Abstract
To reduce healthcare cost while simultaneously delivering high-quality health services, developing new portable and/or implantable biomedical devices is of great importance for both health monitoring and clinical treatment. In this chapter, we describe a radically new framework for ultra-low-power biomedical circuit design and optimization. The proposed framework seamlessly integrates data processing algorithms and their customized circuit implementations for co-optimization. The efficacy of the proposed framework is demonstrated by a case study of brain–computer interface (BCI).
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
As reported by United Nations and US Census Bureau, the US population has enormously grown during the past several decades, climbing from 209 million in 1970 to 310 million in 2010. Most importantly, the percentage of senior citizens (more than 65-year old) is expected to reach 21.28% in 2050. With the rapid booming of senior citizen population, the expenditure of healthcare continuously increases at a rate of 5–10% per year in the USA. Such a trend is also observed worldwide over a large number of other countries.
To reduce healthcare cost while simultaneously delivering high-quality health services, developing new portable and/or implantable biomedical devices is of great importance. Billions of US dollars could be saved by reforming today’s healthcare infrastructure with these biomedical devices for various medical applications [8, 24]:
-
Health monitoring: Health condition should be reliably monitored for each person to predict and diagnose chronic diseases at the very early stage. For instance, ECG signals can be continuously measured and automatically classified by a portable biomedical device to diagnose arrhythmia [3, 19, 29].
-
Clinical treatment: Clinical therapy should be reliably delivered for each patient for both preventative care and disease treatment. Taking neuroprosthesis as an example, brain signals are sensed and decoded by an implantable device to control the prosthesis of a patient with neurological disorder [5, 11, 13, 21, 22, 26].
Towards these goals, miniaturized portable and/or implantable biomedical circuits must be designed and deployed to reliably sense, process, and transmit a large amount of physiological data with extremely low-power consumption. These circuits must carry several important “features”:
-
High accuracy: A biomedical device must accurately generate the desired output, such as diagnosis result for arrhythmia [3, 19, 29], and movement direction and velocity for neuroprosthesis [5, 11, 13, 21, 22, 26], that is not contaminated by artifacts, errors, and noises originated from human body and/or external environment [6, 17, 25].
-
Small latency: The response of a biomedical device must be sufficiently fast for a number of real-time applications such as vital sign monitoring [7, 31] and deep brain stimulation [9, 18]. In these cases, physiological data must be locally processed within the biomedical device to ensure fast response time, especially when a reliable wired or wireless communication channel is not available to transmit the data to an external device (e.g., smart phone, cloud server, etc.) for remote processing. Even in the cases where data transmission is possible such as neuroprosthesis control [5, 11, 13, 21, 22, 26], the raw data must be locally processed and compressed before transmission in order to minimize the communication energy.
-
Low power: To facilitate a portable and/or implantable device to continuously operate over a long time without recharging the battery, its power consumption must be minimized. Especially for the implantable applications where power consumption is highly constrained (e.g., less than 100 μW), it is necessary to design an application-specific circuit, instead of relying on general-purpose microprocessors, to meet the tight power budget [12, 15, 16, 27, 28, 30, 32].
-
Flexible reconfigurability: Reconfigurability is needed to customize a biomedical device for different patients and/or different usage scenarios. For instance, the movement decoder of neuroprosthesis should be retrained every day to accommodate the time-varying characteristics of neural sources, recording electrodes and environmental conditions [26]. It, in turn, requires a reconfigurable circuit implementation that can be customized every day.
The aforementioned features, however, are considered to be mutually exclusive today. Taking neuroprosthesis as an example, executing a sophisticated movement decoding algorithm is overly power hungry for portable and/or implantable applications. For this reason, renovating the healthcare infrastructure with portable and/or implantable biomedical devices requires an even higher standard of performance than what can be offered by today’s circuit technology.
In this chapter, we discuss a radically new design framework to seamlessly integrate data processing algorithms and their customized circuit implementations for co-optimization. The proposed framework could bring about numerous opportunities to substantially improve the performance of biomedical circuits. From this point of view, it offers a fundamental infrastructure that enables next-generation biomedical circuit design and optimization for many emerging applications.
