Abstract
In natural conditions, mycorrhizal fungi are surrounded by complex microbial communities, which may trigger various responses, from enhancement of the establishment of mycorrhizal symbiosis to mycelial growth inhibition or cell death. The symbiosis between mycorrhizal soil fungi and higher plants takes advantage of active collaboration with specific helper bacteria. Thus, a symbiosis so far thought of involving two components could be the result of the interaction among at least three different partners. This chapter focuses on the relationship between edible ectomycorrhizal mushrooms and soil bacteria, in particular nitrogen-fixing bacteria associated with Tuber species. The ability of these bacteria to modify nutrient availability during the fructification phase is very important to truffle development. This chapter will also discuss perspectives on the beneficial use of ectomycorrhizal symbiosis with nitrogen-fixing bacteria to develop predictive models that could be used to improve the mycorrhization processes with the further aim of obtaining plants infected with Tuber magnatum Pico, the most economically important truffle species that remains difficult to cultivate.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Arbuscular Mycorrhizal Fungus
- Mycorrhizal Fungus
- Fruit Body
- Ectomycorrhizal Fungus
- Mycorrhizal Symbiosis
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Forests function in a capacity that improves environmental quality and water and carbon stocks and are critical to the conservation of biodiversity. To this end, the application of techniques for controlled mycorrhization may be important for improving reforestation strategies. Indeed, for efficient nutrient uptake, most plant species need to be associated with mycorrhizal fungi that supply minerals, increase growth, and confer stress resistance (Singh et al. 2011). The management of these symbioses in natural and agricultural environments is of significant ecological and economic importance.
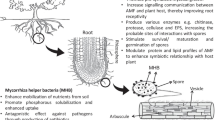
The establishment of a functional symbiosis between plants and microorganisms affects the chemical–biological nature of rhizosphere. Important examples of this include nitrogen-fixing symbiosis between bacteria and leguminous plants and the symbiosis between mycorrhizal soil fungi and numerous plant species. The microbial populations that colonize roots often differ between plants involved, in a symbiotic relationship with an active molecular dialogue between partners. This crosstalk between plant and microbe can result in response mechanisms capable of attracting additional microbial species that are not directly involved in the symbiosis. Recently, some of these species, so far considered alien to the symbiosis, have been identified as important components of the ectomycorrhizal system. The establishment of mycorrhizal symbiosis alters the composition and amount of root exudates, thus changing the microbial equilibrium of the rhizosphere (Duponnois and Garbaye 1992; Barea et al. 1997; Frey-Klett and Garbaye 2005; Frey-Klett et al. 2007). This effect, known as the “mycorrhizosphere effect,” seems to favor the occurrence of particular bacteria involved in the mycorrhization process, called “mycorrhizal helper bacteria” (MHB) (Dakora 2003; Frey-Klett et al. 2007), as well as the growth of bacteria, called “mycorrhizal associated bacteria,” whose role is still unclear (Assigbetse et al. 2005; de Boer et al. 2005).
The bacterial populations present in the fruiting bodies of several species of mycorrhizal fungi exceed numerically those usually found directly in soil for the same bacterial species, whereas the diversity of species observed seems to be greatly reduced (Gazzanelli et al. 1999; Sbrana et al. 2002). These MHB populations have different roles and undoubtedly promote the establishment of mycorrhizal symbiosis and stimulate the development of mycelium resulting in a greater surface contact between the plant root and the fungus mycelium; this is easily verifiable through bacteria and fungi coculture (Founoune et al. 2002; Deveau et al. 2007). While both ectomycorrhizal (Garbaye 1994) and endomycorrhizal (Meyer and Linderman 1986) fungi can interact with different bacterial species, we will focus on the interactions relating ectomycorrhizal fungi.
In the past several years, the potential ability of bacteria associated with ectomycorrhizas to fix atmospheric nitrogen has been suggested (Frey-Klett et al. 2007). This hypothesis originates from the observation that ectomycorrhizas are particularly frequent on plants typical of land ecosystems characterized by a deficiency in combined nitrogen. Based on the hypothesis of the presence of nitrogen-fixing bacteria, previous research allowed for the detection of the occurrence of genes directly involved with nitrogen fixation process.
Several studies have found that potential nitrogen-fixing bacteria can be associated with ectomycorrhizal and arbuscular mycorrhizal fungi (Frey-Klett et al. 2007) and that the highly conserved nifH gene, which encodes for the nitrogenase reductase, is present in Pinus sylvestris and Pinus nigra ectomycorrhizas (Timonen and Hurek 2006; Izumi et al. 2006a).
These experiments suggest a real possibility that bacteria present in mycorrhizal tissues contribute to the nutritional needs of both fungi, and consequently plants, providing them with available nitrogen derived from atmospheric nitrogen (N2). The recent detection of nitrogen-fixing bacteria in fruit bodies of Tuber borchii and Tuber magnatum and in association with soil and ectomycorrhizal fungi led to the hypothesis that nitrogen fixation activity may play an important role in fungal growth and in the development of the ascocarp (Rainey et al. 1990; Barbieri et al. 2005a, 2007; Frey-Klett et al. 2007). However, in order to unequivocally demonstrate this role, more targeted investigations are needed.
In this chapter, we first introduce the major biological characteristics of ectomycorrhizas followed by a description of different types of interaction taking place between bacteria and mycorrhizal fungi. Promotion of plant growth, release of active molecules, and nutritional exchange strategies are discussed within the establishment of plant–bacteria–fungus interactions. Particular attention is focused on interactions between bacteria and ectomycorrhizal fungi belonging to the Tuber genus with established or potentially synergistic properties important to its fruiting and life cycle.
2 Defining the Mycorrhizosphere
The influence of plant assimilates on microbial communities has been defined in relation to the rhizosphere, the narrow zone of soil surrounding living roots. The rhizosphere is characterized by increased microbial activity stimulated by the exudation of organic substances from the root (Grayston et al. 1997). However, since plant roots in natural and semi-natural ecosystems are commonly mycorrhized, the rhizosphere concept has been extended to include the fungal component of the symbiosis, resulting in the term “mycorrhizosphere.” The mycorrhizosphere is the zone influenced by both the root and the mycorrhizal fungus and includes the more specific term “hyphosphere,” which refers only to the zone surrounding individual fungal hyphae. Since mycorrhizas and fungal hyphae are more or less ubiquitous in natural soils, it could be argued that all soil could be included in the term “mycorrhizosphere” (Johansson et al. 2004).
