Abstract
The economical production of ethanol from lignocellulosic materials needs the conversion not only of glucose, which is the sugar of preference of the best performing ethanologenic microorganisms, but also of the rest of sugars found in the fermentation broth, derived from pretreatment and enzymatic steps. This chapter summarizes recent work directed to that objective, by using different modification techniques of microorganisms. After considering the main metabolic pathways for pentoses, the second most abundant kind of fermentable sugars, a review of such modifications taking either Escherichia coli or Saccharomyces cerevisiae as a basis is presented. Although E. coli and S. cerevisiae are the most studied microorganisms through a wide range of techniques, other microorganisms are also being subject of study with the same purpose, and are briefly described at the end of this chapter.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
8.1 Introduction
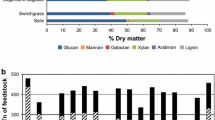
Lignocellulosic materials are mainly composed of cellulose, hemicelluloses, lignin, extractives, and ashes. It has been claimed that they are the most promising sugar source for biofuel production mainly due to their renewable nature and the lack of competition with food or feeding applications. But in contrast to starch or raw materials containing sucrose such as corn or sugarcane, the generation of fermentable sugars from the homogeneous (cellulose) or heterogeneous (hemicelluloses) polysaccharides of lignocellulose need an intense, energy demanding pretreatment step. Cellulose forms the major part of lignocellulosic materials and hence, once the pretreatment and enzymatic hydrolysis steps have been performed, the sugar broth is constituted by glucose as the major sugar. Nevertheless, the fraction of hemicellulose can account for up to 30 % depending on the material, making sugars derived from hemicellulose very important in economic terms; sometimes their transformation into ethanol becomes the threshold of profitability. Among these sugars, xylose is the most abundant, after glucose. Arabinose and cellobiose are also present in significant proportions in certain lignocellulosic materials. Converting xylose and other sugars, in addition to glucose, into ethanol would reduce the overall production costs and make the process a real alternative to fossil fuels from an economic point of view. Additionally, the pretreatment step is also responsible for the presence of inhibitor compounds which may hinder the fermentation process. To overcome these drawbacks, the ideal ethanologenic microorganism should be able to efficiently produce ethanol from different kinds of sugars and be resistant to the presence of both inhibitors and ethanol. There are a number of microorganisms able to naturally ferment a wide variety of sugars, including glucose, xylose, arabinose, and others but, unfortunately, those microorganisms do not perform the same when a mixture of sugars is present. Instead, they assimilate in such a way that one of the sugars is preferable to the others, and this particular one may act in some way to repress the others, thereby reducing the overall capacity of the microorganisms to transform all the sugars. Escherichia coli is one of those microorganisms able to ferment a wide range of sugars and it is the focus of a very intense investigation centered on trying to improve ethanolic fermentation results.
Another strategy to take full advantage of the different sugars issued from the pretreatment of lignocellulose consists in using metabolic engineering to modify microorganisms that ferment glucose with good results, and to repeat this behavior with other sugars. Recombinant DNA technology and evolutionary engineering techniques are being assayed in an attempt to improve both ethanol yield and productivities. In this particular field, Saccharomyces cerevisiae is the main candidate to modification.
As an alternative to use several steps to convert pretreated lignocellulosic materials into ethanol, the process can also be addressed by combining all of them in a single one. This is the fundamental of the so-called Consolidated Bioprocessing (CBP), which can be advantageous from an economic point of view because enzyme production, saccharification, and fermentation are conducted in a single vessel. In CBP, both cellulosic and hemicellulosic materials should be simultaneously fermented. This kind of process requires a highly engineered microorganism able to produce effective hydrolyzing enzymes, for high ethanol titer and productivities, using both hexoses and pentoses from a high solid pretreated material. In addition, it is recommended that this special microorganism exhibits a high resistance to ethanol, fermentation inhibitors, and stressful environments (Hasunuma and Kondo 2012).
This chapter summarizes the main strategies aimed at taking full advantage of all sugar fractions obtained from lignocellulosic materials for ethanol production through the use of modified microorganisms. First, it presents a brief introduction to pentose metabolism so that the main points for modification in microorganisms can be identified. Then, these main points are described based on either E. coli or S. cerevisiae. Finally, the chapter presents a concise review of recent results dealing with ethanol production from lignocellulosic materials using other modified microorganisms.
8.2 Pentose Metabolic Pathways
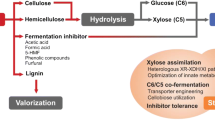
Figure 8.1 shows a simplified diagram of the main sugars, other than glucose, found in lignocellulosic materials hydrolysates, d-xylose, and l-arabinose (Hahn-Hägerdal et al. 2007).
Simplified metabolism of d-xylose and l-arabinose in bacteria and fungi (Hahn-Hägerdal et al. 2007)
The biochemical route for xylose metabolism is the pentose phosphate pathway (PPP), which is present in all cellular organisms, and can be described as a two-step process (Jeffries 2006): conversion of d-glucose 6P into d-ribulose 5P (oxidative phase) and further conversion (non-oxidative phase) into several compounds including d-xylulose 5P, which is the way in which d-xylose enters the PPP. To point out one main difference, the conversion of d-xylose into d-xylulose occurs in bacteria under the action of xylose isomerase, while in yeasts, the process includes reduction and oxidation mediated by the enzymes xylose reductase and xylitol dehydrogenase (Fig. 8.1), with xylitol as an intermediate compound.
