. Adv Exp Med Biol 477:331–341
Mai J, Waisman DM, Sloane BF (2000) Cell surface complex of cathepsin B/annexin II tetramer in malignant progression. Biochim Biophys Acta 1477(1–2):215–230
Manon-Jensen T, Itoh Y, Couchman JR (2010) Proteoglycans in health and disease: the multiple roles of syndecan shedding. FEBS J 277(19):3876–3889
Marfany G, Denuc A (2008) To ubiquitinate or to deubiquitinate: it all depends on the partners. Biochem Soc Trans 36(Pt 5):833–838
Matrisian LM (1999) Cancer biology: extracellular proteinases in malignancy. Curr Biol 9(20):R776–R778
Matteucci E, Giampietro O (2009) Dipeptidyl peptidase-4 (CD26): knowing the function before inhibiting the enzyme. Curr Med Chem 16(23):2943–2951
Mayer K, Schwartz S, Lentze MJ, Kalff JC, Brix K (2006) Extracellular localization of intestinal cathepsins: implications of their actions during post-operative ileus. In: XLI congress of the European society for surgical research. Medimond International Proceedings, Rostock, Germany
Mayer K, Iolyeva ME, Meyer-Grahle U, Brix K (2008) Intestine-specific expression of green fluorescent protein-tagged cathepsin B: proof-of-principle experiments. Biol Chem 389(8):1085–1096
Mayer K, Vreemann A, Qu H, Brix K (2009) Release of endo-lysosomal cathepsins B, D, and L from IEC6 cells in a cell culture model mimicking intestinal manipulation. Biol Chem 390(5–6):471–480
McGrath ME (1999) The lysosomal cysteine proteases. Annu Rev Biophys Biomol Struct 28:181–204
McHugh B, Krause SA, Yu B, Deans AM, Heasman S, McLaughlin P, Heck MM (2004) Invadolysin: a novel, conserved metalloprotease links mitotic structural rearrangements with cell migration. J Cell Biol 167(4):673–686
McIntyre GF, Godbold GD, Erickson AH (1994) The pH-dependent membrane association of procathepsin L is mediated by a 9-residue sequence within the propeptide. J Biol Chem 269(1):567–572
McNeil PL, Steinhardt RA (1997) Loss, restoration, and maintenance of plasma membrane integrity. J Cell Biol 137(1):1–4
Meister T, Niehues R, Hahn D, Domschke W, Sendler M, Lerch MM, Schnekenburger J (2010) Missorting of cathepsin B into the secretory compartment of CI-MPR/IGFII-deficient mice does not induce spontaneous trypsinogen activation but leads to enhanced trypsin activity during experimental pancreatitis–without affecting disease severit. J Physiol Pharmacol 61(5):565–575
Mentlein R (2004) Cell-surface peptidases. Int Rev Cytol 235:165–213
Meusser B, Hirsch C, Jarosch E, Sommer T (2005) ERAD: the long road to destruction. Nat Cell Biol 7(8):766–772
Meyer-Hoffert U (2009) Neutrophil-derived serine proteases modulate innate immune responses. Front Biosci 14:3409–3418
Mizushima N, Levine B, Cuervo AM, Klionsky DJ (2008) Autophagy fights disease through cellular self-digestion. Nature 451(7182):1069–1075
Mockaitis K, Estelle M (2008) Auxin receptors and plant development: a new signaling paradigm. Annu Rev Cell Dev Biol 24:55–80
Mohamed MM, Sloane BF (2006) Cysteine cathepsins: multifunctional enzymes in cancer. Nat Rev Cancer 6(10):764–775
Mondino A, Blasi F (2004) uPA and uPAR in fibrinolysis, immunity and pathology. Trends Immunol 25(8):450–455
Morita M, Kurochkin IV, Motojima K, Goto S, Takano T, Okamura S, Sato R, Yokota S, Imanaka T (2000) Insulin-degrading enzyme exists inside of rat liver peroxisomes and degrades oxidized proteins. Cell Struct Funct 25(5):309–315
Mort JS, Buttle DJ (1997) Cathepsin B. Int J Biochem Cell Biol 29(5):715–720
Mort JS, Recklies AD, Poole AR (1984) Extracellular presence of the lysosomal proteinase cathepsin B in rheumatoid synovium and its activity at neutral pH. Arthritis Rheum 27(5):509–515
Mott JD, Werb Z (2004) Regulation of matrix biology by matrix metalloproteinases. Curr Opin Cell Biol 16(5):558–564
Mukherjee S, Ghosh RN, Maxfield FR (1997) Endocytosis. Physiol Rev 77(3):759–803
Munshi HG, Stack MS (2006) Reciprocal interactions between adhesion receptor signaling and MMP regulation. Cancer Metastasis Rev 25(1):45–56
Murphy G (2009) Regulation of the proteolytic disintegrin metalloproteinases, the ‘Sheddases’. Semin Cell Dev Biol 20(2):138–145
