Abstract
As an integrated part of the complex array of post-translational modifications on histone, methylation mainly occurs on histone lysine and arginine residues and plays pivotal roles in the regulation of chromatin organization and function. Histone methylation is catalyzed by different groups of methyltransferases while methylation marks on different residues as well as different methylation states on the same residue serve as docking sites to recruit a variety of binding proteins harboring specific recognition domains. These methyl-histone binding proteins further recruit additional chromatin modifiers and other proteins to transduce methylation signals into diverse functional outcomes. Here we summarize histone methyltransferases and discuss their specificities and regulations for different methylation reactions. We also discuss specific methyl-histone recognition by different families of binding proteins at multiple molecular layers. Given that the disruption of histone methylation and recognition has been associated with altered gene function in various diseases and human malignancy, understanding the regulation of histone methylation and recognition will also provide molecular basics for therapeutics.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Epigenetics
- Chromatin
- Histone methylation
- Histone methyltransferase
- Methyl-histone binding domain
- Structure and function
1 Introduction
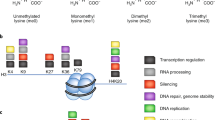
In eukaryotes, about 147 bp DNA wraps around a histone octamer (assembled from H3-H4 tetramer and two H2A-H2B dimers) to form the nucleosome core-particle. This structure is stabilized by histone H1 and can be further folded into higher-order chromatin structures [1]. It has been demonstrated that histone molecules are subject to diverse modifications, including methylation, acetylation, phosphorylation, ubiquitination and many others, which constitute a unique “code” for the regulation of chromatin function and dynamics [2, 3]. Histone methylation was discovered in 1964 [4, 5], however, its regulation and functional significance were revealed in the past 15 years. Through reactions catalyzed by different families of enzymes, histone lysine residues can be mono-, di- or tri-methylated [6] whereas arginines can be mono- or di-methylated (symmetrically or asymmetrically) [7] (Fig. 1). While most methylation occurs on the flexible histone N-terminal tails, several methylations were also detected within the globular domain. It has been well-documented that histone methylation at different residues as well as different methylation states within the same residue can confer a variety of biological functions. Table 1 summarized validated histone methylation sites together with their catalyzing enzymes and the functional outcomes. In this Chapter, we discuss several major histone methyltransferases (HMTases) and molecular mechanisms underlying methylation reactions. In most cases, the addition of methyl groups to histones does not directly affect chromatin structure. It is believed that diverse functions of different histone methylations are mediated through binding proteins harboring specific motifs. Therefore, we also discuss different methyl-histone binding domains and their recognition mechanisms.
Lysine and arginine methylation. (a) Catalyzed by different histone KMTs, lysine residues can be mono-, di-, and tri-methylated. (b) Arginine residue in histones can be mono-methylated by type I, II, III PRMTs. Type I and II PRMTs can further introduce asymmetric and symmetric di-methylation, respectively
2 Histone Methyltransferase
Methylation is one of the most common protein modifications on multiple amino acids, which is catalyzed by protein methyltransferases (MTases) using S-Adenosyl methionine (SAM) as the cofactor and methyl-donor [65]. Histone methylation main occurs on lysine and arginine residues, although glutamine (H2AQ104) methylation has been recently reported which is catalyzed by a nucleolus specific rRNA 2′-O-methyltransferase [66]. Histone lysine and arginine methylations are catalyzed by two major types of MTases, lysine methyltransferase (KMT) and protein arginine methyltransferase (PRMT). While most KMTs catalyze methyl transfer mainly onto histones, PRMTs methylate histones and a wide range of non-histone substrates. These two types of enzymes share little similarity in primary and tertiary structures and reaction mechanisms which are discussed separately in different categories.
2.1 Lysine Methyltransferases (KMTs)
SUV39H1 is the first characterized de novo histone KMT, containing a conserved catalytic motif (~120aa) which was initially identified in three drosophila proteins, Suppressor of variegation [Su(var)3–9], Enhancer of zeste [E(z)]and Trithorax (Trx) [67] and thereafter named as the SET domain [25]. While sequence alignment identified ~50 SET domain-containing proteins in human genome [68], many of them have been shown to possess histone KMT activities. Among the characterized KMTs, human DOT1-like (DOT1L) does not harbor a SET domain but processes the robust catalytic activity [49]. SET domain often localizes in the C-terminus of histone KMTs while this bifurcated motif can be divided into the conserved SET-N, SET-C and a highly variable insertion (SET-I) in the middle. These enzymes also harbor different sets of other domains and can be briefly classified into seven families, including SUV39, SET1, SET2, EZH, SMYD, PRDM, SUV420 and others (Fig. 2).
The SET domain of SUV39 family KMTs are encompassed by two conserved cysteine-rich pre-SET and post-SET domains. These KMTs mainly catalyze H3K9 methylation with the exception of SETMAR which methylates H3K4 and H3K36 [16]. The SET1 and SET2 families KMTs comprise different groups of large enzymes which specifically methylate H3K4 and H3K36 respectively. The SET1 family KMTs lack the pre-SET domain and their enzymatic activities require the formation of complexes with other proteins including RBBP5 and ASH2L [69]. In contrast, the pre-SET domain is replaced by a AWS (Associated with SET) domain in the SET2 family KMTs. This unique AWS-SET-postSET configuration is believed to confer the specific methyl transfer onto H3K36 [42]. In the EZH family KMTs, both pre-SET and post-SET domains are absent. However, these enzymes harbor a conserved CXC domain which is located upstream of the SET domain and is critical for their activity. EZH1/2 also have no detectable activity unless they form the Polycomb Repressive Complex 2 (PRC2) with SUZ12 and EED [70]. Unlike other KMT families, the SUV420H family KMTs only contain the conserved SET and post-SET domains but no other domains. SUV420H1/H2 are able to introduce H4K20me2/3 to H4K20me1 [55].
The SET domain in the SMYD and PRDM families KMTs has unique configurations. SMYD family of KMTs contain a long interposed sequence composed by a MYND domain and a SET-I motif. The MYND domain of SMYD1/2 has been shown to interact with the proline-rich motif in their binding partners while the MYND domain of SMYD3 is believed to bind to DNA [71]. The SMYD family of KMTs also contain the post-SET domain and a conserved C-terminal domain (CTD) with unknown function. Different from other KMT families, SMYDs display diverse substrate specificity [40, 58]. The PRDM family of KMTs harbor a catalytic PR/SET domain which only shares 20% similarity with the SET domain in sequence but displays a similar tertiary structure. Most PRDMs contain multiple zinc-fingers while some also have a zinc knuckle motif. The human PRDM family has 17 members in which six have histone KMT activities [30,31,33, 46].
Other human KMTs do not fit into above categories are SETD7, SETD8 and DOT1L. SETD7 and SETD8 only contain the catalytic SET domain and specifically mediate mono-methylation on H3K4 [72] and H4K20 [73] respectively. DOT1L is a non-SET domain-containing KMT which specifically catalyzes H3K79 methylation. The catalytic domain of DOT1L is similar to the catalytic domain of PRMT which utilizes a Rossmann fold for SAM binding [74, 75]. Furthermore, in vitro methylation catalyzed by DOT1L is a non-processive reaction whereas the reaction mediated by most SET domain-containing KMTs is processive [76]. Although several other proteins have also been reported to possess KMT activities [77, 78], the molecular details underlying these reactions still remain elusive.
