Abstract
Tuberculosis (TB) is the leading cause of death globally as a result of a single infectious disease. A staggering 6 million new cases were reported in the 2014. In order to eradicate the ongoing threat of TB and combat rising rates of TB drug resistance new therapeutics must be developed. In this chapter, we briefly review the history of TB, Mycobacterium tuberculosis (Mtb) cell wall structure, the enzymes involved in synthesizing cell wall, and the trehalose utilization pathways (TUP). We focus on the recent discovery of enzyme Mtb GlgE, a glycosyl hydrolase-like phosphorylase, which has been found to be essential for Mtb viability and the ongoing efforts to design inhibitors against this target.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Mycobacterium tuberculosis
- TB
- Trehalose utilization pathways
- GlgE
- Substrate analogs
- Transition-state inhibitors
- Docking studies
1 Mycobacterium Tuberculosis
1.1 Introduction
Dr. Koch discovered the functioning agent of tuberculosis (TB), Mycobacterium tuberculosis (Mtb) in 1882 [1]. Over 100 years later, TB is the leading cause of death in the world due to an infectious disease. In 2014, 1.5 million people were killed by TB. Mortality, due to TB has decreased by 47% since 1990, and effective diagnosis and treatment of TB saved an estimation of 43 million lives between 2000 and 2014 [2]. Globally, the incident rates of TB have fallen by an average of 1.5% per year since 2000. Most of the deaths are caused by drug-resistant strains [2], which have evolved due to selective pressure, co-infection with HIV, and inconsistent adherence to treatment regimens [3]. In 2014, it was estimated that 480,000 new multidrug-resistant TB (MDR-TB) cases occurred worldwide and approximately 190,000 deaths resulted from MDR-TB. Thus, 3.3% of new TB cases were MDR-TB in 2014 and it is estimated that 9.7% of people with MDR-TB have extensively drug-resistant-TB (XDR-TB). Clearly, new drugs are desperately needed to eradicate TB and combat the rising numbers of drug-resistant infections including MDR-TB and XDR-TB [4,5,6,7].
1.2 Present Drugs for Tuberculosis
TB drug therapy began after Koch’s discovery became widely accepted [1]. The major era of anti-TB drug discovery occurred in the mid 1960s [8]. Streptomycin was the first antibiotic used for treating Mtb, beginning in 1944 [9]. Unfortunately, the pattern of Mtb developing resistance to antibiotics was discovered shortly after [10]. First- and second-line drugs that are commonly utilized to fight Mtb. infections are shown in Table 1 [11, 12].
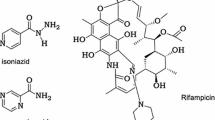
The main reason for identifying new drugs for tuberculosis is the increased prevalence of resistance strains. Further, the current treatment is very complex which results in patient non-compliance. The treatment consists of two phases, the intensive phase and the continuous phase. During the intensive phase patients are administered four first-line drugs for approximately two months. The continuous phase spans four months on isoniazid and rifampin with a three times a week dosing regimen. Situations such as improper medical attention, lack of supplies, and a poor understanding of the strict dosing regimen all contribute to the development of resistant strains.
Drug-resistant strains are categorized as single-drug resistant, MDR and XDR strains. Single-drug resistant should be treatable by a combination of first-line drugs. However, MDR strains are resistant to both isoniazid and rifampin, whereas XDR strains are resistant to isoniazid and rifampin along with any fluoroquinolone and at least one of three injectable second-line drugs (i.e., amikacin, kanamycin, or capreomycin) [13]. Active Mtb infections are also more prominent with the patients affected with HIV, and TB treatments can tremendously interfere with the anti-retroviral drugs used to treat HIV [14]. Presently, researchers are focusing on inhibiting critical cellular process that occurs in either actively dividing or dormant Mtb [15]. Recently GlgE, a maltosyl transferase, in Mtb was identified as a potential new drug target [16].
