Abstract
Due to the long history of genetic analyses in yeasts and their experimental tractability, the yeast genome duplication provides important perspectives on the genome and population-level processes that follow whole-genome duplication (WGD). We discuss the history of the discovery of the Saccharomyces cerevisiae WGD, with special emphasis on the role of comparative genomics in its analysis. We then explore models of the WGD shaped population and species divergence, both at a gene level (e.g., Dobzhansky-Muller incompatibility) and from the perspective of recent work on secondary allopolyploidy in Saccharomyces pastorianus. Finally, we explore the selective forces that act on the WGD-produced paralogs and shape their patterns of loss and retention. In addition to discussing the dosage balance hypothesis as it applies to the yeast WGD, we explore the role of the WGD in shaping several complex metabolic and regulatory phenotypes.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
15.1 Introduction
Researchers have found remnants of ancient whole-genome duplications (WGDs) preserved in the genomes of many and diverse eukaryotes. This book, in fact, is a testament to that diversity, illustrating the sheer number of independent events in plants as well as the evolutionarily basal events in vertebrates and other, more recent WGDs in teleost fishes and frogs. Although we have still not fully validated Susumo Ohno’s claim for the primacy of polyploidization in the generation of new adaptations (Ohno 1970), it is clear that WGD events have had a massive influence on the content and structure of the genomes of their possessors. The next step in exploring Ohno’s hypothesis is to link genome evolution to known changes in function. This goal, however, remains challenging, primarily because our knowledge of how genotype links to phenotype remains woefully incomplete (Pigliucci 2010). However, one group of organisms in which we can at least begin to make such associations is in the polyploid yeasts. Our knowledge of the functional genomics of yeast is drawn primarily from Saccharomyces cerevisiae, which has a well-annotated genome, decades of biochemical, genetic, and cell biology research, a relatively small genome, and a life cycle that lends itself to scalable laboratory analyses. Taken together, these facts have allowed yeast researchers not only to understand the structure of the genome following WGD but also to experimentally evaluate hypotheses regarding the evolution of particular complex phenotypes. As we will stress throughout this chapter, one of the themes that emerges from all of these analyses is the degree to which the outcome of a WGD depends as much on the interactions between genes as on the role of any particular locus. Of equal importance evolutionarily, we now also have data from other polyploid yeast species, which are valuable both as a point of comparison to S. cerevisiae and for their own sakes. This wealth of data affords us insight into the mechanisms that drive the preferential loss and preservation of gene duplicates after polyploidy, lead to the functional divergence of genes, and are behind the evolutionary origins of complex phenotypes.
15.2 Evidence for WGD in Yeast
Although early analyses of genes potentially created by the vertebrate 2R events used phylogenetic approaches (Hughes 1999; Furlong and Holland 2002), most current studies of WGD rely on one or both of two methods: (1) finding numerous blocks of paralogous genes in multiple chromosomes with similar gene orders and (2) clustering homologs into groups by measuring the rate of synonymous substitutions (K s or dS). This second method assumes that the gene pairs created by WGD cluster about some mean K s value (Lynch and Conery 2000). The simultaneous application of both methods has been used to group multiple WGD events within species (e.g., Arabidopsis thaliana and Tetraodon nigroviridis; Jaillon et al. 2004; Van de Peer et al. 2009). However, the yeast genomes present an interesting challenge in this respect because the synonymous substitutions between yeast paralogs produced by WGD (hereafter ohnologs; Wolfe 2000) are often saturated (Byrne and Wolfe 2007). In other words, identical synonymous positions between two ohnologs occur almost as often due to repeated convergent substitutions as due to common ancestry, a fact pointed out by Smith (1987), who attempted to date histone gene duplicates in yeast. While the genomic structure of the core histone genes suggested that they were all duplicated simultaneously, these genes show considerable variation in the numbers of synonymous substitutions separating them. This led Smith (1987) to hypothesize that S. cerevisiae underwent a WGD ancient enough that the duplicates surviving from it had saturated. However, it was not until the genome sequence of S. cerevisiae became available that this speculation could be confirmed (see below). Another similar but subtler problem in using K s as a means of dating duplicate genes is the issue of gene conversion. Gene conversion was presumed to be quite common in yeast (Petes and Hill 1988), even prompting some authors (Gao and Innan 2004) to suggest that estimates of duplication rates based on duplicate divergences were inapplicable due to the homogenization of duplicate loci by conversion. Fortunately, although gene conversion is very common among yeast ribosomal proteins, it does not appear to be a general characteristic of the genome (Evangelisti and Conant 2010). Nonetheless, these various issues collectively meant that comparisons of paralogous sequence divergence were deemed unhelpful as a means to detect WGD in yeast.
15.2.1 Synteny-Based Evidence for WGD in Saccharomyces cerevisiae
Given that paralogous sequence comparisons were generally unhelpful in finding WGD relics, another tactic was to consider gene order. In fact, even before the S. cerevisiae genome sequence was completed in 1996, it was clear to many researchers that it contained numerous, long, homologous clusters of ordered genes (Goffeau et al. 1996). Melnick and Sherman (1993) found ordered homologous gene clusters in chromosomes V and X covering 7.5 kb. Lalo et al. (1993) similarly found ordered homologous gene clusters in chromosomes XIV and III covering 15 kb. When the genome was sequenced, researchers found 18 ordered homologous genes in chromosomes IV and II that covered 120 and 170 kb, respectively (Goffeau et al. 1996). Just how to interpret these redundant regions remained a challenge at that time (Goffeau et al. 1996; Oliver 1996), and, in spite of Smith’s prior hypothesis (Smith 1987), few, if any, of the contemporaneous explanations included an ancient WGD.