2 How Can We Beat the State of the Art?
In this chapter, we attempt to address the following fundamental question: How can we further push the limit of accuracy, latency, power, and reconfigurability to meet the challenging performance required for portable and/or implantable biomedical applications? Historically, algorithm and circuit designs have been considered as two separate steps. Namely, a biomedical data processing algorithm is first developed and validated by its software implementation (e.g., MATLAB, C++, etc.). Next, a circuit is designed to implement the given algorithm. Such a two-step strategy suffers from several major limitations that motivate us to fundamentally rethink the conventional wisdom in this area.
First, since the biomedical data processing algorithms are particularly developed and tuned for their software implementations, they are not fully optimized for circuit implementations. Ideally, data processing algorithms should be customized to mitigate the non-idealities induced by circuit implementations (e.g., nonlinear distortion of analog front-end, quantization error of digital computing, etc.). Second, while a circuit implementation inevitably introduces various non-idealities, these non-idealities can be classified into two broad categories: (1) critical non-idealities that may significantly distort the output of a biomedical circuit, and (2) non-critical non-idealities that can be effectively mitigated or even completely eliminated by the data processing algorithm. A good circuit implementation should optimally budget the available resources (e.g., power) to maximally reduce the critical non-idealities rather than the non-critical ones.
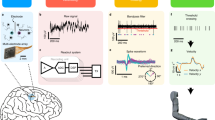
Motivated by these observations, we propose to develop a radically new design framework to seamlessly integrate data processing algorithms and their customized circuit implementations for co-optimization, as shown in Fig. 7.1. Our core idea is to view a biomedical circuit, along with the data processing algorithm implemented by the circuit, as an information processing system. We develop an information-theoretic metric, referred to as information processing capacity (IPC) that extends the conventional communication notion of channel capacity to our application of biomedical data sensing, processing, and transmission. IPC quantitatively measures the amount of information that can be processed by the circuit. Intuitively, IPC is directly correlated to the accuracy of the circuit implementation. If a circuit can accurately process the input data and generate the desired output, its IPC is high. Otherwise, its IPC is low. In the extreme case, if a circuit cannot generate any meaningful output due to large errors, its IPC reaches the lowest value zero.
IPC can efficiently distinguish critical vs. non-critical non-idealities. It is strongly dependent on the critical non-idealities that distort the output, and is independent of the non-critical non-idealities that can be eliminated by the data processing algorithm. Hence, it serves as an excellent “quality” metric that we should maximize in order to determine the optimal data processing algorithm and the corresponding circuit implementation subject to a set of design constraints (e.g., latency, power, reconfigurability, etc.).
It is important to note that our proposed design framework is not simply to combine algorithm and circuit designs. Instead, we aim to develop new methodologies that would profoundly revise today’s data processing algorithms and integrated circuit designs for biomedical applications. In particular, our proposed information-theoretic framework can optimally explore the tradeoffs between accuracy, latency, power, and reconfigurability over all hierarchical levels from algorithm design to circuit implementation. From this point of view, the proposed framework based on IPC offers a fundamental infrastructure that enables next-generation biomedical circuit design and optimization for numerous emerging applications.
3 Information Processing Capacity
In this section, we describe an information-theoretic metric, IPC, to quantitatively measure the amount of information that can be processed by a biomedical circuit. It serves as a “quality” measure, when we co-optimize the algorithms and circuits for data sensing, processing, and transmission of biomedical devices. It, in turn, facilitates us to achieve superior accuracy, latency, power, and reconfigurability over the conventional design strategies.
3.1 Information-Theoretic Modeling
The IPC of a biomedical circuit can be mathematically modeled based on information theory. Without loss of generality, we consider a biomedical circuit, including data sensing, processing, and transmission modules in general, as an information processing system shown in Fig. 7.2a. Since the circuit implementation is not perfect due to non-idealities, the aforementioned information processing system may generate errors where its output y CKT deviates from the desired value.