1.3 Cell Wall of Mycobacterium Tuberculosis (Mtb)
The cell wall of Mtb has two layers. One is the inner layer which consists of membrane proteins in the plasma membrane (Fig. 1), whereas, the outer layer consists of four important main components: the mycolylarabinogalactan (mAG), the peptidoglycan, free glycolipids, and the capsule. The cell wall of Mtb is a sturdy structure which makes it difficult for drugs to pass through [17]. The arabinogalactan and the associated free glycolipid layer together form the mycobacterial outer membrane (MOM). The mAG is made up of D-arabinan, mycolic acids, and D-galactan which are covalently connected [18, 19]. The galactan fragment contains repeating units of galactofuranose in alternating β-(1 → 5) and β-(1 → 6) linkages. Arabinan is attached to the galactan at the fifth position of galactofuranose residue and itself with α-(1 → 3), α-(1 → 5), and β-(1 → 2) linkages. The arabinogalactan is esterified with mycolic acids and further covalently linked to peptigoglycan (PG) by a phosphoryl-N-acetylglucosaminosyl-rhamonse linker [20] which together form the mycolyl arabinogalactan peptidoglycan (mAGP) complex [21,22,23]. The peptidogycan is comprised of alternating units of N-acetylglucosamine and muramic acid residues, which are often acetylated or glycolated [20].
Mycolic acids are made of long chain α-branched-β-hydroxyl fatty acids. The α-branches generally consist of more than 20 carbons, but 60 carbons can also be accomodated by longer meromycolate moities. These are usually functionalized by cis and trans double bonds, cyclopropane rings, methoxy, and keto esters [23,24,25]. These mycolic acids are attached to trehalose monomycolate (TMM), which can also be converted to trehalose dimycolate (TDM). Mycolic acids also form complexes with various sulfolipids, phosphotidylinositol mannoside (PIM), and lipoarabinomannan (LAM) [26]. All these connections and functionalities contribute to the hydrophobicity and impermeability of the mycobacterial cell wall [27].
The LAM complex consists of a PIM anchor, D-mannan, and D-arabinan polysaccheride backbones [28]. These structures are formed by various mannosyl tranferases (ManTs) by the addition of mannose residues to phosphotidylinositol (PI) to form PIM [29, 30]. PIMs can be extended by ManTs to form linear and mature branched lipomannans (LM). LM is subsequently arabinosylated to form mature LAM. The terminal end of LAM can be further glycosylated with mannose to form an mannose cap refered to as manLAM. Together the structures are called the LAM complex. The PI anchor present in the membrane helps to connect the LAM complex with the membrane. This LAM complex also plays a crucial role in maintaining the integrity of the cell wall [31,32,33].
The outer part of the mycobacterial cell wall consists of the capsular layer which is made of polysaccherides and proteins, such as superoxidase dismutase, glutamine synthase, and thioredoxin. The characteristics of the capsular enzymes play crucial roles in bacterium’s response to oxidative stress from the environment.
1.4 Trehalose Utilization Pathways (TUP)
Trehalose (1-O-α-D-glucopyranosyl-α-D-glucopyranoside) is a non-reducing disaccharide which is present in microbial cells, plants, bacterial, insects, and fungi. It protects the organism from stress, typically temperature, dehydration, and oxidative stress [34]. Its mechanism of protecting the cells is not well understood but depends on concentration [35]. Trehalose utilization pathways in Mtb contain several possible drug targets, for example TPP2 [36], Pks13 [37], GlgE [16], MmpL3 [38], and Ag85s [39, 40].
Mtb uses trehalose for protection, energy, and as a precursor for building components of the cell wall [35]. Trehalose can be synthesized in three different pathways (Fig. 2) [41]. The first pathway is the OtsA/TPS pathway, it transforms glucose-6-phosphate to trehalose. OtsA encodes a trehalose phosphate synthase (TPS) which transfers glucose units from UDP-glucose to glucose-6-phosphate yielding trehalose-6-phosphate [42, 43]. In the next step, OtsB2 encodes TPP which dephosphorylates trehalose-6-phosphate forming trehalose. According to mutational studies, OtsA and OtsB genes are essential for the viability [44, 45].
The second pathway is the TreYZ pathway which involves three enzymes TreX, TreY, and TreZ and transforms glycogen to trehalose [46]. Trehalose is also involved in production of TMM via Pks13 [47]. The newly synthesized TMM is passed from the cytoplasm to the periplasm which is taken up by the Ag85 complex and converted into trehalose, mycolylarabinogalactan (mAG), and TDM. TDM is a cell wall envelope glycolipid required for Mtb virulence [40].