However, opinions changed the next year when Wolfe and Shields (1997) presented a thorough, genome sequence-based analysis that gave strong evidence for WGD in S. cerevisiae. To find syntenic regions, they conducted a BLASTP search of amino acid sequences throughout the yeast genome and made a dot plot of the results. They then created gene blocks from these data, where each block was required to have at least three homologous pairs with intergenic distances ≤ 50 kb and conservation of gene order and orientation. This analysis yielded 55 duplicated regions containing a total of 376 pairs of ohnologs. The large number of duplicated regions led Wolfe and Shields to posit two explanations: (1) successive independent gene duplications, and (2) a single duplication of the entire genome, followed by massive gene loss. There were two lines of evidence discounting the first possibility. First, 90 % (50/55) of the gene regions shared the same orientation with respect to the centromeres of the duplicated regions when we would expect independent duplications to be instead randomly distributed about the centromeres. Second, there were no examples of triplicated regions in the S. cerevisiae genome. If the duplications involved several distinct events separated in time, such a pattern would be highly unlikely, because it would require that later duplication events never overlapped with prior ones. Given these arguments, Wolfe and Shields (1997) argued for a single ancient WGD, which they dated to be hundreds of millions of years old (although attempts to conclusively date this have been difficult, due to a lack of fossils and the previously mentioned saturation of substitutions; see Taylor and Berbee 2006; Rolland and Dujon 2011).
15.2.2 Comparative Genomics and Proof of WGD in S. cerevisiae
A number of researchers disputed the claims of Wolfe and Shields (1997), arguing that, because the syntenic regions identified made up only a small part of the genome, independent duplications better explained the genomic structure of S. cerevisiae (Coissac et al. 1997; Mewes et al. 1997; Hughes et al. 2000; Llorente et al. 2000a; Llorente et al. 2000b; Friedman and Hughes 2001; Piskur 2001; Koszul et al. 2004). However, this independent duplication hypothesis became untenable following the genome sequencing of other yeasts that proved to lack these syntenic paralog blocks. These sequences were described by three independent groups. The comparison of S. cerevisiae with Kluyveromyces waltii (Kellis et al. 2004) and the comparison of S. cerevisiae with Ashbya gossypii (Dietrich et al. 2004) involved different genomes, but effectively made the same argument: that the 2:1 mapping of blocks of paralogs from S. cerevisiae to homologous single-copy genes in K. waltii/A.gossypii could best be explained by WGD. This explanation was particularly striking because the doubly conserved synteny blocks cover 90 % of the genome in K. waltii (Kellis et al. 2004) and 96 % of that in A. gossypii (Dietrich et al. 2004). Furthermore, both studies found a large number of 2:1 pairing of centromeres in the species-respective chromosomes. There were 16:8 such pairings between S. cerevisiae and K. waltii and 14:7 between S. cerevisiae and A. gossypii with a subsequent break at the expected centromere position in S. cerevisiae chromosomes X and XII that are syntenic with regions in A. gossypii chromosomes I and III. Finally, and perhaps most strikingly, both groups also showed that the single-copy orthologs of genes from A. gossypii or K. waltii in the genome of S. cerevisiae are interleaved between two paralogous chromosomes in S. cerevisiae that nonetheless retain the relative gene order of the single chromosome in the non-WGD yeast (see Fig. 15.1). Such a pattern is only explicable under the hypothesis of a WGD event followed by massive gene losses.
Consensus view of the evolutionary relationships between the yeast taxa discussed. Black branches indicate relationships described by both Kurtzman and Robnett (2003) and Fitzpatrick et al. (2006). Red branches indicate conflicts between the two phylogenies, in which case Fitzpatrick et al. (2006) is presented. Curved grey branches illustrate the allopolyploidy events between two species (S. cerevisiae and S. bayanus, Z. rouxii, and Z. pseudorouxii). Taxa in blue are reported in text (e.g., S. pastorianus and Z. rouxii ATCC 42981). Stars mark whole-genome duplications. Note that genus names are an imperfect guide to the relationships
The argument of Dujon et al. (2004) is subtly different. They sequenced and analyzed four other genomes. One genome that of Candida glabrata, shares the genome duplication with S. cerevisiae. This was determined by comparing syntenic blocks in S. cerevisiae and C. glabrata with the other three sequenced genomes, Kluveromyces lactis, Debaryomyces hansenii, and Yarrowia lipolytica. Dujon et al. (2004) found 20 distinct blocks of paralogs shared by both S. cerevisiae and C. glabrata. These blocks allowed them to map the WGD onto a phylogeny, rather than do a simple pairwise comparison. Mapping this WGD phylogenetically creates distinct hypotheses as to where in the tree we expect to find polyploid yeasts (c.f., Fig. 15.1), predictions that have been confirmed with each of the subsequently sequenced genomes of known phylogenetic position (Wapinski et al. 2007; Scannell et al. 2011).
15.2.3 Yeast Gene Order Browser
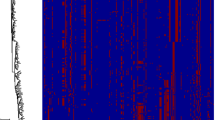
One of the major benefits of studying the yeast WGD is that the relatively slow rates of gene order change in yeast genomes and the compactness of their genomes means that an exhaustive enumeration of all WGD-produced ohnologs is possible. Just such a project was carried out, with the results presented as the web-based Yeast Gene Order Browser (YGOB) (Byrne and Wolfe 2005), which illustrates a number of non- and post-WGD yeasts in a graphical framework (Fig. 15.2). This work has been followed by a reconstruction of the set of genes and their relative orders that existed just prior to the WGD (Gordon et al. 2009) and by a likelihood-based model of post-WGD duplicate loss that attempts to quantify the orthology inferences made by YGOB (Conant and Wolfe 2008a). On the basis of these three projects, the post-WGD evolutionary history of virtually every locus in the S. cerevisiae genome can be traced (Fig. 15.2 is thus illustrative of the predominant pattern seen across the genome).
