Abstract
We present a section boundary detection framework specifically for clinical dictations. Detection is cast as a semi-supervised binary tagging problem and solved using a neural network model composed of a stack of embeddings, unidirectional long-short term memory units (LSTMs), and sigmoid outputs. Physicians’ dictations documenting clinical encounters are typically transcribed using automatic speech recognition (ASR) followed by a post-processor (PP) to transform the raw text into written reports. Section boundary detection can be performed directly upon the raw text to better anticipate the post-processing stage: we describe an architecture for real-time (“live”) ASR use in which sections detected by the tagger are sent individually to a machine translation-based PP (for which continuous execution in real time would not be possible). Our implementation of section detection makes viable the use of a sophisticated machine learning PP in a live dictation paradigm.
N. Sadoughi and G.P. Finley—The authors made equal contributions to this paper.
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Clinical documentation in the form of written reports virtually always follows a structured format, with various types of information primarily constrained to the appropriate (and often clearly identified) sections of the note. Due to the ubiquity of this organizational style, a common first step for information extraction or data mining of clinical text is to detect and classify sections [3, 9].
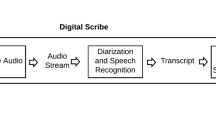
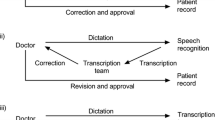
Medical records can be produced and entered into a database using one of several strategies. One means adopted by many physicians is to dictate a description of a clinical encounter, which is then transcribed, converted into a report by either a human transcriber or an automated system, and finally sent back to the physician for approval. For automated systems, automatic speech recognition (ASR) converts the spoken words to their text representations, and then a post-processor (PP) is used for formatting the raw text to create the final report (by adding punctuation, casing, inserting section headers, new lines, numbers for numeral items, etc). In the dictation modality, section headers are sometimes spoken explicitly and sometimes not; in the latter case, the transcriber must rely on context to insert them. In the case of using automated systems, the problem is complicated by possible ASR errors. Thus, detecting actual section boundaries in dictations themselves can be difficult.
To the best of our knowledge, nobody has looked at medical dictation section boundary detection as a stand-alone problem; previous studies focus on converting the dictations to the final report to appear in the electronic medical record (EMR) [5, 8]. While this process does perform section detection implicitly, it does not make that information available at an early stage for other processes to rely on. Determining these boundaries early has several potential advantages.
One possible use case is re-scoring ASR lattices using deep language models (LMs). Widely used ASR frameworks only permit building a decoding graph from n-gram LMs, which cannot model long-term dependencies. Deep recurrent LMs are typically deployed by re-scoring the decoding lattice, possibly changing the best ASR hypothesis obtained by an n-gram LM. Medical dictations typically consist of several minutes of continuous speech, and splitting these long dictations to shorter independent segments can expedite the rescoring process.
Real-time access to the report of the physician’s dictation is very valuable in a fast-paced health care environment, where a patient’s condition can change rapidly. Therefore, another use case for identifying section boundaries, which we investigate in this paper, is to use section breaks to segment the dictation to enable real time use of a PP in a live ASR setting. Live ASR provides physicians with immediate access to the output of the ASR system, as opposed to batch ASR, where the dictation has to pass through a separate system off-line and come back for review later. We recently presented a successful PP based on statistical machine translation [5] called the MTPP, which significantly reduces error rates from a different hybrid rule- and machine learning-based PP but requires significantly more time to run, especially for longer inputs. This added runtime is not an issue in batch ASR, as it is still much faster than the recognition itself. In a live environment, however, the MTPP is too slow to be re-run after every word. Moreover, the MTPP can benefit from having the full context of a finished section during translation, whereas the MTPP hypothesis for a dictation currently in mid-section will often have errors due to unusual right-edge context.
On the other hand, if we split the incoming dictation by sections, a completely dictated section can be sent to the MTPP to run in the background while the physician dictates the next section. After a few seconds, when machine translation has finished, the raw text is replaced with its post-processed output in a manner that is not distracting to the user. In short, our method of section boundary detection at the point of dictation enables the use of sophisticated data-driven natural language processing (NLP) techniques in conjunction with large vocabulary continuous ASR (LVCASR) all in real time.
