Abstract
Analysis of medical images involves robust computational approaches to various optimization problems prior to interpretation of the embedded pathological changes. Major computational efforts are essential in unsupervised learning of structures in various medical images. Some unsupervised learning techniques that take advantage of information theory concepts provide a different perspective on the solution of automated learning methods. This chapter will review a recent approach to clustering examined under an information theoretic framework that efficiently finds a suitable number of clusters representing different tissue characteristics in a medical image. The proposed clustering approach optimizes an objective function which quantifies the quality of particular cluster configurations. Recent developments involving interesting relationships between spectral clustering (SC) and kernel principal component analysis (kPCA) are used in this technique to include the nonlinear domain. In this novel SC approach, the data is mapped to a new space where the points belonging to the same cluster are collinear if the parameters of a radial basis function (RBF) kernel are adequately selected. The effectiveness of this nonlinear approach is demonstrated in the segmentation of uterine cervix color images for early identification of cervical neoplasia, as an aid to cervical cancer diagnosis. The limitations of this method in the segmentation of specific medical images such as brain images with multiple sclerosis lesions and a strategy to overcome them are discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Cervical Cancer
- Cervical Intraepithelial Neoplasia
- Unsupervised Learning
- Spectral Cluster
- Multiple Sclerosis Lesion
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
The discovery of underlying structures in medical images in a robust and efficient manner is of crucial significance in many applications, such as the diagnostic classification of precancerous uterine cervix lesions and the identification of anomalies in magnetic resonance image (MRI) brain structures. Often, with medical images, an adequate ground truth is not available, and unsupervised learning techniques must be used. One of the fundamental questions in the identification process is the number of distinguishable clusters or segments that comprise the image data. The machine learning methods that address these problems form a highly interdisciplinary field, where a non-interacting machine can be developed to learn in either a supervised or unsupervised manner.
With supervised learning, the system is “taught” by providing a labeled training set, which is a sequence of desired output y = {y 1, y 2, …}. The goal of the machine is to learn to produce the correct output (or class label in a classification context) given a new input. Although such a system may be quite fast after training, it can be limited by a subjective bias that depends on the training set, i.e. it may perform poorly when presented with a data pattern that is considerably different from the patterns encountered during training. Furthermore, some applications cannot provide a universally accepted “ground truth” to label training data in the first place. In unsupervised learning, the machine only receives the input data x = {x 1, x 2, …} but obtains neither supervised target outputs nor rewards from its environment. It may seem mysterious that a system can learn to recognize patterns without being informed of the features it should analyze or even the correct number of clusters. With unsupervised learning, the machine is not explicitly taught; instead, its goal is to build representations of the input that can be used for decision making. Instead of relying upon a correctly labeled training set, the machine learns typically via a general objective function or a set of update rules. In one sense, unsupervised learning can be thought of as discovering patterns that are more significant than noise. Hence, a standard example of unsupervised learning is clustering, which becomes segmentation when applied to image data [1].
The two competing formalisms of machine learning methods are discriminative and generative approaches. Generative or informative approaches produce a probability density model over all variables to compute classification in the manner of parametric clustering. In parametric clustering, a predefined distribution for the dataset is assumed, and then sufficient statistics or parameters are calculated. For example, when assuming a multidimensional normal distribution, the sufficient statistics are the mean and the covariance matrix [2]. Furthermore, some information about the statistical dependencies of the data can be gained through unsupervised learning [3, 4]. Another approach is discriminative learning, which lends itself to nonparametric clustering. Discriminative approaches provide a direct attempt to compute the input–output mappings for classification, while eschewing any direct modeling of the underlying distributions. Nonparametric clustering divides the data into groups of points that possess strong internal similarities. A criterion function is used to measure these similarities and to seek the grouping that optimizes the criterion [2] (Jebara 2004).
Another approach to unsupervised learning uses an information theoretic approach, which in a sense transcends both the generative and discriminative approaches. For example, from Table 1 both the maximum mutual information and maximum entropy discrimination approaches to learning are connected to information theory. Information theory addresses the trade-off between memory storage cost and accuracy. Almost all work in unsupervised learning can be viewed in terms of learning a probabilistic model of the data. The machine can build a model of the data which assumes that the data points are independently drawn from some distribution P(x). In general, the true distribution of the data is unknown, but we can learn a model of this distribution, say Q(x). Information theory is used to estimate the accuracy of such a model at a given memory storage cost. Such a model can be used for classification. Furthermore, an information theoretic measure can be used as a similarity criterion for clustering. This approach is appealing, because it can capture descriptions beyond second order (variance) statistics. The major problem of information theoretic based clustering has been the difficulty to evaluate the metric without imposing unrealistic assumptions on the data distributions. Successful implementations carefully estimate the information theoretic measure [2].
In this information theoretic approach to medical image analysis, where the number of significant segments in an image is unknown a priori, we apply a method of information theoretic unsupervised learning. A simple information theoretic approach to clustering, proposed by Sugar and James [5], is called the “jump” method as explained below. The “jump” method performs well for high dimensional Gaussian data, but it behaves unreliably for clusters corresponding to natural image segments. To make this approach more practical, Corona developed an improved “jump” method (IJM) [6]. IJM performs well with synthetic data and has been validated with standard clustering datasets as well as with medical images with known ground truths. In this chapter, we will briefly introduce the original jump method and improvements to address some of its drawbacks. The applications of the IJM will be described in segmentation of medical images. The potential of the IJM in compression and 3-D visualization of medical images will also be addressed.