2.2 Molecular Basis of Lysine Methylation
The representative SET domains of all KMT family members have been crystalized and their tertiary structures reveal several common features which are critical to confer efficient methyl transfer reaction [79]. Overall the SET domain is a highly interwined globular structure with extensive intra-domain interactions, suggesting each motif is critical for the structural integrity. The conserved SET-N and SET-C motifs form three beta-sheets and two beta-sheets with a pseudo-knot structure respectively. The variable SET-I motif also displays a similar structural fold, containing two anti-parallel β-strands and a short α-helix. Together with SET-C, SET-I motif is directly involved in substrate and SAM bindings and thus contributes to the substrate specificity [79]. Furthermore, zinc-chelating was observed in several structures. For example, the pre-SET domain of SUV39H family chelates three zinc ions [80, 81] and the AWS domain of SET2 family chelates two [36, 44], whereas the Zinc-Knuckle domain of PRDM9 chelate one zinc ion [82] and the CXC domain of EZH2 binds to six [70]. These domains do not contact substrate but pack against the SET domain to facilitate its structure stability and enzymatic activity [79]. Zinc-chelating was missing in SU420H1, SETD7 and SETD8 structures, however, a α-helix bundle, a beta-sheet and a long alpha-helix were observed respectively [72, 73, 83], suggesting that they may exert the similar function without zinc binding. Additionally, three cysteines in the post-SET domain of many KMTs chelate one zinc ion together with a conserved cysteine in the SET-C motif and this structure is also critical for the enzymatic activity [84]. Without zinc- chelating, the C-terminal sequences of the SET domain in SETD7 [72] and SETD8 [73] also form the similar structural fold for substrate and SAM binding.
Furthermore, the SET domain interacts with histone substrate and SAM through opposite surfaces. While SAM fits into an open concave pocket, histone peptide exhibits an extended conformation and extensively interactions with the binding groove. In this way, the target lysine is precisely positioned and its side chain can go through a narrow channel to meet with SAM [79, 85]. Different SET domains form different interactions with the backbone of their substrates to specifically define the methylation site. For instance, DIM-5, a neurospora SUV39 family KMT methylates H3K9 (QARK9ST) but not H3K27 (AARK27SA) due to its specific interaction with the side chain of T11 [81]. Moreover, the alkyl component of the lysine side-chain inhabits a hydrophobic environment while the ε-Nitrogen is stabilized by hydrogen bonds with surrounding carbonyl groups and hydroxyl-group [85]. In SETD7 structure, one ε-nitrogen on the side chain of H3K4 forms a hydrogen bond with a conserved Y245 and another with a tightly bound water molecule to prohibit the rotation of the εC-N bond for additional methylation. Accordingly, Y245A mutation enables SETD7 to catalyze H3K4me2/3 [72]. Mutation of another the target lysine binding tyrosine (Y305F) also results in H3K4me2 [84]. While the same Y-to-F mutation in SETD8 [73], MLL [86], G9A [70, 71] leads to similar effects on methylation, the F281Y mutation in DIM-5 disables the catalyzed H3K9me3 [84]. Intriguingly, the ε-N on the target lysine side chain forms a critical hydrogen-bond with S161 in the SET domain of mouse Suv420h2, which makes the enzyme inefficient for trimethylation [83, 87]. Disabling this hydrogen-bond by S161A mutation greatly increases H4K20me3 [83]. Together, the substrate specificity of KMTs is determined by extensive substrate-SET domain interactions whereas the methylation states rely on the accommodation of the ammonium group on the target lysine side chain in the structure.
SMYD proteins share a conserved bilobal structure in which the catalytic core is located in the middle of the N-terminal lobe with the MYND domain and CTD around. While the MYND domain is dispensable for methylation, the SET and post-SET domains form a surface pocket for cofactor binding and a deep pocket of the interface between the SET domain and CTD binds to substrate [71]. Although the overall structures are similar, SMYD1-3 display substantial differences in the size and surrounding structure of the substrate binding pocket, which could be responsible for divergent specificities on substrate and methylation states [88]. In the PRDM family KMTs, the SET domain signature motifs are poorly conserved. However, the overall structure of the PR/SET domain corresponds to the SET domains, in which the central SET domain fold is flanked by pre-SET and post-SET regions. The bindings of cofactor and substrate peptide to the PR/SET domain of Prdm9 are also similar to the SET domains [82]. These findings suggest that the structural similarity to the SET domain confers the lysine methylation activity of both SMYD and PR/SET domains. In contrast, the catalytic domain of DOT1L forms an open α/β structure which is comprised of a seven-stranded β sheet. This structure is distinct from the SET, SMYD or PR/SET domain but similar to several class I SAM-dependent MTases [89]. The active core of DOT1L has an elongated structure, containing a SAM binding pocket and a lysine binding channel. While the SAM binding pocket is critical for methylation, the lysine binding channel allows the accommodation of all three methylation states. The positive charged C-terminal region of the catalytic domain is also critical for the enzymatic activity, likely through binding to negatively charged nucleosomal DNA [75]. Despite of the structure similarity between DOTlL and PRMTs, arginine methylation by DOT1L was not detected.
2.3 Protein Arginine Methyltransferase
Human genome harbors eleven protein arginine methyltransferases (PRMTs), eight of them are able to methylate different histones. Based on methylation products, PRMTs are classified into four different types [7]. Type I enzymes, including PRMT1, 3, 4, 6 and 8 introduce monomethylation on arginine and further proceed to asymmetrical di-methylation (aDMA). PRMT5, 7 and 9 are type II PRMTs which generate the symmetrical dimethylated products (sDMA) after the initial monomethylation. Type III enzymes only catalyze monomethylation but do not proceed. Different from methylation on the terminal guanidine nitrogen atoms catalyzed by above PRMTs, Type IV PRMTs introduce monomethylation of δ-nitrogen. Most PRMTs catalyze aDMA on arginine, which is likely attributed to the higher energetically challenging with sDMA [90]. The characterized PRMTs and their function are also summarized in Table 1.
2.4 Molecular Basis of Arginine Methylation
Different from the SET domain, the conserved 310-aa catalytic core of PRMTs shares a similar structure with a Rossmann-fold domain and a C-terminal β-barrel domain and functions in a homo-dimer [91]. The methylation occurs at the interface of the catalytic core where SAM is accommodated in the Rossmann-fold, whereas the substrate peptide binds to an acidic groove with the side chain of target arginine inserted into a narrow tunnel to meet SAM [92]. Two notable structures were observed at the interface, a double-E loop from the Rossmann fold and a THW loop from the beta-barrel, which are important for methylation. It has been shown that E181D mutation in the double-E loop of Trypanosoma brucei PRMT7 converts this type III enzyme to a type I enzyme to catalyze asymmetric dimethylarginine (aDMA) [93]. Another critical motif for the SAM and target arginine binding is in the N-terminal helix of Rossmann-fold. F379M mutation in this motif of C. elegans PRMT5 partially shifts the reaction towards aDMA. This mutation likely opens up the active site to allow more bulky asymmetrical di-methyl groups [94]. To corroborate the importance of this residue, M48F mutation of rat PRMT1 enables its ability to catalyze symmetrical di-methylation [90], suggesting the Phe or Met residue in the N-terminal helix of Rossmann-fold could define the type of the methylation. However, type II PRMT9 contains a Met at the exact is position. Therefore, this proposed F/M switch could only be the part of the underlying mechanism [91]. In general, the size of the target arginine binding pocket significantly affects the product specificity catalyzed by different PRMTs.
3 Regulation of Histone Methylation
The enzymatic activity of HMTs is often evaluated in an in vitro assay in which the enzyme is incubated with SAM and substrate to catalyze methylation. In this reaction, several HMTases, including EZH2 and MLL failed to show robust activity unless their core protein complexes were used [95, 96]. Different substrates have also been used in this assay, including histone peptides, recombinant histones, histone octamers and nucleosomes [56]. While several KMTs prefer nucleosomes, such as SETD8 [56], SUV420H1/2 [55] and DOT1L [49], many enzymes predominantly methylate recombinant histones or octamers, such as SETD7 [15], G9A and GLP [97]. These observations suggest that the intact catalytic domain is necessary but not sufficient for histone methylation. Therefore, we discuss several regulatory mechanisms at different molecular layers.
