Abstract
Plant genomes contain many duplicated genes, some of which were produced by recursive polyploidizations. These duplicated genes may evolve interactively and even concertedly through homoeologous recombination. Here, we explored likely gene conversion in Brassica rapa and Brassica oleracea. By checking gene colinearity, we detected 4296 duplicated genes existing in both the species, which were produced by whole-genome triplication from their common ancestor. Incongruities of homologous gene tree topologies indicated that 8 % of these duplicated genes were converted by one another after the divergence of B. rapa and B. oleracea. These converted genes are more often from larger duplicated chromosomal blocks, indicating that illegitimate recombination is more likely to occur between larger homoeologous chromosomal regions. This research contributed to understanding genome stability and gene evolution after polyploidization.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
11.1 Introduction
Plant genomes have been widely affected by recursive polyploidizations, which repeatedly double or triple the genome information in a cell over-night (Bowers et al. 2003, 2005; Jaillon et al. 2007; Soltis et al. 2008; Soltis and Soltis 2009; Abrouk et al. 2010; Tang et al. 2010; Jiao et al. 2011, 2012). Though wide-spread gene losses and DNA rearrangements often follow, mostly leading to restoration of diploid heredity, hundreds of duplicated genes are often preserved in colinearity on homoeologous chromosomes or chromosomal segments, retaining valuable traces of these abrupt evolutionary events (Wang et al. 2005; Gaeta et al. 2007; Paterson 2008; Paterson et al. 2009, 2012; Proost et al. 2011; Schnable et al. 2011; Freeling et al. 2012).
Recent research into illegitimate recombination between duplicated genes revealed that many duplicated genes might have been affected by gene conversion, with one copy of a pair of duplicates being converted to the DNA sequence by the other by a unidirectional recombination-like mechanism (Xu et al. 2008; Gaeta and Chris Pires 2009; Wang and Paterson 2011). A comparative analysis between rice and sorghum genomes showed that 12 % of rice duplicated genes and 14 % of sorghum duplicated genes were affected by conversion after the divergence of these lineages (Wang et al. 2007, 2009). Among those converted genes, 40 % were affected to their full gene length and the others in only partial sequence. These conversion events may have occurred tens of millions years ago.
A comparison between rice subspecies indica and japonica found evidence of more recent gene conversion, showing that ~8 % of rice genes may have been converted after the split of the two subspecies about 400,000 years ago (Zhu et al. 2007). One pair of grass chromosomes, e.g., rice chromosomes 11 and 12, their sorghum orthologous chromosomes 5 and 8, and corresponding chromosomes in other grasses, have been affected by prominent conversion (Wang et al. 2011a, b, c). After the split of rice and sorghum, nearly 60 % of rice and sorghum duplicated genes have been converted by their duplicated copies. Evidence from sequence similarity analysis, and independent analysis of Oryza species (including rice) indicated that near the termini of the short arms of Oryza chromosomes 11 and 12, gene conversion may be still ongoing, 70 million years or after the origination of these duplicated genes (Jacquemin et al. 2009; Wang et al. 2011a, b, c).
Analysis of eudicot genomes found more evidence of homo(eo)logous gene conversion. In a tetraploid cotton, Acala Maxxa, 40 % of paralogous genes from its two subgenomes At and Dt differ in sequence from their diploid progenitors. The vast majority of these mutations are convergent, with At genes converted to the Dt state at more than twice the rate (25 %) as the reciprocal (10.6 %) (Paterson et al. 2012). As to conversion between homologous chromosomes, sequencing 40 Arabidopsis F2 plants and their parents showed that small gene conversion tracts, often biased, represented over 90–99 % of all recombination events. Moreover, the rate of alteration of protein sequence caused by gene conversion is reported to be more than 600-times that caused by mutation (Yang et al. 2012).
11.2 Comparative Inference of Gene Conversion in B. rapa and B. oleracea
The existence of large homoeologous blocks provides a chance for homoeologous (ectopic) DNA recombination, which may result in concerted evolution of duplicated genes as inferred previously in grasses (Wang et al. 2009, 2011a, b, c).