Yeast gene order browser (YGOB) screenshots with a window size of six. Each box represents a gene; each color, a chromosome. The gene in focus, the A. gossypii gene ABR086W, is highlighted by an orange border. Each vertical column (“pillar”) represents a single gene prior to the WGD (hence, all genes in a column are homologs, and the paired upper and lower genes, when present, are paralogs). The ancestral order of these genes (pink boxes) just prior to the WGD has also been exhaustively inferred (Gordon et al. 2011). Connectors join nearby genes: a solid bar for adjacent genes, two bars for loci less than five genes apart, and one bar for loci <20 genes apart. The connectors are extended in gray over intervening space. The end of a chromosome or contig is denoted by a brace. Arrows denote transcriptional orientation. The browser also includes a control panel that allows users to select the window size and the gene to focus on. This panel also has buttons for running BLAST searches against YGOB’s database, outputting YGOB data in tabular format, obtaining pairwise K a and K s values among genes, and computing multiple sequence alignments and phylogenetic trees of individual pillars. Species names for each track are labeled at right (Byrne and Wolfe 2005)
15.2.4 Additional Non-Saccharomyces-Specific WGDs
In addition to the ancient WGD that characterizes the Saccharomyces clade (Fig. 15.1), several cases of allopolyploidy have been discovered in yeasts. Some of these occur in species within the Saccharomyces sensu stricto clade (Scannell et al. 2011), while others are independent.
15.2.4.1 Secondary Allopolyploidy in Saccharomyces pastorianus
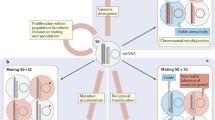
Several cases of allopolyploidy are known from within S. sensu stricto (Dequin and Casaregola 2011). One of the most well studied is that of the lager yeast, S. pastorianus (syn. S. carlsbergensis). It has long been known that the polyploid S. pastorianus and other members of the complex of related lager yeasts are allotetraploids of diploid S. cerevisiae and some other unknown diploid species (Martini and Kurtzman 1985; Kielland-Brandt et al. 1995). However, aside from the general difficulties facing anyone interested in identifying the origins of hybrid genomes, the debate surrounding the origin of the second parental diploid species was further complicated by a difficulty in delimiting species within these groups (Rainieri et al. 2006). The tetraploid S. pastorianus belongs to a group of yeast species which, until recently, was represented as a phylogenetically unresolved species complex including S. pastorianus, S. monacensis (S. pastorianus strain CBS 1503), S. bayanus, and S. bayanus var. uvarum (Casaregola et al. 2001; Rainieri et al. 2006). This taxonomic confusion has recently been partially resolved through the sequencing of the genomes of both S. pastorianus and one of its presumed parental diploid species, S. eubayanus (Nakao et al. 2009). The genome history that has emerged is a complicated story of allopolyploidy followed by genomic transformation forming the related species S. bayanus (Libkind et al. 2011). As summarized by Libkind et al. (2011), S. cerevisiae hybridized with S. eubayanus (a species recently recovered in Patagonia) with subsequent genome doubling producing the allotetraploid progenitor of modern S. pastorianus. Following domestication, smaller regions of the S. pastorianus genome were then apparently transferred into the genome of the diploid parent S. eubayanus (which is nonetheless a descendant of the ancient polyploidy). This hybrid form of S. eubayanus, with contributions from S. pastorianus, then proceeded to interbreed with diploid S. uvarum to produce the modern, diploid, S. bayanus (Fig. 15.3).
Genome evolution in S. pastorianus. A model of the formation of S. pastorianus and the hybrid strains of S. bayanus. First, wild S. eubayanus and ale-type S. cerevisiae hybridized to form an allotetraploid that became the ancestor of the modern (doubly paleopolyploid) S. pastorianus. Second, domestication imposed strong selective pressure for strains with the most desirable brewing properties. Third, in the brewing vats with high densities of S. pastorianus, cell lysis releases large DNA fragments that occasionally transform, fourth, contaminating wild strains of S. eubayanus (which possesses only the ancient WGD shared with S. cerevisiae) because of the lack of pure culture techniques. Fifth, multiple hybridization events between S. eubayanus and wild strains of S. uvarum gave rise to CBS 380T and NBRC 1948. This model does not exclude prior or parallel involvement of S. uvarum in brewing or contamination. Reprinted from Libkind et al. (2011)
15.2.4.2 Allopolyploidy in Zygosaccharomyces rouxii
The spoilage agent and industrial yeast Zygosaccharomyces rouxii strain ATCC 42981 was identified as another allopolyploid by James et al. (2005), Gordon and Wolfe (2008). This hybridization/polyploidy event is significant for two reasons. First, unlike all of the previous examples, it occurs outside of S. sensu stricto. Second, Gordon and Wolfe (2008) determined that most of the paralogs produced by WGD are still present, presumably due to the recentness of the event. Thus, while other yeast genome duplications are ancient and show considerable gene loss and rearrangement (Wolfe and Shields 1997), the Z. rouxii genome retains most of the “new” genes produced by its WGD. Since the survival time of ohnologs has been modeled to follow a power law, most of the duplicates are expected to be lost very rapidly (Maere et al. 2005), suggesting that Z. rouxii represents an example of the early features of genome evolution following WGD.
