Abstract
This chapter comes up with a brief overview of molecular communication models and modulation techniques by reviewing current research works found in the literature. The chapter also provides with an analysis of molecular communication in free diffusion-based molecular communication channel. In this model, the trasmitter nanomachine releases messenger molecules, the molecules diffuse through the channel, and the receiver nanomachine counts the received molecules to decode the information. We consider free diffusion of molecules where no additional force is required. Such a channel is referred to as the diffusion channel and can be modeled by using Ficks law of diffusion. Diffusion coefficient describes the tendency of propagation of the messenger molecules through the medium. Analysis shows that, channel memory offers a significant impact on performance.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Forster Resonance Energy Transfer
- Receive Molecule
- Symbol Error Rate
- Inverse Gaussian Distribution
- Messenger Molecule
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Nanomachines are the blessings of nanotechnology built from individual molecule or arranged set of molecules. Communication between nanomachines can be built through mechanical, acoustic, electromagnetic, and chemical or molecular means [1]. Nanonetwork is the interconnection of nanomachines anticipated to perform collaborative task which an individual nanomachine is unable to do. We are promised with a handful offers by nanonetworks not limited to lab-on-a-chip, health monitoring, drug delivery, regenerative medicine, environment monitoring, waste/population control, pattern and structure formation etc. [2]. However, conventional communication technologies are found inapt for nanonetworks mainly due to the size and power consumption of the components in operation [3]. Molecular communication appears to be a promising approach for nanonetworks. First nanonetwork models are stimulated by molecular communications observed in biological systems. Small-scale devices built from biological materials, posing a high degree of energy efficiency and biocompatibility, are capable of interacting with biological molecules and cells in nano to micrometer scale [2]. However, communication based on molecular signal differs from traditional communications. A summary of the features of molecular communication and classical electromagnetic communication is presented in Table 1. Seemingly, unlike electromagnetic or other traditional communications, molecules serve as information carriers in molecular communication. The number of molecules in transmit and received signals can be considered as the amplitudes of the signals. Therefore, existing information and communication theories will likely not be applicable directly. For example, the terms such as encoding, modulation, demodulaltion, transmission power, signal-to-noise-ratio (SNR), bandwidth, inter-channel interference (ICI), inter-symbol interference (ISI), noise, channel memory etc. are supposed to be treated in dissimilar fashion in molecular communication.
2 How Molecular Communication Works?
2.1 Encoding
In molecular communication, information are encoded onto molecules instead of electromagnetic or acoustic waves. Individual molecule properties or molecule ensemble properties are used for encoding. The information may be encoded based on the three-dimensional structure of the information molecule (e.g. a specific structure of molecule), or on the specific composition of the information molecules (e.g. DNA sequence), or on the concentration of information molecules (i.e. the number of information molecules per unit volume) [2].
2.2 Modulation
2.2.1 On Off Keying (OOK)
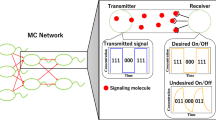
Analogous to classical electromagnetic communication, on-off keying is the process where the transmitter either releases molecules or remains silent over a symbol time period [5]. This is a binary modulation scheme. Decision for choosing 1 or 0 is based upon a threshold number of received molecules.
2.2.2 Concentration Shift Keying (CSK)
Concentration Shift Keying is analogous to amplitude shift keying as in classical electromagnetic communication. In this modulation scheme, information is encoded according to the number of information molecules per unit volume i.e., concentration [6]. Decision for choosing 1 or 0 is based on a threshold of the concentration of received molecules. In binary concentration shift keying, two concentration levels are used and we need one threshold for decision. The scheme can be increased to higher levels depending upon the number of concentration levels. For example, in quadrature concentration shift keying, four concentration levels are used and we need three thresholds for decision. Figure 1 shows the diagram for QCSK modulation where the concentration levels \(c_0\), \(c_1\), \(c_2\), \(c_3\) are used for the symbols \(s_0=(0,0)\), \(s_1=(0,1)\), \(s_2=(1,0)\), \(s_3=(1,1)\), respectively.
2.2.3 Molecule Shift Keying (MoSK)
Bearing a resemblance to frequency shift keying, in this modulation scheme, information is encoded onto different types of molecules. In binary molecular shift keying, two types of molecules are used [6]. Like concentration shift keying, this scheme can also be increased to higher levels depending upon the types of molecules. For example, in quadrature molecular shift keying, four types of molecules are used for encoding. Figure 2 shows the diagram for QMoSK modulation where \(n_1\), \(n_2\), \(n_3\), \(n_4\) molecules are used for the symbols \(s_0=(0,0)\), \(s_1=(0,1)\), \(s_2=(1,0)\), \(s_3=(1,1)\), respectively and \(z_1\), \(z_2\), \(z_3\), \(z_4\) are corresponding threshold number of received molecules.