2 The “Jump” Method
Suppose one has p-dimensional data with n points to be clustered, but the exact number of clusters K is unknown a priori. First, a method is needed to quantify the quality of the clustering given the number of clusters, that is, a measure of the inter-cluster dispersion. One such measure is the minimal achievable distortion, d K . This distortion is defined as the mean squared Mahalanobis distance of every point in the cluster to the nearest cluster’s centroid (or cluster representative), i.e.
where x i is the ith data point that belongs to the kth cluster, c k is the centroid or representative point of the kth cluster, Σ −1 is the inverse covariance matrix for each cluster (assumed to be approximately equal for all clusters), and K is the number of clusters considered. One can create a monotonically decreasing curve by gradually increasing the value of K, and re-estimating Eq. (1) at each step; that is, consider the sequence d K = {d K=1, d K=2,…, d K=K max}. This curve contains important information that can be used to estimate a suitable number of representatives for the data.
Sugar and James developed a simple information theoretic approach where a classic clustering algorithm (e.g., k-means with Σ = I) is used to generate a curve of the number of clusters, K, versus an estimate of the associated distortion, d K [5]. What they observed is that there tends to be a discontinuous “jump” in this curve at the appropriate number of clusters K (see Figs. 1 and 3). The jump operator can be defined as
where Y > 0 is a suitable power transformation acting on d K [5]. It can be theoretically shown that for Gaussian clusters sharing the same covariance matrix, the distortion curve becomes linear for Y = p/2, if the dataset dimensionality p is large enough. Furthermore, if one defines d − Y K = 0 = 0, the detection of the absence of clustering structure (i.e., K = 1) is feasible. Finally, the number of clusters where the largest jump occurs provides the estimate of the number of clusters in the dataset, i.e.
Finally, it should be noted that this “jump” method approach is hierarchical. That is, the relative height arrangement of the jump operator reflects more likely cluster configurations. This means that a proper sub-clustering may show up as a distinct second highest jump operator peak (see Fig. 2).
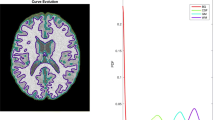
The “jump” method, with linear behavior of the transformed d K curve for a mixture of five spherical Gaussians. The box on the left shows the mixture of Gaussian components. The middle plot exhibits the estimated d K curve. The rightmost plot illustrated the transformed distortion curve of d − Y K with Y = p/2 with a fitted line for K > K *. The jump operator calculates the difference of successive values on the rightmost plot, thus converting the problem into finding a maximum [7]
The hierarchical aspect of the “jump” method. (a) The raw data generated from nine three-dimensional Gaussians that form three super-clusters. (b) The raw distortion curve. (c) The transformed distortion curve with Y = p/2 = 3/2. (d) The jump statistic, defined as the first forward difference of the transformed distortion. Note that the most obvious configuration is the three super-clusters, with the nine individual clusters at a close second in terms of the jump operator [6]
3 The Improved “Jump” Method
3.1 The Major Drawbacks of the “Jump” Method
The major drawbacks of the “jump” method are rooted in the fixing of the value Y = p/2 by applying the restrictive assumptions:
-
Because the k-means algorithm performance is extremely dependent on the choice of the initial cluster centers, the accuracy of the distortion curve may be compromised by a poor initialization; i.e., the standard k-means implementation can get “trapped” in local optima, which yields a poor estimation of d K using k-means and results in very noisy or inaccurate “jump” selections.
-
The method assumes high dimensionality (p → ∞), accurate d K values, well balanced clusters, identity covariance matrix, sufficiently large distance between centroids, etc., and the power transformation can only be determined under these assumptions for underlying Gaussian distributions. Real-world data easily violate these assumptions.
-
If strong correlations are present among variables, or if the clusters have intriguing or complicated shapes, linear algorithms such as k-means will not capture the data essence [6].
3.2 The Improvements to the “Jump” Method
The IJM works by first fixing the cluster number K and then searching for its best associated transformation, finding Y = p eff/2 for each K, where p eff is called the effective dimension:
-
Spectral clustering (SC) and probabilistic initialization techniques can be used to alleviate the suboptimal convergence of the k-means algorithm.
-
Handling realistic data necessitates that the power transform be adaptively modified according to the data at hand. One can make use of the hierarchical property of the “jump” method to create a criterion, the margin, which measures the quality of each of the K partitions. This approach allows the measurement of the operator performance in terms of linearity and the relative “jump” peak height. By maximizing this new objective function, one obtains a suitable power transformation, Y, which ensures a more robust solution to the number of clusters in a dataset.
-
Find a suitable kernel method to map the original dataset onto a “better” space where the mapped data is well behaved, such that the “jump” method can be used there effectively.
