Abstract
Chromosome data and characterization by fluorescent banding, silver nucleolar organizer region staining (AgNOR), and fluorescence in situ hybridization (FISH) are compiled in this chapter, together with estimations of nuclear DNA content of Capsicum species. To date, the diploid chromosome number of 77.8% of the species in the genus has been recorded. The chromosome number distinguishes two groups of species, one with 2n = 2x = 24 and the other with 2n = 2x = 26. Only two clades, Andean and Atlantic Forest, possess the chromosome number of 2n = 26. A physical chromosome map with heterochromatin distribution besides 5S and active and inactive 45S ribosomal genes (rDNA) of 12 Capsicum taxa was constructed using fluorescent banding, AgNOR and FISH. The chromosome banding pattern with fluorochromes chromomycin A3 and 4′-6-diamidino-2-phenylindole (CMA/DAPI) reveals number of bands, distribution and content of heterochromatin, and FISH reports the localization of 5S and active and inactive 45S rDNA. Both methods are specific and, together with morphological characters, are instrumental for identifying taxa in Capsicum. AgNOR method informs the number, size, and position of just active NORs. Additionally, nuclear DNA content was estimated for nine diploid species of Capsicum by flow cytometry. Genome size displays significant variation between but not within species and contributes to their taxonomic grouping.
The Author E. A. Moscone was deceased.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
The genus Capsicum (Solanaceae, subtribe Capsiceae; Olmstead et al. 2008; Särkinen et al. 2013), with 36 variable species (Carrizo García et al. 2016), is a small increasing genus from tropical and temperate areas in America, distributed from southern Mexico to central Argentina. Cultivation of sweet and hot chili peppers has great economic implication, since these vegetables and spices are highly consumed worldwide. The most important cultivated species grow around the world and belong to the Capsicum annuum complex (C. annuum, C. chinense, and C. frutescens), and two other cultivated species are predominantly regionally consumed in Latin America, C. pubescens and C. baccatum (Pickersgill 1997; Scaldaferro et al. 2018). Their wild relatives originated from Central and South America and were domesticated by American natives at least 6000 years before present (Pickersgill 1984; Eshbaugh 1993; Perry et al. 2007; Piperno 2011; Scaldaferro et al. 2018).
Cultivated and wild Capsicum species have been usually characterized by corolla color, conforming ʽwhite- and purple-flowered groups.’ However, this could not be considered to describe the flower color of more distantly wild species and the genus as a whole, since some species exhibit single-colored flowers, i.e., white, cream, yellow, ocher, pink, lilac, or purple–violet (e.g., Capsicum chacoense, C. friburgense, C. rhomboideum), whereas others have different color combinations in lobules, throat, and tube, often including spots of various colors which makes species delimitation complex (e.g., Capsicum coccineum, C. hunzikerianum, C. parvifolium; Hunziker 2001; Barboza and Bianchetti 2005) (see Table 4.1).
Fruit pungency is a characteristic of the genus due to a group of compounds called ʽcapsaicinoidsʼ which produce the organoleptic heat sensation, and are exclusive for Capsicum (Bosland 1996). More than 20 capsaicinoids found in chili peppers (Bosland and Votava 2000) are synthesized in the epidermis of the placenta substances (Stewart et al. 2007). Although capsaicinoid contents and the intensity of pungency are quantitative traits, in the C. annuum complex the presence of capsaicinoids was found to be regulated by the Pun1 gene; however, its expression is distinct among species and cultivars, since it is based on other modifier genes epistatically affected by Pun1 and environmental conditions (Lippert et al. 1966; IBPGR 1983; Bosland and Votava 2000; Hunziker 2001; Lefebvre et al. 2002; Stewart et al. 2005). Sometimes, pungency is missing, as in all species from the x = 13 ʽyellow-flowered groupʼ or Andean clade (Carrizo García et al. 2016; Scaldaferro et al. 2016); e.g., Capsicum dimorphum, C. geminifolium, C. hookerianum, C. lanceolatum, C. lycianthoides, and C. rhomboideum are reported to be completely free of pungency (Barboza pers. com.). This peculiarity of the genus is also absent in Capsicum longidentatum Agra and Barboza (Barboza pers.com.), some accessions of C. chacoense (Eshbaugh 1980; Tewksbury et al. 2006), in one of Capsicum cornutum (as Capsicum dusenii Bitter; Hunziker 1971), and in cultivars of C. annuum var. annuum after human selection.
Meanwhile, chromosome markers have been very important tools to elucidate the evolution and diversification of the genus (Moscone et al. 1993, 1995, 1996a, b, 1999, 2003, 2007; Park et al. 2000; Scaldaferro et al. 2006, 2013, 2016; Romero-da Cruz et al. 2016). The most used markers for chromosome identification in the Capsicum species studied up to now are chromosome banding methods with fluorochromes to reveal heterochromatic regions, the use of silver impregnation to show the exact position of active NORs, and the application of FISH with rDNA probes. These techniques have provided a more defined karyo-systematic analysis, contributing to the comprehension of the diversification and evolution of the genus (Moscone et al. 1993, 1995, 1996a, b, 1999, 2003, 2007; Park et al. 2000; Scaldaferro et al. 2006, 2011, 2013, 2016; Barboza et al. 2011; Romero-da Cruz et al. 2016).
This chapter describes all Capsicum chromosome features studied to date. A list of the recognized Capsicum species and some taxonomically relevant cytogenetic traits are presented in Table 4.1.
4.2 Phylogeny of Chili Peppers
Carrizo García et al. (2016) proposed an informal classification of Capsicum into 11 clades according to three molecular markers, two plastid DNA markers, the maturase K gene (matK) and the psbA-trnH intergenic spacer, and one nuclear gene waxy (GBSSI, granule-bound starch synthase). Based on this grouping, there are only two clades where species possess the chromosome number of 2n = 26, i.e., Andean and Atlantic Forest clades.
C. annuum var. annuum and var. glabriusculum, C. chinense, C. frutescens, and Capsicum galapagoense previously belonged to the ʽwhite-flowered groupʼ and were grouped together again in Annuum clade based on the phylogeny study. Nevertheless, C. chacoense which was also a member of that group but with controversial positions was now nested with C. baccatum and Capsicum praetermissum in Baccatum clade. In Moscone et al. (2007) and Scaldaferro et al. (2013), C. praetermissum was specifically ranked in an intermediate position between Baccatum and Purple corolla clade (Moscone et al. 2007; Scaldaferro et al. 2013).