Murphy G, Gavrilovic J (1999) Proteolysis and cell migration: creating a path? Curr Opin Cell Biol 11(5):614–621
Murphy G, Nagase H (2011) Localizing matrix metalloproteinase activities in the pericellular environment. FEBS J 278(1):2–15
Murphy JE, Padilla BE, Hasdemir B, Cottrell GS, Bunnett NW (2009) Endosomes: a legitimate platform for the signaling train. Proc Natl Acad Sci U S A 106(42):17615–17622
Nagase H, Suzuki K, Itoh Y, Kan CC, Gehring MR, Huang W, Brew K (1996) Involvement of tissue inhibitors of metalloproteinases (TIMPS) during matrix metalloproteinase activation. Adv Exp Med Biol 389:23–31
Nakagawa T, Roth W, Wong P, Nelson A, Farr A, Deussing J, Villadangos JA, Ploegh H, Peters C, Rudensky AY (1998) Cathepsin L: critical role in Ii degradation and CD4 T cell selection in the thymus. Science 280(5362):450–453
Nakanishi H (2003) Microglial functions and proteases. Mol Neurobiol 27(2):163–176
Nakayama K (1997) Furin: a mammalian subtilisin/Kex2p-like endoprotease involved in processing of a wide variety of precursor proteins. Biochem J 327(Pt 3):625–635
Nasmyth K, Peters JM, Uhlmann F (2000) Splitting the chromosome: cutting the ties that bind sister chromatids. Science 288(5470):1379–1385
Navarro A, Anand-Apte B, Parat MO (2004) A role for caveolae in cell migration. FASEB J 18(15):1801–1811
Nichols BJ, Lippincott-Schwartz J (2001) Endocytosis without clathrin coats. Trends Cell Biol 11(10):406–412
Nicotra G, Castino R, Follo C, Peracchio C, Valente G, Isidoro C (2010) The dilemma: does tissue expression of cathepsin D reflect tumor malignancy? The question: does the assay truly mirror cathepsin D mis-function in the tumor? Cancer Biomark 7(1):47–64
Nixon RA, Cataldo AM (1993) The lysosomal system in neuronal cell death: a review. Ann NY Acad Sci 679:87–109
Noel A, Maillard C, Rocks N, Jost M, Chabottaux V, Sounni NE, Maquoi E, Cataldo D, Foidart JM (2004) Membrane associated proteases and their inhibitors in tumour angiogenesis. J Clin Pathol 57(6):577–584
Norman LP, Matters GL, Crisman JM, Bond JS (2003) Expression of meprins in health and disease. Curr Top Dev Biol 54:145–166
Novinec M, Kordis D, Turk V, Lenarcic B (2006) Diversity and evolution of the thyroglobulin type-1 domain superfamily. Mol Biol Evol 23(4):744–755
O’Brien P, O’Connor BF (2008) Seprase: an overview of an important matrix serine protease. Biochim Biophys Acta 1784(9):1130–1145
Olson TS, Dice JF (1989) Regulation of protein degradation rates in eukaryotes. Curr Opin Cell Biol 1(6):1194–1200
Ong PC, McGowan S, Pearce MC, Irving JA, Kan WT, Grigoryev SA, Turk B, Silverman GA, Brix K, Bottomley SP, Whisstock JC, Pike RN (2007) DNA accelerates the inhibition of human cathepsin V by serpins. J Biol Chem 282(51):36980–36986
Ovaere P, Lippens S, Vandenabeele P, Declercq W (2009) The emerging roles of serine protease cascades in the epidermis. Trends Biochem Sci 34(9):453–463
Overall CM, Blobel CP (2007) In search of partners: linking extracellular proteases to substrates. Nat Rev Mol Cell Biol 8(3):245–257
Owen CA (2008) Leukocyte cell surface proteinases: regulation of expression, functions, and mechanisms of surface localization. Int J Biochem Cell Biol 40(6–7):1246–1272
Owen CA, Campbell EJ (1995) Neutrophil proteinases and matrix degradation. The cell biology of pericellular proteolysis. Semin Cell Biol 6(6):367–376
Pagano M, Reboud-Ravaux M (2011) Cryptic activities of fibronectin fragments, particularly cryptic proteases. Front Biosci 16:698–706
Palade GE (1964) The organization of living matter. Proc Natl Acad Sci U S A 52:613–634
Palade GE (1966) Structure and function at the cellular level. JAMA 198(8):815–825
Palade G (1975) Intracellular aspects of the process of protein synthesis. Science 189(4200):347–358
Pappot H, Gardsvoll H, Romer J, Pedersen AN, Grondahl-Hansen J, Pyke C, Brunner N (1995) Plasminogen activator inhibitor type 1 in cancer: therapeutic and prognostic implications. Biol Chem Hoppe Seyler 376(5):259–267
Parkin ET, Turner AJ, Hooper NM (2004) Secretase-mediated cell surface shedding of the angiotensin-converting enzyme. Protein Pept Lett 11(5):423–432