3.3 The Margin Operator of the IJM
The original “jump” method possesses a hierarchical nature, in the sense that the relative heights of the peaks of the jump operator J Y K indicate the degree of validity of the solution (or a valid sub-clustering). With this in mind, one can further define another operator called the margin operator,
For a fixed Y, the margin operator M Y K attempts to quantify the quality of a configuration with K clusters with respect to its closest competitor. Therefore, the larger the value of the margin operator becomes (for any positive Y), the better the cluster configuration is. Thus, by maximizing the margin value with respect to Y, as in
it becomes possible to quantify the quality of a clustering configuration with K partitions in terms of cluster compactness and the degree of decorrelation of its variables. The higher the margin, the better the configuration is with respect to the others. Hence, the margin measures the quality of the configuration. The maximum jump peaks must have margin values greater than unity (see Fig. 3).
After solving the eigenproblem posed by the spectral clustering (SC) algorithm for a number of training points, the cluster indicator vectors (i.e., eigenvectors) and the extended kernel matrix are used to expand the solution to the rest of the points [8]. To find the labels for K clusters, one performs kernel principal component analysis (kPCA), solving for at least the K − 1 eigenvectors associated with the highest eigenvalues. The ith entries of these eigenvectors represent the components of the ith point in a new K − 1 dimensional space. In principle, when using a properly parameterized radial basis function (RBF) kernel, the points in the same cluster become collinear in this new space. Thus, by projecting all points onto the unit hyper sphere, one can use spherical k-means [8] and compute the required values to build the d K curve (see Fig. 4).
The collinearity property in the new space (middle boxes) approximately transforms each of the clusters into lines radiating from the origin. After mapping these collinear points onto the surface of a hyper sphere (rightmost boxes), the clustering problem in most cases reduces in complexity, and an algorithm such as spherical k-means can be applied to find the d K curve and, subsequently, the number of clusters. The quality of the line formation depends on the RBF parameter σ and the shape of the original clusters. One can sweep σ and select the value that generates the best clusters according to the IJM criteria. The p = 3 raw data in the top row is highly correlated with N = 400 in each cluster, and it would be handled poorly by standard k-means. The p = 2 raw data in the bottom row could not be correctly segmented by any linear algorithm such as k-means [6, 7]
Using the d K information, one can calculate the corresponding M * K for each configuration tested (i.e., M * K = 0 , M * K = 1 , …, M * K = K max ). A simple comparison among the values of M * K is sufficient to determine a suitable number of clusters and appropriate parameters for the construction of the kernel matrix (i.e., model selection). The clustering is readily obtained from the identified parameters.
3.4 The Image Segmentation Technique: Application of IJM
In our IJM experiments, the RGB uterine cervix image is first transformed to the L * a * b * space, which is a more perceptual color space (this step is not applied to non-visual medical data) [9]. Then, the image color levels are quantized to only N q levels using the criteria of minimum variance. Next, the color-quantized image is subdivided into windows centered at each pixel, and a normalized histogram is computed for each window. A χ 2 score can then be used to compare the pixels as computed from these histograms [10]. The χ 2 score is defined as
where h i and h j are histograms for the windows located on the ith and jth pixels, and h i (k) indicates the repetition frequency of the kth bin (out of N q bins) in the corresponding histogram for the ith pixel. Finally, a χ 2 (chi-squared) RBF kernel and its kernel matrix can be computed from k(h i , h j ) = exp(−χ 2 ij /σ χ ), where k(h i , h j ) is the element in the ith row and jth column of the kernel matrix, and σ χ is the kernel standard deviation, as determined previously from the model selection.
The information theoretic IJM is applied to image segmentation by examining the local statistics of an indexed version of the image. Corona details many tests of IJM on synthetic data and evaluates this method’s performance on standard clustering benchmark datasets in his dissertation [6]. The performance of the IJM in segmenting uterine cervix images is also presented. This application is important because these kinds of medical images have no established ground truth, and even segmentations derived by experts differ due to subjective bias. It remained for IJM to be validated for an application involving medical images. This validation is carried out with MRI datasets of the human brain that have a labeled ground truth available.
4 IJM Validation Results
The IJM has been shown to perform well with benchmark clustering datasets. This is remarkable because IJM proves to be one of the few methods available that can cluster real-world data without knowing the number of clusters a priori. We desire eventually to apply the IJM to the segmentation of medical images to aid in the creation of automated diagnostic tools. One such application proposed here is the segmentation of uterine cervix images. However, the remaining issue is that the pathological uterine cervix images lack an established ground truth. Therefore, there is a need to validate the application of IJM to medical images, using medical image data with an available ground truth. This validation is done with a standard benchmark MRI dataset.
4.1 MRI Benchmark Test
Experiments are conducted with the standard benchmark MRI data of the human brain (BrainWeb: Simulated Brain Database; http://www.bic.mni.mcgill.ca/brainweb/), where the gray matter (GM), white matter (WM), and cerebrospinal fluid (CSF) are segmented from the background using the proposed technique. This dataset contains MRI cross-sectional images of a normal brain with no noise or field inhomogeneity [11]. The MRI raw data is given in terms of T 1, T 2, and the proton density, which were all available in both 8-bit and 12-bit formats, where the data has been stretched from the minimum to maximum value. These three modalities can be used to generate a pseudo-RGB space [12]. Analysis of the means and variances of the data for the various tissues shows that the space is not fully spanned, so an appropriate pseudo-color transformation can be performed on the raw data to improve the perception between the various tissue types (see Fig. 5; [7]).
