Abstract
Arrestin-3 (β-arrestin2) is the second member of the non-visual arrestin subfamily cloned. Together with arrestin-2 (β-arrestin1), it is responsible for arrestin function outside the visual system. Arrestins 2 and 3 are multi-functional proteins, originally identified as important for termination of signaling through G protein-coupled receptors (GPCRs), but now recognized as versatile regulators of signaling and trafficking of most GPCRs, as well as signaling switches controlling the balance between G protein-dependent and -independent GPCR signaling. Arrestin-3 is notably the least selective arrestin family member in that it binds a wide variety of GPCRs and that binding is less dependent on receptor activation and phosphorylation than for other arrestin subtypes. A recently determined arrestin-3 structure reveals that the basal conformation of arrestin-3 is similar to other arrestins, and that similar molecular interactions stabilize the basal state in all arrestins. A disruption of a structural element in the C-domain of arrestin-3 has been implicated as the cause of the reduced selectivity of arrestin-3. Here we compare the basal arrestin-3 structure with the recently determined structure of an activated receptor-bound arrestin-1.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
The four arrestin subtypes expressed in mammals can be broadly classified into visual and non-visual groups (Gurevich and Gurevich 2006a). The visual arrestins, arrestin-1 (rod arrestin) and arrestin-4 (cone arrestin) are expressed only in photoreceptors and bind rod and cone opsins (Strissel et al. 2006; Hanson et al. 2007; Nikonov et al. 2008). In contrast, arrestin-2 and arrestin-3 (also referred to as β-arrestin1 and β-arrestin2, respectively) are present in most cells, and interact with hundreds of G protein-coupled receptors (GPCRs) (Gurevich and Gurevich 2006b). In most cell types arrestin-2 is the major arrestin family member, being more abundant than arrestin-3 by ~10–20 fold (Gurevich et al. 2002, 2004). In addition to their roles in terminating GPCR signaling, arrestins 2 and 3 also serve as clathrin adaptors, thereby controlling GPCR endocytosis (Laporte et al. 1999; Oakley et al. 1999; Goodman et al. 1996), and as signaling hubs, where they scaffold and direct kinase networks, and as mediating ubiquitination of receptors and other signaling molecules [reviewed in Gurevich and Gurevich (2006b), DeWire et al. (2007), Jean-Charles et al. (2016)]. A key functional difference between arrestin-3 and other family members is its relative receptor promiscuity. Binding studies comparing different arrestins to different functional states of their cognate receptors (active phosphorylated, inactive phosphorylated, active unphosphorylated, and inactive unphosphorylated) revealed arrestin 3 to be significantly less specific than other arrestins . Arrestin-1 is the most specific and shows a >10 fold preference for active, phosphorylated, receptor over inactive, phosphorylated, receptor (Gurevich et al. 1993; Gurevich 1998). Both arrestin-2 and arrestin-3 are less selective , and show an ~2 fold preference for active, phosphorylated, receptor over inactive, phosphorylated, receptor, with arrestin-3 being the least selective (Gurevich et al. 1995). The requirement for receptor phosphorylation can be more effectively relieved through mutagenesis in arrestin-3 than in other arrestins , further supporting the conclusion that arrestin-3 is the most promiscuous arrestin (Gurevich and Gurevich 2006b).
In addition to being more promiscuous, arrestin-3 generally shows higher affinity for GPCRs, higher affinity for clathrin, as well as unique kinase scaffolding activity (Gurevich and Gurevich 2006b; Goodman et al. 1996; DeWire et al. 2007; Kohout et al. 2001; Oakley et al. 2000). Importantly, only arrestin-3, but not highly homologous arrestin-2, facilitates the activation of JNK family kinases (McDonald et al. 2000; Song et al. 2009a; Miller et al. 2001; Breitman et al. 2012; Zhan et al. 2011a, 2013, 2016; Kook et al. 2013). Despite these unique features, the two proteins are partially redundant because knockout a either non-visual arrestin is tolerated, but the double knockout is embryonic lethal (Kohout et al. 2001).