Concerning l-arabinose, its metabolism is more complex, and requires more enzymatic reactions for the transformation into compounds entering the PPP (Hahn-Hägerdal et al. 2007). Nevertheless, the amount of this sugar in lignocelluloses is in general much lower than that of d-xylose, drawing little attention on the arabinose metabolism pathway.
The modification strategies implemented in the processes regarding how microorganisms can get a better use of sugars contained in lignocellulosic materials and improve ethanol yield include several metabolic and evolutionary engineering techniques and advanced genomics, transcriptomics, proteomics, metabolomics techniques, as reported by Gírio et al. (2010). Considering the metabolic pathways depicted in Fig. 8.1, an intense research work has been developed in recent years through the introduction of foreign genes, the elimination of competitive pathways, the disruption of byproducts formation (Jarboe et al. 2007), and the study of the redox imbalance from xylose reductase and xylitol dehydrogenase, xylulokinase, among others (Chu and Lee 2007).
8.3 Escherichia coli-Based Modifications
The nonpathogenic, Gram + species of bacterium E. coli ferments a wide range of monomeric sugars, including all those present into hemicellulosic and cellulosic hydrolysates: xylose, arabinose, glucose, galactose, mannose, and also uronic acids, such as glucuronate, that is obtained from the hydrolysis of some lignocellulosic materials (Alterthum and Ingram 1989). However, the cultivation of wild-type E. coli under fermentative conditions produces a variety of fermentative products: lactate, succinate, acetate, formate, hydrogen, carbon dioxide, and small amounts of ethanol. Furthermore, for several decades E. coli has been the workhorse for the development of genetic and molecular biology tools, including modifications for the production of ethanol through metabolic engineering, i.e., the improvement of cellular activities by manipulations of enzymatic, transport, and regulatory functions of the cell with the use of recombinant DNA technology (Bailey 1991). Therefore, for more than two decades, this microorganism has been the target for metabolic modifications and it has been studied for the production of ethanol (Jarboe et al. 2007; Orencio-Trejo et al. 2010). With the advent of the system and synthetic biology tools, a new generation of E. coli strains are being metabolically engineered for the production of fuel ethanol and the so-called, advanced biofuels including, but not limited to: 1-propanol, n-butanol, isobutanol, isopentanol, pinene, farnesane, bisabolane, fatty-acid methyl esters, fatty-acid ethyl esters, fatty alcohols, polyketide-derived biofuels, aromatic alcohols, and alkenes and alkanes from short (C5) to long carbon chain (up to C20) (Dellomonaco et al. 2011; Huffer et al. 2012; Peralta-Yahya et al. 2012; Rodríguez-Moya and Gonzalez 2010).
The introduction of foreign genes, using plasmids or cloned into the E. coli chromosome, the elimination of competitive pathways, the adaptive evolution, carbon flux distribution, and the disruption of byproducts formation have been the main strategies used to improve sugar utilization, ethanol and inhibitors tolerance, ethanol yields on carbon consumed, and specific and volumetric ethanol productivity (Ingram et al. 1999; Jarboe et al. 2007; Orencio-Trejo et al. 2010). But some drawbacks are still highlighted for this microorganism, such as the low tolerance toward ethanol and the narrow and neutral working pH (6–8) (Gírio et al. 2010). However, several metabolically engineered strains had shown ethanol yields above 95 % of the theoretical maximum, either using pure sugars, sugar mixtures (pentoses and hexoses), and actual lignocellulosic hydrolysates containing acetate (Fernández-Sandoval et al. 2012; Geddes et al. 2011).
In the late 1980s, E. coli was used to drive the expression of the Zymomonas mobilis genes that encode pyruvate decarboxylase (pdc) and alcohol dehydrogenase II (adhII) (Ingram and Conway 1998), allowing the production of ethanol instead of organic acids. E. coli K011, developed by Ingram and coworkers (USA Patent 5,000,000), constitutes one of the breakthroughs in E. coli toward the development of ethanologenic bacteria by means of metabolic engineering. The development consisted in: (i) the chromosomal integration of genes encoding pdc and adhII from Z. mobilis under the control of the pyruvate formate lyase promoter (this promoter drives a strong transcription under fermentative conditions); (ii) deletion of one subunit of the fumarate reductase (to avoid the formation of succinate under fermentative conditions); (iii) and selection of variants (through adaptive evolution in plates with a high concentration on the antibiotic marker used for the integration of pdc and adhII) showing high pyruvate decarboxylase and alcohol dehydrogenase enzymatic activities. The main results attained with E. coli K011 under different operational conditions are summarized in Table 8.1, and Fig. 8.2 shows common modifications performed into wild-type E. coli strains to attain ethanologenic strains.
