Abstract
Evolutionarily conserved and pleiotropic, the translationally controlled tumor protein (TCTP) is a housekeeping protein present in eukaryotic organisms. It plays an important role in regulating many fundamental processes, such as cell proliferation, cell death, immune responses, and apoptosis. As a result of the pioneer work by Adam Telerman and Robert Amson, the critical role of TCTP in tumor reversion was revealed. Moreover, TCTP has emerged as a regulator of cell fate determination and a promising therapeutic target for cancers. The multifaceted action of TCTP depends on its ability to interact with different proteins. Through this interaction network, TCTP regulates diverse physiological and pathological processes in a context-dependent manner. Complete mapping of the entire sets of TCTP protein interactions (interactome) is essential to understand its various cellular functions and to lay the foundation for the rational design of TCTP-based therapeutic approaches. So far, the global profiling of the interacting partners of TCTP has rarely been performed, but many interactions have been identified in small-scale studies in a specific biological system. This chapter, based on information from protein interaction databases and the literature, illustrates current knowledge of the TCTP interactome.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
5.1 Introduction
Translationally controlled tumor protein (TCTP), also termed as tumor protein translationally controlled 1 (TPT1), histamine releasing factor (HRF), p23, or fortilin, was initially discovered in the tumor cells in mice by researchers working on translationally regulated genes (Yenofsky et al. 1983; Macdonald et al. 1995). It was subsequently found that TCTP is an evolutionarily conserved protein that shares a high degree of homology with the protein from plants to mammals (Acunzo et al. 2014; Amson et al. 2013; Bommer and Thiele 2004). Numerous studies have shown that TCTP plays indispensable roles in various physiological processes, including cell proliferation (Gu et al. 2014; Hsu et al. 2007), cell death (Chen et al. 2014; Lucibello et al. 2011; Susini et al. 2008), cell cycle (Burgess et al. 2008; Johnson et al. 2008), the cytoskeleton (Jeon et al. 2016; Jaglarz et al. 2012; Bazile et al. 2009), protein synthesis (Chen et al. 2013a; Rho et al. 2011), immune responses (Tsai et al. 2014; Kaarbo et al. 2013), malignant transformation (Huang et al. 2015), and nuclear reprogramming (Amson et al. 2013; Roque et al. 2016; Wu et al. 2012; Cheng et al. 2012; Sirois et al. 2011). Telerman and colleagues demonstrated that TCTP plays an important role in tumor reversion, which is defined as the process by which cancer cells lose their malignant phenotype (Tuynder et al. 2002, 2004). Our previous study also revealed that downregulation of TCTP in multiple myeloma cells can lead to tumor reversion (Ge et al. 2011). Importantly, increasing evidences suggest that TCTP is a promising therapeutic target for cancer prevention and intervention (Acunzo et al. 2014; Lucibello et al. 2015; Baylot et al. 2012).
The role of TCTP in many cellular functions is the result of its dynamic interactions with numerous cellular proteins. It is well known that tumorigenesis is the consequence of multiple genetic and epigenetic events that induce cell proliferation and the progression of tumor growth. TCTP was implicated in diverse cellular functions due to interactions with other proteins related to tumorigenesis. Therefore, identification and the characterization of the TCTP interacting proteins on a large scale is important for the understanding of its regulatory mechanisms and revealing its functions in tumorigenesis.
5.2 Global Interactome Profiling Methods
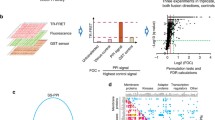
Individual proteins perform their functions through interactions with other proteins and these interactions are crucial for all cellular processes. The knowledge about the entire set of protein interactions (interactome) is essential for our understanding of both the function of individual proteins and the functional organization of the whole cell (Lage 2014). Many experimental high-throughput (HTP) approaches have been developed to determine the protein interactomes in various organisms on a large scale. Through their integration with other “omics” data, interactome datasets have provided valuable information to uncover the functional cellular protein networks and the origin of many diseases. In this chapter, we discuss the emerging and established techniques currently employed to identify the interactome with a particular focus on yeast two-hybrid screens (Y2H) and mass spectrometry (MS)-based approaches. Excellent in-depth reviews on HTP approaches are already available (Mehta and Trinkle-Mulcahy 2016; Lievens et al. 2010).