Arrestin 3 was the final isoform to have its structre reported, with arrestin 1, 2, and 4 published previously (Hirsch et al. 1999; Han et al. 2001; Kang et al. 2009; Milano et al. 2002, 2006; Sutton et al. 2005). Here we review the arrestin-3 structure, with the added benefit of comparison to the recent structure of the rhodopsin-arrestin-1 complex, which offers the first insight into an activated receptor-bound arrestin. The overall fold of arrestin-3 resembles that of the other family members, Fig. 5.1. The key features known to stabilize the basal state, the three-element interaction and the polar core, are present in the now expected form (Fig. 5.2), but the structure identified certain peculiarities characteristic for this subtype.
Structurally important features are conserved in Arrestin 3. a Arrestin 3 with the conserved three-element interaction highlighted as a Van der Waals mesh and the polar core side chains rendered as stick figures. Additionally, the disrupted beta strand in the C-domain of Arrestin 3 is colored red. b and c Expanded overlays of Arrestins 1, 2, and 3, colored as in Fig. 5.1 and rendered from the coordinates listed in Fig. 5.1. Both views are rotated ~90° toward the reader, about a horizontal axis, relative to the view in a. b Conserved three-element interaction within three arrestins . c Near perfect conservation of the polar core among arrestins 1, 2, and 3. b Outermost strand of the arrestin 3 C-domain is also distorted relative to arrestin 2. The disruption is small, ~2 Å, but it disrupts hydrogen bonding to the adjacent strand and likely reduces the energy barrier to activation
Arrestin-3 Structure
As expected, the arrestin-3 adopts a canonical arrestin fold (Figs. 5.1 and 5.2). It is composed of structurally homologous domains, the N-terminal and C-terminal domains, each of which is formed by a 7-stranded beta sandwich. The two domains are connected by a flexible linker and stabilized by inter-domain contacts. The extreme carboxy terminus traverses the C-domain to form critical interactions with the N-domain (three-element interaction, Fig. 5.2b). The last residue visible in the arrestin-3 structure is the arginine in the C-terminus that contributes a salt bridge to a critical functional component, the phosphate sensor (polar core, Fig. 5.2c). Additionally, three residues on this C-terminal peptide form part of the three-element interaction.
Intermolecular interactions hold arrestins in their basal conformations. Two sets of interactions known to be critical are the three-element interaction and the polar core (Fig. 5.2). Disruption of either of these interactions in known to reduce fidelity in the arrestin-receptor interaction, likely by decreasing the activation barrier for the transition to the activated state (Gurevich and Gurevich 2006b; Kovoor et al. 1999; Celver et al. 2002; Song et al. 2009b; Gurevich et al. 2011). The arrestin-3 structure confirmed that alterations to the polar core are not responsible for the increased promiscuity of arrestin-3. The polar core in arrestin-3 includes identical residues in nearly identical orientations to those observed in arrestin-1 and arrestin-2 (Fig. 5.2c). The polar core is essentially unchanged in arrestin-3, indicating that a different structural feature is responsible for the unique functional characteristics of arrestin-3.
The three-element interaction, although somewhat less well studied, is thought to serve as a molecular clamp that prevents release of the C-tail (Gurevich 1998; Vishnivetskiy et al. 2000). Release of the C-tail is a necessary step in arrestin activation (Palczewski et al. 1991; Kang et al. 2015; Shukla et al. 2013). In arrestin-3 , the three-element interaction is present, but it is altered, as compared to arrestin-1 in three ways. The interaction involves three hydrophobic residues on each of three different secondary structural elements: β-strand I contributes Val8, Ile9, and Phe10; α-helix I contributes Leu101, Leu105, and Leu109; and the C-tail contributes Phe386, Val387, and Phe388 (bovine arrestin-1 numbering). In arrestin-3, Val8 is replaced with an Arg, Ile9 is replaced with a Val, and Phe386 is replaced with an Ile (Fig. 5.2b). The possibility that these mutations may result in the C-tail being more weakly held in arrestin-3 than in arrestin-1 has not been tested experimentally. In any event, this change is likely not responsible for the entirety of the differences between the arrestins because arrestin-2 and arrestin-3 have identical three-element interactions (Fig. 5.2b), indicating another structural features must underlie the functional differences between the two non-visual arrestins.