2 Related Work
There are several previous studies on text segmentation and classification in clinical records. For instance, Denny et al. [3] devised an algorithm using NLP techniques and Naïve Bayes algorithm to find section boundaries and identify section headers. Ginter et al. [6] proposed to use hidden Markov models (HMMs) for segmenting and classifying the sections. Tepper et al. [14] presented algorithms based on maximum entropy (ME) and beam search to simultaneously segment and classify sections. Dai et al. [2] presented a conditional random field (CRF) classifier for section header tagging, assuming that the headers are present in the text. These studies were focused on structural reports, whereas our study considers raw medical dictations. There are previous studies which use raw medical dictations as their input. For instance, Jancsary et al. [8] presented a statistical framework based on CRFs for transforming medical dictations into clinical reports, simultaneously identifying structures such as sections, headers and lists. However, our study is focused solely on finding the section boundaries from dictations, which can be useful as a preprocessing step for many of these tasks, by sending the detected sections instead of the full dictation to these PPs.
3 Resources
3.1 Datasets for Section Boundary Detection
For training our tagger, we explore the use of transcripts (i.e., the ideal ASR case), ASR hypotheses, or a combination of both. We use a dataset of 18,769 medical reports, with their parallel ASR hypotheses as given by our ASR system [4]. Our ASR is composed of an acoustic model (AM) built by a neural networks-based methodology to predict phonemes from acoustic features and a LM which is a 3- or 4-gram statistical model tailored with methods of interpolation and pruning, used to address the large medical vocabulary challenge. Since manual transcription is costly, we have a smaller dataset of 9,073 medical reports with their parallel manual transcriptions. We divide these randomly into training and testing sets, making sure that the test sets for both of these corpora are attributed to the same dictations.
We detect the sections in the manually prepared reports using regular expressions (to detect the section headers), which is very reliable for the corpus of reports we used due to regular adherence to a style guide specifying the form of section headers. To find the break points in the corresponding ASR hypotheses, we used dynamic programming to find the word alignments between them. Prior to doing so, we processed reports to represent all punctuation as tokens and to include a dummy token for the section boundary itself. During alignment, we explicitly disallowed “substitution” as an edit to account for this dummy token; it had to be considered an“insertion.” The place of this insertion was then marked as the section boundary in the dictation. (Additionally, we considered certain words to be equivalent for alignment purposes—for example, ‘examination’ and ‘exam’, as it is commonly dictated.)
Figure 1 illustrates an example of an ASR hypothesis and its parallel report with section boundaries marked using this algorithm. Table 1 provides the details of our datasets in terms of the number of documents, total number of tokens, the number of unique tokens (i.e., types), and the number of sections.
Note that ASR hypotheses have the advantage of being more similar to the test domain during real-time application, which can help the tagger contend with common mistakes of ASR, and they are less costly to obtain than transcripts. On the other hand, as transcripts are more accurate than ASR hypotheses, they resemble the reports more, and therefore the alignment algorithm is more accurate for them. Using a mix of both for training the tagger, it was hypothesized, might include the advantages of both.
3.2 Dataset for MTPP Evaluation
We eventually use our tagger to split the dictation into pieces and pass them through the MTPP. Note, however, that the statistical machine translation model used in the MTPP was trained on full dictations that were not segmented by section. Thus, we need to evaluate the quality of the final report when the MTPP is fed with smaller segments.
We hypothesized also that a translation model trained on shorter sections may perform better on sections during testing. For this model (hereafter, MTPP-SEC), we use the same data used to train and tune the MTPP [5], with one key difference: when building parallel training samples, ASR hypothesis samples with more than 20 tokens are passed through the tagger for further splitting. The split sections are aligned to the report phrases using the alignment method explained in Sect. 3.1. The bitexts of the tuning set are likewise split using the same tagger and the alignment method; thus, the hyper-parameters of the MT model are set according to the expectation of translating individual sections.
We use a corpus of 1,172 medical reports, along with the ASR hypotheses of their associated dictations, for evaluation of PP live usage. We divide this corpus into development and test as shown in Table 2. Note that, for setting the hyper-parameter of the MTPP in Sect. 5.4, we rely on the development set; the test set is used only for reporting the results in this paper.