2.1 Occurrence of Ectomycorrhizas
Fossil records indicate that ectomycorrhizal associations emerged at least 50 million years ago (Mya). For example, Hydnangiaceae appeared before 40 Mya according to Ryberg and Matheny (2012), although there is molecular evidence that this emergence dated to more than 180 Mya (Martin et al. 2007). In ectomycorrhizal associations, hyphae of fungal symbionts produce extensive nets of mycelium that extends beyond exploratory roots of plants. The mycelium acquires soil minerals through solubilization, particularly of phosphorus and nitrogen, and exchanges these nutrients with their host plants, which may allocate as much as 10–20 % of their photosynthate to the mycobiont. Ectomycorrhizal (EM) fungi produce a hyphal mantle during the colonization process that tightly covers the root tip, while epidermal (and in some cases also cortical) cells become separated by the development of labyrinth-like hyphae (the Hartig net), which increases the surface contact area with root cells. Thus, in ectomycorrhizas, hyphae remain extracellular, inducing important changes to root morphogenesis, whereas their presence only leads to thin modifications in epidermal or cortical cells (Bonfante 2001).
EM fungi alter the physical, chemical, and microbiological characteristics of the surrounding soil and create a special environment called mycorrhizosphere in which the microbial communities differ from those in the rhizosphere and in other portions of the soil (Izumi et al. 2006b). There is considerable evidence that EM fungi influence the taxonomic composition of communities of soil bacteria, although this effect varies among mycorrhizas of different fungi species and between different root tips colonized with the same species (Burke et al. 2008; Izumi et al. 2008; Kataoka et al. 2008). For example, Brooks and colleagues have demonstrated that EM fungal hyphae may control organic phosphorus resources in soil by selecting against soil bacteria with higher abilities to mobilize phosphorus from organic compounds. This interaction could result in complementary and synergistic modes of action providing a sustained supply of phosphorus from organic and inorganic fonts to the EM host (Brooks et al. 2011).
3 Interactions Between Ectomycorrhizal Fungi and Associated Bacteria
Mycorrhizal symbiosis has long been considered as a bipartite relationship between plant roots and mycorrhizal fungi. However, in natural conditions, ectomycorrhizal fungi physically and metabolically interact with a wide community of soil bacteria with potential unfavorable, neutral, and beneficial consequences (Bowen and Theodorou 1979; Barea et al. 2002; Johansson et al. 2004; Frey-Klett et al. 2007, 2011; Bonfante and Anca 2009).
Bowen and Theodorou (1979) and then Garbaye and Bowen (1989) demonstrated that the rhizosphere microflora could have positive or negative impacts on the mycorrhizal symbiosis, depending on particular bacterial isolates involved. Garbaye and Duponnois (1992) showed that pure bacterial strains such as Pseudomonas spp. and Bacillus spp. stimulated the Pseudotsuga menziesii–Laccaria laccata symbiosis. This was the experimental evidence for the so-called helper effect of those bacteria named MHB. The MHB are not plant-specific, but are clearly selective about the fungal species, so they can be defined as fungus-specific (Garbaye 1994). They exist in arbuscular and ectomycorrhizal systems and now are currently the most investigated group among bacteria interacting with mycorrhizas. Some research has proposed that the ability of Pseudomonas spp. to utilize trehalose may be key for bacterial growth in association with the edible ectomycorrhizal fungus Cantharellus cibarius without causing fungal cell damage (Rangel-Castro et al. 2002). Some MHB isolated from Lactarius deliciosus are able to enhance establishment of mycorrhization with Pinus pinea and Pinus pinaster (Barriuso et al. 2008). More recently, Wu et al. evaluated positively the effects of the co-inoculum between the ectomycorrhizal fungus Boletus edulis and the MHB Bacillus cereus (HB12 or HB59) on the growth and nutrient uptake of Pinus thunbergii (Wu et al. 2011). Experimental evidence suggests that a release of active diffusible molecules occurs prior to physical contact between bacteria and/or mycorrhizal fungi, which is important for the establishment of their interactions. Recent data show the production of volatile organic compounds (VOCs) from all the members of the symbiosis, important for inter- and intra-organism communications (Splivallo et al. 2007; Tarkka and Piechulla 2007). Studies performed on the ectomycorrhizal fungus Laccaria bicolor S238N and the MHB strain Pseudomonas fluorescens BBc6R8 have indicated that both fungal trehalose and bacterial thiamine play a key role in the mutually beneficial interactions of two organisms (Deveau et al. 2010). The main significant functions of MHB are in plant protection against root pathogens, nutrient mobilization from soil minerals and fixation of atmospheric nitrogen in forms available to plants, fungi, and microbes (Frey-Klett et al. 2007). In addition, they may have an active role in ascocarp decomposition and spore dispersal.
3.1 Plant Protection
Mycorrhiza-associated bacteria contribute, together with the fungal symbiont, protection against root pathogens. For example, there is now abundant evidence that some Streptomyces species colonize the rhizospheres of plant roots and even plant tissues (Coombs and Franco 2003), and it has been suggested that antibiotic production by the streptomycete may protect the host plants against phytopathogens (Challis and Hopwood 2003). More recently, Tarkka et al. (2008) illustrated the streptomyces mechanism that induces plant defenses and facilitates symbiosis formation. Moreover, Frey-Klett et al. (2005) revealed a significantly higher proportion of fluorescent Pseudomonads inhibiting the growth of root-pathogenic fungi belonging to the genera Rhizoctonia, Fusarium, Phytophthora, and Heterobasidion in the Douglas-fir L. bicolor ectomycorrhizas than in the surrounding bulk soil. These findings suggest that MHB could have evolved selective mechanisms of interaction with their microbial surroundings, having neutral or positive effects on their host mycorrhizal associations but negative effects on the root pathogens that might threaten their very habitat.
3.2 Nutrient Mobilization
Recent findings suggest that ectomycorrhiza-associated bacteria complement the roles of the external mycelium by mobilizing nutrients from minerals. In fact, these bacteria can solubilize calcium phosphate, rock phosphate, iron phosphate, and aluminium phosphate by secretion of organic acids. For example, different experiments revealed a higher proportion of culturable bacteria able to release iron from the biotite, in Quercus sessiliflora–Scleroderma citrinum ectomycorrhizas than in the bulk soil (Uroz et al. 2007). Other authors demonstrated that the solubilization of rock phosphate is enhanced by formation of mixed biofilms between phosphate-solubilizing saprotrophic fungi and Bradyrhizobium elkanii strain (Jayasinghearachchi and Seneviratne 2005).
