Abstract
Halophilic environments such as solar salterns and salt lakes are enriched in organisms belonging to the domain Archaea. The number of virus-like particles has also been shown to be high. Although most of the described haloarchaeal viruses are head–tail viruses, direct microscopic examination of environmental samples suggests more diversity. In this chapter, we shortly review the existing knowledge of the previously described head–tail viruses and provide more in-depth information on structurally well-characterized icosahedral virus SH1 and the group of enveloped viruses exemplified by HRPV-1 and HHPV-1.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Envelop Virus
- Major Capsid Protein
- Hypersaline Environment
- Major Structural Protein
- Halobacterium Salinarum
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
8.1 Introduction
Organisms thriving in hypersaline environments represent all three domains of life: Archaea, Bacteria, and Eukarya (Oren 2008). It has been shown, however, that the higher the salt concentration of the hypersaline environment, the more dominant are the haloarchaeal species (Benlloch et al. 2002; Casamayor et al. 2002; Pedrós-Alió et al. 2000; Burns et al. 2004a; Santos et al. 2010). The number of virus-like particles (VLPs) is also very high (Guixa-Boixareu et al. 1996; Oren et al. 1997; Pedrós-Alió et al. 2000; Maturrano et al. 2006), and it has been suggested that in hypersaline environment, viruses may be the only predators (Guixa-Boixareu et al. 1996).
Studies on organisms living in extreme environments started already in the late nineteenth century (Oren 2002; Rainey and Oren 2006) and the earliest studies on extreme halophilic environments such as Great Salt Lake and the Dead Sea were reported soon after the first reports (Daniels 1917; Wilkansky 1936). The first haloarchaeal virus was described in 1974 (Torsvik and Dundas 1974).
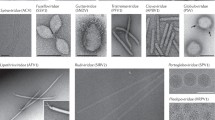
Although a common theme in early studies on halophilic viruses seems to be the discovery of a head–tail virus as a result of a spontaneous lysis of a haloarchaeal culture (Schnabel et al. 1982b; Vogelsang-Wenke and Oesterhelt 1988; Witte et al. 1997) or a discovery of a virus in a flagellar preparation (Torsvik and Dundas 1974), active searches for halophilic viruses were also conducted (Wais et al. 1975; Pauling 1982). All the isolated viruses were of head–tail morphotype and they were studied for the purpose of finding molecular tools for haloarchaea as well as for studying molecular mechanisms of haloarchaeal transcription (Reiter et al. 1988). Unfortunately, many of these early viruses are most probably not available anymore. Prokaryotic viruses have now returned to the spotlight as major players in the biogeochemical cycles and as examples of new viral architectures (Prangishvili et al. 2006; Suttle 2007). This has also activated the search for new haloviruses. Until this day, only approximately 20 haloarchaeal viruses are characterized and most of them represent the head–tail morphology (e.g., ΦH and ΦCh1); and an icosahedral virus (SH1) as well as enveloped viruses (HRPV-1, HHPV-1, His1 and His2) have also been described. These viruses do not show as high diversity in the morphotypes as do the crenarchaeal viruses (Prangishvili et al. 2006), but they show unexpected variability, especially in the nature of the genome and its structure, even between related viruses (Witte et al. 1997; Roine et al. 2010; Vogelsang-Wenke and Oesterhelt 1988). Microscopic examination of environmental samples has also shed light into the potential diversity of the haloarchaeal VLPs, suggesting that although most of the isolated viruses are head–tail viruses, the majority of haloarchaeal viruses are of other morphotypes (Guixa-Boixareu et al. 1996; Oren et al. 1997; Diez et al. 2000; Santos et al. 2007; Sime-Ngando et al. 2010).
In this review, we give a short introduction to the isolated haloarchaeal viruses. The relatively few descriptions of the head–tail halophages (Calvo et al. 1988; Kauri et al. 1991; Kukkaro and Bamford 2009) are not included. We then provide more in-depth information on the structurally well-characterized icosahedral virus SH1 and the group of enveloped viruses exemplified by HRPV-1 and HHPV-1. The detailed structural studies of SH1 have shown evolutionary connections that cannot be revealed by studying the nucleotide or amino acid sequences alone.
8.2 Approaches to Studying Viruses in Hypersaline Environment
As the studies of other environmental samples, the ones on the micro-organisms in extreme environments can methodologically be divided into three different types of approaches. Microscopy of environmental samples gives us an idea of the number of the VLPs leaving the infectivity of these particles an open question (Weinbauer 2004). Viruses having novel particle morphology can also easily pass unrecognized. Environmental sequencing (metagenomics) is a powerful tool in generating new data on the nucleotide sequence space of a sample. As is the case also with the other approaches, it is not devoid of bias due to the sample preparation and analysis of data (Tringe and Rubin 2005; Kunin et al. 2008; Wooley et al. 2010). The third approach, i.e., isolation and cultivation of organisms from the hypersaline environments has been relatively easy, but we face the same problems as in the isolation of organisms from other sources. It has been estimated that only 1–10% of all organisms of an environment can be cultivated in laboratory conditions. An exceptional achievement was the long-lasting project of isolating the most abundant haloarchaeal organism in many hypersaline sources, the square haloarchaeon Haloquadratum walsbyi: It took 24 years from the first microscopic identification by Walsby (1980) to the first reports of an axenic culture (Bolhuis et al. 2004; Burns et al. 2004b). For prokaryotic viruses also the ones with temperate life cycles will easily go unnoticed in the direct screening of viruses from an environmental sample (Stedman et al. 2009).
8.3 Viral Life Cycles in the Hypersaline Environment
The life cycles of haloarchaeal viruses seem to fall in the basic modes of the bacteriophage life cycles (Weinbauer 2004). Most of the head–tail viruses of haloarchaea are temperate, replicating their genome either as part of the host genome (e.g., ΦCh1) or as independent episomes (e.g., ΦH). The increasing NaCl concentrations have shown to change the mode of Hs1 and S5100 infection from lytic to persistent (Torsvik and Dundas 1980; Daniels and Wais 1990). Although this behavior can be easily explained as a strategy of the virus to utilize the cells’ resources for virus production before cell lysis due to dilution of the salt concentration (Daniels and Wais 1990), a critical step-by-step analysis of the interactions at the molecular level will be needed in order to elucidate the causes and the effects. The effect of salt concentration on the adsorption of haloarchaeal viruses and different phages in a gradient of salt was systematically tested in the study by Kukkaro and Bamford (2009). As also reported previously, they concluded that there is no commonly shared correlation between the changes in adsorption rates and a range of salt concentrations used.