In the phylogeny, Capsicum flexuosum was suggested to be close to Bolivian clade and was recognized as the monotypic Flexuosum clade, although Carrizo García et al. (2016) have included C. aff. flexuosum (unknown chromosome number) in the clade, considered as a local variation because results did not support a strong specific separation from typical C. flexuosum.
Capsicum cardenasii, C. eshbaughii, C. eximium, C. pubescens, and C. tovarii were members of the traditional ʽpurple-flowered group.ʼ Then, they were assigned to Purple corolla (C. cardenasii, C. eshbaughii, and C. eximium), Pubescens, and Tovarii clades, respectively.
Another group with Capsicum campylopodium, C. cornutum, C. friburgense, C. hunzikerianum, C. mirabile, C. pereirae, C. recurvatum, C. schottianum, and C. villosum belong to Atlantic Forest clade (x = 13), phylogenetically distant from the above-mentioned groups. This clade has corollas mostly white, with some variations in the throat (golden, violet, brownish, greenish, or purple spots) and in the tube (yellowish or greenish). In C. friburgense, corolla appears completely pink or lilac.
On the other hand, C. rhomboideum, C. lanceolatum, C. geminifolium, C. lycianthoides, C. dimorphum, and C. hookerianum are the most distant taxa, belonging to Andean ʽyellow-flowered group,ʼ sometimes with violet spots in the throat and with x = 13.
Caatingae clade includes Capsicum caatingae and C. parvifolium, both with x = 12 and with similar karyotype formulas. Longidentatum clade, with a single species, C. longidentatum, possesses x = 12 with a karyotype very similar to that of Caatingae clade.
Finally, Bolivian clade, with Capsicum caballeroi, C. ceratocalyx, C. coccineum, and C. minutiflorum, presents lemon yellow flowers, sometimes with violet spots in the throat. This clade has not been studied cytogenetically until now, and therefore, its chromosome number is still unknown; however, its position in the phylogeny suggests that Bolivian species have 2n = 24, as the sister clades Longidentatum, Flexuosum, and Caatingae (Fig. 4.1).
Phylogenetic tree of Capsicum species, based on three molecular markers: matK, psbA-trnH, and waxy (Carrizo García et al. 2016), and the relationships with chromosome numbers
4.3 Basic Chromosome Number
The genus Capsicum has two universal chromosome numbers: 2n = 2x = 24 and 2n = 2x = 26, the latter only found in wild species (Pickersgill 1971, 1991; Moscone 1990, 1993, 1999; Moscone et al. 1996a, 2007; Tong and Bosland 2003; Pozzobon et al. 2006; Scaldaferro et al. 2011, 2013, 2016).
To date, the diploid chromosome number for 77.8% of the recorded Capsicum species (28/36) is known. Among them, 15 species (15/28) have 2n = 2x = 24, whereas 13 species (13/28) possess 2n = 2x = 26 (Table 4.1). The phenomenon of polyploidy has never been significant in Capsicum and was only found in one accession of C. annuum var. glabriusculum with 2n = 4x = 48 (Pickersgill 1977).
The chromosome numbers of the following eight species have not been reported yet: the whole Bolivian clade (C. caballeroi, C. ceratocalyx, C. coccineum, and C. minutiflorum), C. eshbaughii (Purple corolla clade), C. dimorphum, C. hookerianum (Andean clade), and C. hunzikerianum (Atlantic Forest clade).
4.4 Karyotyping of Capsicum Species
Since 1971, Capsicum species have been assessed using different methodological chromosome approaches, including classic and silver staining, fluorescent banding, FISH, and nuclear DNA content estimation (Pickersgill 1971, 1991; Moscone 1990, 1993, 1999; Moscone et al. 1993, 1995, 1996a, b, 1999, 2003, 2007; Park et al. 2000; Scaldaferro et al. 2006, 2011, 2013, 2016; Barboza et al. 2011; Romero-da Cruz et al. 2016).
Although chromosome number has been studied in 28 species, their karyotypes have been obtained only from 24, since the number of chromosomes proceeds from meiosis in some cases (i.e., Capsicum buforum, C. cornutum, C. friburgense, and C. lanceolatum; Tong and Bosland 2003; Pozzobon et al. 2006). Half of cytogenetically studied taxa present intraspecific karyotype variation, differing in karyotype formulas, number and location of active NORs, heterochromatin content (Hc), and banding pattern (Moscone et al. 2007; Scaldaferro et al. 2013, 2016) (Figs. 4.2, 4.3, 4.4, 4.5, 4.6, 4.11, 4.12; Table 4.1).