An information-theoretic framework is proposed to co-optimize data processing algorithms and their customized circuit implementations for higher accuracy, smaller latency, lower power, and better reconfigurability of biomedical devices. (a) circuit implementation with non-idealities, and (b) ideal implementation
To accurately characterize the “error” of the biomedical circuit, we further consider an ideal information processing system that guarantees to provide the correct output y IDEAL, as shown in Fig. 7.2b. In other words, the ideal system is “conceptually” implemented with a circuit with “infinite” precision. It does not carry any non-ideality and, hence, is error-free. The “difference” between y CKT and y IDEAL indicates the non-idealities caused by the circuit implementation. However, quantitatively measuring such a difference is non-trivial, since a biomedical circuit can be applied to a broad range of usage scenarios (e.g., various physiological signals, various users, various environmental conditions, etc.). The comparison between y CKT and y IDEAL must cover all these scenarios where y CKT and y IDEAL are not just two numerical numbers and, hence, we cannot compare their difference by simply subtracting y CKT from y IDEAL. The information-theoretic metric, IPC, quantitatively measures the “quality” of approximating y IDEAL by y CKT. To derive the mathematical representation of IPC, we consider two different cases: (1) discrete output and (2) continuous output.
First, if the outputs y CKT and y IDEAL are discrete values (e.g., the diagnosis result of arrhythmia may be positive or negative), y CKT and y IDEAL can be modeled as two discrete random variables to cover the uncertainties over all usage scenarios. In general, we assume that y CKT and y IDEAL take M possible values {y 1, y 2, ⋅⋅⋅, y M }. The statistics of these two random variables can be described by using their joint probability mass function (PMF) pmf(y CKT, y IDEAL). Table 7.1 shows a simple example for the binary random variables y CKT and y IDEAL (i.e., either TRUE or FALSE) where their statistics are fully described by four probabilities: true positive rate P TP, false negative rate P FN, false positive rate P FP, and true negative rate P TN.
IPC is defined as the mutual information I(y CKT, y IDEAL) between y CKT and y IDEAL [2, 4]:
Intuitively, the IPC metric in (7.1) measures the amount of information carried by y IDEAL that can be learned from y CKT. In one extreme case, if the circuit implementation is perfect, y CKT is identical to y IDEAL and, hence, IPC reaches its maximum. In the other extreme case, if y CKT does not follow y IDEAL at all due to large errors, there is no information about y IDEAL that can be learned from y CKT and, hence, IPC reaches its minimum (i.e., zero).
There are two important clarifications that should be made for IPC. First, instead of directly measuring the information carried by the circuit output y CKT, we take the ideal output y IDEAL as the “reference” and measure the information related to y IDEAL. Since y IDEAL represents all the important information of interest, IPC accurately captures our “goal” and ignores the “don’t cares.” This is the reason why IPC can serve as an excellent quality metric to guide our proposed algorithm/circuit co-optimization.
Second, IPC is different from other simple accuracy metrics that directly measure the difference between y CKT and y IDEAL based on statistical expectations. To understand the reason, we consider the example in Fig. 7.3a for which we may simply define the accuracy as the summation of the true positive rate P TP and the true negative rate P TN. Fig. 7.3b shows how this accuracy metric varies as a function of the false positive rate P FP and the false negative rate P FN where the probabilities for y IDEAL to be TRUE and FALSE are set to pmf(y IDEAL = TRUE) = 0.01 and pmf(y IDEAL = FALSE) = 0.99, respectively. In this example, we set pmf(y IDEAL = TRUE) to be much less than pmf(y IDEAL = FALSE) to mimic the practical scenarios where pmf(y IDEAL = TRUE) and pmf(y IDEAL = FALSE) are highly unbalanced. For example, in the application of arrhythmia diagnosis [3, 19, 29], the probably of being positive (i.e., with arrhythmia) should be much less than the probability of being negative (i.e., without arrhythmia), since arrhythmia is only carried by a small group of unhealthy patients over the entire population.
(a) The conventional accuracy metric is not appropriately influenced by the false negative rate P FN, if pmf(y IDEAL = TRUE) is much less than pmf(y IDEAL = FALSE). (b) The proposed IPC metric is appropriately influenced by both the false positive rate P FP and the false negative rate P FN, even if pmf(y IDEAL = TRUE) and pmf(y IDEAL = FALSE) are highly unbalanced
Studying Fig. 7.3a, we observe that the simple accuracy metric heavily depends on the false positive rate P FP, but weakly depends on the false negative rate P FN, because the probability pmf(y IDEAL = TRUE) is extremely small. If we maximize the aforementioned accuracy metric for algorithm/circuit co-optimization, it would aggressively minimize the false positive rate P FP, thereby resulting in a large false negative rate P FN. Consequently, a large portion of the unhealthy patients with arrhythmia may be mistakenly diagnosed as healthy ones.