The two extreme ends of the viral life cycles are represented by SH1, a virulent haloarchaeal virus, and the enveloped haloarchaeal viruses HRPV-1 and HHPV-1, which extrude continuously from the cells affecting the host cell growth only minimally (Porter et al. 2005; Pietilä et al. 2009; Roine et al. 2010). The difference in these life cycles can be seen in the one-step experiments where a culture of synchronized cells (i.e., cells that are in the same exponential growth phase) is infected at high multiplicity of infection (MOI) in order to get most of the cells infected. The virulent virus SH1 that lyses the cells in the end of the infection cycle causes a dramatic drop in the turbidity of the culture (Fig. 8.1a). In a similar set-up, the enveloped viruses, on the other hand, cause only minor decrease in host cell culture turbidity (Fig. 8.1b). The changes in turbidity also correlate with the amount of living and dividing cells in the culture: The viable cell count of a culture infected with the virulent SH1 has decreased dramatically whereas the cells infected with enveloped viruses can still divide and produce colonies on the plate. In the case of HRPV-1 or HHPV-1, the infected cells form smaller colonies which produce viruses when re-cultured (Pietilä et al. 2009; Roine et al. 2010). In conclusion, the viruses are released from the cells without lysis, but viral infection slows down the division rate of the infected cells.
The one-step growth curves of halophilic euryarchaeal (a) icosahedral virus SH1 and (b) pleomorphic virus HHPV-1 infecting Haloarcula hispanica. The turbidity of uninfected (closed circles) and infected (open circles) cultures are shown. The exponentially growing hosts were infected (white arrows) using multiplicity of infection (MOI) of 40 (in a, SH1) and MOI of 15 (in b, HHPV-1). The free HHPV-1 virions were removed by centrifugation after 1 h post-infection. The beginning of the release of the progeny viruses is marked by black arrows. (a) The lysis of Haloarcula hispanica occurs concomitantly with the SH1 release 5 h post-infection, and the maximum virus yield is approximately 1 × 1012 pfu/ml. (b) HHPV-1 viruses are released continuously from the cells by budding. Virus release starts 3–5 h post-infection and it reaches its maximum (~1 × 1012 pfu/ml) around 25 h post-infection. The cell densities (colony forming units/ml) around 25 h post-infection are indicated by arrowheads: uninfected (black) and infected (white)
8.4 Head–Tail Archaeal Viruses
There are a number of extensive reviews on the research of the head–tail haloviruses with their properties listed and we refer to these for further information (Reiter et al. 1988; Dyall-Smith et al. 2003; Porter and Dyall-Smith 2006; Porter et al. 2007, 2008b). Most of the head–tail viruses were originally reported to infect Halobacterium halobium or Halobacterium curtirubrum (now Halobacterium salinarum) (Porter et al. 2008b; Reiter et al. 1988). Although the status of those strains in terms of the detailed characterization for classification is unclear, many of them are now classified into Halobacterium salinarum (Ventosa and Oren 1996). All the isolated head–tail viruses fall into the groups of myo- and siphoviruses, and no podoviruses have been isolated. Out of the 16 isolated head–tail viruses, only the complete genome sequences of ΦCh1, HF1, HF2 and BJ1 have been published (Pagaling et al. 2007; Porter et al. 2008b). On the basis of this information it is clear, however, that the genomes show similar type of modular structure as the head–tail bacteriophages (Hendrix et al. 2000). The connection between the haloarchaeal and bacterial head–tail viruses is also clear on the basis of gene homologues (Tang et al. 2002; Pagaling et al. 2007; Krupovič et al. 2010). Three dimensional structural modeling of the haloarchaeal major capsid protein gpE of ΦCh1 resulted in a protein fold similar to the bacteriophage HK97 major capsid protein fold showing relatedness also at the higher structural level (Krupovič et al. 2010).
Still to date, the temperate myovirus ΦH isolated from spontaneously lysed cells of Halobacterium halobium R1 (Halobacterium salinarum R1) in 1982 (Schnabel et al. 1982b) is one of the most studied among haloarchaeal viruses. Yet, only partial genome sequence of this virus is available. The distinctive feature of ΦH genome was the observed frequent rearrangements in virus population (Schnabel et al. 1982a, b). The rearrangements were caused by a varying number of copies of the insertion element ISH1.8 as well as insertion of ISH50 (Schnabel et al. 1982a). The variants of the virus containing two copies of ISH1.8 in inverted orientation were shown to experience two types of frequent recombination events: An inversion of the genomic region flanked by the two copies of ISH1.8, the so called L-segment, as well as circularization of the L-segment into a 12-kb plasmid pΦHL (Schnabel 1984; Schnabel et al. 1982a). The pΦHL was shown to confer partial resistance against the ΦH infection (Schnabel 1984). The rest of the research on ΦH deals mostly with the regulation of transcription and super-infection immunity (Schnabel et al. 1984; Gropp et al. 1989, 1992; Stolt and Zillig 1992, 1993b). A ΦH repressor was discovered that forms unusually stable complexes with DNA (Ken and Hackett 1991); also the first archaeal antisense transcript was reported that takes part into the super-infection immunity (Stolt and Zillig 1993a).
Another temperate myovirus ΦCh1 was isolated in 1997 by Angela Witte and her colleagues after spontaneous lysis of the haloalkaliphilic archaeon Natrialba magadii (Witte et al. 1997). ΦCh1 is the only archaeal virus known to contain both DNA and RNA in its virion. The virion-associated RNA is mainly of host-origin varying in length from 100 to 800 nucleotides (Witte et al. 1997). ΦCh1 ORFs that could be assigned to a homologue in the sequence databases were the ORFs of Hbt. salinarum head–tail virus ΦH (Klein et al. 2002). The close relationship between these viruses is surprising as their hosts are phylogenetically distant and in terms of pH they live in totally different environments. As in ΦH, genomic rearrangements have been shown to take place in the ΦCh1 genome. These rearrangements occur in the region encoding a putative site-specific recombinase Int1 (Rössler et al. 2004). These genomic rearrangements are not as versatile as in the ΦH involving mainly an inversion of the int1 gene region that affects the amino acid sequences of two structural proteins of the virion (Rössler et al. 2004). It was suggested that this inversion affects the specificity of the host recognition by changing the putative tail fiber proteins (Rössler et al. 2004). As discussed above for ΦH, variations of the genome are caused by insertion elements. These rearrangements were suggested to reflect the extremely instable nature of the host genome as shown for the Hbt. halobium plasmid pHH1 (Pfeifer et al. 1981) as well as for the whole genome (Sapienza et al. 1982; Simsek et al. 1982; Reiter et al. 1988). Genomic instability comparable to one found in Hbt. salinarium has not been reported for Nab. magadii.
Although there are several other viruses e.g., Hs1, Ja.1, S41, S50.2, S4100, S5100, ΦN, Hh-1, Hh-3, and S45 (Porter et al. 2008b) reported to infect Hbt. salinarum, such genetic divergence of the viral genomes have not been studied. Indirect evidence, however, suggests such a phenomenon for S41 and S50.2 viruses (Daniels and Wais 1998) and genomic rearrangements were also reported for the BJ1 head–tail virus infecting Halorubrum saccharovorum (Pagaling et al. 2007).