The third pathway contains the genetically validated drug target GlgE. This pathway involves the conversion of trehalose into α-1,6-glucans. Briefly, trehalose is converted to maltose by TreS. The resulting maltose is phosphorylated by Mak yielding phosphorylated sugar, maltose-1-phosphate (M1P) [48]. M1P is the substrate for GlgE which forms a linear α-1,4-glucan [16, 49]. The linear α-1,4-glucans are modified with α-1,6 branches by GlgB which is transformed to capsular glucan.
1.5 Importance of GlgE as Drug Target
Kalscheuer et al. showed evidence that GlgE is an essential enzyme present in the α-glucan biosynthesis and showed by four ways that the enzyme meets the characteristic features of a promising new anti-TB target. First, by blocking GlgE activity with a chemical genetic strategy they inhibited the growth of Mtb and established that cell death was caused by a self-poisoning feedback loop. Second, GlgE inhibition lead to Mtb cell death by in vivo studies in lungs and spleens of infected mice. Third, GlgE is not present in humans or in normal gut flora. This implies the drugs made to inhibit GlgE will specifically inhibit Mtb with limited off target effects. Fourth, up to now no one has targeted the α-glucan pathway by chemotherapeutics in the treatment of Mtb. The self-poisoning response is a novel mode of action when compared to the mechanism of action of drugs used presently [16, 49]. Hopefully, a compound can be developed to potently inhibit GlgE which might solve the problem of MDR- and XDR-TB.
1.5.1 Mechanism of GlgE
GlgE is a member of the glycoside hydrolase subfamily GH13_3. The enzyme works by an α-retaining double-displacement mechanism and catalyzes the transfer of maltosyl units as shown in Fig. 3 [50]. The mechanism begins with the side chain of Asp418 attacking M1P generating a β-glycosyl enzyme intermediate. The incoming acceptor glucan is deprotonated by the general acid/base Glu447 side chain and attacks the β-glycosyl enzyme forming linear α-1,4 glucans [51]. Notably, release of phosphate was not observed when β-M1P was used as substrate.
1.6 Synthesis of Ligands for Studying the Mtb GlgE Binding Site
1.6.1 Substrate Analog Maltose-C-Phosphonate
Veleti et al. have synthesized a non-hydrolysable and isosteric analog of M1P, maltose-C-phosphonate (2, MCP) which interacts sufficiently with the substrate binding pocket to produce inhibitory activity. Co-crystallization of the MCP with the homologous enzyme Streptomyces coelicolor (Sco) GlgEI provided information via an X-ray crystal structure that aids in the development of new anti-tuberculosis drugs. The molecule was prepared starting from maltose (1) and employed Wittig and Micheal-Arbuzov chemistry that led to the first inhibitor MCP. MCP moderately inhibited Mtb GlgE with an IC50 = 230 ± 24 µM [52]. MCP was complexed with Sco GlgEI-V279S, an enzyme surrogate for Mtb GlgE, and the crystal structure was solved to 1.9 Å resolution [53]. The results helped to define important interactions between the enzyme and MCP (Fig. 4).
1.6.2 2-Deoxy-2-Fluoro Substrate Analogue
Syson and co-workers [51] reported on the interaction between Sco GlgEI and α-M1P. They performed mutational studies and substituted Asp394 with Ala in order to eliminate hydrolysis of α-M1P over the time scale of protein crystallization. Then they complexed Sco GlgEI-D394A with α-M1P, referred to as the Michaelis complex, and determined the X-ray structure for the complex to 2.55 Å resolution. The structural studies help to define the assignment of the binding subsites. To trap the putative enzyme intermediate they mutated Glu423 with Ala. Sco GlgEI-E423A was complexed with 2-deoxy-2-fluoro-α-D-maltosyl fluoride (1). The X-ray crystal structure was determined to 2.5 Å resolution and showed covalent bond formation between the Asp394 and the C1 carbon. They also used mass spectrometry to show the successive extension of the acceptor by incubating protein with substrate analogue. These crystal structures were the first to show trapping of the glycosyl enzyme intermediate. Their findings provide strong evidence to the support a double-displacement mechanism and assignment of the catalytic nucleophile (Fig. 5).