The same procedure is followed for the MRI experiment as for the uterine cervix images. However, in this case, the number of clusters is selected using the second highest margin, instead of the maximum margin [6, 10]. The reason for this is the hierarchical nature of the results. Put simply, the maximum margin M * K reflects the most obvious cluster configuration in terms of cluster separation. The second highest margin is associated with less obvious and, sometimes, more interesting configurations. For example, the background and foreground often tend to strongly separate into two clusters, while we are actually interested in different types of foreground. Prior to the application of the clustering algorithm, the MRI data is preprocessed [11, 13]. The anatomical labels are used to create a mask that automatically removes most of the skull. The three modalities can be treated as a quasi-RGB image [12]. The original dataset does not span the entire range of possible values, so a transformation (to a pseudo-color space) is made to stretch out the ranges and to increase the perceptibility between the different tissue types. The transformed image is segmented using the procedure previously described (without the L * a * b * transformation). The anatomically labeled data can provide a ground truth to compare with the segmentation results of the IJM.
Model selection methods as applied to an MRI of the human brain. The top plot displays the number of clusters K est using the second best margin scores, where 15 out of 20 are for K = 4. (the best margin scores are always for K = 2. due to the striking difference between the tissue data and the background). The middle plot is of the corresponding margin scores, where the maximum margin at σ = 1.025 is highlighted in red (also for K = 4.). The bottom plot is the cluster balance. Note the 20 trial standard deviation values are on a logarithmic scale [7]
Figure 6 shows an example output of the model selection plots for such an MRI; the number of clusters with the second highest margin scores is K = 4, with a peak value located at the kernel standard deviation of σ = 1.025 in this run. Due to internal randomness of the training and testing sets, the algorithm’s results do somewhat vary from run to run. Recall that the jump method is hierarchical, so the second peak in J Y K can correspond to a valid sub-clustering of the data. For the model selection plots, the horizontal axes represent the standard deviation of the RBF kernel used. The top plots represent the selection of the number of clusters, called K est. The middle plots depict the values for the margin, while the bottom plots show the cluster balance values. The cluster balance is a measure of uniformity of the size of the clusters. Equal-size cluster configurations have a balance equal to one. Very different cluster sizes are associated with a balance close to zero. Figures 7 and 8 illustrate an example of MRI segmentation using the parameters selected from the model selection plot in Fig. 6. The confusion matrix for this experiment is shown in Table 2. The segmentation results using only eight color levels and a window size of three pixels for the four clusters are 100 % correct for the background, 94.58 % correct for the white matter, and only 92.75 and 90.82 % correct for the CSF and gray/glial matter associated segments, respectively. The overall accuracy is 95.48 %.
The original MRI after preprocessing (left), the false-colored cluster labels (middle) and the resulting segmented image (right) [7]
The successful segmentation results from applying the IJM algorithm, with the background (segment 1), the gray matter (segment 2), the white matter (segment 3), and the cerebrospinal fluid (segment 4) [7]
The IJM has been shown to perform at 95 % agreement with a labeled MRI dataset, taking into account hierarchical issues. An MRI of the human brain has a distinct background, so it is most obviously clustered into two segments; the sub-clustering of the brain tissue is the issue of importance. The variation in illumination in the uterine cervix images is handled by factoring in cluster balance. The segmentation of the human brain MRI neglects the consideration of cluster balance, yet it still achieves acceptable results.
4.2 Segmentation of Color Uterine Cervix Images
In the uterine cervical images, changes in three distinct and usually concentric regions are of importance to specialists for early detection of cervical cancer. Because cervical cancer slowly develops, it causes a detectable and treatable precursor condition known as cervical intraepithelial neoplasia (CIN). CIN causes abnormal regions and changes in the cervix that are of interest to specialists for several reasons. The columnar epithelium (CE) region immediately surrounds the central cervix opening, and it is near the center of the image. The CE region is bright red and is usually the darkest in the luminance plane. The acetowhite (AW) region is the most common abnormality immediately surrounding the (CE) region. It appears white or very pale pink and is usually the brightest region in the luminance plane. The outermost region is the squamous epithelium (SE), which has a reddish hue usually between that of AW and CE [6]. The acetowhite region is produced by applying a 5 % acetic acid solution to the cervix, which causes the abnormal cells to become turgid (swollen) and increases their reflectivity relative to the normal CE. They become turgid partly because their cellular machinery is not working normally, causing them to produce different kinds of cytokeratin (cellular structural proteins), hence they are unable to hold their normal shape [14].