On the other hand, Fig. 7.3b shows the relation between our proposed IPC and P FP and P FN. It can be observed that IPC is influenced by both P FP and P FN. Hence, by maximizing IPC, we take both P FP and P FN into account. This simple example demonstrates that when pmf(y IDEAL = TRUE) and pmf(y IDEAL = FALSE) are highly unbalanced, IPC can appropriately guide our proposed algorithm/circuit co-optimization, while the simple accuracy metric fails to work.
Finally, it is worth mentioning that if the outputs y CKT and y IDEAL are continuous values (e.g., movement decoding for neuroprosthesis results in the velocity value that is continuous), y CKT and y IDEAL can be modeled as two continuous random variables and their statistics can be described by the joint probability density function pdf(y CKT, y IDEAL). In this case, IPC can again be defined as the mutual information I(y CKT, y IDEAL) between y CKT and y IDEAL [2, 4]:
In the following sub-sections, we will further discuss how IPC can be used for design and optimization of biomedical circuits.
3.2 Soft Channel Selection
Soft channel selection is an important task that is facilitated by our proposed algorithm/circuit co-optimization based on IPC. We consider the multi-channel biomedical device in Fig. 7.4, where channel selection is one of the most important tasks [1, 10, 23]. Appropriately selecting the important channels and removing the unimportant ones can efficiently minimize the amount of data for sensing, processing, and transmission, thereby substantially reducing the power consumption.
Today’s channel selection is typically considered as a binary decision: a channel is either selected or not selected for recording. With the proposed information-theoretic framework based on IPC, we are now able to make a “soft” decision for each channel, referred to as soft channel selection. Namely, instead of simply including or excluding a given channel, we can finely tune the resolution of the channel (e.g., the number of bits required to represent the signal from the channel). Intuitively, an important channel should be designed with high resolution, while an unimportant channel can be designed with low resolution. The channel resolution is directly correlated to the power consumption of both analog front-end (e.g., sensors, analog filters, ADCs, etc.) and digital computing (e.g., digital filters, data processors, etc.). It, in turn, facilitates us to optimally explore the tradeoff between accuracy and power. In the extreme case, if the resolution of a channel is set to 0-bit, the channel is completely removed and it is equivalent to the conventional binary channel selection in the literature.
To demonstrate the efficacy of our proposed soft channel selection, we consider a preliminary example of movement decoding for neuroprosthesis, where our objective is to decode the movement direction from electrocorticography (ECoG) [26]. Fig. 7.5 compares the optimal IPC for both the conventional binary channel selection and the proposed soft channel selection. Note that the proposed approach successfully reduces the power of the analog front-end by up to 10×. It is important to mention that the proposed idea of soft channel selection can be further extended to other important applications such as data compression and transmission.
3.3 Robust Data Processing
To maximally reduce the power consumption for portable and/or implantable applications, fixed-point arithmetic, instead of floating-point arithmetic, is often adopted to implement data processing algorithms and the word length for fixed-point computing must be aggressively minimized. While fixed-point arithmetic has been extensively studied for digital signal processing during the past several decades [14, 20], it is rarely explored for many emerging data processing tasks that involve sophisticated learning algorithms (e.g., movement decoding for neuroprosthesis). It remains an open question how to revise these algorithms to maximally tolerate the quantization error posed by finite word length. Based upon IPC, data processing algorithms can be completely redesigned to mitigate the quantization error so that these algorithms can be mapped to fixed-point implementations with extremely low resolution.
As shown in Fig. 7.6, a learning algorithm typically consists of two steps: (1) feature extraction and (2) classification (e.g., to determine movement direction for neuroprosthesis) and/or regression (e.g., to determine movement velocity for neuroprosthesis). We propose to maximize the IPC metric of a classification or regression engine subject to the constraint that all arithmetic operations for both feature extraction and classification/regression are quantized. Our reformulated learning algorithm solves a “robust” optimization problem to find the optimal, quantized classifier or regressor that is least sensitive to quantization error. It, in turn, offers superior performance over other conventional approaches where quantization error is not explicitly considered within the learning process.
As an example for illustration purpose, we consider the classification problem of decoding the movement direction from electrocorticography (ECoG) for neuroprosthesis. Fig. 7.7 shows the optimized IPC metric as a function of word length. To achieve the same IPC, our proposed approach can reduce the word length by 2-bit compared to the conventional classifier. Note that the word length of fixed-point arithmetic is directly correlated to the power consumption of its circuit implementation. Hence, reducing word length is of great importance for low-power portable and/or implantable biomedical devices.