2.2.4 Isomer-Based Ratio Shift Keying (IRSK)
IRSK encodes the information based on the ratios of different types of messenger molecules and this type of modulation scheme has high, theoretically infinite, modulation order [7].
Other modulation schemes include in-sequence molecule delivery, time of dispersal of molecules, etc.
2.3 Transmission
Transmitter nanomachine releases the messenger molecules in the environment either by unbinding the messenger molecules from the sender nanomachine or by opening a gate that allows the information molecules to diffuse away [2].
2.4 Signal Propagation
The propagation of messenger molecules from the transmitter to the receiver nanomachine is governed by Brownian motion and is affected by two parameters: drift velocity and the diffusion coefficient [8]. Information molecules in molecular communication propagate through fluid medium (such as blood or water) whereas electromagnetic waves propagate through free space or wire. Apparently, molecular communication is much slower than electromagnetic and other communication processes. Figure 3 shows the trajectory of Brownian motion of a particle in 20 s. We can infer from the figure that molecular communication is not a deterministic process. Unlike electromagnetic or other communication processes, it is not certain that a molecule will reach a certain distance at a certain time. Molecular signal is also a discrete signal. Even a single molecule can be a signal in molecular communication. The diffusion of molecules can be interpreted by using Wiener process in which, the displacement of each molecule during an infinitesimal interval of time dt is modeled with a zero-mean Gaussian distribution with variance 2Ddt [9]:
where \(\mathbf R (t)=\hat{i}x(t)+\hat{j}y(t)+\hat{k}z(t)\) is the position vector of a molecule with unit vector \((\hat{i}, \hat{j}, \hat{k})\) at time t and \(\mathbf v (t)\) is a vector of random variables. \(\mathbf v (t)\) has a multivariate Normal distribution \(\mathbf v (t)\sim \mathcal {N} (\mathbf 0 , \mathbf I )\), where \(\mathbf 0 \) and \(\mathbf I \) are the null and identity matrices. Diffusion coefficient D, measured in m\(^2\)/s, describes the tendency of propagation of the messenger molecules through the medium. It can be written as
where \(k_B\) is the Boltzmann constant in \(JK^{-1}\), T is the absolute temperature of the environment in Kelvin, and b is the drag constant of the propagating molecule inside the given fluid. The constant b is affected by the comparative sizes between the propagating molecule (\(S_m\)) and the molecules of the fluid (\(S_f\)) [10]. If the propagating molecule’s size is similar to the size of the molecules of the fluid then the fluid can be considered as a continuum [11]. Based on these two different conditions, the constant b is calculated as
where \(\eta \) is the viscosity of the fluid and \(\zeta _s\) defines the Stokes’ radius of the propagating molecule. Stokes’ radius of a molecule is defined as the radius of a sphere whose diffusion dynamics are the same as the molecule in question in the same environment (such as fluid type, temperature).
The mean change in the concentration of molecules with time can be formalized by using Fick’s second law of diffusion:
where \(C(\mathbf R ,t)\) is the mean concentration of molecules in \(molecules/m^3\) and t is the time after release from the transmitter, \(\nabla ^{2}\) is the three dimensional Laplacian operator denoted as \(\nabla ^{2}=\frac{\partial ^2}{\partial x^2} +\frac{\partial ^2}{\partial y^2} +\frac{\partial ^2}{\partial z^2} \). By solving the above equation, when an impulse of n messenger molecules (i.e., \(n \delta (t)\)) released at time \(t=0\), the mean concentration of molecules \(C(\mathbf R ,t)\) can be obtained as follows:
where \(\mathbf R _{tx}=\hat{i}x_{tx}+\hat{j}y_{tx}+\hat{k}z_{tx}\) is the position of the transmitter nanomachine. Figure 4 shows the graph for normalized concentration at different diffusion coefficients. We notice from the figure that, the higher the value of the diffusion coefficient, the quicker the normalized concentration reaches its peak.
The first passage time T of a molecule at a certain distance r with a certain diffusion coefficient D is a random variable. The probability density function of first passage time T is [12]
where \(\mathbf r =\mathbf R - \mathbf R _{tx}\).