In addition to isolated haloviruses, the “environmental halophage 1” (EHP-1) has been identified using a metagenomic approach (Santos et al. 2007). Most of the predicted ORFs with matches to database were similar to predicted gene products of ΦCh1. The low G + C content of the 35 kb genome as well as the codon usage were similar to those of Haloquadratum walsbyi (Santos et al. 2007). Using the combination of a metagenomic approach and environmental microscopy, another saline environment, the Lake Retba (Senegal) was analyzed for viral and halophage content (Sime-Ngando et al. 2010). Microscopy of the concentrated salt water sample showed great diversity in the morphotypes of VLPs with less than 1% representing head–tail viruses. Also sequence data revealed very few similarities to the known sequences (Sime-Ngando et al. 2010). The majority of matches found in databases, however, represented the sequences of head–tail viruses BJ1, ΦCh1 and ΦH and a couple of sequences had matches to the enveloped virus HRPV-1 and His1 genomes.
8.5 The Icosahedral Virus SH1
SH1 was isolated in 2005 (Porter et al. 2005) from a salt lake in Western Australia. To date, SH1 is the only described euryarchaeal virus with icosahedral morphology. Under the icosahedral protein capsid there is a membrane enclosing the genome. SH1 is a virulent virus infecting Haloarcula hispanica. As discussed above (Fig. 8.1a), the infected host cells lyse between 5 and 6 h post infection releasing progeny particles (Porter et al. 2005).
8.5.1 The SH1 Genome
The genome is a linear dsDNA molecule of 30,898 bp, and contains 309-bp inverted repeats and 5′ terminal proteins (Bamford et al. 2005b; Porter and Dyall-Smith 2008). Thus, the genome is most probably replicated by a protein-primed mechanism. This replication strategy has been described in great detail for phage Φ29 (Salas 1984). However, the SH1 genome does not appear to have any ORF with canonical DNA polymerase motifs, and most likely uses a host enzyme (Bamford et al. 2005b). The G + C content of the genome is 68.4%, which is close to that of the host Har. hispanica (63.5%). The genome contains 56 predicted ORFs, which are arranged in at least seven polycistronic operons with tightly regulated promoters and containing in most cases a potential 5′ ribosome binding site (Bamford et al. 2005b; Porter et al. 2008a). Late in the infection cycle the SH1 transcription seems to be characterized by extensive counter-transcription for most of the coding transcripts that may have a function in translational regulation (Porter et al. 2008a).
Among the SH1 ORFs only few sequences are similar to previously identified sequences in the databases (Bamford et al. 2005b). This reflects the fact that only a limited number of archaeal sequences have been determined, and that archaeal genes seem to be significantly different from those of bacterial and eukaryotic origin. Recently, a proviral element (IHP) was found to be integrated in the genome of Haloarcula marismortui (Jalasvuori et al. 2009). The IHP element contains genes similar to the SH1 genes encoding the major capsid proteins and the ATPase (see below). The same genetic module is also found in the archaeal plasmid pHH205 of Hbt. salinarum as well as in bacterial viruses (phage P23-77 and IN93) revealing an evolutionary relationship between bacterial and archaeal genetic elements (Jalasvuori et al. 2009). Additionally, the ORF 17 of SH1 encodes a putative protein with ATPase motifs including classical Walker A and B motifs (Walker et al. 1982) as well as a motif common to internal membrane-containing dsDNA viruses e.g., bacteriophages PRD1 and PM2, eukaryotic viruses Paramecium bursaria Chlorella virus 1 (PBCV-1) and Chilo Iridescent Virus (CIV), and archaeal virus Sulfolobus turreted icosahedral virus (STIV) (Bamford et al. 2005a; Strömsten et al. 2005).
8.5.2 SH1 Virion Structure
Production of large amounts of highly purified viral material has enabled us to study the structure of SH1 by quantitative dissociation studies as well as by electron cryo-microscopy (cryo-EM) and image reconstruction at 9.6-Å resolution (Kivelä et al. 2006; Jäälinoja et al. 2008). The icosahedral SH1 virion has a diameter of 78 nm between facets and is composed of approximately 15 virally encoded structural protein species (VP1-VP15), of which 11 have been identified as virion proteins by protein chemistry (Fig. 8.2; Bamford et al. 2005b; Kivelä et al. 2006). Most of the structural protein species have been localized within the virions and the proteins can be divided into those forming the protein capsid, the vertex complex, and those being associated with the viral membrane (Fig. 8.2; Bamford et al. 2005b; Kivelä et al. 2006; Jäälinoja et al. 2008).
Halophilic icosahedral membrane-containing virus SH1. (a) Schematic picture of the virion architecture showing the location of the major structural proteins (VPs). (b) Cryo-electron microscopy based structure of the SH1 virion viewed down twofold axis of symmetry. Part (b) is reproduced from Jäälinoja et al. (2008). Copyright (2008) National Academy of Sciences, USA
The SH1 spikes are horn-like structures projecting almost 20 nm outwards from the capsid surface (Fig. 8.2). These two-fold symmetric structures, composed of at least two proteins VP3 and VP6, are most probably involved in receptor recognition (Jäälinoja et al. 2008). Also protein VP2 is associated with the SH1 vertex structure. The glycine- and serine-rich VP2 protein with heptapeptide repeat patterns characteristic for coiled-coil proteins suggest that it could be an elongated fiber-like protein appropriate for forming a flexible structure (Bamford et al. 2005b).
In SH1, the membrane lying under the capsid follows its icosahedral shape and there are several interactions between the capsomers and the membrane (Jäälinoja et al. 2008; Fig. 8.2). The viral membrane is composed of neutral lipids and three major archaeal phospholipids: phosphatidylglycerol, phosphatidylglycerosulfate, and phosphatidylglycerosulfate methyl ester (Bamford et al. 2005b). The viral lipid composition is different from that of its host cell indicating that the phospholipids are selectively acquired from the host cytoplasmic membrane during virus maturation. The overall structure of SH1 virion resembles the virion morphology of archaeal Sulfolobus turreted icosahedral viruses STIV and STIV-2 (Happonen et al. 2010; Khayat et al. 2005) and bacteriophages PRD1, Bam35, PM2 and P23-77 (Abrescia et al. 2004, 2008; Laurinmäki et al. 2005; Jaatinen et al. 2008) as well as eukaryotic Paramecium bursaria Chlorella virus 1 (PBCV-1; Nandhagopal et al. 2002). They all have a dsDNA genome encapsidated into a proteinaceous membrane residing in an icosahedral protein capsid. The capsid architecture of SH1 is most similar to that of the bacteriophage P23-77 infecting Thermus thermophilus (Jaatinen et al. 2008). Structurally related viruses having hosts from different domains of life support the hypothesis that viruses are ancient and these two viruses might share a common ancestry dating back to the time before the separation of the three domains of life (Bamford 2003; Krupovič and Bamford 2008).