3.1 Regulation by Interacting Proteins
The structure of EZH2 SET domain uncovers an inappropriate position of the SET-I and post-SET domains, which prohibits their interactions with substrate peptide and SAM [98, 99]. Recently, the crystal structures of PRC2 reveal that the extensive protein interactions with EED and SUZ12 optimize the structure of EZH2’s SET-I to form an active catalytic moiety for H3K27 methylation [70]. In the SET domain of MLL, the SET-I motif is highly dynamic. After forming the protein complex with the RBP5-ASH2L heterodimer, extensive protein interactions significantly reduce this inherent flexibility and lock the SET domain in an active conformation to enable efficient methyl transfer [69]. Intriguingly, the intermolecular β-sheet interactions between MLL SET-I and RBBP5(330–344aa) was also observed in the structure of other KMTs. In SUV39 and SET2 family KMTs, the similar interactions are formed intramolecularly between the SET-I and a short fragment upstream of the pre-SET domain. This fragment also exists in EZH2 and functions as SET activation loop (SAL) [70], suggesting it is a conserved structural configuration for the functional SET domain. Together, these novel advances in structure demonstrate that the inherent imperfection of certain SET domains can be corrected through interactions with their binding partners.
Furthermore, interacting proteins can regulate HMTases’ activity through different mechanisms. In the in vitro assays, PRMT5 equally catalyzes symmetrical methylation on both H3R8 and H4R3. However, it preferentially methylates H4R3 after binding to COPR5 [100], suggesting this regulatory protein fine-tunes PRMT5’s substrate specificity. HSP90α, the binding partner of SMYD2 stimulates methylation on H3K4 but not H3K36 [101]. Similarly, the substrate specificity of EZH2 on either H1K26 or H3K27 is modulated by a PRC2 core component EED [63]. A Polycomb-like protein PHF1 also facilitates PRC2-mediated H3K27me3 without affecting H3K27me1/2 [102, 103]. Additionally, SETDB1’s binding partner ATF7IP/AM, an ATFα-associated factor not only augments SETDB1’s enzymatic activity, but also facilitates the conversion of H3K9me2 to H3K9me3 in vitro and in vivo [104]. However, the similar effects were not observed when peptide substrates were used in the in vitro assays [105]. The molecular mechanisms underlying these regulations are largely unknown.
3.2 Regulation by Post-Translational Modifications
Several HMTases are subjected to different post-translational modifications which could also regulate their catalytic activity. For example, PKB/AKT phosphorylate EZH2-Ser21 and this phosphorylation inhibits EZH2 binding to H3 and thus reduces H3K27me3 in vitro and in vivo [106]. In glioblastoma stem-like cells, EZH2 interacts with and methylates STAT3, while the Ser21 phosphorylation facilitates PRC2-catalyzed STAT3-K180 methylation to activate STAT3 signaling [107]. In response to DNA damage, SETD7 has been shown to interact with and methylate SUV39H1 at K105 and K123, leading to a dramatically reduced enzymatic activity of SUV39H1. Since these lysines are close to the chromodomain which is critical for chromatin binding, these methylations could weaken the SUV39H1-substrate interaction [108]. While SUV39H1-K266 acetylation within the SET domain reduces its catalytic activity, SIRT1-mediated deacetylation can restore it by facilitating the interaction between the SET and post-SET domains [109].
Moreover, bacteria-purified SETDB1 failed to methylate histones in the in vitro assays but 293T or Sf9 cell-purified enzymes displayed robust activity [110], indicating post-translational modifications could regulate SETDB1’s activity. Recently we demonstrate that SETDB1 is monoubiquitinated at K867 (K867ub1) within its unique SET-I motif via an E3-independent mechanism. The conjugated-ubiquitin is protected from active deubiquitination, likely through multiple intramolecular interactions. Importantly, the resulting constitutive monoubiquitination is required for SETDB1’s enzymatic activity and function. While most post-translational modifications are dynamically regulated, our findings highlight the constitutive role of K867Ub1 in regulating enzymatic activity of KMTs [97].
3.3 Regulation by Histone Modification
It is well-documented that histone methylation is also modulated by other histone modifications. On the same molecule, H3S10 phosphorylation blocks the access of the adjacent H3K9 for methylation [111]. The similar regulation also exists on different histones and one good example is that H2A and H2B ubiquitination affects H3K4 and H3K79 methylation. Site-specific installation of H2BK120ub1 causes the allosteric regulation of nucleosomes to facilitate DOT1L binding and thus increases the intranucleosomal H3K79me1/2 [112]. H2BK120ub1 also interacts with the N-terminal winged helix motif of ASH2L and promotes H3K4 methylation catalyzed by ASH2L-MLL-RBP5 complex [113]. However, H2AK119ub1 inhibits PRC2-catalyzed H3K27 methylation, suggesting that ubiquitination at different sites trans-regulates different histone methylations [114]. Similarly, the internucleosomal regulations have also been reported. While the SET domain of GLP and G9A preferentially methylates histone octamer, the full-length proteins efficiently catalyze oligonucleosomal methylation because G91/GLP bind to methyl-H3K9 binding on adjacent nucleosomes through their Ankyrin repeats domain [115]. Similarly, PRC2-catalzyed H3K27 methylation is stimulated by EED which binds to methyl-H3K27 on adjacent nucleosomes [116, 117]. In the PRC2 structure, H3K27me3 binding also stabilizes the conformation of EZH2 N-terminal SRM motif and affects the SET-I conformation to facilitate H3K27 methylation [70]. Similar to G9A/GLP, the chromodomain of SUV39H1 recognizes H3K9me3 and this methyl-binding anchors the enzyme to chromatin allosterically to allow further spreading of H3K9me3 [118]. Therefore, pre-existing modifications on histones could dramatically modulate methylation through different mechanisms.
4 Recognition of Histone Methylation
Although histone methylation does not neutralize the positive charge of DNA, several methylations could affect nucleosome structure to facilitate transcription [119]. For example, H3R42 locates at the DNA entry-exit region of nucleosome and addition of asymmetric dimethylation by CARM1 and PRMT6 could reduce nucleosome stability [48]. Structural study reveals that H3K79me2 leads to a subtle reorientation of the surrounding region in nucleosome [120]. Moreover, H3R17me2a and H3K4me1 have been shown to reduce chromatin association of NuRD complex, suggesting that these methylation marks could indirectly regulate accessibility of chromatin [121, 122]. In most cases, however, histone methylation serves as docking site for specific binding proteins which in turn recruit additional chromatin modifiers for diverse functional outcomes [119]. So far at least nine domains have been characterized as methyl-histone binding motifs that are briefly summarized in Table 2 together with their recognition sites.
4.1 The Royal Family of Domains
Several methyl-histone binding motifs, including chromodomain, Tudor, MBT and PWWP belong to the Royal family of domains that are descended from a common ancestor [173]. These domains share a similar structure of barrel-like three-strand β-sheet with a short helix and bind to different methyl-lysines [174]. Existing in ~31 human proteins [175], the chromodomain recognizes methyl-lysines using an aromatic cage formed by three highly conserved residue [176]. The binding site specificity is determined by specific interactions between amino acids around the methyl-lysine and the chromodomain. For example, the chromodomains of HP1 [176, 177] and MPP8 [125, 178, 179] preferentially recognize H3K9me3 [176, 177] while the same domain in PC binds to H3K27me3 [180, 181]. Intriguingly, the CHD family proteins contain tandem chromodomains which bind to a single H3K4me3 mark in a coordinated manner [182, 183]. In most cases, the aromatic cage of chromodomain accommodates trimethylation and binds to mono- or di-methylation with a lower affinity due to less optimal van der Waals and cation-π interaction [149, 184].
The Tudor domain forms four- or five β-strands and bind to methyl-lysine with a similar aromatic cage. The tandem Tudor domains of JMJD2A specifically recognize H3K4me3 [156]. In the Tudor domains of 53BP1, the carboxylate group of Asp1521 forms a hydrogen bond with the nitrogen group of dimethyl-amine to confer stable binding to H4K20me2 but causes a steric hindrance for the trimethyl-amine [149]. Therefore, binding specificity to different methylation states is precisely regulated in different Tudor domains. Although the tandem Tudor domains are often observed, only one of the tandem domains interacts with methyl-lysine, leaving another free of binding.