Using Eq. 6, we find the probability that a molecule reaches a receiver nanomachine within \((\tau ,\tau +T_s )\) as
where \(\tau \) is a given transmission time defined as the delay and \(T_s\) is the time slot.
2.5 Receiving and Decoding
Upon capturing the incoming molecules, the receiver bio-nanomachines biochemically react with the molecules to decode the information [2]. Receptors, capable of binding to a specific type of information molecules, can be used for capturing. Channels can also be used for capturing (e.g., gap-junction channels) [4]. For decoding, the receiver may produce new molecules, perform simple task, or produce another signal [4].
2.6 Noise
Other source(s) may emit the same molecules used to encode the information messages [3] or it can be originated from an undesired reaction occurring between information molecules and other molecules present in the medium. As occurs in traditional communication systems, noise can be overlapped with molecular signals such as concentration level of molecules. If there is a drift, the time an information molecule requires to reach the receiver nanomachine follows an inverse Gaussian distribution [13, 14]. This leads to inverse Gaussian noise in time modulation.
2.7 Inter-symbol Interference (ISI)
Inter-symbol interference (ISI) may occur due to the delay of molecules in the medium [13]. The molecules used to convey information (be the information encoded in the number of molecules emitted, in their concentration, or in the time instant of release etc.) can interact with the transmission medium [15]. The molecules emitted from other source(s) can also be overlapped with intended molecular signals.
2.8 Channel Memory
Some molecules stay longer in the medium and reach the receiver after a delay. The transmitter may release the molecule corresponding to the ‘next’ symbol while the ‘previous’ molecules are still propagating. These delaying molecules arrive out of order introducing memory to the channel [13]. Following phenomena should also be taken into account:
-
Information-carrying molecules may react with other molecules present in the medium and get absorbed.
-
Molecules may replicate by stimulating the generation of new molecules.
-
Spontaneous emission may take place so that molecules may get generated within the medium without any external intervention.
2.9 Delay
Not all the molecules reach the receiver within a specific time slot. Rather, molecules move around in the medium and reach the receiver after a delay. Since the propagation of molecules is Brownian in nature, the molecules may reach the receiver nanomachine after a long time. These delaying molecules may impinge on the later transmissions contributing to ISI or ICI.
3 Molecular Communication Model
Concepts of molecular nanonetworks are inspired by molecular communications observed in biological systems. Various modes and mechanisms of molecular communications found within and between cells are common and ubiquitous methods by which biological nanomachines communicate. Quorum sensing, calcium signaling etc. are few examples of biological nanonetworks that appear in intra-cell, inter-cell and intra-species communication. Modes and mechanisms of molecular communication are categorized based on how signal molecules propagate through the environment. Molecular signals may simply diffuse through the medium or directionally propagate by consuming chemical energy. The former type is categorized as the passive transport-based molecular communication, and the later as active transport-based molecular communication [4].
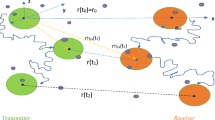
Free diffusion-based molecular communication [16]
3.1 Passive Transport-Based Molecular Communication
Passive transport is the simplest method of propagating signal molecules within a cell and between cells in which, no additional force is required to propagate the molecules [2]. Signal molecules diffuse in all available directions in passive transport-based molecular communication and the process is suitable for highly dynamic and unpredictable environment. However, passive transport requires large number of messenger molecules and it is suitable for smaller size of molecules. Passive transport can be categorized as:
3.1.1 Free Diffusion-Based Molecular Communication
Free diffusion appears to be the most fundamental mechanism that molecular communication relies on to propagate a molecule. Figure 5 shows a free diffusion-based molecular communication model. Signal molecules (e.g., proteins and peptides) are released by the transmitter cells into extracellular environment and the molecules diffuse through the medium freely. Neighboring cells capture the signal molecules by using protein receptors which results in activation of chemical reaction such as increasing metabolism or transcription of cellular proteins. In biology, several examples of this class of molecular communication are observed. One illustration is intracellular metabolites propagating between cells. Another example is DNA binding molecules (e.g., repressors) that propagate over a DNA segment to search for a binding site [17]. Quorum sensing, a communication mechanism of bacterial cells, is referred to as a free diffusion-based molecular communication. In this process, bacteria coordinate certain behaviors such as biofilm formation, virulence, and antibiotic resistance, based on the local density of the population of bacteria [18]. They constitutively produce and secrete certain signaling molecules such as autoinducers (e.g., acyl homoserine lactone) or pheromones. They also have a receptor that can specifically detect the signaling molecule (inducer). Molecular communications of this class have been studied extensively in [5, 19,20,21].