1.6.3 Transition-State Like Inhibitor
In an attempt to mimic the transition state for the reaction catalyzed by GlgE, Veleti, and co-workers designed an inhibitor with a pyrrolidine moiety shown to inhibit other glycosyl hydrolases in the micromolar range [54, 55]. To synthesize 2,5-dideoxy-3-O-α-D-glucopyranosyl-2,5-imino-D-mannitol (5, DDGIM) a convergent synthesis was followed by coupling thioglycoside (2) with 5-azido-3-O-benzyl-5-deoxy-1,2-O-isopropylindene-β-D-fructopyranose (4) [56] followed by global deprotection to afford the target molecule. Pyrrolidine 5 inhibited both Mtb GlgE and Sco GlgEI-V279S with Ki = 237 ± 27 µM and Ki = 102 ± 7.52 µM, respectively [57]. Further studies complexing DDGIM with Sco GlgEI-V279S were conducted by Lindenberger et al. They solved the crystal structure to a resolution of 2.5 Å [53]. The structure showed numerous interactions in the binding pocket, specifically a strong ionic interaction between D394 and the putative secondary ammonium ion of DDGIM (Fig. 6).
1.6.4 2-Deoxy-2,2-Difluoro Substrate Analogue
Extending the use of fluoro sugars for studying structural insight of GlgE, Thanna et al. synthesized α-2-deoxy-2,2-difluoromaltosyl fluoride (2, α-MTF) and β-2-deoxy-2,2-difluoromaltosyl fluoride (3, β-MTF) [58]. These molecules did not inhibit GlgE; however, 2-deoxy-2,2-difluoro-α-maltosyl fluoride provided useful insight from its X-ray crystal structure complex with Sco GlgEI-V279S at 2.3 Å resolution. The complex provided evidence that Glu423 functions as proton donor by showing a hydrogen bond interaction between Glu423 and C1F. Further, Arg392 and axial C2 difluoromethylene moiety of α-MTF interacts by hydrogen bonding suggesting that C2 substitution can be tolerated with hydrogen bond acceptors (Fig. 7).
1.6.5 Proline and Pyrrolidine-Based Phosphonates as Transistion State Inhibitors
Ongoing work in our lab based on iminosugars as potential transition-state inhibitor against GlgE includes proline-based phosphonates (1 and 3) and pyrrolidine-based phosphonates (2 and 4). These compounds inhibited Sco GlgEI-V279S in a range of 45–95 µM. Addition of phosphonates improved enzyme inhibition 2-fold when compared to the previously synthesized compounds [59]. Crystallizing these target compounds with Sco GlgEI-V279S will be helpful for advancing the structure-based approach for identifying improved inhibitors against GlgE (Fig. 8).
1.7 Inhibitors for Mtb GlgE Based on Docking Studies
Structure-based drug design frequently includes the use of molecular docking. It has become increasingly important and complementary to wet laboratory experiments in that it aids in evaluating the complex information of target enzyme with small ligands. The tools sample optimal geometrical arrangements and quantitate the strength of the bonding forces. By using molecular docking, the binding energy between the ligand and the enzyme binding site can be calculated as shown in Eq. 1 [60]. Recent advances in high-performance computational screening methods have contributed to the design of several drugs that have advanced in clinical trials [61, 62]. For example, computational chemistry has informed lead compound development for compounds that prevent myocardial infraction, treat HIV infection, rheumatoid arthritis, and other diseases [63, 64].
1.7.1 Screening of α-Amylase Family of Protein Binders Against Homology Model of Mtb GlgE
Sengupta et al. studied the binding affinity of an α-amylase family of protein binders to a homology model of Mtb Glg. Since an X-ray structure of Mtb GlgE was unavailable at that time they used the homologue Sco GlgEI [50] to prepare a homologous model of Mtb GlgE. They retrieved the amino acid sequence of Mtb GlgE from the protein database of National Center of Biotechnology Information. They utilized Sco GlgEI (PDB ID: 3ZSS) as a template to superimpose the homology model of Mtb GlgE. Subsequently, the energy of homologous model was minimized using CHARMm. The model structure has three binding cavities with one primary binding site (PBS) and two secondary binding site (SBS1 and SBS2) [60]. Sengupta et al. used CDOCKER, which is a molecular dynamics simulated annealing based docking program, and LibDock, which is a fast feature-based algorithm for molecular docking. 3-O-α-D-gluco-pyranosyl-D-fructose (OTU) docked with the best binding affinity to the PBS among all the computationally screened substrates that were taken from the CHEMBL database as shown in Table 2. This study revealed insights into the active site, substrate binding affinities of ligands, and provide the first homologous 3D structure of Mtb GlgE.