Somatic metaphases of Annuum clade (2n = 24) stained with CMA. a C. annuum var. annuum cytotype 2 (NMCA 10272). b C. annuum var. glabriusculum cytotype 1 (NMCA 10955). c C. annuum var. glabriusculum cytotype 2 (NMCA 10983). d C. annuum var. glabriusculum cytotype 3 (Ll. Q. w. no.). e C. annuum var. glabriusculum cytotype 4 (YSG w. no.). f C. annuum var. glabriusculum cytotype 5 (Netherlands 804750009). g C. annuum var. glabriusculum cytotype 6 (PI 511885). h C. annuum var. glabriusculum cytotype 7 (PI 511886). i C. chinense cytotype 1 (GEB et al. 797). j C. frutescens (GEB et al. 795). k C. galapagoense (PI 639682). Identified homologous chromosomes are indicated with the same numbers as those used in the respective ideogram (Fig. 4.11). Arrowheads indicate CMA+ NOR-associated heterochromatin. Scale bar = 10 µm
Somatic metaphases of Baccatum clade (2n = 24) stained with CMA. a C. baccatum var. baccatum (GEB 163). b C. baccatum var. pendulum cytotype 2 (EAM & RN 211). c C. chacoense cytotype 1 (EAM 250). d C. praetermissum cytotype 2 (EFM 05-17). Identified homologous chromosomes are indicated with the same numbers as those used in the respective ideogram (Fig. 4.11). Arrowheads indicate CMA+ NOR-associated heterochromatin. Scale bar = 10 µm
Somatic metaphases of Purple corolla, Pubescens, and Tovarii clades (2n = 24) stained with CMA. a C. cardenasii cytotype 1 (GEB w. no.). b C. cardenasii cytotype 2 (Netherlands 904750136). c C. eximium cytotype 1 (EAM 254). d C. eximium cytotype 2 (EAM 255). e C. pubescens (EAM 257). f C. tovarii cytotype 2 (NMCA 90008). Identified homologous chromosomes are indicated with the same numbers as those used in the respective ideogram (Fig. 4.11). Arrowheads indicate CMA+ NOR-associated heterochromatin. Scale bar = 10 µm
Somatic metaphases of Caatingae, Flexuosum (2n = 24), and Andean clades (2n = 26) stained with CMA. a C. caatingae cytotype 1 (ATH 25233). b C. caatingae cytotype 2 (ATH 25233 bis). c C. flexuosum (GEB et al. 1034). d C. rhomboideum (YSG 20). e C. lycianthoides (GDB 85). Identified homologous chromosomes are indicated with the same numbers as those used in the respective ideogram (Fig. 4.11). Arrowheads indicate CMA+ NOR-associated heterochromatin. Scale bar = 10 µm
Somatic metaphases of Atlantic forest clade (2n = 26) stained with CMA. a C. campylopodium cytotype 1 (ATH 25116). b C. campylopodium cytotype 2 (ATH 25128). c C. mirabile cytotype 1 (ATH 25251). d C. mirabile cytotype 3 (ATH 25238). e C. pereirae cytotype 1 (ATH 26137). f C. recurvatum (GEB et al. 1629). g C. villosum (GEB et al. 1653). Identified homologous chromosomes are indicated with the same numbers as those used in the respective ideogram (Fig. 4.11). Arrowheads indicate CMA+ NOR-associated heterochromatin. Scale bar = 10 µm
According to the base-specific fluorochromes used for chromosome banding, there are four types of constitutive heterochromatin in Capsicum, which depend on composition of the satellite DNA: (1) highly GC-rich heterochromatin CMA+/DAPI−, CMA homogeneously bright and DAPI dull, occurring in NORs of every Capsicum species; (2) highly AT-rich heterochromatin CMA−/DAPI+, CMA dull and DAPI bright, only present in C. campylopodium, C. pereirae, C. praetermissum, and C. pubescens; (3) moderately GC-rich heterochromatin CMA+/DAPIo, CMA bright and DAPI indifferent, and occurs in a variable number of distal and intercalary bands; and (4) CMA+/DAPI+ mixed distal bands CMA and DAPI bright, only observed in C. campylopodium and C. praetermissum (Fig. 4.11).
The species of Purple corolla clade share the karyotype formula but differ in heterochromatin amount: C. eximium exhibits a slightly different chromosome banding pattern with lower Hc than C. cardenasii. In Pubescens, Tovarii, Baccatum, and Annuum clades, chromosome number and karyotype formula of every species have been yet reported, but in C. eshbaughii from Purple corolla clade the chromosome number is still unknown.
Chromosome data from Andean clade species only has been reported for C. lycianthoides and C. rhomboideum. Both karyotype formulas are quite similar: They have small chromosomes compared to other clades, little Hc, and only one pair of NORs.
Atlantic Forest clade comprises 11 species; to date, chromosomes of only eight species in this group have been cytogenetically studied. All of them possess x = 13, with more asymmetrical karyotypes than those in Andean clade and with higher frequencies of sm, st, or t chromosomes. Their chromosome complements show longer haploid karyotype length (HKL) than Andean clade, being twice the complement length than in C. rhomboideum in every case (Table 4.1).
The NOR-associated heterochromatin is CMA+/DAPI−; however, it sometimes appears as CMA+/DAPIo and includes the distal satellite and a small portion of the respective arm adjacent to the NOR. Nucleolar organizer regions are situated on short or long arms, although more often on short arms, and appear as constrictions or gaps in fluorochrome-banded chromosomes. Sometimes, the centromeric heterochromatin is visible only as faint CMA+/DAPIo paired dots, as previously reported for the cultivated taxa of Capsicum (Moscone et al. 1996a). The amplitude ranges of Hc (indicated as percentage of HKL) vary broadly (from 1.80 to 38.91) in the genus and correlate positively with the HKL in most of the taxa examined (Fig. 4.13). C. annuum and C. tovarii have the lowest and highest Hc, respectively, but among clades, Annuum remains the one with lowest Hc, whereas the species with the highest Hc prevail in Atlantic Forest (Table 4.1).
Species with 2n = 24 show rather uniform and comparatively most symmetrical karyotypes, since most of them have the 11 m + 1 st karyotype formula, although 11 m + 1 sm is also frequent. In contrast, among 2n = 26 species karyotype formulas are more asymmetrical, having nine different karyotypes among nine taxa.
4.5 Mapping of the 45S and 5S Ribosomal RNA Genes
Cytotaxonomy commonly uses number and distribution of secondary constrictions, AgNOR bands, satellites, and 45S rDNA loci as morphological karyotype characters (Baeza and Schrader 2005; Xu et al. 2007; García et al. 2009, among hundreds of studied species). All of those characters are particularly associated with the highly preserved ribosomal 45S RNA genes, described as markers of transcriptional or active 45S rDNA genes (e.g., secondary constrictions, AgNOR bands, and satellites) and of non-functional rDNA sites or inactive 45S rDNA loci (Kovarik et al. 2008).
Recently, a physical chromosome map of 12 Capsicum taxa was constructed employing AgNOR banding, which reports the number, size, and position of active NORs and FISH; used together, both methods inform about 5S and active and inactive 45S rRNA genes, revealing the functional 45S rRNA genes in most species of the genus (Scaldaferro et al. 2016).
In Capsicum, AgNORs are frequently associated with satellites that not always differentially dye with silver nitrate. Nucleolar organizer regions appear as constrictions in chromosomes stained with fluorescent dyes in every case (Scaldaferro et al. 2013). The NORs and their associated heterochromatin are rich in GC base pairs (Moscone et al. 1996a, 2007; Scaldaferro et al. 2013) as is the prevalence in plants (Sinclair and Brown 1971). Therefore, in the genus all NORs are considered descendants of the same initial NOR due to an identical base pair constitution (Berg and Greilhuber 1993).