The yeast two-hybrid (Y2H) assay has been used for over 25 years and remains the most popular choice for researchers investigating interactomes (Rajagopala 2015). This assay is a genetic complementation technique where the proteins to be tested for interaction (referred to as “bait” and “prey”) are fused to the DNA-binding domain and the activation domain of the transcription factor (Bruckner et al. 2009). The proteins are co-expressed in a yeast strain reconstituting transcription factor activity, which drives the expression of a reporter gene (Lentze and Auerbach 2008). Large-scale Y2H strategies have been applied to map the human interactome and to generate protein interactome in a number of model organisms (Rajagopala 2015; Zhang et al. 2010; Yu et al. 2008; Li et al. 2004). There are many different variations of the Y2H assay developed such as the recruitment of the bait and prey to the cytosol, plasma membrane, and endoplasmic reticulum or using multiple baits (Koegl and Uetz 2007; Stellberger et al. 2010).
MS-based proteomics has become a widely used technology to identify protein–protein interactions (PPIs) during the past decade (Smits and Vermeulen 2016). The workflows of the commonly used MS-based interaction proteomics are based on affinity-purification MS (AP-MS) of the protein of interest using specific antibodies. The application of quantitative proteomics such as quantitative immunoprecipitation combined with knockdown (QUICK) for protein enrichments from crude lysates to discriminate bona fide interactors from background proteins has proved to be particularly useful (Ge et al. 2010; Selbach and Mann 2006; Zheng et al. 2012; Chen et al. 2013b). Recently, many different MS-based global interactome profiling approaches have been developed, such as proximity-ligation technology based on engineered ascorbate peroxidase (APEX) labeling and global interactome profiling based on the co-behavior of proteins in biochemical fractionations (Havugimana et al. 2012; Kristensen et al. 2012) or perturbation experiments (Christoforou et al. 2016).
An alternative method that has proved invaluable in protein interactome research is protein arrays, which are miniaturized parallel assay systems that contain small amounts of purified proteins in a high-density format (Phizicky et al. 2003). They allow the simultaneous determination of a variety of analytes from small amounts of samples in a single experiment (Tao et al. 2007; Tao and Zhu 2006). This technique has undergone considerable developments since it was first introduced (Chen et al. 2013b; Yang et al. 2016). This has become one of the most powerful multiplexed detection platforms, which can be used for identification of protein interactome, antibody classification, and protein functional analysis.
5.3 The Current Knowledge of the TCTP Interactome
The results of TCTP interactome analysis imply that TCTP interacting proteins belong to a range of functional groups (Li et al. 2016), including nucleic acid-binding proteins, cytoskeletal proteins, chaperones, enzyme modulators, and transferases, which is consistent with the multifunctional nature of TCTP.
5.3.1 Chaperone Proteins
Chaperone proteins from the highly conserved HSP70 family were identified to be interacting partners of TCTP, and HSPA9 was subsequently confirmed as a TCTP-binding partner (Li et al. 2016). HSP70 family members have key regulatory roles in a variety of cellular stress responses (Murphy 2013; Daugaard et al. 2007). The study of over-expression of HSPA9 in tumor cells demonstrated its function in protecting cells from oxidative damage (Liu et al. 2005; Orsini et al. 2004). Interestingly, TCTP is upregulated during oxidative stress and has been implicated as an antioxidant protein (Lucibello et al. 2011; Oikawa et al. 2002; Rupec et al. 1998). Thus, it is likely that the HSPA9–TCTP complex can function together in resistance to intracellular oxidative stress. The complex of TCTP and HSP70 family may also be involved in anti-apoptotic processes (Fig. 5.1a), which is similar to the HSP27–TCTP complex (Baylot et al. 2012).