3.3 Nitrogen Fixation
Nitrogen, a critical component of many biomolecules, is essential for growth and development of all organisms. Although roughly 78 % of the Earth’s atmosphere is composed by nitrogen, it is in a form (N2) that is not biologically accessible. However, the ability to fix atmospheric nitrogen via the nitrogenase enzyme complex has evolved in some bacteria. Many eukaryotic organisms are able to obtain fixed nitrogen through their symbiotic interactions with nitrogen-fixing prokaryotes. In the last decade, special attention has been paid to potential nitrogen fixation by bacteria associated with ectomycorrhizas. Several studies have found that potential nitrogen-fixing bacteria can be associated with ectomycorrhizal and arbuscular mycorrhizal fungi (Frey-Klett et al. 2007; Kretzer et al. 2009). The presence of nitrogen-fixing bacteria in these diverse ectomycorrhizal types clearly supports their potential for improving plant nutrition. Using molecular methods, several authors found DNA sequences of the nifH gene (encoding nitrogenase) in P. sylvestris–S. bovinus and P. nigra–Suillus variegatus ectomycorrhizas (Timonen and Hurek 2006; Izumi et al. 2006a). Furthermore, Barbieri et al. showed the predominance of α-Proteobacteria represented by Sinorhizobium/Ensifer and Rhizobium/Agrobacterium groups as well as by nitrogen-fixing Bradyrhizobium spp. in T. borchii and T. magnatum ascocarps (Barbieri et al. 2005a, 2007). More recently, Sphingobium sp. strain TMG 022C was isolated from the ascocarp of white truffle T. magnatum. This strain is capable of growing in nitrogen-depleted conditions and may thus be important in mycelial nutrition and ascocarp decomposition (Pavić et al. 2011). These results suggest that diazotrophic bacteria present in ectomycorrhizal tissues contribute to nitrogen input in forest ecosystems by directly providing nitrogen of atmospheric origin to the two partners of the symbiosis, thus bypassing classic nitrogen mineralization pathways.
3.4 Specificity in Cell-to-Cell Communication
Mechanisms used by prokaryotes to communicate with eukaryotic cells are an important topic that has been widely investigated, particularly in regard to pathogenesis. Bacteria that cause disease may express virulence genes in order to successfully invade a host cell (Alfano and Collmer 2004). However, little information is available on fungal–bacterial signaling. The crosstalk may involve a combination of bacterial type III secretion system (TTSS) as well as host cell cytoskeleton rearrangement, quorum sensing, stress responses, competence, conjugation, motility, sporulation, biofilm formation, antibiotic, and volatile microbial organic compounds (MVOCs) production (Cotter and DiRita 2000; Miller and Bassler 2001). Many symbionts, such as nodulating rhizobia, rely on TTSS for communication (Freiberg et al. 1997; Marie et al. 2001). Moreover, a significant number of bacteria occurring in the Laccaria proxima mycorrhizosphere have been described for their TTSS (Warmink and van Elsas 2008). These data could help to identify in the TTSS a specific trigger in which bacteria and fungi become associated. Among soil bacteria, the quorum sensing mechanisms are again well described for Rhizobium sp., nodulation (Rodelas et al. 1999), nitrogen fixation, and symbiosome development (Daniels et al. 2002).
In this regard, as discussed above within Tuber-associated bacteria, another molecular determinant for fungal–bacterial attachment is described by a specific secreted protein, a lectin that binds Rhizobium sp. (Cerigini et al. 2008).
Moreover, some bacterial metabolites may favor hyphal growth. For example, during its interaction with Amanita muscaria, Streptomyces sp. AcH505 produces high levels of auxofuran, a secondary metabolite that promotes the extension of the fungal mycelium. In another case, unidentified volatile substances synthesized by some bark beetle-associated bacteria induce the growth of their symbiotic fungi (Adams et al. 2009).
In contrast, an example of specificity crosstalk between Tuber and associated bacteria showing a negative effect on mycelia growth is demonstrated by MVOCs produced by Staphylococcus pasteurii, which is strongly antagonistic toward T. borchii mycelium when grown together (Barbieri et al. 2005b). This bacterial strain, found on the roots of micropropagated plantlets, appears to selectively inhibit the growth of T. borchii, but not that of Hebeloma radicosum (Bull.) Ricken, supporting the specificity in cell-to-cell communication between bacteria and ectomycorrhizal fungal partners (Zambonelli et al. 2009).
4 Truffle Life Cycle: A New System for Studying the Interactions Between Edible Ectomycorrhizal Fungi and N2-Fixing Bacteria
Ectomycorrhizal symbiosis is often a necessary event that precedes fruit body formation, such as that of truffles. Truffles of many Tuber species are edible and in great demand because of their particular organoleptic properties. The presence of nitrogen-fixing bacteria recently shown in the white truffle fruit bodies and in association with soil and other ectomycorrhizal fungi led to the hypothesis that nitrogen fixation activity may play an important role in fungal growth and in the development of the ascocarp.
Although there are many studies concerning interactions between ectomycorrhizal fungi and bacteria, the underlying mechanisms behind these associations are generally not well understood, and their functional properties still require further experimental confirmation.
4.1 Truffle Life-Cycle Phases and Bacterial Interactions
Truffles belonging to the Pezizales establish ectomycorrhizal symbiosis with the roots of gymnosperms and angiosperms (Pegler et al. 1993). The truffle life cycle is highly complex and includes morphogenetic changes which can be summarized in three different stages: in the first stage, spore germination generates a filamentous mycelium that successively establishes a symbiotic association with the plant roots leading, in the second stage, to the formation of the mycorrhizas where the fungus and the plant are in an intimate symbiotic relationship. In the final stage, an hypogeous ascoma containing asci and ascospores is formed (Trappe 1979; Harley and Smith 1993). Hypogeous mycorrhizal fungi constantly interact with microorganisms present in the soil, but questions remain about when and how bacteria and mycorrhizal fungi interact and when and how bacteria become associated with truffles and mycelium.
Recent molecular approaches demonstrate that prokaryotes are associated not only with the extra radical hyphae of mycorrhizal fungi, but they also occur in their mycorrhizas and ascocarps, i.e., the Cytophaga/Flexibacter strain found in T. borchii (Barbieri et al. 2005a), suggesting that bacteria occur within the symbiotic fungi during their life cycle.
