Abstract
The fast development and capabilities of high-throughput ‘omics’ technologies have provided new insights into the complexity of endometriosis and enabled the identification of novel diagnostic biomarkers. In this chapter, we take a closer look at high-throughput genomics, transcriptomics, epigenomics, proteomics, and metabolomics studies applied in endometriosis research. We summarise the existing information concerning ‘omics’ studies applied to blood, endometrium, endometriotic lesions, and body fluids in order to describe the potential disease-specific biomarkers. Also, we discuss the importance of sample collection, proper study design, data processing, and analysis in high-throughput studies. And finally, future perspectives in endometriosis biomarker research will be provided.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Conventional methods have not been very successful in endometriosis-specific biomarker discovery, and to date there are no reliable non-invasive or minimally invasive diagnostic markers for endometriosis. Therefore, there is a considerable need for non-invasive biomarkers, because due to the non-specific symptoms , the average delay between the onset of symptoms and the surgical diagnosis is almost 7 years [1]. The delayed diagnosis may in turn lead to more severe complications and is associated with remarkable healthcare costs [2]. Objective and reliable non-invasive diagnostic biomarkers would not only avoid the unnecessary laparoscopy in suspicious cases but would also make it possible to get the diagnosis of endometriosis earlier and thus provide an easy strategy for monitoring the disease treatment efficacy and recurrence [3]. However, despite extensive research in this field during the past 10 years, there are still no reliable non-invasive diagnostic markers for endometriosis [4, 5], and numerous women with nonspecific complaints, such as infertility and pelvic pain, undergo diagnostic laparoscopy. Thus, ‘omicsʼ-level studies using both easily assessable materials like blood, urine, and menstrual blood but also endometrium and endometriotic lesions are one of the top research priorities in the field.
The high-throughput techniques provide massive data from the genome (variability in DNA sequence in the genome, i.e. genomics), epigenome (epigenetic modifications of DNA, i.e. epigenomics), transcriptome (variability in composition and abundance of mRNA and miRNA levels, i.e. transcriptomics), proteome (variability in composition and abundance of the proteins, i.e. proteomics), and metabolome (variability in composition and abundance of metabolites, i.e. metabolomics/metabonomics). The major advantage of ‘omics’ studies is that the data can be collected without existing hypotheses, and a primary research question is not always needed (first experiment-then-hypothesise approach) [6]. This could be particularly useful when studying complex diseases with unknown pathogenesis, such as endometriosis. There are still many missing pieces in the puzzle of endometriosis, and the new ‘omicsʼ studies promise to add new biological knowledge transferrable into the development of disease-specific biomarkers. The considerable increase (15 publication in 1999–2006, 104 publication in 2007–2016, altogether 118 studies) in ‘omicsʼ research is a definite sign that the ‘omicsʼ revolution in endometriosis is actively ongoing.
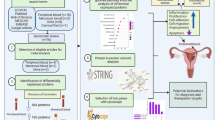
In this chapter, we take a closer look at the high-throughput studies applied in endometriosis research, namely, genomics, transcriptomics, epigenomics, proteomics, and metabolomics (Fig. 1). We summarise the existing information concerning endometrium, endometriotic lesions, blood, and body fluids in order to describe the potential disease-specific biomarkers. Also, future perspectives of single-cell ‘omicsʼ in endometriosis biomarker research will be provided. And finally, we discuss the importance of sample collection and proper study design in high-throughput studies.
‘Omicsʼ publications in endometriosis studies. Literature search was performed in PubMed up to December 2016. Only publications that were in English were considered. The keyword ‘endometriosis’ was one-by-one searched with terms: ‘endometrium + microarrayʼ, ‘miRNA + microarrayʼ, ‘sequencingʼ, ‘microarrayʼ, ‘gene expression + microarrayʼ, ‘exome sequencingʼ, ‘GWASʼ, ‘CNVʼ, ‘genomicsʼ, ‘proteomicsʼ, ‘metabolomicsʼ, ‘DNA methylation + microarrayʼ, ‘DNA alterations + microarrayʼ, and ‘proteomeʼ. Some of the eligible studies were identified using the reference list of appropriate review articles. In total 118 ‘omicsʼ studies were included into this review chapter
Search for Endometriosis Biomarkers: ‘Omics’ Studies and Endometrium
Endometrium is not just a uniform tissue that undergoes cyclical changes under the influence of endogenous hormones, cytokines, and chemokines but an assortment of different cells, each with their own special functions responsible for tissue differentiation, desquamation, and regeneration. It is evident that eutopic endometrium of women with endometriosis functions normally and has almost comparable responsiveness to steroid hormones; however, there is evidence from epigenomic, transcriptomic, and proteomic studies that endometrial tissue from patients with endometriosis and healthy women is differently regulated at the molecular level. Therefore, understanding the complex mechanisms controlling the changes within the endometrium is crucial to find endometrial biomarkers for endometriosis.
Genomic studies focusing only on eutopic endometria of endometriosis patients have not been very popular, and to date, only two studies have investigated somatic DNA mutations in endometrium (Table 1). Guo et al. found a number of individual chromosomal losses and gains in laser capture microdissection (LCM)-harvested endometrial epithelial cells and hypothesised that these genomic alterations could be the proximate cause of endometriosis [7]. Li et al. conducted whole-exome sequencing of blood DNA and LCM-harvested endometrial cells from eutopic and ectopic endometria of 16 endometriosis patients and eutopic endometria of 5 healthy women [8]. They found that DNA originating from healthy endometria contains thousands of somatic mutations that are absent in blood DNA. Furthermore, the general somatic mutation spectrum in endometria of women with and without endometriosis was very similar and authors proposed that most of the mutations are probably benign and irrelevant to endometriosis pathogenesis [8].
Aberrant DNA methylation is shown to contribute to many human diseases, and there is accumulating data from DNA methylation studies that methylation alterations in certain genes could contribute to the pathogenesis of endometriosis (reviewed [9]). So far, three studies have applied genome-wide microarray-based DNA methylation analysis to eutopic endometria of endometriosis patients [10,11,12] (Table 1). The study by Naqvi et al. described several aberrantly methylated and expressed genes, among them MGMT, DUSP22, CDCA2, ID2, TNFRSF1B, ZNF681, and IGSF21 have previously not been associated with endometriosis [10]. Although several formerly known genes with altered methylation levels, including MAFB, HOXD10, and HOXD11, were highlighted, alterations in DNA methylation levels in other genes (PR-B, CYP19A1, SF1, COX2, and ER-β) previously associated with endometriosis were not confirmed. The study by Saare et al. showed that the endometrial DNA methylation profiles were highly similar between endometria of patients and controls but largely influenced by the menstrual cycle phases [11]. Authors suggested that DNA methylation differences are likely not the main reason for endometriosis development, but it is crucially important to take into account the normal epigenetic changes across the menstrual cycle when looking for disease-specific methylation differences in endometrium. A subsequent study by Houshdaran et al. compared endometrial DNA methylation patterns and associated gene expression levels in endometriosis patients and healthy controls across the menstrual cycle and found a small number of differentially methylated loci between the patients and controls [12]. The differences in endometrial DNA methylome were most contrasting between the patients and controls in the mid-secretory phase (137 CpG sites, corresponding to 125 loci), followed by proliferative (58 CpG sites, corresponding to 58 loci) and early-secretory phase (39 CpG sites, corresponding to 36 loci). Interestingly, there were no overlapping differentially methylated genes in all three genome-wide studies [10,11,12]. Based on these results, it can be proposed that the normal physiological fluctuations during the menstrual cycle may have larger impact on endometrial DNA methylation signature than disease/non-disease status, and thus, the DNA methylation changes in endometria of patients is probably not the primary cause for endometriosis development.
Several transcriptome studies have used mRNA microarray technology to resolve the question whether there are any differences between endometria of patients with endometriosis and healthy women [13,14,15,16,17,18,19,20,21,22,23,24,25] (Table 1). While a majority of these studies have yielded numerous candidate genes, the amount of genes which have consistently been shown as up- or downregulated has remained small. Aghajanova and Giudice provided evidence that also the endometria from patients with different endometriosis stages have differences on the molecular level [18]. Further, the authors proposed that the influence of menstrual cycle phase on endometrial transcriptome could be larger than the presence or absence of endometriosis. Still, dysregulation of progesterone and/or cyclic adenosine monophosphate (cAMP) -regulated genes and genes related to thyroid hormone action and metabolism between endometria of patients with different endometriosis stages and menstrual cycle phases was found. Also, upregulation of epidermal growth factor receptor (EGFR) and extracellular matrix proteoglycan versican (VCAN) during the early secretory phase was found in severe versus mild disease [18]. The pathway analysis of differently expressed genes in endometria of patients with severe endometriosis exhibited dysregulation of PI3K/AKT, JAK/STAT, SPK/JNK, and MAPK pathways that have been associated with endometriosis pathogenesis in several studies [26]. The study conducted by Tamaresis et al., comparing endometria of patients and controls, found 18 upregulated and 11 downregulated genes in all three studied menstrual phases, and also a number of genes were dysregulated in patients with different stages of the disease [19]. They used gene expression data of 148 women to develop a molecular classifier that distinguishes endometria of women with and without endometriosis and found that the best performing classifiers, enabling identification of endometriosis with 90–100% accuracy, were mostly menstrual phase specific and utilised relatively few genes to determine the presence and severity of the disease. Multiple pathways were found to be activated in the proliferative and early secretory phase endometrium (JAK/STAT, EGF/PGF/DGF, PI3K-AKT signalling, p53 signalling, integrin-mediated cell adhesion) of women with moderate-severe endometriosis compared to minimal-mild endometriosis [19], and this was in good concordance with the previous results [18]. Dysregulation of the RAS/RAF/MAPK and PI3 kinase signalling pathway genes, which participate in a wide variety of cellular functions and cell survival, is identified in several studies [18,19,20, 25], referring to a link between these pathways and disease pathogenesis. Ahn et al. noticed that based on the unsupervised hierarchic clustering analysis, the overall gene expression signature of endometria from patients and controls was similar [23]. Still, 91 differentially expressed genes involved in regulation of decidualization, cellular adhesion, cytokine-cytokine receptor interaction, apoptosis, and complement pathway were found. In the latest study by Zhou et al., mid-secretory endometria from patients and controls were analysed and 357 differentially expressed mRNAs were found to be involved in signalling pathways such as the JNK/MAPK, PI3K-AKT, p53, adherens junction, and calcium signalling pathway [20]. In addition to studies reporting distinct endometrial molecular signatures of endometriosis patients, there are also evidence that endometrial receptivity gene signature during the implantation window is similar in patients with endometriosis and healthy women [22, 27].