Proposed model depicting the molecular mechanism of the validated TCTP-binding proteins. (a) When exposed to cellular stress, HSPA9–TCTP may form complexes to resist apoptosis, which is similar to HSP27–TCTP complex. (b) When exposed to irradiation or ROS, which may result in DSB, the TCTP–XRCC6–XRCC5 complexes will form and exert their function in DSB sensing and repairing. (c) The TCTP–PRDX1 complex may prevent Akt-driven transformation by similar mechanism as TCTP–PTEN in a mild oxidative stress. However, under high doses of oxidative stress, TCTP may also dissociate from oxidized PRDX1 and inactivated. The TCTP–YBX1 complexes are associated with PI3K–Akt signaling pathway which regulates tumorigenesis, malignant transformation, and mTOR signaling. The complexes may also activate mTOR signaling directly and further regulate cell proliferation, cell growth, and cell cycle. (d) The ACTB–TCTP and TUBA1C–TCTP complexes may probably have similar functions as the interaction of TCTP between F-actin, G-actin, and tubulin and exert their function in cell morphology, tumorigenesis, cell proliferation, cell growth, and cell cycle. Red: TCTP protein. Yellow: the novel TCTP-binding proteins; Pink: TCTP-binding proteins that identified previously. Blue: molecules contribute to those functional pathways. The solid line indicates the reported regulation relationships, while the dash line indicates the conjectural regulation relationships
5.3.2 Nucleic Acid-Binding Proteins
A large number of nucleic acid-binding proteins are the regulatory proteins of transcription and translation. Among them, XRCC6 (Ku70) has been demonstrated to play a critical role in genomic stability maintenance by binding to TCTP and XRCC5 (Ku80) (Zhang et al. 2012; Wang et al. 2015; Gullo et al. 2006). When DNA double-strand breaks (DSB) occur, TCTP accumulates at the damage sites, co-localizing with XRCC6 and XRCC5 (Ku80) and forms complexes for DSB repairs (Fig. 5.1b). However, the levels of XRCC5 and XRCC6 in nuclei are reduced in the absence of TCTP (Zhang et al. 2012), suggesting that TCTP can act as a chaperone. Moreover, Gurdon et al. demonstrated that TCTP directly binds to the promoter region of oct4 and acts as a transcription factor for this gene (Koziol et al. 2007). They further showed that TCTP also indirectly activates nanog transcription by binding to a distant site from its promoter (Koziol et al. 2007 ). Together, the interactome analysis further confirmed that TCTP is a critical transcription and translation regulator and may fulfill its functions by binding to other proteins.
5.3.3 Cytoskeletal Proteins
The cytoskeleton is a highly dynamic system comprising of different groups of structural proteins including tubulin, actin, and intermediate filaments to form polymers and associated proteins with diverse regulatory functions (Petrasek and Schwarzerova 2009). TCTP has been reported to be associated with cytoskeleton proteins and many related cellular processes (Bazile et al. 2009; Gachet et al. 1999; Tsarova et al. 2010). For example, TCTP is involved in regulating cell shape, probably via complex interactions with both F-actin and the microtubule cytoskeleton (Bazile et al. 2009). TCTP also associates with microtubules during specific phases of the cell cycle by binding to tubulin (Gachet et al. 1999). TCTP can release the binding of cofilin to G-actin and transfer the active cofilin to F-actin, increasing the cofilin-activity cycle in invasive tumor cells (Tsarova et al. 2010). The proteomics analysis reveals 15 TCTP interacting cytoskeleton proteins and sheds new light on the role of TCTP in cytoskeleton-related functions like cell morphology, tumorigenesis, cell proliferation, cell growth, and cell cycle (Fig. 5.1d).