From the perspective of the bacterium, there are a number of ways in which it may establish an interaction with the ectomycorrhizal fungi, but many questions have not yet been answered (1) Is there a specific stage of the fungal life cycle that attracts bacteria? (2) How do bacteria adhere, colonize, and infect fungal tissues and remain associated with the fungus? (3) What are the advantages for the host to maintain this association?
It is important to assess whether there are potential helper bacteria for the truffle ontogenetic cycle. Some of our studies are planned to address this, and a recent molecular approach based on 16S rRNA ribosomal genes (rDNAs) has allowed us to identify a large number of microbial species associated with the most important species of white truffle T. borchii and T. magnatum at different stages of maturation, greatly expanding the scenario of the microbial ecology that characterizes truffle development and maturation. Indeed, the molecular approaches used for both T. borchii and T. magnatum fruit bodies generated a high variety of 16S rDNA sequences from six phylogenetic groups: α-, β-, γ-Proteobacteria; Bacteroidetes; and low G + C and high G + C Gram-positive bacteria. However, the largest portion of operational taxonomic units (OTUs) within these species was identified as α-Proteobacteria with close relatives among the Sinorhizobium/Ensifer, Rhizobium/Agrobacterium, and Bradyrhizobium groups. The prevalence of α-Proteobacteria associated with T. magnatum ascocarps, regardless of their stage of maturation, was confirmed by the quantitative FISH approach (85 % of α-Proteobacteria among all FISH-positive cells). This method demonstrated the occurrence of rhizobia-like bacterial cells, detected by hybridization of the specific Rhizobium probe RHI1247. The most representative clones among the α-Proteobacteria were closely related to B. elkanii (Barbieri et al. 2005a, 2007). This OTU was consistently present within each ascoma analyzed. B. elkanii was previously described as an inoculant for soybean, which is capable of fixing atmospheric nitrogen in symbiotic interactions with leguminous plants (Rumjanek et al. 1993). No information is currently available on the specific interaction of this species with ectomycorrhizal fungi.
Preliminary studies on the survey of α-Proteobacteria occurring in the ectomycorrhizas collected in productive experimental truffle grounds by 16S rDNA sequencing retrieval also revealed the presence of Bradyrhizobium spp. within T. borchii ectomycorrhizas (unpublished data). These results provide the first evidence that potential nitrogen-fixing bacteria may be involved in the entire life cycle of truffle and that the nitrogenase expression may be a common feature of this symbiosis (Fig. 8.1).
Neighbor-joining phylogenetic analysis of the NifH amino acid-deduced sequences extracted from ascomata of both Tuber magnatum and Tuber borchii RNA overlapping with the 16S rDNA sequence analyses from the RNA extracted from the same truffle species and 16S rDNA sequences from DNA extracted from T. borchii ectomycorrhizas collected in productive experimental truffle grounds. Sequences obtained were compared with homologous sequences obtained by t/BLASTX algorithm. Bootstrap analyses were based on 1,000 re-samplings of the sequence alignment. The sequence of Frankia sp. was included as the out-group
4.2 Toward Molecular Signaling for Truffle–Bacterial Attachment
Although the interaction between fungi and bacteria has been extensively described in several different contexts, including agriculture and clinical environment to food microbiology (Frey-Klett et al. 2011), more studies are needed in these areas to acquire significant progress in the interaction between bacteria and ectomycorrhizal fungi belonging to the Tuber genus and in different phases of the truffle life cycle.
One of the few examples of molecular signaling for fungal–bacterial attachment in truffles is given by the TBF-1 protein secreted by T. borchii in the ascocarp, a species-specific lectin that binds Rhizobium sp. (Cerigini et al. 2008).
The participation of lectins in the Rhizobium–Leguminosae association has been extensively investigated, and it was proposed that the interaction occurs between lectins present in the roots of the leguminous plants and the bacterial surface polysaccharides (Hamblin and Kent 1973; Bohlool and Schidt 1974; Kijne et al. 1997; Hirsch 1999; Fraysse et al. 2003; Laus et al. 2006). The TBF-1 is expressed only during the fructification phase (De Bellis et al. 1998), when the presence of nitrogen-fixing bacteria could be required for fruiting body development, and fungus bacteria mediator molecules, such as lectins, may be involved in these mutualistic associations. Rhizobium spp. produces different surface polysaccharides which are either secreted or included into the capsule surrounding the cell (Laus et al. 2006; Skorupska et al. 2006). The rhizobial exopolysaccharides (EPS), species-specific complex heteropolysaccharides, are exported outside the cell, and they seemed to be involved in the attachment of some rhizobacteria to arbuscular mycorrhizal hyphal structures (Bianciotto et al. 2001).
In a recent paper (Cerigini et al. 2008), we employed some of the Rhizobium strains isolated from T. borchii and T. magnatum ascomata and a few reference Rhizobium strains, which showed a mucoid phenotype, for crude EPS extracts testing to evaluate their ability to bind TBF-1 in an hemagglutination inhibition assay. One interesting result was that TBF-1 lectin binded only to EPS extracted by the Rhizobium strains from T. borchii ascoma. The correlation between the specificity of the lectin and its ability to recognize the correspondent bacteria was very strict: it did not react either with specific Rhizobium symbionts of Leguminosae nor with Rhizobia isolated from the phylogenetically related organism T. magnatum.
The high binding specificity of TBF-1 towards the rhizobial surface polysaccharides led us to suppose an active role of the protein to attract and to sort these bacteria during the fructification phase. The ability of this protein to selectively bind the respective T. borchii-associated Rhizobia supports the hypothesis of its involvement in species-specific interaction with soil bacteria.
In which phase of the fungal biological cycle these bacteria interact, mycelium propagation, ectomycorrhizal establishment, or ascoma formation, still remains to be discovered. Since TBF-1 lectin is expressed only in the fructification phase, it could be hypothesized that rhizobia first infect the developing ascocarp and remain attached to the hypha which emerge from germinating truffle spores. Further investigation will be needed to confirm this hypothesis, and although truffle is not fully culturable in vitro, T. borchii is one of the ideal experimental models because the entire biological cycle is already available in vitro for this species (e.g., mycelial culture, pre-symbiosis, and ectomycorrhizal synthesis as well as experimental truffle-producing areas for ascocarp collection).