Several microarray-based microRNA (miRNA) studies concentrating on eutopic endometria have been performed [28,29,30,31,32]. Burney et al. studied eutopic endometria from patients and controls to reveal a disease-specific endometrial miRNA signature [28]. They found six downregulated miRNAs from miR-9 and miR-34 families in eutopic endometria of endometriosis patients and suggested that downregulation of miR-34 family could be involved in maintaining the molecular fingerprint in proliferative endometrium and mediate the delayed proliferative to secretory transition observed in women with moderate-severe endometriosis [28]. A following study by Laudanski et al. reported a lower expression of miR-483-5p and miR-629* in the eutopic endometrium of women with advanced ovarian endometriosis compared to controls [29]. They suggested that expression changes of these miRNAs are a consequence of an early defect in the physiological activity of the proliferative endometrium, ultimately resulting in the overgrowth of this tissue outside the uterus [29]. Subsequently, Laudanski et al. utilised a more comprehensive array and reported the presence of 136 upregulated miRNAs in the eutopic endometrium of patients with endometriosis compared with the healthy women [30]. However, after validation, only three out of 11 validated miRNAs revealed borderline significance. In the study by Braza-Boils et al., both eutopic endometria from patients and controls and endometriotic tissues were studied, and only five miRNAs were found to be differentially expressed in eutopic endometria of endometriosis patients compared to healthy endometrium [31]. Thirty-six downregulated miRNAs in endometria of patients were also reported by Shi et al. [32]. However, the comparison of all results from aforementioned miRNA studies showed a minute overlap, and only two miRNAs (miR-9* [28, 32] and miR-636 [31, 32]) were reported in at least two studies. Therefore, as different miRNA studies have reported different candidate miRNAs, the potential application of endometrial miRNAs as endometriosis biomarkers is still limited. Clearly, our knowledge about the endometrial miRNome and its physiological and pathophysiological significance in association with endometriosis is scarce and remains to be unravelled.
The functional interpretation and understanding of the proteome is one of the current challenges in biology due to the presence of sequence variations, alternative splicing, and epigenetic and post-translation modifications [33, 34]. The complexity of the proteome is illustrated by the fact that there is a poor correlation between the transcript levels and the abundance of the corresponding proteins [35, 36]. Proteomic research in endometriosis is currently a ‘hot topicʼ, and a number of endometrial proteome studies have been performed in endometriosis patients [13, 35, 37,38,39,40,41,42,43,44] (Table 1). A long list of potential disease-related proteins has been proposed but only a few of them, like vimentin, peroxiredoxin, HSP70, HSP90, annexins, actins, and 14-3-3 family proteins (phosphoserine- or phosphothreonine-binding proteins), are consistently identified as differentially expressed in patients in at least three different studies. A recent excellent review by Siva et al. summarises the current situation in the proteomic research field—so far no clear biomarker or therapeutic targets have been discovered [45]. Nevertheless, as the protein synthesis is the final result of the gene expression and is directly linked to the phenotype, the endometrial proteome studies do hold a great promise for future biomarker discovery.
Taken together, the large-scale ‘omicsʼ studies have provided clear evidence that the endometrial genome, epigenome , transcriptome , and proteome are differently regulated in endometriosis. Although the concordance between different ‘omics’ studies has been moderate, some potential biomarkers such as miR-9 and miR-636 family; disease-related pathways PI3K/AKT, JAK/STAT, SPK/JNK, and MAPK from transcriptome studies; and proteins like vimentin, peroxiredoxin, HSP70, HSP90, annexins, actins, and 14-3-3 family members from proteome studies have been proposed.
‘Omicsʼ Studies of Endometriotic Lesions and Possible Biomarkers
When it comes to biomarkers research, endometriotic lesions are a less-favoured study object than endometrial biopsies , as lesions do not provide direct non-invasive or minimally invasive biomarkers for clinical use. Nevertheless, the studies using lesions are crucial for detecting molecular alterations involved in the disease development and pathogenesis and thereby provide valuable information for biomarker research.
Microarrays and single nucleotide polymorphism (SNP) genotyping technologies together with recent advances in high-throughput sequencing have led to a rapid progress in genomic studies in endometriosis and provided new evidence about the genetic background of the disease. However, genome-wide studies have provided no clear consensus about the somatic DNA alterations either in endometriotic lesions and/or eutopic endometria. A number of studies have reported chromosomal alterations, more frequently gains in chromosomes 1p, 3p, 6q, 17q, and Xq and losses in chromosomes 1p, 5p, and 6q [46,47,48,49,50], while other studies have found no chromosomal aberrations in ectopic endometrial tissue or eutopic endometrium [51, 52], thus raising a question about the relevance of DNA genomic imbalance in the pathogenesis of endometriosis (Table 1). Saare et al. used SNP microarrays instead of traditional array comparative genomic hybridization (aCGH)] to compare the same patients’ blood, endometria, and LCM-harvested cells of endometriotic lesions and found no evidence of disease-specific somatic DNA copy number alterations (SCNAs) [52]. The authors suggested that some SCNAs identified in previous studies may be related to the detection methodology (CGH or array-CGH) as it has been shown that some G-C-rich chromosomal regions (1p and 16p and chromosomes 19 and 22) tend to give false-positive results [50, 53].
The identification of epigenetic biomarkers for endometriosis diagnostics is definitely an emerging, challenging, and still largely uncovered field of investigation. In the recent years, researchers have turned their major interest from transcriptome to epigenome , and a few large-scale epigenome studies, describing the lesion-specific DNA methylation profiles, have been performed (Table 1). The genome-wide DNA methylation profiles of endometriotic lesions or stromal cells originating from lesions have been described in three studies [54,55,56]. Borghese et al. published the first study describing the global DNA methylation profile of different endometriotic lesions, including ovarian endometriomas, deep infiltrating endometriosis, and superficial endometriosis, and showed that global methylation pattern was similar in different lesion types and eutopic endometria [55]. When global methylation data of eutopic endometria was compared to ovarian endometriomas in combination with gene expression data, 35 genes were found to share alterations both in methylation and expression patterns [55]. Specific regions were consistently hypermethylated (or hypomethylated) in all subtypes of the disease, and other regions were strictly altered in one endometriosis type only, and variation in methylation was more likely to occur at discreet loci across the genome. The later study by Dyson et al. found more than four thousand differentially methylated CpGs when stromal cells from eutopic endometria were compared to stromal cells from endometriomas, and the authors concluded that endometriotic cells possess a unique epigenetic fingerprint [54]. The analysis of differentially methylated and expressed genes identified 403 genes that were aberrantly methylated and differentially expressed in endometriosis, among them are genes from the HOXA cluster, ESR1, NR5A1, and GATA family transcription factors [54]. Although a different study design (entire lesions vs. cultivated stromal cells) and platforms were applied to interrogate DNA methylation, both investigations [54, 55] reported different methylation of ADAP1, HPCAL1, PRKAG2, PRKCZ, RIPK1, SEC61A1, ZNF22, and HOXD10 genes. Most recent study by Yamagata et al. analysed stromal cell cultures from endometriomas and eutopic endometria of patients and controls and found that methylation profiles of eutopic endometria were very similar but significantly different from stromal cells originating from endometriomas [56]. The genes with altered methylation in endometriomas were related to signal transduction, molecular functions of receptors and signalling molecules, and cytokine-cytokine receptor interactions and development. Comparison of datasets from Yamagata et al. and Dyson et al. revealed four overlapping genes: HOXD10, BST2, GATA4, and TCF21, but when all three DNA methylation studies were compared to each other, only HOXD10 was seen to be differentially methylated.
The first transcriptome studies in endometriosis applying microarray technology were conducted already in 2002, and since then, many studies have been carried out to reveal the specific gene expression profile of endometriotic lesions. Transcriptome studies in lesions can be divided into two groups—studies performed in 2002–2007 [57,58,59,60,61,62,63,64,65] that used less comprehensive microarrays [up to 23 thousand (23K) probes] and studies conducted in the recent years using advanced large-scale microarrays covering 44K or 60K probes [23, 30, 51, 66,67,68,69,70,71] (Table 1). Although the list of candidates in each study contains a remarkable number of dysregulated genes in ectopic endometrial tissue compared to eutopic endometria, there is little concordance in the reported genes between studies. However, altered expression of genes belonging to RAS, MAPK, and PI3K signalling pathways was proposed in several studies (reviewed in [26]). Khan et al. found 50 differently expressed genes associated with immunological, neurocrine, and endocrine functions and gynaecological cancers (CHEK1, ERBB family, laminin gamma, and Ki-67), but there was no overt oncogenic potential in endometriotic tissue [66]. Also, they reported a list of 28 novel genes that were not previously associated with endometriosis, representing potential markers for ovarian endometriosis. The following studies by Monsivais et al., Crispi et al., and Suryawanshi et al. found many dysregulated genes that belong mostly to tissue and organ development pathways [68], pathways regulating metabolism and action of prostaglandins and glucocorticoids [72], and complement pathway [24]. Sun et al. used a microarray comprising of probes for long non-coding RNAs (lncRNA) and mRNAs and found hundreds of dysregulated lncRNAs and thousands of mRNA transcripts in ectopic endometrial tissues compared to paired eutopic endometrial tissues [67]. Authors proposed that many dysregulated lncRNAs may participate in biological pathways related to endometriosis through cis- and trans-regulation of target protein-coding genes. In the latest study by Ahn et al., a large number of differentially expressed genes involved in cytokine-cytokine receptor interaction, cellular adhesion, immune cell recruitment, apoptosis, cell signalling, T-cell cytotoxicity, and regulation of inflammatory responses were found [23].
To date, eight high-throughput miRNA studies describing the miRNome of the whole endometriotic lesion biopsies or cultured stromal cells from lesions have been performed [31, 32, 73,74,75,76,77,78] (Table 1). Each study has identified a subset of miRNAs that has been differently expressed in ectopic lesions compared to eutopic endometria. As there is a large variability between studies in the terms of design, analysis methods, and selection of controls, the concordance has been moderate, and only 22% of reported miRNAs are consistent between studies [79]. In addition to microarrays, next-generation miRNA sequencing technology has also been applied in endometriosis studies [73, 76]. Hawkins et al. compared specimens from endometrioma and normal endometrium and found several miRNAs that were upregulated (miR-29c, miR-100, miR-193a-5p, miR-202, miR-485-3p, miR-509-3-5p, miR-708, and miR-720) or downregulated (miR-10a, miR-34c-5p, miR-141, miR-200a/b/c, miR-203, miR-375, miR-429, miR-449b, miR-504, and miR-873) in endometriomas [73]. The following study by Saare et al. investigated paired samples of peritoneal endometriotic lesions and matched healthy surrounding tissues together with eutopic endometria of the same patients and found five miRNAs (miR-34c-5p, miR-449a, miR-200a, miR-200b, and miR-141) that were significantly overexpressed in lesions compared to healthy surrounding tissues [76]. Although majority of these miRNAs were reported to be associated with endometriosis pathogenesis in Hawkins et al. [73] and Ohlsson Teague et al. [75] studies, Saare et al. [76] concluded that these miRNAs rather reflect the presence of endometrial cells in the peritoneal tissue than are associated with pathologic events. Furthermore, the authors suggested that the miRNA profile of peritoneal endometriotic lesions is largely masked by the surrounding peritoneal tissue present in biopsy samples, challenging the discovery of an accurate lesion-specific miRNA profile [76]. Still, it should be pointed out that according to the results of all these miRNome studies, there was only one single miRNA (miR-200b) that was differentially expressed in all six studies in whole lesions [32, 73,74,75,76,77] but not in endometriotic stromal cells [78]. miR-200b, member of the miR-200 family, could be an attractive molecular marker that can be easily linked to disease pathogenesis because of its important role in cell migration and epithelial-mesenchymal transition (EMT). It could be proposed that altered expression of miR-200b in lesions changes the well-balanced network of EMT and leads the endometrial epithelial cells to acquire mesenchymal phenotype with higher migratory capacity.