5.3.4 Other Functions
PRDX1 and YBX1 were also validated to be TCTP-binding partners. PRDX1 was the first antioxidant protein reported to protect other proteins from inactivation through interaction (Neumann et al. 2009). When exposed to mild oxidative stress, PRDX1 is upregulated and binds to phosphatase and tensin homolog (PTEN) to protect it from oxidation-induced inactivation (Neumann et al. 2009). However, under high doses of oxidative stress, PTEN irreversibly dissociates from oxidized PRDX1 and becomes inactivated, resulting in hyperactivation of Akt signaling (Neumann et al. 2009; Stambolic et al. 1998, 2000; Backman et al. 2004). Notably, TCTP upregulation has been detected in surviving cells after oxidative stress (Lucibello et al. 2011). Conversely, when exposed to a strong oxidative stress, cancer cells caused a downregulation of TCTP, followed by cell death (Lucibello et al. 2011). Therefore, the binding of TCTP and PRDX1 may also be involved in antioxidant pathways, and the TCTP–PRDX1 complex may prevent Akt-driven transformation by a similar mechanism as PTEN under mild oxidative stress (Fig. 5.1c).
YBX1 has been implicated in numerous cellular processes similar to TCTP. The pleiotropic functions of YBX1 and TCTP indicate that the TCTP–YBX1 complex may be involved in vital signaling pathways. YBX1 is closely related to the PI3K/Akt/mTOR signaling pathway (Dazert and Hall 2011). It transcriptionally activates the expression of PIK3CA in basal-like breast cancer cells (Astanehe et al. 2009). Serine phosphorylation of the YBX1 102 residue relies on Akt kinase activity (Sutherland et al. 2005; Basaki et al. 2007; Sinnberg et al. 2012). The inhibition of the PI3K pathway can also reduce the expression of YBX1 (Sinnberg et al. 2012). Through experiments of YBX1 silencing, Lee et al. confirmed that the reduction of YBX1 resulted in decreasing of mTOR protein levels (Lee et al. 2008). Interestingly, the translation of TCTP mRNA is regulated by PI3-K/Akt/mTOR signaling, and a positive feedback loop between TCTP and mTOR contributes to tumor formation (Bommer et al. 2015; Kobayashi et al. 2014). By interacting with each other, TCTP and YBX1 may work cooperatively in the PI3K/Akt/mTOR pathway, which regulates tumorigenesis, malignant transformation, and mTOR signaling. The complexes may also be directly related to mTOR signaling and may further influence the glycolysis pathway to regulate cell proliferation, cell growth, and cell cycle (Fig. 5.1c).
5.4 Concluding Remarks
As described above, the TCTP interactome is complex, multifunctional, and has many missing pieces. There are still many unexplained functions that have not yet been attributed to a specific TCTP interactor. Much work is still required to understand the TCTP interactome and its physiological significance. Large-scale approaches, such as next generation Y2H, tandem-affinity purification coupled to MS, and protein arrays might lead to the identification of the new interactors of TCTP and the entire TCTP interactome. The identification of bona fide TCTP interactors can reveal novel functional properties of TCTP. A complete identification of the TCTP interactome will give us a better understanding of the role of TCTP, its mechanism of action, and its associations with the interacting proteins to affect diverse biological and pathological processes. It is critical to identify the key hubs and nodes in the TCTP interaction networks as well as to obtain a detailed molecular characterization of the TCTP interactome. This knowledge will facilitate the development of agents to perturb (or mimic) these interactions. As an example of the potential of this approach, the identification of the interaction between TCTP and YBX1 gives us an idea to find the therapeutic small molecule compound (inhibitor) which can specifically block the interaction (Li et al. 2016). Therefore, elucidating the mechanisms of action of TCTP interactome should provide substantial benefits for the discovery of novel drug targets and biomarkers of disease.