FISH method has resulted in an essential tool for physical gene mapping. Ribosomal genes are highly repetitive sequences or tandem arrangements found in a small number of sites (loci) in the species genome. In the genus Capsicum, FISH of 5S and 45S rRNA genes shows disparity in number, size, and location among the species (Park et al. 2000; Scaldaferro et al. 2006, 2016). Physically, 5S locus maps in a single preserved position, principally intercalary in a metacentric median chromosome. This 5S rDNA distribution could be parsimoniously explained if the common ancestor of the genus was bearer of a single intercalary 5S locus on a medium-sized to large chromosome. Until now, established linkage maps in Capsicum have not included the 5S rRNA gene (Livingstone et al. 1999; Lefebvre et al. 2002; Paran et al. 2004).
The number and position of 45S rDNA loci are useful characters for morphological identification of similar chromosome sites and operate as evolutionary markers between species. In Capsicum, the number of 45S rDNA sites is remarkably variable, ranging widely from a unique pair in C. rhomboideum up to 30 pairs in C. villosum (Figs. 4.7, 4.8, 4.12; Table 4.1), and both number and position of 45S loci remain constant within each species, with some exceptions; e.g., in C. annuum, there are from 1 to 6 sites, 14 to 15 sites in C. baccatum, and from 8 to 18 sites in C. cardenasii. Although a relative constancy is observed in the 45S loci as a whole, the smaller landmarks are more variable, as the major sites hold number and position constant within each species and cytotype. These last sites are principally concomitant with that NORs that have been previously identified by AgNOR banding, and therefore are the active sites (Scaldaferro et al. 2006, 2016). In general, diploid plant genera species bear one pair of NOR (Raina and Khoshoo 1971), but in very few cases diploid taxa contain more than one pair of NOR. In situ, hybridization studies have identified several other rDNA loci but on chromosomes that are devoid of NORs. Hence, the signals at those sites are generally considered to be inactive sites that do not synthesize ribosomal RNA. Even in those diploid species with more than two NORs, only two remain active, as generally found using FISH (Raina and Mukai 1999).
Localization of 45S (red) and 5S (green) rDNA loci inferred via FISH in Annuum and Baccatum clades. a C. annuum var. annuum cytotype 1 (EAM 251). b C. annuum var. annuum cytotype 2 (EAM 204). c C. chinense cytotype 1 (GEB et al. 807). d C. frutescens (GEB et al. 795). e C. baccatum var. baccatum (GEB 163). f C. baccatum var. pendulum cytotype 2 (EAM 247). g C. baccatum var. umbilicatum (EAM 253). h C. chacoense cytotype 1 (EAM 250). i C. praetermissum cytotype 2 (EFM 05-17). Identified homologous chromosomes are indicated with the same numbers as those used in the respective ideogram (Fig. 4.12). Arrowheads indicate active NOR. Scale bar = 10 µm
Localization of 45S (red) and 5S (green) rDNA loci inferred via FISH in Purple corolla, Pubescens, Tovarii, Flexuosum, Andean, and Atlantic Forest clades. a C. cardenasii cytotype 1 (GEB w. no.). b C. cardenasii cytotype 2 (Netherlands 904750136). c C. eximium cytotype 2 (EAM 255). d C. pubescens (EAM 257). e C. tovarii cytotype 2 (NMCA 90008). f C. flexuosum (GEB et al. 1034). g C. rhomboideum (YSG 20). h C. recurvatum (GEB et al. 915). i C. villosum (GEB et al. 1653). Identified homologous chromosomes are indicated with the same numbers as those used in the respective ideogram (Fig. 4.12). Arrowheads indicate active NOR. Scale bar = 10 µm
Genomic evolution in Capsicum has involved considerable changes in number and distribution of the 45S gene family, including locus loss and gain, and sequence spreading. Other mechanisms that generate variations in size, number, and position of rDNA sites are structural rearrangements, such as inversions and translocations, homologous and non-homologous unequal crossing over, gene conversion, and transpositional events (Hall and Parker 1995; Sharma and Raina 2005). Evidence suggests that positioning and remodeling of rDNA sites could be related to the rDNA gene shuffling or transposable elements playing an important role in plant genome evolution (Dubcovsky and Dvorák 1995; Raskina et al. 2004; Datson and Murray 2006).
In Capsicum, 45S FISH signals mostly correspond to specific fluorescent banding, although they do not coincide absolutely in number, location, or size (Moscone et al. 2007; Scaldaferro et al. 2013, 2016). Accordingly, there would be a relationship between 45S rDNA probes and GC-rich heterochromatic regions. Park et al. (2012) have studied in detail the evolution of constitutive heterochromatin in Capsicum. They showed an expansion of this genome structure 20.0–7.5 million years ago in pepper through a massive accumulation of single-type Ty3/Gypsy-like elements from the Del subgroup. Interestingly, derivatives of the Del elements played important roles in the expansion of constitutive heterochromatic regions. This process represents a characteristic mechanism for genome amplification in plant species through expansion of constitutive heterochromatic regions, which does not involve a genome-wide duplication event. Most recently, Qin and Yu (2014) explained that LTR expansion promoted the large genome size in Capsicum. Our findings about the localization of 45S probes and their relationship with heterochromatic regions and active NORs also suggest their additional role in Capsicum genome diversity.
4.6 AgNOR Mapping
Silver impregnation is used to reliably detect active NORs (Ag-I; Bloom and Goodpasture 1976; Kodama et al. 1980). Active NORs vary in number from one to four pairs among Capsicum species. The maximum number of NORs (four pairs) only appears in species with 2n = 24; instead, taxa with 2n = 26 present 1–2 pairs maximum. Few species have only one pair: C. annuum var. glabriusculum (cytotypes 1, 3, and 5) and C. rhomboideum; instead, nine species present two pairs: C. annuum var. annuum (cytotype 2) and C. annuum var. glabriusculum (cytotypes 2 and 4), C. chinense, C. frutescens, C. eximium (cytotype 2), C. cardenasii (cytotypes 1 and 2), C. flexuosum, C. praetermissum, C. recurvatum, and C. villosum. The only species exhibiting three pairs is C. tovarii (cytotype 2). Finally, C. annuum var. glabriusculum (cytotypes 6 and 7; Fig. 4.9) and C. baccatum var. baccatum and var. pendulum show four pairs of NORs in their diploid complements (Fig. 4.9).