An additional difference between arrestin-3 and arrestin-2 is a disruption in the packing of the C-domain. Residues 257–260, which form part of the outer most “strand” in the C-domain, are displaced relative to arrestin-2. The displacements are relatively small (1.3–2.0 Å), but are large enough to disrupt the β-sheet hydrogen bonding pattern, resulting in the loss of 2 hydrogen bonds (Zhan et al. 2011b). Despite the relatively small size of the distortion, it seems important for three reasons: it is substantially larger than the mean coordinate error of the structure; it occurs in a region known to be important for receptor binding; and the C-domain (including this strand) is essentially invariant in all arrestin structures (Hirsch et al. 1999; Han et al. 2001; Kang et al. 2009; Milano et al. 2002, 2006; Zhan et al. 2011b; Vishnivetskiy et al. 2004).
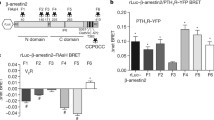
The importance of this distortion was tested by exchanging the strand with the homologous strand from arrestin-2. A bioluminescence resonance energy transfer-based arrestin recruitment experiment designed to determine the fraction of receptor-bound arrestin in the cell revealed that substitution of the distorted strand into arrestin-2 conferred arrestin-3 like features on arrestin-2 (Zhan et al. 2011b). β2AR binding of these chimeras and the wild type arrestins was compared in an arrestin recruitment assay (Zhan et al. 2011b). This assay revealed that distortion of the C-domain sheet promoted binding to inactive and active β2AR (Zhan et al. 2011b), a hallmark feature of arrestin-3 (Gurevich et al. 1995; Kovoor et al. 1999; Celver et al. 2002). Conversely, the introduction of the arrestin-2 residues into arrestin-3 reduced binding to unstimulated β2AR (Zhan et al. 2011b).
Comparisons to the Rhodopsin-Arrestin-1 Complex
The recently determined structure of rhodopsin bound to arrestin-1 (Kang et al. 2015) allows analysis of how differences in the three-element interaction and the disrupted β-strand in the C-domain might be important for the conformational changes associated with receptor binding . Figure 5.3 shows a comparison of receptor-bound arrestin-1 and the basal structure of arrestin-3. The most significant changes are the re-structuring of the finger loop (residues 64–77), the release of the C-tail from the N-domain, and the rotation of the C-domain relative to the N-domain. The finger loop intercalates into the rhodopsin, fitting into the cavity between the cytoplasmic ends of the transmemebrane helices that opens upon GPCR activation (Farrens et al. 1996). This interaction, however, does not appear to require a substantial conformation change on the part of arrestin . The finger loop is the loop between the 4th and 5th strands of the N-domain and is structurally variable in arrestins. An interdomain conformational change is not likely required for it to adopt the active conformation.
Arrestin 3 is configured to transition to the receptor bound state. a Structural rearrangements that occur upon receptor binding. Arrestin 1 shown in salmon undergoes significant conformational changes upon rhodopsin (grey) binding (the arrestin-rhodopsin complex is rendered from (pdb 4zwj). The finger loop, highlighted with a black ellipse, undergoes a loop to helix transition and inserts into the helical bundle of the receptor. The C-domain rotates ~20° relative to the N-domain. The arrestin 3 structure, overlayed based on an alignment to the N-domain, is shown in blue, with the outermost strand of the C-domain colored red. The homologous strand in arrestin 1 undergoes an ~9 Å displacement upon activation, as shown in b, in which the view is rotated 90° toward the reader about a vertical axis c an overlay of arrestin-1, yellow, with arrestin-3, blue, showing the connection between the finger loop and the loosely packed outer strand in arrestin-3. Loose packing of this strand may accomadate the conformational change associated with activation
The rotation of the C-domain by ~20°, in contrast, requires a complete reorganization of the interdomain interface and a coincident translation of the beta-sandwich. In arrestin-1, this reorganization results in a repacking of the outermost strand, such that it forms beta strand hydrogen bonds with the neighboring strand for only 7 residues, as compared to 11 residues in the basal state. If arrestin-3 undergoes a similar conformational change upon receptor binding, the loose packing of this strand in the basal state may better accommodate the conformational change associated with activation (Fig. 5.3c). This would explain apparently lower energy barrier, which results in high binding to inactive phosphorylated receptor and wide variety of GPCRs.