Model selection methods as applied to a uterine cervix image. The top plot displays the number of clusters K est with the overall best margin scores, where 15 out of 20 are for K = 3. The middle plot is of the corresponding margin scores, where the maximum margin at σ = 0.886 is highlighted in red (also for K = 3). The bottom plot is of the cluster balance. Note that the 20 trial standard deviation values are on a logarithmic scale [7]
The IJM was also used to segment color uterine cervix images into these three regions for an abnormal case. The model selection results are shown in the form of plots (see Fig. 9) as described in the previous subsection. Figure 10 shows an example of uterine cervix image segmentation. The model selection plots indicate that K = 3 is a good configuration, which is located at σ = 0.866. The segmentation using these parameters is shown in Fig. 11. It should also be noted that in the generation of this segmentation cluster balance was also factored in. Cluster balance is a measure of size similarity between clusters. For MRI data, one does not expect the clusters to similar in size to each other, while for uterine cervix images one might. Thus, the cluster balance was calculated for each configuration and multiplied by the margin to determine the overall quantity to be maximized. This balancing tends to stabilize the segmentation of the uterine cervix images against smaller sub-clusters of the same region, such as the brightest region in Segment 2 of Fig. 11.
The original uterine cervix image after preprocessing (left) and the segmented image (right) [7]
Demonstration of uterine cervix image segmentation using the model selection in IJM [7]
The direction of ongoing and future work is two-pronged. One is to apply the IJM to the segmentation of MRIs of abnormal human brains, and the other is to apply the IJM to the segmentation of cell images. Abnormal brain features tend to be small and distinct. This poses quite a challenging problem for the current implementation of the IJM. Evaluations have identified limitations of the IJM in such cases. To overcome these limitations, an iterative hierarchical approach is suggested along with a dynamic window size. These limitations and this proposed solution are detailed below.
In addition, much work has been done in automating the entire process for diagnosing cervical cancer. In addition to segmenting the uterine cervix images, there is also a need to preprocess and segment micrograph slides of cell smears. Many methods have been applied to this problem, and there is an aim to bring the IJM to bear on the preprocessing and segmentation of such cell images. This technique could aid in the development of a diagnostic tool for cervical cancer.
5 MRIs with MS Lesions
5.1 The IJM Failure with MRIs with MS Lesions
After successfully segmenting MRIs of a normal brain with the IJM, the next step of the validation process is to attempt to segment the MRI of a brain with multiple sclerosis (MS) lesions. This is a more challenging segmentation problem not just because of the existence of another cluster (K = 5 is expected), but also because the MS lesions are relatively small compared to the other tissue areas. Upon performing experiments, indeed five clusters are weakly indicated with the third highest margin (K = 8 is the found for second highest margin), however, the segmentation results are incorrect (see Figs. 12 and 13). The gray matter segment splits into the two additional transition zones between gray matter/white matter, and the gray matter/CSF, respectively, to form five clusters; the MS lesions remain undistinguished. The bulk of the MS lesion is included with the CSF cluster, while the transition zone between the MS lesion and the white matter is included with the transition zone between the gray and white matter. This failure prompted a study of the limitations of the implementation of the IJM.
The original MRI with MS lesions after preprocessing (left), the false-colored cluster labels (middle), and the resulting segmented image (right) [15]
The failed segmentation results from applying the IJM algorithm, with the background (segment 1), the gray matter/white matter transition zone (segment 2), the white matter (segment 3), the gray matter/CSF transition zone (segment 4), and the cerebrospinal fluid (CSF) (segment 5) [15]
5.2 Limitations of the Proposed Image Segmentation Technique
In the above segmentation algorithm, the image color levels are quantized to only N q levels; this color-quantized image is subdivided into windows centered at each pixel, and a normalized histogram is computed for each window. The estimation of the distortion and hence the margin operator is then calculated from these histograms. One limitation of this implementation is that the number of quantization levels must be less than or equal to the number of pixels in a window. Two issues are at the root of the failure of this IJM implementation. One issue is due to the size of the window and the other is due to the number of quantization levels.
Relying on the window to determine local statistics is good for a relatively smoothly varying image, such as the uterine cervix images. However, for images with a sharp background, discontinuities, or rapidly changing edges, the local statistics can become peculiar in the transition zones between regions. If such boundaries are long and crenelated like those between the internal tissues of the brain, then local statistics from the windows can “trick” the algorithm into detecting another statistically significant segment. This new segment is the aggregate of the transition zone, which in the case of the MRIs can have an area larger than the area of the MS lesion tissue. This pseudo-clustering tendency also helps to explain an unexpected observation. Sometimes improving the perception contrast between regions can actually lead to poorer segmentation results. Again this is due to having more sharp transition zones. Decreasing the window size will minimize this effect for all pixels, except for those directly on the edge boundary.
Reducing the window size can mitigate the transition zone effect, but this can have an unintended consequence; it reduces the number of quantization levels. This reduction is significant because the ability to discern between regions in an image is limited by pixel indexing. Limiting quantization goes back to the core issue of this application of information theory, storage efficiency versus the ability to discriminate. As it turns out, the normal brain MRIs are handled well with a 3 × 3 window with eight quantization levels (labeled as zero through seven). Examining the raw index image shows that the different tissues are distinctly labeled. However, examining the MS lesion image reveals that the MS lesions are not distinctly labeled (with index 19) until there are at least 20 quantization levels. This number of levels is required to accommodate the transition zones between regions and the MS lesion tissue. All the various color regions that are comparable in size to the MS lesion area would bring the effective cluster number to at least seven or eight. Thus the MS lesions are not even distinguishable unless at least a 5 × 5 window is used. However, using a window size and number of quantization levels so large also turns the transition regions into statistically significant pseudo-clusters. The current IJM implementation here fails in the proper segmentation of the MS lesions. This issue was not previously encountered when applying the IJM to synthetic test data and medical images.