Silver-stained somatic metaphases of Capsicum. a–b C. annuum var. annuum: a NMCA 10272 cytotype 2 and b NMCA 10544 cytotype 2. c–i C. annuum var. glabriusculum: c NMCA 10955 cytotype 1, d NMCA 10983 cytotype 2, e LQ w. no. cytotype 3, f YSG w. no. cytotype 4, g Netherlands 804750009 cytotype 5, h PI 511885 cytotype 6, i PI 511886 cytotype 7. j C. chinense cytotype 1 GEB 807. k C. frutescens GEB, FC, MM 795. l C. baccatum var. baccatum GEB 163. m C. baccatum var. pendulum EAM and RN 211 cytotype 2. n C.eximium EAM 255 cytotype 2. o–p C. cardenasii: o Netherlands 904750136 cytotype 1, p AAC w. no. cytotype 2. q C. flexuosum GEB, FC, EMa 1034. r C. praetermissum EFM 05-17 cytotype 2. s C. rhomboideum YSG 20. t C. recurvatum GEB, MM, RSc, RM 915. u C. tovarii NMCA 90008 cytotype 2. v C. villosum GEB, EFi, AG, GB 1653.M, macrosatellite; T, tandem satellite. Arrows indicate AgNOR. Scale bar = 10 µm
Mostly, the NORs in Capsicum are positioned on the short arm of the respective chromosomes, although some taxa exhibit one NOR on the long arm of different chromosome pairs, e.g., C. annuum var. glabriusculum (cytotypes 2, 4, 6, and 7), C. eximium (cytotype 2), C. cardenasii (cytotypes 1 and 2), C. tovarii (cytotype 2), and C. villosum. The sizes of NOR-associated satellites are diverse among species, individuals, and frequently among cells from the same plant. According to Battaglia terminology (Battaglia 1955), microsatellites, macrosatellites, and tandem satellites are registered in varying proportions among species (Fig. 4.9; see M NOR-bearing chromosomes and T NOR-bearing chromosomes). Also in some cases, size of NORs varies between homologues.
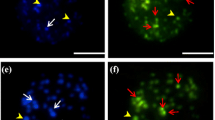
In all cases, the maximum number of nucleoli seen in interphase is coincident with the maximum number of NORs found in metaphase (Fig. 4.10). Although the correspondence between the size of NORs in metaphase and the size of nucleoli in the interphase nuclei is a well-established phenomenon in plants (Burger and Knälmann 1980; Hizume et al. 1982; Linde-Laursen 1984), no size correlation has been observed in Capsicum (Moscone et al. 1995; Scaldaferro et al. 2016).
Silver-stained interphase nucleus of Capsicum. a–b C. annuum var. annuum: a NMCA 10272 cytotype 2 and b NMCA 10544 cytotype 2. c–i C. annuum var. glabriusculum: c NMCA 10955 cytotype 1, d NMCA 10938 cytotype 2, e LQ w. no. cytotype 3, f YSG w. no. cytotype 4, g Netherlands 804750009 cytotype 5, h PI 511885 cytotype 6, i PI 511886 cytotype 7. j C. chinense GEB 807. k C. frutescens GEB, FC, MM 795. l C. baccatum var. baccatum GEB 163. m C. baccatum var. pendulum EAM and RN 211. n C. eximium EAM 255 cytotype 2. o–p C. cardenasii: o Netherlands 904750136 cytotype 1, p AAC w. no. cytotype 2. q C. flexuosum GEB, FC, EMa 1034. r C. praetermissum EFM 05-17 cytotype 2. s C. rhomboideum YSG 20. t C. recurvatum GEB, MM, RSc, RM 915. u C. tovarii NMCA 90008 cytotype 2. v C. villosum GEB, EFi, AG, GB 1653. Scale bar = 10 m
4.7 DNA Content of Capsicum Species
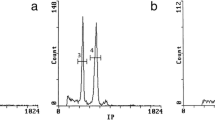
Moscone et al. (2003) estimated nuclear DNA content in nine diploid species of Capsicum by flow cytometry, using ethidium bromide to stain the DNA (internal standard, Hordeum vulgare, 1C = 5.063 pg) (Table 4.1). Additionally, two samples were analyzed using Feulgen densitometry (C. annuum var. annuum and C. pubescens; standard, Allium cepa, 1C = 16.75 pg). Very similar relative values were obtained from both staining methods. The 1C values ranged from 3.34 to 3.43 pg (3273–3361 Mbp) in C. chacoense and the C. annuum complex to 4.53–5.77 pg (4439–5655 Mbp) in C. campylopodium and C. caatingae. Genome size displayed significant variation between but not within species (except in C. campylopodium) and contributed to their taxonomic grouping (Moscone et al. 2003) (Figs. 4.11 and 4.12).
Ideograms of Capsicum taxa showing heterochromatic fluorochrome banding pattern after staining with CMA/DAPI. Solid black blocks indicate CMA+/DAPIo or bands of homogeneous aspect; solid gray blocks indicate DAPI+/CMA− bands of homogeneous aspect; spotted blocks indicate CMA+/DAPI+ bands of mottled appearance. Active 45S rDNA sites are indicated by a constriction. Chromosomes that have the same number on the ideogram are not necessarily homologous for the different taxa. In each ideogram, chromosomes with similar measures without markers were grouped. m, metacentric; sm, submetacentric; st, subtelocentric; t, telocentric. Scale bar = 5 µm
Ideograms of Capsicum taxa showing the distribution of 45S (red blocks) and 5S (green circles) rDNA loci. Euchromatic regions appear in light blue. Active 45S rDNA sites are indicated by a constriction. Chromosomes that have the same number on the ideogram are not necessarily homologous for the different taxa. In each ideogram, chromosomes with similar measures without markers were grouped. m, metacentric; sm, submetacentric; st, subtelocentric; t, telocentric. Some ideograms are extracted from Scaldaferro et al. (2016). Scale bar = 5 µm
Quantity and distribution of heterochromatin in Capsicum suggested that this type of chromatin may have been gained by addition rather than by euchromatin transformation. As a consequence of the proportion change of repeated DNA sequences in the nuclear genome (particularly tandem repeats or satellite DNAs that make up heterochromatic C-bands on the chromosomes), the DNA content in angiosperms varies (Flavell 1986; Raina and Bisht 1988; Bennett and Leitch 1995; Greilhuber 1995). Another previous work showed a strong positive correlation between genome size and Hc in Capsicum, with higher range of variation in the latter parameter (Scaldaferro et al. 2013) (Fig. 4.13).