8.6 Enveloped Viruses Infecting Haloarchaea
The characterization of the new group of pleomorphic viruses was a result of our efforts in looking for new viruses from the hypersaline environments. The first member of this group, the Halorubrum pleomorphic virus 1 (HRPV-1) is also the first archaeal virus reported to have a single-stranded DNA (ssDNA) genome (Pietilä et al. 2009). The group includes at the moment also a dsDNA virus, Haloarcula hispanica pleomorphic virus 1 (HHPV-1, Fig. 8.3a) as well as two putative proviruses, the pHK2, a genetic element previously reported to be a plasmid (Holmes et al. 1995) and a provirus in the genome of Haloferax volcanii (Roine et al. 2010, Fig. 8.3b). The genomes of HRPV-1 and HHPV-1 are circular molecules of 7,048 nt and 8,082 bp, in size, respectively. Previously identified spindle-shaped viruses His1 and His2 (genus Salterprovirus; Bath et al. 2006) are flexible particles containing approximately 15 kb linear dsDNA genomes. The chloroform sensitivity as well as the low buoyant density of His1 and His2 suggest they belong to enveloped viruses (Bath et al. 2006). Although the genome types of pleomorphic viruses and spindle-shaped viruses are different, limited homology is shared at the amino acid sequence level between His2 virus and HHPV-1 and HRPV-1 (see below; Bath et al. 2006). It is worth emphasizing that even though the two pleomorphic viruses are closely related (see below), they contain different types of genomes: HRPV-1 contains a ssDNA genome whereas the HHPV-1 contains a dsDNA genome (Pietilä et al. 2009; Roine et al. 2010). These results conflicts with current viral classification that is based on the genome types and the mode of replication (Baltimore 1971), because in the absence of the sequence data these two viruses having two different genome types would be classified as unrelated viruses (Roine et al. 2010).
The virion architecture and genome alignment of the new group of pleomorphic viruses. (a) A schematic of the pleomorphic virus architecture pointing out the predicted orientation of the two major structural proteins (left). A sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) analysis of the purified HRPV-1 and HHPV-1 virions is shown on the right. Molecular weight standards (kDa) are shown on the right. (b) Genomic comparison of the viruses HRPV-1 and HHPV-1 as well as the putative proviral elements pHK2 and Hfx. volcanii genomic region. The newly identified genomic region in Hmc. mukohataei (DSM12286) shows the same features as the Hfx. volcanii proviral region with a tRNAArg in the 5′ region followed by the ORF1 and a putative integrase in the end of the proviral element. The tRNAArg in Hmc. mukohataei is a split-tRNA. The percentages show the identity of the amino acid sequences of the pairwise alignments. The Hmuk_0952 contains the DUF1424 conserved domain common to proteins involved in replication. It also shows 30% identity with the pHK2 ORF1 which is not indicated in the picture
8.6.1 Genomic Comparison
Similar protein patterns of HRPV-1 and HHPV-1 with two major structural proteins already suggested relatedness (Fig. 8.3a). At the nucleotide sequence level these two genomes only show homology along a very short stretch. However, the genomes are co-linear along the whole genomes and at the amino acid sequence level the highest identity can be found between the small major structural protein VP3 (Fig. 8.3b). The co-linearity can also be found between the viruses and the putative proviruses (Fig. 8.3b; Roine et al. 2010). This region includes ORFs 1, 2, 6 (HHPV-1 ORF5), 7 (HHPV-1 ORF6) and 9 (HHPV-1 ORF7 and HRPV-1 gene 8). ORFs 1 encode putative replication initiation proteins showing rather low identity between each other. They are predicted, however, to serve the same function (Roine et al. 2010). The region starting from ORF2 and ending in ORF9 (in pHK2 and in the Hfx. volcanii genomic region) show amino acid sequence identity higher than 20%. In the genomic alignment, we can see small ORFs that may have been lost in HHPV-1. HRPV-1 still seems to have a homolog of ORF5, but has lost ORF8 that can be found in pHK2 and Hfx. volcanii proviral elements. At the genomic level, HRPV-1 seems to be a closer relative to pHK2 proviral element than to HHPV-1 as judged by the higher number of homologous ORFs as well as by the higher identity of the proteins VP3 and VP4 sequences (Fig. 8.3b, Table 8.1).
The pHK2 and Hfx. volcanii proviral elements are almost identical along the conserved ORF2 to ORF9 region (Fig. 8.3b). After that there have been genomic rearrangements, still preserving the last three predicted ORFs. The translated polypeptides of these three ORFs show high identity. The pHK2 ORF11 and ORF13 and the corresponding ORFs in the Hfx. volcanii genome also encode putative polypeptides that by homology are predicted to be a ΦH-type repressor and an integrase like protein, respectively (Fig. 8.3b).
The pHK2 region spanning from ORF4 to ORF9 can also be found in the genome of the spindle-shaped virus His2 (Bath et al. 2006; Pietilä et al. 2009). The major structural protein of His2 (VP1 encoded by gp29) shows 25% identity to the pHK2 ORF4 encoded polypeptide, which is higher than the identity between the HHPV-1 protein VP4 and pHK2 ORF4 encoded protein (Table 8.1). The rest of the His2 ORFs encoded putative polypeptides show lower homology to their predicted counterparts.
In addition to the homologous genetic elements mentioned above, there are regions in almost all haloarchaeal genomes that show homology to different sets of these genes (Table 8.2). After a closer examination of the genomic databases, an almost complete proviral element was found from the genome of Halomicrobium mukohataei (GenBank CP001688.1). This element contains all the other homologous ORFs found in the Hfx. volcanii proviral element except for the ORF encoding the putative ΦH repressor like protein (Fig. 8.3b). The alignment of this element with the haloarchaeal enveloped viruses and other putative proviral elements further supports the observation that the central region containing the genes encoding the major structural proteins, are the most conserved part of the viruses (Roine et al. 2010).
8.6.2 Particle Architecture and Structural Proteins
The virion structure of the pleomorphic viruses is relatively simple with the genome enveloped by protein-containing membrane. The envelope lipids are acquired from the host cytoplasmic membrane non-selectively and thus the lipid composition is approximately the same as in the host (Pietilä et al. 2010; Roine et al. 2010). There are two major structural proteins, VP3 and VP4 (Fig. 8.3). In addition, protein VP8 has been identified as a minor structural component in HRPV-1 (Pietilä et al. 2009). The smaller major structural protein VP3 is predicted to be a membrane associated protein facing the interior of the virion (Pietilä et al. 2010) whereas VP4 is the spike protein protruding outside and being anchored to the virion envelope by a C-terminal trans-membrane domain (Pietilä et al. 2009, 2010). The HRPV-1 particles are approximately 44 × 55 nm in size and not as uniform in shape as the virions containing a rigid icosahedral protein coat. This can be seen as a very diffuse light scattering band in a rate zonal sucrose gradient, for example. However, the particles of HRPV-1 seem to be more uniform in shape than the HHPV-1 particles when samples are studied by negative staining and transmission electron microscopy (TEM). This may be explained by the fact that the HRPV-1 VP4 is glycosylated but the HHPV-1 VP4 is not (Pietilä et al. 2010; Roine et al. 2010) and thus the surface of the HHPV-1 particle is more prone to the pressure by the different negative stains. Some negative stains seem to radically reduce the infectivity of HHPV-1 (Roine et al. 2010) and at the same time change the morphology of the virion from flexible into round particles that are more uniform in shape (Kukkaro, personal communication). To conclude, the definition of a pleomorphic particle, in our opinion, describes best this virus morphology.