Paulick MG, Bogyo M (2008) Application of activity-based probes to the study of enzymes involved in cancer progression. Curr Opin Genet Dev 18(1):97–106
Pelkmans L, Helenius A (2002) Endocytosis via caveolae. Traffic 3(5):311–320
Pellman D, Christman MF (2001) Separase anxiety: dissolving the sister bond and more. Nat Cell Biol 3(9):E207–E209
Peters C, von Figura K (1994) Biogenesis of lysosomal membranes. FEBS Lett 346(1):108–114
Peters C, Braun M, Weber B, Wendland M, Schmidt B, Pohlmann R, Waheed A, von Figura K (1990) Targeting of a lysosomal membrane protein: a tyrosine-containing endocytosis signal in the cytoplasmic tail of lysosomal acid phosphatase is necessary and sufficient for targeting to lysosomes. EMBO J 9(11):3497–3506
Pham CT (2006) Neutrophil serine proteases: specific regulators of inflammation. Nat Rev Immunol 6(7):541–550
Pham CT (2008) Neutrophil serine proteases fine-tune the inflammatory response. Int J Biochem Cell Biol 40(6–7):1317–1333
Pickart CM (1997) Targeting of substrates to the 26S proteasome. FASEB J 11(13):1055–1066
Pickart CM (2001) Mechanisms underlying ubiquitination. Annu Rev Biochem 70:503–533
Pickart CM, Eddins MJ (2004) Ubiquitin: structures, functions, mechanisms. Biochim Biophys Acta 1695(1–3):55–72
Pickart CM, Fushman D (2004) Polyubiquitin chains: polymeric protein signals. Curr Opin Chem Biol 8(6):610–616
Pillay CS, Elliott E, Dennison C (2002) Endolysosomal proteolysis and its regulation. Biochem J 363(Pt 3):417–429
Piper RC, Katzmann DJ (2007) Biogenesis and function of multivesicular bodies. Annu Rev Cell Dev Biol 23:519–547
Platta HW, Erdmann R (2007) Peroxisomal dynamics. Trends Cell Biol 17(10):474–484
Podgorski I (2009) Future of anticathepsin K drugs: dual therapy for skeletal disease and atherosclerosis? Future Med Chem 1(1):21–34
Pohlmann R, Boeker MW, von Figura K (1995) The two mannose 6-phosphate receptors transport distinct complements of lysosomal proteins. J Biol Chem 270(45):27311–27318
Pop C, Salvesen GS (2009) Human caspases: activation, specificity, and regulation. J Biol Chem 284(33):21777–21781
Potempa J, Sroka A, Imamura T, Travis J (2003) Gingipains, the major cysteine proteinases and virulence factors of Porphyromonas gingivalis: structure, function and assembly of multidomain protein complexes. Curr Protein Pept Sci 4(6):397–407
Predescu SA, Predescu DN, Palade GE (2001) Endothelial transcytotic machinery involves supramolecular protein-lipid complexes. Mol Biol Cell 12(4):1019–1033
Pungercar JR, Caglic D, Sajid M, Dolinar M, Vasiljeva O, Pozgan U, Turk D, Bogyo M, Turk V, Turk B (2009) Autocatalytic processing of procathepsin B is triggered by proenzyme activity. FEBS J 276(3):660–668
Rachner TD, Khosla S, Hofbauer LC (2011) Osteoporosis: now and the future. Lancet 377(9773):1276–1287
Ramsay AJ, Hooper JD, Folgueras AR, Velasco G, Lopez-Otin C (2009) Matriptase-2 (TMPRSS6): a proteolytic regulator of iron homeostasis. Haematologica 94(6):840–849
Rao JS (2003) Molecular mechanisms of glioma invasiveness: the role of proteases. Nat Rev Cancer 3(7):489–501
Rawlings ND (2010) Peptidase inhibitors in the MEROPS database. Biochimie 92(11):1463–1483
Rawlings ND, Tolle DP, Barrett AJ (2004) Evolutionary families of peptidase inhibitors. Biochem J 378(Pt 3):705–716
Rechsteiner M (1990) PEST sequences are signals for rapid intracellular proteolysis. Semin Cell Biol 1(6):433–440
Rechsteiner M, Hill CP (2005) Mobilizing the proteolytic machine: cell biological roles of proteasome activators and inhibitors. Trends Cell Biol 15(1):27–33
Reed SI (2003) Ratchets and clocks: the cell cycle, ubiquitylation and protein turnover. Nat Rev Mol Cell Biol 4(11):855–864
Reinheckel T, Grune T, Davies KJ (2000) The measurement of protein degradation in response to oxidative stress. Methods Mol Biol 99:49–60
Reinheckel T, Deussing J, Roth W, Peters C (2001) Towards specific functions of lysosomal cysteine peptidases: phenotypes of mice deficient for cathepsin B or cathepsin L. Biol Chem 382(5):735–741
Reiser J, Adair B, Reinheckel T (2010) Specialized roles for cysteine cathepsins in health and disease. J Clin Invest 120(10):3421–3431