1.7.2 Screening the ZINC Database Against Sco GlgEI
Billones et al. worked on structure-based inhibitors by screening virtual natural products against GlgE. They used the X-ray data from closely related, Sco GlgEI [65] complexed with maltose (PDB code: 3ZT5). The natural products catalogs were obtained from the ZINC database and CDOCKER software was used to dock the molecules. To modify the chosen lead compounds, the de novo evolution protocol was used. They also studied the carcinogenicity, mutagenicity, and biodegradability using a toxicity prediction (extensible) protocol. Among all the screened compounds, they observed three compounds with higher binding energy when compared to natural substrate M1P. The compounds were derived from ZINC39010596, these compounds found to be non-carcinogenic, non-mutagenic, and biodegradable (Table 3).
1.7.3 Screening the ZINC Database and Anti-Tuberculosis Compounds Database Against the Mtb GlgE
Sengupta et al. reported on a pharmacophore-based virtual screening, docking, and molecular dynamics simulations on Mtb GlgE [66]. They presented ligand and structure-based pharmacophore models showing a PBS and SBS2 which were constructed based on the 3D homology model of Mtb GlgE. The structure were screened against the ZINC database and an anti-tuberculosis compounds database (ATD). They identified 23 molecules from ZINC and ATD with better binding affinity than the natural substrate, M1P. The four top molecules that are the best at binding the PBS or SBS2 are shown in Table 4.
2 Summary
Tuberculosis is a communicable disease which is taking millions of lives every year. Current treatment options are complex, lengthy, and clearly difficult to complete. As result, drug-resistance rates are rising. Discovering new drug targets like GlgE, which may rapidly kill the organism, will help in eradicating TB. So far, efforts to identify inhibitors against GlgE have been structure-based. It is expected that gaining an understanding of the detailed binding interactions between the enzyme and its various ligands will inform improved inhibitor design and ultimately lead to new antibiotics.
References
Koch R (1882) Lecture on the discovery of Mycobacterium tuberculosis
World Health Organisation (2015) The Global tuberculosis report 2015. World Health Organization, Geneva
Brennan P, Young D, Robertson B (2008) Handbook of anti-tuberculosis agents. Tuberculosis 88(2):85–170
Chiang CY, Centis R, Migliori GB (2010) Drug-resistant tuberculosis: past, present, future. Respirology 15(3):413–432
Crofton J (1960) Tuberculosis undefeated. Brit Med J 2(5200):679
Mitchison D (2000) Role of individual drugs in the chemotherapy of tuberculosis. Int J Tuberc Lung Dis 4(9):796–806
Udwadia ZF, Amale RA, Ajbani KK, Rodrigues C (2012) Totally drug-resistant tuberculosis in India. Clin Infect Dis 54(4):579–581
Ginsberg A (2011) Research spotlight: The TB alliance: overcoming challenges to chart the future course of TB drug development. Future Med Chem 3(10):1247–1252
Jones D, Metzger H, Schatz A, Waksman SA (1944) Control of gram-negative bacteria in experimental animals by streptomycin. Science 100(2588):103–105
Waksman SA, Reilly HC, Schatz A (1945) Strain specificity and production of antibiotic substances: V. strain resistance of bacteria to antibiotic substances, especially to streptomycin. P Natl Acad Sci USA 31(6):157
Lehmann J (1946) Para-aminosalicylic acid in the treatment of tuberculosis. The Lancet 247(6384):15–16
Janin YL (2007) Antituberculosis drugs: ten years of research. Bioorg Med Chem 15(7):2479–2513