15.3 WGD and Speciation
An important potential outcome of polyploidy is in altering patterns of speciation. This change can happen in at least two ways. First, the WGD can relax selective constraints resulting in an adaptive radiation by means of ecological speciation. Another, more neutral mechanism, is a special case of the Dobzhansky-Muller (DM) process of speciation, in which species lose reciprocal paralogs following some period of isolation (Lynch and Force 2000; Werth and Windham 1991; see also Chap. 1, this volume). WGD potentially increases the probability of this simply by increasing the number of paired genes in a genome. The fertility of hybrids is 0.75n, where n is the number of reciprocal losses of essential genes among populations (Werth and Windham 1991). Clearly, for any significant number of reciprocal losses (such as occur after WGD), the number of viable, fertile offspring of a crossing of two such populations is negligible. Both phylogenetic and experimental studies of the DM process after WGD have been carried out in yeast. Scannell et al. (2006) showed that the number of reciprocal gene losses in several species of yeast sharing the S. cerevisiae WGD was sufficient to induce such inviability. This observation suggests that a DM mechanism was partly responsible for the multiple speciation events among the Saccharomyces species (e.g., S. cerevisiae, S. bayanus, and C. glabrata) following WGD.
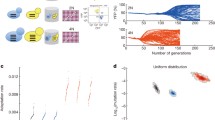
An advantage of studying the DM process in yeast is the ability to experimentally create and cross artificial polyploids. This possibility has been highlighted in experimental studies of reproductive isolation. Polyploid yeasts have been allowed to evolve in different selective environments (Dettman et al. 2007) and in neutral environments subject to random mutagenesis (Maclean and Greig 2011). These two experiments have shown that moderate reproductive isolation, coupled with reciprocal gene loss, results in a clear loss of fitness when independently derived polyploids are crossed. Similarly, Lee et al. (2008) showed that hybrids of S. cerevisiae and S. bayanus were less fit than their parental phenotypes, due primarily to incompatibility between their nuclear and mitochondrial genomes. Chou et al. (2010) extended this analysis, providing another pair of mitochondrial and nuclear genes and posited nuclear-mitochondrial incompatibility as a common mechanism in species formation. In another twist, Anderson et al. (2010) demonstrated the existence of alleles with depressed hybrid fitness in low-glucose environments, which argues for a model in which neutral changes in paired genes are followed by strong selection, a sequence of events that promotes rapid reproductive isolation. Kao et al. (2010), however, argue against the existence of a small number of so-called speciation genes, instead claiming that genome scans provide no evidence of any single paired dominant or recessive genic incompatibilities. They instead argue that following WGD, many changes in loci of little effect resulted in lowered fitness due, in part, to the rewiring of transcriptional and metabolic networks.
Another debate that has emerged in this field is whether these changes are due primarily to the decrease in the fertility of hybrids (Xu and He 2011) or a decrease in their viability (Greig 2008). This question ultimately amounts to a debate about what stage in the yeast life cycle the genetic incompatibilities occur—sporulation or clonal growth, and whether the decrease in fitness is the result of competition for resources or offspring. The discontinuity between these ideas likely represents an opportunity to explain speciation as a process across different genomic and temporal scales, and we would speculate that the process of DM incompatibility induces selection for the evolution of some form of prezygotic barrier.
15.4 Changes in Genome Content and Complexity Post-WGD
Duplicate retention and evolutionary models. In addition to such population processes as speciation, WGD also altered many other aspects of the S. cerevisiae lifestyle. For instance, several pairs of ohnologs have been shown to have undergone various types of functional divergence, allowing the study of some of the proposed mechanisms of duplicate divergence after duplication (Conant and Wolfe 2008b). In an elegant series of experiments, van Hoof (2005a) showed that two ohnologs, ORC1 and SIR3, have distinct and non-overlapping functions (in DNA replication and gene silencing, respectively). Strikingly, however, the mutual ortholog of these genes from the non-WGD yeast S. kluyveri is able to complement both functions, constituting a clear example of subfunctionalization. An apparently similar case, involving the S. cerevisiae ohnolog pair GAL1 and GAL3, which presently functions, respectively, as an enzyme and as a transcriptional regulator, was complicated by the discovery of an adaptive conflict between the shared regulator and enzymatic function of their ortholog in the non-WGD K. lactis. Thus, although the K. lactisGAL1 gene does indeed serve the functions of both GAL1 and GAL3 in S. cerevisiae, it does so in a suboptimal way, being unable to tune its expression to both roles simultaneously (Hittinger and Carroll 2007). This conflict illustrates an important point about subfunctionalization, namely that the original neutral model of subfunctionalization proposed by Force and coauthors (Force et al. 1999) is not the only possible mechanism for such functional partitioning (Des Marais and Rausher 2008). Other examples of divergence among ohnologs where the mechanism of that divergence is less clear include ribosomal proteins (Ni and Snyder 2001; Komili et al. 2007; Kim et al. 2009), glucose sensors (Özcan et al. 1998), and glycolysis enzymes (Boles et al. 1997).