The MBT domain is a larger motif (~100aa) containing 2–4 repeats and exists in nine human proteins. All human MBT domains harbor the conserved aromatic residues, indicating they can bind to methyl-lysines. However, the MBT domain preferentially recognizes mono- and di-methylated lysines [185] with poor site specificity. It is likely due to the lack of interactions with amino acids around the methyl-lysine [186, 187]. The PWWP domain (100–150aa) folds into a five-strand β-sheet packed against a helical bundle with significant variations in β2 and β3 while many have the conserved aromatic cage [188]. It has been shown that the PWWP domains of Pdp1 and DNMT3A/3B specifically recognize H4K20me [189] and H3K36me3 [128, 190] respectively.
4.2 Other Methyl-Lysine Binding Domains
In addition to the Royal family of domains, several other motifs can recognize different methyl-lysines as well, including PHD finger, CW, Ankyrin repeat, WD repeat and BAH. The PHD finger domain forms two anti-parallel beta strands and one C-terminal alpha-helix, which are stabilized by two zinc ions chelated by a consensus C4HC3 sequence [191]. This motif exists in multiple chromatin-associated proteins and many have been shown to recognize methylated histones [130, 192]. Due to a favorable accommodation of H3R2, most PHD finger domains bind to methyl-H3K4 [150, 151]. For example, the PHD domains of BPTF and ING2 recognize H3K4me3 through an aromatic cage similar to the Royal family domains [150, 151]. Similarly, CW domain also centers two anti-parallel beta-strands and chelates one Zinc ion with the consensus C4 sequence [164]. The CW domain of ZCWPW1 preferentially recognizes H3K4me3 [163] through an aromatic cage. Among the seven human CW domain-containing proteins, four contain at least two conserved aromatic residues and can bind to H3K4me3 peptide in vitro [164].
The Ankyrin repeat domain is a widely distributed motif for protein-protein interactions. Intriguingly, the Ankyrin repeat domains of G9a and GLP can bind to H3K9me2 [160]. Similar to the chromodomain, three aromatic residues and a Glu are involved in the binding. However, the size of the aromatic cage cannot accommodate trimethyl groups. Distinctly, the WD repeats form a seven-bladed β-propeller, in which three scattered aromatic residues are responsible for methyl-lysine binding. It has been shown that the WD40 repeat domain of a PRC2 component EED recognizes multiple methyl-lysines, including H1K26me3, H3K9me3, H3K27me3 and H4K20me3 with similar affinity [116, 117]. The MLL complex subunit WDR5 contains seven WD40 repeats which have been reported to bind to methyl-H3K4 [193]. Existing in many chromatin-associated proteins, the BAH domain folds into a beta-rich structure. It has been demonstrated that the BAH domain of a DNA replication protein ORC1 specifically recognizes H4K20me2 through a four aromatic residues cage [168]. Due to a hydrogen bond between methyl-ammonium and side carboxylate chain of a nearby Glu, this binding favors H4K20me2 over H4K20me3 [168]. The BAH domain in BAHD1 has also been reported to recognize H3K27me3 [169].
4.3 Recognition of Arginine Methylation
Among methyl-histone binding domains, multiple Tudor domain-containing proteins also bind to methyl-arginine in various proteins, suggesting they can recognize methyl-arginine in histones [194]. The extended Tudor domain of SND1 has been shown to interact with peptides harboring H4R3 methylation [195]. Similar to methyl-lysine recognition, methyl-H4R3 binding also involves the aromatic cage of the Tudor domain of SND1. However, two aromatic residues pack to the guanidium planar in parallel, whose distance to the ammonium group is shorter than methyl-lysine binding [195]. While some Tudor domains recognize aDMA and sDMA on histones with comparable affinity, others display clear preference. In a peptide pull-down assay, the Tudor domain of TDRD3 preferentially recognizes aDMA on histone H3 tail [166]. Structural study reveals that such selectivity is rendered by a unique hydrogen bond between the unmodified amino group and the hydroxyl group of a Tyr in the aromatic cage [196]. Unexpectedly, the Glu-rich region in PELP1 has also been shown to bind to H3K4me2, H3K9me2 [197] and arginine methylated in vitro [198], suggesting there are more unidentified methyl-histone binding motifs.
4.4 Modulation of Methyl-Histone Binding
The methyl-histone binding by different domains is also subject to multiple regulations. Because of the extensive interaction between the binding domain and amino acids adjacent to the methylated residue, post-translational modifications on these amino acids could have drastic effects. For example, H3S10 phosphorylation, one of the most prominent modification on mitotic chromosome, inhibits HP1 binding to H3K9me3 [199, 200]. Similarly, H3K4me3 binding by different domains are blocked when H3R2 is asymmetrically dimethylated [10]. Intriguingly, modification adjacent to the methyl-lysine has also been shown to facilitate the recognition by binding domains. For example, the structure of RAG2 PHD finger domain complexed with H3K4me3 peptide uncovers an additional binding pocket. Therefore, H3K4me3 binding is increased when H3R2 is symmetrically methylated on the same molecule [201]. Furthermore, methyl-histone binding are regulated by other interacting partners. The structure of the ternary complex of Pygo PHD finger, the BCL9/Legless HD1 domain and H3K4me peptides demonstrates that the efficient H3K4me2 recognition requires the PHD-HD1 complex in instead of the PHD domain alone [155]. In addition to interacting proteins, ncRNA TUG1 can switch H3K9me3 binding by the PC2 chromodomain to H4R3me2s and H3K27me2. Through unknown mechanisms, another ncRNA NEAT2 can convert PC2’s H3K9me3 binding to H2AK5ac and H2AK13ac binding [202].
5 Conclusion Remark
As a key component of the proposed “histone code”, histone methylation is precisely regulated in cells and plays pivotal roles in the regulation of all chromatin-based processes. Histone methylation “code” is introduced by different groups of HMTases and recognized by various methyl-histone binding proteins. These proteins coordinate with various transcription factors, chromatin modifying proteins, signal pathway cascades and non-coding RNAs to constitute a large sophisticated network for diverse functional outcomes. It has been acknowledged that many histone methylating and binding proteins are altered in human diseases, including various cancers. Accordingly, numerous small molecule modulators have been developed and characterized for the pharmaceutical intervention of these diseases [203]. Multiple inhibitors targeting different HMTases have also been applied to various clinical trials. Therefore, a thorough understanding of the molecular basics underlying histone methylation and recognition will not only shed lights on their physiological functions, but also facilitate the development of therapeutic strategies for human diseases.