Fermentation pathways in wild-type E. coli and common modifications in metabolically engineered ethanologenic strains. Genes encoding enzymes are indicated by italics, the “X” sign indicate deleted genes to reduce byproducts. Ethanol production pathway (pdc and adhB) from Z. mobilis is shown by dashed arrows. adhE alcohol dehydrogenase, frdABCD fumarate reductase, ldhA lactate dehydrogenase, pflB pyruvate formate lyase, pta phosphotransacetylase, ackA acetate kinase A, pdc pyruvate decarboxylase from Z. mobilis, adhB alcohol dehydrogenase B from Z. mobilis
Although E. coli K011 produced ethanol with specific rates as high as those of yeasts (S. cerevisiae), its ethanol tolerance was lower than that of the yeasts, hence metabolic evolution was applied for improving ethanol tolerance and ethanol production, leading to E. coli strain LY01 (Yomano et al. 1998; Jarboe et al. 2007). Nevertheless, the performance of both K011 and LY01 relied on costly nutritional supplementation which is not justified when producing biofuels. To address this issue, a new medium was developed (AM1, Martínez et al. 2007) along with a new development of homo-ethanologenic strains, such as LY168 (Yomano et al. 2009). This strain contained a deletion of the methylglyoxal synthase gene (mgsA) that resulted in the co-metabolism of pentose and hexose sugars to ethanol and was able to utilize hemicellulose sugars from hydrolysates as carbon source. In order to increase the substrate range, further modifications to LY168 resulted in the strain LY180 (Miller et al. 2009). LY180 was then subjected to serial transfers in hemicellulose hydrolysates obtained from the steam explosion pretreatment of sugarcane bagasse using dilute sulfuric acid (and later dilute phosphoric acid) to increase the tolerance of the strain to the fermentation inhibitors resulting from pretreatment. The inhibitor tolerant strain obtained (MM160) was able to ferment slurries of dilute phosphoric acid pretreated sugarcane bagasse with ethanol yields of over 0.20 g/g (Geddes et al. 2011). Additional serial transfers of MM160 in hemicellulose hydrolysates resulted in a superior strain (MM170) with an improved yield of 0.27 g/g (Nieves et al. 2011).
Vinuselvi and Lee (2012) assayed a combination of genetic and evolutionary engineering strategies for improving simultaneous utilization of cellobiose and xylose. The recombinant E. coli was capable of utilizing around 6 g/L of cellobiose and 2 g/L of xylose in approximately 36 h, whereas wild-type E. coli was unable to utilize xylose completely in the presence of 6 g/L of glucose even after 75 h.
Recent reports have demonstrated the development of a new generation of homo-ethanologenic strains that can grow and produce ethanol efficiently in the presence of acetate (Fernández-Sandoval et al. 2012); ferment cellobiose and glucose mixtures simultaneously (Muñoz-Gutiérrez et al. 2012); produce cellulases and xylanases allowing the direct fermentation of pretreated corn stover cellulose into ethanol (Ryu and Karim 2011); ferment alginate from brown macroalgae into ethanol (Wargacki et al. 2012); ferment pectin-rich lignocellulosic biomass, specifically sugar beet pulp, into ethanol (Edwards et al. 2011).
Concerning the transformation of actual hydrolysates into ethanol, some authors have determined the performance of different E. coli strains. Thus, the effects of both ethanol and degradation products have been identified as the main reasons for the loss of fermentability of E. coli K011 for transforming sugars contained in corn stover Ammonia Fiber Explosion (AFEX) pretreated xylose hydrolysate into ethanol (Jin et al. 2012). However, using a new generation of metabolic engineered strains (E. coli MM160 and MM170), Geddes et al. (2011) and Nieves et al. (2011) have studied the fermentation of sugarcane bagasse slurries, obtained by phosphoric acid pretreatment, into ethanol. After applying simultaneous saccharification and fermentation, with a previous liquefaction step, these researchers have demonstrated a yield of 340 l of ethanol per dry metric ton of untreated bagasse. Furthermore, the fermentation of these slurries has been scaled up to 80-L fermentors (Nieves et al. (2011), and Ingram and coworkers in partnership with Buckeye Technologies Inc. are developing, for the first time, a biorefinery demonstration plant in Perry, Florida, USA, with the aim to “produce up to 400 gal of fuel ethanol (and 2,000 pounds of organic acids for biopolymers) each day”, using lignocellulosic biomass and the E. coli strains described in this paragraph.
Some studies addressed the behavior of modified-ethanologenic E. coli strains in salt-rich media, because hydrolysates from agricultural residues are frequently rich in salts, which can exert an inhibitory effect on the growth of the bacteria. For example, Qureshi et al. (2006) determined that ethanologenic E. coli FBR5 could tolerate up to 40 g/L of NaCl, although the microorganism exhibited some inhibition for concentrations above 10 g/L. Saha et al. (2011) successfully used recombinant E. coli FBR5 on hydrolysates from sulfuric acid pretreated wheat straw. 41.1 g ethanol/L were produced in 168 h at pH 7.0 and 35 °C. The authors claim that this is the first report showing greater than 4 % ethanol production from lignocellulose by the strain.
It is also worth noting that E. coli, which had long been considered incapable of utilizing glycerol as raw material for ethanol production, was recently metabolically engineered by overexpressing the aldehyde dehydrogenase and deleting the lactate dehydrogenase genes, which resulted in a high yield of ethanol (20.7 g/L), associated with a productivity of 0.22 g/L/h (Durnin et al. 2009). Further studies with glycerol have allowed proposing the use of this residual and inexpensive chemical from the biodiesel industry as a platform for the production of several biofuels and chemical with the concomitant development of E. coli metabolical engineered strains (Dellomonaco et al. 2011; Rodríguez-Moya and Gonzalez 2010).