References
Breitman M, Kook S, Gimenez LE, Lizama BN, Palazzo MC, Gurevich EV, Gurevich VV (2012) Silent scaffolds: inhibition of c-Jun N-terminal kinase 3 activity in cell by dominant-negative arrestin-3 mutant. J Biol Chem 287:19653–19664
Celver J, Vishnivetskiy SA, Chavkin C, Gurevich VV (2002) Conservation of the phosphate-sensitive elements in the arrestin family of proteins. J Biol Chem 277:9043–9048
DeWire SM, Ahn S, Lefkowitz RJ, Shenoy SK (2007) β-arrestins and cell signaling. Annu Rev Physiol 69:483–510
Farrens DL, Altenbach C, Yang K, Hubbell WL, Khorana HG (1996) Requirement of rigid-body motion of transmembrane helices for light activation of rhodopsin. Science 274:768–770
Goodman OB, Krupnick JG, Santini F, Gurevich VV, Penn RB, Gagnon AW, Keen JH, Benovic JL (1996) Beta-arrestin acts as a clathrin adaptor in endocytosis of the beta2-adrenergic receptor. Nature 383:447–450
Gurevich VV (1998) The selectivity of visual arrestin for light-activated phosphorhodopsin is controlled by multiple nonredundant mechanisms. J Biol Chem 273:15501–15506
Gurevich EV, Gurevich VV (2006a) Arrestins: ubiquitous regulators of cellular signaling pathways. Genome Biol 7:236
Gurevich VV, Gurevich EV (2006b) The structural basis of arrestin-mediated regulation of G-protein-coupled receptors. Pharmacol Ther 110:465–502
Gurevich VV, Richardson RM, Kim CM, Hosey MM, Benovic JL (1993) Binding of wild type and chimeric arrestins to the m2 muscarinic cholinergic receptor. J Biol Chem 268:16879–16882
Gurevich VV, Dion SB, Onorato JJ, Ptasienski J, Kim CM, Sterne-Marr R, Hosey MM, Benovic JL (1995) Arrestin interactions with G protein-coupled receptors. Direct binding studies of wild type and mutant arrestins with rhodopsin, beta 2-adrenergic, and m2 muscarinic cholinergic receptors. J Biol Chem 270:720–731
Gurevich EV, Benovic JL, Gurevich VV (2002) Arrestin2 and arrestin3 are differentially expressed in the rat brain during postnatal development. Neuroscience 109:421–436
Gurevich EV, Benovic JL, Gurevich VV (2004) Arrestin2 expression selectively increases during neural differentiation. J Neurochem 91:1404–1416
Gurevich VV, Hanson SM, Song X, Vishnivetskiy SA, Gurevich EV (2011) The functional cycle of visual arrestins in photoreceptor cells. Prog Retin Eye Res 30:405–430
Han M, Gurevich VV, Vishnivetskiy SA, Sigler PB, Schubert C (2001) Crystal structure of beta-arrestin at 1.9 A: possible mechanism of receptor binding and membrane translocation. Structure 9:869–880
Hanson SM, Gurevich EV, Vishnivetskiy SA, Ahmed MR, Song X, Gurevich VV (2007) Each rhodopsin molecule binds its own arrestin. Proc Natl Acad Sci U S A 104:3125–3128
Hirsch JA, Schubert C, Gurevich VV, Sigler PB (1999) The 2.8 A crystal structure of visual arrestin: a model for arrestin’s regulation. Cell 97:257–269
Jean-Charles P-Y, Rajiv V, Shenoy SK (2016) Ubiquitin-related roles of β-arrestins in endocytic trafficking and signal transduction. J Cell Physiol 231:2071–2080
Kang DS, Kern RC, Puthenveedu MA, von Zastrow M, Williams JC, Benovic JL (2009) Structure of an arrestin2-clathrin complex reveals a novel clathrin binding domain that modulates receptor trafficking. J Biol Chem 284:29860–29872