To summarise, transcriptome studies of endometriotic lesions have already provided some clues about the pathogenesis of endometriosis, though the concern still remains about using whole-tissue biopsies instead of pure populations of endometrial epithelial and stromal cells from lesions to reveal transcriptome changes inside the lesion.
Only a few studies on endometriotic tissue proteomics have been conducted so far [80,81,82,83,84] (Table 1). However, it could be hypothesised that if disease-specific proteins with high concentration in affected tissues exist and are secreted from lesions into the blood stream, they could also be monitored in the body fluids [81] and thereby could offer potential for discovery of non-invasive markers. The results of proteome studies have shown that some proteins are differentially expressed (e.g. SM-22α and Rab37), modified (e.g. haptoglobin and Rho-GDIα,), or localised (e.g. haptoglobin) in endometriotic lesions compared to eutopic endometria of patients or healthy women [80]. Also, significant increase in transforming growth factor β-1, calponin-1, and emilin-1 [81] in ovarian endometriomas has been reported. In the recent study by Kasvandik et al., metabolic reprogramming of ectopic endometrial stromal cells with extensive upregulation of glycolysis and downregulation of oxidative respiration was noticed [82].
The above-discussed examples from genomic, epigenomic, transcriptomic, and proteomic levels have provided inconsistent evidence about the possible molecular changes occurring in the endometriotic lesions. The genomic studies in endometriotic lesions have reported chromosomal alterations more frequently in chromosomes 1p, 3p, 5p, 6q, 17q, and Xq; epigenome studies have provided evidence on altered DNA methylation in HOXD10; transcriptome studies have found altered expression of genes belonging to RAS, MAPK, and PI3K signalling pathways, and proteome studies have found upregulation of glycolysis and downregulation of oxidative respiration and differently expressed proteins like SM-22α, Rab37, haptoglobin, and Rho-GDIα. Although the ‘omicsʼ studies in endometriotic lesions require invasive procedures and will not provide biomarkers directly translatable into the clinical practice, the knowledge obtained from these studies enables more complex insight into the possible mechanisms of endometriosis pathogenesis.
‘Omicsʼ Studies on Blood and Body Fluids and Novel Endometriosis Biomarkers
The ultimate goal of ‘omicsʼ studies in endometriosis is to find robust and specific non-invasive biomarkers with acceptable sensitivity and specificity and preferably from easily assessable sources like blood and body fluids. Endometriosis biomarkers have been sought from peripheral blood (whole blood, plasma, serum), menstrual blood, endometrial fluid, peritoneal fluid and urine samples, and more than 100 markers, among them annexin V, VEGF, CA-125 and sICAM-1/or glycodelin, glycoproteins, inflammatory and non-inflammatory cytokines, and angiogenic and growth factors have been reported but with inconsistent and contradictory results [4, 5, 85, 86].
The first large-scale genomic study from blood was published in 2010, and since then, numbers of SNP microarray-based genome wide association studies (GWAS) from genomic DNA, together with following replication studies, have been conducted to reveal associations between common SNPs and endometriosis [87,88,89,90,91,92,93,94,95]. Previous meta-analysis included more than 11,506 patients and 32,678 controls and found six loci (rs12700667 on 7p15.2, rs7521902 near WNT4, rs10859871 near VEZT, rs1537377 near CDKN2B-AS1, rs7739264 near ID4, and rs13394619 in GREB1) that had consistent effects across different populations [96]. In a recent study by Steinthorsdottir et al., 1840 women with endometriosis and 129,016 controls were included into GWAS, and in addition to the previously reported susceptibility loci [96], also two new loci on 4q12 (rs17773813) upstream of KDR encoding vascular endothelial growth factor receptor 2 (VEGFR2) and rs519664 in TTC39B gene on 9p22 were identified [95].
Although GWAS have provided valuable information about novel candidate genes and genome regions, the effect sizes for the associated variants are quite moderate (odds ratios between 1.0 and 1.2), and apparently these common variants do not provide any molecular markers for direct diagnostic or prognostic tests. However, it is very likely that in the case of a common diseases, such as endometriosis, the rare variants (minor allele frequency <0.05) could contribute to the risk of the disease [97].
In addition to GWAS , high-resolution SNP arrays provide the opportunity to assess inherited copy number variations (CNVs) that normally exist in all tissues and may potentially contribute to genetic predisposition of common diseases. Although many disease-related CNVs have been described, large population-based CNV studies have also found substantial variability in CNV distribution in healthy individuals [98,99,100], challenging the findings of CNVs responsible for the development of a complex diseases. Genomic CNVs in endometriosis have been investigated so far only in two large-scale studies [101, 102]. Chettier et al. conducted a case-control study of 2126 surgically confirmed endometriosis cases and 17,974 controls of European ancestry and found no significant differences in CNV counts, excess of large CNVs, and gene-based CNVs between controls and patients [101]. However, the locus-specific analysis revealed 22 rare CNVs that were detected in 6.9% of the affected women compared to 2.1% of the general population. Three out of 22 CNVs passed a genome-wide P-value threshold, namely, a deletion at SGCZ on 8p22, a deletion in MALRD1 on 10p12.31, and a deletion at 11q14.1 [101]. Recently, six sub-telomeric putative novel CNV loci in regions 1p36.33, 16p13.3, 19p13.3, 20p13, 17q25.3, and 20q13.33 from pooled DNA samples of 100 patients and 50 controls were reported by Mafra et al. [102]. Though the genomic studies have not been very successful in uncovering the pathogenesis of endometriosis or finding disease-specific biomarkers, it is very likely that the availability of more advanced methodologies (exome sequencing and whole-genome sequencing) will provide more detailed information about the genomic background of endometriosis in the near future.
Transcriptome studies have focused on determining circulating miRNAs in blood serum or plasma. miRNAs are considered as good candidates for biomarkers because cell-free miRNAs are shown to be stable in different body fluids [103]. In healthy individuals, the miRNA profile in serum is similar to that of circulating blood cells, but in the case of physiological or pathological changes, the levels of miRNAs in serum may differ [103]. Thus, alterations in the miRNA levels may possibly be used as biomarkers for endometriosis [104]. Although more than 200 potential non-invasive miRNA biomarkers have been proposed for endometriosis [105,106,107,108,109,110], the results are still inconsistent between studies, and only 12 miRNAs (miR-548b-5p, miR-92a, miR-320d, miR-139-3p, miR-122, miR-145*, miR-15b, miR-21, miR-572, miR-9*, miR-199a-5p, miR-342-3p) have been found to be differentially expressed in at least two studies. Not a single miRNA alteration has been confirmed in all studies. As there are only six global miRNA studies published to date with moderate numbers of participants involved, thus the real diagnostic potential of miRNAs in endometriosis is not fully discovered, and further studies, using additionally more advanced sequencing techniques to fully describe the wide spectrum of miRNAs, are needed.
Proteomics analyses have been extensively conducted to identify endometriosis-specific biomarkers from blood plasma [111, 112], serum [37, 44, 113,114,115,116,117,118,119,120,121,122,123,124], urine [125,126,127,128], endometrial fluid aspirate [129], menstrual blood [130], and peritoneal fluid [128, 131,132,133,134,135,136,137]. Although the number of serum-based studies is quite remarkable, the correlation between the different studies is relatively small. Nevertheless, some of the studies have identified a signature of peptides/proteins that could discriminate patients from controls with relatively high sensitivity and specificity. Jing et al. found two proteins (5830 m/z and 8865 m/z) in serum samples that were significantly more abundant in patients than in healthy women, and the signature of these two proteins offered diagnostic potential with a sensitivity of 86.7% and specificity of 96.7% [115]. A following study by Zheng et al. found three peptide peaks (5988,7; 7185,3; 8929,8 m/z) in serum that distinguished endometriosis patients in test and training sets with a sensitivity of 89.3–91.4% and a specificity of 90–95% [113]. A study conducted by Long et al. reported 13 serum proteins with significantly different levels between controls and patients and proposed one promising peptide (4180 Da) with 100% specificity and 100% sensitivity [114]. In the most recent study by Dutta et al., two proteins, HP and A1BG, were found to be effective for the diagnosis of stage II, III, and IV endometriosis, with a sensitivity of 68–92% and a specificity of 84–96% [124]. In addition to the large number of serum proteome studies, a few plasma studies have been published [111, 112]. Fassbender et al. found that a model based on five protein/peptide peaks (2058; 2456; 3883; 14,694 and 42,065 m/z) discriminated ultrasonography-negative endometriosis with a sensitivity of 88% and a specificity of 84% [111]. A study by Liu et al. found 20 protein peaks that were up- or downregulated in the plasma of endometriosis patients [112]. Overall, the blood-based proteomic studies have brought out several potential biomarkers with relatively high sensitivity and specificity. However, it should be pointed out that the methods most commonly used in biomarker identification, SELDI-TOF-MS and MALDI-TOF-MS, have several shortcomings like limited mass range, and these methods do not provide direct protein identities. Therefore, for further biomarker discovery, other methods that enable identification of proteins (such as tandem mass spectrometry) are needed [138].
From the clinical perspective, urine that is simple to collect would be the most preferable source for disease-specific biomarkers . Similarly to other fluids, urine contains peptides and proteins that may reflect disease status and can be easily measured by proteomic methods. Indeed, proteomic studies from urine [125,126,127,128] have provided promising results. The study by Tokushige et al. found that cytokeratin-19 levels were not influenced by the menstrual cycle phase or disease severity and were elevated in patients with endometriosis [126]. The following study by El-Kasti et al. identified six peptides influenced by disease severity and menstrual cycle phase when controls were compared with moderate-severe endometriosis patients, and seven peptides when patients with minimal-mild disease were compared to moderate-severe endometriosis patients [125]. The study by Cho et al. found 22 protein spots with differential expression in patients, among them is urinary vitamin D-binding protein (VDBP) . However, the diagnostic potential of VDBP in endometriosis alone or combined with serum CA-125 remained moderate [127]. No specific urine proteomic biomarkers that could discriminate patients and controls were found in a study by Williams et al. [128].
The full potential of menstrual blood as a source of biomarkers is yet to be found. Although the collection of menstrual blood is fairly complicated, it could reflect the physiological and molecular environment of the pathologically altered endometrial cells of endometriosis patients more precisely compared to the peripheral blood (reviewed in [138]). However, there is only one study using menstrual blood for biomarkers research published so far [130]. This study identified three differentially expressed proteins as endometriosis-specific markers: collapsin response mediator protein 2, ubiquitin carboxyl-terminal hydrolase isozyme L1, and myosin regulatory light polypeptide 9 [130]. Additionally, the study reported higher expression of stem cell marker gene transcripts (Oct-4, CXCR4, SOX2, and c-MET) in the menstrual blood of patients with endometriosis [130]. The higher expression of stem cell markers in menstrual blood of women with endometriosis may indicate the importance of these markers in implantation process of endometriotic lesion.