8.4 Saccharomyces cerevisiae-Based Modifications
Because S. cerevisiae is one of the best glucose-fermenting microorganisms but it lacks the ability to ferment xylose, several strategies have been identified to enable it to utilize xylose. These strategies correspond to the two pathways depicted in Fig. 8.1. The first one (reductive-oxidative pathway) is moduled by xylose reductase (XR), to catalyze the transformation of xylose to xylitol, and xylitol dehydrogenase (XDH), which catalyzes further conversion to xylulose. The second pathway implies the use of xylose isomerase to directly convert xylose to xylulose, which in turn enters the pentose phosphate pathway (Parachin et al. 2011) by means of the action of xylulokinase (XK).
It is worth mentioning that even if S. cerevisiae possesses the genes encoding the enzymes XR, XDH, and XK, it is not able to ferment xylose even when the genes were overexpressed, Hahn-Hägerdal et al. (2007). So a big research effort in the last four decades has been devoted to express those genes into S. cerevisiae so that it is able to efficiently utilize xylose.
It has previously been suggested that xylose is taken up by both high- and low-affinity systems of glucose transporters (Fig. 8.3), but the uptake is increased in the presence of low glucose concentrations (Olofsson et al. 2008).
Simplified scheme of sugar transport and metabolism in S. cerevisiae (Olofsson et al. 2008). 1. Low- and intermediate-affinity hexose transporters. 2. High-affinity hexose transporters. Abbreviations: PPP pentose phosphate pathway, XR xylose reductase, XDH xylitol dehydrogenase, XK xylulokinase, GK glucokinase, PGI phosphoglucose isomerase, PFK phosphofructokinase, AD aldolase, TPI triose phosphate isomerase, GDH glyceraldehyde-3-P dehydrogenase, GPD glycerol-3-P dehydrogenase, GPP glycerol-3-phosphatase, PDC pyruvate decarboxylase, ALD acetaldehyde dehydrogenase, ADH alcohol dehydrogenase
One of the first attempts to get S. cerevisiae utilizing xylose was based on the fact that it can ferment xylulose. For this reason, it was thought that expressing xylose-isomerase (XI) would be a promising modification, and the strain TBM3050, carrying XI from Thermus thermophiles was produced (strain TMB3050). As the yeast Pichia stipitis was known for its ability to ferment xylose with minimal amounts of xylitol as coproduct, it was selected for the first successful xylose-utilizing of S. cerevisiae, which was produced by expressing P. stipitis genes encoding XR and XDH. The ethanol yields were however low, and xylitol was also produced, which was attributed to the need of a high activity of both XR and XDH.
A complete description of xylose consumption process by different S. cerevisiae strains, including ethanol and xylitol yields anaerobic and oxygen-limited batch cultures which can be found in Hahn-Hägerdal et al. (2007), and in Chu and Lee (2007). Although much work has been dedicated to lab scale study on pentose-fermenting S. cerevisiae, there are still some issues to be addressed, especially those related with industrial uses, as pointed out by Matsushika et al. (2009).
A review of more recent work on modified S. cerevisiae for ethanol production from xylose or pentoses in general is presented below (Table 8.2).
Recently, Liu and Hu (2010) reported on a S. cerevisiae strain via combined approaches of recombinant DNA technology, chemical mutagenesis, and evolutionary adaptation for an efficient xylose utilization and ethanol fermentation. A haploid derivative of an industrial-fermenting strain was first engineered to express the genes from P. stipitis encoding XR and XDH, and XK. Then, the recombinant strain was submitted to ethyl-methanesulfonate mutagenesis followed by adaptive evolution, resulting in a single isolate with improved xylose utilization characteristics.
Although large research efforts have been devoted toward modifying S. cerevisiae to allow xylose consumption and ethanol production, a number of issues have limited the success of the process, including poor xylose uptake, cofactor imbalance, insufficiency in the pentose phosphate pathway, deregulation of the ethanologenic enzymes, and specially the regulation of metabolism in the eukaryotic yeasts, less known than that of bacteria (Hahn-Hägerdal et al. 2007). Efficient utilization of xylose appears to require complex global changes in cellular processes (Liu and Hu 2010). The need for novel tools and approaches to overcome the major remaining difficulties, like engineering simultaneous, exogenous sugar metabolism has also been emphasized. Beyond catabolic pathways, the focus must shift toward non-traditional aspects of cellular engineering such as host molecular transport capability, catabolite sensing, and stress response mechanisms (Young et al. 2010).
Direct evolution is one of the strategies to improve the performance of S. cerevisiae for utilizing all sugars derived from pretreatment of LCM. Using pine hydrolysates, Hawkins and Doran-Peterson (2011) proposed a combination of adaptation by inoculation first into a solids loading of 7 % `w/v for 24 h, followed by a 10 % v/v inoculum into 17.5 % w/v solids. Under such conditions, the final strain (AJP50) produced ethanol at more than 80 % of the maximum theoretical yield after 72 h of fermentation, and more than 90 % of the maximum theoretical yield after 120 h of fermentation. This improvement in comparison with results by the starting industrial strain (XR122n) was attributed to the ability of AJP50 to rapidly convert furfural and hydroxymethylfurfural to their less-toxic alcohol derivatives.
In a recent work, Zhou et al. (2012) described the metabolic engineering of a S. cerevisiae strain, including overexpression of the Piromyces xylose isomerase gene (XYLA), P. stipitis xylulose kinase (XYL3) and genes of the non-oxidative pentose phosphate pathway (PPP). The engineered strain, named H131-A3, was used to initialize a three-stage process of evolutionary engineering, first through aerobic and anaerobic sequential batch cultivation followed by growth in a xylose-limited chemostat. These authors claimed that the evolved strain H131-A3-ALCS produced the best result on xylose utilization and ethanol production by S. cerevisiae reported to-date.