Zhang Y, Yew W (2009) Mechanisms of drug resistance in Mycobacterium tuberculosis [State of the art series. Drug-resistant tuberculosis. Edited by CY. Chiang. Number 1 in the series]. Intl J Tuberc Lung Dis 13(11):1320–1330
Falzon D, Jaramillo E, Schünemann H, Arentz M, Bauer M, Bayona J, Blanc L, Caminero J, Daley C, Duncombe C (2011) WHO guidelines for the programmatic management of drug-resistant tuberculosis: 2011 update. Eur Respir J 38(3):516–528
Lamichhane G (2011) Novel targets in M. tuberculosis: search for new drugs. Trends Mol Med 17(1):25–33
Kalscheuer R, Syson K, Veeraraghavan U, Weinrick B, Biermann KE, Liu Z, Sacchettini JC, Besra G, Bornemann S, Jacobs WR Jr (2010) Self-poisoning of Mycobacterium tuberculosis by targeting GlgE in an [alpha]-glucan pathway. Nat Chem Biol 6(5):376–384
Daffé M, Draper P (1997) The envelope layers of mycobacteria with reference to their pathogenicity. Adv Microb Physiol 39:131–203
Barsom EK, Hatfull GF (1996) Characterization of a Mycobacterium smegmatis gene that confers resistance to phages L5 and D29 when overexpressed. Mol Microbiol 21(1):159–170
Tam P-H, Lowary TL (2009) Recent advances in mycobacterial cell wall glycan biosynthesis. Curr Opin Chem Biol 13(5):618–625
Crick DC, Mahapatra S, Brennan PJ (2001) Biosynthesis of the arabinogalactan-peptidoglycan complex of Mycobacterium tuberculosis. Glycobiology 11(9):107R–118R
Kremer L, Dover LG, Morehouse C, Hitchin P, Everett M, Morris HR, Dell A, Brennan PJ, McNeil MR, Flaherty C (2001) Galactan biosynthesis in Mycobacterium tuberculosis Identification of a bifunctional UDP-galactofuranosyltransferase. J Biol Chem 276(28): 26430–26440
Alderwick L, Birch H, Mishra A, Eggeling L, Besra G (2007) Structure, function and biosynthesis of the Mycobacterium tuberculosis cell wall: arabinogalactan and lipoarabinomannan assembly with a view to discovering new drug targets. Biochem Soc T 35(5):1325–1328
Besra GS, Brennan PJ (1997) The mycobacterial cell envelope. J Pharm Pharmacol 49(S1):25–30
Dover LG, Alderwick LJ, Brown AK, Futterer K, Besra GS (2007) Regulation of cell wall synthesis and growth. Curr Mol Med 7(3):247–276
Besra GS, Khoo K-H, McNeil MR, Dell A, Morris HR, Brennan PJ (1995) A new interpretation of the structure of the mycolyl-arabinogalactan complex of Mycobacterium tuberculosis as revealed through characterization of oligoglycosylalditol fragments by fast-atom bombardment mass spectrometry and 1H nuclear magnetic resonance spectroscopy. Biochemistry 34(13):4257–4266
Kaur D, Guerin ME, Škovierová H, Brennan PJ, Jackson M (2009) Biogenesis of the cell wall and other glycoconjugates of Mycobacterium tuberculosis. Adv Appl Microbiol 69:23–78
Favrot L, Ronning DR (2012) Targeting the mycobacterial envelope for tuberculosis drug development. Expert Rev Anti Infect Ther 10(9):1023–1036
Nigou J, Gilleron M, Puzo G (2003) Lipoarabinomannans: from structure to biosynthesis. Biochimie 85(1):153–166
Kaur D, Berg S, Dinadayala P, Gicquel B, Chatterjee D, McNeil MR, Vissa VD, Crick DC, Jackson M, Brennan PJ (2006) Biosynthesis of mycobacterial lipoarabinomannan: role of a branching mannosyltransferase. P Natl Acad Sci 103(37):13664–13669
Besra GS, Morehouse CB, Rittner CM, Waechter CJ, Brennan PJ (1997) Biosynthesis of mycobacterial lipoarabinomannan. J Biol Chem 272(29):18460–18466
Mishra AK, Driessen NN, Appelmelk BJ, Besra GS (2011) Lipoarabinomannan and related glycoconjugates: structure, biogenesis and role in Mycobacterium tuberculosis physiology and host–pathogen interaction. FEMS Microbiol Rev 35(6):1126–1157