References
Woodcock CL, Ghosh RP (2010) Chromatin higher-order structure and dynamics. Cold Spring Harb Perspect Biol 2:a000596
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403:41–45
Ruthenburg AJ, Li H, Patel DJ, Allis CD (2007) Multivalent engagement of chromatin modifications by linked binding modules. Nat Rev Mol Cell Biol 8:983–994
Allfrey VG, Faulkner R, Mirsky AE (1964) Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci U S A 51:786–794
Murray K (1964) The occurrence of epsilon-N-methyl lysine in histones. Biochemistry 3:10–15
Strahl BD, Ohba R, Cook RG, Allis CD (1999) Methylation of histone H3 at lysine 4 is highly conserved and correlates with transcriptionally active nuclei in Tetrahymena. Proc Natl Acad Sci U S A 96:14967–14972
Bedford MT (2007) Arginine methylation at a glance. J Cell Sci 120:4243–4246
Guccione E et al (2007) Methylation of histone H3R2 by PRMT6 and H3K4 by an MLL complex are mutually exclusive. Nature 449:933–937
Hyllus D et al (2007) PRMT6-mediated methylation of R2 in histone H3 antagonizes H3 K4 trimethylation. Genes Dev 21:3369–3380
Iberg AN et al (2008) Arginine methylation of the histone H3 tail impedes effector binding. J Biol Chem 283:3006–3010
Schurter BT et al (2001) Methylation of histone H3 by coactivator-associated arginine methyltransferase 1. Biochemistry 40:5747–5756
Migliori V et al (2012) Symmetric dimethylation of H3R2 is a newly identified histone mark that supports euchromatin maintenance. Nat Struct Mol Biol 19:136–144
Wysocka J, Myers MP, Laherty CD, Eisenman RN, Herr W (2003) Human Sin3 deacetylase and trithorax-related Set1/Ash2 histone H3-K4 methyltransferase are tethered together selectively by the cell-proliferation factor HCF-1. Genes Dev 17:896–911
Lee JH, Tate CM, You JS, Skalnik DG (2007) Identification and characterization of the human Set1B histone H3-Lys4 methyltransferase complex. J Biol Chem 282:13419–13428
Wang H et al (2001) Purification and functional characterization of a histone H3-lysine 4-specific methyltransferase. Mol Cell 8:1207–1217
Lee SH et al (2005) The SET domain protein Metnase mediates foreign DNA integration and links integration to nonhomologous end-joining repair. Proc Natl Acad Sci U S A 102:18075–18080
Blazer LL et al (2016) PR domain-containing protein 7 (PRDM7) is a histone 3 lysine 4 trimethyltransferase. J Biol Chem 291:13509–13519
Powers NR et al (2016) The meiotic recombination activator PRDM9 trimethylates both H3K36 and H3K4 at recombination hotspots in vivo. PLoS Genet 12:e1006146
Hayashi K, Yoshida K, Matsui Y (2005) A histone H3 methyltransferase controls epigenetic events required for meiotic prophase. Nature 438:374–378
Tan X, Rotllant J, Li H, De Deyne P, Du SJ (2006) SmyD1, a histone methyltransferase, is required for myofibril organization and muscle contraction in zebrafish embryos. Proc Natl Acad Sci U S A 103:2713–2718
Hamamoto R et al (2004) SMYD3 encodes a histone methyltransferase involved in the proliferation of cancer cells. Nat Cell Biol 6:731–740
Van Aller GS et al (2012) Smyd3 regulates cancer cell phenotypes and catalyzes histone H4 lysine 5 methylation. Epigenetics 7:340–343
Blythe SA, Cha SW, Tadjuidje E, Heasman J, Klein PS (2010) Beta-catenin primes organizer gene expression by recruiting a histone H3 arginine 8 methyltransferase, Prmt2. Dev Cell 19:220–231
Pal S, Vishwanath SN, Erdjument-Bromage H, Tempst P, Sif S (2004) Human SWI/SNF-associated PRMT5 methylates histone H3 arginine 8 and negatively regulates expression of ST7 and NM23 tumor suppressor genes. Mol Cell Biol 24:9630–9645
Rea S et al (2000) Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406:593–599
O'Carroll D et al (2000) Isolation and characterization of Suv39h2, a second histone H3 methyltransferase gene that displays testis-specific expression. Mol Cell Biol 20:9423–9433
Ogawa H, Ishiguro K, Gaubatz S, Livingston DM, Nakatani Y (2002) A complex with chromatin modifiers that occupies E2F- and Myc-responsive genes in G0 cells. Science 296:1132–1136
Tachibana M, Sugimoto K, Fukushima T, Shinkai Y (2001) Set domain-containing protein, G9a, is a novel lysine-preferring mammalian histone methyltransferase with hyperactivity and specific selectivity to lysines 9 and 27 of histone H3. J Biol Chem 276:25309–25317
Yang L et al (2002) Molecular cloning of ESET, a novel histone H3-specific methyltransferase that interacts with ERG transcription factor. Oncogene 21:148–152
Falandry C et al (2010) CLLD8/KMT1F is a lysine methyltransferase that is important for chromosome segregation. J Biol Chem 285:20234–20241
Kim KC, Geng L, Huang S (2003) Inactivation of a histone methyltransferase by mutations in human cancers. Cancer Res 63:7619–7623
Pinheiro I et al (2012) Prdm3 and Prdm16 are H3K9me1 methyltransferases required for mammalian heterochromatin integrity. Cell 150:948–960
Eom GH et al (2009) Histone methyltransferase PRDM8 regulates mouse testis steroidogenesis. Biochem Biophys Res Commun 388:131–136
Bauer UM, Daujat S, Nielsen SJ, Nightingale K, Kouzarides T (2002) Methylation at arginine 17 of histone H3 is linked to gene activation. EMBO Rep 3:39–44
Wu H et al (2011) Histone methyltransferase G9a contributes to H3K27 methylation in vivo. Cell Res 21:365–367
Qiao Q et al (2011) The structure of NSD1 reveals an autoregulatory mechanism underlying histone H3K36 methylation. J Biol Chem 286:8361–8368
Kuo AJ et al (2011) NSD2 links dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol Cell 44:609–620
Asangani IA et al (2012) Characterization of the EZH2-MMSET histone methyltransferase regulatory axis in cancer. Mol Cell 49:80–93
Li Y et al (2009) The target of the NSD family of histone lysine methyltransferases depends on the nature of the substrate. J Biol Chem 284:34283–34295
Brown MA, Sims RJ 3rd, Gottlieb PD, Tucker PW (2006) Identification and characterization of Smyd2: a split SET/MYND domain-containing histone H3 lysine 36-specific methyltransferase that interacts with the Sin3 histone deacetylase complex. Mol Cancer 5:26
Sun XJ et al (2005) Identification and characterization of a novel human histone H3 lysine 36-specific methyltransferase. J Biol Chem 280:35261–35271
Kim DW, Kim KB, Kim JY, Seo SB (2011) Characterization of a novel histone H3K36 methyltransferase setd3 in zebrafish. Biosci Biotechnol Biochem 75:289–294
Eom GH et al (2011) Histone methyltransferase SETD3 regulates muscle differentiation. J Biol Chem 286:34733–34742
An S, Yeo KJ, Jeon YH, Song JJ (2011) Crystal structure of the human histone methyltransferase ASH1L catalytic domain and its implications for the regulatory mechanism. J Biol Chem 286:8369–8374
Tanaka Y, Katagiri Z, Kawahashi K, Kioussis D, Kitajima S (2007) Trithorax-group protein ASH1 methylates histone H3 lysine 36. Gene 397:161–168
Eram MS et al (2014) Trimethylation of histone H3 lysine 36 by human methyltransferase PRDM9 protein. J Biol Chem 289:12177–12188
Yu Y et al (2012) Histone H3 lysine 56 methylation regulates DNA replication through its interaction with PCNA. Mol Cell 46:7–17
Casadio F et al (2013) H3R42me2a is a histone modification with positive transcriptional effects. Proc Natl Acad Sci U S A 110:14894–14899
Feng Q et al (2002) Methylation of H3-lysine 79 is mediated by a new family of HMTases without a SET domain. Curr Biol 12:1052–1058
Ng HH et al (2002) Lysine methylation within the globular domain of histone H3 by Dot1 is important for telomeric silencing and Sir protein association. Genes Dev 16:1518–1527
Wang H et al (2001) Methylation of histone H4 at arginine 3 facilitating transcriptional activation by nuclear hormone receptor. Science 293:853–857