Concerning the application of recombinant S. cerevisiae on lignocellulose hydrolysates, several strategies have also been applied. For example, the fermentability of ammonia-pretreated corn stover detoxified hydrolysate was significantly improved by using immobilized cells of recombinant S. cerevisiae in Ca-alginate (Zhao and Xia 2010). The composition of the detoxified and concentrated hydrolysate included 72 g/L xylose and 14.3 g/L arabinose and the ethanol yield based on fermentable sugars was 0.41 g/g within 72 h in batch fermentation.
Strain S. cerevisiae Y5 is a newly lab-developed patented microorganism. The strain has the ability to metabolize furfural, tolerate fermentation inhibitors, and efficiently metabolize glucose to produce ethanol. It was used for ethanol production from enzymatic hydrolysates of non-detoxified steam-exploded corn stover, with and without a nitrogen source, and decreasing inoculum size (Li et al. 2011). Results showed that ethanol yields as high as 94.5 % of the theoretical yield were obtained after 24 h, with an inoculum size of 10 % (v/v) and nitrogen source (corn steep liquor) of 40 mL/L.
8.5 Other Modified Microorganisms
In addition to E. coli and S. cerevisiae, a number of other microorganisms have also been extensively studied for ethanol production from lignocellulosic materials. Some examples of these recent studies are summarized in Table 8.3.
Zymomonas mobilis is a bacterium that can ferment certain sugars to ethanol via the Entner–Doudoroff (ED), glyceraldehyde-3-phosphate-to-pyruvate (GP), and pyruvate-to-ethanol (PE) pathways. It has been proposed as a promising alternative to other fermenting microorganisms as ethanol yields as high as 97 % of the theoretical maximum along with high production rates have been reported (Hayashi et al. 2011). This advantageous behavior is a consequence of a favorable energy balance from the ED pathway, compared to Embden–Meyerhof–Parnas pathway used by both E. coli and S. cerevisiae (Gírio et al. 2010).
However, Z. mobilis exhibits a relatively low tolerance to inhibitors generated during the conversion process. This fact, along with the lack of the complete pentose metabolism pathway necessary for fermentation of lignocellulosic hydrolysates (Davis et al. 2005), are the reasons for several engineering works on this microorganism trying to improve ethanol yields. The introduction of the xylose metabolism pathway has been one of the main strategies, although with limited results (Chandel et al. 2011). In another study, a threefold higher ethanol concentration was produced when a fragment from Enterobacter cloacae conferring cellulase activity was cloned in Z. mobilis (Vasan et al. 2011) in comparison with results with the native E. cloacae.
Bacillus subtilis is also a promising microorganism known for producing ethanol from lignocellulose, as it has xylose isomerase and xylulokinase for metabolizing xylose. However, the wild type lacks a specific xylose transporter and hence it is not feasible to use xylose as a sole carbon source. In an attempt to improve xylose transportation, the arabinose: H+ symporter, AraE protein from B. subtilis was expressed in B. subtilis 168 (Park et al. 2012). B. subtilis has also been engineered to produce ethanol and depolymerize cellulose (Romero et al. 2007; You et al. 2011).
Clostridium acetobutylicum is an anaerobic bacterium that is known for its excellent capacity to produce ABE (acetone, butanol, and ethanol) solvents. Nevertheless, it shows inefficient pentose consumption when fermenting sugar mixtures. As a strategy to overcome this fact, a predicted glcG gene, encoding enzyme II of the d-glucose phosphoenolpyruvate-dependent phosphotransferase system (PTS), was first disrupted in the ABE-producing model strain C. acetobutylicum ATCC 824, which resulted in a greatly improved d-xylose and l-arabinose consumption in the presence of d-glucose. Further overexpression of the xylose pathway resulted in an engineered strain (824glcG-TBA) that was able to efficiently conferment mixtures of d-glucose, d-xylose, and l-arabinose, with better results than the results tied to the wild-type strain (Xiao et al. 2011).
Thermoanaerobacterium saccharolyticum is a potential catalyst for lignocellulose conversion that can naturally hydrolyze xylan and ferment all monosaccharides and disaccharides found in lignocellulosic materials. Encoding the urease gene resulted in one of the highest titers reported in this microorganism. In addition, it is evident that the use of urea instead of ammonium salts can be advantageous here because of the lower cost (Shaw et al. 2012). Ethanologenic strains of T. saccharolyticum have been obtained through metabolical engineering and adaptive evolution and 37 g/L of ethanol were produced using simultaneous saccharification and fermentation from Avicel (Shaw et al. 2008). Lynd and coworkers from Dartmouth College claim that this is one of the best microorganism to develop CBP.