Reiss K, Saftig P (2009) The “a disintegrin and metalloprotease” (ADAM) family of sheddases: physiological and cellular functions. Semin Cell Dev Biol 20(2):126–137
Reudelhuber TL, Brechler V, Jutras I, Mercure C, Methot D (1998) Proteolytic and non-proteolytic activation of prorenin. Adv Exp Med Biol 436:229–238
Reverter D, Sorimachi H, Bode W (2001) The structure of calcium-free human m-calpain: implications for calcium activation and function. Trends Cardiovasc Med 11(6):222–229
Reyes-Turcu FE, Ventii KH, Wilkinson KD (2009) Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annu Rev Biochem 78:363–397
Riedl SJ, Salvesen GS (2007) The apoptosome: signalling platform of cell death. Nat Rev Mol Cell Biol 8(5):405–413
Riedl SJ, Shi Y (2004) Molecular mechanisms of caspase regulation during apoptosis. Nat Rev Mol Cell Biol 5(11):897–907
Riese RJ, Chapman HA (2000) Cathepsins and compartmentalization in antigen presentation. Curr Opin Immunol 12(1):107–113
Rivett AJ (1998) Intracellular distribution of proteasomes. Curr Opin Immunol 10(1):110–114
Rivett AJ, Hearn AR (2004) Proteasome function in antigen presentation: immunoproteasome complexes, Peptide production, and interactions with viral proteins. Curr Protein Pept Sci 5(3):153–161
Rochefort H, Garcia M, Glondu M, Laurent V, Liaudet E, Rey JM, Roger P (2000) Cathepsin D in breast cancer: mechanisms and clinical applications, a 1999 overview. Clin Chim Acta 291(2):157–170
Rockwell NC, Thorner JW (2004) The kindest cuts of all: crystal structures of Kex2 and furin reveal secrets of precursor processing. Trends Biochem Sci 29(2):80–87
Romisch K (2005) Endoplasmic reticulum-associated degradation. Annu Rev Cell Dev Biol 21:435–456
Rosenberg GA (2009) Matrix metalloproteinases and their multiple roles in neurodegenerative diseases. Lancet Neurol 8(2):205–216
Rosenblum JS, Kozarich JW (2003) Prolyl peptidases: a serine protease subfamily with high potential for drug discovery. Curr Opin Chem Biol 7(4):496–504
Rothman JE, Wieland FT (1996) Protein sorting by transport vesicles. Science 272(5259):227–234
Ruchaud S, Carmena M, Earnshaw WC (2007) Chromosomal passengers: conducting cell division. Nat Rev Mol Cell Biol 8(10):798–812
Sachse M, Urbe S, Oorschot V, Strous GJ, Klumperman J (2002) Bilayered clathrin coats on endosomal vacuoles are involved in protein sorting toward lysosomes. Mol Biol Cell 13(4):1313–1328
Saftig P, Reiss K (2011) The “A Disintegrin And Metalloproteases” ADAM10 and ADAM17: novel drug targets with therapeutic potential? Eur J Cell Biol 90(6–7):527–535
Saftig P, Hunziker E, Wehmeyer O, Jones S, Boyde A, Rommerskirch W, Moritz JD, Schu P, von Figura K (1998) Impaired osteoclastic bone resorption leads to osteopetrosis in cathepsin-K-deficient mice. Proc Natl Acad Sci U S A 95(23):13453–13458
Salvesen GS (2002) Caspases and apoptosis. Essays Biochem 38:9–19
Salvesen GS, Dixit VM (1997) Caspases: intracellular signaling by proteolysis. Cell 91(4):443–446
Salvesen GS, Duckett CS (2002) IAP proteins: blocking the road to death’s door. Nat Rev Mol Cell Biol 3(6):401–410
Sameni M, Cavallo-Medved D, Dosescu J, Jedeszko C, Moin K, Mullins SR, Olive MB, Rudy D, Sloane BF (2009) Imaging and quantifying the dynamics of tumor-associated proteolysis. Clin Exp Metastasis 26(4):299–309
Sato K, Kawashima S (2001) Calpain function in the modulation of signal transduction molecules. Biol Chem 382(5):743–751
Sato H, Takino T (2010) Coordinate action of membrane-type matrix metalloproteinase-1 (MT1-MMP) and MMP-2 enhances pericellular proteolysis and invasion. Cancer Sci 101(4):843–847
Scamuffa N, Calvo F, Chretien M, Seidah NG, Khatib AM (2006) Proprotein convertases: lessons from knockouts. FASEB J 20(12):1954–1963
Schaller A, Ryan CA (1996) Systemin–a polypeptide defense signal in plants. Bioessays 18(1):27–33
Schaschke N, Assfalg-Machleidt I, Machleidt W, Moroder L (1998) Substrate/propeptide-derived endo-epoxysuccinyl peptides as highly potent and selective cathepsin B inhibitors. FEBS Lett 421(1):80–82