Microenvironment of the peritoneal cavity is thought to be one important factor influencing the capability of endometrial cells to implant into the peritoneal cavity. Further, there is strong evidence that dysregulation of peritoneal immunological and proinflammatory systems, and also alterations in angiogenesis processes may play a crucial role in the progression of the disease. Therefore, a number of proteome studies of peritoneal fluid have been conducted, and several potential biomarkers have been proposed [128, 131,132,133,134,135,136]. Silicano et al. studied peritoneal fluid in patients with different disease stages and found a pattern of peptides corresponding to fibrinogen alpha chain that were more frequently present in women with moderate-severe endometriosis [131]. Study by Wölfler et al. identified 11 differentially regulated proteins that might have an impact on the development and establishment of endometriotic lesions [136]. Ferrero et al. found nine proteins mostly involved in the immune response (e.g. serotransferrin, complement C3, serum amyloid P-component, alpha-1-antitrypsin, and clusterin) with significantly higher expression in infertile endometriosis patients than in infertile controls [135]. Williams et al. detected a number of proteins with metabolic functions, such as proteins involved in glucose metabolism (phosphoglycerate kinase-1, fructose-bisphosphate aldolase A, transaldolase, triosephosphate isomerase, malate dehydrogenase) and glutathione S-transferase P which is involved in detoxification [128]. Summing up, although studies in peritoneal fluid have provided some insight into the pathogenesis of endometriosis, the concordance between the results of different studies is non-existent, and to date, there are no specific reliable biomarkers for diagnostic purposes. Also, the diagnostic potential of peritoneal fluid biomarkers in clinical practice is debatable due to the invasive nature of the peritoneal fluid collection.
Beside other ‘omicsʼ technologies, metabolomics has great potential to become a new frontier in endometriosis biomarkers research, as global changes in measurable, low-molecular-weight products of metabolism are thought to be good indicators of health status. Although the concentration of circulating metabolites provides integrative information about the tissue function within the larger context of the organism, the global metabolic profile is influenced by a number of dependent variables, such as environmental factors, altered activities or levels of enzymes, genetic factors, and lifestyle factors including diet, drugs, exercise, microbiota, hormonal homeostasis, and age (see review [139]), thereby challenging the finding of disease-specific metabolites. To date, only a few global metabolome studies using either blood serum or plasma of patients and controls have been conducted [140,141,142,143]. These studies have proposed several potential metabolites with a good diagnostic potential for endometriosis. Vouk et al. proposed a model including hydroxyl sphingomyelin (SMOH C16:1) and phosphatidylcholine/ether-phospholipid ratio (PCaa C36:2/PCae C34:2) that discriminates the ovarian endometriosis patients from controls with a sensitivity of 90% and a specificity of 84% [142]. Dutta et al. found 13 metabolites that discriminated minimal-mild endometriosis patients from healthy women with a sensitivity of 82% and specificity of 91% [141]. The study by Jana et al. found 15 metabolites showing a sensitivity and specificity of 92.83% and 100%, respectively [140]. In the most recent study by Ghazi et al., several metabolites such as 2-methoxyestradiol, 2-methoxyestrone, dehydroepiandrostion, androstenedione, and cholesterol showed significant increase in the endometriosis group compared to control group [143]. As all these four studies reported different metabolites that discriminate diseased and healthy women with high sensitivity and specificity, future studies including larger number of participants and different types of endometriosis are needed.
The research in the field of non-invasive biomarkers has been comprehensive and continuously ongoing, and valuable knowledge will be obtained piece by piece. The results from transcriptome , proteome , and metabolome studies are encouraging and hold a great promise for endometriosis biomarker discovery.
Perspectives in Single-Cell ‘Omics’
The remarkable progress in ‘omicsʼ technologies using DNA or RNA from small amount of cells or even single cells has revolutionised our conceptual understanding of biological diversity of human cells and has allowed to take a closer look into the single-cell genome , proteome , epigenome , and proteome in health and disease [144].
So far all ‘omics’ studies in endometriosis have operated on the level of systemic or multicellular analysis, and results of these studies inevitably show any changes in a greatly diluted fashion, and therefore the potential of single cell technologies in endometriosis research is yet to be realised. The traditional approaches in endometriotic lesion research have not provided any clear consensus about the genetic, transcriptomic, and epigenetic changes inside the lesions, and therefore, the ‘omicsʼ information from single cells or from homogenous cell populations from lesions (e.g. endometrial epithelial and stromal cells, stem cells, endothelial cells, monocytes, NK cells, lymphocytes, and dendritic cells) could offer new prospects to reveal the true disease-specific molecular changes (Fig. 2).
The major challenge of using single-cell technologies is not related to the research methodology or data analysis per se but is rather associated with obtaining specific single cells or cell populations from lesion biopsy. There are already some good examples of using fresh tissue biopsies for single-cell RNA sequencing [145,146,147], and the methodology can be transferred from these studies to endometriosis research. Also, there is great potential to use combinations of fluorescently labelled antibodies (e.g. CD10, CD9, and CD13 are previously shown to be markers of endometrial stromal and epithelial cells [148]) and fluorescence-activated cell sorting (FACS) for isolating single cells or cell populations from lesions. Furthermore, cell populations could be isolated from lesions using LCM technique, but as DNA and RNA quality obtained by this methodology varies a lot [149, 150], this method would probably not be the first choice. In addition to the lesion and endometrial single-cell studies, specific cells originating from blood or body fluids (like monocytes, NK cells, lymphocytes, granulocytes, endothelial cells, and progenitor cells) could offer new interesting perspectives to uncover pathologic changes related to the disease.
Although the advantages of single-cell ‘omicsʼ research in endometriosis are not utilised yet, it could be assumed that traditional transcriptome and epigenome studies will progress from the multicell level to single-cell level in the nearest future .
Systems Biology and Integrative ‘Omics’ Studies in Endometriosis
The number of ‘omicsʼ studies in endometriosis is rapidly increasing, and the massive amount of ‘omicsʼ data is becoming an immense challenge to the researchers. However, no single ‘omicsʼ analysis can fully resolve the complexity of the biology behind the disease development. Therefore, one of the main tasks for systems biology is the integration of large amounts of different types of data in order to understand the functional principles and dynamics of cellular systems and unravel the complexity of molecular networks in health and disease. Integration of multiple layers of ‘omicsʼ data requires network analysis and annotation of the involved pathways to capture meaningful information from genomics, epigenomics, transcriptomics, proteomics, metabolomics, and other ‘omicsʼ studies. After integration of single ‘omicsʼ data with computational engineering, modelling, and simulation and visualisation tools, new theoretical models can be created that provide comprehensive interpretation of integrated data (Fig. 3).
In endometriosis research, the first attempts to use systems biology and integrated data and knowledge from different ‘omicsʼ studies have been done [12, 13, 54,55,56, 67, 73, 151]. Most of these studies have combined epigenome data with gene expression data in order to reveal correlations between DNA methylation and gene expression. Borghese et al. found 35 genes that shared alterations both in methylation and gene expression levels [55]. The following study by Yamagata et al. observed a relationship between altered methylation and mRNA expression in 75 genes from cultured stromal cells from eutopic endometria and endometrioma, including steroidogenesis-related genes NRA1, STAR, STAR6, and HSD17B2 [56]. The results of this integrative analysis suggested that aberrant DNA methylation of the key steroidogenesis-related genes causes aberrant gene expression leading to the development of endometriosis. The transcriptome and epigenome data interaction modelling of stromal cells originating from eutopic and ectopic endometria of patients together with stromal cells from healthy endometria was performed by Dyson et al. [54]. They identified hundreds of genes, correspondent to thousands of differentially methylated CpGs that were aberrantly methylated and differentially expressed, among them the HOXA cluster, ESR2, NR5A1, and PGR genes. Furthermore, the authors classified the list of these genes by protein function and found that the only proteins that reached a statistical significance were transcription factors, such as GATA transcription factors and transcriptional coregulators of the GATA family. They proposed that GATA2 could regulate genes important for the decidualization process, whereas GATA6 promotes an endometriotic phenotype via regulation of steroidogenesis in endometriotic cells [54]. Sun et al. combined transcriptomic data of eutopic and ectopic endometria from mRNA and lncRNA microarray and found hundreds of lncRNAs that were co-expressed with thousands of mRNAs [67]. Hawkins et al. performed the first transcriptome-miRNome analysis of endometriomas and eutopic endometrium in order to narrow down genes that were functionally targeted by miRNAs [73]. The combined analysis revealed several potential biologically important pathways involved in cellular development, connective tissue development and function, and cellular growth and proliferation including TGFβ and mitogen-activated kinase 1 [73]. Completely novel approach to uncover regulatory changes in endometriosis was applied by Yang et al. [151]. They used integrative analysis of gene expression data from two different datasets combined with data from transcription factor (TF) gene regulatory interactions database (identified from ChiP-seq or ChIP-chip experiments available from ChEA database) to find endometriosis-associated TFs. This data was further combined with data from protein-protein interaction database (Interologous Interaction Database (I2D) and data from Ravasi et al. [152]) to create integrated regulatory network for understanding molecular mechanisms involved in endometriosis [151]. Authors identified a network of known TFs such as androgen receptor and estrogen receptor α and β participating in endometriosis pathogenesis. Also, several new TFs, such as FOXA2 and TFAP2C, were identified and validated in mRNA and protein level [151].
In conclusion, the ‘omics’ studies in endometriosis have provided the researchers with a substantial amount of data, but clearly, the full potential of this data is not entirely utilised. Although the amount of ‘omics’ datasets in publicly available repositories (Gene Expression Omnibus – GEO, etc.) is considerable, researchers still prefer using their own small datasets [153]. While the ‘omicsʼ studies in endometriosis are relatively small in size, future studies should use the advantage of the publicly available pre-existing data to raise the power, credibility, and reliability of the findings and also to find new biomarkers without the wet-lab costs .
Study Design in ‘Omics’ Studies in Endometriosis
The outcomes of ‘omicsʼ studies have also highlighted the shortcomings related to study design that may be the reason for the poor overlap between the results from different studies.
Large variability in tissue collection, processing, and storage methodology, poor description of patient phenotype data, small size of study groups, and differences between ‘omicsʼ platforms, together with data analysis and interpretation differences, are likely to result in bias and measurement error between studies. Standardised sample collection is crucial in biomarker discovery, as small differences in the processing and handling of biological samples can lead to the pre-analytical bias and might have a huge effect on analytical reliability and reproducibility [154]. The importance of harmonising standard operating procedures for collecting phenotypical data, and for sample collection, processing and storage is discussed in detail in the publications of the World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonisation Project (WERF EPHect) [155,156,157]. The main goals of the harmonised data and sample collection are not only to facilitate large-scale international collaborations and decrease cross-centre variability but also to collect information for future studies addressing specific research questions on different patient subtypes. The minimum and standard recommendations for surgical phenotype data collection include information about menstrual cycle, hormone treatment, history of previous endometriosis surgery, as well as any imaging findings before the procedure and the type and duration of the procedure; the extent, exact location, and colour of endometriotic lesions; and video/photo documentation of surgery [157]. In addition, the WERF EPHect provides precise recommendations for collecting, processing, and storing fluid biospecimens (plasma, serum, saliva, urine, endometrial/peritoneal fluid, and menstrual effluent) and tissue specimens (ectopic and eutopic endometrium, peritoneum, and myometrium) and for collecting biospecimen data, including information about the menstrual phase on the day of the sample collection, menstrual cycle regularity, timing of the next menstrual cycle, administration of any premedication or anaesthetics, and also information about the weight, height, and waist and hip circumference [155, 156]. It is strongly advised to follow the WERF EPHect published guidelines for designing new studies, as the well-characterised phenotypic datasets could be used to answer various current and future research questions in endometriosis.