4 Approach
We approach the problem of detecting section boundaries as a binary sequence tagging problem over tokens, where a boundary token is labeled as 1 and all others 0. To accomplish this, we use the popular RNN extension, LSTM [7]. LSTM solves the problem with vanishing or exploding gradient in vanilla RNN by using a cell unit which keeps track of both short- and long- term dependencies. As there are short- and long- term dependencies in medical dictations which can help in detecting the section boundaries, LSTM is a natural fit for modeling this problem. Our model is composed of an embedding layer, followed by an LSTM layer, and finally an output layer of sigmoid units shared across all the time frames. The model is trained to minimize binary cross entropy with Adam optimizer [10]. We build all models using Keras [1] with a TensorFlow backend.
For many applications, researchers have obtained better results with bidirectional LSTMs, which use separate forward and backward recurrent layers whose outputs are concatenated as the input to the top layer, thus allowing consideration of future context when decoding. This is not appropriate for our problem, however, as we have to operate on incomplete dictations where future frames are not available. Therefore, we rely on unidirectional LSTMs (although note that, in pilot experiments, we achieved slightly better results when using bidirectional LSTMs). This allows the tagger to be updated one frame at a time, enabling very fast real-time decoding. A simplified architecture of our model is given in Fig. 2.
We experimented with initializing the embedding layer either randomly (emb: random) or with pre-trained embeddings. In the case of pre-trained embeddings, we use the original implementation of the word2vec package [11, 12] and consider continuous bag-of-words embeddings with a size of 200, trained for 15 iterations over a window of 8 words with no minimum count. When using pre-trained embeddings, we experimented with fixing the embeddings (emb: freeze) or allowing back-propagation to fine-tune them during training (emb: fine-tune).
5 Experiments
We evaluate the performance of the tagger in terms of precision, recall and F1-score of the split point. Pevzner and Hearst [13] proposed an alternative metric for assessing segmentation, called WindowDiff (WD), which uses a sliding window to count the errors, penalizing the near misses with a lower weight. When the detected segmentation matches the ground truth segmentation, WD becomes zero. Since this is a segmentation problem, we also report WD, setting the window size as the average segment lengths of the ground truth.
5.1 Word Embeddings
We compare the three embedding settings (random, freeze, fine-tune) when training the tagger on the ASR hypotheses train set. Table 3 shows the evaluation on the test set of ASR hypotheses. According to the results, the model performs better in terms of precision, recall, F1-score, and WD when using the pre-trained embeddings, compared to using random initialization (proportion tests for WD, random vs. freeze: \(\chi ^2=829.3, p<0.001\) and random vs. fine-tune: \(\chi ^2=892.7, p<0.001\)). Moreover, fine-tuning the pre-trained embeddings may also slightly enhance the result.
5.2 Impact of ASR Errors
Note that ASR errors can contribute to errors made by the alignment algorithm (Sect. 3.1). These errors can then propagate to the tagger, as the training samples may not be consistent. Furthermore, during inference, ASR errors unseen during training can also increase the errors made by the tagger. Therefore, we perform another set of experiments using the transcription sets. Due to the embeddings results described in Sect. 5.1, we exclusively used pre-trained embeddings with fine-tuning during training.
Precision, recall, and F1-scores are summarized in Table 4. The results of testing the models on transcripts are only provided for analysis purposes, since they are not the input of the model in real-world application. It appears that the alignment algorithm finds more sections when using transcripts compared to ASR hypotheses—refer to the last two rows of Table 1—suggesting that ASR errors prevent the alignment from finding some of the sections in the ASR hypotheses. We speculate this may be the reason that the model trained on transcripts gives higher recall on the ASR hypotheses test set, and the model trained on ASR hypotheses gives lower recall on the transcripts. On the other hand, the models trained on ASR hypotheses or the mix of ASR hypotheses and transcripts have the advantage of overcoming some of the more common ASR mistakes, and therefore give higher precision and lower WD on ASR hypotheses (proportion tests for WD, training on hyp vs. tra: \(\chi ^2 =378.2, p<0.001\), and for training on hyp vs. both: \(\chi ^2 =343.9, p<0.001\)). The best model for ASR hypotheses seems to be the one trained with both datasets (hyp + tra), which gives better performance in terms of all metrics compared with the model trained with only ASR hypotheses and higher precision and F1-score compared to the model trained on transcripts. We rely on this model for experiments with the PP in Sect. 5.4.