The large major structural proteins (VP4) show very low sequence similarity to each other (Table 8.1). We have hypothesized that the VP4 protein is responsible for the receptor recognition and fusion of the envelope with the host cytoplasmic membrane (Pietilä et al. 2009; Roine et al. 2010). The proteins recognizing the host receptors are usually very diverse due to the selection pressure of the host (Lubbers et al. 1995; Brüssow and Desiere 2001; Saren et al. 2005). HRPV-1 and HHPV-1 VP4 polypeptides are both N-terminally processed and during the morphogenesis of the viral particle, the protein is translocated to the extracellular face of the host membrane (Pietilä et al. 2010). The VP4 molecules also share some putative structural features such as a predicted C-terminal membrane anchor preceded by predicted coiled-coil domain. As coiled-coil domains are usually involved in multimeric interactions, it is probable that the VP4 is multimeric at some stage of host recognition or entry. It is highly probable that VP4 protein interacts with VP3, which contains four predicted trans-membrane domains (Pietilä et al. 2009). Two of these have been experimentally verified suggesting a role as a matrix forming protein (Pietilä et al. 2009, 2010). The interaction of VP3 with the genomic DNA was not excluded, although the experimental results did not indicate any strong interaction to occur (Pietilä et al. 2010). Whether the VP3 proteins act solely as structural components or are active also in the fusion process needs to be determined.
Another predicted protein product showing relatively high homology between the identified viruses and the proviral elements are the ORF9 encoded products, the VP8 protein of HRPV-1 and the putative ORF33 of His2 virus. Many of these show the conserved motifs of P-loop NTPases (Bath et al. 2006; Pietilä et al. 2009), i.e., the Walker A and Walker B elements (Walker et al. 1982). Experimental evidence, however, is needed to show that they are functional. We propose that they may function in the assembly of the viral particles.
As discussed above, the putative ORF1 encoded proteins are all predicted to be replication initiation proteins. The conserved motifs 2 and 3 (Ilyina and Koonin 1992) found in plasmids and viruses replicating their genomes by rolling circle replication (RCR) can also be found in all of them (Roine et al. 2010). This is in good agreement with the fact that the HRPV-1 and HHPV-1 genomes are circular and that pHK2 was initially described as a haloplasmid.
8.7 Conclusions
Haloarchaeal viruses are among the least studied viruses comprising currently only around 20 characterized isolates. Our systematic approach for screening new host isolates and subsequently viruses for them has shown that there are still new types of viruses to be found. Although not as versatile in the particle architecture as the crenarchaeal viruses (Prangishvili et al. 2006), the description of the new haloarchaeal viruses have significantly contributed to the general knowledge of the viral universe. This is true especially in the case of the pleomorphic viruses HRPV-1 and HHPV-1 that are closely related, but have different genome types. The higher order viral classification divides viruses according to the genome type and replication strategy (Baltimore 1971). The pleomorphic viruses thus represent a deviation from this scheme and points to a reconsideration of the higher order taxonomy.
The virion architecture of the haloarchaeal pleomorphic viruses is to some extent reminiscent of the Plasmaviridae family of viruses L2 and L172 that infect Acholeplasma laidlawii (Dybvig et al. 1985). These viruses are enveloped and pleomorphic in structure and have a similar structural protein pattern between each other and also between the haloarchaeal enveloped viruses (Dybvig et al. 1985). L2 contains a circular 12 kb dsDNA genome whereas the L172 genome is a circular ssDNA molecule 14 kb in size. However, only the L2 genome sequence is available (Maniloff et al. 1994) and the relatedness of the plasma viruses cannot be confirmed. The structural relatedness of the plasmaviruses and the pleomorphic viruses would not be surprising when taking into account the fact that both SH1 and the head–tail viruses of haloarchaea have relatives among the bacteriophages.
Our studies together with the ones by Mike Dyall-Smith have shown that the early isolation of only head–tail haloarchaeal viruses does not reflect the virus population in halophilic environment. The isolation of many enveloped viruses correlates better with the results of microscopical examination of environmental samples, suggesting that in terms of morphotypes there is much diversity to be found (Guixa-Boixareu et al. 1996; Oren et al. 1997; Diez et al. 2000; Santos et al. 2007; Sime-Ngando et al. 2010). Although our culture-dependent studies support the observations of the environmental microscopy, not all the VLPs seen by microscopy are necessarily viruses. Viruses are genetic elements capable of packaging their genome into a particle. They are also infective agents and consequently obey Koch’s postulates. This means that the viruses produce new infective progeny highly similar to the one used for the infection. Unfortunately, this can be proven only by the culture-dependent approach. Environmental sequencing has expanded our knowledge of the sequence space to be found from the hypersaline environment. Several limitations in the preparation of the sample such as viruses that do not pass the 0.2-μm filter (Hendrix 2009) or are dissociated by chloroform, and sequences that are toxic to the cloning hosts used pose serious challenges if the entire viral fraction of the sample is to be determined (Tringe and Rubin 2005; Kunin et al. 2008; Wooley et al. 2010). Consequently, the environmental sequencing approach also has a bias that may not be any less significant than the bias in the other approaches. In addition, the sequences that show no matches to databases gain their full value only when similar sequences have been identified in a virus characterized in detail. This usually involves the isolation of the virus. As when mapping the sequences in other environmental niches, the mapping of the hypersaline environment should include all the possible types of approaches from the measurement of the bulk biological processes such as primary production and physico-chemical conditions to environmental microscopy and sequencing as well as the culture-dependent approach of isolating the infective viruses.
It is clear that prokaryotic viruses are important players in the genomic plasticity of their hosts. In hypersaline environment, it is generally agreed that the amount of viruses and their hosts is high (Guixa-Boixareu et al. 1996; Oren et al. 1997; Maturrano et al. 2006; Pedrós-Alió et al. 2000). The lysis of cells due to viral infection, however, was shown to be low (Guixa-Boixareu et al. 1996). This suggests that the popular “kill the winner” phenomenon would not fully operate in hypersaline environments. Rodríguez-Valera and others (Rodriguez-Valera et al. 2009), on the other hand, propose a “kill the winner” type of control by viruses over the dominating host population preserving the microdiversity of the microbial population and generating a pan genome comprised of numerous different, but closely related clones (Papke et al. 2004; Rodriguez-Valera et al. 2009). The existence of proviruses and the numerous remnants of them that are homologous to the pleomorphic viruses suggest a very intimate relationship between the virus and the haloarchaeal host. The proposed high number of the enveloped viruses suggests that these viruses have a great impact in generating the microdiversity using a strategy more benign than the “kill the winner” strategy is. In such an extreme environment it is conceivable that the relationship of the predator and prey are closely connected and dependent on each other. Support for this can also be found in the observed changes of the head–tail virus life cycles in which the higher salinity caused a switch from lysis into proviral mode. As seen in other prokaryotes, proviruses may have dramatic effects on the behavior of their hosts (Brüssow et al. 2004). More studies, however, are needed to elucidate the complex interactions monitoring population dynamics of the viruses and their hosts by taking advantage of the detailed molecular biology data available.