4 Case Study: Brain–Computer Interface
Brain–computer interface (BCI) has been considered as a promising communication technique for patients with neuromuscular impairments. For instance, neural prosthesis provides a direct control pathway from brain to external prosthesis for paralyzed patients. It can offer substantially improved quality of life to these patients. To create a neural prosthesis, we must appropriately measure the brain signals and then accurately decode the movement information from the measured signals [5, 11, 13, 21, 22, 26].
A variety of signal processing algorithms have been proposed for movement decoding in the literature. Most of these algorithms first extract the important features to compactly represent the information carried by the brain signals. Next, the extracted features are provided to a classification and/or regression engine to decode the movement information of interest. While most movement decoding algorithms in the literature are implemented with software on microprocessors, there is a strong need to migrate these algorithms to hardware in order to reduce the power consumption for practical BCI applications.
4.1 System Design
Fig. 7.8shows a simplified block diagram for the proposed hardware implementation of BCI. It consists of three major components:
-
Signal normalization: The magnitude of brain signals varies from subject to subject and from channel to channel. Hence, representing brain signals by fixed-point arithmetic requires a large word length (i.e., a large number of bits). In order to minimize the word length and, consequently, the power consumption for fixed-point computation, we must appropriately normalize the brain signal from each channel.
-
Feature extraction: There are many different feature extraction approaches for movement decoding of BCI. For instance, given the brain signal recorded from a particular channel, we can apply discrete cosine transform (DCT) and consider the DCT coefficients as the features for decoding [28].
-
Classification: Once all features are extracted for multiple channels, they are further combined to decode the movement information. For instance, all features can be linearly combined by a linear classifier to determine the movement direction of interest. Here, a variety of linear classification algorithms (e.g., linear discriminant analysis, support vector machine, etc.) can be used, where the classifier training is performed offline. The on-chip classification engine performs the multiply-and-accumulate operations to determine the final output (i.e., the movement direction) from the features.
4.2 Experimental Results
We consider the ECoG data set collected from a human subject with tetraplegia due to spinal cord injury [26]. The ECoG signals are recorded with a high-density 32-electrode grid over the hand and arm area of the left sensorimotor cortex. The sampling frequency is 1.2 kHz. The human subject is able to voluntarily activate his sensorimotor cortex using attempted movements.
Our objective is to decode the binary movement direction (i.e., left or right) from a single trial that is 300 ms in length. The ECoG data set contains 70 trials for each movement direction (i.e., 140 trials in total). For movement decoding, 7 important channels with 6 features per channel (i.e., 42 features in total) are selected based on the Fisher criterion. A linear classifier is trained and implemented with 8-bit fixed-point arithmetic to decode the movement direction.
The BCI system is implemented with a Xilinx FPGA Zynq-7000 board. For testing and comparison purposes, we further implement a reference design based on the conventional technique [30]. In this sub-section, we compare the performance between our proposed hardware implementation and the reference design.
We estimate the power and energy consumption for both the proposed and the reference designs by using Xilinx Power Analyzer, where the clock frequency is set to 0.5 MHz. Table 7.2 compares the power consumption for these two different designs. Note that the proposed design achieves more than 56× energy reduction over the reference design. Table 7.3 further shows the power consumption for different functional blocks of the proposed design. Note that feature extraction dominates the overall power consumption for our proposed hardware implementation. Hence, additional efforts should be pursued to further reduce the power consumption of feature extraction in our future research.
To validate the proposed design on the Xilinx Zynq-7000 board, we first load our hardware design to the FPGA chip through the programming interface. Next, the ECoG data set is copied to an SD card that is connected to the Zynq-7000 board. When running the movement decoding flow, a single trial of the ECoG signals is first loaded to the SRAM block inside the FPGA chip. Next, these signals are passed to the functional blocks of signal normalization, feature extraction and classification for decoding. The decoding results are read back to an external computer through an RS-232 serial port on the Zynq-7000 board so that we can verify the decoding accuracy. Fig. 7.9 shows a photograph of the Xilinx FPGA Zynq-7000 board where the RS-232 port and the programming interface are both highlighted.