Schechter I (2005) Mapping of the active site of proteases in the 1960s and rational design of inhibitors/drugs in the 1990s. Curr Protein Pept Sci 6(6):501–512
Schenk S, Quaranta V (2003) Tales from the crypt[ic] sites of the extracellular matrix. Trends Cell Biol 13(7):366–375
Schmid SL (1993) Toward a biochemical definition of the endosomal compartment. Studies using free flow electrophoresis. Subcell Biochem 19:1–28
Schwartz DC, Hochstrasser M (2003) A superfamily of protein tags: ubiquitin, SUMO and related modifiers. Trends Biochem Sci 28(6):321–328
Schweitzer K, Naumann M (2010) Control of NF-kappaB activation by the COP9 signalosome. Biochem Soc Trans 38(Pt 1):156–161
Scott CC, Gruenberg J (2011) Ion flux and the function of endosomes and lysosomes: pH is just the start: the flux of ions across endosomal membranes influences endosome function not only through regulation of the luminal pH. Bioessays 33(2):103–110
Scott CJ, Taggart CC (2010) Biologic protease inhibitors as novel therapeutic agents. Biochimie 92(11):1681–1688
Seglen PO, Berg TO, Blankson H, Fengsrud M, Holen I, Stromhaug PE (1996) Structural aspects of autophagy. Adv Exp Med Biol 389:103–111
Seidah NG (2011) What lies ahead for the proprotein convertases? Ann NY Acad Sci 1220:149–161
Seidah NG, Chretien M (1999) Proprotein and prohormone convertases: a family of subtilases generating diverse bioactive polypeptides. Brain Res 848(1–2):45–62
Seidah NG, Prat A (2002) Precursor convertases in the secretory pathway, cytosol and extracellular milieu. Essays Biochem 38:79–94
Seidah NG, Day R, Marcinkiewicz M, Benjannet S, Chretien M (1991) Mammalian neural and endocrine pro-protein and pro-hormone convertases belonging to the subtilisin family of serine proteinases. Enzyme 45(5–6):271–284
Seiki M (2002) The cell surface: the stage for matrix metalloproteinase regulation of migration. Curr Opin Cell Biol 14(5):624–632
Selkoe DJ, Wolfe MS (2007) Presenilin: running with scissors in the membrane. Cell 131(2):215–221
Shi Z, Stack MS (2007) Urinary-type plasminogen activator (uPA) and its receptor (uPAR) in squamous cell carcinoma of the oral cavity. Biochem J 407(2):153–159
Silverman GA, Bird PI, Carrell RW, Church FC, Coughlin PB, Gettins PG, Irving JA, Lomas DA, Luke CJ, Moyer RW, Pemberton PA, Remold-O’Donnell E, Salvesen GS, Travis J, Whisstock JC (2001) The serpins are an expanding superfamily of structurally similar but functionally diverse proteins. Evolution, mechanism of inhibition, novel functions, and a revised nomenclature. J Biol Chem 276(36):33293–33296
Simionescu N, Simionescu M, Palade GE (1972) Permeability of intestinal capillaries. Pathway followed by dextrans and glycogens. J Cell Biol 53(2):365–392
Singhvi A, Garriga G (2009) Asymmetric divisions, aggresomes and apoptosis. Trends Cell Biol 19(1):1–7
Sloane BF, Yan S, Podgorski I, Linebaugh BE, Cher ML, Mai J, Cavallo-Medved D, Sameni M, Dosescu J, Moin K (2005) Cathepsin B and tumor proteolysis: contribution of the tumor microenvironment. Semin Cancer Biol 15(2):149–157
Sloane BF, Sameni M, Podgorski I, Cavallo-Medved D, Moin K (2006) Functional imaging of tumor proteolysis. Annu Rev Pharmacol Toxicol 46:301–315
Smirnova IV, Ho GJ, Fenton JW II, Festoff BW (1994) Extravascular proteolysis and the nervous system: serine protease/serpin balance. Semin Thromb Hemost 20(4):426–432
Smith HW, Marshall CJ (2010) Regulation of cell signalling by uPAR. Nat Rev Mol Cell Biol 11(1):23–36
Sohail A, Sun Q, Zhao H, Bernardo MM, Cho JA, Fridman R (2008) MT4-(MMP17) and MT6-MMP (MMP25), a unique set of membrane-anchored matrix metalloproteinases: properties and expression in cancer. Cancer Metastasis Rev 27(2):289–302
Sol-Church K, Picerno GN, Stabley DL, Frenck J, Xing S, Bertenshaw GP, Mason RW (2002) Evolution of placentally expressed cathepsins. Biochem Biophys Res Commun 293(1):23–29
Song L, Rape M (2008) Reverse the curse–the role of deubiquitination in cell cycle control. Curr Opin Cell Biol 20(2):156–163
Song Z, Steller H (1999) Death by design: mechanism and control of apoptosis. Trends Cell Biol 9(12):M49–M52