11.2.1 Rationale to Infer Gene Conversion
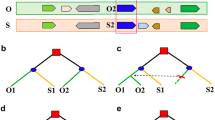
Annotated genes from Brassica rapa and Brassica oleracea were from sequencing project websites (Wang et al. 2011a, b, c; Liu et al. 2014). To find colinear homologs within a plant or between two plants, we run BLASTP to find homologous genes. Homologs with E-values smaller than 1e-10 were taken as input for ColinearScan, which was adopted to infer DNA blocks containing 10 or more colinear genes. By checking chromosome numbers, it was not difficult to define orthologs between B. rapa and B. oleracea. By using an approach described previously, we defined three subgenomes A, B, and C, and found paralogs in each plant. If there was no gene loss, at corresponding locations there would be three colinear genes in each plant produced by the genome triplication, namely, Br-A, Br-B, and Br-C in B. rapa, and their respective orthologs, Bo-A, Bo-B, and Bo-C in B. oleracea, forming homologous gene sextet. However, due to wide-spread gene loss after the genome triplication, often we could not find sextets of homoeologs.
To infer gene conversion, based on sextets or incomplete groups, we defined homologous gene quartets, two paralogs in a plant and their respective orthologs in the other plant. Then we inferred synonymous nucleotide substitution rates (Ks) between them. We anticipated that orthologs were more similar than paralogs, in that speciation was after genome triplication. However, if paralogs in a genotype were more similar than orthologs, we considered that the paralogs might have been affected by gene conversion. Bootstrapping tests were repeated 100 times. To estimate Ks, we first aligned proteins of a homologous quartet with CLUSTALW, and after removing gaps, the protein alignment was then translated into cDNA alignment in codons. Ks were estimated by using the Nei-Gojobori approach implemented by BioPerl.
11.2.2 Characterization of Gene Conversion
By using ColinearScan to find gene colinearity and by checking sequence similarity between chromosomal regions, we inferred paralogous genes within B. rapa and B. oleracea genomes, respectively, and inferred orthologous genes between them. Here, we checked triplicated genes that were preserved in both Brassica species, which form homologous gene sextets. For genes in each sextet, we checked each quartet of homologs within them. We compared gene similarity or tree topology. We anticipated that the paralogs (duplicated genes) were more diverged than their respective orthologs. If not, we inferred that the paralogs might have been affected by gene conversion. We removed possible redundancy when counting converted gene pairs.
We detected 4296 homoeologous pairs of genes, involving 8592 (20.6 % of) B. rapa genes and 8592 (24.7 % of) B. oleracea genes. Most of these reside in 23 large duplicated blocks in B. rapa (Fig. 11.1a) and 19 large duplicated blocks in B. oleracea (Fig. 11.1b), distributed throughout the chromosomes. In total, we found that ~8 % of duplicates (368 and 343) in B. rapa and B. oleracea have been affected by gene conversion (Table 11.1). The conversion tracts vary in size, ranging from a few base pairs to full gene lengths.
11.2.3 Unbalanced Gene Conversion Among Chromosomes
Different chromosomes have been unequally affected by gene conversion (Fig. 11.1c, d). In B. rapa, the most affected chromosomes are Br01, Br04, and Br05, with >10 % of paralogs affected, whereas in B. oleracea, the most affected chromosomes are Bo01 and Bo06, with >10 % of paralogs affected. In contrast, no paralogous pair from between Br09 and Br01, Bo08 or Bo09, Bo04 and Bo09 has been affected. Genes residing in bigger chromosomes with more colinear homoeologs are more likely to be affected by conversion (Fig. 11.2). This means larger duplicated regions on these chromosomes may facilitate the occurrence of homoeologous recombination due to preserving more DNA homology.