3.1.2 Molecular Diffusion with Drift
In this model, molecules undergo Brownian motion with drift. The propagation of molecules depends upon the drift velocity and diffusion coefficient. Information molecules may have drift velocity during release. The molecules may also undergo directional drift which continuously propagates molecules in the direction of drift [8, 13]. If there is a drift from transmitter to receiver, the first arrival time of the particles has the Inverse Gaussian distribution, leading to the additive Inverse Gaussian (IG) Noise channel [13]. This type of molecular communication is observed in human body. Cells in the body secrete hormonal substances which circulate with the flow of the blood stream and propagate to the distant target cells distributed throughout the body [2]. The process may also represent the active mode of molecular communication by which motor proteins carry and directionally propagate molecules from a sender bio-nanomachine to a receiver bio-nanomachine [22, 23].
3.1.3 Molecular Diffusion with Reactions by Amplifiers
Amplifiers, situated in the medium, may chemically or chemically react with molecules that propagate through the medium and produce the same type of molecules which propagates in the medium. Thus they increase the reliability of molecular propagation by increasing the number of propagating information molecules. This class of molecular communication may be enabled by using protein molecules that are responsible for amplifying calcium ions, adenosine triphosphate (ATP), and cyclic adenosine monophosphate (CAMP) [24].
3.1.4 Gap Junction Mediated Diffusion-Based Molecular Communication
This is basically a cell to cell communication method in which the diffusion of signal molecules can be guided through cell-cell communication channel [24]. Gap junction channels, analogous to the plasmodesmata that join plant cells, are physical channels between two adjacent cells allowing only connected cells to communicate. One gap junction channel is composed of two connexons (or hemichannels). Two adjacent cells are connected through the cytoplasm of the two cells. In vertebrates, gap junction hemichannels are primarily homo- or hetero-hexamers of connexin proteins while invertebrate gap junctions are composed of proteins from the hypothetical innexin family. However, pannexin family functions as single-membrane channels that communicate with the extracellular environment and have been shown to pass calcium and adenosine triphosphate (ATP). Figure 6 shows gap junction mediated reaction-diffusion based molecular communication. Nanomachines are engineered organisms or biological devices whose behavior is programmed to achieve application specific goals, and chemically communicate over a cell-cell communication medium. To support molecular communication between nanomachines, the communication medium may provide various networking mechanisms and services such as signal amplification and switching [25].
Gap junction mediated reaction-diffusion based molecular communication [4]
3.1.5 Reaction-Diffusion-Based Molecular Communication
Diffusion of signal molecules may biochemically react to form a different mode of communication letting the propagation of impulses [4]. Quick variation of the concentration of the signal molecules results in an impulse that propagates through the environment. For example, endoplasmic reticulum (ER) in a cell gathers and stores calcium ions. The cell, if stimulated, releases the stored calcium ions from the ER and the calcium ions diffuse to adjacent cells through cell-cell junction channels. A chain reaction of calcium stimulation occurs because the diffused calcium stimulates the adjacent cells. A short impulse of calcium is created, since the cell pumps calcium within the cell back into the ER and suppresses further stimulation. Likewise, neurons also produce ion impulses that propagate over the length of the neuron.
3.2 Active Transport-Based Molecular Communication
In active transport mechanism, signal molecules are directionally transported to specific locations and can propagate over longer distances as compared to diffusion-based passive transport. The mechanism is suitable for propagating large signal molecules and requires fewer molecules. However, the process requires a regular supply of messenger molecules in order to overcome the chemical reactions between the messenger molecules and the molecules in the environment. Active transport can be categorized as:
3.2.1 Molecular Motor-Based Molecular Communication
Within a biological cell, molecular motors are found to be used in this type of molecular communication. Kinesin appears as one example of many molecular motors that may facilitate such transport [23]. Molecular motor is a protein or a protein complex that converts chemical energy (e.g., ATP hydrolysis) into mechanical work allowing the transportation of signal molecules or large vesicles (e.g., liposomes, cell organelles) that contain signal molecule [26,27,28,29].
3.2.2 Bacterial Motor-Based Molecular Communication
Based on chemical concentration in the medium, bacteria can move directionally or it can also exchange DNA through the process of conjugation [4]. In this process, two types of bacteria transfer Deoxyribonucleic acid (DNA) chromosome through a pilus (i.e., a projection from the sender bacterium to the receiver bacterium forming a bridge for transmitting DNA). Sender bacterium here is with an F-plasmid (i.e., a genetic sequence that enhances the transfer of genetic information) while a receiver bacterium is without F-plasmid. The receiver bacterium, in this process, acquires DNA that produces some useful cellular functionality (e.g., protein production, antibiotic resistance). The receiver may also release pheromones creating a chemical gradient that leads a sender bacterium toward a receiver bacterium. Figure 7 shows bacterial motor-based molecular communication.