5.3 The Proposed Solution: An Iterative Hierarchical Scheme
One means to deal with the problematic transition zones is to divide them before they become a problem. Hence, this approach is to “divide and conquer.” This scheme would involve utilizing the hierarchical information of the IJM in a different manner. Recall, that for the MRIs, the largest margin indicated K = 2 clusters. This result suggests that the correct approach is to first segment the MRI into two clusters, “cleaving” the background from the brain tissue cluster. Next, one reapplies the IJM to further segment the brain tissue super-cluster using a masked image, etc. In such an iterative scheme, the various segments can be progressively cleaved from each other, dividing the transition zones along the way and making them less significant. This process can be continued until some parameter threshold or statistical feature of the subsequent segmented image signals the iterative program to stop. This process could be automated to iteratively segment the image according to the number of clusters with the largest margin as obtained from the IJM, using the full power of its hierarchical character. In this manner, the quantization restriction can be partially decoupled from the transition zone problem.
5.4 An Adaptive Window Size
The problem with iteratively segmenting an image is that the shapes of the future iterations of image masks are unknown. Consider the cases of a small “peninsula” or “island” of pixels with fewer neighbors than the window size (see Fig. 14). Another case would be an edge with a reduced number of neighbors. These samples near the mask boundary would degrade the subsequent statistical analysis of the histograms. Furthermore, as the sub-clusters get smaller and demonstrate more complicated edges, the performance of the IJM will deteriorate. In a sense, the transition zone problem is replaced with a mask edge effect problem.
The root of this problem is that for pixels near an already cleaved boundary, the effective window size has changed. In order to counter this effect, the window size must be adaptive for the segmentation of the masked images. The purpose of the window is to provide a consistent pool of neighboring pixels in order to characterize the local statistics. When segmenting a single image into multiple clusters, every sample pixel has the same window size; those on the edge of the image can be used with simple symmetric padding. However, this uniformity is broken in the subsequent steps of the iterative hierarchical scheme, and the effects of quasi-symmetric extrapolations could actually amplify the transitions near boundaries. In order to carry out the original purpose of the window, its size must be adaptive for the masked images. The suggested implementation enlarges the original square window into a successively more circular shape until a sufficient number of neighboring pixels are sampled.
The following description outlines the procedure to adaptively enlarge the window size. A pixel contained within the mask is selected. The initial window is applied, and the number of existing neighbors is counted. If the number falls below the default window size, then the window size is enlarged. Allow at most three enlargements or until the default window size is passed. Randomly remove any excess pixels or randomly duplicate the pixels found until the window has the same number of pixels as the default window size. Now the histogram statistics for almost every sample pixel derived from the segment mask should be comparable. In this manner, the effects of transition zones are again reduced, because more of the pixels in the “bulk” are included in the histograms of sample pixels near boundaries. This procedure should also allow relatively small clusters like the MS lesion tissue to be robustly detectable, provided sufficient quantization levels. Hence, using an iterative hierarchical scheme and an adaptive window size should allow the proper segmentation of MRIs containing MS lesions using the IJM with at least 20 quantization levels.
6 Application to Pap Test Images
Another medical segmentation imaging problem of interest is the processing of cytological images in separating the nucleus from the cytoplasm. An example of this is seen in the micrographs of stained cells derived from the Papanicolaou (Pap) smear test. In the diagnosis of cervical cancer, cervical uterine photographic images are examined, and if a suspicious region is identified, then it is swabbed. The collected cells are then smeared onto a slide, stained, magnified through a microscope, and photographed. Precancerous or cancerous cells often exhibit a condition called polyploidy, where they have multiple copies of the same chromosome in their nuclei. Polyploidy usually causes their nuclei to be larger than normal. Hence, an automated system that can measure the ratio of the area of the nuclei to cytoplasm in Pap test images could be useful as a diagnostic aid. There are many approaches to such a system; one is reviewed below.
Current routine clinical tests for cervical cancer include the cytology-based Pap smear test and HPV testing to identify DNA or RNA in cervical cells. Because the recently developed HPV vaccination does not protect against all HPV types, cervical screening tests are still required [16]. Because cervical cancer develops slowly and has a precursor condition known as dysplasia or CIN, it can be detected through screening of at-risk women and treated to prevent the development of invasive cancer [17]. However, the true sensitivity of the conventional Pap smear has been reported to be as low as 51 %, requiring annual screenings to detect CIN, making the Pap smear impractical to implement in resource-poor regions [18, 19]. The development of automated diagnostic tools could increase the universal effectiveness and availability of the Pap test.
