Abstract
In recent years, many pre-processing filters have been developed in order to enhance anatomical structures on chest CT images. These filters are typically based on the analysis of the multiscale second-order local information of the image, that helps identify structures with even (tubes) or odd (surfaces) symmetries. Therefore, they often require specific parameter tuning to enhance the different structures. Moreover, while the filters seem to be able to isolate the structure of interest, they do not provide information about the sub-voxel location of the feature. In this work, we present a novel method for vessel, airway, and fissure strength computation on chest CT images using convolutional neural networks. A scale-space particle segmentation is used to isolate training points for vessels, airways, and fissures which are then used to train an 8-layer neural network with 3 convolutional layers which define high order local information of the image. The network returns a probability map of each feature and provides information on the feature offset from the voxel sampling center, allowing for sub-voxel location of the different structures. The proposed method has been evaluated on clinical CT images and compared to other methods for feature enhancement available in the literature. Results show that the proposed method outperforms competing algorithms in terms of enhancement and is also unique in providing subvoxel information.
This study was supported by the National Institutes of Health NHLBI awards R01HL116931 and R01HL116473.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
1 Introduction
Enhancement of anatomical structures on CT images is often used as initial step of medical imaging techniques to isolate the structure of interest from the surrounding background. For chest CT images, several methods have been proposed to enhance the vascular and airway trees [1,2,3,4], as well as fissures [5].
Conventional methods are based on the idea that the anatomical structures have either an even (tubes) or odd (surfaces) symmetry. For this reason, the multiscale second-order local information of the image (Hessian matrix) is analyzed to compute the feature strength. This idea was first introduced by the authors in [1], who proposed a method for vessel enhancement based on the geometrical shape of the structure. Although filters using the Hessian matrix provide good results, CT feature image enhancement remains a challenge. In particular, conventional filters often need parameter tuning, and while they show good sensitivity (i.e. they properly enhance the structure of interest), they tend to suffer of poor specificity. To address this, [6] proposed a Hessian-based method for enhancement of nodules, vessels, and airway walls that simultaneously enhances objects of a specific shape while suppressing others. [2] proposed a method that determines the medialness of tubular structures based on the Hessian matrix and then uses a model of cylindrical vessels to derive the optimal parameters for the detection. [7] presented an enhancement algorithm for plate-like structures through a sheetness measure computed based on Frangi’s idea. In an attempt to reduce the sensitivity to noise of Hessian-based filters and enhance small and thin vessels, [3] proposed a method that computes a directional filter bank that facilitates and improves the Hessian analysis.
A main drawback of conventional Hessian-based methods is the assumption that all voxels of a structure are characterized by a single strong shape modeling that structure. For this reason, information at specific points, such as bifurcations or curved ends, is often lost. Therefore, [8] proposed an approach for vessel detection based on a non-parametric model that does not use image derivatives. The method assumes that the intensities inside at least one conical-shaped region around the voxel of interest do not vary much. The method proposed by [4] also tries to preserve the general vascular structures by measuring the strength in terms of the strain energy density, which is tuned to a specific vessel shape computed by means of the Hessian eigenvalues. Other approaches exploit learning techniques to determine optimal filter for feature detection [9].
In this paper, we present a novel method for vessel, airway, and fissure strength computation on chest CT images that aims at breaking the limitations of Hessian-based methods. The idea is to exploit high order derivatives of the image to improve both sensitivity and specificity of the filter and preserve information at critical points, i.e. bifurcations and curved ends. Since their initial development, convolutional neural network (CNN) [10] algorithms have been proved really powerful to outperform conventional methods on challenging problems. However, this approach has not been considered yet to solve problems as CT feature enhancement.
We propose a CNN network composed of 3 convolutional layers and no max-pooling that can learn high order kernel functions that are optimal for feature enhancement. This way, no parameter tuning is involved, and the shape of the structure of interest does not need to be defined a-priori, allowing for simultaneous enhancement of several features. The network returns a probability map of vessels, airways, and fissures. We further exploit the potential phase information that is encoded by the convolutional kernels to estimate the feature subvoxel offset by means of a fully connected regression layer. The method has been compared to six different algorithms using clinical CT cases from the COPDGene study [11], and the experimental results show that the proposed method significantly improves both the sensitivity and the specificity for each of the considered features and outperforms competing approaches as assessed by AUC analysis.
