Abstract
We have established a novel in situ protein analysis pipeline, which is built upon highly sensitive, multichannel immunofluorescent staining of paraffin sections of human and xenografted tumor tissue. Specimens are digitized using slide scanners equipped with suitable light sources and fluorescence filter combinations. Resulting digital images are subsequently subjected to quantitative image analysis using a primarily object-based approach, which comprises segmentation of single cells or higher-order structures (e.g., blood vessels), cell shape approximation, measurement of signal intensities in individual fluorescent channels and correlation of these data with positional information for each object. Our approach could be particularly useful for the study of the hypoxic tumor microenvironment as it can be utilized to systematically explore the influence of spatial factors on cell phenotypes, e.g., the distance of a given cell type from the nearest blood vessel on the cellular expression of hypoxia-associated biomarkers and other proteins reflecting their specific state of activation or function. In this report, we outline the basic methodology and provide an outlook on possible use cases.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Tumor microenvironment
- Tumor hypoxia
- Multiparametric image analysis
- Immunofluorescence
- Immunohistochemistry
1 Introduction
A thorough characterization of disease mechanisms (e.g., the relevance of certain driver mutations) is an absolute requirement for the success of targeted strategies in cancer therapy. In a series of publications from the Cancer Genome Atlas Research Network (TCGA, e.g., [1]) this has been achieved using modern genome analysis methods . As an additional layer of complexity, however, solid malignant tumors are exceptionally heterogeneous, leading to a situation where a tailored therapeutic approach may be successful in some tumor subvolumes but ineffective in others. An important part of this tumor heterogeneity is caused by the pathological architecture of tumor microvessels and enlarged diffusion distances leading to hypoxia, nutrient deprivation, extracellular acidosis and metabolic waste product accumulation. This characteristic solid tumor microenvironment brings along substantial changes in gene expression, the proteome and a resultant phenotype of tumor cells, which may mediate resistance not only against ionizing irradiation and cytotoxic chemotherapy but also against molecularly targeted and immune-stimulatory approaches. In order to develop successful strategies to overcome this major obstacle against cancer therapy, an ex vivo analysis of the microenvironment of human tumors is necessary both to select patients for intensified or de-escalated treatment protocols and to develop specific countermeasures. A method suitable to contribute to the accomplishment of this goal is outlined in this report.
2 Dimensions of Tumor Heterogeneity
Tumor heterogeneity exists at least at three levels: (i) genetic or clonal, (ii) cellular and (iii) architectural. Clonal heterogeneity , the result of an evolutionary process originally outlined for tumor cell populations by Peter Nowell [2], has now been unequivocally proven to exist using a combination of whole-exome multiregion spatial sequencing, single nucleotide polymorphism analysis and messenger RNA expression profiling [3]. Diverging genomic constitution of individual manifestations of a tumor within a single patient, i.e., the primary tumor and metastases in different organs, can indeed partially explain incomplete or mixed responses to therapy, which are often observed in the clinic. A prominent example is the development of T790 M resistance mutations against small molecule inhibition of EGFR in lung cancer [4]. Some manifestations of therapeutic resistance are, however, not explained by genomic maladaptation. Instead, they have been shown to arise from functional mechanisms. The term cellular heterogeneity refers to the fact that solid malignant tumors are complex tissues composed of many different cell types, which have been demonstrated to actively contribute to tumor growth instead of acting as a passive barrier to cancer cell invasion or as innocent bystanders. Among these cancer promoting cells are activated fibroblasts (“cancer-associated fibroblasts”, [5]), endothelial cells [6], macrophages [7] and regulatory T-cells [8]. Contrary to the depiction of malignant tumors in many textbooks and review articles as rather homogeneous masses of tumor cells (e.g., [9]), the true cellular complexity of solid malignancies is immediately apparent under the microscope and thus well known to the pathologist, but surprisingly less appreciated by many other oncology disciplines. Finally, architectural heterogeneity refers to the different degree of abnormalities of higher-order structures. A prominent example of particular relevance to the topic of this communication is the different degree of the abnormality of the tumor blood vessels. It ranges from greatly increased vascular density in isolated “hotspots” to large, almost avascular areas in which substantial hypoxia develops and from nearly normal configuration of individual tumor microvessels to highly irregular arrangement, orientation and ultrastructure. Many other important aspects also fall in this category, including – but not limited – to the individual hierarchy and localization of the stem cell compartment, patterns of cancer cell invasion (e.g., pushing borders vs. infiltrating single cells) and the degree of immune cell infiltration and activity. Both cellular and architectural heterogeneity can be assessed using morphometric and immunohistochemical methods, e.g., by identifying certain cell types using antibodies against characteristic cell surface markers. In fact, these methods have already experienced broad application in clinically oriented basic oncology research. However, the definition of cellular phenotypes, the identification of activation states and the exploration of spatial patterns of different cell types relative to each other or in relation to higher order structures of the tissue has been severely hampered by the fact that multiplex staining, although certainly possible [10, 11], has always been notoriously difficult to achieve with traditional immunohistochemical methods and even harder to standardize for large research projects. We have thus developed novel approaches to overcome traditional obstacles and enable a truly multiparametric analysis of the tumor microenvironment.