Karkhanis V et al (2012) Protein arginine methyltransferase 7 regulates cellular response to DNA damage by methylating promoter histones H2A and H4 of the polymerase delta catalytic subunit gene, POLD1. J Biol Chem 287:29801–29814
Tweedie-Cullen RY et al (2012) Identification of combinatorial patterns of post-translational modifications on individual histones in the mouse brain. PLoS One 7:e36980
Feng Y et al (2013) Mammalian protein arginine methyltransferase 7 (PRMT7) specifically targets RXR sites in lysine- and arginine-rich regions. J Biol Chem 288:37010–37025
Schotta G et al (2004) A silencing pathway to induce H3-K9 and H4-K20 trimethylation at constitutive heterochromatin. Genes Dev 18:1251–1262
Fang J et al (2002) Purification and functional characterization of SET8, a nucleosomal histone H4-lysine 20-specific methyltransferase. Curr Biol 12:1086–1099
Nishioka K et al (2002) PR-Set7 is a nucleosome-specific methyltransferase that modifies lysine 20 of histone H4 and is associated with silent chromatin. Mol Cell 9:1201–1213
Foreman KW et al (2011) Structural and functional profiling of the human histone methyltransferase SMYD3. PLoS One 6:e22290
Ancelin K et al (2006) Blimp1 associates with Prmt5 and directs histone arginine methylation in mouse germ cells. Nat Cell Biol 8:623–630
Waldmann T et al (2011) Methylation of H2AR29 is a novel repressive PRMT6 target. Epigenetics Chromatin 4:11
Binda O et al (2013) SETD6 monomethylates H2AZ on lysine 7 and is required for the maintenance of embryonic stem cell self-renewal. Epigenetics 8:177–183
Kogure M et al (2013) The oncogenic polycomb histone methyltransferase EZH2 methylates lysine 120 on histone H2B and competes ubiquitination. Neoplasia 15:1251–1261
Kuzmichev A, Jenuwein T, Tempst P, Reinberg D (2004) Different EZH2-containing complexes target methylation of histone H1 or nucleosomal histone H3. Mol Cell 14:183–193
Weiss T et al (2010) Histone H1 variant-specific lysine methylation by G9a/KMT1C and Glp1/KMT1D. Epigenetics Chromatin 3:7
Polevoda B, Sherman F (2007) Methylation of proteins involved in translation. Mol Microbiol 65:590–606
Tessarz P et al (2014) Glutamine methylation in histone H2A is an RNA-polymerase-I-dedicated modification. Nature 505:564–568
Jenuwein T, Laible G, Dorn R, Reuter G (1998) SET domain proteins modulate chromatin domains in eu- and heterochromatin. Cell Mol Life Sci 54:80–93
Cheng X, Collins RE, Zhang X (2005) Structural and sequence motifs of protein (histone) methylation enzymes. Annu Rev Biophys Biomol Struct 34:267–294
Li Y et al (2016) Structural basis for activity regulation of MLL family methyltransferases. Nature 530:447–452
Jiao L, Liu X (2015) Structural basis of histone H3K27 trimethylation by an active polycomb repressive complex 2. Science 350:aac4383
Spellmon N, Holcomb J, Trescott L, Sirinupong N, Yang Z (2015) Structure and function of SET and MYND domain-containing proteins. Int J Mol Sci 16:1406–1428
Xiao B et al (2003) Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature 421:652–656
Couture JF, Collazo E, Brunzelle JS, Trievel RC (2005) Structural and functional analysis of SET8, a histone H4 Lys-20 methyltransferase. Genes Dev 19:1455–1465
Sawada K et al (2004) Structure of the conserved core of the yeast Dot1p, a nucleosomal histone H3 lysine 79 methyltransferase. J Biol Chem 279:43296–43306
Min J, Feng Q, Li Z, Zhang Y, Xu RM (2003) Structure of the catalytic domain of human DOT1L, a non-SET domain nucleosomal histone methyltransferase. Cell 112:711–723
Chen X, Liu H, Shim AH, Focia PJ, He X (2008) Structural basis for synaptic adhesion mediated by neuroligin-neurexin interactions. Nat Struct Mol Biol 15:50–56
Cao F et al (2010) An Ash2L/RbBP5 heterodimer stimulates the MLL1 methyltransferase activity through coordinated substrate interactions with the MLL1 SET domain. PLoS One 5:e14102
Patel A, Vought VE, Dharmarajan V, Cosgrove MS (2011) A novel non-SET domain multi-subunit methyltransferase required for sequential nucleosomal histone H3 methylation by the mixed lineage leukemia protein-1 (MLL1) core complex. J Biol Chem 286:3359–3369
Xiao B, Wilson JR, Gamblin SJ (2003) SET domains and histone methylation. Curr Opin Struct Biol 13:699–705
Wu H et al (2010) Structural biology of human H3K9 methyltransferases. PLoS One 5:e8570
Zhang X et al (2002) Structure of the Neurospora SET domain protein DIM-5, a histone H3 lysine methyltransferase. Cell 111:117–127
Wu H et al (2013) Molecular basis for the regulation of the H3K4 methyltransferase activity of PRDM9. Cell Rep 5:13–20
Southall SM, Cronin NB, Wilson JR (2014) A novel route to product specificity in the Suv4-20 family of histone H4K20 methyltransferases. Nucleic Acids Res 42:661–671
Zhang X et al (2003) Structural basis for the product specificity of histone lysine methyltransferases. Mol Cell 12:177–185
Dillon SC, Zhang X, Trievel RC, Cheng X (2005) The SET-domain protein superfamily: protein lysine methyltransferases. Genome Biol 6:227
Southall SM, Wong PS, Odho Z, Roe SM, Wilson JR (2009) Structural basis for the requirement of additional factors for MLL1 SET domain activity and recognition of epigenetic marks. Mol Cell 33:181–191
Wu H et al (2013) Crystal structures of the human histone H4K20 methyltransferases SUV420H1 and SUV420H2. FEBS Lett 587:3859–3868
Xu S, Zhong C, Zhang T, Ding J (2011) Structure of human lysine methyltransferase Smyd2 reveals insights into the substrate divergence in Smyd proteins. J Mol Cell Biol 3:293–300
Nguyen AT, Zhang Y (2011) The diverse functions of Dot1 and H3K79 methylation. Genes Dev 25:1345–1358
Gui S et al (2014) A remodeled protein arginine methyltransferase 1 (PRMT1) generates symmetric dimethylarginine. J Biol Chem 289:9320–9327
Fuhrmann J, Clancy KW, Thompson PR (2015) Chemical biology of protein arginine modifications in epigenetic regulation. Chem Rev 115:5413–5461
Zhang X, Zhou L, Cheng X (2000) Crystal structure of the conserved core of protein arginine methyltransferase PRMT3. EMBO J 19:3509–3519
Debler EW et al (2016) A glutamate/aspartate switch controls product specificity in a protein arginine methyltransferase. Proc Natl Acad Sci U S A 113:2068–2073
Sun L et al (2011) Structural insights into protein arginine symmetric dimethylation by PRMT5. Proc Natl Acad Sci U S A 108:20538–20543
Kuzmichev A, Nishioka K, Erdjument-Bromage H, Tempst P, Reinberg D (2002) Histone methyltransferase activity associated with a human multiprotein complex containing the enhancer of Zeste protein. Genes Dev 16:2893–2905
Cao R et al (2002) Role of histone H3 lysine 27 methylation in polycomb-group silencing. Science 298:1039–1043
Sun L, Fang J (2016) E3-independent constitutive monoubiquitination complements histone methyltransferase activity of SETDB1. Mol Cell 62:958–966
Wu H et al (2013) Structure of the catalytic domain of EZH2 reveals conformational plasticity in cofactor and substrate binding sites and explains oncogenic mutations. PLoS One 8:e83737
Antonysamy S et al (2013) Structural context of disease-associated mutations and putative mechanism of autoinhibition revealed by X-ray crystallographic analysis of the EZH2-SET domain. PLoS One 8:e84147
Lacroix M et al (2008) The histone-binding protein COPR5 is required for nuclear functions of the protein arginine methyltransferase PRMT5. EMBO Rep 9:452–458
Abu-Farha M et al (2008) The tale of two domains: proteomics and genomics analysis of SMYD2, a new histone methyltransferase. Mol Cell Proteomics 7:560–572
Cao R et al (2008) Role of hPHF1 in H3K27 methylation and Hox gene silencing. Mol Cell Biol 28:1862–1872