References
Acunzo J et al (2014) TCTP as therapeutic target in cancers. Cancer Treat Rev 40:760–769
Amson R et al (2013) TPT1/TCTP-regulated pathways in phenotypic reprogramming. Trends Cell Biol 23:37–46
Astanehe A et al (2009) The transcriptional induction of PIK3CA in tumor cells is dependent on the oncoprotein Y-box binding protein-1. Oncogene 28:2406–2418
Backman SA et al (2004) Early onset of neoplasia in the prostate and skin of mice with tissue-specific deletion of pten. Proc Natl Acad Sci USA 101:1725–1730
Basaki Y et al (2007) Akt-dependent nuclear localization of Y-box-binding protein 1 in acquisition of malignant characteristics by human ovarian cancer cells. Oncogene 26:2736–2746
Baylot V et al (2012) Targeting TCTP as a new therapeutic strategy in castration-resistant prostate cancer. Mol Ther 20:2244–2256
Bazile F et al (2009) Complex relationship between TCTP, microtubules and actin microfilaments regulates cell shape in normal and cancer cells. Carcinogenesis 30:555–565
Bommer UA, Thiele BJ (2004) The translationally controlled tumour protein (TCTP). Int J Biochem Cell Biol 36:379–385
Bommer UA et al (2015) Growth-factor dependent expression of the translationally controlled tumour protein TCTP is regulated through the PI3-K/akt/mTORC1 signalling pathway. Cell Signal 27:1557–1568
Bruckner A et al (2009) Yeast two-hybrid, a powerful tool for systems biology. Int J Mol Sci 10:2763–2788
Burgess A et al (2008) Chfr interacts and colocalizes with TCTP to the mitotic spindle. Oncogene 27:5554–5566
Chen K et al (2013a) TCTP increases stability of hypoxia-inducible factor 1alpha by interaction with and degradation of the tumour suppressor VHL. Biol Cell 105:208–218
Chen Y et al (2013b) Bcl2-associated athanogene 3 interactome analysis reveals a new role in modulating proteasome activity. Mol Cell Proteomics 12:2804–2819
Chen K et al (2014) Long-term artificial selection reveals a role of TCTP in autophagy in mammalian cells. Mol Biol Evol 31:2194–2211
Cheng X et al (2012) Translationally controlled tumor protein (TCTP) downregulates Oct4 expression in mouse pluripotent cells. BMB Rep 45:20–25
Christoforou A et al (2016) A draft map of the mouse pluripotent stem cell spatial proteome. Nat Commun 7:8992
Daugaard M et al (2007) The heat shock protein 70 family: highly homologous proteins with overlapping and distinct functions. FEBS Lett 581:3702–3710
Dazert E, Hall MN (2011) mTOR signaling in disease. Curr Opin Cell Biol 23:744–755
Gachet Y et al (1999) The growth-related, translationally controlled protein P23 has properties of a tubulin binding protein and associates transiently with microtubules during the cell cycle. J Cell Sci 112(Pt 8):1257–1271
Ge F et al (2010) Identification of novel 14-3-3zeta interacting proteins by quantitative immunoprecipitation combined with knockdown (QUICK). J Proteome Res 9:5848–5858
Ge F et al (2011) Quantitative proteomic analysis of tumor reversion in multiple myeloma cells. J Proteome Res 10:845–855
Gu X et al (2014) TCTP promotes glioma cell proliferation in vitro and in vivo via enhanced beta-catenin/TCF-4 transcription. Neuro-Oncology 16:217–227
Gullo C et al (2006) The biology of ku and its potential oncogenic role in cancer. Biochim Biophys Acta Rev Cancer 1765:223–234
Havugimana PC et al (2012) A census of human soluble protein complexes. Cell 150:1068–1081
Hsu YC et al (2007) Drosophila TCTP is essential for growth and proliferation through regulation of dRheb GTPase. Nature 445:785–788
Huang H et al (2015) Poly(ADP-ribose) glycohydrolase silencing down-regulates TCTP and cofilin-1 associated with metastasis in benzo(a)pyrene carcinogenesis. Am J Cancer Res 5:155–167
Jaglarz MK et al (2012) Association of TCTP with centrosome and microtubules. Biochem Res Int 2012:541906
Jeon HJ et al (2016) TCTP regulates spindle microtubule dynamics by stabilizing polar microtubules during mouse oocyte meiosis. Biochim Biophys Acta 1863:630–637