Klebsiella pneumoniae is a Gram-negative bacterium that has been described as capable of fermenting glycerol, a by-product of biodiesel. Due to the large amount of glycerol generated (equivalent to approximately 10 % of biodiesel produced) and its need to be treated before discharge, a great interest has emerged regarding the use of glycerol as raw material for the production of industrially valuable materials like ethanol. The estimated cost of ethanol produced from glycerol, considering both the feedstock demand and operational costs, is 40 % less than the estimated cost of production from corn-derived sugars (which is seen as an additional advantage). Oh et al. (2011) have isolated a γ-irradiated mutant strain of K. pneumoniae (GEM167) exhibiting high production of ethanol from glycerol. Ethanol production level was improved to 25.0 g/L upon overexpression of Z. mobilispdc and adhII genes encoding pyruvate decarboxylase (Pdc) and aldehyde dehydrogenase (Adh), respectively in the mutant strain GEM167.
8.6 Conclusions
The conversion of sugars contained in lignocellulosic materials into ethanol is a promising way for partially substituting fossil fuels. As a consequence of the pretreatment step in the bioconversion process, a mixture of hexose and pentose sugars, together with sugar degradation and inhibitory products, is produced and this must be considered in the subsequent hydrolysis and fermentation steps. No matter what the process configuration is, the corresponding microorganism should produce ethanol with both high yields and productivity, with low medium and operational requirements, and should at the same time be tolerant of ethanol and resistant to inhibitors. Unfortunately, such an ideal ethanologenic microorganism is not available at the moment. Nevertheless, a great deal of research is being devoted to modify, adapt, and engineer yeasts and bacteria for such a purpose. Recombinant DNA technology and evolutionary engineering techniques, direct evolution, introduction of foreign genes, elimination of competitive pathways, disruption of byproducts, and formation are some of the many strategies being assayed that should lead to ethanol production from lignocellulosic materials at an industrial scale in the near future.
References
Alterthum F, Ingram LO (1989) Efficient ethanol production from glucose, lactose, and xylose by recombinant Escherichia coli. Appl Environ Microbiol 55:1943–1948
Bailey JE (1991) Toward a science of metabolic engineering. Science 252:1668–1675
Chandel AK, Chandrasekhar G, Radhika K, Ravinder R, Ravindra P (2011) Bioconversion of pentose sugars into ethanol: a review and future directions. Biotechnol Mol Biol Rev 6:8–20
Chu BCH, Lee H (2007) Genetic improvement of Saccharomyces cerevisiae for xylose fermentation. Biotechnol Adv 25:425–441
Davis L, Jeon YJ, Svenson C, Rogers P, Pearce J, Peiris P (2005) Evaluation of wheat stillage for ethanol production by recombinant Zymomonas mobilis. Biomass Bioenergy 29:49–59
Dellomonaco C, Clomburg JM, Miller EN, Gonzalez R (2011) Engineered reversal of the ß-oxidation cycle for the synthesis of fuels and chemicals. Nature 476:355–359
Dien BS, Hespell RB, Ingram LO, Bothast RJ (1997) Conversion of corn milling fibrous co-products into ethanol by recombinant Escherichia coli strains KO11 and SL40. World J Microbiol Biotech 13:619–625
Durnin G, Clomburg J, Yeates Z, Alvarez PJJ, Zygourakis K, Campbell P, Gonzalez R (2009) Understanding and harnessing the microaerobic metabolism of glycerol in Escherichia coli. Biotechnol Bioeng 103:148–161
Edwards MC, Henriksen ED, Yomano LP, Gardner BC, Sharma LN, Ingram LO, Peterson JD (2011) Addition of genes for cellobiase and pectinolytic activity in Escherichia coli for fuel ethanol production from pectin-rich lignocellulosic biomass. Appl Environ Microbiol 77:5184–5191
Fernández-Sandoval MT, Huerta-Beristain G, Trujillo-Martinez B, Bustos P, González V, Bolivar F, Gosset G, Martinez A (2012) Laboratory metabolic evolution improves acetate tolerance and growth on acetate of ethanologenic Escherichia coli under non aerated conditions in glucose-mineral medium. Appl Microbiol Biotechnol 96:1291–1300
Geddes CC, Mullinnix MT, Nieves IU, Peterson JJ, Hoffman RW, York SW, Yomano LP, Miller EN, Shanmugam KT, Ingram LO (2011) Simplified process for ethanol production from sugarcane bagasse using hydrolysate-resistant Escherichia coli strain MM160. Bioresour Technol 102(3):2702–2711
Gírio FM, Fonseca C, Carvalheiro F, Duarte LC, Marques S, Bogel-Łukasik R (2010) Hemicelluloses for fuel ethanol: a review. Bioresour Technol 101:4775–4800
Hahn-Hägerdal B, Karhumaa K, Jeppsson M, Gorwa-Grauslund MF (2007) Metabolic engineering for pentose utilization. Adv Biochem Eng Biotechnol 108:147–177
Hasunuma T, Kondo A (2012) Development of yeast cell factories for consolidated bioprocessing of lignocellulose to bioethanol through cell surface engineering. Biotechnol Adv 30:1207–1218