Relationship between haploid karyotype length (HKL) and heterochromatin amount (expressed as percentage of HKL) in Capsicum. 1 C. rhomboideum; 2 C. rhomboideum; 3 C. galapagoense; 4 C. annuum var. glabriusculum cytotype 2; 5 C. chacoense cytotype 1; 6 C. annuum var. glabriusculum cytotype 1; 7 and 8 C. chinense cytotype 1 and 2, respectively; 9 C. chacoense cytotype 1; 10 C. frutescens; 11 C. baccatum var. baccatum; 12 C. tovarii cytotype 2; 13 C. annuum var. annuum cytotype 1; 14 C. eximium cytotype 1; 15 C. cardenasii cytotype 1; 16 C. eximium cytotype 2; 17 C. annuum var. glabriusculum cytotype 7; 18 C. tovarii cytotype 1; 19 C. annuum var. annuum cytotype 2; 20 C. chacoense cytotype 2; 21 C. praetermissum cytotype 1; 22 and 24 C. baccatum var. pendulum cytotype 1 and 2; 23 C. baccatum var. umbilicatum; 25 C. pereirae cytotype 1; 26 C. recurvatum; 27 C. villosum; 28 C. pereirae cytotype 2; 29 C. praetermissum cytotype 2; 30 C. caatingae cytotype 2; 31 C. pubescens; 32 C. cardenasii cytotype 2; 33 C. caatingae cytotype 1; 34, 38, and 39 C. mirabile cytotype 1, 2 and 3, respectively; 35 and 36 C. campylopodium cytotype 1 and 2, respectively; 37 C. schottianum; 40 C. flexuosum. The line (r = 0.65; P\0.0001) represents the correlation among data.
4.8 Concluding Remarks and Future Prospects
This chapter compiles all the chromosome features that have been studied in the American genus Capsicum until now, also included DNA content data. This is an innovative approach since the genus is treated based on the last phylogeny from Carrizo García et al. (2016), which allowed to relate chromosome similarities within each clade, and chromosome diversity among different clades. Our group continues working on the genus, and new sequencing technologies (Harrison and Kidner 2011; Macas et al. 2011; Buggs et al. 2012; Egan et al. 2012) would facilitate comprehensive studies of Capsicum genome. (Harrison and Kidner 2011; Macas et al. 2011; Buggs et al. 2012; Egan et al. 2012). Therefore, with the rapidly advancing sequencing technology and cytogenetic analysis, we will gain knowledge that could be compared with data from this chapter.
References
Baeza C, Schrader O (2005) Comparative karyotype analysis in Haplopappus Cass. and Grindelia Willd. (Asteraceae) by double FISH with rRNA specific genes. Plant Syst Evol 251:161–172
Barboza GE, Bianchetti LB (2005) Three new species of Capsicum (Solanaceae) and a key to the wild species from Brazil. Syst Bot 30(4):863–871
Barboza GE, Agra MF, Romero MV, Scaldaferro MA, Moscone EA (2011) New endemic species of Capsicum (Solanaceae) from the Brazilian caatinga: comparison with the re-circumscribed C. parvifolium. Syst Bot 36(3):768–781
Battaglia E (1955) Chromosome morphology and terminology. Caryologia 8:179–187
Bennett MD, Leitch IJ (1995) Nuclear DNA amounts in angiosperms. Ann Bot 76:85–91
Berg C, Greilhuber J (1993) Cold-sensitive chromosome regions and heterochromatin in Cestrum (Solanaceae): C. srtigillatum, C. fasciculatum, and C. elegans. Plant Syst Evol 185:133–151
Bloom SE, Goodpasture C (1976) An improved technique for selective silver staining of nucleolar organizer regions in human chromosomes. Hum Genet 34:199–206. https://doi.org/10.1007/BF00278889 PMID:63440
Bosland PW (1996) Capsicums: innovative use of an ancient crop. In: Janick J (ed) Progress in new crops. ASHS Press, Arlington, pp 479–487
Bosland PW, Votava EJ (2000) Peppers: vegetable and spice Capsicums. Crops production science in horticulture 12. CAB Intl Publishing, Wallingford, UK, p 204
Buggs RJA, Chamala S, Wu W, Tate JA, Schnable PS, Soltis DE, Soltis PS, Barbazuk WB (2012) Rapid, repeated, and clustered loss of duplicate genes in allopolyploid plant populations of independent origin. Curr Biol 22:248–252
Burger EC, Knälmann M (1980) Koinzidenz von Feulgen-Achromasie, in situ hybridisierung und silberbandenfärbung in vier nukleolusorganisatoren von Vicia sativa. Eur J Cell Biol 21:313–318
Carrizo García C, Barfuss MHJ, Sehr EM Barboza GE, Samuel R, Moscone EA, Ehrendorfer F (2016) Phylogenetic relationships, diversification and expansion of chili peppers (Capsicum, Solanaceae). Ann Bot 118:35–51
Datson PM, Murray BG (2006) Ribosomal DNA locus evolution in Nemesia: transposition rather than structural rearrangement as the key mechanism? Chromosome Res 14:845–857
Dubcovsky J, Dvorák J (1995) Ribosomal RNA multigene loci: nomads of the triticeae genomes. Genetics 140:1367–1377
Egan AN, Schlueter J, Spooner DM (2012) Applications of next-generation sequencing in plant biology. Am J Bot 99:175–185
Eshbaugh WH (1980) The taxonomy of the genus Capsicum (Solanaceae). Phytologia 47(3):153–165
Eshbaugh WH (1993) History and exploitation of a serendipitous new crop discovery. In: Janick J, Simon JE (eds) New crops. Wiley, New York, pp 132–139
Flavell RB (1986) Repetitive DNA and chromosome evolution in plants. Philos T R Soc Lon B 312:227–242
García S, Garnatje T, McArthur ED, Pellicer J, Siljak-Yakovlev S, Vallès J (2009) Ribosomal DNA, heterochromatin, and correlation with genome size in diploid and polyploid North American endemic sagebrushes (Artemisia, Asteraceae). Genome 52:1012–1024
Greilhuber J (1995) Chromosomes of the monocotyledons (general aspects). In: Rudall PJ, Cribb PJ, Cutler DF, Humphries CJ (eds) Monocotyledons: systematics and evolution. Royal Botanic Gardens, Kew, pp 379–414