5 Summary
In this chapter, we describe a new design framework for ultra-low-power biomedical circuits. The key idea is to co-optimize data processing algorithms and their circuit implementations based on an information-theoretic metric: IPC. The proposed design framework has been demonstrated by a case study of BCI. Our experimental results show that the proposed design achieves more than 56× energy reduction over a reference design. As an important aspect of our future research, we will further apply the proposed design framework to other emerging biomedical applications.
References
M. Arvaneh, C. Guan, K.K. Ang, C. Quek, Optimizing the channel selection and classification accuracy in EEG-based BCI. IEEE Trans. Biomed. Eng. 58, 1865–1873 (2011). doi:10.1109/TBME.2011.2131142
C. Bishop, Pattern Recognition and Machine Learning (Springer, Berlin, 2006)
P. de Chazal, R.B. Reilly, A patient-adapting heartbeat classifier using ECG morphology and heartbeat interval features. IEEE Trans. Biomed. Eng. 53, 2535–2543 (2006). doi:10.1109/TBME.2006.883802
T. Cover, J. Thomas, Elements of Information Theory (Interscience, Wiley, 2006)
J.P. Donoghue, Bridging the brain to the world: a perspective on neural interface systems. Neuron 60, 511–521 (2008). doi:10.1016/j.neuron.2008.10.037
C. Guo, X. Li, S. Taulu, W. Wang, D.J. Weber, Real-time robust signal space separation for magnetoencephalography. IEEE Trans. Biomed. Eng. 57, 1856–1866 (2010). doi:10.1109/TBME.2010.2043358
S. Khalid, D. Clifton, L. Clifton, L. Tarassenko, A two-class approach to the detection of physiological deterioration in patient vital signs with clinical label refinement. IEEE Trans. Inf. Technol. Biomed. 16, 1231–1238 (2012). doi:10.1109/TITB.2012.2212202
J. Ko, C. Lu, M.B. Srivastava, J.A. Stankovic, A. Terzis, M. Welsh, Wireless sensor networks for healthcare. Proc IEEE 98, 1947–1960 (2010). doi:10.1109/JPROC.2010.2065210
M.L. Kringelbach, N. Jenkinson, S.L. Owen, T.Z. Aziz, Translational principles of deep brain stimulation. Nature 8, 523–635 (2007). doi:10.1038/nrn2196
T.N. Lal, M. Schroder, T. Hinterberger, J. Weston, M. Bogdan, N. Birbaumer, B. Scholkopf, Support vector channel selection in BCI. IEEE Trans. Biomed. Eng. 51, 1003–1010 (2004). doi:10.1109/TBME.2004.827827
M.A. Lebedev, M.A. Nicolelis, Brain-machine-interface: past, present and future. Trends Neurosci. 29, 536–546 (2006). doi:10.1016/j.tins.2006.07.004
K.H. Lee, S.Y. Kung, N. Verma, Low-energy formulations of support vector machine kernel functions for biomedical sensor applications. J Signal Process Syst 69, 339–349 (2012). doi:10.1007/s11265-012-0672-8
E.C. Leuthardt, G. Schalk, J.R. Wolpaw, J.G. Ojemann, D.W. Moran, A brain-computer interface using electrocorticographic signals in humans. J. Neural Eng. 1, 63–71 (2004). doi:10.1088/1741-2560/1/2/001
W. Meyer-Baese, Digital Signal Processing with Field Programmable Gate Arrays (Springer, Berlin, 2007)
C. Mora Lopez, D. Prodanov, D. Braeken, I. Gligorijevic, W. Eberle, C. Bartic, R. Puers, G. Gielen, A multichannel integrated circuit for electrical recording of neural activity, with independent channel programmability. IEEE Trans Biomed Circuits Syst 6, 101–110 (2012). doi:10.1109/TBCAS.2011.2181842
R. Muller, S. Gambini, J.M. Rabaey, A 0.013 mm2, 5 μW, DC-coupled neural signal acquisition IC with 0.5 V supply. J Solid-State Circuits 47, 232–243 (2012). doi:10.1109/JSSC.2011.2163552
K. Nazarpour, Y. Wongsawat, S. Sanei, J.A. Chambers, S. Oraintara, Removal of the eye-blink artifacts from EEGs via STF-TS modeling and robust minimum variance beamforming. IEEE Trans. Biomed. Eng. 55, 2221–2231 (2008). doi:10.1109/TBME.2008.919847