Sarma K, Margueron R, Ivanov A, Pirrotta V, Reinberg D (2008) Ezh2 requires PHF1 to efficiently catalyze H3 lysine 27 trimethylation in vivo. Mol Cell Biol 28:2718–2731
Wang H et al (2003) mAM facilitates conversion by ESET of dimethyl to trimethyl lysine 9 of histone H3 to cause transcriptional repression. Mol Cell 12:475–487
Basavapathruni A et al (2016) Characterization of the enzymatic activity of SETDB1 and its 1:1 complex with ATF7IP. Biochemistry 55:1645–1651
Cha TL et al (2005) Akt-mediated phosphorylation of EZH2 suppresses methylation of lysine 27 in histone H3. Science 310:306–310
Kim E et al (2013) Phosphorylation of EZH2 activates STAT3 signaling via STAT3 methylation and promotes tumorigenicity of glioblastoma stem-like cells. Cancer Cell 23:839–852
Wang D et al (2013) Methylation of SUV39H1 by SET7/9 results in heterochromatin relaxation and genome instability. Proc Natl Acad Sci U S A 110:5516–5521
Vaquero A et al (2007) SIRT1 regulates the histone methyl-transferase SUV39H1 during heterochromatin formation. Nature 450:440–444
Schultz DC, Ayyanathan K, Negorev D, Maul GG, Rauscher FJ 3rd (2002) SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev 16:919–932
Duan Q, Chen H, Costa M, Dai W (2008) Phosphorylation of H3S10 blocks the access of H3K9 by specific antibodies and histone methyltransferase. Implication in regulating chromatin dynamics and epigenetic inheritance during mitosis. J Biol Chem 283:33585–33590
McGinty RK, Kim J, Chatterjee C, Roeder RG, Muir TW (2008) Chemically ubiquitylated histone H2B stimulates hDot1L-mediated intranucleosomal methylation. Nature 453:812–816
Wu L et al (2013) ASH2L regulates ubiquitylation signaling to MLL: trans-regulation of H3 K4 methylation in higher eukaryotes. Mol Cell 49:1108–1120
Whitcomb SJ et al (2012) Histone monoubiquitylation position determines specificity and direction of enzymatic cross-talk with histone methyltransferases Dot1L and PRC2. J Biol Chem 287:23718–23725
Liu N et al (2015) Recognition of H3K9 methylation by GLP is required for efficient establishment of H3K9 methylation, rapid target gene repression, and mouse viability. Genes Dev 29:379–393
Margueron R et al (2009) Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 461:762–767
Xu C et al (2010) Binding of different histone marks differentially regulates the activity and specificity of polycomb repressive complex 2 (PRC2). Proc Natl Acad Sci U S A 107:19266–19271
Muller MM, Fierz B, Bittova L, Liszczak G, Muir TW (2016) A two-state activation mechanism controls the histone methyltransferase Suv39h1. Nat Chem Biol 12:188–193
Tessarz P, Kouzarides T (2014) Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 15:703–708
Lu X et al (2008) The effect of H3K79 dimethylation and H4K20 trimethylation on nucleosome and chromatin structure. Nat Struct Mol Biol 15:1122–1124
Wu J, Cui N, Wang R, Li J, Wong J (2012) A role for CARM1-mediated histone H3 arginine methylation in protecting histone acetylation by releasing corepressors from chromatin. PLoS One 7:e34692
Nishioka K et al (2002) Set9, a novel histone H3 methyltransferase that facilitates transcription by precluding histone tail modifications required for heterochromatin formation. Genes Dev 16:479–489
Lachner M, O'Carroll D, Rea S, Mechtler K, Jenuwein T (2001) Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410:116–120
Vezzoli A et al (2010) Molecular basis of histone H3K36me3 recognition by the PWWP domain of Brpf1. Nat Struct Mol Biol 17:617–619
Kokura K, Sun L, Bedford MT, Fang J (2010) Methyl-H3K9-binding protein MPP8 mediates E-cadherin gene silencing and promotes tumour cell motility and invasion. EMBO J 29:3673–3687
Wen H et al (2014) ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression. Nature 508:263–268
Bernstein E et al (2006) Mouse polycomb proteins bind differentially to methylated histone H3 and RNA and are enriched in facultative heterochromatin. Mol Cell Biol 26:2560–2569
Dhayalan A et al (2010) The Dnmt3a PWWP domain reads histone 3 lysine 36 trimethylation and guides DNA methylation. J Biol Chem 285:26114–26120
Zhao Q et al (2009) PRMT5-mediated methylation of histone H4R3 recruits DNMT3A, coupling histone and DNA methylation in gene silencing. Nat Struct Mol Biol 16:304–311
Shi X et al (2006) ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442:96–99
Klymenko T et al (2006) A polycomb group protein complex with sequence-specific DNA-binding and selective methyl-lysine-binding activities. Genes Dev 20:1110–1122
Kim S et al (2016) Mechanism of histone H3K4me3 recognition by the plant homeodomain of inhibitor of growth 3. J Biol Chem 291:18326–18341
Hung T et al (2009) ING4 mediates crosstalk between histone H3 K4 trimethylation and H3 acetylation to attenuate cellular transformation. Mol Cell 33:248–256
Champagne KS et al (2008) The crystal structure of the ING5 PHD finger in complex with an H3K4me3 histone peptide. Proteins 72:1371–1376
Musselman CA et al (2009) Binding of the CHD4 PHD2 finger to histone H3 is modulated by covalent modifications. Biochem J 423:179–187
Chang PY et al (2010) Binding of the MLL PHD3 finger to histone H3K4me3 is required for MLL-dependent gene transcription. J Mol Biol 400:137–144
Sims RJ 3rd et al (2005) Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. J Biol Chem 280:41789–41792
Karagianni P, Amazit L, Qin J, Wong J (2008) ICBP90, a novel methyl K9 H3 binding protein linking protein ubiquitination with heterochromatin formation. Mol Cell Biol 28:705–717
Fischle W, Franz H, Jacobs SA, Allis CD, Khorasanizadeh S (2008) Specificity of the chromodomain Y chromosome family of chromodomains for lysine-methylated ARK(S/T) motifs. J Biol Chem 283:19626–19635
Wang GG et al (2009) Haematopoietic malignancies caused by dysregulation of a chromatin-binding PHD finger. Nature 459:847–851
Sun Y et al (2009) Histone H3 methylation links DNA damage detection to activation of the tumour suppressor Tip60. Nat Cell Biol 11:1376–1382
Iwase S et al (2007) The X-linked mental retardation gene SMCX/JARID1C defines a family of histone H3 lysine 4 demethylases. Cell 128:1077–1088
Zhang P et al (2006) Structure of human MRG15 chromo domain and its binding to Lys36-methylated histone H3. Nucleic Acids Res 34:6621–6628
Wen H et al (2010) Recognition of histone H3K4 trimethylation by the plant homeodomain of PHF2 modulates histone demethylation. J Biol Chem 285:9322–9326
Moore SA, Ferhatoglu Y, Jia Y, Al-Jiab RA, Scott MJ (2010) Structural and biochemical studies on the chromo-barrel domain of male specific lethal 3 (MSL3) reveal a binding preference for mono- or dimethyllysine 20 on histone H4. J Biol Chem 285:40879–40890
Kim D et al (2010) Corecognition of DNA and a methylated histone tail by the MSL3 chromodomain. Nat Struct Mol Biol 17:1027–1029
Feng W, Yonezawa M, Ye J, Jenuwein T, Grummt I (2010) PHF8 activates transcription of rRNA genes through H3K4me3 binding and H3K9me1/2 demethylation. Nat Struct Mol Biol 17:445–450
Vermeulen M et al (2007) Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell 131:58–69
Botuyan MV et al (2006) Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell 127:1361–1373
Li H et al (2006) Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature 442:91–95
Wysocka J et al (2006) A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442:86–90
Nady N et al (2011) Recognition of multivalent histone states associated with heterochromatin by UHRF1 protein. J Biol Chem 286:24300–24311
Liu Y, Subrahmanyam R, Chakraborty T, Sen R, Desiderio S (2007) A plant homeodomain in RAG-2 that binds Hypermethylated lysine 4 of histone H3 is necessary for efficient antigen-receptor-gene rearrangement. Immunity 27:561–571