5.3 Comparison with Baseline
CRFs have been used by previous studies for section header tagging [2, 8]. As a baseline for our section tagger, we train a CRF model, using uni-gram and bi-gram features, with a minimum count of one optimized with L2 regularized stochastic gradient descent. We set the L2 regularization coefficient as 0.01, and train the model for 100 epochs. This model achieves a precision of 0.832, recall of 0.587, F1-score of 0.688 and WD of 0.1580. This evaluations shows that the our RNN based section tagger outperforms the CRF baseline (proportion tests for WD, \(\chi ^2 =6631.4: p<0.001\)).
5.4 Evaluation of Live Use of MTPP
We perform experiments to evaluate the effect of using the MTPP in a live ASR setting, in which the detected sections are passed through the MTPP separately, compared to batch mode, in which the full report is passed through the MTPP. Recall that the MTPP is trained on bitexts that have not been separated by section, and therefore splitting the task may influence the translation accuracy of the final report. Therefore, we also evaluate the use of MTPP-SEC and compare that with MTPP.
We use the tagger to detect the section boundaries, send these sections as input for the PP sequentially, and concatenate the results to yield the full report. We use a test set of 597 unseen ASR hypotheses with their parallel reports (Sect. 3.2) and measure the accuracy of PP in terms of post-processor error rate (PER). PER is measured similar to word error rate (WER) but takes punctuation, case, and certain whitespace characters into account. PER is a very harsh and honest metric as it combines ASR errors with errors in post-processing, thus providing some estimation of amount of manual work that would be needed to correct the output.
Table 5 provides the PER for different conditions. As expected, applying the MTPP independently on sections degrades the quality of the final report, which is reflected in the increase in PER (proportion test: \(\chi ^2=138.05, p < 0.001\)). Note also, however, that when the MTPP is applied on sections separately, it reduces the processing time required per token (z-test: \(z=60.0, p<0.001\)), as the running time of the MTPP is not linear with respect to the input length. (Splitting into sections also carries the advantage that processing the sections can be easily parallelized; our tests do not take this into account.) MTPP-SEC increases the PER compared to MTPP (proportion test: \(\chi ^2=26.00, p < 0.001\)), suggesting that the model trained on shorter segments may have some shortcomings. Firstly, building the phrase table based on discrete sections may result in MTPP-SEC not capturing some of edits around section boundaries—and these are precisely the places where many edits occur (formatting headers, e.g.). Compared with MTPP, the size of phrase table is reduced (17M vs 35M entries), which does lead to reduced processing time per token (z-test: \(z=4.9, p<0.001\)). Secondly, MTPP-SEC relies on parallel sections from ASR hypotheses and reports, which are subject to tagging and alignment errors.
Since MTPP-SEC does not improve the PER compared to MTPP, in order to compensate for the errors introduced by the MTPP, we propose to include a window of n tokens from both preceding and following sections. Since these overlapping tokens are processed twice, we use a post-editing process to remove the extra tokens at the end of the previously formatted section and the extra tokens in the beginning of the newly formatted section. We experimented with n from \(\{3, 4, ..., 7\}\), choosing \(n=5\), since after that PER does not decrease much on the development set, but the processing time per token increases. This reduces the PER on the test set very effectively (proportion test: \(\chi ^2=61.59, p<0.001\)) at the cost of slightly increased processing time (z-test: \(z=2.0, p=0.04\)), as some of the tokens are fed to MTPP twice. Adding overlaps is an effective way to mitigate increased MT errors because many of the edits made during the translation process occur around these boundaries—formatting the header, adding newlines around boundaries, starting numerical items from the beginning of a section, etc. Providing just a few tokens of context helps the MTPP achieve much better results.