Johnson TM et al (2008) Plk1 activation by Ste20-like kinase (slk) phosphorylation and polo-box phosphopeptide binding assayed with the substrate translationally controlled tumor protein (TCTP). Biochemistry 47:3688–3696
Kaarbo M et al (2013) TCTP is an androgen-regulated gene implicated in prostate cancer. PLoS One 8:e69398
Kobayashi D et al (2014) Translationally controlled tumor protein is a novel biological target for neurofibromatosis type 1-associated tumors. J Biol Chem 289:26314–26326
Koegl M, Uetz P (2007) Improving yeast two-hybrid screening systems. Brief Funct Genomics Proteomics 6:302–312
Koziol MJ et al (2007) Tpt1 activates transcription of oct4 and nanog in transplanted somatic nuclei. Curr Biol 17:801–807
Kristensen AR et al (2012) A high-throughput approach for measuring temporal changes in the interactome. Nat Methods 9:907–909
Lage K (2014) Protein-protein interactions and genetic diseases: the interactome. Biochim Biophys Acta 1842:1971–1980
Lee C et al (2008) Targeting YB-1 in HER-2 overexpressing breast cancer cells induces apoptosis via the mTOR/STAT3 pathway and suppresses tumor growth in mice. Cancer Res 68:8661–8666
Lentze N, Auerbach D (2008) The yeast two-hybrid system and its role in drug discovery. Expert Opin Ther Targets 12:505–515
Li S et al (2004) A map of the interactome network of the metazoan C. elegans. Science 303:540–543
Li S et al (2016) Characterization of the translationally controlled tumor protein (TCTP) interactome reveals novel binding partners in human cancer cells. J Proteome Res 15:3741–3751
Lievens S et al (2010) Large-scale protein interactome mapping: strategies and opportunities. Expert Rev Proteomics 7:679–690
Liu Y et al (2005) Effect of GRP75/mthsp70/PBP74/mortalin overexpression on intracellular ATP level, mitochondrial membrane potential and ROS accumulation following glucose deprivation in PC12 cells. Mol Cell Biochem 268:45–51
Lucibello M et al (2011) TCTP is a critical survival factor that protects cancer cells from oxidative stress-induced cell-death. Exp Cell Res 317:2479–2489
Lucibello M et al (2015) Phospho-TCTP as a therapeutic target of dihydroartemisinin for aggressive breast cancer cells. Oncotarget 6:5275–5291
Macdonald SM et al (1995) Molecular-identification of an ige-dependent histamine-releasing factor. Science 269:688–690
Mehta V, Trinkle-Mulcahy L (2016) Recent advances in large-scale protein interactome mapping. F1000Res 5
Murphy ME (2013) The HSP70 family and cancer. Carcinogenesis 34:1181–1188
Neumann CA et al (2009) Peroxiredoxin 1 and its role in cell signaling. Cell Cycle 8:4072–4078
Oikawa K et al (2002) Dioxin stimulates synthesis and secretion of IgE-dependent histamine-releasing factor. Biochem Biophys Res Commun 290:984–987
Orsini F et al (2004) The life span determinant p66Shc localizes to mitochondria where it associates with mitochondrial heat shock protein 70 and regulates trans-membrane potential. J Biol Chem 279:25689–25695
Petrasek J, Schwarzerova K (2009) Actin and microtubule cytoskeleton interactions. Curr Opin Plant Biol 12:728–734
Phizicky E et al (2003) Protein analysis on a proteomic scale. Nature 422:208–215
Rajagopala SV (2015) Mapping the protein-protein interactome networks using yeast two-hybrid screens. Adv Exp Med Biol 883:187–214
Rho SB et al (2011) Anti-apoptotic protein TCTP controls the stability of the tumor suppressor p53. FEBS Lett 585:29–35
Roque CG et al (2016) Tumor protein Tctp regulates axon development in the embryonic visual system. Development 143(7):1134–1148
Rupec RA et al (1998) Isolation of a hypoxia-induced cDNA with homology to the mammalian growth-related protein p23. Oncol Res 10:69–74
Selbach M, Mann M (2006) Protein interaction screening by quantitative immunoprecipitation combined with knockdown (QUICK). Nat Methods 3:981–983
Sinnberg T et al (2012) MAPK and PI3K/AKT mediated YB-1 activation promotes melanoma cell proliferation which is counteracted by an autoregulatory loop. Exp Dermatol 21:265–270