4.3 Can Nitrogen-Fixing Bacteria Play a Role in the Nutrition of Fruiting Bodies in Truffles?
Extending previous research from our group, we have recently addressed the problem of the molecular link between nif gene expression and the energetics of N2 fixation in truffles. In our efforts to identify N2-fixing bacteria within truffles, we also demonstrated nitrogen fixation activity throughout the study of bacterial nitrogenase gene expression and by assay of enzymatic activity in T. magnatum. The nitrogenase activity, evaluated by an acetylene reduction assay, was 0.5–7.5 μmol C2H4 h−1 g−1, comparable with early nodules of legumes associated with specific nitrogen-fixing bacteria. The phylogenetic analysis of nitrogenase gene nifH from T. magnatum ascomata at different stages of maturation revealed the presence of α-Proteobacteria belonging to Bradyrhizobium spp., and expression of Bradyrhizobia nifH genes was also detected (Barbieri et al. 2010). Since Bradyrhizobium spp. is found abundantly within the mature ascoma, the conditions these bacteria encounter in the colonized tissue may approach those required for free-living nitrogen fixation (Agarwal and Keister 1983; Casella et al. 1988). Indeed, T. magnatum ascomata develop in soil exposed to variable temperatures during a period from late summer to autumn/winter, where drought and moisture alternate. These seasonal fluctuations affect their development.
Nitrogenase activity in the specific association of T. magnatum with diazotrophic bacteria has important implications for the biology of this fungus. Although the saprotrophic strategy of ascocarp development is debated (Barry et al. 1994; Zeller et al. 2008), T. magnatum ascomata can grow in soils that appear to have low numbers of mycorrhizas (Bertini et al. 2006). Moreover, direct linkages among belowground mycelium, mycorrhizas, and ascocarps in T. magnatum truffle ground are not evident (Zampieri et al. 2010). Under these conditions, enhanced nutrient availability by nitrogen-fixing bacteria might have a beneficial effect on the development and maturation of the fruit bodies.
The Italian white truffle is highly regarded by chefs and gourmets and commands very high market prices, but repeated attempts to cultivate it have been unsuccessful (Murat et al. 2005; Hall et al. 2007). Traditional cultivation techniques, which involve planting Tuber colonized plants in suitable sites, do not consider the potential effects of nitrogen-fixing bacteria on truffle development (Zambonelli and Iotti 2006). The recent studies on Tuber-associated bacteria provide new insights into the role of the nitrogen-fixing bacteria, occurring within fungal tissue during the maturation of T. magnatum ascomata, and suggest new possibilities for improving the cultivation technique of this prized delicacy.
The ability of nitrogen fixation demonstrated for the white truffle T. magnatum has been recently analyzed in other species, and preliminary results from our laboratory show that nitrogenase expression of α-Proteobacteria is also associated with T. borchii, Tuber aestivum, and Tuber melanosporum (unpublished data). This evidence leads to hypothesize that a common mechanism in nitrogen utilization occurs during truffle development and maturation.
5 Conclusions and Perspectives
The studies described in this chapter are useful in advancing knowledge in plant–fungal–bacteria interactions and helpful in guiding future investigations on nitrogen-fixing bacteria associated with ectomycorrhizal fungi, with particular attention to the genus Tuber. Indeed, the expression analyses of both nitrogenase gene and enzyme activity in T. magnatum ascocarps and the occurrence of nitrogen-fixing bacteria in other truffle species suggest that diazotrophic bacteria may contribute to nitrogen input in forest ecosystems. These microbes can therefore be assigned to the recently revisited mycorrhizal helper bacteria group, given that they positively interact with the functioning of the mycorrhizal symbiosis. Nevertheless, direct confirmation of nitrogen fixation and quantification of the net nitrogen input remain to be performed and improved understanding of the regulation of the nitrogenase genes expression is needed.
Current evidence suggests that mutualistic fungal–bacterial interactions are more widespread than expected and that their influence may be crucial in ecosystems. The possibility that the competitiveness of ectomycorrhizal fungi in the soil and perhaps the composition of fungal communities could be influenced by nitrogen-fixing bacteria is an interesting scientific speculation, which may have serious implications for micropropagated plant production in forestry and agronomy.
References
Adams AS, Currie CR, Cardoza Y, Klepzig KD, Raffa KF (2009) Effects of symbiotic bacteria and tree chemistry on the growth and reproduction of bark beetle fungal symbionts. Can J For Res 39:1133–1147. doi:10.1139/X09-034
Agarwal AK, Keister DL (1983) Physiology of ex planta nitrogenase activity in Rhizobium japonicum. Appl Environ Microbiol 45:1592–1601
Alfano JR, Collmer A (2004) Type III secretion system effector proteins: double agents in bacterial disease and plant defense. Annu Rev Phytopathol 42:385–414. doi:10.1146/annurev.phyto.42.040103.110731
Assigbetse K, Gueye M, Thioulouse J, Duponnois R (2005) Soil bacterial diversity responses to root colonization by an ectomycorrhizal fungus are not root-growth-dependent. Microb Ecol 50:350–359. doi:10.1007/s00248-004-0229-x