References
Abrescia NG, Cockburn JJ, Grimes JM, Sutton GC, Diprose JM, Butcher SJ, Fuller SD, San Martin C, Burnett RM, Stuart DI, Bamford DH, Bamford JK (2004) Insights into assembly from structural analysis of bacteriophage PRD1. Nature 432:68–74
Abrescia NG, Grimes JM, Kivelä HM, Assenberg R, Sutton GC, Butcher SJ, Bamford JK, Bamford DH, Stuart DI (2008) Insights into virus evolution and membrane biogenesis from the structure of the marine lipid-containing bacteriophage PM2. Mol Cell 31:749–761
Baltimore D (1971) Expression of animal virus genomes. Bacteriol Rev 35:235–241
Bamford DH (2003) Do viruses form lineages across different domains of life? Res Microbiol 154:231–236
Bamford DH, Grimes JM, Stuart DI (2005a) What does structure tell us about virus evolution? Curr Opin Struct Biol 15:655–663
Bamford DH, Ravantti JJ, Rönnholm G, Laurinavičius S, Kukkaro P, Dyall-Smith M, Somerharju P, Kalkkinen N, Bamford JK (2005b) Constituents of SH1, a novel lipid-containing virus infecting the halophilic euryarchaeon Haloarcula hispanica. J Virol 79:9097–9107
Bath C, Cukalac T, Porter K, Dyall-Smith ML (2006) His1 and His2 are distantly related, spindle-shaped haloviruses belonging to the novel virus group, Salterprovirus. Virology 350:228–239
Benlloch S, López-López A, Casamayor EO, Øvreås L, Goddard V, Daae FL, Smerdon G, Massana R, Joint I, Thingstad F, Pedrós-Alió C, Rodríguez-Valera F (2002) Prokaryotic genetic diversity throughout the salinity gradient of a coastal solar saltern. Environ Microbiol 4:349–360
Bolhuis H, te Poele EM, Rodríguez-Valera F (2004) Isolation and cultivation of Walsby’s square archaeon. Environ Microbiol 6:1287–1291
Brüssow H, Desiere F (2001) Comparative phage genomics and the evolution of Siphoviridae: insights from dairy phages. Mol Microbiol 39:213–222
Brüssow H, Canchaya C, Hardt WD (2004) Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiol Mol Biol Rev 68:560–602
Burns DG, Camakaris HM, Janssen PH, Dyall-Smith ML (2004a) Combined use of cultivation-dependent and cultivation-independent methods indicates that members of most haloarchaeal groups in an Australian crystallizer pond are cultivable. Appl Environ Microbiol 70:5258–5265
Burns DG, Camakaris HM, Janssen PH, Dyall-Smith ML (2004b) Cultivation of Walsby’s square haloarchaeon. FEMS Microbiol Lett 238:469–473
Calvo C, de la Paz AG, Bejar V, Quesada E, Ramos-Cormenzana A (1988) Isolation and characterization of phage F9-11 from a lysogenic Deleya halophila strain. Curr Microbiol 17:49–53
Casamayor EO, Massana R, Benlloch S, Øvreås L, Diez B, Goddard VJ, Gasol JM, Joint I, Rodríguez-Valera F, Pedrós-Alió C (2002) Changes in archaeal, bacterial and eukaryal assemblages along a salinity gradient by comparison of genetic fingerprinting methods in a multipond solar saltern. Environ Microbiol 4:338–348
Daniels LL (1917) On the flora of Great Salt Lake. Am Nat 51:499–506
Daniels LL, Wais AC (1990) Ecophysiology of bacteriophage S5100 infecting Halobacterium cutirubrum. Appl Environ Microbiol 56:3605–3608
Daniels LL, Wais AC (1998) Virulence in phage populations infecting Halobacterium curtirubrum. FEMS Microbiol Ecol 25:129–134
Diez B, Antón J, Guixa-Boixereu N, Pedrós-Alió C, Rodríguez-Valera F (2000) Pulsed-field gel electrophoresis analysis of virus assemblages present in a hypersaline environment. Int Microbiol 3:159–164
Dyall-Smith M, Tang SL, Bath C (2003) Haloarchaeal viruses: how diverse are they? Res Microbiol 154:309–313
Dybvig K, Nowak JA, Sladek TL, Maniloff J (1985) Identification of an enveloped phage, mycoplasma virus L172, that contains a 14-kilobase single-stranded DNA genome. J Virol 53:384–390
Gropp F, Palm P, Zillig W (1989) Expression and regulation of Halobacterium halobium phage ΦH genes. Can J Microbiol 35:182–188
Gropp F, Grampp B, Stolt P, Palm P, Zillig W (1992) The immunity-conferring plasmid pΦHL from the Halobacterium salinarium phage ΦH: nucleotide sequence and transcription. Virology 190:45–54
Guixa-Boixareu N, Calderon-Paz JI, Heldal M, Bratbak G, Pedrós-Alió C (1996) Viral lysis and bacterivory as prokaryotic loss factors along a salinity gradient. Aquat Microb Ecol 11:215–227
Happonen LJ, Redder P, Peng X, Reigstad LJ, Prangishvili D, Butcher SJ (2010) Familial relationships in hyperthermo- and acidophilic archaeal viruses. J Virol 84:4747–4754
Hendrix RW (2009) Jumbo bacteriophages. Curr Top Microbiol Immunol 328:229–240
Hendrix RW, Lawrence JG, Hatfull GF, Casjens S (2000) The origins and ongoing evolution of viruses. Trends Microbiol 8:504–508
Holmes ML, Pfeifer F, Dyall-Smith ML (1995) Analysis of the halobacterial plasmid pHK2 minimal replicon. Gene 153:117–121
Ilyina TV, Koonin EV (1992) Conserved sequence motifs in the initiator proteins for rolling circle DNA replication encoded by diverse replicons from eubacteria, eucaryotes and archaebacteria. Nucleic Acids Res 20:3279–3285
Jäälinoja HT, Roine E, Laurinmäki P, Kivelä HM, Bamford DH, Butcher SJ (2008) Structure and host-cell interaction of SH1, a membrane-containing, halophilic euryarchaeal virus. Proc Natl Acad Sci USA 105:8008–8013
Jaatinen ST, Happonen LJ, Laurinmäki P, Butcher SJ, Bamford DH (2008) Biochemical and structural characterisation of membrane-containing icosahedral dsDNA bacteriophages infecting thermophilic Thermus thermophilus. Virology 379:10–19
Jalasvuori M, Jaatinen ST, Laurinavičius S, Ahola-Iivarinen E, Kalkkinen N, Bamford DH, Bamford JK (2009) The closest relatives of icosahedral viruses of thermophilic bacteria are among viruses and plasmids of the halophilic archaea. J Virol 83:9388–9397
Kauri T, Ackermann H-W, Goel U, Kushner DJ (1991) A bacteriophage of a moderately halophilic bacterium. Arch Microbiol 156:435–438