One of the concerns related to the design of ‘omicsʼ studies is the definition of the endometriotic lesion . Majority of the studies have compared biopsied lesion samples and endometria; however, endometriotic lesions often contain only a small proportion of endometrial glands and stromal cells and a large proportion of surrounding tissue. Histological evaluation of biopsies is routinely used in everyday practice, and several studies have demonstrated that approximately 30–50% of surgical specimens removed during laparoscopy are not confirmed by histological assessment [158,159,160,161]. Even in the case of histologically confirmed biopsied endometriotic lesions, the proportion of endometrial tissue is variable. As different tissue types have their own molecular signature, the comparison between endometrial tissue and lesions that contain a mixed population of endometrial and peritoneal or ovarian tissue will not reveal disease-specific alterations but may rather reflect molecular signature of the surrounding tissue [76]. Therefore, future ‘omicsʼ studies of lesions should focus on pure cell populations or single-cell analyses instead of studying the entire endometriotic lesion to reveal the true molecular signature of endometriotic cells.
Selecting proper controls for biomarker discovery studies is one of the key issues that need to be resolved. Currently, endometriosis studies have applied different strategies for choosing controls and have included either fertile women undergoing laparoscopic sterilisation, women undergoing laparoscopy because of infertility, hospital-based controls (women with various indications for laparoscopic procedure), and also self-reported disease-free population-based controls [162]. Selecting controls for searching biomarkers from blood and body fluids is problematic, and there is never one ideal control group, as all above-mentioned options have their pros and cons. For example, laparoscopically confirmed endometriosis-free controls usually suffer from other diseases; a self-reported disease-free population-based control group could contain a number of undiagnosed cases and thereby dilute the disease risk factor effects.
The cost of the high-throughput technologies is relatively high, the sample sizes in ‘omicsʼ studies are rather small, and therefore the studies are underpowered to take into account the variance of individual measurements. Thus, it is strongly encouraged to maximise the sample sizes in ‘omicsʼ studies by collaborating with other workgroups and sharing either phenotypically well-described samples or data. In addition, there are other study type-specific standards for minimum information, including MIAME (Minimum Information about a Microarray Experiment), MIAPE (Minimum Information about a Proteomics Experiment), MIGS-MIMS (Minimum Information about a Genome/Metagenome Sequence), MIMIx (Minimum Information about a Molecular Interaction eXperiment), MINISEQE (Minimum Information about a high-throughput Nucleotide Sequencing Experiment), and CIMR (Core Information for Metabolomics Reporting). These guidelines must be followed to ensure the interpretability of the experimental results generated using ‘omics’ technologies [163, 164].
In conclusion, a good study design in endometriosis ‘omicsʼ studies includes setting an innovative study hypothesis, defining phenotypically well-described controls/cases, calculating study power that takes into account individual measurement variance, acceptable false-positive rate, and desired power of the used platform, collecting phenotypical data and biospecimens according to the guidelines of WERF EPHect, identifying risk factors and confounders, assessing sample quality and quantity, following the protocols for ‘omicsʼ technologies according to the specific guidelines (MIAME, MIAPE etc.), considering technical duplicates and statistical methods, describing databases for data analysis, validating results using alternative technologies, presenting data (e.g. GEO database), and addressing limitations/strengths of the study.
Conclusions and Future Perspectives
The ‘omicsʼ revolution in endometriosis research is ongoing, and around 120 studies applying the high-throughput ‘omicsʼ technologies have been published to date. Significant advances in ‘omicsʼ technologies have been made to discover potential biomarkers for endometriosis; however, most of the results have not been replicated in other studies, and the practical value of the proposed biomarkers is still limited. Though the genomic studies in endometriosis have not been very successful for finding potential biomarkers, the transcriptomic studies of endometriosis have provided some clues about the potential disease related pathways. Also, the results from proteome studies have been encouraging and hold a great promise for non-invasive biomarker discovery. Furthermore, great perspectives for future endometriosis biomarkers discovery are related to metabolomics and epigenomics as these fields are still poorly covered and harbour immense opportunities. Also, the advantages of single-cell transcriptome and epigenome studies should be carefully considered when planning future research.
Abbreviations
- aCGH:
-
Array-based comparative genomic hybridization
- CNV:
-
Copy number variation
- CpG:
-
C-phosphate-G-site
- 2D-DIGE:
-
Two-dimensional difference gel electrophoresis
- ESI-MS/MS:
-
Electrospray ionisation tandem mass spectrometry
- GWAS:
-
Genome wide association study
- H-NMR spectroscopy:
-
Proton nuclear magnetic resonance spectroscopy
- LCM:
-
Laser capture microdissection
- lncRNA:
-
Long non-coding RNA
- MALDI-TOF-MS:
-
Matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry
- miRNome:
-
Full spectrum of expressed miRNAs
- NMR:
-
Nuclear magnetic resonance spectroscopy
- SCNA:
-
Somatic copy number alteration
- SELDI-TOF-MS:
-
Surface-enhanced laser desorption/ionisation time-of-flight mass spectrometry
- SNP:
-
Single nucleotide polymorphism
- WERF EPHect:
-
World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonisation Project
References
Rogers PA, D’Hooghe TM, Fazleabas A, Giudice LC, Montgomery GW, Petraglia F, et al. Defining future directions for endometriosis research: workshop report from the 2011 World Congress of Endometriosis in Montpellier, France. Reprod Sci. 2013;20(5):483–99.
Nnoaham KE, Hummelshoj L, Webster P, d’Hooghe T, de Cicco NF, de Cicco NC, et al. Impact of endometriosis on quality of life and work productivity: a multicenter study across ten countries. Fertil Steril. 2011;96(2):366–73 e8.
Palmer SS, Barnhart KT. Biomarkers in reproductive medicine: the promise, and can it be fulfilled? Fertil Steril. 2013;99(4):954–62.
Gupta D, Hull ML, Fraser I, Miller L, Bossuyt PM, Johnson N, et al. Endometrial biomarkers for the non-invasive diagnosis of endometriosis. Cochrane Database Syst Rev. 2016;4:CD012165.
Nisenblat V, Bossuyt PM, Shaikh R, Farquhar C, Jordan V, Scheffers CS, et al. Blood biomarkers for the non-invasive diagnosis of endometriosis. Cochrane Database Syst Rev. 2016;5:CD012179.
Ozdemir V, Suarez-Kurtz G, Stenne R, Somogyi AA, Someya T, Kayaalp SO, et al. Risk assessment and communication tools for genotype associations with multifactorial phenotypes: the concept of “edge effect” and cultivating an ethical bridge between omics innovations and society. Omics. 2009;13(1):43–61.
Guo SW, Wu Y, Strawn E, Basir Z, Wang Y, Halverson G, et al. Genomic alterations in the endometrium may be a proximate cause for endometriosis. Eur J Obstet Gynecol Reprod Biol. 2004;116(1):89–99.
Li X, Zhang Y, Zhao L, Wang L, Wu Z, Mei Q, et al. Whole-exome sequencing of endometriosis identifies frequent alterations in genes involved in cell adhesion and chromatin-remodeling complexes. Hum Mol Genet. 2014;23(22):6008–21.
Izawa M, Taniguchi F, Terakawa N, Harada T. Epigenetic aberration of gene expression in endometriosis. Front Biosci (Elite Ed). 2013;5:900–10.
Naqvi H, Ilagan Y, Krikun G, Taylor HS. Altered genome-wide methylation in endometriosis. Reprod Sci. 2014;21(10):1237–43.
Saare M, Modhukur V, Suhorutshenko M, Rajashekar B, Rekker K, Soritsa D, et al. The influence of menstrual cycle and endometriosis on endometrial methylome. Clin Epigenetics. 2016;8:2.
Houshdaran S, Nezhat CR, Vo KC, Zelenko Z, Irwin JC, Giudice LC. Aberrant endometrial DNA methylome and associated gene expression in women with endometriosis. Biol Reprod. 2016;95(5):93.
Fassbender A, Verbeeck N, Bornigen D, Kyama CM, Bokor A, Vodolazkaia A, et al. Combined mRNA microarray and proteomic analysis of eutopic endometrium of women with and without endometriosis. Hum Reprod. 2012;27(7):2020–9.
Sherwin JR, Sharkey AM, Mihalyi A, Simsa P, Catalano RD, D’Hooghe TM. Global gene analysis of late secretory phase, eutopic endometrium does not provide the basis for a minimally invasive test of endometriosis. Hum Reprod. 2008;23(5):1063–8.
Burney RO, Talbi S, Hamilton AE, Vo KC, Nyegaard M, Nezhat CR, et al. Gene expression analysis of endometrium reveals progesterone resistance and candidate susceptibility genes in women with endometriosis. Endocrinology. 2007;148(8):3814–26.
Absenger Y, Hess-Stumpp H, Kreft B, Kratzschmar J, Haendler B, Schutze N, et al. Cyr61, a deregulated gene in endometriosis. Mol Hum Reprod. 2004;10(6):399–407.
Kao LC, Germeyer A, Tulac S, Lobo S, Yang JP, Taylor RN, et al. Expression profiling of endometrium from women with endometriosis reveals candidate genes for disease-based implantation failure and infertility. Endocrinology. 2003;144(7):2870–81.
Aghajanova L, Giudice LC. Molecular evidence for differences in endometrium in severe versus mild endometriosis. Reprod Sci. 2011;18(3):229–51.
Tamaresis JS, Irwin JC, Goldfien GA, Rabban JT, Burney RO, Nezhat C, et al. Molecular classification of endometriosis and disease stage using high-dimensional genomic data. Endocrinology. 2014;55(12):4986–99.
Zhou M, Fu J, Xiao L, Yang S, Song Y, Zhang X, et al. miR-196a overexpression activates the MEK/ERK signal and represses the progesterone receptor and decidualization in eutopic endometrium from women with endometriosis. Hum Reprod. 2016;31(11):2598–608.
Wang Y, Li Y, Yang Z, Liu K, Wang D. Genome-wide microarray analysis of long non-coding RNAs in eutopic secretory endometrium with endometriosis. Cell Physiol Biochem. 2015;37(6):2231–45.
Garcia-Velasco JA, Fassbender A, Ruiz-Alonso M, Blesa D, D’Hooghe T, Simon C. Is endometrial receptivity transcriptomics affected in women with endometriosis? A pilot study. Reprod Biomed Online. 2015;31(5):647–54.
Ahn SH, Khalaj K, Young SL, Lessey BA, Koti M, Tayade C. Immune-inflammation gene signatures in endometriosis patients. Fertil Steril. 2016;106(6):1420–31 e7.
Suryawanshi S, Huang X, Elishaev E, Budiu RA, Zhang L, Kim S, et al. Complement pathway is frequently altered in endometriosis and endometriosis-associated ovarian cancer. Clin Cancer Res. 2014;20(23):6163–74.