Barbieri E, Bertini L, Rossi I, Ceccaroli P, Saltarelli R, Guidi C, Zambonelli A, Stocchi V (2005a) New evidence for bacterial diversity in the ascoma of the ectomycorrhizal fungus Tuber borchii Vittad. FEMS Microbiol Lett 247:23–35. doi:10.1016/j.femsle.2005.04.027
Barbieri E, Gioacchini AM, Zambonelli A, Bertini L, Stocchi V (2005b) Determination of microbial volatile organic compounds from Staphylococcus pasteuri against Tuber borchii using solid-phase microextraction and gas chromatography/ion trap mass spectrometry. Rapid Commun Mass Spectrom 19:3411–3415. doi:10.1002/rcm.2209
Barbieri E, Guidi C, Bertaux J, Frey-Klett P, Garbaye J, Ceccaroli P, Saltarelli R, Zambonelli A, Stocchi V (2007) Occurrence and diversity of bacterial communities in Tuber magnatum during truffle maturation. Environ Microbiol Rep 9:2234–2246. doi:10.1111/j.1462-2920.2007.01338.x
Barbieri E, Ceccaroli P, Saltarelli R, Guidi C, Potenza L, Basaglia M, Fontana F, Baldan E, Casella S, Ryahi O et al (2010) New evidence for nitrogen fixation within the Italian white truffle Tuber magnatum. Fungal Biol 114:936–942. doi:10.1016/j.funbio.2010.09.001
Barea JM, Azcón-Aguilar C, Azcón R (1997) In: Gange AC, Brown VK (eds) Interactions between mycorrhizal fungi and rhizosphere microorganisms within the context of sustainable soil-plant systems. Blackwell Science, Cambridge
Barea JM, Azcon R, Azcon-Aguilar C (2002) Mycorrhizosphere interactions to improve plant fitness and soil quality. Antonie Van Leeuwenhoek 81:343–351. doi:10.1023/A:1020588701325
Barriuso J, Ramos Solano B, Lucas JA, Lobo AP, García-Villaraco A, Gutiérrez Mañero FJ (2008) Ecology, genetic diversity and screening strategies of plant growth promoting rhizobacteria (PGPR). In: Ahmad I, Pichtel J, Hayat S (eds) Plant-bacteria interactions strategies and techniques to promote plant growth. Wiley, Weinheim
Barry D, Staunton S, Callot G (1994) Mode of the absorption of water and nutrients by ascocarps of Tuber Melanosporum and Tuber aestivum—a radioactive-tracer technique. Can J Bot 72:317–322. doi:10.1139/b94-041
Bertini L, Rossi I, Zambonelli A, Amicucci A, Sacchi A, Cecchini M, Gregori G, Stocchi V (2006) Molecular identification of Tuber magnatum ectomycorrhizae in the field. Microbiol Res 161:59–64. doi:10.1016/j.micres.2005.06.003
Bianciotto V, Andreotti S, Balestrini R, Bonfante P, Perotto S (2001) Extracellular polysaccharides are involved in the attachment of Azospirillum brasilense and Rhizobium leguminosarum to arbuscular mycorrhizal structures. Eur J Histochem 45:39–49
Bohlool BB, Schidt EL (1974) Lectins: a possible basis for specificity in the Rhizobium-legume root module symbiosis. Science 188:269–271. doi:10.1126/science.185.4147.269
Bonfante P (2001) At the interface between mycorrhizal fungi and plants: the structural organization of cell wall, plasma membrane and cytoskeleton. In: Hock B (ed) Mycota, IX Fungal associations. Springer, Berlin
Bonfante P, Anca IA (2009) Plants, mycorrhizal fungi, and bacteria: a network of interactions. Annu Rev Microbiol 63:363–383. doi:10.1146/annurev.micro.091208.073504
Bowen GD, Theodorou C (1979) Interactions between bacteria and ectomycorrhizal fungi. Soil Biol Biochem 11:119–126
Brooks DD, Chan R, Starks ER, Grayston SJ, Jones MD (2011) Ectomycorrhizal hyphae structure components of the soil bacterial community for decreased phosphatase production. FEMS Microbiol Ecol 76:245–255. doi:10.1111/j.1574-6941.2011.01060.x
Burke DJ, Dunham SM, Kretzer AM (2008) Molecular analysis of bacterial communities associated with the roots of Douglas fir (Pseudotsuga menziesii) colonized by different ectomycorrhizal fungi. FEMS Microbiol Ecol 65:299–309. doi:10.1111/j.1574-6941.2008.00491.x
Casella S, Shapleigh JP, Lupi F, Payne WJ (1988) Nitrite reduction in bacteroids of Rhizobium “hedysari” strain HCNT 1. Arch Microbiol 149:384–388
Cerigini E, Palma F, Barbieri E, Buffalini M, Stocchi V (2008) The Tuber borchii fruiting body-specific protein TBF-1, a novel lectin which interacts with associated Rhizobium species. FEMS Microbiol Lett 284:197–203. doi:10.1111/j.1574-6968.2008.01197.x
Challis GL, Hopwood DA (2003) Synergy and contingency as driving forces for the evolution of multiple secondary metabolite production by Streptomyces species. Proc Natl Acad Sci USA 100:14555–14561. doi:10.1073/pnas.1934677100
Coombs JT, Franco CMM (2003) Isolation and identification of actinobacteria from surface-sterilized wheat roots. Appl Environ Microbiol 69:5603–5608. doi:10.1128/Aem.69.9.5603-5608.2003
Cotter PA, DiRita VJ (2000) Bacterial virulence gene regulation: an evolutionary perspective. Annu Rev Microbiol 54:519–565. doi:10.1146/annurev.micro.54.1.519
Dakora FD (2003) Defining new roles for plant and rhizobial molecules in sole and mixed plant cultures involving symbiotic legumes. New Phytol 158:39–49. doi:10.1046/j.1469-8137.2003.00725.x
Daniels R, De Vos DE, Desair J, Raedschelders G, Luyten E, Rosemeyer V, Verreth C, Schoeters E, Vanderleyden J, Michiels J (2002) The cin quorum sensing locus of Rhizobium etli CNPAF512 affects growth and symbiotic nitrogen fixation. J Biol Chem 277:462–468. doi:10.1074/jbc.M106655200
De Bellis R, Agostini D, Piccoli G, Vallorani L, Potenza L, Polidori E, Sisti D, Amoresano A, Pucci P, Arpaia G et al (1998) The tbf-1 gene from the white truffle Tuber borchii codes for a structural cell wall protein specifically expressed in fruit body. Fungal Genet Biol 25:87–99. doi:S1087184598910921
de Boer W, Folman LB, Summerbell RC, Boddy L (2005) Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol Rev 29:795–811. doi:10.1016/j.femsre.2004.11.005
