Abstract
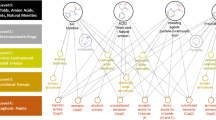
The computerised design of potentially bioactive molecules or the computer recognition of a new biological property of an existing molecule can lead to orders of magnitude more suggestions than are reasonable to explore experimentally. Thus we added to the ALADDIN Control Language new commands that describe the transformation of one chemical structure into another. Atoms or groups can be removed or replaced and bonds can be made, broken, or changed in bond order. We used this language to describe transformations that remove extraneous substituents from molecules suggested for synthesis as potential dopaminergic agonists1 and demonstrated that such transformations reduce the number of compounds to be investigated by 66–85% without loss of geometric information. Similarity such transformations lead to the identification of the structural family to which an existing compound belongs.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Martin, Y.C. ‘Computer Design of Potentially Bioactive Molecules by Geometric Searching with ALADDIN’. Tetrahedron Comput. Methodol 1990, 3, 15–25.
Martin, Y.C.; Bures, M.G.; Willett, P. ‘Searching Databases of Three-dimensional Structures’. In Reviews in Computational Chemistry; Boyd, D.B.; Lipkowitz, K., Eds; VCH: New York, 1990; Vol. 1, pp. 213–263.
Van Drie, J.H.; Weininger, D.; Martin, Y.C. ‘ALADDIN: An Integrated Tool for Computer-assisted Molecular Design and Pharmacophore Recognition from Geometric, Steric, and Substructure Searching of Three-dimensional Molecular Structures’. J. Comput.-Aided Mol. Des. 1989, 3, 225–251.
Bures, M.G.; Black-Schaefer, C.; Gardner, G. ‘The Discovery of Novel Auxin Transport Inhibitors by Molecular Modelling and Three-dimensional Pattern Analysis’. J. Comput.- Aided Mol. Des. 1991, 5, 323–334.
Martin, Y.C.; Danaher, E.A. ‘Molecular Modelling of Receptor-Ligand Interactions’. In Receptor Pharmacology and Function; Williams, M.; Glennon, R.A.; Timmermans, P.B.M.W.M., Eds; Marcel Dekker: New York, 1989; pp. 137–171.
Rusinko III, A.; Skell, J.M.; Balducci, R; McGarity, C.M.; Pearlman, R.S. ‘CONCORD, A Program for the Rapid Generation of High Quality Approximate 3-Dimensional Molecular Structures’. Tripos Associates, St. Louis, Missouri, 1988.
Anon. Daylight Chemical Information Systems, Irvine, California.
Hansch, C.; Leo, A. The 1989 edition of the Pomoma College Medicinal Chemistry Project database of measured octanol-water partition coefficients and pKa’s distributed by Daylight Chemical Information Systems, Irvine, California.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 1993 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Martin, Y.C., van Drie, J.H. (1993). Identifying Unique Core Molecules from the Output of a 3-D Database Search. In: Warr, W.A. (eds) Chemical Structures 2. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-78027-1_28
Download citation
DOI: https://doi.org/10.1007/978-3-642-78027-1_28
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-78029-5
Online ISBN: 978-3-642-78027-1
eBook Packages: Springer Book Archive