Matsuzaki S, Canis M, Vaurs-Barriere C, Boespflug-Tanguy O, Dastugue B, Mage G. DNA microarray analysis of gene expression in eutopic endometrium from patients with deep endometriosis using laser capture microdissection. Fertil Steril. 2005;84(Suppl 2):1180–90.
Giudice LC, Hamilton A, Lessey B. Transcriptomics. In Aplin JD, Fasleabas AT, Glasser SR, Giudice LC, editors. The endometrium: molecular, cellular and clinical perspectives. 2nd ed. Longon. Informa Healthcare; 2008. p. 193–222
Altmäe S, Aghajanova L. What do we know about endometrial receptivity in women with endometriosis? A molecular perspective. Reprod Biomed Online. 2015;31(5):581–3.
Burney RO, Hamilton AE, Aghajanova L, Vo KC, Nezhat CN, Lessey BA, et al. MicroRNA expression profiling of eutopic secretory endometrium in women with versus without endometriosis. Mol Hum Reprod. 2009;15(10):625–31.
Laudanski P, Charkiewicz R, Kuzmicki M, Szamatowicz J, Charkiewicz A, Niklinski J. MicroRNAs expression profiling of eutopic proliferative endometrium in women with ovarian endometriosis. Reprod Biol Endocrinol. 2013;11:78.
Laudanski P, Charkiewicz R, Tolwinska A, Szamatowicz J, Charkiewicz A, Niklinski J. Profiling of selected microRNAs in proliferative eutopic endometrium of women with ovarian endometriosis. Biomed Res Int. 2015;2015:760698.
Braza-Boils A, Mari-Alexandre J, Gilabert J, Sanchez-Izquierdo D, Espana F, Estelles A, et al. MicroRNA expression profile in endometriosis: its relation to angiogenesis and fibrinolytic factors. Hum Reprod. 2014;29(5):978–88.
Shi XY, Gu L, Chen J, Guo XR, Shi YL. Downregulation of miR-183 inhibits apoptosis and enhances the invasive potential of endometrial stromal cells in endometriosis. Int J Mol Med. 2014;33(1):59–67.
Cox J, Mann M. Quantitative, high-resolution proteomics for data-driven systems biology. Annu Rev Biochem. 2011;80:273–99.
Altmäe S, Esteban FJ, Stavreus-Evers A, Simon C, Giudice L, Lessey BA, et al. Guidelines for the design, analysis and interpretation of ‘omics’ data: focus on human endometrium. Hum Reprod Update. 2014;20(1):12–28.
Fassbender A, Simsa P, Kyama CM, Waelkens E, Mihalyi A, Meuleman C, et al. TRIzol treatment of secretory phase endometrium allows combined proteomic and mRNA microarray analysis of the same sample in women with and without endometriosis. Reprod Biol Endocrinol. 2010;8:123.
Ning K, Fermin D, Nesvizhskii AI. Comparative analysis of different label-free mass spectrometry based protein abundance estimates and their correlation with RNA-Seq gene expression data. J Proteome Res. 2012;11(4):2261–71.
Zhang H, Niu Y, Feng J, Guo H, Ye X, Cui H. Use of proteomic analysis of endometriosis to identify different protein expression in patients with endometriosis versus normal controls. Fertil Steril. 2006;86(2):274–82.
Stephens AN, Hannan NJ, Rainczuk A, Meehan KL, Chen J, Nicholls PK, et al. Post-translational modifications and protein-specific isoforms in endometriosis revealed by 2D DIGE. J Proteome Res. 2010;9(5):2438–49.
Rai P, Kota V, Deendayal M, Shivaji S. Differential proteome profiling of eutopic endometrium from women with endometriosis to understand etiology of endometriosis. J Proteome Res. 2010;9(9):4407–19.
Fowler PA, Tattum J, Bhattacharya S, Klonisch T, Hombach-Klonisch S, Gazvani R, et al. An investigation of the effects of endometriosis on the proteome of human eutopic endometrium: a heterogeneous tissue with a complex disease. Proteomics. 2007;7(1):130–42.
Ten Have S, Fraser I, Markham R, Lam A, Matsumoto I. Proteomic analysis of protein expression in the eutopic endometrium of women with endometriosis. Proteomics Clin Appl. 2007;1(10):1243–51.
Ding X, Wang L, Ren Y, Zheng W. Detection of mitochondrial biomarkers in eutopic endometria of endometriosis using surface-enhanced laser desorption/ionization time-of-flight mass spectrometry. Fertil Steril. 2010;94(7):2528–30.
Kyama CM, Mihalyi A, Gevaert O, Waelkens E, Simsa P, Van de Plas R, et al. Evaluation of endometrial biomarkers for semi-invasive diagnosis of endometriosis. Fertil Steril. 2011;95(4):1338–43 e1–3.
Wang L, Zheng W, Ding XY, Yu JK, Jiang WZ, Zhang SZ. Identification biomarkers of eutopic endometrium in endometriosis using artificial neural networks and protein fingerprinting. Fertil Steril. 2010;93(7):2460–2.
Siva AB, Srivastava P, Shivaji S. Understanding the pathogenesis of endometriosis through proteomics: recent advances and future prospects. Proteomics Clin Appl. 2014;8(1–2):86–98.
Yang W, Zhang Y, Fu F, Li R. High-resolution array-comparative genomic hybridization profiling reveals 20q13.33 alterations associated with ovarian endometriosis. Gynecol Endocrinol. 2013;29(6):603–7.
Silveira CG, Abrao MS, Dias JA Jr, Coudry RA, Soares FA, Drigo SA, et al. Common chromosomal imbalances and stemness-related protein expression markers in endometriotic lesions from different anatomical sites: the potential role of stem cells. Hum Reprod. 2012;27(11):3187–97.
Gogusev J, Bouquet de Joliniere J, Telvi L, Doussau M, du Manoir S, Stojkoski A, et al. Detection of DNA copy number changes in human endometriosis by comparative genomic hybridization. Hum Genet. 1999;105(5):444–51.
Wu Y, Strawn E, Basir Z, Wang Y, Halverson G, Jailwala P, et al. Genomic alterations in ectopic and eutopic endometria of women with endometriosis. Gynecol Obstet Invest. 2006;62(3):148–59.
Veiga-Castelli LC, Silva JC, Meola J, Ferriani RA, Yoshimoto M, Santos SA, et al. Genomic alterations detected by comparative genomic hybridization in ovarian endometriomas. Braz J Med Biol Res. 2010;43(8):799–805.
Zafrakas M, Tarlatzis BC, Streichert T, Pournaropoulos F, Wolfle U, Smeets SJ, et al. Genome-wide microarray gene expression, array-CGH analysis, and telomerase activity in advanced ovarian endometriosis: a high degree of differentiation rather than malignant potential. Int J Mol Med. 2008;21(3):335–44.
Saare M, Soritsa D, Vaidla K, Palta P, Remm M, Laan M, et al. No evidence of somatic DNA copy number alterations in eutopic and ectopic endometrial tissue in endometriosis. Hum Reprod. 2012;27(6):1857–64.
Karhu R, Kahkonen M, Kuukasjarvi T, Pennanen S, Tirkkonen M, Kallioniemi O. Quality control of CGH: impact of metaphase chromosomes and the dynamic range of hybridization. Cytometry. 1997;28(3):198–205.
Dyson MT, Roqueiro D, Monsivais D, Ercan CM, Pavone ME, Brooks DC, et al. Genome-wide DNA methylation analysis predicts an epigenetic switch for GATA factor expression in endometriosis. PLoS Genet. 2014;10(3):e1004158.
Borghese B, Barbaux S, Mondon F, Santulli P, Pierre G, Vinci G, et al. Research resource: genome-wide profiling of methylated promoters in endometriosis reveals a subtelomeric location of hypermethylation. Mol Endocrinol. 2010;24(9):1872–85.
Yamagata Y, Nishino K, Takaki E, Sato S, Maekawa R, Nakai A, et al. Genome-wide DNA methylation profiling in cultured eutopic and ectopic endometrial stromal cells. PLoS One. 2014;9(1):e83612.
Arimoto T, Katagiri T, Oda K, Tsunoda T, Yasugi T, Osuga Y, et al. Genome-wide cDNA microarray analysis of gene-expression profiles involved in ovarian endometriosis. Int J Oncol. 2003;22(3):551–60.
Hu WP, Tay SK, Zhao Y. Endometriosis-specific genes identified by real-time reverse transcription-polymerase chain reaction expression profiling of endometriosis versus autologous uterine endometrium. J Clin Endocrinol Metab. 2006;91(1):228–38.
Wu Y, Kajdacsy-Balla A, Strawn E, Basir Z, Halverson G, Jailwala P, et al. Transcriptional characterizations of differences between eutopic and ectopic endometrium. Endocrinology. 2006;147(1):232–46.
Matsuzaki S, Canis M, Vaurs-Barriere C, Pouly JL, Boespflug-Tanguy O, Penault-Llorca F, et al. DNA microarray analysis of gene expression profiles in deep endometriosis using laser capture microdissection. Mol Hum Reprod. 2004;10(10):719–28.
Konno R, Yamada-Okabe H, Fujiwara H, Uchiide I, Shibahara H, Ohwada M, et al. Role of immunoreactions and mast cells in pathogenesis of human endometriosis—morphologic study and gene expression analysis. Hum Cell. 2003;16(3):141–9.
Lebovic DI, Baldocchi RA, Mueller MD, Taylor RN. Altered expression of a cell-cycle suppressor gene, Tob-1, in endometriotic cells by cDNA array analyses. Fertil Steril. 2002;78(4):849–54.
Eyster KM, Klinkova O, Kennedy V, Hansen KA. Whole genome deoxyribonucleic acid microarray analysis of gene expression in ectopic versus eutopic endometrium. Fertil Steril. 2007;88(6):1505–33.
Eyster KM, Boles AL, Brannian JD, Hansen KA. DNA microarray analysis of gene expression markers of endometriosis. Fertil Steril. 2002;77(1):38–42.
Mettler L, Salmassi A, Schollmeyer T, Schmutzler AG, Pungel F, Jonat W. Comparison of c-DNA microarray analysis of gene expression between eutopic endometrium and ectopic endometrium (endometriosis). J Assist Reprod Genet. 2007;24(6):249–58.
Khan MA, Sengupta J, Mittal S, Ghosh D. Genome-wide expressions in autologous eutopic and ectopic endometrium of fertile women with endometriosis. Reprod Biol Endocrinol. 2012;10:84.
Sun PR, Jia SZ, Lin H, Leng JH, Lang JH. Genome-wide profiling of long noncoding ribonucleic acid expression patterns in ovarian endometriosis by microarray. Fertil Steril. 2014;101(4):1038–46 e7.
Crispi S, Piccolo MT, D'Avino A, Donizetti A, Viceconte R, Spyrou M, et al. Transcriptional profiling of endometriosis tissues identifies genes related to organogenesis defects. J Cell Physiol. 2013;228(9):1927–34.
Monsivais D, Dyson MT, Yin P, Navarro A, Coon JS, Pavone ME, et al. Estrogen receptor beta regulates endometriotic cell survival through serum and glucocorticoid-regulated kinase activation. Fertil Steril. 2016;105(5):1266–73.