Sorimachi H, Ishiura S, Suzuki K (1997) Structure and physiological function of calpains. Biochem J 328(Pt 3):721–732
Sorimachi H, Hata S, Ono Y (2010) Expanding members and roles of the calpain superfamily and their genetically modified animals. Exp Anim 59(5):549–566
Sounni NE, Noel A (2005) Membrane type-matrix metalloproteinases and tumor progression. Biochimie 87(3–4):329–342
Squier TC (2006) Redox modulation of cellular metabolism through targeted degradation of signaling proteins by the proteasome. Antioxid Redox Signal 8(1–2):217–228
Stadtmueller BM, Hill CP (2011) Proteasome activators. Mol Cell 41(1):8–19
Stamenkovic I (2003) Extracellular matrix remodelling: the role of matrix metalloproteinases. J Pathol 200(4):448–464
Steiner DF (1969) Proinsulin and the biosynthesis of insulin. N Engl J Med 280(20):1106–1113
Steiner DF (1998) The proprotein convertases. Curr Opin Chem Biol 2(1):31–39
Steiner H, Haass C (2000) Intramembrane proteolysis by presenilins. Nat Rev Mol Cell Biol 1(3):217–224
Steiner DF, Quinn PS, Chan SJ, Marsh J, Tager HS (1980) Processing mechanisms in the biosynthesis of proteins. Ann NY Acad Sci 343:1–16
Sterchi EE, Stocker W, Bond JS (2008) Meprins, membrane-bound and secreted astacin metalloproteinases. Mol Aspects Med 29(5):309–328
Sternlicht MD, Werb Z (2001) How matrix metalloproteinases regulate cell behavior. Annu Rev Cell Dev Biol 17:463–516
Stetler-Stevenson WG (2008) The tumor microenvironment: regulation by MMP-independent effects of tissue inhibitor of metalloproteinases-2. Cancer Metastasis Rev 27(1):57–66
Storr SJ, Carragher NO, Frame MC, Parr T, Martin SG (2011) The calpain system and cancer. Nat Rev Cancer 11(5):364–374
Strongin AY (2010) Proteolytic and non-proteolytic roles of membrane type-1 matrix metalloproteinase in malignancy. Biochim Biophys Acta 1803(1):133–141
Stubbs MT, Bode W (1994) Coagulation factors and their inhibitors. Curr Opin Struct Biol 4(6):823–832
Sun Y, Kucej M, Fan HY, Yu H, Sun QY, Zou H (2009) Separase is recruited to mitotic chromosomes to dissolve sister chromatid cohesion in a DNA-dependent manner. Cell 137(1):123–132
Szabo R, Bugge TH (2008) Type II transmembrane serine proteases in development and disease. Int J Biochem Cell Biol 40(6–7):1297–1316
Tai HC, Schuman EM (2008) Ubiquitin, the proteasome and protein degradation in neuronal function and dysfunction. Nat Rev Neurosci 9(11):826–838
Takahashi T, Tang J (1981) Cathepsin D from porcine and bovine spleen. Methods Enzymol 80(Pt C):565–581
Tallant C, Marrero A, Gomis-Ruth FX (2010) Matrix metalloproteinases: fold and function of their catalytic domains. Biochim Biophys Acta 1803(1):20–28
Tanaka Y, Tanaka R, Kawabata T, Noguchi Y, Himeno M (2000) Lysosomal cysteine protease, cathepsin B, is targeted to lysosomes by the mannose 6-phosphate-independent pathway in rat hepatocytes: site-specific phosphorylation in oligosaccharides of the proregion. J Biochem 128(1):39–48
Tatnell PJ, Fowler SD, Bur D, Lees WE, Kay J (1998) Cathepsin E. The best laid plans of mice and men. Adv Exp Med Biol 436:147–152
Tedelind S, Poliakova K, Valeta A, Hunegnaw R, Yemanaberhan EL, Heldin NE, Kurebayashi J, Weber E, Kopitar-Jerala N, Turk B, Bogyo M, Brix K (2010) Nuclear cysteine cathepsin variants in thyroid carcinoma cells. Biol Chem 391(8):923–935
Tedelind S, Jordans S, Resemann H, Blum G, Bogyo M, Fuhrer D, Brix K (2011) Cathepsin B trafficking in thyroid carcinoma cells. Thyroid Res 4(Suppl 1):S2
Tepel C, Bromme D, Herzog V, Brix K (2000) Cathepsin K in thyroid epithelial cells: sequence, localization and possible function in extracellular proteolysis of thyroglobulin. J Cell Sci 113(Pt 24):4487–4498
Thomas G (2002) Furin at the cutting edge: from protein traffic to embryogenesis and disease. Nat Rev Mol Cell Biol 3(10):753–766
Tooze J, Hollinshead M, Hensel G, Kern HF, Hoflack B (1991) Regulated secretion of mature cathepsin B from rat exocrine pancreatic cells. Eur J Cell Biol 56(2):187–200
Travis J (1988) Structure, function, and control of neutrophil proteinases. Am J Med 84(6A):37–42
Travis J, Fritz H (1991) Potential problems in designing elastase inhibitors for therapy. Am Rev Respir Dis 143(6):1412–1415