Bacterial motor-based molecular communication [4]
3.3 Energy Model
3.3.1 Energy Model via Diffusion
Energy budget of the transmitting and receiving units is one of the factors that limits over the performance of the communication system. In this model, a system is considered in which each unit is able to produce and store energy [11]. Some of the produced energy is used by routine activities of the unit and the rest is available for communication purposes.
3.3.2 Forster Resonance Energy Transfer (FRET) Model
Forster resonance energy transfer (FRET) is a nonradiative energy transfer process between fluorescent molecules based on the dipole-dipole interactions of molecules. Energy is transferred rapidly from a donor to an acceptor molecule in a close proximity such as 0–10 nm without radiation of a photon [30].
4 Free Diffusion Molecular Communication Channel
4.1 Capacity Performance
Channel capacity, one of the key measures of the channel, is the upper bound of the information rate that can be reliably transmitted over a communication channel. We calculate the capacity of a diffusion-based communication system. The channel is said to have memory from one previous symbol. Such a channel is termed as first-order memory channel [31]. We consider OOK modulation scheme where the transmitter nanomachine either releases molecules or remains silent over a symbol period. The emitted molecules diffuse through the medium and arrive at the receiver. The receiver counts the number of molecules in order to retrieve the information. The decision is made on the basis of a threshold number of received molecules. No equalizer to mitigate inter-symbol interference (ISI) is considered in this model because the performance depends only on the number of received molecules. Let, \(n_0\) molecules are released for symbol \(x=1\), and 0 molecules for \(x=0\). Transmission time is divided into equal sized slots in which a single symbol (1 or 0) can be sent. The receiving time slots are chosen such a way that they are equal to the symbol durations and denoted by \(T_s\). \(n(r,T_s)\) is the number of molecules received at a distance r at time \(T_s\). The decision for choosing between 1 and 0 is denoted by z, where z is a designated number of molecules (threshold). If \(n(r,T_s)\ge z\) then we say that 1 is received, and if \(n(r,T_s)<z\) then we say that 0 is received. The probability of a molecule hitting the receiver is denoted by \(P_{hit}(r,T_s)\). Under the assumption that each molecule propagates independently, if \(n_0\) molecules are sent within a symbol duration, the number of molecules received within the current symbol duration \(n(r,T_s)\sim Binomial (n_0,P_{hit} (r,T_s))\) is a random variable and follows a binomial distribution. The first hitting time is \(\tau \), as we call delay, \(T_s\) is in the time interval from \(\tau \) to \(\tau +T_s\). The number of molecules \(n(r,T_s)\) received is the summation of molecules from current and previous symbols. We notice that the current symbol duration falls in the time interval from \(T_s\) to \(2T_s\) of the previous symbol. If \(n_p\) is the number of left over molecules belong to the previous symbol, then \(n_p\sim Binomial (n_0,P_{hit} (r,T_p ))\), where \(T_p\) is in the time interval from \(T_s\) to \(2T_s\). If the first hitting time is \(\tau \), then \(T_p\) is in the time interval from \(\tau +T_s\) to \(\tau +2T_s\). The number of molecules received within two symbol durations [11]
Clearly, \(N_{hit}\) is nothing but the number of molecules from current symbol received in the time interval from 0 to \(2T_s\) or from \(\tau \) to \(\tau +2T_s\).