C.O. Oluigbo, A. Salma, A.R. Rezai, C. Oluigbo, Deep brain stimulation for neurological disorders. IEEE Rev. Biomed. Eng. 5, 88–99 (2012). doi:10.1109/RBME.2012.2197745
S. Osowski, L.T. Hoai, T. Markiewicz, Support vector machine-based expert system for reliable heartbeat recognition. IEEE Trans. Biomed. Eng. 51, 582–589 (2004). doi:10.1109/TBME.2004.824138
W. Padgett, D. Anderson, Fixed-point Signal Processing (Morgan and Claypool Publishers, Williston, 2009)
G. Pfurtscheller, C. Neuper, Motor imagery and direct brain-computer communication. Proc EEE 89, 1123–1134 (2001). doi:10.1109/5.939829
A.B. Schwartz, X.T. Cui, D.J. Weber, D.W. Moran, Brain-controlled interfaces: movement restoration with neural prosthetics. Neuron 52, 205–220 (2006). doi:10.1016/j.neuron.2006.09.019
P. Shenoy, K.J. Miller, J.G. Ojemann, R.P. Rao, Generalized features for electrocorticographic BCIs. IEEE Trans. Biomed. Eng. 55, 273–280 (2008). doi:10.1109/TBME.2007.903528
X.F. Teng, Y.T. Zhang, C.C. Poon, P. Bonato, Wearable medical systems for p-health. IEEE Rev. Biomed. Eng. 1, 62–74 (2008). doi:10.1109/RBME.2008.2008248
N.V. Thakor, Y. Zhu, Applications of adaptive filtering to ECG analysis: noise cancellation and arrhythmia detection. IEEE Trans. Biomed. Eng. 38, 785–794 (1991). doi:10.1109/10.83591
W. Wang, J.L. Collinger, A.D. Degenhart, E.C. Tyler-Kabara, A.B. Schwartz, D.W. Moran, D.J. Weber, B. Wodlinger, R.K. Vinjamuri, R.C. Ashmore, J.W. Kelly, M.L. Boninger, An electrocorticographic brain interface in an individual with tetraplegia. PLoS One 8, e55344 (2013). doi:10.1371/journal.pone.0055344
E.S. Winokur, M.K. Delano, C.G. Sodini, A wearable cardiac monitor for long-term data acquisition and analysis. IEEE Trans. Biomed. Eng. 60, 189–192 (2013). doi:10.1109/TBME.2012.2217958
M. Won, H. Albalawi, L. Li, D.E. Thomas, Low-power hardware implementation of movement decoding for brain computer interface with reduced-resolution discrete cosine transform. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2014, 1626–1629 (2014). doi:10.1109/EMBC.2014.6943916
C. Ye, B.V. Kumar, M.T. Coimbra, Heartbeat classification using morphological and dynamic features of ECG signals. IEEE Trans. Biomed. Eng. 59, 2930–2941 (2012). doi:10.1109/TBME.2012.2213253
J. Yoo, L. Yan, D. El-Damak, M. Altaf, A.H. Shoeb, A.P. Chandrakasan, An 8-channel scalable EEG acquisition SoC with patient-specific seizure classification and recording processor. J Solid-State Circuits 48, 214–228 (2013). doi:10.1109/JSSC.2012.2221220
K. Yousef, M.R. Pinsky, M.A. DeVita, S. Sereika, M. Hravnak, Characteristics of patients with cardiorespiratory instability in a step-down unit. Am. J. Crit. Care 21, 344–350 (2012). doi:10.4037/ajcc2012797
X. Zou, X. Xu, L. Yao, Y. Lian, A 1-V 450-nW fully integrated programmable biomedical sensor interface chip. J Solid-State Circuits 44, 1067–1077 (2009). doi:10.1109/JSSC.2009.2014707
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Li, X., (Shawn) Blanton, R.D., Grover, P., Thomas, D.E. (2017). Ultra-Low-Power Biomedical Circuit Design and Optimization: Catching the Don’t Cares. In: Chattopadhyay, A., Chang, C., Yu, H. (eds) Emerging Technology and Architecture for Big-data Analytics. Springer, Cham. https://doi.org/10.1007/978-3-319-54840-1_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-54840-1_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-54839-5
Online ISBN: 978-3-319-54840-1
eBook Packages: EngineeringEngineering (R0)