11.2.4 Gene Conversion Occurs Correspondingly in Two Brassica Species
Gene conversion often occurs in both Brassica species in a corresponding manner, that is, if a duplicated gene pair were affected by gene conversion in one species, so were their counterparts in the other species. Most homoeologous quartets (~92 %) were found to be converted in both species. Only 53, or about one-sixth, of homoeologous gene quartets showed evidence of independent concerted evolution, i.e. were inferred to have experienced independent conversion events in B. rapa or B. oleracea. That is, it is likely that 5/6 the events are likely to have occurred shortly after the triplication but before the lineages diverged, or co-occurred independently in each lineage.
11.2.5 Biased Gene Conversion Among Different Subgenomes
Previous publication (Wang et al. 2011a, b, c) revealed three subgenomes that formed the present genomes of B. rapa and B. oleracea, and here we characterize gene conversion between different subgenomes. As to the analysis, there is an occurrence bias of gene conversion among subgenomes. About 40–44 % of conversion events involved paralogs on subgenomes A and B in both species, substantially more than between other subgenome combinations (Table 11.2). However, this increase parallels gene numbers in the respective subgenomes, with the percentages of converted paralogs from any two subgenomes being similar. This suggests that gene conversion is related to homologous gene density, which determines the likelihood of illegitimate recombination to occur.
11.3 Gene Conversion and Genome Stability
11.3.1 Increased Genome Stability and Complexity After Polyploidization
Wide-spread and recursive polyploidizations have affected all flowering and seed plants, and may have been an important driving force of their evolution, especially the likely rapid divergence and speciation of lineages to form large groups of related species (Bowers et al. 2003; Ziolkowski et al. 2006; Soltis and Soltis 2009; Jiao et al. 2012). This should be a direct result of genome instability after whole-genome duplication/triplication (Marfil et al. 2006; Mazowita et al. 2006). These large-scale genome addition events duplicated/triplicated DNA content overnight, adding much genomic complexity and increasing interactions between chromosomes. Such interactions may include physically by DNA binding, knotting, splitting and breaking; and genetically by pairing, clustering, recombining, and segregating. A drive to recover diploid heredity may be the paramount source of force. Anyway, the majority of land plants favors diploid heredity and are adapted to finish a cycle of meiosis each year. Increased complexity will lead a lot of outcomes genetically, and the first among them is genomic instability.
Genomic instability is often accompanied by wide-spread gene losses, chromosomal rearrangement, and recombination between homo(eo)logous chromosomes or chromosomal segments (Wang et al. 2005; Feldman et al. 2012). If a polyploid came to recover diploid heredity, with one-to-one pairing of homologous chromosomes rather than pairing among multiple homo(eo)logous chromosomes, it may eventually regain much of its genomic stability. However, small scale chromosomal rearrangement may still continue to occur. The analysis of grass genomes indicated that the majority of genomic changes occurred before the divergence of major grass clades. For example, after the divergence of rice and sorghum, only ~2–3 % of genes were lost, resulting in minimal erosion of gene colinearity along orthologous chromosomes, in contrast to the loss of at least 65 % of genes duplicated in their common ancestor (Wang et al. 2005; Paterson et al. 2009). For another example, the majority of chromosomal rearrangement occurred before their divergence, and only a few such rearrangements can be identified in the sorghum lineage after its split with rice (Murat et al. 2010).
11.3.2 Homoeologous Recombination Is a Driving Force for Genomic Evolution
Homoeologous recombination is also a phenomenon of genomic instability, and can last much longer than other changes discussed above. As a result of this kind of illegitimate recombination, gene conversion transfers genetic information in a unidirectional manner. As gene conversion mechanisms proposed, it would increase DNA substitution rates, and therefore may play a role as a driving force of evolution (Chen et al. 2007; Wang and Paterson 2011). This has been attested to by comparative analysis of grass genes (Wang et al. 2009; Wang and Paterson 2011). After the ease of major genomic changes, homoeologous recombination and gene conversion can still occur millions of years after ancestral polyploidization (Wang et al. 2011a, b, c; Paterson et al. 2012). This has been evidenced from the analysis of both monocot to dicot plants. A particularly striking finding involves genes at the very end of rice chromosomes 11 and 12 and their counterparts in other grasses (Wang et al. 2007; Jacquemin et al. 2009; Paterson et al. 2012).