Kang Y, Zhou XE, Gao X, He Y, Liu W, Ishchenko A, Barty A, White TA, Yefanov O, Han GW, Xu Q, de Waal PW, Ke J, Tan MHE, Zhang C, Moeller A, West GM, Pascal BD, Van Eps N, Caro LN, Vishnivetskiy SA, Lee RJ, Suino-Powell KM, Gu X, Pal K, Ma J, Zhi X, Boutet S, Williams GJ, Messerschmidt M, Gati C, Zatsepin NA, Wang D, James D, Basu S, Roy-Chowdhury S, Conrad CE, Coe J, Liu H, Lisova S, Kupitz C, Grotjohann I, Fromme R, Jiang Y, Tan M, Yang H, Li J, Wang M, Zheng Z, Li D, Howe N, Zhao Y, Standfuss J, Diederichs K, Dong Y, Potter CS, Carragher B, Caffrey M, Jiang H, Chapman HN, Spence JCH, Fromme P, Weierstall U, Ernst OP, Katritch V, Gurevich VV, Griffin PR, Hubbell WL, Stevens RC, Cherezov V, Melcher K, Xu HE (2015) Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser. Nature 523:561–567
Kohout TA, Lin FS, Perry SJ, Conner DA, Lefkowitz RJ (2001) Beta-arrestin 1 and 2 differentially regulate heptahelical receptor signaling and trafficking. Proc Natl Acad Sci U S A 98:1601–1606
Kook S, Zhan X, Kaoud TS, Dalby KN, Gurevich VV, Gurevich EV (2013) Arrestin-3 binds c-Jun N-terminal kinase 1 (JNK1) and JNK2 and facilitates the activation of these ubiquitous JNK isoforms in cells via scaffolding. J Biol Chem 288:37332–37342
Kovoor A, Celver J, Abdryashitov RI, Chavkin C, Gurevich VV (1999) Targeted construction of phosphorylation-independent beta-arrestin mutants with constitutive activity in cells. J Biol Chem 274:6831–6834
Laporte SA, Oakley RH, Zhang J, Holt JA, Ferguson SSG, Caron MG, Barak LS (1999) The β2-adrenergic receptor/βarrestin complex recruits the clathrin adaptor AP-2 during endocytosis. Proc Natl Acad Sci USA 96:3712–3717
McDonald PH, Chow CW, Miller WE, Laporte SA, Field ME, Lin FT, Davis RJ, Lefkowitz RJ (2000) Beta-arrestin 2: a receptor-regulated MAPK scaffold for the activation of JNK3. Science 290:1574–1577
Milano SK, Pace HC, Kim Y-M, Brenner C, Benovic JL (2002) Scaffolding functions of arrestin-2 revealed by crystal structure and mutagenesis. Biochemistry 41:3321–3328
Milano SK, Kim Y-M, Stefano FP, Benovic JL, Brenner C (2006) Nonvisual arrestin oligomerization and cellular localization are regulated by inositol hexakisphosphate binding. J Biol Chem 281:9812–9823
Miller WE, McDonald PH, Cai SF, Field ME, Davis RJ, Lefkowitz RJ (2001) Identification of a motif in the carboxyl terminus of beta -arrestin2 responsible for activation of JNK3. J Biol Chem 276:27770–27777
Nikonov SS, Brown BM, Davis JA, Zuniga FI, Bragin A, Pugh EN, Craft CM (2008) Mouse cones require an arrestin for normal inactivation of phototransduction. Neuron 59:462–474
Oakley RH, Laporte SA, Holt JA, Barak LS, Caron MG (1999) Association of beta-arrestin with G protein-coupled receptors during clathrin-mediated endocytosis dictates the profile of receptor resensitization. J Biol Chem 274:32248–32257
Oakley RH, Laporte SA, Holt JA, Caron MG, Barak LS (2000) Differential affinities of visual arrestin, beta arrestin1, and beta arrestin2 for G protein-coupled receptors delineate two major classes of receptors. J Biol Chem 275:17201–17210