6 Conclusion
We have shown that section detection in medical dictations is a tractable problem with deep learning methods. Splitting sections at the point of dictation, as opposed to the report itself, is not a task that has been well explored in the literature. Nevertheless, it has many potential applications; in the present study we investigated the use of section detection to partition a dictation into sections in a live medical ASR setting, passing each section through a sophisticated post-processor for formatting while the speaker continues dictating the next section. This can be done with minimal impact on error rates, thus enabling the use of a non-instantaneous ASR post-processing stage in a live paradigm, giving access to all the respective advantages of machine translation-driven post-processing and real-time ASR.
References
Chollet, F.: Keras (2015). https://github.com/fchollet/keras
Dai, H.J., Syed-Abdul, S., Chen, C.W., Wu, C.C.: Recognition and evaluation of clinical section headings in clinical documents using token-based formulation with conditional random fields. BioMed Res. Int. 2015, 10 (2015)
Denny, J.C., Spickard III, A., Johnson, K.B., Peterson, N.B., Peterson, J.F., Miller, R.A.: Evaluation of a method to identify and categorize section headers in clinical documents. J. Am. Med. Inform. Assoc. 16(6), 806–815 (2009)
Edwards, E., Salloum, W., Finley, G.P., Fone, J., Cardiff, G., Miller, M., Suendermann-Oeft, D.: Medical speech recognition: reaching parity with humans. In: Karpov, A., Potapova, R., Mporas, I. (eds.) SPECOM 2017. LNCS (LNAI), vol. 10458, pp. 512–524. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-66429-3_51
Finley, G., Salloum, W., Sadoughi, N., Edwards, E., Robinson, A., Brenndoerfer, M., Axtmann, N., Miller, M., Suendermann-Oeft, D.: Automated preamble detection in dictated medical reports. In: Proceedings of the 2018 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (2018)
Ginter, F., Suominen, H., Pyysalo, S., Salakoski, T.: Combining hidden Markov models and latent semantic analysis for topic segmentation and labeling: method and clinical application. Int. J. Med. Inform. 78(12), e1–e6 (2009)
Hochreiter, S., Schmidhuber, J.: Long short-term memory. Neural Comput. 9(8), 1735–1780 (1997)
Jancsary, J., Matiasek, J., Trost, H.: Revealing the structure of medical dictations with conditional random fields. In: Proceedings of the Conference on Empirical Methods in Natural Language Processing, pp. 1–10. Association for Computational Linguistics (2008)
Jiang, M., Wu, Y., Shah, A., Priyanka, P., Denny, J.C., Xu, H.: Extracting and standardizing medication information in clinical text-the medex-uima system. AMIA Summits Transl. Sci. Proc. 2014, 37 (2014)
Kingma, D.P., Ba, J.: Adam: a method for stochastic optimization. arXiv preprint arXiv:1412.6980 (2014)
Mikolov, T., Chen, K., Corrado, G., Dean, J.: Efficient estimation of word representations in vector space. arXiv preprint arXiv:1301.3781 (2013)
Mikolov, T., Yih, W.T., Zweig, G.: Linguistic regularities in continuous space word representations. In: Proceedings of the 2013 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, pp. 746–751 (2013)
Pevzner, L., Hearst, M.A.: A critique and improvement of an evaluation metric for text segmentation. Comput. Linguist. 28(1), 19–36 (2002)
Tepper, M., Capurro, D., Xia, F., Vanderwende, L., Yetisgen-Yildiz, M.: Statistical section segmentation in free-text clinical records. In: LREC, pp. 2001–2008 (2012)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Switzerland AG
About this paper
Cite this paper
Sadoughi, N. et al. (2018). Detecting Section Boundaries in Medical Dictations: Toward Real-Time Conversion of Medical Dictations to Clinical Reports. In: Karpov, A., Jokisch, O., Potapova, R. (eds) Speech and Computer. SPECOM 2018. Lecture Notes in Computer Science(), vol 11096. Springer, Cham. https://doi.org/10.1007/978-3-319-99579-3_58
Download citation
DOI: https://doi.org/10.1007/978-3-319-99579-3_58
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-99578-6
Online ISBN: 978-3-319-99579-3
eBook Packages: Computer ScienceComputer Science (R0)