Travis J, Salvesen GS (1983) Human plasma proteinase inhibitors. Annu Rev Biochem 52:655–709
Trombetta ES, Mellman I (2005) Cell biology of antigen processing in vitro and in vivo. Annu Rev Immunol 23:975–1028
Tschopp J, Martinon F, Burns K (2003) NALPs: a novel protein family involved in inflammation. Nat Rev Mol Cell Biol 4(2):95–104
Tsukuba T, Okamoto K, Yasuda Y, Morikawa W, Nakanishi H, Yamamoto K (2000) New functional aspects of cathepsin D and cathepsin E. Mol Cells 10(6):601–611
Turk V, Bode W (1991) The cystatins: protein inhibitors of cysteine proteinases. FEBS Lett 285(2):213–219
Turk B, Turk V (2009) Lysosomes as “suicide bags” in cell death: myth or reality? J Biol Chem 284(33):21783–21787
Turk B, Turk V, Turk D (1997) Structural and functional aspects of papain-like cysteine proteinases and their protein inhibitors. Biol Chem 378(3–4):141–150
Turk B, Turk D, Turk V (2000) Lysosomal cysteine proteases: more than scavengers. Biochim Biophys Acta 1477(1–2):98–111
Turk V, Turk B, Turk D (2001) Lysosomal cysteine proteases: facts and opportunities. EMBO J 20(17):4629–4633
Turk D, Turk B, Turk V (2003) Papain-like lysosomal cysteine proteases and their inhibitors: drug discovery targets? Biochem Soc Symp 70:15–30
Turner AJ, Nalivaeva NN (2007) New insights into the roles of metalloproteinases in neurodegeneration and neuroprotection. Int Rev Neurobiol 82:113–135
Uddin MN, Nabi AH, Nakagawa T, Ichihara A, Inagami T, Suzuki F (2008) Non-proteolytic activation of prorenin: activation by (pro)renin receptor and its inhibition by a prorenin prosegment, “decoy peptide”. Front Biosci 13:745–753
Uhlmann F (2003) Separase regulation during mitosis. Biochem Soc Symp 70:243–251
Urban S (2010) Taking the plunge: integrating structural, enzymatic and computational insights into a unified model for membrane-immersed rhomboid proteolysis. Biochem J 425(3):501–512
Urban S, Freeman M (2002) Intramembrane proteolysis controls diverse signalling pathways throughout evolution. Curr Opin Genet Dev 12(5):512–518
Urban S, Lee JR, Freeman M (2001) Drosophila rhomboid-1 defines a family of putative intramembrane serine proteases. Cell 107(2):173–182
Van den Eynde BJ, Morel S (2001) Differential processing of class-I-restricted epitopes by the standard proteasome and the immunoproteasome. Curr Opin Immunol 13(2):147–153
van Goor H, Melenhorst WB, Turner AJ, Holgate ST (2009) Adamalysins in biology and disease. J Pathol 219(3):277–286
van Meer G, de Kroon AI (2011) Lipid map of the mammalian cell. J Cell Sci 124(Pt 1):5–8
Vandenabeele P, Galluzzi L, Vanden Berghe T, Kroemer G (2010) Molecular mechanisms of necroptosis: an ordered cellular explosion. Nat Rev Mol Cell Biol 11(10):700–714
Vanlangenakker N, Vanden Berghe T, Krysko DV, Festjens N, Vandenabeele P (2008) Molecular mechanisms and pathophysiology of necrotic cell death. Curr Mol Med 8(3):207–220
Varki A (1993) Biological roles of oligosaccharides: all of the theories are correct. Glycobiology 3(2):97–130
Varki A (1998) Factors controlling the glycosylation potential of the Golgi apparatus. Trends Cell Biol 8(1):34–40
Vasiljeva O, Reinheckel T, Peters C, Turk D, Turk V, Turk B (2007) Emerging roles of cysteine cathepsins in disease and their potential as drug targets. Curr Pharm Des 13(4):387–403
Victor BC, Sloane BF (2007) Cysteine cathepsin non-inhibitory binding partners: modulating intracellular trafficking and function. Biol Chem 388(11):1131–1140
Vierstra RD (2009) The ubiquitin-26S proteasome system at the nexus of plant biology. Nat Rev Mol Cell Biol 10(6):385–397
Villadangos JA, Bryant RA, Deussing J, Driessen C, Lennon-Dumenil AM, Riese RJ, Roth W, Saftig P, Shi GP, Chapman HA, Peters C, Ploegh HL (1999) Proteases involved in MHC class II antigen presentation. Immunol Rev 172:109–120
Vreemann A, Qu H, Mayer K, Andersen LB, Stefana MI, Wehner S, Lysson M, Farcas AM, Peters C, Reinheckel T, Kalff J, Brix K (2009) Cathepsin B release from rodent intestine mucosa due to mechanical injury results in extracellular matrix damage in early post-traumatic phases. Biol Chem 390(5–6):481–492