Several combinations of imaging modalities and classification algorithms have been used by researchers to detect some of the abnormalities due to CIN [20]. However, the problem of isolating and segmenting specific abnormalities, such as mosaicism and punctations, is not addressed by most methods. In previous work, an automated computer vision system has been developed for the detection and characterization of CIN in an attempt to overcome the inadequacies of existing approaches used for automated diagnosis of CIN [9]. This system is capable of automatically segmenting the cervix region of interest (ROI) from raw cervicographic images; removing specular reflection (SR); segmenting the cervix ROI into acetowhite (AW), columnar epithelium (CE), and squamous epithelium (SE); classifying AW regions into AW, mosaic, or punctation tiles; segmenting mosaic and punctation from AW tiles; and assessing of disease severity [9, 18].
We intend to utilize the IJM [6] for precise segmentation of the macro-features as well as a hybrid watershed model for segmentation of cytological cell images acquired by Pap test to assess abnormalities in the cell nuclei [21]. The significance of including the cell structure analysis is to provide a cross-validation of cervical abnormalities assessed by two different approaches, namely from the analysis of the color cervix images and of the cervical cell images. Such an automated system for analyzing both cervix images and cervical cell images would provide an improved diagnostic tool for cross-validating the results from both imaging modalities. Based upon the validation of the IJM unsupervised learning algorithm, the high cost of generating manual ground truth from specialists, who are not in general readily available in resource-poor regions, can be reduced [18]. With the proposed modifications to its implementation discussed in Sects. 5.3 and 5.4, it should be possible to segment these images using the IJM. The clusters in the Pap test images have sharp edges and are better balanced than the MS lesion case in the MRIs.
6.1 A Hybrid Watershed Method for Generating Cervical Cell Profiles
A combination of global histogram thresholding and local watershed segmentation was used to generate the cervical cell profiles for final separation of the cell nuclei from the cell cytoplasm to assess the abnormalities in the cells. The preprocessing steps are described in detail in reference [21]. The selection of useful cervical cell images to be analyzed can be automatically achieved by the IJM clustering, and subsequently the cell nuclei can be precisely segmented. Figure 15 shows some examples of cervical cell segmentation results with this technique.
6.2 Experimental Results
All uterine cervix images and cervical cell cytology images were provided by the National Library of Medicine from the data collected by the National Cancer Institute at the National Institutes of Health. Figure 15 shows typical segmentation results of cervical cell images with semiautomated and manual segmentation, where corresponding Dice coefficients [22] are used as the segmentation quality measures. Dice’s coefficient is defined as:
in which S Auto is the pixel set from automatic segmentation, and S Manu is the pixel set from manual segmentation. The proposed IJM algorithm is expected to improve these segmentation results by automatically identifying and discarding the background and objects of no interest [18].
7 Conclusion
7.1 Further Development
The application of information theoretic techniques such as the IJM to medical image analysis is promising. The current outlook is to successfully apply the IJM to the segmentation of MRIs of abnormal human brains, such as brains with MS lesions. One scheme could involve scanning through the MRI slices for MS lesions using IJM. Once found, a simple Gaussian model of the pixel values can be constructed and applied to the full volume, allowing an estimation of the total MS lesion volume that can be tracked over time. Alternatively, a three-dimensional voxel segmentation of the MS lesion could be developed by applying the IJM.
Finally, the IJM also is promising in its application to segmenting cell images, such as those derived from the Pap smear test. In the preprocessing step, the IJM could aid in discarding “junk” slide images from those containing useful images. When useful images are found, the IJM offers an additional method to segment such images into nucleus and cytoplasm clusters. Cell overlaps could even be identified to correct the estimation of the cytoplasm area. Then the ratio of nuclear to cytoplasm area can be calculated and used as a diagnostic aid. These steps combined with the uterine cervix image segmentation described earlier can be used as components of an automated, integrated diagnostic system for cervical cancer.
7.2 Outlook for Application
The IJM effectively performs an optimized transformation of the distortion-rate curve D(R) to reveal a distinct signature of the cluster configurations in an unlabeled image. This improved clustering technique, based on an information theoretic approach, demonstrates high accuracy in segmentation of the standard benchmark MRI data of a normal human brain.
Limitations of the current implementation of the IJM are found that cause it to fail to correctly segment the MRI data of a human brain with MS lesions. The root source of the failure was identified as the joint effect of the requirement for a relatively high number of quantized levels (20) and the problematic transition zones between tissues that can be sources of pseudo-clusters. An iterative hierarchical scheme with adaptive window sizes is proposed to make the IJM more robust in its segmentation.
This chapter discusses an element of a computer vision system with embedded novel segmentation models for analyzing both cervix and cervical cell images for detection of precancerous lesions. Application of the proposed IJM in segmentation of uterine cervix images, with ground truth unavailable in most cases, has potential to identify abnormal tissue characteristics at an early stage, thus providing a diagnostic aid to assessment of uterine cervical cancer. Such a system allows cross-validation of the CIN abnormalities detected by two different testing methods and may reduce frequent false-positive results obtained by the routine Pap test. This system may also provide a low-cost yet valuable analysis tool to aid in preventive assessment of cervical cancer in resource-poor regions [18].