According to the probabilistic theory, a Binomial distribution (Binomial(n, p)) can be approximated with a normal distribution \(\mathcal {N}\sim (np,np(1-p))\), when p is not close to one or zero and np is large enough. Therefore Eq. 9, can be approximated as
There are four such different cases for decoding the received symbol. From Eq. 10, we obtain \(N_{hit} \sim \mathcal {N}(n_0P_2,n_0P_2[1-P_2 ])\), where \(P_2=P_{hit}(r,2T_s)\). Probability that 1 is received when current and previous symbols are 1
where \(\mathcal {Q}(.)\) is the tail probability of the Gaussian probability distribution function. The first and second superscripts in \(P^{11}\) denote current and previous symbol molecules respectively. Similarly
where \(P_1=P_{hit}(r,T_s)\). Thus, the probability of error \(p^1\) when the symbol 1 is sent is
Similarly, the probability of error \(p^0\) when the symbol 0 is sent is
Capacity as a function of normalized diffusion length [31]
Equations 13 and 14 represent the cross-over probabilities for transmit symbols 1 and 0 respectively. Capacity is defined as
with mutual information \(I(X;Y)=I(Y;X)\) and \(I(Y;X)=H(Y)-H(Y|X)\). H(Y) is entropy of the output y and H(Y|X) is conditional entropy of y. In diffusion process, any number of molecules k at distance r at time t (\(t=T_s\) or \(t=2T_s\)) is \(k(r,t)=n_0\cdot erfc\left( \frac{r}{\sqrt{4Dt}}\right) \) and the probability of finding k molecules at distance r at time t (\(t=T_s\) or \(t=2T_s\)), \(P(k)= {n_0 \atopwithdelims ()k} p^k q^{n_0-k}=\frac{n_0!}{k!(n_0-k)!} p^k q^{n_0-k}\), where \(q=(1-p)\) and p stands for the success for a binomial distribution. The probability of first passage time t, as we defined as delay, is less than or equal to \(\tau \), \(p=P(t<\tau )=erfc\left( \frac{r}{\sqrt{4D\tau }}\right) \), where \(\tau \) is maximum delay for a certain diffusion coefficient D and distance r. Apparently, this probability p characterizes the channel. On the basis of this probability p, we can find \(P_1\) and \(P_2\). Figure 8 shows the capacity as a function of normalized diffusion length. The term \(2\sqrt{Dt}\) is called the diffusion length which provides a measure of how far the concentration has propagated by diffusion in time t. The diffusion length is normalized to the distance between the transmitter and receiver. Capacity increases as the normalized diffusion length increases. However, the capacity was found to start decreasing after some normalized diffusion length.
4.2 Bit Error Rate (BER) Performance
4.2.1 Memoryless Channel
Let us consider that \(n_0\) molecules are released for the symbol \(x=1\). The number of received molecules \(n_x(r,T_s)\) is given by [32]
where \(T_s\) is in the time interval from \(\tau \) to \(\tau +T_s\). Letting \(p=p(r,T_s)\) and \(q=1-p\), the probability of error for the symbol \(x=1\) is
For the symbol \(x=0\), no molecule is sent. The receiver will not receive any molecule and there is no error in this case. Therefore,
So, BER for memoryless channel is
4.2.2 Memory Channel
In molecular communication, some molecules stay longer in the medium and impinge on later transmissions [33]. The received molecules are an amalgamation of molecules from current and previous symbols. Let us consider fourth-order memory channel for a test case i.e., the effect of four previous symbols over current symbol is considered. When received, for every transmit symbol \(x \in \big \{0,1\big \}\), one of \(2^5=32\) cases may occur with equal probability; 16 cases for \(x=0\) and 16 cases for \(x=1\) as is shown in Table 2. The number of received molecules \(N_{px}\) for a channel with memory is
where \(n_p\) is the number of left over molecules from previous symbols to the current symbol duration. The first and second subscripts in \(N_{px}\) denote previous and current symbols respectively. We write \(n_p\) as
where \(x_1\) is previous symbol, \(x_2\) is second previous symbol and so on. We notice from Fig. 9 that time slot \((\tau ,\tau +T_s)\) of current symbol falls in the time slot \((\tau +T_s,\tau +2T_s)\) of the previous symbol, \((\tau +2T_s,\tau +3T_s)\) of the second previous symbol, \((\tau +3T_s,\tau +4T_s)\) of the third previous symbol and so on. We have
Receiving history of transmitted molecules [33]
where \(T_1\) is the time interval between \(\tau +T_s\) and \(\tau +2T_s\), \(T_2\) is between \(\tau +2T_s\) and \(\tau +3T_s\) and so on. Let us consider a case when the transmit symbol is \(x=0\). The symbols that contribute to the received molecules are 10110 (case 14 in Table 2). The rightmost digit is the current symbol. Every digit represents the contributor of molecules from that particular symbol. From Eqs. 20 and 21, the number of received molecules is
The probability of error for transmit symbol \(x=0\) is
BER for different channels [33]
Similarly, for the case of transmit symbol \(x=1\) (case 30 in Table 2), the probability of error is
where \(N_{px}=n_x+n_{x_1}+n_{x_2}+n_{x_4}\). If \(p^0\) and \(p^1\) are the probabilities of errors for \(x=0\) and \(x=1\), respectively, then
Therefore, BER for a channel with memory is
BER for fourth-order memory channel [33]
Figure 10 compares BER performances in various channels when the delay is 20 \(\upmu \)s and the distance between the transmitter and receiver is 20 \(\upmu \)m. Other parameters are shown in the Table 3. We observe that the performance degrades as the order of the channel memory increases. BER performances in fourth-order memory channel for different delays are reported in Fig. 11. As shown in the figure, performance degrades as the delay increases.