Watts C (2012) The endosome-lysosome pathway and information generation in the immune system. Biochim Biophys Acta 1824(1):14–21
Watts C, Matthews SP, Mazzeo D, Manoury B, Moss CX (2005) Asparaginyl endopeptidase: case history of a class II MHC compartment protease. Immunol Rev 207:218–228
Weihofen A, Martoglio B (2003) Intramembrane-cleaving proteases: controlled liberation of proteins and bioactive peptides. Trends Cell Biol 13(2):71–78
Whisstock JC, Bottomley SP (2006) Molecular gymnastics: serpin structure, folding and misfolding. Curr Opin Struct Biol 16(6):761–768
Whisstock JC, Bottomley SP (2008) Structural biology: serpins’ mystery solved. Nature 455(7217):1189–1190
White JM (2003) ADAMs: modulators of cell-cell and cell-matrix interactions. Curr Opin Cell Biol 15(5):598–606
Wiederanders B, Kaulmann G, Schilling K (2003) Functions of propeptide parts in cysteine proteases. Curr Protein Pept Sci 4(5):309–326
Wiertz EJ, Jones TR, Sun L, Bogyo M, Geuze HJ, Ploegh HL (1996a) The human cytomegalovirus US11 gene product dislocates MHC class I heavy chains from the endoplasmic reticulum to the cytosol. Cell 84(5):769–779
Wiertz EJ, Tortorella D, Bogyo M, Yu J, Mothes W, Jones TR, Rapoport TA, Ploegh HL (1996b) Sec61-mediated transfer of a membrane protein from the endoplasmic reticulum to the proteasome for destruction. Nature 384(6608):432–438
Wojcik C, DeMartino GN (2003) Intracellular localization of proteasomes. Int J Biochem Cell Biol 35(5):579–589
Wolf DH, Hilt W (2004) The proteasome: a proteolytic nanomachine of cell regulation and waste disposal. Biochim Biophys Acta 1695(1–3):19–31
Wolfe MS, Kopan R (2004) Intramembrane proteolysis: theme and variations. Science 305(5687):1119–1123
Woodward JK, Holen I, Coleman RE, Buttle DJ (2007) The roles of proteolytic enzymes in the development of tumour-induced bone disease in breast and prostate cancer. Bone 41(6):912–927
Wyllie AH, Kerr JF, Currie AR (1980) Cell death: the significance of apoptosis. Int Rev Cytol 68:251–306
Yamamoto K (1995) Cathepsin E and cathepsin D: biosynthesis, processing and subcellular location. Adv Exp Med Biol 362:223–229
Yanagida M (2009) Clearing the way for mitosis: is cohesin a target? Nat Rev Mol Cell Biol 10(7):489–496
Yao C (2010) Major surface protease of trypanosomatids: one size fits all? Infect Immun 78(1):22–31
Yu DM, Wang XM, McCaughan GW, Gorrell MD (2006) Extraenzymatic functions of the dipeptidyl peptidase IV-related proteins DP8 and DP9 in cell adhesion, migration and apoptosis. FEBS J 273(11):2447–2460
Yu DM, Yao TW, Chowdhury S, Nadvi NA, Osborne B, Church WB, McCaughan GW, Gorrell MD (2010) The dipeptidyl peptidase IV family in cancer and cell biology. FEBS J 277(5):1126–1144
Zaidi N, Hermann C, Herrmann T, Kalbacher H (2008) Emerging functional roles of cathepsin E. Biochem Biophys Res Commun 377(2):327–330
Zhao JH, Yang CT, Wu JW, Tsai WB, Lin HY, Fang HW, Ho Y, Liu HL (2008) RING domains functioning as E3 ligases reveal distinct structural features: a molecular dynamics simulation study. J Biomol Struct Dyn 26(1):65–74
Zhou A, Webb G, Zhu X, Steiner DF (1999) Proteolytic processing in the secretory pathway. J Biol Chem 274(30):20745–20748
Zhou A, Carrell RW, Murphy MP, Wei Z, Yan Y, Stanley PL, Stein PE, Broughton Pipkin F, Read RJ (2010) A redox switch in angiotensinogen modulates angiotensin release. Nature 468(7320):108–111
Zogg T, Brandstetter H (2011) Complex assemblies of factors IX and X regulate the initiation, maintenance, and shutdown of blood coagulation. Prog Mol Biol Transl Sci 99:51–103
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Wien
About this chapter
Cite this chapter
Brix, K., Scott, C.J., Heck, M.M.S. (2013). Compartmentalization of Proteolysis. In: Brix, K., Stöcker, W. (eds) Proteases: Structure and Function. Springer, Vienna. https://doi.org/10.1007/978-3-7091-0885-7_3
Download citation
DOI: https://doi.org/10.1007/978-3-7091-0885-7_3
Published:
Publisher Name: Springer, Vienna
Print ISBN: 978-3-7091-0884-0
Online ISBN: 978-3-7091-0885-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)