Borghese B, Mondon F, Noel JC, Fayt I, Mignot TM, Vaiman D, et al. Gene expression profile for ectopic versus eutopic endometrium provides new insights into endometriosis oncogenic potential. Mol Endocrinol. 2008;22(11):2557–62.
Hever A, Roth RB, Hevezi P, Marin ME, Acosta JA, Acosta H, et al. Human endometriosis is associated with plasma cells and overexpression of B lymphocyte stimulator. Proc Natl Acad Sci U S A. 2007;104(30):12451–6.
Monsivais D, Bray JD, Su E, Pavone ME, Dyson MT, Navarro A, et al. Activated glucocorticoid and eicosanoid pathways in endometriosis. Fertil Steril. 2012;98(1):117–25.
Hawkins SM, Creighton CJ, Han DY, Zariff A, Anderson ML, Gunaratne PH, et al. Functional microRNA involved in endometriosis. Mol Endocrinol. 2011;25(5):821–32.
Filigheddu N, Gregnanin I, Porporato PE, Surico D, Perego B, Galli L, et al. Differential expression of microRNAs between eutopic and ectopic endometrium in ovarian endometriosis. J Biomed Biotechnol. 2010;2010:369549.
Ohlsson Teague EM, Van der Hoek KH, Van der Hoek MB, Perry N, Wagaarachchi P, Robertson SA, et al. MicroRNA-regulated pathways associated with endometriosis. Mol Endocrinol. 2009;23(2):265–75.
Saare M, Rekker K, Laisk-Podar T, Soritsa D, Roost AM, Simm J, et al. High-throughput sequencing approach uncovers the miRNome of peritoneal endometriotic lesions and adjacent healthy tissues. PLoS One. 2014;9(11):e112630.
Yang RQ, Teng H, Xu XH, Liu SY, Wang YH, Guo FJ, et al. Microarray analysis of microRNA deregulation and angiogenesis-related proteins in endometriosis. Genet Mol Res. 2016;15(2).
Abe W, Nasu K, Nakada C, Kawano Y, Moriyama M, Narahara H. miR-196b targets c-myc and Bcl-2 expression, inhibits proliferation and induces apoptosis in endometriotic stromal cells. Hum Reprod. 2013;28(3):750–61.
Hull ML, Nisenblat V. Tissue and circulating microRNA influence reproductive function in endometrial disease. Reprod Biomed Online. 2013;27(5):515–29.
Chehna-Patel N, Sachdeva G, Gajbhiye R, Warty N, Khole V. “Spot”-ting differences between the ectopic and eutopic endometrium of endometriosis patients. Fertil Steril. 2010;94(6):1964–71.
Vehmas AP, Muth-Pawlak D, Huhtinen K, Saloniemi-Heinonen T, Jaakkola K, Laajala TD, et al. Ovarian endometriosis signatures established through discovery and directed mass spectrometry analysis. J Proteome Res. 2014;13(11):4983–94.
Kasvandik S, Samuel K, Peters M, Eimre M, Peet N, Roost AM, et al. Deep quantitative proteomics reveals extensive metabolic reprogramming and cancer-like changes of ectopic endometriotic stromal cells. J Proteome Res. 2016;15(2):572–84.
Marianowski P, Szymusik I, Malejczyk J, Hibner M, Wielgos M. Proteomic analysis of eutopic and ectopic endometriotic tissues based on isobaric peptide tags for relative and absolute quantification (iTRAQ) method. Neuro Endocrinol Lett. 2013;34(7):717–21.
Kyama CM, T’Jampens D, Mihalyi A, Simsa P, Debrock S, Waelkens E, et al. ProteinChip technology is a useful method in the pathogenesis and diagnosis of endometriosis: a preliminary study. Fertil Steril. 2006;86(1):203–9.
May KE, Conduit-Hulbert SA, Villar J, Kirtley S, Kennedy SH, Becker CM. Peripheral biomarkers of endometriosis: a systematic review. Hum Reprod Update. 2010;16(6):651–74.
Vodolazkaia A, El-Aalamat Y, Popovic D, Mihalyi A, Bossuyt X, Kyama CM, et al. Evaluation of a panel of 28 biomarkers for the non-invasive diagnosis of endometriosis. Hum Reprod. 2012;27(9):2698–711.
Uno S, Zembutsu H, Hirasawa A, Takahashi A, Kubo M, Akahane T, et al. A genome-wide association study identifies genetic variants in the CDKN2BAS locus associated with endometriosis in Japanese. Nat Genet. 2010;42(8):707–10.
Adachi S, Tajima A, Quan J, Haino K, Yoshihara K, Masuzaki H, et al. Meta-analysis of genome-wide association scans for genetic susceptibility to endometriosis in Japanese population. J Hum Genet. 2010;55(12):816–21.
Painter JN, Anderson CA, Nyholt DR, Macgregor S, Lin J, Lee SH, et al. Genome-wide association study identifies a locus at 7p15.2 associated with endometriosis. Nat Genet. 2011;43(1):51–4.
Albertsen HM, Chettier R, Farrington P, Ward K. Genome-wide association study link novel loci to endometriosis. PLoS One. 2013;8(3):e58257.
Pagliardini L, Gentilini D, Vigano P, Panina-Bordignon P, Busacca M, Candiani M, et al. An Italian association study and meta-analysis with previous GWAS confirm WNT4, CDKN2BAS and FN1 as the first identified susceptibility loci for endometriosis. J Med Genet. 2013;50(1):43–6.
Sundqvist J, Xu H, Vodolazkaia A, Fassbender A, Kyama C, Bokor A, et al. Replication of endometriosis-associated single-nucleotide polymorphisms from genome-wide association studies in a Caucasian population. Hum Reprod. 2013;28(3):835–9.
Nyholt DR, Low SK, Anderson CA, Painter JN, Uno S, Morris AP, et al. Genome-wide association meta-analysis identifies new endometriosis risk loci. Nat Genet. 2012;44(12):1355–9.
Sapkota Y, Fassbender A, Bowdler L, Fung JN, Peterse D, O D, et al. Independent replication and meta-analysis for endometriosis risk loci. Twin Res Hum Genet. 2015:1–8.
Steinthorsdottir V, Thorleifsson G, Aradottir K, Feenstra B, Sigurdsson A, Stefansdottir L, et al. Common variants upstream of KDR encoding VEGFR2 and in TTC39B associate with endometriosis. Nat Commun. 2016;7:12350.
Rahmioglu N, Nyholt DR, Morris AP, Missmer SA, Montgomery GW, Zondervan KT. Genetic variants underlying risk of endometriosis: insights from meta-analysis of eight genome-wide association and replication datasets. Hum Reprod Update. 2014;20(5):702–16.
Visscher PM, Brown MA, McCarthy MI, Yang J. Five years of GWAS discovery. Am J Hum Genet. 2012;90(1):7–24.
Teo SM, Ku CS, Naidoo N, Hall P, Chia KS, Salim A, et al. A population-based study of copy number variants and regions of homozygosity in healthy Swedish individuals. J Hum Genet. 2011;56(7):524–33.
Sebat J, Lakshmi B, Troge J, Alexander J, Young J, Lundin P, et al. Large-scale copy number polymorphism in the human genome. Science. 2004;305(5683):525–8.
Pinto D, Marshall C, Feuk L, Scherer SW. Copy-number variation in control population cohorts. Hum Mol Genet. 2007;16(Spec no. 2):R168–73.
Chettier R, Ward K, Albertsen HM. Endometriosis is associated with rare copy number variants. PLoS One. 2014;9(8):e103968.
Mafra F, Mazzotti D, Pellegrino R, Bianco B, Barbosa CP, Hakonarson H, et al. Copy number variation analysis reveals additional variants contributing to endometriosis development. J Assist Reprod Genet. 2017;34(1):117–24.
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18(10):997–1006.
Cortez MA, Calin GA. MicroRNA identification in plasma and serum: a new tool to diagnose and monitor diseases. Expert Opin Biol Ther. 2009;9(6):703–11.
Suryawanshi S, Vlad AM, Lin HM, Mantia-Smaldone G, Laskey R, Lee M, et al. Plasma microRNAs as novel biomarkers for endometriosis and endometriosis-associated ovarian cancer. Clin Cancer Res. 2013;19(5):1213–24.
Hsu CY, Hsieh TH, Tsai CF, Tsai HP, Chen HS, Chang Y, et al. miRNA-199a-5p regulates VEGFA in endometrial mesenchymal stem cells and contributes to the pathogenesis of endometriosis. J Pathol. 2014;232(3):330–43.
Jia SZ, Yang Y, Lang J, Sun P, Leng J. Plasma miR-17-5p, miR-20a and miR-22 are down-regulated in women with endometriosis. Hum Reprod. 2013;28(2):322–30.
Wang WT, Zhao YN, Han BW, Hong SJ, Chen YQ. Circulating microRNAs identified in a genome-wide serum microRNA expression analysis as noninvasive biomarkers for endometriosis. J Clin Endocrinol Metab. 2013;98(1):281–9.
Wang L, Huang W, Ren C, Zhao M, Jiang X, Fang X, et al. Analysis of serum microRNA Profile by Solexa sequencing in women with endometriosis. Reprod Sci. 2016;23(10):1359–70.
Cosar E, Mamillapalli R, Ersoy GS, Cho S, Seifer B, Taylor HS. Serum microRNAs as diagnostic markers of endometriosis: a comprehensive array-based analysis. Fertil Steril. 2016;106(2):402–9.
Fassbender A, Waelkens E, Verbeeck N, Kyama CM, Bokor A, Vodolazkaia A, et al. Proteomics analysis of plasma for early diagnosis of endometriosis. Obstet Gynecol. 2012;119(2 Pt 1):276–85.
Liu H, Lang J, Zhou Q, Shan D, Li Q. Detection of endometriosis with the use of plasma protein profiling by surface-enhanced laser desorption/ionization time-of-flight mass spectrometry. Fertil Steril. 2007;87(4):988–90.
Zheng N, Pan C, Liu W. New serum biomarkers for detection of endometriosis using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Int Med Res. 2011;39(4):1184–92.
Long X, Jiang P, Zhou L, Zhang W. Evaluation of novel serum biomarkers and the proteomic differences of endometriosis and adenomyosis using MALDI-TOF-MS. Arch Gynecol Obstet. 2013;288(1):201–5.
Jing J, Qiao Y, Suginami H, Taniguchi F, Shi H, Wang X. Two novel serum biomarkers for endometriosis screened by surface-enhanced laser desorption/ionization time-of-flight mass spectrometry and their change after laparoscopic removal of endometriosis. Fertil Steril. 2009;92(4):1221–7.
Nabeta M, Abe Y, Haraguchi R, Kito K, Kusanagi Y, Ito M. Serum anti-PDIK1L autoantibody as a novel marker for endometriosis. Fertil Steril. 2010;94(7):2552–7.
Nabeta M, Abe Y, Kagawa L, Haraguchi R, Kito K, Ueda N, et al. Identification of anti-alpha-enolase autoantibody as a novel serum marker for endometriosis. Proteomics Clin Appl. 2009;3(10):1201–10.
Nabeta M, Abe Y, Takaoka Y, Kusanagi Y, Ito M. Identification of anti-syntaxin 5 autoantibody as a novel serum marker of endometriosis. J Reprod Immunol. 2011;91(1–2):48–55.