Strohmeier GR, Fenton MJ (1999) Roles of lipoarabinomannan in the pathogenesis of tuberculosis. Microbes Infect 1(9):709–717
Gaitonde V, Sucheck SJ (2015) Antitubercular drugs based on carbohydrate derivatives. carbohydrate chemistry: state of the art and challenges for drug development: an overview on structure, biological roles, synthetic methods and application as therapeutics, 441
Singer MA, Lindquist S (1998) Multiple effects of trehalose on protein folding in vitro and in vivo. Mol Cell 1(5):639–648
Erdei É, Molnár M, Gyémánt G, Antal K, Emri T, Pócsi I, Nagy J (2011) Trehalose overproduction affects the stress tolerance of Kluyveromyces marxianus ambiguously. Bioresource Technol 102(14):7232–7235
Paul MJ, Primavesi LF, Jhurreea D, Zhang Y (2008) Trehalose metabolism and signaling. Annu Rev Plant Biol 59:417–441
Gavalda S, Bardou F, Laval F, Bon C, Malaga W, Chalut C, Guilhot C, Mourey L, Daffé M, Quémard A (2014) The polyketide synthase Pks13 catalyzes a novel mechanism of lipid transfer in mycobacteria. Chem Bio 21(12):1660–1669
Li W, Upadhyay A, Fontes FL, North EJ, Wang Y, Crans DC, Grzegorzewicz AE, Jones V, Franzblau SG, Lee RE (2014) Novel insights into the mechanism of inhibition of MmpL3, a target of multiple pharmacophores in Mycobacterium tuberculosis. Antimicrob Agents Chemother 58(11):6413–6423
Harth G, Lee B-Y, Wang J, Clemens DL, Horwitz MA (1996) Novel insights into the genetics, biochemistry, and immunocytochemistry of the 30-kilodalton major extracellular protein of Mycobacterium tuberculosis. Infect Immun 64(8):3038–3047
Jackson M, Raynaud C, Lanéelle M A, Guilhot C, Laurent‐Winter C, Ensergueix D, Gicquel B, Daffé M Inactivation of the antigen 85C gene profoundly affects the mycolate content and alters the permeability of the Mycobacterium tuberculosis cell envelope. Mol Microbiol 31(5):1573–1587
De Smet KA, Weston A, Brown IN, Young DB, Robertson BD (2000) Three pathways for trehalose biosynthesis in mycobacteria. Microbiology 146(1):199–208
Gibson RP, Turkenburg JP, Charnock SJ, Lloyd R, Davies GJ (2002) Insights into trehalose synthesis provided by the structure of the retaining glucosyltransferase OtsA. Chem Biol 9(12):1337–1346
Cole S, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon S, Eiglmeier K, Gas S, Barry CR (1998) Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393(6685):537–544
Sassetti CM, Boyd DH, Rubin EJ (2003) Genes required for mycobacterial growth defined by high density mutagenesis. Mol Microbiol 48(1):77–84
Murphy HN, Stewart GR, Mischenko VV, Apt AS, Harris R, McAlister MS, Driscoll PC, Young DB, Robertson BD (2005) The OtsAB pathway is essential for trehalose biosynthesis in Mycobacterium tuberculosis. J Biol Chem 280(15):14524–14529
Kalscheuer R, Weinrick B, Veeraraghavan U, Besra GS, Jacobs WR (2010) Trehalose-recycling ABC transporter LpqY-SugA-SugB-SugC is essential for virulence of Mycobacterium tuberculosis. P Natl Acad Sci 107(50):21761–21766
Portevin D, de Sousa-D’Auria C, Houssin C, Grimaldi C, Chami M, Daffé M, Guilhot C (2004) A polyketide synthase catalyzes the last condensation step of mycolic acid biosynthesis in mycobacteria and related organisms. P Natl Acad Sci 101(1):314–319
Fraga J, Maranha A, Mendes V, Pereira PJB, Empadinhas N, Macedo-Ribeiro S (2015) Structure of mycobacterial maltokinase, the missing link in the essential GlgE-pathway. Sci Rep 5
Kalscheuer R, Jacobs WR Jr (2010) The significance of GlgE as a new target for tuberculosis. Drug News Perspect 23(10):619–624