5 Related Works
BER is the main performance measure of a communication system. Capacity and delay profile etc. characterize the channel. However, very few papers have been found in literature dealing with BER. Symbol error rate (SER) with respect to drift velocity has been studied in Inverse Gaussian (IG) noise channel [13]. They reported that SER decreases as the drift velocity increases. Leeson, et al. studied forward error correction for molecular communications [34]. They calculated BER versus molecules per bit. The authors reported that, upto a certain limit, performance improves as the number of molecules per bit increases. Many authors have worked with channel capacity. A mathematical expression for the capacity in molecular communication nanonetworks has been provided when the propagation of the information relies on the free diffusion of molecules [35]. It was observed that the capacity was of the order of \(10^{36}\) bits/s which is extremely high in comparison to the classical electromagnetic communication. This is due to the fact that, the authors considered thermal entropy and information entropy to be equal which might not be the case in practical scenario. Nakano et al. analyzed capacity of molecular communication with Brownian motion. They modeled the channel as a time slotted binary channel [36]. The authors found that the capacity is largely influenced by the molecules life expectancy. In energy model it was observed that as the energy budget increases, the achievable data rate increases. They also reported that only the previous symbol has significant ISI effect on the current symbol [11]. FRET-based molecular communication model stands as a promising solution to high rate nanoscale communication. The authors also showed the potential of the model for long-range nanonetworks by serially connecting the channels using relay nanomachines. A comparison of information transfer between Brownian motion and molecular motors is reported in [22]. The authors showed that active transport is best when the available number of vesicles is small, and Brownian motion is best when the number of vesicles is large.
6 Conclusion
We introduce molecular communication models and modulation techniques. A free diffusion-based molecular communication channel has also been analyzed where the first passage time of a molecule has been used as delay. Capacity was found to increase with normalized diffusion length. However, after a certain limit, it starts decreasing. Calculation shows that, BER performance degrades as the memory order increases. BER performance was also found to degrade as the delay increases. Enormous research works have been found analyzing the capacity and BER with respect to fluid velocity, number of molecules per bit, transmission distance, diffusion coefficient etc. However, unlike classical electromagnetic communication, a unique quality measure like signal-to-noise-ratio (SNR) is yet to be defined in molecular communication. Coding is yet to be studied exclusively. Transmitter power and transmitter/receiver diversity should also be studied. No mentionable work is found to deal with multiplexing, equalizer, and filtering etc.
References
Freitas RA (1999) Nanomedicine, vol 1: Basic capabilities. Landes Bioscience Georgetown, TX (1999)
Nakano T, Moore MJ, Wei F, Vasilakos AV, Shuai J (2012) Molecular communication and networking: opportunities and challenges. IEEE Trans Nanobiosci 11:135–148
Akyildiz IF, Brunetti F, Blázquez C (2008) Nanonetworks: a new communication paradigm. Comput. Netw. 52:2260–2279
Sawai H (2011) Biological functions for information and communication technologies: theory and inspiration. Springer
Moore MJ, Suda T, Oiwa K (2009) Molecular communication: modeling noise effects on information rate. IEEE Trans Nanobiosci 8:169–180
Kuran M, Yilmaz HB, Tugcu T, Akyildiz IF (2011) Modulation techniques for communication via diffusion in nanonetworks. In: 2011 IEEE International Conference on Communications (ICC), pp 1–5
Kim NR, Chae CB (2013) Novel modulation techniques using isomers as messenger molecules for nano communication networks via diffusion. IEEE J Sel Areas Commun 31:847–856
Kadloor S, Adve R (2009) A framework to study the molecular communication system. In: Proceedings of 18th International Conference on Computer Communication Networks, pp 1–6
ShahMohammadian H, Messier GG, Magierowski S (2012) Optimum receiver for molecule shift keying modulation in diffusion-based molecular communication channels. Nano Commun Netw 3:183–195