Deveau A, Palin B, Delaruelle C, Peter M, Kohler A, Pierrat JC, Sarniguet A, Garbaye J, Martin F, Frey-Klett P (2007) The mycorrhiza helper Pseudomonas fluorescens BBc6R8 has a specific priming effect on the growth, morphology and gene expression of the ectomycorrhizal fungus Laccaria bicolor S238N. New Phytol 175:743–755. doi:10.1111/j.1469-8137.2007.02148.x
Deveau A, Brule C, Palin B, Champmartin D, Rubini P, Garbaye J, Sarniguet A, Frey-Klett P (2010) Role of fungal trehalose and bacterial thiamine in the improved survival and growth of the ectomycorrhizal fungus Laccaria bicolor S238N and the helper bacterium Pseudomonas fluorescens BBc6R8. Environ Microbiol Rep 2:560–568. doi:10.1111/j.1758-2229.2010.00145.x
Duponnois R, Garbaye J (1992) Some mechanisms involved in growth stimulation of ectomycorrhizal fungi by bacteria. Can J Bot 68:2148–2152
Founoune H, Duponnois R, Ba AM, Sall S, Branget I, Lorquin J, Neyra M, Chotte JL (2002) Mycorrhiza helper bacteria stimulated ectomycorrhizal symbiosis of Acacia holosericea with Pisolithus alba. New Phytol 153:81–89. doi:10.1046/j.0028-646X.2001.00284.x
Fraysse N, Couderc F, Poinsot V (2003) Surface polysaccharide involvement in establishing the rhizobium-legume symbiosis. Eur J Biochem 270:1365–1380. doi:3492
Freiberg C, Fellay R, Bairoch A, Broughton WJ, Rosenthal A, Perret X (1997) Molecular basis of symbiosis between Rhizobium and legumes. Nature 387:394–401. doi:10.1038/387394a0
Frey-Klett P, Garbaye J (2005) Mycorrhiza helper bacteria: a promising model for the genomic analysis of fungal-bacterial interactions. New Phytol 168:4–8. doi:10.1111/j.1469-8137.2005.01553.x
Frey-Klett P, Chavatte M, Clausse ML, Courrier S, Le Roux C, Raaijmakers J, Martinotti MG, Pierrat JC, Garbaye J (2005) Ectomycorrhizal symbiosis affects functional diversity of rhizosphere fluorescent pseudomonads. New Phytol 165:317–328. doi:10.1111/j.1469-8137.2004.01212.x
Frey-Klett P, Garbaye J, Tarkka M (2007) The mycorrhiza helper bacteria revisited. New Phytol 176:22–36. doi:10.1111/j.1469-8137.2007.02191.x
Frey-Klett P, Burlinson P, Deveau A, Barret M, Tarkka M, Sarniguet A (2011) Bacterial-fungal interactions: hyphens between agricultural, clinical, environmental, and food microbiologists. Microbiol Mol Biol Rev 75:583–609. doi:10.1128/MMBR.00020-11
Garbaye J (1994) Helper bacteria—a new dimension to the mycorrhizal symbiosis. New Phytol 128:197–210
Garbaye J, Bowen GD (1989) Stimulation of ectomycorrhizal infection of Pinus radiata by some microorganisms associated with the mantle of ectomycorrhizas. New Phytol 112:383–388
Garbaye J, Duponnois R (1992) Specificity and function of mycorrhization helper bacteria (MHB) associated with the Pseudotsuga menziesii-Laccaria laccata symbiosis. Symbiosis 14:335–344
Gazzanelli G, Malatesta M, Pianetti A, Baffone W, Stocchi V, Citterio B (1999) Bacteria associated to fruit bodies of the ecto-mycorrhizal fungus Tuber borchii Vittad. Symbiosis 26:211–222. doi:10.1023/A:1020949531639
Grayston SJ, Vaughan D, Jones D (1997) Rhizosphere carbon flow in trees, in comparison with annual plants: the importance of root exudation and its impact on microbial activity and nutrient availability. Appl Soil Ecol 5:29–56. doi:10.1016/S0929-1393(96)00126-6
Hall IR, Brown G, Zambonelli A (2007) Taming the truffle. The history, lore, and science of the ultimate mushroom. Timber, Portland, OR
Hamblin J, Kent SP (1973) Possible role of phytohaemagglutinin in Phaseolus vulgaris L. Nat New Biol 245:28–30
Harley JL, Smith SE (1993) Mycorrhizal symbiosis. Academic, London
Hirsch AM (1999) Role of lectins (and rhizobial exopolysaccharides) in legume nodulation. Curr Opin Plant Biol 2:320–326. doi:10.1016/S1369-5266(99)80056-9
Izumi H, Anderson IC, Alexander IJ, Killham K, Moore ERB (2006a) Diversity and expression of nitrogenase genes (nifH) from ectomycorrhizas of Corsican pine (Pinus nigra). Environ Microbiol 8:2224–2230. doi:10.1111/j.1462-2920.2006.01104.x
Izumi H, Anderson IC, Alexander IJ, Killham K, Moore ERB (2006b) Endobacteria in some ectomycorrhiza of Scots pine (Pinus sylvestris). FEMS Microbiol Ecol 56:34–43. doi:10.1111/j.1574-6941.2005.00048.x
Izumi H, Cairney JWG, Killham K, Moore E, Alexander IJ, Anderson IC (2008) Bacteria associated with ectomycorrhizas of slash pine (Pinus elliottii) in south-eastern Queensland, Australia. FEMS Microbiol Lett 282:196–204. doi:10.1111/j.1574-6968.2008.01122.x
Jayasinghearachchi HS, Seneviratne G (2005) Fungal solubilization of rock phosphate is enhanced by forming fungal-rhizobial biofilms. Soil Biol Biochem 38:405–408. doi:10.1016/j.soilbio.2005.06.004
Johansson JF, Paul LR, Finlay RD (2004) Microbial interactions in the mycorrhizosphere and their significance for sustainable agriculture. FEMS Microbiol Ecol 48:1–13. doi:10.1016/j.femsec.2003.11.012
Kataoka R, Taniguchi T, Ooshima H, Futai K (2008) Comparison of the bacterial communities established on the mycorrhizae formed on Pinus thunbergii root tips by eight species of fungi. Plant Soil 304:267–275. doi:10.1007/s11104-008-9548-x
Kijne JW, Bauchrowitz MA, Diaz CL (1997) Root lectins and Rhizobia. Plant Physiol 115:869–873. doi:115/3/869
Kretzer AM, King ZR, Bai S (2009) Bacterial communities associated with tuberculate ectomycorrhizae of Rhizopogon spp. Mycorrhiza 19:277–282. doi:10.1007/s00572-008-0213-2
Laus MC, Logman TJ, Lamers GE, Van Brussel AA, Carlson RW, Kijne JW (2006) A novel polar surface polysaccharide from Rhizobium leguminosarum binds host plant lectin. Mol Microbiol 59:1704–1713. doi:10.1111/j.1365-2958.2006.05057.x
Marie C, Broughton WJ, Deakin WJ (2001) Rhizobium type III secretion systems: legume charmers or alarmers? Curr Opin Plant Biol 4:336–342. doi:10.1016/S1369-5266(00)00182-5