11.3.3 Gene Conversion and Homoeologous Block Length
Here, we revealed a correlation of longer lengths of duplicated blocks (or larger numbers of genes) with higher conversion rates, which agrees with previous findings in grasses. More colinear genes often mean higher DNA similarity between duplicated regions, which would increase the likelihood of homoeologous pairing. The chance of pairing is definitely much less between homoeologous than homologous chromosomes. Once it occurs, it would have some genetic outcomes, such as relatively low-level genomic instability, DNA mutations, and conversion.
11.4 Conclusion
Here, by performing comparative genomic analysis, we characterized gene conversion in B. rapa and B. oleracea. Gene conversion as a result of homoeologous recombination is a long lasting driving force of plant evolution. Widespread and recursive polyploidizations have played a pivotal role in the evolution, divergence and speciation of land plants. After the ease of genome shock (McClintock 1984) often in the early days after polyploidization, characterized by wide-spread gene losses and chromosomal rearrangements, genomes may recover much stability and return to diploid heredity. Though occurring at lower levels in later stages than early days after polyploidization, homoeologous recombination and gene conversion may last for a very long time, continuing to play a driving force in genomic evolution and genetic innovation.
References
Abrouk M, Murat F, Pont C, Messing J, Jackson S et al (2010) Palaeogenomics of plants: synteny-based modelling of extinct ancestors. Trends Plant Sci 15:479–487
Bowers JE, Chapman BA, Rong J, Paterson AH (2003) Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 422:433–438
Bowers JE, Arias MA, Asher R, Avise JA, Ball RT, et al (2005) Comparative physical mapping links conservation of microsynteny to chromosome structure and recombination in grasses. Proc Natl Acad Sci USA 102:13206–13211
Chen JM, Cooper DN, Chuzhanova N, Ferec C, Patrinos GP (2007) Gene conversion: mechanisms, evolution and human disease. Nat Rev Genet 8:762–775
Feldman M, Levy AA, Fahima T, Korol A (2012) Genomic asymmetry in allopolyploid plants: wheat as a model. J Exp Bot 63:5045–5059
Freeling M, Woodhouse MR, Subramaniam S, Turco G, Lisch D, Schnable JC (2012) Fractionation mutagenesis and similar consequences of mechanisms removing dispensable or less-expressed DNA in plants. Curr Opin Plant Biol 15:131–139
Gaeta RT, Chris Pires J (2009) Homoeologous recombination in allopolyploids: the polyploid ratchet. New Phytol 186:18–28
Gaeta RT, Pires JC, Iniguez-Luy F, Leon E, Osborn TC (2007) Genomic changes in resynthesized Brassica napus and their effect on gene expression and phenotype. Plant Cell 19:3403–3417
Jacquemin J, Laudie M, Cooke R (2009) A recent duplication revisited: phylogenetic analysis reveals an ancestral duplication highly-conserved throughout the Oryza genus and beyond. BMC Plant Biol 9:146
Jaillon O, Aury JM, Noel B, Policriti A, Clepet C et al (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–467
Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, et al (2011) Ancestral polyploidy in seed plants and angiosperms. Nature 473:97–100
Jiao Y, Leebens-Mack J, Ayyampalayam S, Bowers JE, McKain MR et al (2012) A genome triplication associated with early diversification of the core eudicots. Genome Biol 13:R3
Liu S, Liu Y, Yang X, Tong C, Edwards D, et al (2014) The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat Commun 5:3930
Marfil CF, Masuelli RW, Davison J, Comai L (2006) Genomic instability in Solanum tuberosum × Solanum kurtzianum interspecific hybrids. Genome 49:104–113
Mazowita M, Haque L, Sankoff D (2006) Stability of rearrangement measures in the comparison of genome sequences. J Comput Biol 13:554–566