2 Materials and Methods
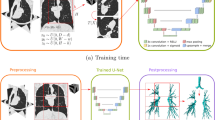
Figure 1 shows the flow diagram of the proposed method to train a CNN to automatically extract feature probabilities from CT images. In this section, we first introduce the construction of the neural network and the specific training set. Then, we introduce the dataset used for evaluation and the experimental setup.
Overview of the proposed method for training CNN to enhance features. Training samples are extracted by particles deployment followed by manual editing. Around each particles, patches of 16 \(\times \) 16 \(\times \) 16 voxels are extracted and given in input to a CNN that produces a probability map for vessels, airways and fissures.
2.1 Feature Likelihood Detection and Localization
To extract the probability of each voxel to belong to vessels, airways, or fissures, we implemented an 8-layer 3D network, which consists of three convolutional layers separated and followed by drop-out layers (to avoid overfitting), and two fully-connected layers. As the idea is to approximate the high local order information of the image, no max pooling, a non-linear down-sampling of the patches, is implemented. The network computes the probability of the central voxel of patches of 16\(\times \)16\(\times \)16 voxels, a size chosen to include enough neighborhood information for big structures, such as the trachea, without losing specificity for small and thin features. We use a Nesterov-momentum update with a softmax function as output non-linearity, and we train on a NVIDIA Titan X GPU machine, using the deep learning framework Keras [12] on top of TensorFlow [13], for 60 epochs with a learning rate of 0.01 and batch size of 128.
We also implemented a regression method that uses the weights trained for classification, and we changed the last fully connected layer to provide the sub-voxel coordinate of the sample of interest.
2.2 Training Set Definition
We used eight cases from the COPDGene study, acquired with an inspiratory protocol, to train the network. Training points were obtained by first performing scale-space particle segmentation [14] for the structures, and manually removing false positive. We also added negative samples to the training set, created from voxels inside the lung that were both close and far from all the structures. Since negative samples and vessels outnumber airways and fissures, we used negative samples and vessels only from four training cases, and we added airway and fissure samples from four additional cases. Moreover, among all available negative points and vessels, we randomly selected a subset of 8,000 negatives (equally divided into far and close points) and 6,000 vessels (including branch points) per case. In summary, the network was trained with a total of 116,147 points, of which 32,000 negatives, 24,000 vessels, 26,412 airways, and 33,735 fissures.
2.3 Experimental Setup
We evaluated our algorithm on 18 cases from the COPDGene study that were not used for the training. In order to create an accurate reference standard, we tweaked parameters to make particles very sensitive, and we manually removed the false positives. Although this approach has limitations, we think that choosing parameters to be extremely sensitive followed by manual touch-up, provides a sensible approach to evaluation.
As with the training set, we considered only a sub-set of negative and vessel samples for evaluation. To evaluate our approach on potentially confounding factors like emphysema, we added to the negative set samples with low intensities, created using a simple thresholding of the image that did not include features of interest. To create the sub-set of samples, we tried to maintain the same distribution of voxels belonging to each structure as in a CT image by randomly selecting 60,000 negative points (equally divided into far, close, and low-intensity points) and 40,000 vessel points per case, while all available fissure and airway points were used. A total of 1,936,154 testing points were used for evaluation: 1, 080, 000 negative points, 720, 000 vessel points, 51, 969 airway points and 84, 185 fissure points. The unbalance ratio between points roughly represents the nominal proportion of each feature in the lung.
Our results were compared with six traditional Hessian-based filters ([1, 2, 4, 7]. Filters as described in [1, 4], originally developed for tubular structures (vessels and airways), were properly modified to also enhance fissures. The method in [2] was implemented only for vessels and airways. In order to compare our method to algorithms specifically developed for surface enhancement, we also implemented the filter described in [7]. Moreover, we combined the methods of [1, 4] and those of [4, 7] by implementing a sheetness equation and adding a step-edge suppressing term to specifically enhance fissures. We applied the different filters to all cases and we evaluated them on the points extracted by particles. The quantitative evaluations were performed by plotting the receiver operator characteristic (ROC) of all considered methods after binarizing the enhanced images with varying global thresholds.