References
Ghabhramani, Z.: “Unsupervised Learning.” Advanced Lectures on Machine Learning, pp. 72–112. Springer, New York (2004)
Gokcay, E., Principe, J.C.: Information theoretic clustering. IEEE Trans. Pattern Anal. Mach. Intell. 24(2), 158–171 (2002)
Bartlett, M.S.: Face Image Analysis by Unsupervised Learning. Kluwer Academic, Boston (2001)
Becker, S., Plumbley, M.: Unsupervised neural network learning procedures for feature extraction and classification. J. Appl. Intell. 6, 1–21 (1996)
Sugar, C.A., James, G.M.: Finding the number of clusters in a dataset: an information-theoretic approach. J. Am. Stat. Assoc. 98(463), 750–763 (2003)
Corona, E.: Unsupervised learning methods: an efficient clustering framework with integrated model selection. Ph.D. Dissertation, Texas Tech University, Lubbock (2012)
Corona, E., Hill, J., Ao, J., Nutter, B., Mitra, S.: An information theoretic approach to automated medical image segmentation. In: Proceeding of SPIE, vol. 8669. Medical Imaging 2013: Image Processing (2013)
Alzate, C., Suykens, J.A.K.: Multiway spectral clustering with out-of-sample extensions through weighted kernel PCA. IEEE Trans. Pattern Anal. Mach. Intell. 32(2), 335–347 (2010)
Srinivasan, Y., Corona, E., Nutter, B., Mitra, S., Bhattacharya, S.: A unified model-based image analysis framework for automated detection of precancerous lesions in digitized uterine cervix images. IEEE J. Sel. Top. Signal Process. 3(1), 101–111 (2009) (Special Issue on Digital Image Processing Techniques for Oncology)
Fowlkes, C., Belongie, S., Chung, F., Malik, J.: Spectral grouping using the Nystrom method. IEEE Trans. Pattern Anal. Mach. Intell. 26(2), 214–225 (2004)
Kwan, R., et~al.: An extensible MRI simulator for post-processing evaluation. In: BrainWeb: Simulated Brain Database. Proceedings of VBC'96, pp. 135–140. Springer (1996). Available online: http://www.bic.mni.mcgoll.ca/brainweb/
Lecoeur, J., Wang, F., Chen, L.M., Li, R., Avison, M., Dawant, B.: Sub millimeter coregistration of functional maps across imaging sessions. In: Proceedings of ISMRM Annual Conference, vol. 1625, Montreal, May 2011
Ge, Z., Mitra, S.: Optimized statistical modeling of MS lesions on MRI voxel outliers for monitoring the effects of drug therapy. Proc. SPIE 4684, 873–882 (2002)
Maddox, P., Szarewski, A., Dyson, J., Cuzick, J.: Cytokeratin expression and acetowhite change in cervical epithelium. J. Clin. Pathol. 47(1), 15–17 (1994)
Hill, J.E.: Application of information theoretic unsupervised learning to medical image analysis. Master Thesis, Texas Tech University, Lubbock (2013)
Ferlay, J., et~al.: GLOBOCAN 2002: Cancer Incidence, Mortality and Prevalence Worldwide. IARC CancerBase No. 5. Version 2.0. Online. Available: http://www.iarc.fr/ (2002)
Koss, L.G.: The Papanicolaou test for cervical cancer detection. A triumph and tragedy. JAMA 261, 773–774 (1987)
Corona, E., Hill, J., Ao, J., Nutter, B., Mitra, S.: A novel unsupervised learning model for automated detection of precancerous abnormalities in uterine cervix with unified analysis of cervical cells and digital uterine cervix images. In: First IEEE Healthcare Technology Conference: Translational Engineering in Health & Medicine, 7–9 November 2012. IEEE HIC 2012 Paper Abstract, Paper ThCT2.13 (2012)
Ferris, D.G.: Cervicography-an adjunct to Papanicolaou screening. Am. Fam. Physician 50, 363–370 (1994)
Gordon, S., Zimmerman, G., Long, R., Antani, S., Jeronimo, J., Greenspan, H.: Content analysis of uterine cervix images: initial steps towards content based indexing and retrieval of cervigrams. Proc. SPIE Conf. Med. Imaging 6144, 1549–1556 (2006)
Ao, J., Mitra, S., Long, R., Nutter, B., Antani, S.: A hybrid watershed method for cell image segmentation. In: Proceedings of IEEE Southwest Symposium on Image Analysis and Interpretation, Santa Fe, 22–24 April 2012
Dice, L.R.: Measures of the amount of ecologic association between species. Ecology 26(3), 297–302 (1945)
Acknowledgments
This research was partially supported by the Intramural Research Program of the National Institutes of Health (NIH), National Library of Medicine (NLM), and Lister Hill National Center for Biomedical Communications (LHNCBC).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Hill, J., Corona, E., Ao, J., Mitra, S., Nutter, B. (2014). Information Theoretic Clustering for Medical Image Segmentation. In: Saha, P., Maulik, U., Basu, S. (eds) Advanced Computational Approaches to Biomedical Engineering. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-41539-5_2
Download citation
DOI: https://doi.org/10.1007/978-3-642-41539-5_2
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-41538-8
Online ISBN: 978-3-642-41539-5
eBook Packages: Computer ScienceComputer Science (R0)