The dosage balance hypothesis (DBH). In addition to facilitating the above work, the wealth of functional data from S. cerevisiae also provides an excellent opportunity to test hypotheses explaining the differences in gene retention patterns after WGD and small-scale duplications (hereafter SSD). Chief among these is probably the DBH (Papp et al. 2003; Freeling and Thomas 2006; Birchler and Veitia 2007; Freeling 2009), which states that, in eukaryotes, there is selection operating to disfavor duplications of central network genes due to the imbalance in network stoichiometry that results. This situation is reversed for WGD because in that case, the loss of a second copy of a gene introduces imbalances relative to the remaining duplicated genes. In keeping with the DBH, several classes of genes are over-retained after several evolutionarily ancient WGD events, including that in yeast. They include ribosomal proteins and protein kinases and transcription factors (Seoighe and Wolfe 1999; Blanc and Wolfe 2004; Maere et al. 2005; Aury et al. 2006; Conant and Wolfe 2008a). Similarly, genes that tend to have been fixed by WGD are less likely to have undergone SSD in other yeast species (Wapinski et al. 2007). However, duplicates produced by WGD have more protein interactions (Guan et al. 2007; Hakes et al. 2007), more phosphorylation sites (Amoutzias et al. 2010), and tend to be highly expressed (Seoighe and Wolfe 1999). Although genes retained in duplicate after WGD are rarely essential on an individual basis (Guan et al. 2007), this dispensability appears to be due to functional compensation by the other ohnolog (Deluna et al. 2008). Thus, it appears that while ohnologs are less likely to be essential than their SSD counterparts today, their ancestral genes were actually at least as essential as current single-copy genes (Deluna et al. 2008).
System-level changes produced by WGD. Of course, one of the unique features of polyploidy relative to SSD is the possibility of coordinated changes among multiple sets of ohnologs. At the simplest level, we have previously illustrated examples of what appears to be network subfunctionalization where a number of ohnologs collectively divided two expression domains among themselves (Conant and Wolfe 2006b). A more complex and interesting example is the role of the WGD (Piškur et al. 2006) in shaping S. cerevisiae’s propensity for aerobic glucose fermentation (the Crabtree effect; Geladé et al. 2003; Johnston and Kim 2005), a novel and somewhat paradoxical phenotype. There is a general association between the presence of the WGD and the Crabtree effect across yeast species (Merico et al. 2007). As a result, we and others have argued that dosage effects among the glycolysis enzymes post-WGD helped to increase flux through glycolysis (Blank et al. 2005; Kuepfer et al. 2005; Conant and Wolfe 2007; Merico et al. 2007; van Hoek and Hogeweg 2009). Such increased flux likely could only be accommodated through fermentation pathways, given the complex spatial organization of the competing respiratory pathway (Conant and Wolfe 2007). Supporting this hypothesis is an elegant computational analysis by van Hoek and Hogeweg (2009) showing that future WGD events in modern S. cerevisiae could also be expected to provide a selective advantage in glucose-rich environments through the preferential retention of duplicated glycolysis enzymes. Note that the apparently “wasteful” fermentation can actually be selectively advantageous in the context of rich but ephemeral resource patches (Pfeiffer et al. 2001; Pfeiffer and Schuster 2005), a phenomenon that has been experimentally confirmed in yeast (MacLean and Gudelj 2006). Such a change in the yeast lifestyle likely led to other, later changes in the genome. One suggestive example concerns the decoupling of cytosolic and mitochondrial ribosomal protein expression post-WGD (Ihmels et al. 2005). Prior to WGD, bakers’ yeast was likely similar to other yeasts in having a strong association in the expression of the two types of ribosomal proteins. After WGD, however, cis-regulatory element evolution diverged in the two groups of genes (Ihmels et al. 2005), allowing S. cerevisiae to express only cytosolic proteins at high levels during fermentation, an important refinement in a fermentative lifestyle.
Connecting the DBH to large-scale evolutionary changes following WGD, Conant (2010) and (Fusco et al. 2010) found transcriptional regulatory motifs to be over-retained in ohnologs. Modeling network evolution after WGD, these authors find the network enriched for transcription factors and particular network motifs. Duplicated transcription factors still show some relics of the WGD, being more likely to share targets than are random transcription factors, but on the whole show considerable divergence post-WGD (Conant 2010). Given this rapid regulatory evolution, it may not be easy to ascertain the role of WGD in the evolution of the modern S. cerevisiae regulatory network. Nonetheless, the retention of many transcription factors that have acquired distinct sets of target genes may imply that the WGD served to “relax” the regulatory complexity of this organism, which may have implications for its future ability to adapt (as seen for the GAL1/GAL3 example).
15.5 Conclusions
The S. cerevisiae WGD has been implicated in a number of evolutionarily complex events. At a minimum, a set of duplicated genes of identical age is a powerful system for exploring duplicate gene evolution (van Hoof 2005b; Conant and Wolfe 2006a; Fares et al. 2006; Kim and Yi 2006). However, we also suggest that, as with the GAL1/GAL3 example, we will not fully understand the biology of S. cerevisiae until we account for how the WGD has altered both the individual roles of particular genes and their relationships to each other. We have outlined some of the areas of yeast biology that we think were altered by this genome-doubling event: there remain others yet to be discovered. Similarly, the presence of other WGD events, of varying ages, allows us to study how these events unfold over various timescales, including, potentially, on the timescale of laboratory experiments in evolution.