3 Multiparametric Immunohistochemistry in Registered Serial Sections (MIRSS)
High precision microtomes (in the hands of experienced technicians) enable the preparation of very thin (i.e., 3–5 μm) serial paraffin sections as raw material for immunohistochemistry (IHC). When combining this technique with digitization of specimens using high resolution slide scanners and advanced elastic image registration algorithms (e.g., [12]), we have shown that it is possible to follow individual cells through several serial sections and use different IHC markers for each (Fig. 14.1). We have recently demonstrated the feasibility and relevance of this approach in an analysis of the expression of hypoxia-associated markers in squamous cell carcinomas of the vulva, which provided important insights into the pathophysiology of this disease [13]. Although fascinating, this technique has a number of weaknesses, which have to be taken into account. First, the method is quite labor intensive when manual staining is performed. Furthermore, high precision sectioning can be very demanding and sometimes even impossible in “difficult” tissue leading to artefacts which thwart successful image registration. Even though individual sections are thin, a substantial number of sections are needed for multiplex staining, and this can be a problem when only limited amounts of tissue are available. Most importantly, however, segmentation of individual cells, as the prerequisite of single-cell based analyses , turned out to be quite challenging in some types of tissue. Methodological work from our laboratory showed that segmentation of single cells using fluorescence staining of cell nuclei, although far from trivial, provides much more reproducible results.
Multiparametric Immunohistochemistry in Registered Serial Sections (MIRSS). Examples from squamous cell carcinoma of the vulva stained for (a) Ki67, (b) GLUT-1, (c) CXCL-12/SDF-1, (d) CXCR-4, (e) CA IX, (f) CD 34, (g) podoplanin and (h) αSMA. Image registration has been achieved down to the level of individual cells, which can be traced through consecutive slices
4 Multichannel Immunofluorescence Staining Combined with a Quantitative Single-Cell Based Analysis
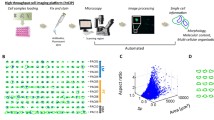
In 2007, Tóth and Mezey [14] demonstrated the feasibility of multiplex immunofluorescence staining in paraffin sections based on fluorochrome tyramide-mediated signal amplification and repeated cycles of primary antibody denaturation and peroxidase quenching by heat treatment. Among those who embraced this approach were researchers working with the multispectral imaging technology, which employs liquid crystal tuneable filters instead of conventional fluorescence filters. Multispectral imaging enables a high degree of multiplexing through linear unmixing of individual fluorochrome signatures from stacks of images, each representing only a small “window” of the entire wavelength spectrum (“lambda stacks”), albeit at the cost of a certain amount of sensitivity. However, the advantage of high signal strength resulting from tyramide amplification also benefits multichannel fluorescence staining of paraffin sections when using conventional filter technology. Most formalin-fixed and paraffin embedded tissue sections show at least some autofluorescence, which poses much less of a problem when the specific signal is very strong. In 2014, our laboratory started to systematically explore multiplex staining using the fluorochrome tyramide-based approach. The technology turned out to be stable and reproducible, compatible with most antibodies tested (although in our hands strictly limited to monoclonal antibodies) and sensitive [15]. Some drawbacks remain: multichannel immunofluorescence is still labor-intensive when performed as a manual procedure, as a four-plex stain takes two full working days to prepare. Furthermore, conventional fluorochrome tyramide staining sometimes lacks the utmost degree of sensitivity which can be achieved when DAB-based IHC is combined with tyramide-based signal ampflication. It may, however, be possible to overcome the latter problem, since (patented and commercially available) “enhanced tyramide” reagents, according to our preliminary testing, may eventually be as sensitive as the most sensitive conventional brightfield approaches. Cell segmentation, object identification, distance calculations and intensity measurements are carried out in the open source image analysis package CellProfiler [16], while data analysis in a manner similar to flow cytometry is carried out in a specialized commercial software (FCS Express Plus 5, De Novo Software, Glendale, CA, USA).
5 Exploring a New Paradigm of Hypoxia-Mediated Treatment Resistance
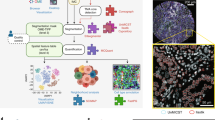
Based on published findings of downregulation of EGFR by the hypoxia-inducible PHD-3 [17, 18], we have applied the above-mentioned novel methodology to the study of a possible downregulation of EGFR by hypoxia in cancers of the head and neck region [19]. Indeed, downregulation of EGFR in diffusion-limited areas of the tumors, which often expressed the hypoxia-associated marker CA IX, was repeatedly found (Fig. 14.2). These data could form a new paradigm for the regulation of cancer-associated signalling pathways by the tumor microenvironment.