Musselman CA et al (2012) Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1. Nat Struct Mol Biol 19:1266–1272
Fiedler M et al (2008) Decoding of methylated histone H3 tail by the Pygo-BCL9 Wnt signaling complex. Mol Cell 30:507–518
Huang Y, Fang J, Bedford MT, Zhang Y, Xu RM (2006) Recognition of histone H3 lysine-4 methylation by the double tudor domain of JMJD2A. Science 312:748–751
Lee J, Thompson JR, Botuyan MV, Mer G (2008) Distinct binding modes specify the recognition of methylated histones H3K4 and H4K20 by JMJD2A-tudor. Nat Struct Mol Biol 15:109–111
Brien GL et al (2012) Polycomb PHF19 binds H3K36me3 and recruits PRC2 and demethylase NO66 to embryonic stem cell genes during differentiation. Nat Struct Mol Biol 19:1273–1281
Ballare C et al (2012) Phf19 links methylated Lys36 of histone H3 to regulation of polycomb activity. Nat Struct Mol Biol 19:1257–1265
Collins RE et al (2008) The ankyrin repeats of G9a and GLP histone methyltransferases are mono- and dimethyllysine binding modules. Nat Struct Mol Biol 15:245–250
Badeaux AI et al (2012) Loss of the methyl lysine effector protein PHF20 impacts the expression of genes regulated by the lysine acetyltransferase MOF. J Biol Chem 287:429–437
Hirano Y et al (2012) Lamin B receptor recognizes specific modifications of histone H4 in heterochromatin formation. J Biol Chem 287:42654–42663
He F et al (2010) Structural insight into the zinc finger CW domain as a histone modification reader. Structure 18:1127–1139
Liu Y et al (2016) Family-wide characterization of histone binding abilities of human CW domain-containing proteins. J Biol Chem 291:9000–9013
Bian C et al (2011) Sgf29 binds histone H3K4me2/3 and is required for SAGA complex recruitment and histone H3 acetylation. EMBO J 30:2829–2842
Yang Y et al (2010) TDRD3 is an effector molecule for arginine-methylated histone marks. Mol Cell 40:1016–1023
Wang W et al (2011) Nucleolar protein Spindlin1 recognizes H3K4 methylation and stimulates the expression of rRNA genes. EMBO Rep 12:1160–1166
Kuo AJ et al (2012) The BAH domain of ORC1 links H4K20me2 to DNA replication licensing and Meier-Gorlin syndrome. Nature 484:115–119
Zhao D et al (2016) The BAH domain of BAHD1 is a histone H3K27me3 reader. Protein Cell 7:222–226
Trojer P et al (2007) L3MBTL1, a histone-methylation-dependent chromatin lock. Cell 129:915–928
Guo Y et al (2009) Methylation-state-specific recognition of histones by the MBT repeat protein L3MBTL2. Nucleic Acids Res 37:2204–2210
Jacquet K et al (2016) The TIP60 complex regulates bivalent chromatin recognition by 53BP1 through direct H4K20me binding and H2AK15 acetylation. Mol Cell 62:409–421
Maurer-Stroh S et al (2003) The Tudor domain ‘Royal Family’: Tudor, plant agenet, chromo, PWWP and MBT domains. Trends Biochem Sci 28:69–74
Yap KL, Zhou MM (2010) Keeping it in the family: diverse histone recognition by conserved structural folds. Crit Rev Biochem Mol Biol 45:488–505
Yap KL, Zhou MM (2011) Structure and mechanisms of lysine methylation recognition by the chromodomain in gene transcription. Biochemistry 50:1966–1980
Nielsen PR et al (2002) Structure of the HP1 chromodomain bound to histone H3 methylated at lysine 9. Nature 416:103–107
Jacobs SA, Khorasanizadeh S (2002) Structure of HP1 chromodomain bound to a lysine 9-methylated histone H3 tail. Science 295:2080–2083
Chang Y, Horton JR, Bedford MT, Zhang X, Cheng X (2011) Structural insights for MPP8 chromodomain interaction with histone H3 lysine 9: potential effect of phosphorylation on methyl-lysine binding. J Mol Biol 408:807–814
Li J et al (2011) Structural basis for specific binding of human MPP8 chromodomain to histone H3 methylated at lysine 9. PLoS One 6:e25104
Fischle W et al (2003) Molecular basis for the discrimination of repressive methyl-lysine marks in histone H3 by Polycomb and HP1 chromodomains. Genes Dev 17:1870–1881
Min J, Zhang Y, Xu RM (2003) Structural basis for specific binding of Polycomb chromodomain to histone H3 methylated at Lys 27. Genes Dev 17:1823–1828
Pray-Grant MG, Daniel JA, Schieltz D, Yates JR 3rd, Grant PA (2005) Chd1 chromodomain links histone H3 methylation with SAGA- and SLIK-dependent acetylation. Nature 433:434–438
Flanagan JF et al (2005) Double chromodomains cooperate to recognize the methylated histone H3 tail. Nature 438:1181–1185
Kamps JJ et al (2015) Chemical basis for the recognition of trimethyllysine by epigenetic reader proteins. Nat Commun 6:8911
Nady N et al (2012) Histone recognition by human malignant brain tumor domains. J Mol Biol 423:702–718
Li H et al (2007) Structural basis for lower lysine methylation state-specific readout by MBT repeats of L3MBTL1 and an engineered PHD finger. Mol Cell 28:677–691
Min J et al (2007) L3MBTL1 recognition of mono- and dimethylated histones. Nat Struct Mol Biol 14:1229–1230
Qin S, Min J (2014) Structure and function of the nucleosome-binding PWWP domain. Trends Biochem Sci 39:536–547
Wang Y et al (2009) Regulation of Set9-mediated H4K20 methylation by a PWWP domain protein. Mol Cell 33:428–437
Baubec T et al (2015) Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520:243–247
Sanchez R, Zhou MM (2011) The PHD finger: a versatile epigenome reader. Trends Biochem Sci 36:364–372
Pena PV et al (2006) Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 442:100–103
Wysocka J et al (2005) WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121:859–872
Gayatri S, Bedford MT (2014) Readers of histone methylarginine marks. Biochim Biophys Acta 1839:702–710
Liu K et al (2010) Structural basis for recognition of arginine methylated Piwi proteins by the extended Tudor domain. Proc Natl Acad Sci U S A 107:18398–18403
Sikorsky T et al (2012) Recognition of asymmetrically dimethylated arginine by TDRD3. Nucleic Acids Res 40:11748–11755
Nair SS et al (2010) PELP1 is a reader of histone H3 methylation that facilitates oestrogen receptor-alpha target gene activation by regulating lysine demethylase 1 specificity. EMBO Rep 11:438–444
Mann M, Cortez V, Vadlamudi R (2013) PELP1 oncogenic functions involve CARM1 regulation. Carcinogenesis 34:1468–1475
Hirota T, Lipp JJ, Toh BH, Peters JM (2005) Histone H3 serine 10 phosphorylation by Aurora B causes HP1 dissociation from heterochromatin. Nature 438:1176–1180
Fischle W et al (2005) Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438:1116–1122
Yuan CC et al (2012) Histone H3R2 symmetric dimethylation and histone H3K4 trimethylation are tightly correlated in eukaryotic genomes. Cell Rep 1:83–90
Yang L et al (2011) ncRNA- and Pc2 methylation-dependent gene relocation between nuclear structures mediates gene activation programs. Cell 147:773–788
Arrowsmith CH, Bountra C, Fish PV, Lee K, Schapira M (2012) Epigenetic protein families: a new frontier for drug discovery. Nat Rev Drug Discov 11:384–400
Acknowledgements
This work was supported by grant from the National Cancer Institute (R01CA172774) to Jia Fang. This work was also supported in part by Core Facilities at the H. Lee Moffitt Cancer Center & Research Institute, an NCI designated Comprehensive Cancer Center.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Sun, L., Fang, J. (2017). The Molecular Basis of Histone Methylation. In: Kaneda, A., Tsukada, Yi. (eds) DNA and Histone Methylation as Cancer Targets. Cancer Drug Discovery and Development. Humana Press, Cham. https://doi.org/10.1007/978-3-319-59786-7_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-59786-7_6
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-59784-3
Online ISBN: 978-3-319-59786-7
eBook Packages: MedicineMedicine (R0)