Palczewski K, Pulvermüller A, Buczyłko J, Hofmann KP (1991) Phosphorylated rhodopsin and heparin induce similar conformational changes in arrestin. J Biol Chem 266:18649–18654
Shukla AK, Manglik A, Kruse AC, Xiao K, Reis RI, Tseng W-C, Staus DP, Hilger D, Uysal S, Huang L-Y, Paduch M, Tripathi-Shukla P, Koide A, Koide S, Weis WI, Kossiakoff AA, Kobilka BK, Lefkowitz RJ (2013) Structure of active β-arrestin-1 bound to a G-protein-coupled receptor phosphopeptide. Nature 497:137–141
Song X, Coffa S, Fu H, Gurevich V (2009a) How does arrestin assemble MAP kinases into a signaling complex? J Biol Chem
Song X, Vishnivetskiy SA, Gross OP, Emelianoff K, Mendez A, Chen J, Gurevich EV, Burns ME, Gurevich VV (2009b) Enhanced arrestin facilitates recovery and protects rods lacking rhodopsin phosphorylation. Curr Biol 19:700–705
Strissel KJ, Sokolov M, Trieu LH, Arshavsky VY (2006) Arrestin translocation is induced at a critical threshold of visual signaling and is superstoichiometric to bleached rhodopsin. J Neurosci 26:1146–1153
Sutton RB, Vishnivetskiy SA, Robert J, Hanson SM, Raman D, Knox BE, Kono M, Navarro J, Gurevich VV (2005) Crystal structure of cone arrestin at 2.3A: evolution of receptor specificity. J Mol Biol 354:1069–1080
Vishnivetskiy SA, Schubert C, Climaco GC, Gurevich YV, Velez MG, Gurevich VV (2000) An additional phosphate-binding element in arrestin molecule. Implications for the mechanism of arrestin activation. J Biol Chem 275:41049–41057
Vishnivetskiy SA, Hosey MM, Benovic JL, Gurevich VV (2004) Mapping the arrestin-receptor interface. Structural elements responsible for receptor specificity of arrestin proteins. J Biol Chem 279:1262–1268
Zhan X, Kaoud TS, Dalby KN, Gurevich VV (2011a) Nonvisual arrestins function as simple scaffolds assembling the MKK4-JNK3α2 signaling complex. Biochemistry 50:10520–10529
Zhan X, Gimenez LE, Gurevich VV, Spiller BW (2011b) Crystal structure of arrestin-3 reveals the basis of the difference in receptor binding between two non-visual subtypes. J Mol Biol 406:467–478
Zhan X, Kaoud TS, Kook S, Dalby KN, Gurevich VV (2013) JNK3 enzyme binding to arrestin-3 differentially affects the recruitment of upstream mitogen-activated protein (MAP) kinase kinases. J Biol Chem 288:28535–28547
Zhan X, Stoy H, Kaoud TS, Perry NA, Chen Q, Perez A, Els-Heindl S, Slagis JV, Iverson TM, Beck-Sickinger AG, Gurevich EV, Dalby KN, Gurevich VV (2016) Peptide mini-scaffold facilitates JNK3 activation in cells. Nat Publ Gr 6:21025
Acknowledgements
This review is dedicated to the memory of Carsten Schubert. Carsten contributed extensively to the arrestin 1 and arrestin 2 structures. He was helpful and generous with his ideas and time. He passed away on his 51st birthday. May his memory be a comfort.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Spiller, B.W., Zhan, X., Gurevich, V.V. (2017). Arrestin-3: The Structural Basis of Lower Receptor Selectivity. In: Gurevich, V. (eds) The Structural Basis of Arrestin Functions. Springer, Cham. https://doi.org/10.1007/978-3-319-57553-7_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-57553-7_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-57552-0
Online ISBN: 978-3-319-57553-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)