Martin F, Kohler A, Duplessis S (2007) Living in harmony in the wood underground: ectomycorrhizal genomics. Curr Opin Plant Biol 10:204–210. doi:10.1016/j.pbi.2007.01.006
Meyer JR, Linderman RG (1986) Selective influence on population of rhizosphere or rhizoplane bacteria and actinomycetes by mycorrhizas formed by Glomus fasciculatum. Soil Boil Biochem 18:191–196
Miller MB, Bassler BL (2001) Quorum sensing in bacteria. Annu Rev Microbiol 55:165–199. doi:10.1146/annurev.micro.55.1.16555/1/165
Murat C, Vizzini A, Bonfante P, Mello A (2005) Morphological and molecular typing of the below-ground fungal community in a natural Tuber magnatum truffle-ground. FEMS Microbiol Lett 245:307–313. doi:10.1016/j.femsle.2005.03.019
Pavić A, Stanković S, Marjanović Ž (2011) Biochemical characterization of a sphingomonad isolate from the ascocarp of white truffle (Tuber magnatum Pico). Arch Biol Sci 63:697–704. doi:10.2298/ABS1103697P
Pegler DN, Spooner BM, Young TWK (1993) British truffles. A revision of British hypogeous fungi. Royal Botanic Garden, Kew
Rainey PB, Cole ALJ, Fermor TR, Wood DA (1990) A model system for examining involvement of bacteria in basidiome initiation of Agaricus-Bisporus. Mycol Res 94:191–195
Rangel-Castro JI, Levenfors JJ, Danell E (2002) Physiological and genetic characterization of fluorescent Pseudomonas associated with Cantharellus cibarius. Can J Microbiol 48:739–748. doi:10.1139/W02-062
Rodelas B, Lithgow JK, Wisniewski-Dye F, Hardman A, Wilkinson A, Economou A, Williams P, Downie JA (1999) Analysis of quorum-sensing-dependent control of rhizosphere-expressed (rhi) genes in Rhizobium leguminosarum bv. viciae. J Bacteriol 181:3816–3823
Rumjanek NG, Dobert RC, van Berkum P, Triplett EW (1993) Common soybean inoculant strains in Brazil are members of Bradyrhizobium elkanii. Appl Environ Microbiol 59:4371–4373
Ryberg M, Matheny PB (2012) Asynchronous origins of ectomycorrhizal clades of Agaricales. Proc R Soc B Biol Sci 279:2003–2011. doi:10.1098/rspb.2011.2428
Sbrana C, Agnolucci M, Bedini S, Lepera A, Toffanin A, Giovannetti M, Nuti MP (2002) Diversity of culturable bacterial populations associated to Tuber borchii ectomycorrhizas and their activity on T. borchii mycelial growth. FEMS Microbiol Lett 211:195–201. doi:S0378109702007127
Singh LP, Gill SS, Tuteja N (2011) Unraveling the role of fungal symbionts in plant abiotic stress tolerance. Plant Signal Behav 6:175–191. doi:10.4161/psb.6.2.14146
Skorupska A, Janczarek M, Marczak M, Mazur A, Krol J (2006) Rhizobial exopolysaccharides: genetic control and symbiotic functions. Microb Cell Fact 5:7. doi:10.1186/1475-2859-5-7
Splivallo R, Novero M, Bertea CM, Bossi S, Bonfante P (2007) Truffle volatiles inhibit growth and induce an oxidative burst in Arabidopsis thaliana. New Phytol 175:417–424. doi:10.1111/j.1469-8137.2007.02141.x
Tarkka MT, Piechulla B (2007) Aromatic weapons: truffles attack plants by the production of volatiles. New Phytol 175:381–383. doi:10.1111/j.1469-8137.2007.02165.x
Tarkka MT, Lehr NA, Hampp R, Schrey SD (2008) Plant behavior upon contact with Streptomycetes. Plant Signal Behav 3:917–919
Timonen S, Hurek T (2006) Characterization of culturable bacterial populations associating with Pinus sylvestris-Suillus bovinus mycorrhizospheres. Can J Microbiol 52:769–778. doi:10.1139/W06-016
Trappe JM (1979) The orders, families, and genera of hypogeus Ascomycotina (truffles and their relatives). Mycotaxon 9:297–340
Uroz S, Calvaruso C, Turpaul MP, Pierrat JC, Mustin C, Frey-Klett P (2007) Effect of the mycorrhizosphere on the genotypic and metabolic diversity of the bacterial communities involved in mineral weathering in a forest soil. Appl Environ Microbiol 73:3019–3027. doi:10.1128/Aem.00121-07
Warmink JA, van Elsas JD (2008) Selection of bacterial populations in the mycosphere of Laccaria proxima: is type III secretion involved? ISME J 2:887–900. doi:10.1038/ismej.2008.41
Wu XQ, Hou L-L, Sheng J-M, Ren J-H, Zheng L, Chen D, Ye J-R (2011) Effects of ectomycorrhizal fungus Boletus edulis and mycorrhiza helper Bacillus cereus on the growth and nutrient uptake by Pinus thunbergii. Biol Fertil Soils. doi:10.1007/s00374-011-0638-1
Zambonelli A, Iotti M (2006) The pure culture of Tuber mycelia and their use in the cultivation of the truffles. Actes du premier Symposium sur les Champignons hypogés du Bassin Méditerranéen, RABAT, Kabar L
Zambonelli A, Iotti M, Barbieri E, Amicucci A, Stocchi V, Peintner U, Hall I (2009) The microbial communities and fruiting of edible ectomycorrhizal mushrooms. Yunnan Zhiwu Yanjiu IWEMM5 16:81–85
Zampieri E, Murat C, Cagnasso M, Bonfante P, Mello A (2010) Soil analysis reveals the presence of an extended mycelial network in a Tuber magnatum truffle-ground. FEMS Microbiol Ecol 71:43–49. doi:10.1111/j.1574-6941.2009.00783.x
Zeller B, Brechet C, Maurice JP, Le Tacon F (2008) Saprotrophic versus symbiotic strategy during truffle ascocarp development under holm oak. A response based on (13)C and (15)N natural abundance. Ann For Sci. doi:10.1051/Forest:2008037
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Barbieri, E., Ceccaroli, P., Palma, F., Agostini, D., Stocchi, V. (2012). Ectomycorrhizal Helper Bacteria: The Third Partner in the Symbiosis. In: Zambonelli, A., Bonito, G. (eds) Edible Ectomycorrhizal Mushrooms. Soil Biology, vol 34. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-33823-6_8
Download citation
DOI: https://doi.org/10.1007/978-3-642-33823-6_8
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-33822-9
Online ISBN: 978-3-642-33823-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)