Multichannel immunofluorescence staining of a squamous cell carcinoma of the head and neck region stained for EGFR (red, Alexa 647), CA IX (green, FITC), CD 34 (white, Alexa 546) and cell nuclei (blue, DAPI). Preferential clustering of EGFR positive cells is observed in the vicinity of tumor blood vessels, while expression of EGFR is weak or absent in more distant, CA IX-positive tumor areas. Scale bar in the lower right corner, 500 μm
6 Conclusion
Using a newly developed combination of advanced immunofluorescence and image analysis methods, we are currently able to detect up to five different antigens in the same tumor section. These antigens may be utilized to select pivotal intratumoral structures (e.g., blood vessels) and hypoxic cell populations (e.g., tumor cells, stromal cells) by staining for suitable cell surface and hypoxia biomarkers and thus allow us to investigate the spatial relationships between these entities. Our methodology is of broad relevance and we have already successfully employed it to investigate possible mechanisms of resistance mediated by the tumor microenvironment, e.g., against anti-EGFR therapy.
References
Cancer Genome Atlas Network (2015) Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517(7536):576–582
Nowell PC (1976) The clonal evolution of tumor cell populations. Science 194(4260):23–28
Gerlinger M, Rowan AJ, Horswell S et al (2012) Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 366(10):883–892
Chen ZY, Zhong WZ, Zhang XC et al (2012) EGFR mutation heterogeneity and the mixed response to EGFR tyrosine kinase inhibitors of lung adenocarcinomas. Oncologist 17(7):978–985
Madar S, Goldstein I, Rotter V (2013) ‘Cancer associated fibroblasts’ – more than meets the eye. Trends Mol Med 19(8):447–453
Carmeliet P, Jain RK (2011) Molecular mechanisms and clinical applications of angiogenesis. Nature 473(7347):298–307
Ruffell B, Coussens LM (2015) Macrophages and therapeutic resistance in cancer. Cancer Cell 27(4):462–472
Zou W (2006) Regulatory T cells, tumour immunity and immunotherapy. Nat Rev Immunol 6(4):295–307
Talmadge JE, Fidler IJ (2010) AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res 70(14):5649–5669
van der Loos CM (1999) Immunoenzyme multiple staining methods. In: Royal Microscopical Society microscopy handbooks, vol 45. Bios Scientific Publishers, Oxford
Mayer A, Höckel M, Schlischewsky N et al (2013) Lacking hypoxia-mediated downregulation of E-cadherin in cancers of the uterine cervix. Br J Cancer 108(2):402–408
Arganda-Carreras I, Sorzano COS, Marabini R et al. (2006) Consistent and elastic registration of histological sections using vector-spline regularization. In: Beichel RR, Sonka M (eds) Computer vision approaches to medical image analysis: second international ECCV workshop, CVAMIA 2006 Graz, Austria, May 12, 2006 Revised papers. Springer, Berlin\Heidelberg, pp 85–95. doi:10.1007/11889762_8
Mayer A, Schmidt M, Seeger A et al (2014) GLUT-1 expression is largely unrelated to both hypoxia and the Warburg phenotype in squamous cell carcinomas of the vulva. BMC Cancer 14(1):1–9
Toth ZE, Mezey E (2007) Simultaneous visualization of multiple antigens with tyramide signal amplification using antibodies from the same species. J Histochem Cytochem 55(6):545–554
Mayer A, Brieger J, Vaupel P et al (2015) Multispectral, multiplexed analysis of the tumor microenvironment in FFPE tissue of head and neck cancer. Strahlenther Onkol 191:S45–S45
Lamprecht MR, Sabatini DM, Carpenter AE (2007) CellProfiler: free, versatile software for automated biological image analysis. BioTechniques 42(1):71–75
Henze AT, Garvalov BK, Seidel S et al (2014) Loss of PHD3 allows tumours to overcome hypoxic growth inhibition and sustain proliferation through EGFR. Nat Commun 5:5582
Garvalov BK, Foss F, Henze AT et al (2014) PHD3 regulates EGFR internalization and signalling in tumours. Nat Commun 5:5577
Mayer A, Zahnreich S, Brieger J et al (2016) Downregulation of EGFR in hypoxic, diffusion-limited areas of squamous cell carcinomas of the head and neck. Br J Cancer 115:1351-1358
Acknowledgment
The authors would like to thank Sysmex GmbH (Norderstedt, Germany) for providing access to slide scanners and scanning services.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Mayer, A., Vaupel, P. (2017). Multiparametric Analysis of the Tumor Microenvironment: Hypoxia Markers and Beyond. In: Halpern, H., LaManna, J., Harrison, D., Epel, B. (eds) Oxygen Transport to Tissue XXXIX. Advances in Experimental Medicine and Biology, vol 977. Springer, Cham. https://doi.org/10.1007/978-3-319-55231-6_14
Download citation
DOI: https://doi.org/10.1007/978-3-319-55231-6_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-55229-3
Online ISBN: 978-3-319-55231-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)