Sirois I et al (2011) Caspase-3-dependent export of TCTP: a novel pathway for antiapoptotic intercellular communication. Cell Death Differ 18:549–562
Smits AH, Vermeulen M (2016) Characterizing protein-protein interactions using mass spectrometry: challenges and opportunities. Trends Biotechnol 34:825–834
Stambolic V et al (1998) Negative regulation of PKB/akt-dependent cell survival by the tumor suppressor PTEN. Cell 95:29–39
Stambolic V et al (2000) High incidence of breast and endometrial neoplasia resembling human cowden syndrome in pten(+/−) mice. Cancer Res 60:3605–3611
Stellberger T et al (2010) Improving the yeast two-hybrid system with permutated fusions proteins: the varicella zoster virus interactome. Proteome Sci 8:8
Susini L et al (2008) TCTP protects from apoptotic cell death by antagonizing bax function. Cell Death Differ 15:1211–1220
Sutherland BW et al (2005) Akt phosphorylates the Y-box binding protein 1 at Ser102 located in the cold shock domain and affects the anchorage-independent growth of breast cancer cells. Oncogene 24:4281–4292
Tao SC, Zhu H (2006) Protein chip fabrication by capture of nascent polypeptides. Nat Biotechnol 24:1253–1254
Tao SC et al (2007) Applications of protein microarray technology. Comb Chem High Throughput Screen 10:706–718
Tsai MJ et al (2014) TCTP is essential for beta-cell proliferation and mass expansion during development and beta-cell adaptation in response to insulin resistance. Endocrinology 155:392–404
Tsarova K et al (2010) Identification of a cofilin-like actin-binding site on translationally controlled tumor protein (TCTP). FEBS Lett 584:4756–4760
Tuynder M et al (2002) Biological models and genes of tumor reversion: cellular reprogramming through tpt1/TCTP and SIAH-1. Proc Natl Acad Sci USA 99:14976–14981
Tuynder M et al (2004) Translationally controlled tumor protein is a target of tumor reversion. Proc Natl Acad Sci USA 101:15364–15369
Wang W et al (2015) The proteomic investigation reveals interaction of mdig protein with the machinery of DNA double-strand break repair. Oncotarget 6:28269–28281
Wu D et al (2012) Upregulation of TCTP expression in human skin squamous cell carcinoma increases tumor cell viability through anti-apoptotic action of the protein. Exp Ther Med 3:437–442
Yang L et al (2016) Identification of serum biomarkers for gastric cancer diagnosis using a human proteome microarray. Mol Cell Proteomics 15:614–623
Yenofsky R et al (1983) Regulation of messenger-RNA utilization in mouse erythroleukemia-cells induced to differentiate by exposure to dimethylsulfoxide. Mol Cell Biol 3:1197–1203
Yu H et al (2008) High-quality binary protein interaction map of the yeast interactome network. Science 322:104–110
Zhang Y et al (2010) Plant protein-protein interaction network and interactome. Curr Genomics 11:40–46
Zhang J et al (2012) Role of the translationally controlled tumor protein in DNA damage sensing and repair. Proc Natl Acad Sci USA 109:E926–E933
Zheng P et al (2012) QUICK identification and SPR validation of signal transducers and activators of transcription 3 (Stat3) interacting proteins. J Proteome 75:1055–1066
Acknowledgements
This work was supported by the National Key Research and Development Program (2016YFA0501304), the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant No. XDB14030202).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Li, S., Ge, F. (2017). Current Understanding of the TCTP Interactome. In: Telerman, A., Amson, R. (eds) TCTP/tpt1 - Remodeling Signaling from Stem Cell to Disease. Results and Problems in Cell Differentiation, vol 64. Springer, Cham. https://doi.org/10.1007/978-3-319-67591-6_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-67591-6_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-67590-9
Online ISBN: 978-3-319-67591-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)