Hall KJ, Parker JS (1995) Stable chromosome fission associated with rDNA mobility. Chromosome Res 3:417–422
Harrison N, Kidner CA (2011) Next-generation sequencing and systematics: what can a billion base pairs of DNA sequence data do for you? Taxon 60:1552–1566
Hizume M, Tanaka A, Shigematsu H (1982) Detection of nucleolar organizing regions in the chromosomes of Nigella damacena [i.e. damascena]. Experientia 38:238–239
Hunziker AT (1971) Estudios sobre Solanaceae. VII. Contribución al conocimiento de Capsicum y géneros afines (Witheringia; Acnistus, Aihenaea, etc.). Tercera parte. Kurtziana 6:241–259
Hunziker AT (2001) Genera solanacearum: the genera of Solanaceae illustrated, arranged according to a new system. ARG Gantner Verlag K-G, Liechtenstein
International Board for Plant Genetic Resources (IBPGR) (1983) Genetic resources of Capsicum, a global plan of action. IBPGR Executive Secretariat, Rome, Italy
Kodama Y, Yoshida MC, Sasaki M (1980) An improved silver staining technique for nucleolus organizer regions by using nylon cloth. Jap J Hum Genet 25:229–233. https://doi.org/10.1007/BF01997700
Kovarik A, Dadejova M, Lim YK, Chase MW, Clarkson JJ, Knapp S, Leitch AR (2008) Evolution of rDNA in Nicotiana allopolyploids: a potential link between rDNA homogenization and epigenetics. Ann Bot 101:815–823
Lefebvre V, Pflieger S, Thabuis A, Caranta C, Blattes A, Chauvet J-C, Daubeze AM, Palloix A (2002) Towards the saturation of the pepper linkage map by alignment of three intraspecific maps including known-function genes. Genome 45:839–854
Linde-Laursen IB (1984) Nucleolus organizer polymorphism in barley, Hordeurn vulgare L. Hereditas 100:33–43
Lippert LF, Smith PG, Bergh BO (1966) Cytogenetics of the vegetable crops. Garden pepper, Capsicum sp. Bot Rev (Lancaster) 32:24–55
Livingstone KD, Lackney VK, Blauth JR, van Wijk R, Jahn MK (1999) Genome mapping in Capsicum and the evolution of genome structure in the Solanaceae. Genetics 152:1183–1202
Macas J, Kejnovský E, Neumann P, Novak P, Koblížková A, Vyskot B (2011) Next generation sequencing-based analysis of repetitive DNA in the model dioceous plant Silene latifolia. PLoS ONE 6:e27335
Moscone EA (1990) Chromosome studies on Capsicum (Solanaceae) I. Karyotype analysis in C. chacoënse. Brittonia 42:147–154
Moscone EA (1993) Estudios cromosómicos en Capsicum (Solanaceae) II. Análisis cariotípico en C. parvifolium y C. annuum var. annuum. Kurtziana 22:9–18
Moscone EA (1999) Análisis cariotípico en Capsicum baccatum var. umbilicatum (Solanaceae) mediante bandeos AgNOR y de fluorescencia. Kurtziana 27:225–232
Moscone EA, Lambrou M, Hunziker AT, Ehrendorfer F (1993) Giemsa C-banded karyotypes in Capsicum (Solanaceae). Plant Syst Evol 186:213–229
Moscone EA, Loidl J, Ehrendorfer F, Hunziker AT (1995) Analysis of active nucleolus organizing regions in Capsicum (Solanaceae) by silver staining. Am J Bot 82:276–287
Moscone EA, Lambrou M, Ehrendorfer F (1996a) Fluorescent chromosome banding in the cultivated species of Capsicum (Solanaceae). Plant Syst Evol 202:37–63
Moscone EA, Matzke MA, Matzke AJM (1996b) The use of combined FISH/GISH in conjunction with DAPI counterstaining to identify chromosomes containing transgene inserts in amphidiploid tobacco. Chromosoma 105:231–236
Moscone EA, Klein F, Lambrou M, Fuchs J, Schweizer D (1999) Quantitative karyotyping and dual-color FISH mapping of 5S and 18S-25S rDNA probes in the cultivated Phaseolus species (Leguminosae). Genome 42:1224–1233
Moscone EA, Baranyi M, Ebert I, Greilhuber J, Ehrendorfer F, Hunziker AT (2003) Analysis of nuclear DNA content in Capsicum (Solanaceae) by flow cytometry and Feulgen densitometry. Ann Bot 92:21–29
Moscone EA, Scaldaferro MA, Grabiele M, Cecchini NM, Sanchez García Y, Jarret R, Daviña JR, Ducasse DA, Barboza GE, Ehrendorfer F (2007) The evolution of chili peppers (Capsicum – Solanaceae): a cytogenetic perspective. Proc PAA/Solanaceae Conf Acta Hortic 745:137–169 (International Society for Horticultural Science, Leuvem, Belgium, ISSN 0567-7572)
Olmstead RG, Bohs L, Migid HA, Santiago-Valentin E, Garcia VF, Collier SM (2008) A molecular phylogeny of Solanaceae. Taxon 57:1159–1181
Paran I, van der Voort JR, Lefebvre V, Jahn M, Landry L, van Schriek M, Tanyolac B, Caranta C, Ben Chaim A, Livingstone K, Palloix A, Peleman J (2004). An integrated genetic linkage map of pepper (Capsicum spp.). Mol Breed 13:251–261
Park Y-K, Park K-C, Park C-H, Kim N-S (2000) Chromosomal localization and sequence variation of 5S rRNA gene in five Capsicum species. Mol Cells 10:18–24
Park M, Park J, Kim S, Kwon J‐K, Park HM, Bae IH, Yang T‐J, Lee Y‐H, Kang B‐C, Choi D (2012) Evolution of the large genome in Capsicum annuum occurred through accumulation of single-type long terminal repeat retrotransposons and their derivatives. Plant J https://doi.org/10.1111/j.1365-313x.2011.04851.x
Perry DR, Zarrillo S, Holst I, Pearsall DM, Piperno DR, Berman MJ, Cooke RG, Rademaker K, Ranere AJ, Raymond JS, Sandweiss DH, Scaramelli F, Tarble K, Zeidler JA (2007) Starch fossils and the domestication and dispersal of chili peppers (Capsicum spp. L.) in the Americas. Science 315:986–988
Pickersgill B (1971) Relationships between weedy and cultivated forms in some species of chili peppers (genus Capsicum). Evolution 25:683–691
Pickersgill B (1977) Taxonomy and the origin and evolution of cultivated plants in the New World. Nature 268:591–595