Hawkins GM, Doran-Peterson J (2011) A strain of Saccharomyces cerevisiae evolved for fermentation of lignocellulosic biomass displays improved growth and fermentative ability in high solids concentrations and in the presence of inhibitory compounds. Biotechnol Biofuels 4:49
Hayashi T, Furuta Y, Furukawa K (2011) Respiration-deficient mutants of Zymomonas mobilis show improved growth and ethanol fermentation under aerobic and high temperature conditions. J Biosci Bioeng 111:414–419
Huffer S, Roche CM, Blanch HW, Clark DS (2012) Escherichia coli for biofuel production: bridging the gap from promise to practice. Trends Biotechnol 30:538–545
Ingram LO, Alterthum F, Conway T (1991) Ethanol production by Escherichia coli strains co-expressing Zymomonas pdc and adh genes. USA Patent number 5(000):000
Ingram LO, Conway T (1998) Expression of different levels of ethanologenic enzymes from Zymomonas mobilis in recombinant strains of Escherichia coli. Appl Environ Microbiol 54:397–404
Ingram LO, Aldrich HC, Borges ACC, Causey TB, Martinez A, Morales F, Saleh A, Underwood SA, Yomano LP, York SW, Zaldivar J, Zhou S (1999) Enteric bacterial catalyst for fuel ethanol production. Biotechnol Prog 15:855–866
Jarboe LR, Grabar TB, Yomano LP, Shanmugan KT, Ingram LO (2007) Development of ethanologenic bacteria. Adv Biochem Eng Biotechnol 108:237–261
Jeffries TW (2006) Engineering yeasts for xylose metabolism. Cur Opinion Biotechnol 17:320–326
Jin M, Balan V, Gunawan C, Dale BE (2012) Quantitatively understanding reduced xylose fermentation performance in AFEX™ treated corn stover hydrolysate using Saccharomyces cerevisiae 424A (LNH-ST) and Escherichia coli KO11. Bioresour Technol 111:294–300
Lee SM, Jellison T, Alper HS (2012) Directed evolution of xylose isomerase for improved xylose catabolism and fermentation in the yeast Saccharomyces cerevisiae. Appl Environ Microbiol 78:5708–5716
Leite AR, Guimaraes WV, Fernandes de Araújo E, Silva DO (2000) Fermentation of sweet whey by recombinant Escherichia coli KO11. Braz J Microbiol 31:212–215
Li Y, Gao K, Tian S, Zhang S, Yang X (2011) Evaluation of Saccharomyces cerevisiae Y5 for ethanol production from enzymatic hydrolysate of non-detoxified steam-exploded corn stover. Bioresour Technol 102:10548–10552
Liu E, Hu Y (2010) Construction of a xylose-fermenting Saccharomyces cerevisiae strain by combined approaches of genetic engineering, chemical mutagenesis and evolutionary adaptation. Biochem Eng J 48:204–210
Madhavan A, Tamalampudi S, Srivastava A, Fukuda H, Bisaria VS, Kondo A (2009) Alcoholic fermentation of xylose and mixed sugars using recombinant Saccharomyces cerevisiae engineered for xylose utilization. Appl Microbiol Biotechnol 82:1037–1047
Martínez A, Grabar TB, Shanmugam KT, Yomano LP, York SW, Ingram LO (2007) Low salt medium for lactate and ethanol production by recombinant Escherichia coli B. Biotechnol Lett 29:397–404
Matsushika A, Inoue H, Kodaki T, Sawayama S (2009) Ethanol production from xylose in engineered Saccharomyces cerevisiae strains: current state and perspectives. Appl Microbiol Biotechnol 84:37–53
Miller EN, Jarboe LR, Yomano LP, York SW, Shanmugam KT, Ingram LO (2009) Silencing of NADPH-dependent oxidoreductases (yqhd and dkga) in furfural-resistant ethanologenic Escherichia coli. Appl Environ Microbiol 75(13):4315–4323
Muñoz-Gutiérrez I, Oropeza R, Gosset G, Martinez A (2012) Cell surface display of a ß-glucosidase employing the type V secretion system on ethanologenic Escherichia coli for the fermentation of cellobiose to ethanol. J Ind Microbiol Biotechnol 39:1141–1152
Nieves IU, Geddes CC, Mullinnix MT, Hoffman RW, Tong Z, Castro E, Shanmugam KT, Ingram LO (2011) Injection of air into the headspace improves fermentation of phosphoric acid pretreated sugarcane bagasse by Escherichia coli MM170. Bioresour Technol 102:6959–6965
Oh BR, Seo JW, Heo SY, Hong WK, Luo LH, Joe M, Park DH, Kim CH (2011) Efficient production of ethanol from crude glycerol by a Klebsiella pneumonia mutant strain. Bioresour Technol 102:3918–3922
Orencio-Trejo M, Utrilla J, Fernández-Sandoval MT, Huerta-Beristain G, Gosset G, Martinez A (2010) Engineering the Escherichia coli fermentative metabolism. Adv Biochem Eng Biotechnol 121:71–107
Olofsson K, Bertilsson M, Lidén G (2008) A short review on SSF—an interesting process option for ethanol production from lignocellulosic feedstocks. Biotechnol Biofuels 1:7
Parachin NS, Bergdahl B, van Niel EWJ, Gorwa-Grauslund MF (2011) Kinetic modelling reveals current limitations in the production of ethanol from xylose by recombinant Saccharomyces cerevisiae. Metab Eng 13:508–517
Park YC, Jun SY, Seo JH (2012) Construction and characterization of recombinant Bacillus subtilis JY123 able to transport xylose efficiently. J Biotechnol 161:402–406
Peralta-Yahya PP, Zhang F, del Cardayre SB, Keasling JD (2012) Microbial engineering for the production of advanced biofuels. Nature 488:320–328