Tyrrell H, Harris K (1984) Diffusion in liquids: a theoretical and experimental study. Butterworth-Heinemann
Kuran M, Yilmaz HB, Tugcu T, Zerman B (2010) Energy model for communication via diffusion in nanonetworks. Nano Commun Netw 1:86–95
Redner S (2001) A guide to first-passage processes. Cambridge University Press
Srinivas KV, Eckford AW, Adve RS (2012) Molecular communication in fluid media: the additive inverse Gaussian noise channel. IEEE Trans Inf Theory 58:4678–4692
Khormuji MN (2011) On the capacity of molecular communication over the AIGN channel. In: 2011 45th annual Conference on Information Sciences and Systems (CISS), pp 1–4
Miorandi D (2011) A stochastic model for molecular communications. Nano Commun Netw 2:205–212
Einolghozati A, Sardari M, Beirami A, Fekri F (2011) Capacity of discrete molecular diffusion channels. In: 2011 IEEE international symposium on Information Theory Proceedings (ISIT), pp 723–727
Berg HC (1993) Random walks in biology. Princeton University Press
De Kievit TR, Iglewski BH (2000) Bacterial quorumsensing in pathogenic relationships. Infecti Immun 68:4839–4849
Eckford AW (2007) Nanoscale communication with brownian motion. In: Proceedings of 41st annual conference on information sciences and systems, pp 160–165
Eckford AW (2007) Achievable information rates for molecular communication with distinct molecules. In: Proceedings of workshop computer communications from biological systems: theory and applications, pp 313–315
Okaie Y, Nakano T (2012) Nanomachine placement strategies for detecting Brownian molecules in nanonetworks. In: Proceedings of IEEE Wireless Communication Networking Conference (WCNC), pp 1755–1759
Eckford AW, Farsad N, Hiyama S, Moritani Y (2010) Microchannel molecular communication with nanoscale carriers: Brownian motion versus active transport. In: Proceedings of IEEE international conference on nanotechnologies, pp 854–858
Eckford AW (2009) Timing information rates for active transport molecular communication. In: Nano-Networks, pp 24–28
Alberts B, Bray D, Lewis J, Raff M, Roberts K, Watson JD (1994) Molecular biology of the cell (1994) Garland. New York, pp 139–194
Nakano T, Suda T, Koujin T, Haraguchi T, Hiraoka Y (2007) Molecular communication through gap junction channels: system design, experiments and modeling. In: Proceedings of International Conference on Bio-Inspired Models of Network, Information and Computing Systems, BIONETICS, pp 139–146 (2007)
Hiyama H, Moritani Y, Suda T (2009) A biochemically engineered molecular communication system. Nano Networks, pp 85–94
Oiwa K, Sakakibara H (2005) Recent progress in dynein structure and mechanism. Current Opin Cell Biol 17:98–103
Shima T, Kon T, Imamula K, Ohkura R, Sutoh K (2006) Two modes of microtubule sliding driven by cytoplasmic dynein. Proc Natl Acad Sci 103:17736–17740
Toba S, Oiwa K (2006) Swing or embrace. New aspects of motility inspired by dynein structure in situ. Bioforum Eur 10:14–16
Kuscu M, Akan OB (2012) A physical channel model and analysis for nanoscale molecular communication with Forster Resonance Energy Transfer (FRET). IEEE Trans Nanotechnol 11:200–207
Kabir MH, Kwak KS (2013) Molecular nanonetwork channel model. Electron Lett 49:1285–1287
Socolofsky SA, Jirka GH (2005) CVEN 489-501: Special Topics in Mixing and Transport Processes in the Environment. In: EngineeringLectures. 5th edn., vol 3136. Coastal and Ocean Engineering Division, Texas A&M University, MS, p 77843
Kabir MH, Kwak KS (2014) Effect of memory on BER in molecular communication. Electron Lett 50:71–72
Leeson MS, Higgins MD (2012) Forward error correction for molecular communications. Nano Commun Netw 3:161–167
Pierobon M, Akyildiz IF (2011) Information capacity of diffusion-based molecular communication in nanonetworks. In: INFOCOM, 2011 Proc IEEE, pp 506–510
Nakano T, Okaie Y, Jian-Qin L (2012) Channel model and capacity analysis of molecular communication with Brownian motion. IEEE Commun Lett 16:797–800
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korean Government (MEST) (No. 2010-0018116).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Kabir, H., Kwak, K.S. (2017). Physical Channel Model for Molecular Communications. In: Suzuki, J., Nakano, T., Moore, M. (eds) Modeling, Methodologies and Tools for Molecular and Nano-scale Communications. Modeling and Optimization in Science and Technologies, vol 9. Springer, Cham. https://doi.org/10.1007/978-3-319-50688-3_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-50688-3_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-50686-9
Online ISBN: 978-3-319-50688-3
eBook Packages: EngineeringEngineering (R0)