Gajbhiye R, Sonawani A, Khan S, Suryawanshi A, Kadam S, Warty N, et al. Identification and validation of novel serum markers for early diagnosis of endometriosis. Hum Reprod. 2012;27(2):408–17.
Faserl K, Golderer G, Kremser L, Lindner H, Sarg B, Wildt L, et al. Polymorphism in vitamin D-binding protein as a genetic risk factor in the pathogenesis of endometriosis. J Clin Endocrinol Metab. 2011;96(1):E233–41.
Seeber B, Sammel MD, Fan X, Gerton GL, Shaunik A, Chittams J, et al. Proteomic analysis of serum yields six candidate proteins that are differentially regulated in a subset of women with endometriosis. Fertil Steril. 2010;93(7):2137–44.
Zhang H, Feng J, Chang XH, Li ZX, Wu XY, Cui H. Effect of surface-enhanced laser desorption/ionization time-of-flight mass spectrometry on identifying biomarkers of endometriosis. Chin Med J (Engl). 2009;122(4):373–6.
Wang L, Zheng W, Yu JK, Jiang WZ, Mu L, Zhang SZ. Artificial neural networks combined with surface-enhanced laser desorption/ionization mass spectra distinguish endometriosis from healthy population. Fertil Steril. 2007;88(6):1700–2.
Dutta M, Subramani E, Taunk K, Gajbhiye A, Seal S, Pendharkar N, et al. Investigation of serum proteome alterations in human endometriosis. J Proteomics. 2015;114:182–96.
El-Kasti MM, Wright C, Fye HK, Roseman F, Kessler BM, Becker CM. Urinary peptide profiling identifies a panel of putative biomarkers for diagnosing and staging endometriosis. Fertil Steril. 2011;95(4):1261–6.
Tokushige N, Markham R, Crossett B, Ahn SB, Nelaturi VL, Khan A, et al. Discovery of a novel biomarker in the urine in women with endometriosis. Fertil Steril. 2011;95(1):46–9.
Cho S, Choi YS, Yim SY, Yang HI, Jeon YE, Lee KE, et al. Urinary vitamin D-binding protein is elevated in patients with endometriosis. Hum Reprod. 2012;27(2):515–22.
Williams KE, Miroshnychenko O, Johansen EB, Niles RK, Sundaram R, Kannan K, et al. urine, peritoneal fluid and omental fat proteomes of reproductive age women: endometriosis-related changes and associations with endocrine disrupting chemicals. J Proteomics. 2014;13:194–205.
Ametzazurra A, Matorras R, Garcia-Velasco JA, Prieto B, Simon L, Martinez A, et al. Endometrial fluid is a specific and non-invasive biological sample for protein biomarker identification in endometriosis. Hum Reprod. 2009;24(4):954–65.
Hwang JH, Oh JJ, Wang T, Jin YC, Lee JS, Choi JR, et al. Identification of biomarkers for endometriosis in eutopic endometrial cells from patients with endometriosis using a proteomics approach. Mol Med Rep. 2013;8(1):183–8.
Siciliano RA, Mazzeo MF, Spada V, Facchiano A, d’Acierno A, Stocchero M, et al. Rapid peptidomic profiling of peritoneal fluid by MALDI-TOF mass spectrometry for the identification of biomarkers of endometriosis. Gynecol Endocrinol. 2014;30(12):872–6.
Wang L, Ding XY, Yu JK, Zhang SZ, Zheng W. Biomarkers of peritoneal fluid in endometriosis identified by surface-enhanced laser desorption/ionization time-of-flight. Clin Exp Obstet Gynecol. 2014;41(1):72–4.
Ferrero S, Gillott DJ, Remorgida V, Anserini P, Leung KY, Ragni N, et al. Proteomic analysis of peritoneal fluid in women with endometriosis. J Proteome Res. 2007;6(9):3402–11.
Ferrero S, Gillott DJ, Remorgida V, Anserini P, Ragni N, Grudzinskas JG. Peritoneal fluid proteome in women with different ASRM stages of endometriosis. Gynecol Endocrinol. 2008;24(8):433–41.
Ferrero S, Gillott DJ, Remorgida V, Anserini P, Ragni N, Grudzinskas JG. Proteomic analysis of peritoneal fluid in fertile and infertile women with endometriosis. J Reprod Med. 2009;54(1):32–40.
Wolfler MM, Meinhold-Heerlein IM, Sohngen L, Rath W, Knuchel R, Neulen J, et al. Two-dimensional gel electrophoresis in peritoneal fluid samples identifies differential protein regulation in patients suffering from peritoneal or ovarian endometriosis. Fertil Steril. 2011;95(8):2764–8.
Williams KE, Miroshnychenko O, Johansen EB, Niles RK, Sundaram R, Kannan K, et al. Urine, peritoneal fluid and omental fat proteomes of reproductive age women: endometriosis-related changes and associations with endocrine disrupting chemicals. J Proteomics. 2015;113:194–205.
Rizner TL. Noninvasive biomarkers of endometriosis: myth or reality? Expert Rev Mol Diagn. 2014;14(3):365–85.
Beger RD. A review of applications of metabolomics in cancer. Metabolites. 2013;3(3):552–74.
Jana SK, Dutta M, Joshi M, Srivastava S, Chakravarty B, Chaudhury K. 1H NMR based targeted metabolite profiling for understanding the complex relationship connecting oxidative stress with endometriosis. Biomed Res Int. 2013;2013:329058.
Dutta M, Joshi M, Srivastava S, Lodh I, Chakravarty B, Chaudhury K. A metabonomics approach as a means for identification of potential biomarkers for early diagnosis of endometriosis. Mol Biosyst. 2012;8(12):3281–7.
Vouk K, Hevir N, Ribic-Pucelj M, Haarpaintner G, Scherb H, Osredkar J, et al. Discovery of phosphatidylcholines and sphingomyelins as biomarkers for ovarian endometriosis. Hum Reprod. 2012;27(10):2955–65.
Ghazi N, Arjmand M, Akbari Z, Mellati AO, Saheb-Kashaf H, Zamani Z. (1)H NMR- based metabolomics approaches as non- invasive tools for diagnosis of endometriosis. Int J Reprod Biomed (Yazd). 2016;14(1):1–8.
Saliba AE, Westermann AJ, Gorski SA, Vogel J. Single-cell RNA-seq: advances and future challenges. Nucleic Acids Res. 2014;42(14):8845–60.
Usoskin D, Furlan A, Islam S, Abdo H, Lonnerberg P, Lou D, et al. Unbiased classification of sensory neuron types by large-scale single-cell RNA sequencing. Nat Neurosci. 2015;18(1):145–53.
Patel AP, Tirosh I, Trombetta JJ, Shalek AK, Gillespie SM, Wakimoto H, et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science. 2014;344(6190):1396–401.
Krjutskov K, Katayama S, Saare M, Vera-Rodriguez M, Lubenets D, Samuel K, et al. Single-cell transcriptome analysis of endometrial tissue. Hum Reprod. 2016;31(4):844–53.
Kato K, Yoshimoto M, Adachi S, Yamayoshi A, Arima T, Asanoma K, et al. Characterization of side-population cells in human normal endometrium. Hum Reprod. 2007;22(5):1214–23.
Cummings M, McGinley CV, Wilkinson N, Field SL, Duffy SR, Orsi NM. A robust RNA integrity-preserving staining protocol for laser capture microdissection of endometrial cancer tissue. Anal Biochem. 2011;416(1):123–5.
Aaltonen KE, Ebbesson A, Wigerup C, Hedenfalk I. Laser capture microdissection (LCM) and whole genome amplification (WGA) of DNA from normal breast tissue—optimization for genome wide array analyses. BMC Res Notes. 2011;4:69.
Yang H, Kang K, Cheng C, Mamillapalli R, Taylor HS. Integrative analysis reveals regulatory programs in endometriosis. Reprod Sci. 2015;22(9):1060–72.
Ravasi T, Suzuki H, Cannistraci CV, Katayama S, Bajic VB, Tan K, et al. An atlas of combinatorial transcriptional regulation in mouse and man. Cell. 2010;140(5):744–52.
Baker M. Gene data to hit milestone. Nature. 2012;487(7407):282–3.
Jackson DH, Banks RE. Banking of clinical samples for proteomic biomarker studies: a consideration of logistical issues with a focus on pre-analytical variation. Proteomics Clin Appl. 2010;4(3):250–70.
Fassbender A, Rahmioglu N, Vitonis AF, Vigano P, Giudice LC, D'Hooghe TM, et al. World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonisation Project: IV. Tissue collection, processing, and storage in endometriosis research. Fertil Steril. 2014;10(2):1244–53.
Rahmioglu N, Fassbender A, Vitonis AF, Tworoger SS, Hummelshoj L, D’Hooghe TM, et al. World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonization Project: III. Fluid biospecimen collection, processing, and storage in endometriosis research. Fertil Steril. 2014;102(5):1233–43.
Becker CM, Laufer MR, Stratton P, Hummelshoj L, Missmer SA, Zondervan KT, et al. World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonisation Project: I. Surgical phenotype data collection in endometriosis research. Fertil Steril. 2014;102(5):1211–22.
Moen MH, Halvorsen TB. Histologic confirmation of endometriosis in different peritoneal lesions. Acta Obstet Gynecol Scand. 1992;71(5):337–42.
Stegmann BJ, Sinaii N, Liu S, Segars J, Merino M, Nieman LK, et al. Using location, color, size, and depth to characterize and identify endometriosis lesions in a cohort of 133 women. Fertil Steril. 2008;89(6):1632–6.
Stratton P, Winkel C, Premkumar A, Chow C, Wilson J, Hearns-Stokes R, et al. Diagnostic accuracy of laparoscopy, magnetic resonance imaging, and histopathologic examination for the detection of endometriosis. Fertil Steril. 2003;79(5):1078–85.
Fernando S, Soh PQ, Cooper M, Evans S, Reid G, Tsaltas J, et al. Reliability of visual diagnosis of endometriosis. J Minim Invasive Gynecol. 2013;20(6):783–9.
Zondervan KT, Cardon LR, Kennedy SH. What makes a good case-control study? Design issues for complex traits such as endometriosis. Hum Reprod. 2002;17(6):1415–23.
Brazma A, Hingamp P, Quackenbush J, Sherlock G, Spellman P, Stoeckert C, et al. Minimum information about a microarray experiment (MIAME)-toward standards for microarray data. Nat Genet. 2001;29(4):365–71.
Chervitz SA, Deutsch EW, Field D, Parkinson H, Quackenbush J, Rocca-Serra P, et al. Data standards for Omics data: the basis of data sharing and reuse. Methods Mol Biol. 2011;719:31–69.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Saare, M., Peters, M., Aints, A., Laisk-Podar, T., Salumets, A., Altmäe, S. (2017). OMICs Studies and Endometriosis Biomarker Identification. In: D'Hooghe, T. (eds) Biomarkers for Endometriosis. Springer, Cham. https://doi.org/10.1007/978-3-319-59856-7_12
Download citation
DOI: https://doi.org/10.1007/978-3-319-59856-7_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-59854-3
Online ISBN: 978-3-319-59856-7
eBook Packages: MedicineMedicine (R0)