References
Amoutzias GD, He Y, Gordon J, Mossialos D, Oliver SG, Van de Peer Y (2010) Posttranslational regulation impacts the fate of duplicated genes. Proc Natl acad sci U.S.A 107:2967–2971
Anderson JB, Funt J, Thompson DA et al (2010) Determinants of divergent adaptation and Dobzhansky-Muller interaction in experimental yeast populations. Curr Biol 20:1383–1388
Aury JM, Jaillon O, Duret L et al (2006) Global trends of whole-genome duplications revealed by the ciliate Paramecium tetraurelia. Nature 444:171–178
Birchler JA, Veitia RA (2007) The gene balance hypothesis: from classical genetics to modern genomics. Plant Cell 19:395–402
Blanc G, Wolfe KH (2004) Functional divergence of duplicated genes formed by polyploidy during Arabidopsis evolution. Plant Cell 16:1679–1691
Blank LM, Lehmbeck F, Sauer U (2005) Metabolic-flux and network analysis of fourteen hemiascomycetous yeasts. FEMS Yeast Res 5:545–558
Boles E, Schulte F, Miosga T, Freidel K, Schlüter E, Zimmermann FK, Hollenberg CP, Heinisch JJ (1997) Characterization of a glucose-repressed pyruvate kinase (Pyk2p) in Saccharomyces cerevisiae that is catalytically insensitive to fructose-1-6-biphosphate. J Bacteriol 179:2987–2993
Byrne KP, Wolfe KH (2005) The yeast gene order browser: combining curated homology and syntenic context reveals gene fate in polyploid species. Genome Res 15:1456–1461
Byrne KP, Wolfe KH (2007) Consistent patterns of rate asymmetry and gene loss indicate widespread neofunctionalization of yeast genes after whole-genome duplication. Genetics 175:1341–1350
Casaregola S, Nguyen HV, Lapathitis G, Kotyk A, Gaillardin C (2001) Analysis of the constitution of the beer yeast genome by PCR, sequencing and subtelomeric sequence hybridization. Int J Syst Evol Microbiol 51:1607–1618
Chou J-Y, Hung Y-S, Lin K-H, Lee H-Y, Leu J-Y (2010) Multiple molecular mechanisms cause reproductive isolation between three yeast species. PLoS Biol 8:e1000432
Coissac E, Maillier E, Netter P (1997) A comparative study of duplications in bacteria and eukaryotes: the importance of telomeres. Mol Biol Evol 14:1062–1074
Conant GC (2010) Rapid reorganization of the transcriptional regulatory network after genome duplication in yeast. Proc R Soc B 277:869–876
Conant GC, Wolfe KH (2006a) Functional partitioning of yeast co-expression networks after genome duplication. PLoS Biol 4:e109
Conant GC, Wolfe KH (2006b) Functional partitioning of yeast co-expression networks after genome duplication. PLoS Biol 4:e109
Conant GC, Wolfe KH (2007) Increased glycolytic flux as an outcome of whole-genome duplication in yeast. Mol Syst Biol 3:129
Conant GC, Wolfe KH (2008a) Probabilistic cross-species inference of orthologous genomic regions created by whole-genome duplication in yeast. Genetics 179:1681–1692
Conant GC, Wolfe KH (2008b) Turning a hobby into a job: how duplicated genes find new functions. Nat Rev Genet 9:938–950
Deluna A, Vetsigian K, Shoresh N, Hegreness M, Colon-Gonzalez M, Chao S, Kishony R (2008) Exposing the fitness contribution of duplicated genes. Nat Genet 40(5):676–681
Dequin S, Casaregola S (2011) The genomes of fermentative Saccharomyces. CR Biol 334:687–693
Des Marais DL, Rausher MD (2008) Escape from adaptive conflict after duplication in an anthocyanin pathway gene. Nature 454:762–765
Dettman JR, Sirjusingh C, Kohn LM, Anderson JB (2007) Incipient speciation by divergent adaptation and antagonistic epistasis in yeast. Nature 447:585–588
Dietrich FS, Voegeli S, Brachat S et al (2004) The Ashbya gossypii genome as a tool for mapping the ancient Saccharomyces cerevisiae genome. Science 304:304–307
Dujon B, Sherman D, Fischer G et al (2004) Genome evolution in yeasts. Nature 430:35–44
Evangelisti AM, Conant GC (2010) Nonrandom survival of gene conversions among yeast ribosomal proteins duplicated through genome doubling. Genome Biol Evol 2:826–834
Fares MA, Byrne KP, Wolfe KH (2006) Rate asymmetry after genome duplication causes substantial long-branch attraction artifacts in the phylogeny of Saccharomyces species. Mol Biol Evol 23:245–253
Fitzpatrick D, Logue M, Stajich J, Butler G (2006) A fungal phylogeny based on 42 complete genomes derived from supertree and combined gene analysis. BMC Evol Biol 6:99
Force A, Lynch M, Pickett FB, Amores A, Yan Y, Postlethwait J (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Freeling M (2009) Bias in plant gene content following different sorts of duplication: tandem, whole-genome, segmental, or by transposition. Annu Rev Plant Biol 60:433–453
Freeling M, Thomas BC (2006) Gene-balanced duplications, like tetraploidy, provide predictable drive to increase morphological complexity. Genome Res 16:805–814
Friedman R, Hughes AL (2001) Gene duplication and the structure of eukaryotic genomes. Genome Res 11:373–381
Furlong RF, Holland PWH (2002) Were vertebrates octoploid? Philos Trans R Soc Lond B 357:531–544
Fusco D, Grassi L, Bassetti B, Caselle M, Lagomarsino MC (2010) Ordered structure of the transcription network inherited from the yeast whole-genome duplication. BMC Syst Biol 4:77
Gao LZ, Innan H (2004) Very low gene duplicaiton rate in the yeast genome. Science 306:1367–1370