Qureshi N, Dien BS, Nichols NN, Saha BC, Cotta MA (2006) Genetically engineered Escherichia coli for ethanol production from xylose. Substrate and product inhibition and kinetic parameters. Trans IChemE C, 84(C2):114–122
Rodríguez-Moya M, Gonzalez R (2010) Systems biology approaches for the microbial production of biofuels. Biofuels 1:291–310
Romero S, Merino E, Bolívar F, Gosset G, Martinez A (2007) Metabolic engineering of Bacillus subtilis for ethanol production: Lactate dehydrogenase plays a key role in the fermentative metabolism. Appl Environ Microbiol 73:5190–5198
Runquist D, Hahn-Hagerdal B, Bettiga M (2010) Increased ethanol productivity in xylose-utilizing Saccharomyces cerevisiae via a randomly mutagenized xylose reductase. Appl Environ Microbiol 76:7796–7802
Ryu S, Karim MN (2011) A whole cell biocatalyst for cellulosic ethanol production from dilute acid-pretreated corn stover hydrolyzates. Appl Microbiol Biotechnol 91:529–542
Saha BC, Nichols NN, Cotta MA (2011) Ethanol production from wheat straw by recombinant Escherichia coli strain FBR5 at high solid loading. Bioresour Technol 102:10892–10897
Sakamoto T, Hasunuma T, Hori Y, Yamada R, Kondo A (2012) Direct ethanol production from hemicellulosic materials of rice straw by use of an engineered yeast strain codisplaying three types of hemicellulolytic enzymes on the surface of xylose-utilizing Saccharomyces cerevisiae cells. J Biotechnol 158:203–210
Shaw AJ, Podkaminer KK, Desai SG, Bardsley JS, Rogers SR, Thorne PG, Hogsett DA, Lynd LR (2008) Metabolic engineering of a thermophilic bacterium to produce ethanol at high yield. Proc Nat Acad Sci USA 105(37):13769–13774
Shaw AJ, Covalla SF, Miller BB, Firliet BT, Hogsett DA, Herring CD (2012) Urease expression in a Thermoanaerobacterium saccharolyticum ethanologen allows high titer ethanol production. Metabolic Eng 14:528–532
Vasan PT, Piriya PS, Prabhu PIG, Vennison SJ (2011) Cellulosic ethanol production by Zymomonas mobilis harboring an endoglucanase gene from Enterobacter cloacae. Bioresource Technol 102:2585–2589
Vinuselvi P, Lee SK (2012) Engineered Escherichia coli capable of co-utilization of cellobiose and xylose. Enzyme Microb Technol 50:1–4
Wargacki AJ, Leonard E, Win MN, Regitsky DD, Santos CNS, Kim PB, Cooper SR, Raisner RM, Herman A, Sivitz AB, Lakshmanaswamy A, Kashiyama Y, Baker D, Yoshikuni Y (2012) An engineered microbial platform for direct biofuel production from brown macroalgae. Science 335:308–313
Wisselink HW, Toirkens MJ, Wu Q, Pronk JT, van Maris AJA (2009) Novel evolutionary engineering approach for accelerated utilization of glucose, xylose, and arabinose mixtures by engineered Saccharomyces cerevisiae strains. Appl Environ Microbiol 75:907–914
Xiao H, Gu Y, Ning Y, Yang Y, Mitchell WJ, Jiang W, Yang S (2011) Confirmation and elimination of xylose metabolism bottlenecks in glucose phosphoenolpyruvate-dependent phosphotransferase system-deficient Clostridium acetobutylicum for simultaneous utilization of glucose, xylose, and arabinose. Appl Environ Microbiol 77:7886–7895
Yasuda M, Miura A, Shiragami T, Matsumoto J, Kamei I, Ishii Y, Ohta K (2012) Ethanol production from non-pretreated napiergrass through a simultaneous saccharification and fermentation process followed by a pentose fermentation with Escherichia coli KO11. J Biosci Bioeng 114:188–192
Yomano LP, York SW, Ingram LO (1998) Isolation and characterization of ethanol-tolerant mutants of Escherichia coli KO11 for fuel ethanol production. J Ind Microbiol Biotechnol 20:132–138
Yomano LP, York SW, Shanmugam KT, Ingram LO (2009) Deletion of methlylglyoxal synthase gene (mgsA) increased sugar co-metabolism in ethanol-producing Escherichia coli. Biotechnol Lett 31:1389–1398
You C, Zhang XZ, Sathitsuksanoh N, Lynd LR, Percival Zhang Y-H (2011) Enhanced microbial utilization of recalcitrant cellulose by an ex vivo cellulosome microbe complex. Appl Environ Microbiol 78:1437–1444
Young E, Lee SM, Halper H (2010) Optimizing pentose utilization in yeast: the need for novel tools and approaches. Biotechnol Biofuels 3:24
Zhao J, Xia L (2010) Ethanol production from corn stover hemicellulosic hydrolysate using immobilized recombinant yeast cells. Biochem Eng J 49:28–32
Zhou H, Cheng J, Wanga BL, Fink GR, Stephanopoulos G (2012) Xylose isomerase overexpression along with engineering of the pentose phosphate pathway and evolutionary engineering enable rapid xylose utilization and ethanol production by Saccharomyces cerevisiae. Metab Eng 14:611–622
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Castro, E. (2013). Other Ethanologenic Microorganisms. In: Faraco, V. (eds) Lignocellulose Conversion. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-37861-4_8
Download citation
DOI: https://doi.org/10.1007/978-3-642-37861-4_8
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-37860-7
Online ISBN: 978-3-642-37861-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)