Syson K, Stevenson CE, Rejzek M, Fairhurst SA, Nair A, Bruton CJ, Field RA, Chater KF, Lawson DM, Bornemann S (2011) Structure of Streptomyces maltosyltransferase GlgE, a homologue of a genetically validated anti-tuberculosis target. J Biol Chem 286(44):38298–38310
Syson K, Stevenson CE, Rashid AM, Saalbach G, Tang M, Tuukkanen A, Svergun DI, Withers SG, Lawson DM, Bornemann S (2014) Structural insight into how streptomyces coelicolor maltosyl transferase GlgE binds α-Maltose-1-phosphate and forms a maltosyl-enzyme intermediate. Biochemistry 53(15):2494–2504
Veleti SK, Lindenberger JJ, Ronning DR, Sucheck SJ (2014) Synthesis of a C-phosphonate mimic of maltose-1-phosphate and inhibition studies on Mycobacterium tuberculosis GlgE. Bioorg Med Chem 22(4):1404–1411
Lindenberger JJ, Veleti SK, Wilson BN, Sucheck SJ, Ronning DR (2015) Crystal structures of Mycobacterium tuberculosis GlgE and complexes with non-covalent inhibitors. Sci Rep 5
Provencher L, Steensma DH, Wong C-H (1994) Five-membered ring azasugars as potent inhibitors of α-L-rhamnosidase (naringinase) from Penicillium decumbens. Bioorg Med Chem 2(11):1179–1188
de Melo EB, da Silveira Gomes A, Carvalho I (2006) α- and β-Glucosidase inhibitors: chemical structure and biological activity. Tetrahedron 62(44):10277–10302
Izquierdo I, Plaza MT, Yáñez V (2007) Polyhydroxylated pyrrolidines: synthesis from d-fructose of new tri-orthogonally protected 2, 5-dideoxy-2, 5-iminohexitols. Tetrahedron 63(6):1440–1447
Veleti SK, Lindenberger JJ, Thanna S, Ronning DR, Sucheck SJ (2014) Synthesis of a Poly-hydroxypyrolidine-Based inhibitor of Mycobacterium tuberculosis GlgE. J Org Chem 79(20):9444–9450
Thanna S, Lindenberger JJ, Gaitonde VV, Ronning DR, Sucheck SJ (2015) Synthesis of 2-deoxy-2, 2-difluoro-α-maltosyl fluoride and its X-ray structure in complex with Streptomyces coelicolor GlgEI-V279S. Org Biomol Chem 13(27):7542–7550
Veleti SK, Petit C, Ronning DR, Sucheck SJ (2016) Synthesis and inhibition studies of proline and pyrolidene-based phosphonates to inhibit Streptomyces Coelicolor (Sco) GlgE (manuscript in preparation)
Sengupta S, Roy D, Bandyopadhyay S (2014) Modeling of a new tubercular maltosyl transferase, GlgE, study of its binding sites and virtual screening. Mol Biol Rep 41(6):3549–3560
Arooj M, Sakkiah S, Kim S, Arulalapperumal V, Lee KW (2013) A combination of receptor-based pharmacophore modeling & QM techniques for identification of human chymase inhibitors. PLoS One 8(4):e63030
Silverman RB, Holladay MW (2014) The organic chemistry of drug design and drug action
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2012) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 64:4–17
Cramer CJ (2013) Essentials of computational chemistry: theories and models. Wiley, New Jersey
Billones JB, Valle AMF (2014) Structure-based design of inhibitors against maltosyltransferase GlgE. Orient J Chem 30(3):1137–1145
Sengupta S, Roy D, Bandyopadhyay S (2015) Structural insight into Mycobacterium tuberculosis maltosyl transferase inhibitors: pharmacophore-based virtual screening, docking, and molecular dynamics simulations. J Biomol Struct Dyn 33(12):2655–2666
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG
About this chapter
Cite this chapter
Veleti, S.K., Sucheck, S.J. (2018). Glycoconjugate-Based Inhibitors of Mycobacterium Tuberculosis GlgE. In: Witczak, Z., Bielski, R. (eds) Coupling and Decoupling of Diverse Molecular Units in Glycosciences. Springer, Cham. https://doi.org/10.1007/978-3-319-65587-1_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-65587-1_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-65586-4
Online ISBN: 978-3-319-65587-1
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)