Ken R, Hackett NR (1991) Halobacterium halobium strains lysogenic for phage ΦH contain a protein resembling coliphage repressors. J Bacteriol 173:955–960
Khayat R, Tang L, Larson ET, Lawrence CM, Young M, Johnson JE (2005) Structure of an archaeal virus capsid protein reveals a common ancestry to eukaryotic and bacterial viruses. Proc Natl Acad Sci USA 102:18944–18949
Kivelä HM, Roine E, Kukkaro P, Laurinavičius S, Somerharju P, Bamford DH (2006) Quantitative dissociation of archaeal virus SH1 reveals distinct capsid proteins and a lipid core. Virology 356:4–11
Klein R, Baranyi U, Rössler N, Greineder B, Scholz H, Witte A (2002) Natrialba magadii virus ΦCh1: first complete nucleotide sequence and functional organization of a virus infecting a haloalkaliphilic archaeon. Mol Microbiol 45:851–863
Krupovič M, Bamford DH (2008) Virus evolution: how far does the double β-barrel viral lineage extend? Nat Rev Microbiol 6:941–948
Krupovič M, Forterre P, Bamford DH (2010) Comparative analysis of the mosaic genomes of tailed archaeal viruses and proviruses suggests common themes for virion architecture and assembly with tailed viruses of bacteria. J Mol Biol 397:144–160
Kukkaro P, Bamford DH (2009) Virus-host interactions in environments with a wide range of ionic strengths. Environ Microbiol Rep 1:71–77
Kunin V, Copeland A, Lapidus A, Mavromatis K, Hugenholtz P (2008) A bioinformatician’s guide to metagenomics. Microbiol Mol Biol Rev 72:557–578
Laurinmäki PA, Huiskonen JT, Bamford DH, Butcher SJ (2005) Membrane proteins modulate the bilayer curvature in the bacterial virus Bam35. Structure 13:1819–1828
Lubbers MW, Waterfield NR, Beresford TP, Le Page RW, Jarvis AW (1995) Sequencing and analysis of the prolate-headed lactococcal bacteriophage c2 genome and identification of the structural genes. Appl Environ Microbiol 61:4348–4356
Maniloff J, Kampo GJ, Dascher CC (1994) Sequence analysis of a unique temperature phage: mycoplasma virus L2. Gene 141:1–8
Maturrano L, Santos F, Rosselló-Mora R, Antón J (2006) Microbial diversity in Maras salterns, a hypersaline environment in the Peruvian Andes. Appl Environ Microbiol 72:3887–3895
Nandhagopal N, Simpson AA, Gurnon JR, Yan X, Baker TS, Graves MV, Van Etten JL, Rossmann MG (2002) The structure and evolution of the major capsid protein of a large, lipid-containing DNA virus. Proc Natl Acad Sci USA 99:14758–14763
Oren A (2002) Halophilic microorganisms and their environments. Kluwer Academic Publishers, Dordrecht
Oren A (2008) Microbial life at high salt concentrations: phylogenetic and metabolic diversity. Saline Systems 4:2
Oren A, Bratbak G, Heldal M (1997) Occurrence of virus-like particles in the Dead Sea. Extremophiles 1:143–149
Pagaling E, Haigh RD, Grant WD, Cowan DA, Jones BE, Ma Y, Ventosa A, Heaphy S (2007) Sequence analysis of an Archaeal virus isolated from a hypersaline lake in Inner Mongolia, China. BMC Genomics 8:410
Papke RT, Koenig JE, Rodríguez-Valera F, Doolittle WF (2004) Frequent recombination in a saltern population of Halorubrum. Science 306:1928–1929
Pauling C (1982) Bacteriophages of Halobacterium halobium: isolated from fermented fish sauce and primary characterization. Can J Microbiol 28:916–921
Pedrós-Alió C, Caldéron-Paz JI, MacLean MH, Medina G, Marrasé C, Gasol JM, Guixa-Boixereu N (2000) The microbial food web along salinity gradients. FEMS Microbiol Ecol 32:143–155
Pfeifer F, Weidinger G, Goebel W (1981) Genetic variability in Halobacterium halobium. J Bacteriol 145:375–381
Pietilä MK, Roine E, Paulin L, Kalkkinen N, Bamford DH (2009) An ssDNA virus infecting archaea: a new lineage of viruses with a membrane envelope. Mol Microbiol 72:307–319
Pietilä MK, Laurinavicius S, Sund J, Roine E, Bamford DH (2010) The single-stranded DNA genome of novel archaeal virus Halorubrum pleomorphic virus 1 is enclosed in the envelope decorated with glycoprotein spikes. J Virol 84:788–798
Porter K, Dyall-Smith M (2006) The isolation and study of viruses of halophilic microorganisms. In: Rainey FA, Oren A (eds) Methods in microbiology, vol 35, Extremophiles. Elsevier/Academic, London, pp 681–702
Porter K, Dyall-Smith ML (2008) Transfection of haloarchaea by the DNAs of spindle and round haloviruses and the use of transposon mutagenesis to identify non-essential regions. Mol Microbiol 70:1236–1245
Porter K, Kukkaro P, Bamford JK, Bath C, Kivelä HM, Dyall-Smith ML, Bamford DH (2005) SH1: a novel, spherical halovirus isolated from an Australian hypersaline lake. Virology 335:22–33
Porter K, Russ BE, Dyall-Smith ML (2007) Virus-host interactions in salt lakes. Curr Opin Microbiol 10:418–424
Porter K, Russ BE, Yang J, Dyall-Smith ML (2008a) The transcription programme of the protein-primed halovirus SH1. Microbiology 154:3599–3608
Porter K, Russ BE, Thorburn AN, Dyall-Smith ML (2008b) Viruses infecting Euryarchaea. In: Mahy BWJ, van Regenmortel MHV (eds) Encyclopedia of virology, vol 5. Elsevier, Oxford, pp 411–423
Prangishvili D, Forterre P, Garrett RA (2006) Viruses of the Archaea: a unifying view. Nat Rev Microbiol 4:837–848
Rainey FA, Oren A (2006) Extremophile microorganisms and the methods to handle them. In: Rainey FA, Oren A (eds) Methods in microbiology, vol 35, Extremophiles. Elsevier/Academic, London, pp 1–25
Reiter WD, Zillig W, Palm P (1988) Archaebacterial viruses. Adv Virus Res 34:143–188
Rodriguez-Valera F, Martin-Cuadrado AB, Rodriguez-Brito B, Pašić L, Thingstad TF, Rohwer F, Mira A (2009) Explaining microbial population genomics through phage predation. Nat Rev Microbiol 7:828–836
Roine E, Kukkaro P, Paulin L, Laurinavičius S, Domanska A, Somerharju P, Bamford DH (2010) New, closely related haloarchaeal viral elements with different nucleic acid types. J Virol 84:3682–3689