The proposed method does not require any parameter to be set. For all other methods we used 10 steps with 0.5–3.0 mm scales for vessels, as proposed by [4], 0.7–6.0 mm for airways, and 0.5–4.0 mm for fissures. [2] represents the only method that does not require other parameters to be set. For [1, 4, 7] we set all the fixed parameters as proposed in the original papers. The only parameter that needs optimizing is the noise-suppressing parameter C. For this parameter, we set C = 120 for vascular images, as this is the value found as optimal in [4], C = 50 for airways and C = 5 for fissures, found after grid search optimization. All results for the CNN-based algorithm were computed on a single NVIDIA Titan X GPU machine, with batch size equals to 5000.
3 Results
Table 1 shows the performance of the proposed approach compared to the others in terms of area under the ROC curve (AUC). As shown, the proposed algorithm achieves an AUC of 99.76% for vessels, 98.73% for airways, and 99.40% for fissures, against 98.22%, 93.87%, and 97.24% of the competing best methods, respectively. The ROC curves are plotted in Fig. 2 to show the filters performance.
Left: 3D rendering of vessels (top), airways (middle), and fissures (bottom) probabilities obtained from a clinical case (threshold = 0.7). Right: comparison of strengths overlaying a cropped CT image obtained with the proposed method for all features (top) and Xiao, Frangi, and Descoteaux-Xiao for vessels, airways, and fissures, respectively (bottom). Vessels are shown in red, airways in green, and fissures in blue.
A 3D rendering of the detected features, obtained by filtering a CT with our approach and doing a simple thresholding at 0.7 for all structures, is presented in Fig. 3. The features probabilities strengths obtained with the current method and with the best competing algorithm for each feature, are also shown in Fig. 3 overlaying the cropped CT image. As shown, while the competing methods tend to pick several false positives, the proposed approach is very specific without compromising sensitivity, making it a good prior candidate for post-processing operations, such as thresholding, connected component, or feature detection.
An accurate analysis of the probabilities obtained for full clinical cases showed that while the network is extremely sensitive, it sometimes confuses airway walls for vessels and fissures. However, this is not a surprising result, since airway walls were not included as negative samples and have structure and intensities similar to vessels and fissures.
Figure 4a shows iso-surfaces for the convolutional filters that provides the biggest mean activation for fissure, airways and vessel, respectively, for each layer. It can be observed that the filters are highly non-symmetric and high-pass corresponding to anisotropic high order derivatives of the input signal.
Finally, to prove the idea that the analysis of the image with the proposed network contains enough information to define the sub-voxel offset of the considered feature, the box-plot of the L2 norm obtained comparing predicted offsets to those provided by particles, is presented in Fig. 4b. A mean error of about 0.1 voxel units is obtained from the analysis of the three features indicating the adequate performance in resolving the sub-voxel localization of the feature of interest.
(a) Isosurfaces for the convolutional kernels with the highest activations for vessel, airways and fissures. The isosurfaces were up-sampled (for proper showing) and are for the 25% (red), 50% (green) and 75% (blue) level of the highest filter value. (b) Boxplot of the L2 error obtained for offset prediction in comparison to particles.
4 Conclusion
In this paper, a novel enhancement filter is proposed to help identify vessels, airways, and fissures on chest CT images. Conventional methods are typically based on the analysis of second-order local information of the image and need a strong parameter tuning. We propose that analysis of the high-order derivative helps improve the results, avoid the definition of a shape model a-priori, and eliminate parameter tuning. To this end, we used a CNN-based network which uses three convolutional layers and no max-pooling. The network is able to simultaneously enhance vessels, airways, and fissures making the filter one of a kind. Moreover, due to the large patch size chosen, the proposed CNN shows multiscale capabilities as it detects both very large and small features like trachea and small airways and vessels.
Results show that the proposed method outperforms the most typical approaches used in the literature for all features, significantly improving the specificity-sensitivity ratio, as it preserves most true positive while suppressing unwanted noise. This is further confirmed by the fact that even though the proposed filter is not meant for feature segmentation, the low false positives rate suggests that the extracted probabilities could be used as a reference for post-processing operations, such as segmentation or detection.
In addition to feature strength, the CNN estimates the sub-voxel location of the features. This represents a novelty compared to competing methods that only provide strength information. Sub-voxel location of features can be of high interest for specific applications. However, it should be noted that the sub-voxel location was trained from particle sub-voxel offsets which is obtained by means of second-order derivative analysis using high-order splines. This may affect the accuracy of the true sub-voxel location of the feature.