McClintock B (1984) The significance of responses of the genome to challenge. Science 226:792–801
Murat F, Xu JH, Tannier E, Abrouk M, Guilhot N et al (2010) Ancestral grass karyotype reconstruction unravels new mechanisms of genome shuffling as a source of plant evolution. Genome Res 20:1545–1557
Paterson AH (2008) Paleopolyploidy and its impact on the structure and function of modern plant genomes. Genome Dyn 4:1–12
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, et al (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457:551–556
Paterson AH, Wendel JF, Gundlach H, Guo H, Jenkins J, et al (2012) Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492:423–427
Proost S, Pattyn P, Gerats T, Van de Peer Y (2011) Journey through the past: 150 million years of plant genome evolution. Plant J 66:58–65
Schnable JC, Springer NM, Freeling M (2011) Differentiation of the maize subgenomes by genome dominance and both ancient and ongoing gene loss. Proc Natl Acad Sci USA 108:4069–4074
Soltis PS, Soltis DE (2009) The role of hybridization in plant speciation. Annu Rev Plant Biol 60:561–588
Soltis DE, Bell CD, Kim S, Soltis PS (2008) Origin and early evolution of angiosperms. Ann N Y Acad Sci 1133:3–25
Tang H, Bowers JE, Wang X, Paterson AH (2010) Angiosperm genome comparisons reveal early polyploidy in the monocot lineage. Proc Natl Acad Sci USA 107:472–477
Wang XY, Paterson AH (2011) Gene conversion in angiosperm genomes with an emphasis on genes duplicated by polyploidization. Genes (Basel) 2:1–20
Wang X, Shi X, Hao B, Ge S, Luo J (2005) Duplication and DNA segmental loss in the rice genome: implications for diploidization. New Phytol 165:937–946
Wang X, Tang H, Bowers JE, Feltus FA, Paterson AH (2007) Extensive concerted evolution of rice paralogs and the road to regaining independence. Genetics 177:1753–1763
Wang X, Tang H, Bowers JE, Paterson AH (2009) Comparative inference of illegitimate recombination between rice and sorghum duplicated genes produced by polyploidization. Genome Res 19:1026–1032
Wang X, Tang H, Paterson AH (2011a) Seventy million years of concerted evolution of a homoeologous chromosome pair, in parallel, in major Poaceae lineages. Plant Cell 23:27–37
Wang X, Wang H, Wang J, Sun R, Wu J et al (2011b) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43:1035–1039
Wang XY, Tang HB, Paterson AH (2011c) Seventy million years of concerted evolution of a homoeologous chromosome pair, in parallel, in major poaceae lineages. Plant Cell 23:27–37
Xu S, Clark T, Zheng H, Vang S, Li R et al (2008) Gene conversion in the rice genome. BMC Genom 9:93
Yang S, Yuan Y, Wang L, Li J, Wang W, et al (2012) Great majority of recombination events in Arabidopsis are gene conversion events. Proc Natl Acad Sci USA 109:20992–20997
Zhu Q, Zheng X, Luo J, Gaut BS, Ge S (2007) Multilocus analysis of nucleotide variation of Oryza sativa and its wild relatives: severe bottleneck during domestication of rice. Mol Biol Evol 24:875–888
Ziolkowski PA, Kaczmarek M, Babula D, Sadowski J (2006) Genome evolution in Arabidopsis/Brassica: conservation and divergence of ancient rearranged segments and their breakpoints. Plant J 47:63–74
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Wang, J., Guo, H., Jin, D., Wang, X., Paterson, A.H. (2015). Comparative Analysis of Gene Conversion Between Duplicated Regions in Brassica rapa and B. oleracea Genomes. In: Wang, X., Kole, C. (eds) The Brassica rapa Genome. Compendium of Plant Genomes. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-662-47901-8_11
Download citation
DOI: https://doi.org/10.1007/978-3-662-47901-8_11
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-47900-1
Online ISBN: 978-3-662-47901-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)