Rössler N, Klein R, Scholz H, Witte A (2004) Inversion within the haloalkaliphilic virus ΦCh1 DNA results in differential expression of structural proteins. Mol Microbiol 52:413–426
Salas M (1984) A new mechanism for the initiation of replication of Φ29 and adenovirus DNA: priming by the terminal protein. Curr Top Microbiol Immunol 109:89–106
Santos F, Meyerdierks A, Peña A, Róssello-Mora R, Amann R, Antón J (2007) Metagenomic approach to the study of halophages: the environmental halophage 1. Environ Microbiol 9:1711–1723
Santos F, Yarza P, Parro V, Briones C, Antón J (2010) The metavirome of a hypersaline environment. Environ Microbiol 12:2965–2976
Sapienza C, Rose MR, Doolittle WF (1982) High-frequency genomic rearrangements involving archaebacterial repeat sequence elements. Nature 299:182–185
Saren AM, Ravantti JJ, Benson SD, Burnett RM, Paulin L, Bamford DH, Bamford JK (2005) A snapshot of viral evolution from genome analysis of the tectiviridae family. J Mol Biol 350:427–440
Schnabel H (1984) An immune strain of Halobacterium halobium carries the invertible L segment of phage ΦH as a plasmid. Proc Natl Acad Sci USA 81:1017–1020
Schnabel H, Schramm E, Schnabel R, Zillig W (1982a) Structural variability in the genome of phage ΦH of Halobacterium halobium. Mol Gen Genet 188:370–377
Schnabel H, Zillig W, Pfaffle M, Schnabel R, Michel H, Delius H (1982b) Halobacterium halobium phage ΦH. EMBO J 1:87–92
Schnabel H, Palm P, Dick K, Grampp B (1984) Sequence analysis of the insertion element ISH1.8 and of associated structural changes in the genome of phage ΦH of the archaebacterium Halobacterium halobium. EMBO J 3:1717–1722
Sime-Ngando T, Lucas S, Robin A, Tucker KP, Colombet J, Bettarel Y, Desmond E, Gribaldo S, Forterre P, Breitbart M, Prangishvili D (2011) Diversity of virus-host systems in hypersaline Lake Retba, Senegal. Environ Microbiol 13: no. doi:10.1111/j.1462-2920.2010.02323.x
Simsek M, DasSarma S, RajBhandary UL, Khorana HG (1982) A transposable element from Halobacterium halobium which inactivates the bacteriorhodopsin gene. Proc Natl Acad Sci USA 79:7268–7272
Stedman KM, Porter K, Dyall-Smith ML (2009) The isolation of viruses infecting Archaea. In: Wilhelm SW, Weinbauer MG, Suttle CA (eds) Manual of aquatic viral ecology. American Society of Limnology and Oceanography, Waco, TX, pp 57–64
Stolt P, Zillig W (1992) In vivo studies on the effects of immunity genes on early lytic transcription in the Halobacterium salinarium phage ΦH. Mol Gen Genet 235:197–204
Stolt P, Zillig W (1993a) In vivo and in vitro analysis of transcription of the L region from the Halobacterium salinarium phage ΦH: definition of a repressor-enhancing gene. Virology 195:649–658
Stolt P, Zillig W (1993b) Antisense RNA mediates transcriptional processing in an archaebacterium, indicating a novel kind of RNase activity. Mol Microbiol 7:875–882
Strömsten NJ, Bamford DH, Bamford JK (2005) In vitro DNA packaging of PRD1: a common mechanism for internal-membrane viruses. J Mol Biol 348:617–629
Suttle CA (2007) Marine viruses – major players in the global ecosystem. Nat Rev Microbiol 5:801–812
Tang SL, Nuttall S, Ngui K, Fisher C, Lopez P, Dyall-Smith M (2002) HF2: a double-stranded DNA tailed haloarchaeal virus with a mosaic genome. Mol Microbiol 44:283–296
Torsvik T, Dundas ID (1974) Bacteriophage of Halobacterium salinarium. Nature 248:680–681
Torsvik T, Dundas ID (1980) Persisting phage infection in Halobacterium salinarium str. 1. J Gen Virol 47:29–36
Tringe SG, Rubin EM (2005) Metagenomics: DNA sequencing of environmental samples. Nat Rev Genet 6:805–814
Ventosa A, Oren A (1996) Halobacterium salinarum nom. corrig., a name to replace Halobacterium salinarium (Elazari-Volcani) and to include Halobacterium halobium and Halobacterium cutirubrum. Int J Syst Bacteriol 46:361
Vogelsang-Wenke H, Oesterhelt D (1988) Isolation of a halobacterial phage with a fully cytosine-methylated genome. Mol Gen Genet 211:407–414
Wais AC, Kon M, MacDonald RE, Stollar BD (1975) Salt-dependent bacteriophage infecting Halobacterium cutirubrum and H. halobium. Nature 256:314–315
Walker JE, Saraste M, Runswick MJ, Gay NJ (1982) Distantly related sequences in the α- and β-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J 1:945–951
Walsby AE (1980) A square bacterium. Nature 238:69–71
Weinbauer MG (2004) Ecology of prokaryotic viruses. FEMS Microbiol Rev 28:127–181
Wilkansky B (1936) Life in the Dead Sea. Nature 138:467
Witte A, Baranyi U, Klein R, Sulzner M, Luo C, Wanner G, Kruger DH, Lubitz W (1997) Characterization of Natronobacterium magadii phage ΦCh1, a unique archaeal phage containing DNA and RNA. Mol Microbiol 23:603–616
Wooley JC, Godzik A, Friedberg I (2010) A primer on metagenomics. PLoS Comput Biol 6:e1000667
Acknowledgments
This work was supported by Academy of Finland Centre of Excellence Program in Virus Research grant 11296841 (2006–2011), by the University of Helsinki Three Year Grant 2010–2012 to E.R., and by the Academy of Finland grant (1127665) to H.M.O. We acknowledge Dennis Bamford for critical comments on our manuscript and Petra Kukkaro for contributing to the studies on the transmission electron microscopy of the pleomorphic viruses.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Roine, E., Oksanen, H.M. (2011). Viruses from the Hypersaline Environment. In: Ventosa, A., Oren, A., Ma, Y. (eds) Halophiles and Hypersaline Environments. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-20198-1_8
Download citation
DOI: https://doi.org/10.1007/978-3-642-20198-1_8
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-20197-4
Online ISBN: 978-3-642-20198-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)