Pickersgill B (1984) Migration of chili peppers, Capsicum spp, in the Americas. In: Store P (ed) Pré-Columbian plant migration. Papers of the Peabody Museum of archaelogy and ethnology 76. Harvard University Press, Cambridge, pp 105–123
Pickersgill B (1991) Cytogenetics and evolution of Capsicum L. In: Tsuchiya T, Gupta PK (eds) Chromosome engineering in plants: genetics, breeding, evolution. Part B. Elsevier, Amsterdam, pp 139–160
Pickersgill B (1997) Genetic resources and breeding of Capsicum spp. Euphytica 96:129–133
Piperno DR (2011) The origins of plant cultivation and domestication in the new world tropics. Patterns, process, and new developments. Curr Anthropol 52:S453–470 https://doi.org/10.1086/659998 (The Wenner-Gren Foundation for Anthropological Research)
Pozzobon MT, Schifino-Wittmann MT, Bianchetti LB (2006) Chromosome numbers in wild and semidomesticated Brazilian Capsicum L. (Solanaceae) species: do x = 12 and x = 13 represent two evolutionary lines? Bot J Linn Soc 151:259–269
Qin C, Yu (2014) Whole-genome sequencing of cultivated and wild peppers provides insights into Capsicum domestication and specialization. Proc Nat Acad Sci USA 111(14):5135–5140
Raina SN, Bisht MS (1988) DNA amounts and chromatin compactness in Vicia. Genetica 77:65–77
Raina SN, Khoshoo TN (1971) Cytogenetics of tropical bulbous ornamentals III: mitotic mosaicism in 3x Crinum augustum. Theor Appl Genet 41:375–378
Raina SN, Mukai Y (1999) Detection of a variable number of 18S–5.8S–26S and 5S ribosomal DNA loci by fluorescence in situ hybridisation in diploid and tetraploid Arachis species. Genome 42:52–59
Raskina O, Belyayev A, Nevo E (2004) Activity of the En/Spm-like transposons in meiosis as a base for chromosome repatterning in a small, isolated, peripheral population of Aegilops speltoides Tausch. Chromosome Res 12:153–161
Romero-da Cruz MV, Urdampilleta JD, Forni Martins ER, Moscone EA (2016) Cytogenetic markers for the characterization of Capsicum annuum L. cultivars. Plant Biosyst https://doi.org/10.1080/11263504.2015.1103798
Särkinen T, Bohs L, Olmstead RG, Knapp S (2013) A phylogenetic framework for evolutionary study of the nightshades (Solanaceae): a dated 1000-tip tree. BMC Evol Biol 13:214
Scaldaferro MA, Seijo GJ, Acosta MC, Barboza GE, Ducasse DA, Moscone EA (2006) Genomic characterization of the germplasm in peppers (Capsicum—Solanaceae) by fluorescent in situ hybridization. Plant Sci 43(4):291–297 (National Centre for Agrarian Sciences, Sofia, Bulgaria; ISSN 0568-465X)
Scaldaferro MA, Barboza GE, Moscone EA (2011) Estudios citogenéticos en especies andinas silvestres de Capsicum (Solanaceae). In: Sociedad Argentina de Genética (ed) XL Congreso Argentino de Genética, III Simposio Latinoamericano de Citogenética y Evolución, I Jornadas Regionales SAG-NEA. Corrientes, Argentina, Sept 2011. J Basic Appl Genet XLI:31
Scaldaferro MA, Grabiele M, Moscone EA (2013) Heterochromatin type, amount and distribution in wild species of chili peppers (Capsicum-Solanaceae). Genet Res Crop Evol 60(2):693–709. https://doi.org/10.1007/s10722-012-9867-x
Scaldaferro MA, Romero da Cruz MV, Cecchini NM, Moscone EA (2016) FISH and AgNor-mapping of the 45S and 5S rRNA genes in wild and cultivated Capsicum species (Solananceae). Genome 59:95–113. https://doi.org/10.1139/gen-2015-0099
Scaldaferro MA, Barboza GE, Acosta MC (2018) Evolutionary history of the chili pepper Capsicum baccatum L. (Solanaceae): domestication in South America and natural diversification in the Seasonally Dry Tropical Forests. Biol J Linn Soc 124(3):466–478 https://doi.org/10.1093/biolinnean/bly062
Sharma S, Raina SN (2005) Organization and evolution of highly repeated satellite DNA sequences in plant chromosomes. Cytogenet Genome Res 109:15–26
Sinclair JH, Brown DD (1971) Retention of common nucleotide sequences in the ribosomal deoxyribonucleic acid of eukaryotes and some of their physical characteristics. Biochemistry 10:2761–2769
Stewart C Jr, Mazourek M, Stellari GM, O’Connell M, Jahn M (2007) Genetic control of pungency in C. chinense via the Pun1 locus. J Exp Bot 58(5):979–991
Stewart C Jr, Kang B-C, Liu K, Mazourek M, Moore SL, Yoo EY, Kim B-D, Paran I, Jahn MM (2005) The Pun1 gene for pungency in pepper encodes a putative acyltransferase. Plant J 42:675–688
Tewksbury JJ, Manchego C, Haak DC, Levey DJ (2006) Where did the chili get its spice? Biogeography of capsaicinoid production in ancestral wild chili species. J Chem Ecol 32:547–564
Tong N, Bosland PW (2003) Observations on interspecific compatibility and meiotic chromosome behavior of Capsicum buforum and C. lanceolatum. Genet Res Crop Evol 50:193–199
Xu YH, Yang F, Cheng YL, Ma L, Wang JB, Li LJ (2007) Comparative analysis of rDNA distribution in metaphase chromosomes of Cucurbitaceae species. Hereditas 29:614–620
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Scaldaferro, M.A., Moscone, E.A. (2019). Cytology and DNA Content Variation of Capsicum Genomes. In: Ramchiary, N., Kole, C. (eds) The Capsicum Genome. Compendium of Plant Genomes. Springer, Cham. https://doi.org/10.1007/978-3-319-97217-6_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-97217-6_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-97216-9
Online ISBN: 978-3-319-97217-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)