Geladé R, Van De Velde S, Van Dijck P, Thevelein JM (2003) Multi-level response of the yeast genome to glucose. Genome Biol 4:233
Goffeau A, Barrell B, Russey H et al (1996) Life with 6000 genes. Science 274:562–567
Gordon JL, Byrne KP, Wolfe KH (2009) Additions, losses and rearrangements on the evolutionary route from a reconstructed ancestor to the modern Saccharomyces cerevisiae genome. PLoS Genet 5:e1000485
Gordon JL, Byrne KP, Wolfe KH (2011) Mechanisms of chromosome number evolution in yeast. PLoS Genet 7:e1002190
Gordon JL, Wolfe KH (2008) Recent allopolyploid origin of Zygosaccharomyces rouxii strain ATCC 42981. Yeast 25:449–456
Greig D (2008) Reproductive isolation in Saccharomyces. Heredity 102:39–44
Guan Y, Dunham MJ, Troyanskaya OG (2007) Functional analysis of gene duplications in Saccharomyces cerevisiae. Genetics 175:933–943
Hakes L, Pinney JW, Lovell SC, Oliver SG, Robertson DL (2007) All duplicates are not equal: the difference between small-scale and genome duplication. Genome Biol 8:R209
Hittinger CT, Carroll SB (2007) Gene duplication and the adaptive evolution of a classic genetic switch. Nature 449:677–681
Hughes AL (1999) Phylogenies of developmentally important proteins do not support the hypothesis of two rounds of genome duplication early in vertebrate history. J Mol Evol 48:565–576
Hughes TR, Roberts CJ, Dai H et al (2000) Widespread aneuploidy revealed by DNA microarray expression profiling. Nat Genet 25:333–337
Ihmels J, Bergmann S, Gerami-Nejad M, Yanai I, McClellan M, Berman J, Barkai N (2005) Rewiring of the yeast transcriptional network through the evolution of motif usage. Science 309:938–940
Jaillon O, Aury J-M, Brunet F et al (2004) Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature 431:946–957
James SA, Bond CJ, Stratford M, Roberts IN (2005) Molecular evidence for the existence of natural hybrids in the genus Zygosaccharomyces. FEMS Yeast Res 5:747–755
Johnston M, Kim J-H (2005) Glucose as a hormone: Receptor-mediated glucose sensing in the yeast Saccharomyces cerevisiae. Biochem Soc Trans 33:247–252
Kao KC, Schwartz K, Sherlock G (2010) A genome-wide analysis reveals no nuclear Dobzhansky-Muller pairs of determinants of speciation between S. cerevisiae and S. paradoxus, but suggests more complex incompatibilities. PLoS Genet 6:e1001038
Kellis M, Birren BW, Lander ES (2004) Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 428:617–624
Kielland-Brandt MC, Nilsson-Tillgren T, Gjermansen C, Holmberg S, Pedersen MB (1995) Genetics of brewing yeasts. In: Rose AH, Wheals AE, Harrison JS (eds) The Yeasts, vol 6, 2nd edn. Academic, London, pp 223–254
Kim S-H, Yi SV (2006) Correlated asymmetry of sequence and functional divergence between duplicate proteins of Saccharomyces cerevisiae. Mol Biol Evol 23:1068–1075
Kim T-Y, Ha CW, Huh W-K (2009) Differential subcellular localization of ribosomal protein L7 paralogs in Saccharomyces cerevisiae. Mol Cells 27:539–546
Komili S, Farny NG, Roth FP, Silver PA (2007) Functional specificity among ribosomal proteins regulates gene expression. Cell 131:557–571
Koszul R, Caburet S, Dujon B, Fischer G (2004) Eucaryotic genome evolution through the spontaneous duplication of large chromosomal segments. EMBO J 23:234–243
Kuepfer L, Sauer U, Blank LM (2005) Metabolic functions of duplicate genes in Saccharomyces cerevisiae. Genome Res 15:1421–1430
Kurtzman C, Robnett C (2003) Phylogenetic relationships among yeasts of the ‘Saccharomyces complex’ determined from multigene sequence analyses. FEMS Yeast Res 3:417–432
Lalo D, Stettler S, Mariotte S, Slominski PP, Thuriaux P (1993) Two yeast chromosomes are related by a fossil duplication of their centromeric regions. C R Acad Sci 316:367–373
Lee H-Y, Chou J-Y, Cheong L, Chang N-H, Yang S-Y, Leu J-Y (2008) Incompatibility of nuclear and mitochondrial genomes causes hybrid sterility between two yeast species. Cell 135:1065–1073
Libkind D, Hittinger CT, Valério E, Gonçalves C, Dover J, Johnston M, Gonçalves P, Sampaio JP (2011) Microbe domestication and the identification of the wild genetic stock of lager-brewing yeast. Proc Nat Acad Sci 108:14539–14544
Llorente B, Durrens P, Malpertuy A et al (2000a) Genomic exploration of the hemiascomycetous yeasts: 20. Evolution of gene redundancy compared to Saccharomyces cerevisiae. FEBS Lett 487:122–133
Llorente B, Malpertuy A, NeuvÈglise C et al (2000b) Genomic exploration of the hemiascomycetous yeasts: 18. Comparative analysis of chromosome maps and synteny with Saccharomyces cerevisiae. FEBS Lett 487:101–112
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290:1151–1155
Lynch M, Force AG (2000) The origin of interspecific genomic incompatibility via gene duplication. Am Nat 156:590–605
Maclean CJ, Greig D (2011) Reciprocal gene loss following experimental whole-genome duplication causes reproductive isolation in yeast. Evolution 65:932–945
MacLean RC, Gudelj I (2006) Resource competition and social conflict in experimental populations of yeast. Nature 441:498–501
Maere S, De Bodt S, Raes J, Casneuf T, Van Montagu M, Kuiper M, Van de Peer. Y (2005) Modeling gene and genome duplications in eukaryotes. Proc Nat Acad Sci U S A 102:5454–5459