Our approach has a higher computational complexity, due to the use of high-order operators, than conventional second-order derivatives at multiple scales. However, the added complexity is well within the current computational capabilities to perform large scale deployment of this technique, as a typical high resolution case (512 \(\times \) 512 \(\times \) 653) takes around 42 min to be computed.
Finally, although the study we have done shows encouraging preliminary results, an interesting idea for future work is to evaluate the performance of the algorithm when varying scan parameters like slice thickness, reconstruction kernel, dose etc., to show the ability of CNN-based method to generalize results.
References
Frangi, A.F., Niessen, W.J., Vincken, K.L., Viergever, M.A.: Multiscale vessel enhancement filtering. In: Wells, W.M., Colchester, A., Delp, S. (eds.) MICCAI 1998. LNCS, vol. 1496, pp. 130–137. Springer, Heidelberg (1998). doi:10.1007/BFb0056195
Krissian, K., Malandain, G., Ayache, N., et al.: Model-based detection of tubular structures in 3D images. Comput. Vis. Image Underst. 80(2), 130–171 (2000)
Truc, P.T.H., Khan, M.A.U., Lee, Y.K., et al.: Vessel enhancement filter using directional filter bank. Comput. Vis. Image Underst. 113(1), 101–112 (2009)
Xiao, C., Staring, M., Shamonin, D., et al.: A strain energy filter for 3D vessel enhancement with application to pulmonary CT images. Med. Image Anal. 15(1), 112–124 (2011)
van Rikxoort, E.M., van Ginneken, B., Klik, M., et al.: Supervised enhancement filters: application to fissure detection in chest CT scans. IEEE Trans. Med. Imaging 27(1), 1–10 (2008)
Li, Q., Sone, S., Doi, K.: Selective enhancement filters for nodules, vessels, and airway walls in two- and three-dimensional CT scans. Med. Phys. 30(8), 2040–2051 (2003)
Descoteaux, M., Audette, M., Chinzei, K., et al.: Bone enhancement filtering: application to sinus bone segmentation and simulation of pituitary surgery. Comput. Aided Surg. 11(5), 247–255 (2006)
Qian, X., Brennan, M.P., Dione, D.P., et al.: A non-parametric vessel detection method for complex vascular structures. Med. Image Anal. 13(1), 49–61 (2009)
Gonzalez, G., Fleurety, F., Fua, P.: Learning rotational features for filament detection. IEEE Conference on Computer Vision and Pattern Recognition, pp. 1582–1589 (2009)
LeCun, Y., Bengio, Y., Hinton, G.: Deep learning. Nature 521(7553), 436–444 (2015)
Regan, E.A., Hokanson, J.E., Murphy, J.R., et al.: Genetic epidemiology of COPD (COPDGene) study design. COPD: J. Chronic Obstructive Pulm. Dis. 7(1), 32–43 (2011)
Chollet, F.: https://github.com/fchollet/keras. Accessed 20 Feb 2017
Abadi, M., Agarwal, A., Barham, P., et al.: Tensorflow: large-scale machine learning on heterogeneous distributed systems. arXiv preprint arXiv:1603.04467 (2016)
Kindlmann, G.L., Estépar, R.S.J., Smith, S.M., et al.: Sampling and visualizing creases with scale-space particles. IEEE Trans. Vis. Comput. Graph. 15(6), 1415–1424 (2009)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this paper
Cite this paper
Nardelli, P., Ross, J.C., Estépar, R.S.J. (2017). CT Image Enhancement for Feature Detection and Localization. In: Descoteaux, M., Maier-Hein, L., Franz, A., Jannin, P., Collins, D., Duchesne, S. (eds) Medical Image Computing and Computer-Assisted Intervention − MICCAI 2017. MICCAI 2017. Lecture Notes in Computer Science(), vol 10434. Springer, Cham. https://doi.org/10.1007/978-3-319-66185-8_26
Download citation
DOI: https://doi.org/10.1007/978-3-319-66185-8_26
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-66184-1
Online ISBN: 978-3-319-66185-8
eBook Packages: Computer ScienceComputer Science (R0)