Martini AV, Kurtzman CP (1985) Deoxyribonucleic acid relatedness among species of the genus Saccharomyces sensu stricto. Int J Syst Bacteriol 35:508–511
Melnick L, Sherman F (1993) The gene clusters ARC and COR on chromosomes 5 and 10, respectively, of Saccharomyces cerevisiae share a common ancestry. J Mol Biol 233:372–388
Merico A, Sulo P, Piškur J, Compagno C (2007) Fermentative lifestyle in yeasts belonging to the Saccharomyces complex. FEBS J 274:976–989
Mewes H, Albermann K, Bähr M et al (1997) Overview of the yeast genome. Nature 387:7–65
Nakao Y, Kanamori T, Itoh T, Kodama Y, Rainieri S, Nakamura N, Shimonaga T, Hattori M, Ashikari T (2009) Genome sequence of the lager brewing yeast, an interspecies hybrid. DNA Res 16:115–129
Ni L, Snyder M (2001) A genomic study of the bipolar bud site selection pattern in Saccharomyces cerevisiae. Mol Biol Cell 12:2147–2170
Ohno S (1970) Evolution by gene duplication. Springer, New York
Oliver SG (1996) From DNA sequence to biological function. Nature 379:597–600
Özcan S, Dover J, Johnston M (1998) Glucose sensing and signaling by two glucose receptors in the yeast Saccharomyces cerevisiae. EMBO J 17:2566–2573
Papp B, Pal C, Hurst LD (2003) Evolution of cis-regulatory elements in duplicated genes of yeast. Trends Genet 19:417–422
Petes T, Hill C (1988) Recombination between repeated genes in microorganisms. Annu Rev Genet 22:147–168
Pfeiffer T, Schuster S (2005) Game-theoretical approaches to studying the evolution of biochemical systems. Trends Biochem Sci 30:20–25
Pfeiffer T, Schuster S, Bonhoeffer S (2001) Cooperation and competition in the evolution of ATP-producing pathways. Science 292:504–507
Pigliucci M (2010) Genotype-phenotype mapping and the end of the ‘genes as blueprint’ metaphor. Philos Trans R Soc Lond B 365:557–566
Piskur J (2001) Origin of the duplicated regions in the yeast genomes. Trends Genet 17:302–303
Piškur J, Rozpedowska E, Polakova S, Merico A, Compagno C (2006) How did Saccharomyces evolve to become a good brewer? Trends Genet 22:183–186
Rainieri S, Kodama Y, Kaneko Y, Mikata K, Nakao Y, Ashikari T (2006) Pure and mixed genetic lines of Saccharomyces bayanus and Saccharomyces pastorianus and their contribution to the lager brewing strain genome. Appl Environ Microbiol 72:3968–3974
Rolland T, Dujon B (2011) Yeasty clocks: dating genomic changes in yeasts. CR Biol 334:620–628
Scannell DR, Byrne KP, Gordon JL, Wong S, Wolfe KH (2006) Multiple rounds of speciation associated with reciprocal gene loss in polyploid yeasts. Nature 440:341–345
Scannell DR, Zill OA, Rokas A, Payen C, Dunham MJ, Eisen MB, Rine J, Johnston M, Hittinger CT (2011) The awesome power of yeast evolutionary genetics: new genome sequences and strain resources for the Saccharomyces sensu stricto genus. G3: Genes, Genomes, Genetics 1:11–25
Seoighe C, Wolfe KH (1999) Yeast genome evolution in the post-genome era. Curr Opin Microbiol 2:548–554
Smith M (1987) Molecular evolution of the Saccharomyces cerevisiae histone gene loci. J Mol Evol 24:252–259
Taylor JW, Berbee ML (2006) Dating divergences in the fungal tree of life: review and new analyses. Mycologia 98:838–849
Van de Peer Y, Fawcett J, Proost S, Sterk L, Vandepoele K (2009) The flowering world: a tale of duplications. Trends Plant Sci 14:680–688
van Hoek MJ, Hogeweg P (2009) Metabolic adaptation after whole genome duplication. Mol Biol Evol 26:2441–2453
van Hoof A (2005a) Conserved functions of yeast genes support the duplication, degeneration and complementation model for gene duplication. Genetics 171:1455–1461
van Hoof A (2005b) Conserved functions of yeast genes support the duplication, degeneration and complementation model for gene duplication. Genetics 171:1455–1461
Wapinski I, Pfeffer A, Friedman N, Regev A (2007) Natural history and evolutionary principles of gene duplication in fungi. Nature 449:54–61
Werth CR, Windham MD (1991) A model for divergent, allopatric speciation of polyploid pteridophytes resulting from silencing of duplicate-gene expression. Am Nat 137:515–526
Wolfe KH (2000) Robustness-it’s not where you think it is. Nat Genet 25:3–4
Wolfe KH, Shields D (1997) Molecular evidence for an ancient duplication of the entire yeast genome. Nature 387:708–713
Xu M, He X (2011) Genetic incompatibility dampens hybrid fertility more than hybrid viability: yeast as a case study. PLoS ONE 6:e18341
Acknowledgments
We would like to thank Michaël Bekaert, Patrick Edger, and Chris Pires for helpful discussions. This work was supported by the National Library of Medicine Biomedical and Health Informatics Training Fellowship [LM007089-19] (CMH) and the Reproductive Biology Group of the Food for the twenty-first century program at the University of Missouri (GCC).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Hudson, C.M., Conant, G.C. (2012). Yeast as a Window into Changes in Genome Complexity Due to Polyploidization. In: Soltis, P., Soltis, D. (eds) Polyploidy and Genome Evolution. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-31442-1_15
Download citation
DOI: https://doi.org/10.1007/978-3-642-31442-1_15
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-31441-4
Online ISBN: 978-3-642-31442-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)